Abstract
Antibiotics play a key role in the management of infectious diseases in humans, animals, livestock, and aquacultures all over the world. The release of increasing amount of antibiotics into waters and soils creates a potential threat to all microorganisms in these environments. This review addresses issues related to the fate and degradation of antibiotics in soils and the impact of antibiotics on the structural, genetic and functional diversity of microbial communities. Due to the emergence of bacterial resistance to antibiotics, which is considered a worldwide public health problem, the abundance and diversity of antibiotic resistance genes (ARGs) in soils are also discussed. When antibiotic residues enter the soil, the main processes determining their persistence are sorption to organic particles and degradation/transformation. The wide range of DT50 values for antibiotic residues in soils shows that the processes governing persistence depend on a number of different factors, e.g., physico-chemical properties of the residue, characteristics of the soil, and climatic factors (temperature, rainfall, and humidity). The results presented in this review show that antibiotics affect soil microorganisms by changing their enzyme activity and ability to metabolize different carbon sources, as well as by altering the overall microbial biomass and the relative abundance of different groups (i.e., Gram-negative bacteria, Gram-positive bacteria, and fungi) in microbial communities. Studies using methods based on analyses of nucleic acids prove that antibiotics alter the biodiversity of microbial communities and the presence of many types of ARGs in soil are affected by agricultural and human activities. It is worth emphasizing that studies on ARGs in soil have resulted in the discovery of new genes and enzymes responsible for bacterial resistance to antibiotics. However, many ambiguous results indicate that precise estimation of the impact of antibiotics on the activity and diversity of soil microbial communities is a great challenge.
Introduction
Antibiotics are complex molecules with different functional groups in their chemical structures and are divided into several classes (Figure 1, Table 1) depending on the mechanisms of action, i.e., inhibition of cell wall synthesis, alteration of cell membranes, inhibition of protein synthesis, inhibition of nucleic acids synthesis, competitive antagonism, and antimetabolite activity (Kümmerer, 2009). Antibiotics are widely prescribed for treatment of infectious diseases in humans and animals. Moreover, they are used in livestock to increase meat production by preventing infections or outbreaks of diseases and promoting growth at a global scale. The production of antibiotics is still increasing, and the total annual usage has reached from 100,000 to 200,000 tons worldwide (Gelband et al., 2015). Between 2000 and 2015 antibiotic consumption in 76 countries around the world, expressed in defined daily doses (DDDs), increased 65% and, in 2015, reached 42 billion DDDs. Among high-income countries, the leading consumers of antibiotics in 2015 were the United States, France, and Italy. Leading consumers of antibiotics between low and middle-income countries were India, China, and Pakistan (Klein et al., 2018). It has been predicted that in 2030 global antibiotics consumption will be 200% higher than in 2015, with the greatest increase coming from low and middle-income countries. There are significant differences in trends in the antibiotic consumption in European countries. According to the Antimicrobial Consumption—Annual Epidemiological Report for 2016 published by the European Center for Disease Prevention and Control (ECDC), a statistically significant trend of increasing antibiotics usage was observed for Greece and Spain from 2012 to 2016, while over the same time period a statistically significant decreasing antibiotics usage trends were observed for Finland, Luxembourg, Norway and Sweden (ECDC, 2018). The most prescribed antibiotic classes in the US and EU are penicillins, macrolides, cephalosporins, and fluoroquinolones (CDC Centers for Disease, 2015; ECDC, 2018). More detailed information about consumption of different antibiotics in some countries of the EU and US was presented by Singer et al. (2016).
Table 1
| Class | Antibiotic | Chemical formula | Molecular weight (g/mol) | Water solubility (mg/L) | Log KOW | K d (L/kg) | K OC (L/kg) |
|---|---|---|---|---|---|---|---|
| Aminoglycosides | Gentamicin | C21H43N5O7 | 477.6 | 100,000 | −3.1 | – | – |
| Streptomycin | C21H39N7O12 | 581.6 | 12,800 | −6.4 | 8–290 | 580–11,000 | |
| Diaminopyrimidines | Trimethoprim | C14H18N4O3 | 290.3 | 400 | 0.91 | 7.40 | 4,600 |
| Fluoroquinolones | Ciprofloxacin | C17H18FN3O3 | 331.3 | 30,000 | 0.28 | 427–4,844 | 1,127–61,000 |
| Difloxacin | C21H19F2N3O3 | 399.4 | 1,330 | 0.89 | – | – | |
| Enrofloxacin | C19H22FN3O3 | 359.4 | >53.9 | 0.7 | 0.54–5,612 | 39–768,740 | |
| Norfloxacin | C16H18FN3O3 | 319.3 | 177,900 | −1.03 | 591–5,791 | 310 | |
| Ofloxacin | C18H20FN3O4 | 361.4 | 10,800 | 0.35 | 1,471–4,325 | 44,140 | |
| Glycopeptides | Vancomycin | C66H75Cl2N9O24 | 1449.3 | >1,000 | −3.1 | 0.3–0.7 | – |
| Ionophores | Lasalocid | C34H54O8 | 590.8 | 750 | – | 9–280 | 2.9–4.2 |
| Monensin | C36H62O11 | 670.9 | 0.003 | 5.43 | 0.5–65 | 2.1–3.8 | |
| β-Lactams | Amoxicillin | C16H19N3O5S | 365.4 | 3,430 | 0.87 | – | 865.5 |
| Cephapirin | C17H17N3O6S2 | 423.5 | 1,030 | −1.15 | 0.21–3.83 | – | |
| Cefuroxime | C16H16N4O8S | 424.4 | 145 | −0.16 | – | 12.4–15.5 | |
| Penicillin G | C16H18N2O4S | 334.4 | 210 | 1.83 | – | 2.68 | |
| Lincosamides | Clindamycin | C18H33ClN2O5S | 424.9 | 30.6 | 2.16 | – | 70 |
| Lincomycin | C18H34N2O6S | 406.5 | 927 | 0.2 | – | 59 | |
| Macrolides | Azithromycin | C38H72N2O12 | 748.9 | 2.37 | 4.02 | 2.18 | 59,900 |
| Clarithromycin | C38H69NO13 | 747.9 | 1.7 | 3.16 | 262–400 | 150 | |
| Erythromycin | C37H67NO13 | 733.9 | 2,000 | 3.06 | 130 | 10 | |
| Tylosin | C46H77NO17 | 916.1 | 5,000 | 1.63 | 5.4–172,480 | 110–95,532 | |
| Sulfonamides | Sulfachloropyridazine | C10H9ClN4O2 | 284.7 | 8,200 | 0.31 | 0.90–3.5 | 41–170 |
| Sulfadiazine | C10H10N4O2S | 250.3 | 77 | −0.09 | 1.40–14 | 37–125 | |
| Sulfadimethoxine | C12H14N4O4S | 310.3 | 343 | 1.63 | 0.7–4.60 | 89–323 | |
| Sulfadoxine | C12H14N4O4S | 310.3 | 2700 | 0.7 | 0.6–4.9 | 1.8–31.3 | |
| Sulfamethoxazole | C10H11N3O3S | 253.3 | 610 | 0.89 | 0.6–4.9 | 1.2–94.9 | |
| Sulfamethazine | C12H14N4O2S | 278.3 | 1,500 | 0.89 | 0.23–206 | 60–208 | |
| Sulfamonomethoxine | C11H12N4O3S | 280.3 | 10,000 | 0.70 | 0.6–4.9 | 60–200 | |
| Sulfapyridine | C11H11N3O2S | 249.3 | 268 | 0.35 | 1.60–7.40 | 80–308 | |
| Tetracyclines | Chlortetracycline | C22H23ClN2O8 | 478.6 | 630 | −0.62 | 1,280–2,386 | 794 |
| Doxycycline | C22H24N2O8 | 444.4 | 630 | −0.02 | – | – | |
| Oxytetracycline | C22H24N2O9 | 460.4 | 1,000 | −0.9 | 417–1,026 | 2,872–93,317 | |
| Tetracycline | C22H24N2O8 | 444.4 | 231 | −1.19 | 417–1,026 | 400–93,320 |
Basic description and physico-chemical properties of the selected antibiotics.
Kd, distribution coefficient; KOC, soil organic carbon-water partitioning coefficient; KOW, octano-water partition coefficient.
Data obtained from McFarland et al. (1997); Nowara et al. (1997); Rabølle and Spliid (2000); Thiele (2000); Kümmerer (2001); Tolls (2001); Boxall et al. (2002, 2006); Hamscher et al. (2002, 2005); Thiele-Bruhn (2003); Jacobsen et al. (2004); Kay et al. (2004, 2005); Thiele-Bruhn et al. (2004); Halling-Sørensen et al. (2005); Kumar et al. (2005a, 2012); Schmitt et al. (2005); Thiele-Bruhn and Beck (2005); Sarmah et al. (2006); Martínez-Carballo et al. (2007); Sassman and Lee (2007); Stoob et al. (2007); Aust et al. (2008, 2010); Park and Choi (2008); Sukul et al. (2008); Zhang and Dong (2008); Karci and Balcioglu (2009); Kuchta et al. (2009); Li et al. (2009, 2010a,b, 2011, 2015); Muñoz et al. (2009); Conkle et al. (2010); Hu et al. (2010); Vazquez-Roig et al. (2010); Watanabe et al. (2010); Yang et al. (2010, 2016); Zhao et al. (2010); Fan et al. (2011); Lin and Gan (2011); Rosendahl et al. (2011); Zhou et al. (2011, 2013b); Leal et al. (2012); Pinna et al. (2012); Shi et al. (2012); Bak et al. (2013); Huang et al. (2013); Kang et al. (2013); Wu et al. (2013, 2014); Awad et al. (2014); Chen et al. (2014); Ho et al. (2014); Pan et al. (2014); Rutgersson et al. (2014); Van Doorslaer et al. (2014); Wegst-Uhrich et al. (2014); Gao et al. (2015); Hou et al. (2015); Liu et al. (2015); Wang and Wang (2015); DeVries and Zhang (2016); Pan and Chu (2016, 2017b); Tasho and Cho (2016); Zhang et al. (2016b, 2017a).
Figure 1
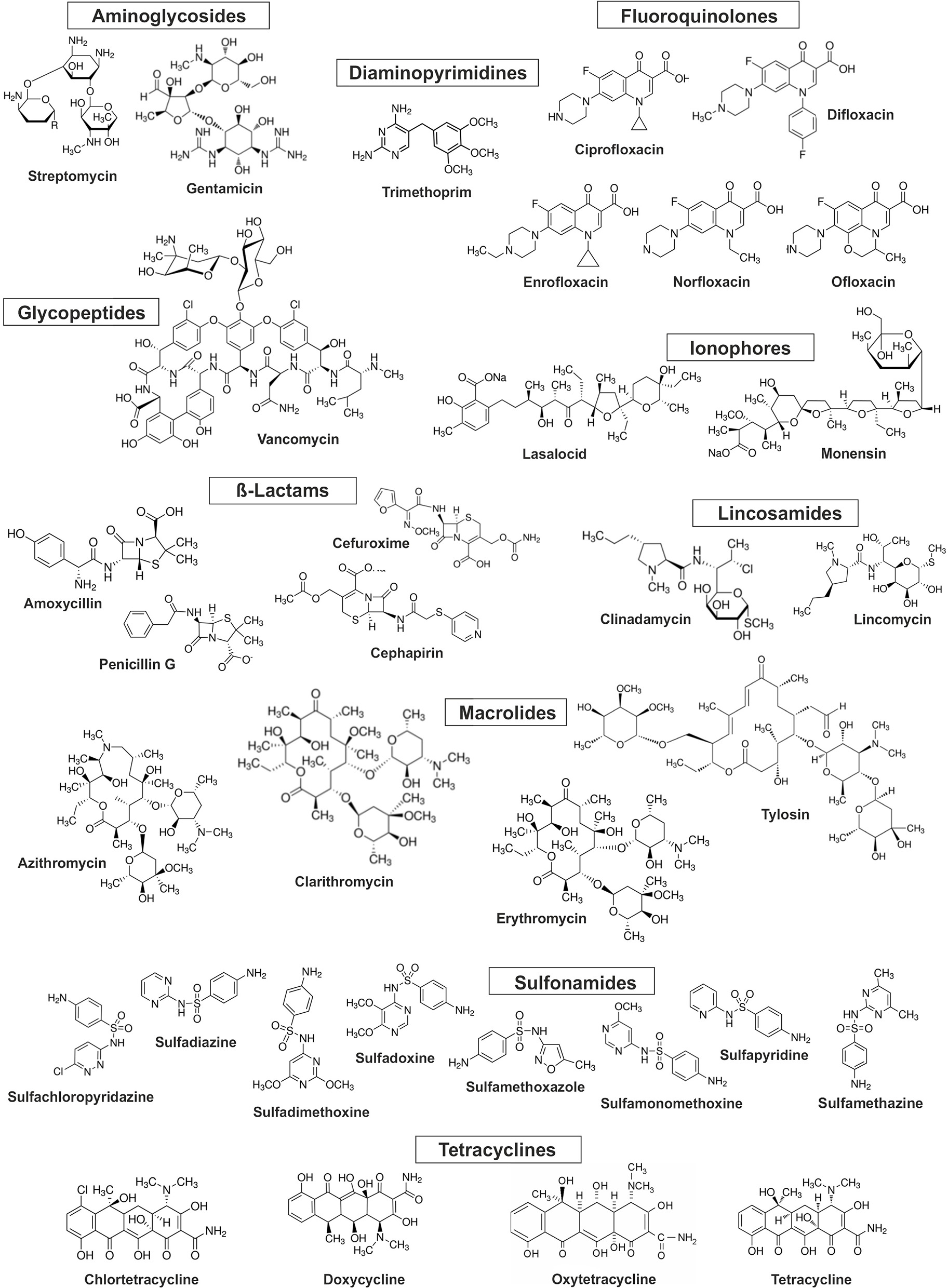
Chemical structure of the selected antibiotics.
Despite their benefits, a continuous release of antibiotics into the environment and their potential adverse impact on living organisms is of great concern (Fatta-Kassinos et al., 2011; De la Torre et al., 2012; Larsson, 2014; Barra Caracciolo et al., 2015; Brandt et al., 2015). Because the majority of antibiotics are not completely metabolized in the bodies of humans and animals, a high percentage of administered drugs is discharged into water and soil through municipal wastewater, animal manure, sewage sludge, and biosolids (nutrient-rich organic materials resulting from the treatment of sewage) that are frequently used to irrigate and fertilize agricultural lands (Bouki et al., 2013; Daghrir and Drogui, 2013; Wu et al., 2014) (Figure 2). It has been reported that 75–80, 50–90, and 60% of the doses of tetracyclines, erythromycin, and lincomycin, respectively, are excreted in urine and feces (Kumar et al., 2005a; Sarmah et al., 2006). Reported antibiotic concentrations in wastewater vary significantly and range from nanograms to micrograms per mL (Gulkowska et al., 2008; Michael et al., 2013; Kulkarni et al., 2017). Though some wastewater treatment processes can degrade antibiotics, there is notable variability in antibiotic removal rates. This can be attributed to differences in treatment processes, such as nature of influent, treatment plant capacity, and the type of technology used (Forsberg et al., 2012; Wu et al., 2014).
Figure 2
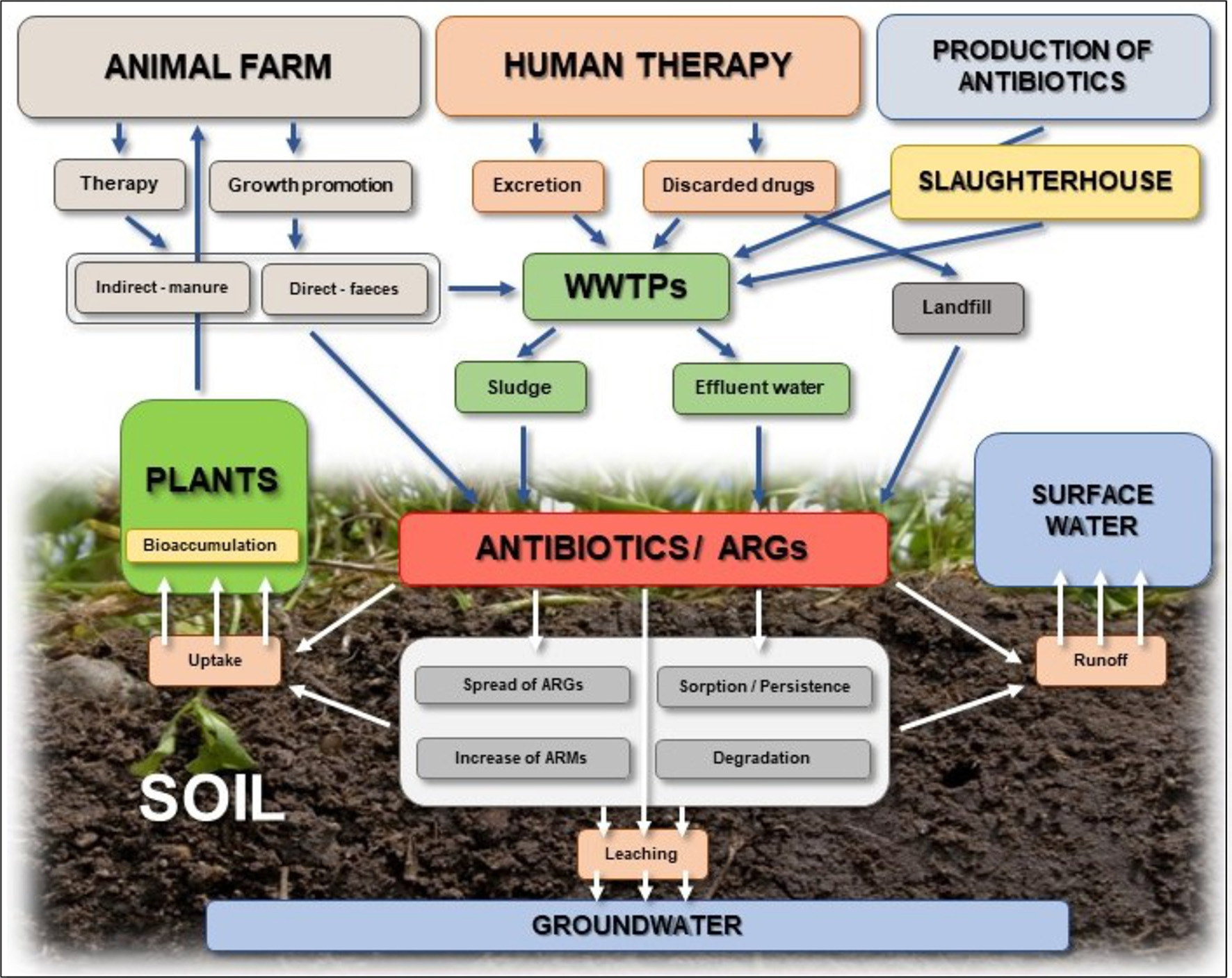
Sources and fate of antibiotics in the soil environment.
The concentrations of antibiotic residues in manure, sewage sludge, biosolids, and soil show large variations (Table 2) and depend on the type of drug, metabolism of the drug in animals, the duration of treatment and the time of sampling relative to the treatment period. Tetracyclines have the highest concentrations and are most frequently reported antibiotic residues in manure (Pan et al., 2011; Chen et al., 2012; Massé et al., 2014). Other groups of antibiotics with considerable concentrations in manure are fluoroquinolones (Zhao et al., 2010; Van Doorslaer et al., 2014) and sulfonamides (Martínez-Carballo et al., 2007). Among the macrolide antibiotics, the highest concentration in manure was measured for tylosin (Dolliver et al., 2008). Compared to manure, biosolids contain much lower amounts of antibiotics (Jones-Lepp and Stevens, 2007) (Table 2).
Table 2
| Class | Antibiotic | Concentration | References |
|---|---|---|---|
| MANURE, μg/kg | |||
| Fluoroquinolones | Ciprofloxacin | 45,000 | Zhao et al., 2010 |
| Enrofloxacin | 1,420 | ||
| Fleroxacin | 99,000 | ||
| Norfloxacin | 225,000 | ||
| Macrolides | Tylosin | 7,000–8,100 | Dolliver et al., 2008; Berendsen et al., 2015 |
| Sulfonamides | Sulfadiazine | 91,000 | Martínez-Carballo et al., 2007 |
| Sulfadimidine | 20,000 | ||
| Tetracyclines | Chlortetracycline | 764,000 | Massé et al., 2014 |
| Oxytetracycline | 354,000 | Chen et al., 2012 | |
| Tetracycline | 98,000 | Pan et al., 2011 | |
| SEWAGE SLUDGE, μg/kg dw | |||
| Diaminopyrimidines | Trimethoprim | 133 | Göbel et al., 2005 |
| Fluoroquinolones | Ciprofloxacin | 426 (8,905) | Lillenberg et al., 2010; Li et al., 2013 |
| Macrolides | Azithromycin | 1.3–158 | Göbel et al., 2005; Li et al., 2013 |
| Sulfonamides | Sulfadimethoxine | 0–20 (22.7) | Lillenberg et al., 2010; Li et al., 2013 |
| Tetracyclines | – | 8,326 | Cheng et al., 2014 |
| BIOSOLIDS, μg/kg dw | |||
| Lincosamides | Lincomycin | 2.6 | Ding et al., 2011 |
| Macrolides | Azithromycin | 14 (6,500) | Jones-Lepp and Stevens, 2007 |
| Erythromycin | 41 (6,500) | Kinney et al., 2006 | |
| Sulfonamides | 650 | US EPA, 2009 | |
| Tetracyclines | Oxytetracycline | 743.6 (8,700) | US EPA, 2009; Ding et al., 2011 |
| SOIL, μg/kg | |||
| Fluoroquinolones | Ciprofloxacin | 5,600 | Thiele-Bruhn, 2003; Martínez-Carballo et al., 2007; Karci and Balcioglu, 2009; Hu et al., 2010; Van Doorslaer et al., 2014; Pan and Chu, 2017b |
| Difloxacin | 21.5 | ||
| Enrofloxacin | 1,347.6 | ||
| Norfloxacin | 2,160 | ||
| Ofloxacin | 898 | ||
| Ionophores | Monensin | 0.0004 | |
| Lincosamides | Lincomycin | 0.117 | |
| Macrolides | Enrofloxacin | 22.93 | Thiele-Bruhn, 2003; Leal et al., 2012; Tasho and Cho, 2016; Pan and Chu, 2017b |
| Erythromycin | 7.2 | ||
| Tylosin | 1,250 | ||
| Sulfonamides | Sulfachloropyridazine | 52.9 | Thiele-Bruhn, 2003; Dolliver et al., 2007; Karci and Balcioglu, 2009; Hu et al., 2010; Carter et al., 2014; Pan and Chu, 2017b |
| Sulfadiazine | 85.5 | ||
| Sulfadimethoxine | 40.4 | ||
| Sulfadoxine | 9.1 | ||
| Sulfamethoxazole | 54.5 | ||
| Sulfamethazine | 200–25,000 | ||
| Sulfamonomethoxine | 5.37 | ||
| Sulfapyridine | 5.11 | ||
| Tetracyclines | Chlortetracycline Doxycycline Oxytetracycline Tetracycline |
12,900 728 50,000 2,683 |
Hamscher et al., 2002; Thiele-Bruhn, 2003; Karci and Balcioglu, 2009; Hu et al., 2010; Liu et al., 2016; Tasho and Cho, 2016; Pan and Chu, 2017b; Łukaszewicz et al., 2018 |
Maximum reported concentrations of selected antibiotics detected in manure, sewage sludge, biosolids, and soil.
dw, dry weight.
Scientists have long been aware of potential problems from the presence of antibiotics in soil. Determined antibiotic concentrations in soil matrices have ranged from a few nanograms to milligrams per kg of soil (Table 2). The highest concentrations are usually found in areas treated with manure or used for livestock (Kay et al., 2004; Zhou et al., 2013a; Hou et al., 2015; DeVries and Zhang, 2016). The concentrations of oxytetracycline and chlortetracycline in some agricultural lands may reach extremely high levels, whereas the concentrations of ciprofloxacin, norfloxacin, and tetracycline are typically significantly lower. Accurate quantification of antibiotics and their transformation products in the soil is of the utmost importance and requires advanced analytical methods, such as high-performance liquid chromatography with tandem mass spectrometry (HPLC/MS) (Aga et al., 2016).
Elevated concentrations of antibiotics in the soil selects for preferential outgrowth of antibiotic-resistant bacteria, which results in changes to antibiotics sensitivity of entire microbial populations (Halling-Sørensen et al., 2005; Binh et al., 2007; Ghosh and LaPara, 2007; Kyselková et al., 2013; Ma et al., 2014; Wepking et al., 2017; Atashgahi et al., 2018) (Figure 3). Even very low concentrations of antibiotics in the soil [below the minimum inhibitory concentration (MIC)], creates conditions for genetic changes in bacterial genomes and transfer of antibiotic resistance genes (ARGs) and associated mobile genetic elements (MGEs), such as plasmids, transposons, and genomic islands, between and among microbial populations (Ghosh et al., 2009; Heuer et al., 2011a,b; Du and Liu, 2012; Keen and Patrick, 2013; Tang et al., 2015; Grenni et al., 2018). In addition, co-selection and expression of resistance genes on MGEs may promote the spread of ARGs, even between distantly related bacterial species (Wellington et al., 2013). Autochthonous bacteria in soil may also represent a reservoir of resistance genes in the environment that can be transferred to the bacteria that colonize the human body (Zhang et al., 2015; Zhou et al., 2017). Such genes, e.g., tetracycline-resistance genes, were found in three different soils collected from Yunnan, Sichuan, and Tibet in China (Wang et al., 2017).
Figure 3
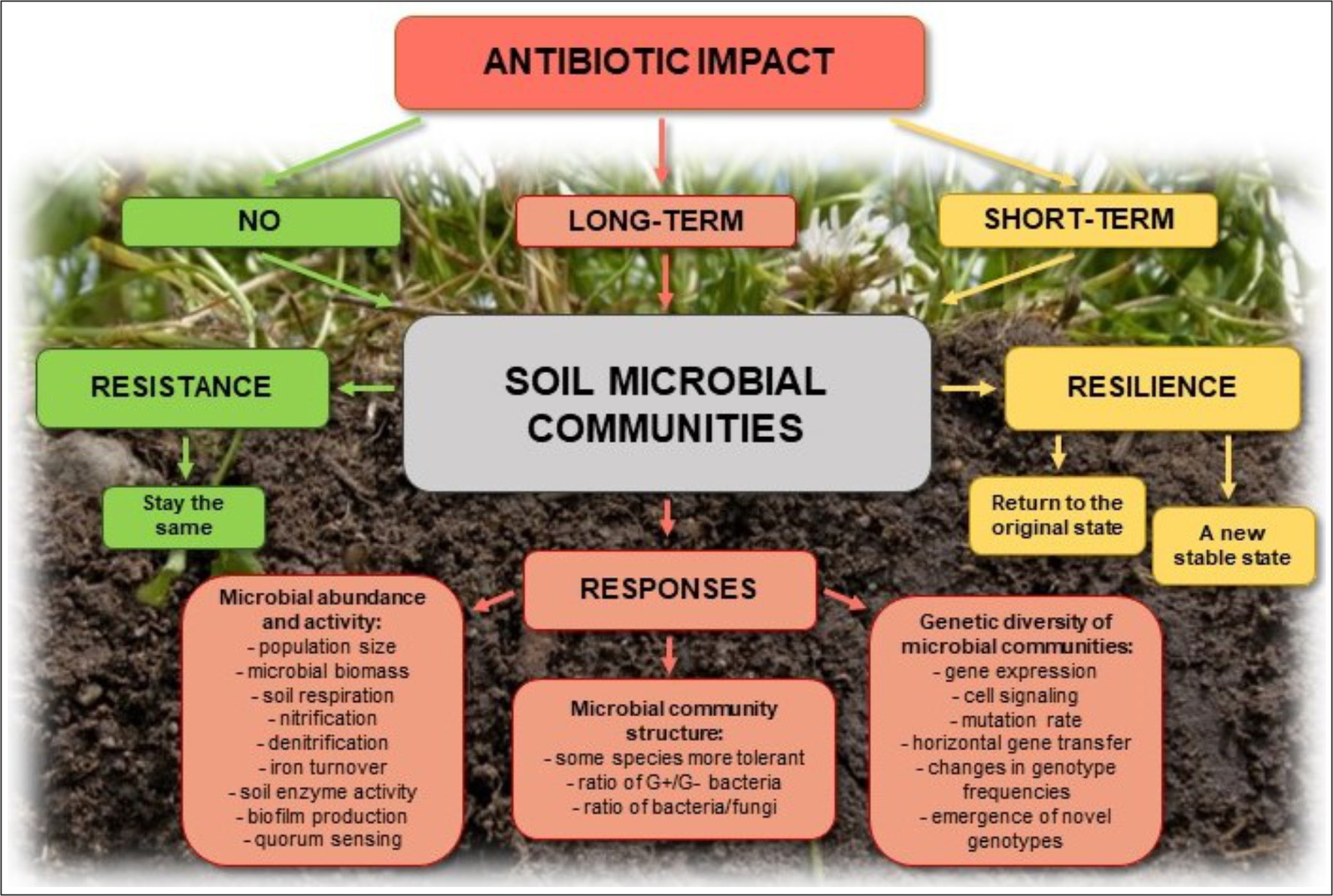
Potential effects of antibiotics on soil microbial communities and their possible responses.
Apart from the selection of antibiotic-resistant microorganisms and the spread of resistance genes in the soil environment, antibiotics may also affect the abundance of soil microorganisms (Pinna et al., 2012; Akimenko et al., 2015; Xu et al., 2016), overall microbial activity (Schmitt et al., 2005; Brandt et al., 2009; Cui et al., 2014; Liu et al., 2015), enzyme activity (Liu et al., 2009; Wei et al., 2009; Chen et al., 2013; Ma et al., 2016), and carbon mineralization and nitrogen cycling (Thiele-Bruhn, 2005; Rosendahl et al., 2012). The impacts of antibiotics on the functional, structural and genetic diversity of soil microorganisms have also been reported (Zielezny et al., 2006; Hammesfahr et al., 2008; Unger et al., 2013; Reichel et al., 2015; Cycoń et al., 2016b; Xu et al., 2016).
The aim of this review is to evaluate recent literature on (1) the degradation of antibiotics in soil and (2) their impact on the microbial community function, (3) the structural and genetic diversity as well as (4) the abundance and diversity of ARGs in soil.
Fate of Antibiotics in Soil
Degradation Rate of Antibiotics in Soil
In the soil environment, antibiotics may be subject to different abiotic and/or biotic processes, including transformation/degradation (Reichel et al., 2013; Cui et al., 2014; Manzetti and Ghisi, 2014; Duan et al., 2017), sorption-desorption (Tolls, 2001; Lin and Gan, 2011; Kong et al., 2012; Leal et al., 2013; Vaz et al., 2015; Martínez-Hernández et al., 2016), uptake by plants (Kumar et al., 2005b; Dolliver et al., 2007; Grote et al., 2007; Kuchta et al., 2009; Carter et al., 2014), and runoff and transport into groundwater (Carlson and Mabury, 2006; Davis et al., 2006; Kuchta et al., 2009; Park and Huwe, 2016; Pan and Chu, 2017a) (Figure 2). Hydrolysis is generally considered one of the most important pathways for abiotic degradation of antibiotics. β-lactams are especially susceptible to hydrolytic degradation, whereas macrolides and sulfonamides are known to be less susceptible to hydrolysis (Braschi et al., 2013; Mitchell et al., 2015). Photo-degradation, which contributes to degradation of antibiotics (e.g., quinolones and tetracyclines) spread on the soil surface during application of manure and slurry to agricultural land, in another important abiotic degradation process (Thiele-Bruhn and Peters, 2007). Antibiotics may also be degraded via reductive or oxidative transformation; however, data on these processes are still scarce.
As many authors have suggested that environmental transformation or degradation depends on the molecular structure (Figure 1) and physicochemical properties (Table 1) of antibiotics are the most important properties governing the fate of different antibiotics in soils (Thiele-Bruhn, 2003; Sassman and Lee, 2007; Crane et al., 2010; Leal et al., 2013; Awad et al., 2014; Pan and Chu, 2016; Zhang et al., 2017a). Since many abiotic and biotic factors affect the degradation of antibiotics, different groups of these pharmaceuticals differ in their rates of degradation in soils, as is evidenced by the large range of half-lives in soil, between < 1 and 3,466 days (Figure 4). For example, Braschi et al. (2013) found that amoxicillin was easily degradable, with a half-life of 0.43–0.57 days. Similarly, a study by Liu et al. (2014) revealed that chlortetracycline (10 mg/kg soil) was quickly degraded in soil with a DT50 value < 1 day. In contrast, an outdoor mesocosm study showed long-term persistence of azithromycin, ofloxacin, and tetracycline in soils, with half-lives of 408−3466, 866–1733, and 578 days, respectively (Walters et al., 2010). Detailed information about the degradation rates of antibiotics and obtained DT50 values in different soils is presented in Table 3. Notable, even for antibiotics within the same groups or, in some cases for particular antibiotics, DT50 values differ significantly. Observed differences in persistence or similar compounds are probably due to variations in soil composition, doses of antibiotics and conditions used in these studies. However, based on a review of the literature, it can be concluded that fluoroquinolones, macrolides, and tetracyclines are characterized by high DT50 or half-life values (Figure 4).
Table 3
| Antibiotic | Type of soil | Main properties of soil | Experimental conditions | Concentration (mg/kg soil) | Half-life/DT50 or comments | References |
|---|---|---|---|---|---|---|
| Amoxycillin | No data | Sand 37%, silt 32%, clay 31%, pH 8.2, OC 7.7 g/kg | At field capacity conditions | 0.2 | Half-life 0.43 day | Braschi et al., 2013 |
| Sand 40%, silt 44%, clay 16%, pH 5.0, OC 21.8 g/kg | Half-life 0.57 day | |||||
| Azithromycin | Silt loam (Canada) | Sand 18%, silt 67%, clay 15%, pH 7.5, OM 3.4%, CEC 13.2 cmol+/kg | 30°C, 15% moisture content, 90 days | 1 | DT50 12.82 days (soil with a history of AZT application at 10 mg/kg soil) | Topp et al., 2016 |
| Sandy clay loam | Sand 53%, silt 27%, clay 20%, | Outdoor mesocosm study, | 0.025 | Half-life 408–990 days | Walters et al., 2010 | |
| Ciprofloxacin | (USA) | pH 5.6, OC 1.7% | average temp. 14°C, soil moisture 14.6–35.1%, 3 years, biosolids addition in a ratio 1:2 | 0.542 | Half-life 1,153–3,466 days | |
| Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | 100 | 100% was removed after 20 days | Ma et al., 2014 | |
| Ustic Cambisol (China) | Sand 12%, silt 54%, clay 34%, pH 7.9, OC 36.76 g/kg, CEC 13.82 cmol+/kg | 25°C, 60% WHC, 40 days | 1, 5, and 50 | Almost 75, 62, and 40% of antibiotic at concentrations of 1, 5, and 50 mg/kg soil, respectively, were degraded within 40 days | Cui et al., 2014 | |
| Agricultural soil (Germany) | Sand 11%, silt 68%, clay 21%, pH 6.6, WHC 37.5%, OC 2.1% | 20°C, 60% WHC, 93 days, sludge addition at 1.8 g/kg | 20 | 0.9% of the initial concentration was mineralized | Girardi et al., 2011 | |
| Clarithromycin | Silt loam (Canada) | Sand 18%, silt 67%, clay 15%, pH 7.5, OM 3.4%, CEC 13.2 cmol+/kg | 30°C, 15% moisture content, 90 days | 1 | DT50 36.48 days (control soil—with no history of CLA application), 15.85 days (soil with a history of antibiotic application at 0.1 mg/kg soil), and 9.51 days (soil with history of antibiotic application at 10 mg/kg soil) | Topp et al., 2016 |
| Clindamycin | Different soil types (Czech Republic) | Sand 15.85–100%, silt 0–76.74%, clay 0–14.7%, pH 5.30–8.71, OC 0.08–2.58% | 20°C, 61 days | 2 | 44–98% of the initial concentration was degraded | Koba et al., 2017 |
| Sandy loam (USA) | pH 6.1 | 30, 20, and 4°C, 30 days, manure addition | 5.6 | 56, 12, and 0% were degraded within 30 days at 30, 20, and 4°C | Gavalchin and Katz, 1994 | |
| Chlortetracycline | Agricultural soil (China) | pH 6.92, OC 6.8 g/kg, CEC 35.2 cmol+/kg | 25°C, 60% WHC, 49 days | 150 | DT50 27.6 and 26.6–26.7 days in non-amended and manure-amended soils, respectively | Li et al., 2010a |
| pH 4.55, OC 16.4 g/kg, CEC 60.0 cmol+/kg | DT50 30.0 and 25.9–30.8 days in non-amended and manure-amended soils, respectively | |||||
| Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | 1–300 | DT50 >20 days | Liu et al., 2009 | |
| No data (China) | Sand 42.95%, silt 43.43%, clay 13.65%, pH 7.6, OC 20.7 g/kg | 25°C, 50% WHC, 45 days, DOM addition at 40 mg C/kg | 10 and 100 | DT50 < 1 and 4.7 days for 10 and 100 mg/kg, respectively | Liu et al., 2014 | |
| Clay loam (China) | Sand 30.4%, silt 34.1%, clay 35.5%, pH 5.7, OC 18.2 g/kg, CEC 9.87 cmol+/kg | 25°C, 60% WHC, 21 days | 10 | <30% dissipated in the first 7 days and < 55% after 21 days | Liu et al., 2012a | |
| Silt loam (gray brown Luvisol) | Sand 18%, silt 67%, clay 15%, pH 7.5, OM 3.4%, CEC 13.2 cmol+/kg | 30°C, 15% WHC, 7 days | 10 | DT50 3.3 days (in soil with history of exposure) and 2.8 days (in soil with no history exposure) | Topp et al., 2013 | |
| Farm field soil (Canada) | No data | Room temp., soil moisture 20%, 47 days | 1 h | Half-life 20 days in a laboratory study; half-life 21 days (24 days with manure addition) in a field study | Carlson and Mabury, 2006 | |
| No data (China) | Sand 42.95%, silt 43.43%, clay 13.65%, pH 7.6, OC 20.7 g/kg | 25°C, 50% WHC, 45 days | 1, 10, and 100 | DT50 < 1, < 1, and 5 days for 1, 10, and 100 mg/kg, respectively | Liu et al., 2015 | |
| Inceptisol (China) | Sand 21.5%, silt 71.1%, clay 7.4%, pH 6.8, OM 3.1%, WHC 27%, CEC 10.6 cmol+/kg | 25°C, 60% WHC, 35 days | 1 and 100 | Half-life 1.58 and 6.07 days for 10 and 100 mg/kg, respectively | Fang et al., 2016 | |
| Loamy sand (Denmark) | Sand 75.4%, silt 10.7%, clay 11.3%, pH 6.1, OC 1.6% | Field experiment, manure addition, 155 days | 0.03 and 0.05 | Half-life 25 days | Halling-Sørensen et al., 2005 | |
| Sand (Denmark) | Sand 87.6%, silt 4.8%, clay 35.2%, pH 4.3, OC 1.4% | Half-life 34 days | ||||
| Difloxacin | Luvisol (Germany) | Sand 6%, silt 78%, clay 16%, pH 6.3, OC 1.2%, CEC 11.4 cmol+/kg | 21°C, 63 days, slurry addition at 40 ml/kg | 0.452 | Residual concentration declined to 0.258 mg/kg soil after 63 days | Reichel et al., 2013 |
| Doxycycline | Silty clay loam (China) | Sand 9%, silt 63.7%, clay 27.3%, pH 7, OM 1% | 25°C, 50–65% WHC, manure addition at 10 g/kg, 56 days | 10 | Almost 92% of the initial concentration was degraded during 49 days | Wang et al., 2016 |
| Sandy clay loam (USA) | Sand 53%, silt 27%, clay 20%, pH 5.6, OC 1.7% | Outdoor mesocosm study, average temp. 14°C, soil moisture 14.6–35.1%, 3 years, biosolids addition in a ratio 1:2 | 0.017 | Half-life 533–578 days | Walters et al., 2010 | |
| Erythromycin | Sandy loam (Germany) | Sand 72%, silt 23%, clay 5%, pH 7.2, WHC 34.4%, OC 1.69% | 20°C, water content 12%, 120 days | 2 | Half-life 20 days; 98% was degraded | Schlüsener et al., 2006 |
| Sandy loam (USA) | pH 6.1 | 30, 20, and 4°C, 30 days, manure addition | 5.6 | 100, 75, and 0% were degraded within 30 days at 30, 20, and 4°C | Gavalchin and Katz, 1994 | |
| Silt loam (Canada) | Sand 18%, silt 67%, clay 15%, pH 7.5, OM 3.4%, CEC 13.2 cmol+/kg | 30°C, 15% moisture content, 90 days | 1 | DT50 65.93 days (control soil—with no history of application), 4.36 days (soil with a history of application at 0.1 mg/kg soil), and 0.94 days (soil with a history of application at 10 mg/kg soil) | Topp et al., 2016 | |
| Clay loam (China) | pH 6.45, OC 0.8%, WHC 50% | 25°C, 70% WHC, 90 days | 0.1 | DT50 6.4 (non-sterile soil) and 40.8 days (sterile soil) under aerobic conditions; DT50 11.0 (non-sterile soil) and 57.8 days (sterile soil) under anaerobic conditions | Pan and Chu, 2016 | |
| 0.05, 0.1, and 0.2 | DT50 3.01–16.9 days (non-sterile soil) under aerobic conditions | |||||
| Enrofloxacin | Sandy loam (UK) | pH 5.4, OC 1.3% | 10 | 30.3% was degraded during 56 days | Martens et al., 1996 | |
| Lasalocid | Silty clay–clay loam (Slovenia) | Sand 22.4%, Silt 49.0%, Clay 28.6%, pH 7.1, OM 4.1%, OC 2.4% | Field experiment, 21 days, manure addition at 10, 20 or 30 t/ha | 3.01 | Half-life 3.1 days (regardless of the treatment and soil depth) | ŽiŽek et al., 2015 |
| Clay loam–silty clay loam (Slovenia) | Sand 19.8%, Silt 49.6%, Clay 30.6%, pH 7.1, OM 4.1%, OC 2.4% | |||||
| Silty clay loam (Slovenia) | Sand 15.8%, Silt 54.4%, Clay 29.8, pH 7.1, OM 4.6%, OC 2.7% | |||||
| Sand (USA) | Clay 11%, pH 7.0, OC 0.87%, CEC 4.4 cmol+/kg | 23°C, 30 days | 2.1 | Half-life 1.5 days | Sassman and Lee, 2007 | |
| Clay loam (USA) | Clay 21%, pH 7.5, OC 2.91%, CEC 26.5 cmol+/kg | Half-life 3.6 days (4.3 days with manure at 20 mg/kg) | ||||
| Monensin | Farm field soil (Canada) | No data | Room temp., soil moisture 20%, 47 days | 1 | Half-life 13.5 days in a laboratory study; half-life 3.8 days (3.3 days with manure addition) | Carlson and Mabury, 2006 |
| Sand (USA) | Clay 11%, pH 7.0, OC 0.87%, CEC 4.4 cmol+/kg | 23°C, 30 days | 2.1 | Half-life 1.3 days | Sassman and Lee, 2007 | |
| Clay loam (USA) | Clay 21%, pH 7.5, OC 2.91%, CEC 26.5 cmol+/kg | Half-life 2 days (1.6 days with manure at 20 mg/kg) | ||||
| Clay loam (China) | pH 6.45, OC 0.8%, WHC 50% | 25°C, 70% WHC, 90 days | 0.1 | DT50 2.91 (non-sterile soil) and 40.8 days (sterile soil) under aerobic conditions; DT50 5.6 (non-sterile soil) and 53.4 days (sterile soil) under anaerobic conditions | Pan and Chu, 2016 | |
| 0.05, 0.1, and 0.2 | DT50 1.8–6.93 days (non-sterile soil) under aerobic conditions | |||||
| Norfloxacin | Clay loam (China) | pH 6.45, OC 0.8%, WHC 50% | 25°C, 70% WHC, 90 days | 0.1 | DT50 2.91 (non-sterile soil) and 40.8 days (sterile soil) under aerobic conditions; DT50 5.6 (non-sterile soil) and 53.4 days (sterile soil) under anaerobic conditions | Pan and Chu, 2016 |
| 0.05, 0.1, and 0.2 | DT50 1.8–6.93 days (non-sterile soil) under aerobic conditions | |||||
| Silty clay loam (China) | Sand 9%, silt 63.7%, clay 27.3%, pH 7, OM 1% | 25°C, 50–65% WHC, manure addition at 10 g/kg, 56 days | 10 | Almost 47% of initial concentration was degraded within 49 days | Wang et al., 2016 | |
| Acidic Soil (China) | Sand 29%, silt 39%, clay 32%, pH 4.3, OM 2.4%, CEC 9.5 cmol+/kg | 25°C, 50% WHC, 42 days | 5, 10, and 30 | Half-life 31, 48, and 62 days (without manure) for 5, 10, and 30 mg/kg, respectively; half-life 24–39 days (with manure 3−9%) for 10 mg/kg | Yang et al., 2012 | |
| Sandy clay loam (USA) | Sand 53%, silt 27%, clay 20%, pH 5.6, OC 1.7% | Outdoor mesocosm study, average temp. 14°C, soil moisture 14.6–35.1%, 3 years biosolids addition in a ratio 1:2 | 0.045 | Half-life 990–1,386 days | Walters et al., 2010 | |
| Ofloxacin | 0.470 | Half-life 866–1,733 days | ||||
| Oxytetracycline | Soil from a wheat field (China) | pH 8.45, OM 19.13 g/kg, CEC 14.84 cmol+/kg, OTC 1.65 mg/kg | 20–25°C day/15°C night, 70% WHC, 60 days | 300 | 85.6 and 87.3% were degraded in soil with compost (10%) and compost (10%) + biochar (2%), respectively | Duan et al., 2017 |
| Agricultural soil (China) | pH 6.92, OC 6.8 g/kg, CEC 35.2 cmol+/kg | 25°C, 60% WHC, 49 days | 150 | DT50 30.2 and 38.2–39.7 days in non-amended and manure-amended soils, respectively | Li et al., 2010a | |
| pH 4.55, OC 16.4 g/kg, CEC 60.0 cmol+/kg | DT50 39.4 and 35.9–41.3 days in non-amended and manure-amended soils, respectively | |||||
| Sandy loam (UK) | Sand 69–80%, silt 6–21%, clay 4–10%, pH 6.2–6.6, OC 1.3% | Field experiment, 127 days | 0.3 | DT50 21.7 days | Blackwell et al., 2007 | |
| Sandy loam (USA) | OM 0.92%, pH 7.2 | 25°C, moisture 20%, 62 days | 50 | Half-life 33 and 56 days in manure-amended and non-amended soils, respectively | Wang and Yates, 2008 | |
| Alfisol (China) | Sand 7.7%, silt 77.5%, clay 14.8%, pH 6.24, OM 2.4%, CEC 12.3 cmol+/kg, OTC 37.3 μg/kg | 25°C, 60% WHC, 120 days | 1–30 | 86.6, 89.6, 93.7, and 95.4% of antibiotic at concentrations of 1, 3.6, 10, and 30 mg/kg soil, respectively, were degraded within 120 days | Ma et al., 2016 | |
| Penicillin G | Sandy loam (USA) | pH 6.1 | manure addition | 5.6 | No degradation within 30 days | Gavalchin and Katz, 1994 |
| Sulfachloropyridazine | Silt Loam (USA) | Sand 19.9%, silt 56.6%, clay 23.6%, pH 7.5, OC 1.8 | 25°C, 40 days | 1, 10, and 100 | Half-life 20, 20, and 22 days (non-sterile soil) for 1, 10, and 100 mg/kg, respectively; half-life 68 days (sterile soil) for 10 mg/kg | Accinelli et al., 2007 |
| Sand (USA) | Sand 93.5%, silt 2.7%, clay 3.8%, pH 7.2, OC 0.94% | Half-life 27, 26, and 28 days (non-sterile soil) for 1, 10, and 100 mg/kg, respectively; half-life 71 days (sterile soil) for 10 mg/kg | ||||
| Sandy loam (UK) | Sand 69–80%, silt 6–21%, clay 4–10%, pH 6.2–6.6, OC 1.3% | Field experiment, 127 days | 0.65 | DT50 3.5 days | Blackwell et al., 2007 | |
| Clay loam (UK) | Sand 42.63%, silt 32.26%, clay 25.11%, pH 6.8, OC 2.2%, | Field experiment | 0.2 | DT50 29 days | Kay et al., 2004 | |
| Sulfadimethoxine | Silt loam (USA) | Sand 8.0%, silt 65.1%, clay 26.9%, pH 5.54, OC 1.44%, moisture 1.8% | 25°C, 70 days, manure addition at 5% | 1, 25, 50, and 100 | DT50 3, 5.8, 6.8, and 11 days for 1, 25, 50, and 100 mg/kg, respectively | Wang et al., 2006 |
| Sulfadiazine | Luvisol (Germany) | Sand 60 g/kg, silt 220 g/kg, clay 30 g/kg, pH 6.3, OC 12.2 g/kg, | 10°C, 30% WHC, 218 days | 2.2 | DT50 19 days (CaCl2 fraction), 24 days (MeOH fraction), and 290 days (residual fraction) | Förster et al., 2009 |
| Cambisol (Germany) | Sand 750 g/kg, silt 780 g/kg, clay 160 g/kg, pH 6.0, OC 9.9 g/kg | 2.7 | DT50 15 days (CaCl2 fraction), 13 days (MeOH fraction), and 490 days (residual fraction) | |||
| Agricultural soil (China) | Sand 36.96%, silt 58.76%, clay 4.28%, pH 7.6, OC 4.71 g/kg, CEC 7.0 cmol+/kg | 21°C, 49 days, WHC 60%, manure addition at 4% | 4, 10, and 20 | DT50 8.48, 8.97, and 10.22 days (non-sterile soil), and 30.09, 26.55, and 21.21 days (sterile soil) for 4, 10, and 20 mg antibiotic/kg, respectively | Zhang et al., 2017a | |
| Loamy sand (Germany) | Sand 73.3%, silt 23.1%, clay 3.6%, pH 5.5, OC 1% | 10°C, 50% WHC, manure addition at 40 mg/g, 61 days | 10 and 100 | DT50 < 1 and 8.5 days for 10 and 100 mg/kg, respectively | Hammesfahr et al., 2008 | |
| Silt loam (Germany) | Sand 6.4%, silt 78.2%, clay 15.4%, pH 7.2, OC 2.1% | DT50 < 1 and 5.6 days for 10 and 100 mg/kg, respectively | ||||
| Endogleyic Cambisol (Germany) | Sand 73.3%, silt 23.1%, clay 3.6%, pH 4.8, OC 1% | 10°C, 50% WHC, manure addition at 20, 40, and 80 g/kg, 32 days | 10 and 100 | At day 1, the recovery rate ranged between 27 and 45%; on day 32 15−18% were extracted in higher treatment and 7−10% in the lower treatment | Hammesfahr et al., 2011 | |
| Luvisol (China) | Sand 58.4%, silt 21.7%, clay 19.9%, pH 6.24, OM 3.56%, CEC 5.38 cmol+/kg | 25°C, 25% WHC, manure addition at 40 mg/kg, Cu addition at 0, 20, and 200 mg/kg, 28 days | 10 and 100 | Degradation rate constant k (day) 0.115, 0.108, 0.087, and 0.041, 0.035, 0.028 at 10 mg/kg + 0, 20, and 200 mg Cu/kg, and antibiotic at 100 mg/kg + 0, 20, and 200 mg Cu/kg, respectively | Xu et al., 2016 | |
| Luvisol (Germany) | Sand 6%, silt 78%, clay 16%, pH 6.3, OC 1.2%, CEC 11.4 cmol+/kg | 21°C, 63 days, slurry addition at 40 ml/kg | 0.256 | < 0.002 mg antibiotic/kg soil was detected within 63 days | Reichel et al., 2013 | |
| Silty loam (Germany) | Sand 4.3%, silt 82.9%, clay 12.8%, pH 6.7, OC 1% | 60 days | 0.71 | DT50 4.8 days | Sittig et al., 2014 | |
| Loamy sand (Germany) | Sand 69.7%, silt 26.3%, clay 4%, pH 5.7, OC 0.9% | 0.68 | DT50 8.6 days | |||
| Sulfamonomethoxine | Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | 100 | 71.8% was removed within 20 days | Ma et al., 2014 |
| Sulfamethoxazole | Sand (USA) | Sand 91%, silt 5%, clay 4%, pH 9.23, OC 0.16%, CEC 8.2 cmol+/kg | 21°C, 75% WHC, 84 days | 0.04 | Half-life 11.4 (non-sterile soil) and 58.7 days (sterile soil) under aerobic conditions; half-life 18.3 (non-sterile soil) under anaerobic conditions | Lin and Gan, 2011 |
| Medium loam (USA | Sand 91%, silt 5%, clay 4%, pH 9.23, OC 0.16%, CEC 8.2 cmol+/kg | Half-life 9.0 and 15.3 days (non-sterile soil) under aerobic and anaerobic conditions, respectively | ||||
| Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | 1–300 | DT50 2–5 days | Liu et al., 2009 | |
| Clay loam (China) | Sand 30.4%, silt 34.1%, clay 35.5%, pH 5.7, OC 18.2 g/kg, CEC 9.87 cmol+/kg | 25°C, 60% WHC, 21 days | 10 | More than 90% dissipated in the first 7 days | Liu et al., 2012a | |
| Silty clay loam (China) | Sand 9%, silt 63.7%, clay 27.3%, pH 7, OM 1% | 25°C, 50–65% WHC, manure addition at 10 g/kg, 56 days | 10 | Almost 80% of initial concentration was degraded during 49 days | Wang et al., 2016 | |
| Silt loam topsoil (TS)/subsoil (SS) (New Zealand) | Sand 9%/12.3%, silt 54%/62.8%, clay 37%/24.9%, pH 6.7/5.7, OC 5%/0. | 25°C, 60% WHC, 36 days | 0.5 | DT50 11 (TS) and 34.7 (SS) days (sterile soil); DT50 9.2 (TS) and 11.8 (SS) days (non-sterile soil) | Srinivasan and Sarmah, 2014 | |
| Clay loam topsoil (TS) and subsoil (SS) (New Zeland) | Sand 13.7%/13.4%, silt 51%/40.3%, clay 30.4%/46.2%, pH 5.8/5.1, OC 4%/0.8% | DT50 13 (TS) and 22.4 (SS) days (sterile soil); DT50 4.3 (TS) and 4.2 (SS) days (non-sterile soil) | ||||
| Silt loam topsoil (TS) and subsoil (SS) (New Zeland) | Sand 34%, silt 48%, clay 17%, pH 5.7/6.6, OC 8.2%/1.7% | DT50 18.1 (TS) and 22.7 (SS) days (sterile soil); DT50 13.3 (TS) and 12.4 (SS) days (non-sterile soil) | ||||
| Loamy sand (Netherlands) | Sand 78.9%, silt 10.4%, clay 7%, pH 4.9, OC 3.7% | 25°C, 35% WHC, 5 weeks | 1–500 | Initial concentrations were reduced to 153, 1.5, and 0.04 mg/kg soil at 500, 20, and 1 mg/kg soil within 5 weeks | Demoling et al., 2009 | |
| Different soil types (Czech Republic) | Sand 15.85–100%, silt 0–76.74%, clay 0–14.7%, pH 5.30–8.71, OC 0.08–2.58% | 20°C, 61 days | 2 | 25–99% of initial concentration was degraded | Koba et al., 2017 | |
| Agricultural soil (China) | Sand 36.96%, silt 58.76%, clay 4.28%, pH 7.6, OC 4.71 g/kg, CEC 7.0 cmol+/kg | 21°C, 49 days, WHC 60%, manure addition at 4% | 4, 10, and 20 | DT50 13.68, 10.28, and 10.81 days (non-sterile soil), and 22.99, 33.24, and 22.79 days (sterile soil) for 4, 10, and 20 mg/kg, respectively | Zhang et al., 2017a | |
| Sulfamethazine | Silt Loam (USA) | Sand 19.9%, silt 56.6%, clay 23.6%, pH 7.5, OC 1.8 | 25°C, 40 days | 1, 10, and 100 | Half-life 17, 18, and 16 days (non-sterile soil) for 1, 10, and 100 mg/kg, respectively; half-life 78 days (sterile soil) for 10 mg/kg | Accinelli et al., 2007 |
| Sand (USA) | Sand 93.5%, silt 2.7%, clay 3.8%, pH 7.2, OC 0.94% | Half-life 22, 23, and 23 days (non-sterile soil) for 1, 10, and 100 mg/kg, respectively; half-life 77 days (sterile soil) for 10 mg/kg | ||||
| Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | 1–300 | DT50 2–5 days | Liu et al., 2009 | |
| Silt loam (Canada) | Sand 18%, silt 67%, clay 15%, pH 7.5, OM 3.4%, CEC 13.2 cmol+/kg | 30°C, 15% WHC, 7 days | 100 ng/g SMZ + 10,000 dpm/g [U-phenyl-14C]-SMZ | DT50 1.3 days (in soil with history of exposure) and 5.3 days (in soil with no history of exposure) | Topp et al., 2013 | |
| Silt loam (Korea) | pH 6.0, OM 2.36% | 25°C, 70% WHC, 56 days, poultry manure addition (1%) | 20 and 100 | The concentration of antibiotic at 100 mg/kg was 255.5 and 129.8 μg/kg while at 20 mg/kg was 62.1 and 31.5 μg/kg at the beginning and on day 56, respectively. | Awad et al., 2016 | |
| Clay loam (China) | pH 6.45, OC 0.8%, WHC 50% | 25°C, 70% WHC, 90 days | 0.1 | DT50 24.8 (non-sterile soil) and 49.5 days (sterile soil) under aerobic conditions; DT50 34.7 (non-sterile soil) and 57.8 days (sterile soil) under anaerobic conditions | Pan and Chu, 2016 | |
| 0.05, 0.1, and 0.2 | DT50 16.90–53.31 days (non-sterile soil) under aerobic conditions | |||||
| Sand (Australia) | Sand 89%, silt 3%, clay 8%, pH 6.25, moisture 0.6%, OC 1%, CEC 5.2 cmol+/kg | 23°C (day) and 15°C (night), 60% WHC, 40 days | DT50 0.99 days | Carter et al., 2014 | ||
| Streptomycin | Sandy loam (USA) | pH 6.1 | Manure addition | 5.6 | No degradation within 30 days | Gavalchin and Katz, 1994 |
| Tetracycline | Agricultural soil (China) | pH 6.92, OC 6.8 g/kg, CEC 35.2 cmol+/kg | 25°C, 60% WHC, 49 days | 150 | DT50 20.9 and 26.7–29.1 days in non-amended and manure-amended soils, respectively | Li et al., 2010a |
| pH 4.55, OC 16.4 g/kg, CEC 60.0 cmol+/kg | DT50 21.7 and 20.6–26.2 days in non-amended and manure-amended soils, respectively | |||||
| Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | 1–300 | DT50 > 20 days | Liu et al., 2009 | |
| Sandy clay loam (USA) | Sand 53%, silt 27%, clay 20%, pH 5.6, OC 1.7% | Outdoor mesocosm study, average temp. 14°C, soil moisture 14.6–35.1%, 3 years, biosolids addition in a ratio 1:2 | 0.021 | Half-life 578 days | Walters et al., 2010 | |
| Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | 100 | 100% was removed within 20 days | Ma et al., 2014 | |
| Clay loam (China) | pH 6.45, OC 0.8%, WHC 50% | 25°C, 70% WHC, 90 days | 0.1 | DT50 31.5 (non-sterile soil) and 57.8 days (sterile soil) under aerobic conditions; DT50 43.3 (non-sterile soil) and 86.6 days (sterile soil) under anaerobic conditions | Pan and Chu, 2016 | |
| 0.05, 0.1, and 0.2 | DT50 14.1–69.3 days (non-sterile soil) under aerobic conditions | |||||
| Trimethoprim | Sand (USA) | Sand 91%, silt 5%, clay 4%, pH 9.23, OC 0.16%, CEC 8.2 cmol+/kg | 21°C, 75% WHC, 84 days | 0.04 | Half-life 26.0 and 26.1 days (non-sterile soil) under aerobic and anaerobic conditions, respectively | Lin and Gan, 2011 |
| Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | 1–300 | DT50 2–5 days | Liu et al., 2009 | |
| Different soil types (Czech Republic) | Sand 15.85–100%, silt 0–76.74%, clay 0–14.7%, pH 5.30–8.71, OC 0.08–2.58% | 20°C, 61 days | 2 | 13–84% of initial concentration was degraded | Koba et al., 2017 | |
| Sandy loam (USA) | Sand 60%, silt 22%, clay 18%, pH 7, WHC 15%, OM 1.6% | 20°C, 30 days | 50 | Half-life 7–8 days | Hu and Coats, 2007 | |
| Sandy loam (Germany) | Sand 72%, silt 23%, clay 5%, pH 7.2, WHC 34.4%, OC 1.69% | 20°C, water content 12%, 120 days | 2 | Half-life 8 days; 100% was degraded | Schlüsener et al., 2006 | |
| Sandy loam (USA) | pH 6.1 | 30, 20, and 4°C, 30 days, manure addition | 5.6 | 100, 100, and 60% were degraded during 30 days at 30, 20, and 4°C | Gavalchin and Katz, 1994 | |
| Sand (Denmark) | Sand 90.7%, silt 2.8%, clay 4.1%, pH 6.8, WHC 15%, OC 1.2% | 25°C, 15% WHC, 33 days | 2,000 | 100% of TYL was degraded within13 days | Westergaard et al., 2001 | |
| Tylosin | Silt loam (Canada) | Sand 18%, silt 67%, clay 15%, pH 7.5, OM 3.4%, CEC 13.2 cmol+/kg | 30°C, 15% WHC, 7 days | 10 | DT50 2 days (in soil with a history of exposure) and 10.2 days (in soil with no history exposure) | Topp et al., 2013 |
| Farm field soil (Canada) | No data | Room temp., soil moisture 20%, 47 days | 1 | Half-life 4.4 days in a laboratory study; half-life 6.1 days (4.5 days with manure addition) in a field study | Carlson and Mabury, 2006 | |
| Loamy sand (Denmark) | Sand 75.4%, silt 10.7%, clay 11.3%, pH 6.1, OC 1.6% | Field experiment, manure addition, 155 days | 0.03 and 0.05 | Half-life 67 days | Halling-Sørensen et al., 2005 | |
| Sand (Denmark) | Sand 87.6%, silt 4.8%, clay 35.2%, pH 4.3, OC 1.4% | Half-life 49 days | ||||
| Silt loam (China) | OC 18.2 g/kg, pH 5.7 | 25°C, 60% WHC, 22 days | 1–300 | DT50 8 days | Liu et al., 2009 | |
| Sand (Denmark) | Sand 90.7%, silt 2.8%, clay 4.1%, pH 6.8, WHC 15%, OC 1.2% | 25°C, 60 days | 2,000 | Completely dissipated within 13 days; all degradation products disappeared after 17 days | Müller et al., 2002 | |
| Sand (Denmark) | Sand 87.6%, silt 4.8%, clay 5.2%, pH 6.3, OC 1.4% | Manure addition | 100 | 50% was degraded within 4.2 days | Ingerslev and Halling-Sørensen, 2001 | |
| Sandy loam (Denmark) | Sand 75.4%, silt 10.8%, clay 11.3%, pH 6.8, OC 1.6% | 50% was degraded within 5.7 days | Ingerslev and Halling-Sørensen, 2001 | |||
| Vancomycin | Sandy loam (Poland) | Sand 67%, silt 24%, clay 9%, pH 6.9, WHC 43%, OC 1.6%, CEC 10 cmol/kg | 22°C, 50% WHC, 90 days | 1 and 10 | DT50 16 days; dissipation was independent of the concentration used. | Cycoń et al., 2018 |
Detailed data on the degradation of the selected antibiotics in soils with different characteristics.
CEC, cation exchange capacity; DOM, dissolved organic matter; OC, organic carbon; OM, organic matter; URE, urease; WHC, water holding capacity.
Figure 4
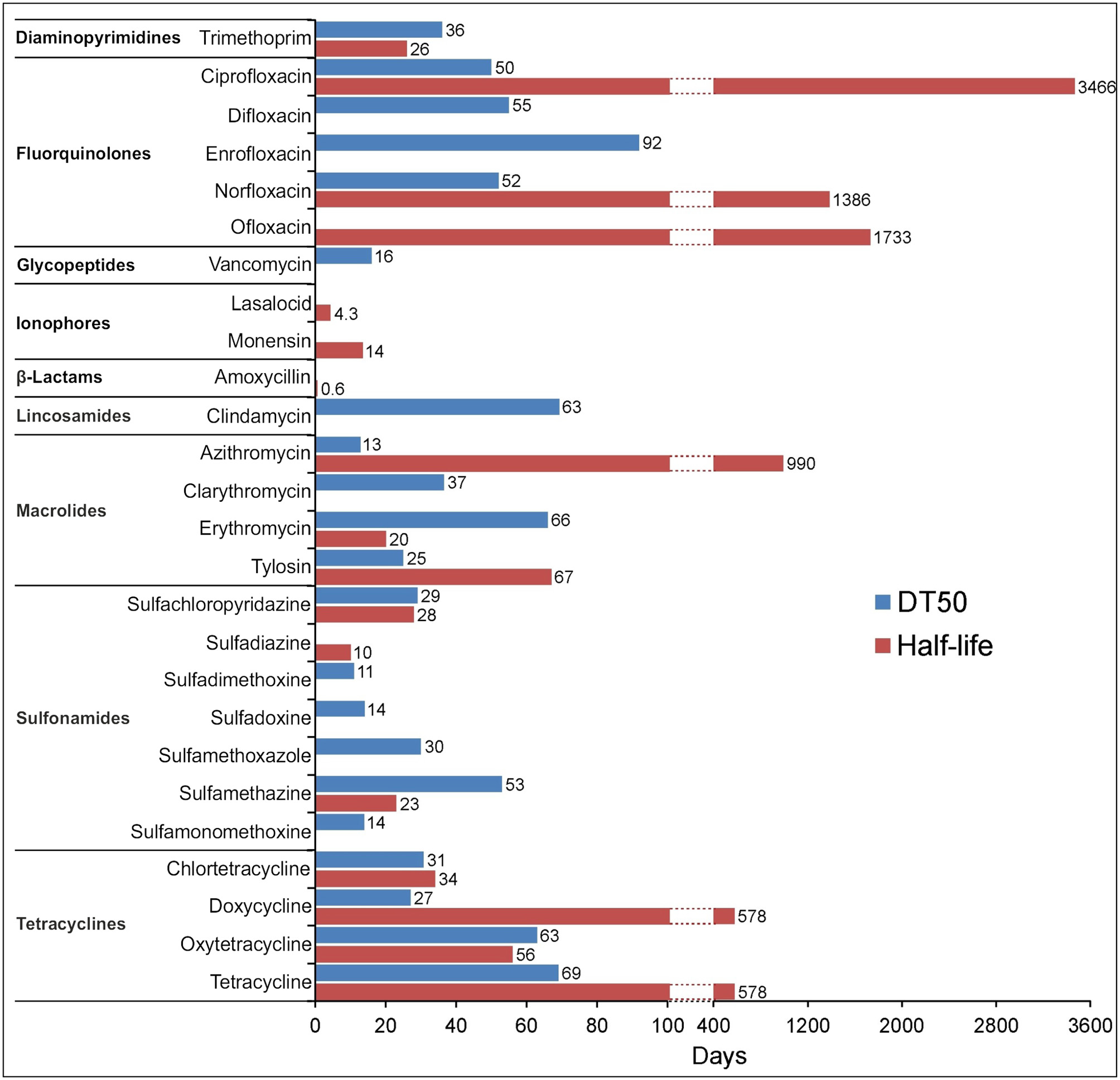
Maximum DT50 and half-life values for the selected antibiotics obtained from available degradation studies (Table 3) irrespective of the type of soil and the concentration of antibiotic used. The absence of bars in some cases means no data available.
The sorption coefficient (Koc) is a very important parameter for estimating environmental distribution and environmental exposure level of antibiotics. As was proposed by Crane et al. (2010), antibiotics with values of Koc > 4,000 l/kg are non-mobile and degradable to a very low degree in soils (very persistent) and the length of time needed for the degradation of 50% of an initial dose is >60 days. In contrast, antibiotics characterized by values of Koc < 15 l/kg are highly mobile and are more easily degraded; these can be classified as compounds with low persistence in soils (DT50 < 5 days). The relatively low persistence of some antibiotics in soils may be related to their low affinity to various organic and non-organic soil components. In contrast, due to their properties tetracyclines, fluoroquinolones, macrolides, and sulfonamides, bind strongly to soil components and form stable residues. For example, Hammesfahr et al. (2008) showed that the recovery rate of sulfadiazine on day 1 of an experiment was 27 and 45% of the initial antibiotic dosages of 10 and 100 mg/kg soil, respectively. Similarly, sulfamethazine applied at concentrations of 20 and 100 mg/kg soil strongly absorbed to soil components and measured concentrations of antibiotic were 62.1 and 31.5 μg/kg soil at the beginning of the experiment and, 255.5 and 129.8 μg/kg soil on day 56, respectively (Awad et al., 2016). Due to the binding of antibiotics to soil particles or the occurrence in a form of complexes, sorbed antibiotics often cannot be detected and may lose their antibacterial activity (Kümmerer, 2009). The adsorption and desorption of antibiotics are also associated with other soil parameters, such as pH and water content. For example, sulfonamides show a decrease in sorption with an increase of soil pH, whereas the binding of macrolides by soil components increases with a decrease of pH. In the first case, the sorption behavior is consistent with changes in the fraction of ionization of the sulfonamide as it converts from its cationic form to the neutral and anionic forms. The behavior of sulfonamides contrasts with tetracyclines and fluoroquinolones, which interact with soil primarily through cation exchange, surface complexation and cation bridging sorption mechanisms. In general, decreases in pH resulted in increased sorption of the cationic forms of antibiotics, suggesting that electrostatic interactions are the favored sorption mechanism for sulfonamides and macrolides (Schauss et al., 2009; Hammesfahr et al., 2011; Kong et al., 2012; Teixidó et al., 2012; Sittig et al., 2014; Wegst-Uhrich et al., 2014; Fernández-Calviño et al., 2015; Wang and Wang, 2015;Liu et al., 2017).
In addition to abiotic processes, microbial degradation may contribute to disappearance of antibiotics in soil. Some bacteria that degrade antibiotics have been isolated from antibiotics-contaminated soils. For example, strains belonging to the genera Microbacterium (Topp et al., 2013), Burkholderia (Zhang and Dick, 2014), Stenotrophomonas (Leng et al., 2016), Labrys (Mulla et al., 2018), Ochrobactrum (Zhang et al., 2017b; Mulla et al., 2018), and Escherichia (Mulla et al., 2018; Wen et al., 2018) were capable of degrading sulfamethazine, penicillin G, tetracycline, erythromycin and doxycycline in liquid cultures, respectively. Other bacteria belonging to the genera Klebsiella (Xin et al., 2012), Acinetobacter, Escherichia (Zhang et al., 2012), Microbacterium (Kim et al., 2011), Labrys (Amorim et al., 2014), and Bacillus (Rafii et al., 2009; Erickson et al., 2014) that were capable of degrading chloramphenicol, sulfapyridine, sulfamethazine, ciprofloxacin, norfloxacin, and ceftiofur have been isolated from patients, sediments, sludge, animal feces, and seawater. In particular, a Microbacterium sp. exhibited degradation of sulfamethazine in soil and, when introduced into agricultural soil, increased the mineralization of that antibiotic by 44–57% (Hirth et al., 2016). The central role of microorganisms in antibiotic degradation or transformation in soil has been confirmed by results of many studies carried out in sterile and non-sterile soils. As depicted in Table 3, the half-life or DT50 values of antibiotics were much lower in soils with autochthonous microorganisms compared to those obtained from sterilized soils. For example, Pan and Chu (2016) showed that when applied to soil at a dosage of 0.1 mg/kg soil, erythromycin disappeared faster in non-sterile soil compared to sterile soil, with DT50 of 6.4 and 40.8 days, respectively. Sulfachloropyridazine applied at 10 mg/kg soil was degraded almost three times faster in soils with autochthonous microorganisms (half-life 20–26 days) compared to sterile soils (half-life 68–71 days) (Accinelli et al., 2007). Zhang et al. (2017a) also showed that microbial activity plays a major role in the biotransformation of sulfadiazine in soil s, with a DT50 of 8.48, 8.97, and 10.22 days (non-sterile soil) and 30.09, 26.55, and 21.21 days (sterile soil) for concentrations of 4, 10, and 20 mg/kg soil, respectively. A similar phenomenon has also been observed for other antibiotics, such as norfloxacin (Pan and Chu, 2016), sulfamethoxazole (Lin and Gan, 2011; Srinivasan and Sarmah, 2014; Zhang et al., 2017a), sulfamethazine (Accinelli et al., 2007; Pan and Chu, 2016), and tetracycline (Lin and Gan, 2011; Pan and Chu, 2016).
Special attention should be paid to the analytical methods for extraction and determination of antibiotics residue when evaluating and comparing degradation experiments. Some techniques may not always be capable of differentiating between degradation and sorption, and insufficient or improper extraction procedures may result in incorrect interpretation of the fate of antibiotics in the soil as antibiotics that are tightly bound to soil particles could be erroneously considered to have been transformed or degraded.
Factors Affecting the Degradation of Antibiotics in Soil
Degradation of antibiotics depends not only on the catabolic activity of soil microorganisms, but also, to a large extent, on the properties of soil, i.e., organic matter content, pH, moisture, temperature, oxygen status, and soil texture (Table 3). For example, Li et al. (2010a) observed differences in the degradation rate of oxytetracycline in two agricultural soils with different characteristics. The calculated DT50 for this antibiotic reached values of 30.2 and 39.4 days for soils with a low or high organic carbon content, respectively. Soil type was also found to significantly affect antibiotics degradation, as demonstrated by Koba et al. (2017) who studied the degradation rate of clindamycin, sulfamethoxazole, and trimethoprim in 12 different soils. The authors characteristics, 44–98, 25–99, and 13–84% of clindamycin, sulfamethoxazole and trimethoprim (2 mg/kg soil), respectively, were degraded within 61 days during the experiment. A similar dependence of the rate of antibiotics degradation on soil type of has also been found for chlortetracycline (Halling-Sørensen et al., 2005; Li et al., 2010a), sulfachloropyridazine (Accinelli et al., 2007), sulfadiazine (Förster et al., 2009; Sittig et al., 2014), sulfamethoxazole (Accinelli et al., 2007; Lin and Gan, 2011; Srinivasan and Sarmah, 2014), and tetracycline (Li et al., 2010a). Another study revealed a significant decrease in the rate of chlortetracycline, erythromycin, and tylosin degradation introduced at a dosage of 5.6 mg/kg soil into a sandy loam soil. These degradation experiments showed that 56, 12, and 0%, 100, 75, and 0%, or 100, 100, and 60% of the initial concentration of chlortetracycline, erythromycin, and tylosin were degraded at temperatures of 30, 20, and 4°C, respectively, within 30 days (Gavalchin and Katz, 1994). In addition, Srinivasan and Sarmah (2014) showed that a lower temperature resulted in a reduced decomposition rate of sulfamethoxazole, irrespective of the soil depth. Some published data have also shown the influence of oxygen in soils on the degradation of antibiotics. For example, Pan and Chu (2016) showed that the half-lives for erythromycin, norfloxacin, sulfamethazine, and tetracycline, all of which had been applied at 0.1 mg/kg soil, increased from 6.4, 2.9, 24.8, and 31.5 days under aerobic conditions to 11.0, 5.6, 34.7, and 43.3 days under anaerobic conditions, respectively. Anaerobic conditions caused an increase in the half-life for sulfamethoxazole by seven days compared to 11.4 days for aerobic degradation of this antibiotic. In turn, no significant effects of incubation conditions on degradation rate were shown for trimethoprim, for which the calculated half-life was about 26 days under both aerobic and anaerobic conditions (Lin and Gan, 2011). The degradation of antibiotics may also be influenced by the pH and moisture content of soils. Weerasinghe and Towner (1997) showed that the half-life of for virginiamycin was negatively correlated with the pH of soils and ranged from 87 to 173 days for different agricultural soils. A study by Wang et al. (2006) revealed that the half-life of sulfadimethoxine in soil decreased from 10.4 days to 6.9, and again to 4.9 days when the soil moisture content was increased from 15, to 20, and 25%, respectively. A similar effect of temperature was shown in the case of the degradation of norfloxacin (Yang et al., 2012).
Degradation of antibiotics strongly depends on their concentration in soil. Increasing dosages of ciprofloxacin (from 1 to 5 and 50 mg/kg soil) led to a reduction of degradation from 75, to 62, and 40% within 40 days (Cui et al., 2014). A similar tendency was also shown by Demoling et al. (2009), who applied sulfamethoxazole at 500, 20, and 1 mg/kg soil and observed a reduction of its removal from 153, to 1.5, and 0.04 mg/kg after 5 weeks. The same trend was reported in degradation of azithromycin, ofloxacin, tetracycline (Walters et al., 2010), sulfadimethoxine (Wang et al., 2006), chlortetracycline (Fang et al., 2016), and SDZ (Zhang et al., 2017a). These results suggest that high concentrations of antibiotics may prolong their persistence in soils, due to the inhibition of the activity of soil microorganisms (Yang et al., 2009; Pan and Chu, 2016). However, application of unrealistically high concentrations tends to overestimate half-lives, which may not reflect realistic situations (Pan and Chu, 2016). The history of antibiotics application also plays a role in the further disappearance of antibiotics in soils. Repeated application of the macrolide antibiotics clarithromycin and erythromycin into soil resulted in a decrease of the DT50 values from 36.48 and 69.93 days to 15.85 and 4.36 days (soil with a history of clarithromycin and erythromycin application at 0.1 mg/kg soil) and to 9.51 and 0.94 days (soil with a history of clarithromycin and erythromycin application at 10 mg/kg soil), respectively (Topp et al., 2016). This phenomenon of pre-adaptation of soil microorganisms was also found for the degradation of chlortetracycline, sulfamethazine and tylosin (Topp et al., 2013).
Most published data have also indicated that the addition of different organic compounds, such as manure, biosolids, slurry, sludge, and compost, into soils may contribute to changes in the rate of antibiotics degradation (Table 3). Sassman and Lee (2007) showed that the addition of manure (20 mg/kg soil) increased the half-life of lasalocid from 3.6 to 4.3 days. In contrast, Yang et al. (2012) showed that norfloxacin (10 mg/kg soil) was degraded faster (half-life 24–39 days) in soil with manure (3–9%) compared to the control soil (half-life 48 days). In turn, Li et al. (2010a) found no significant effects from the addition of manure on degradation of chlortetracycline in soil.
Impact of Antibiotics on Soil Microorganisms
Soil microorganisms perform many vital processes and participate in the maintenance of soil health and quality. They play a crucial role in organic matter turnover, nutrients release, and stabilization of the soil structure and ensure soil fertility. Moreover, many microorganisms act as biological control agents by inhibiting the growth of pathogens (Varma and Buscot, 2005). The homeostasis of soil may be disturbed by biotic and abiotic factors, including bacteriophages, predation, competition, pesticide, heavy metals, toxic hydrocarbons, and antibiotics (Cycoń et al., 2011; Cycoń and Piotrowska-Seget, 2015, 2016; Sułowicz and Piotrowska-Seget, 2016; Xu et al., 2016; Chen et al., 2017; Wepking et al., 2017; Orlewska et al., 2018a). The high antimicrobial activity of antibiotics in soil should differentially inhibit the growth of soil microorganisms and thus influence the soil microbial community composition, which may result in alterations of the ecological functionality of the soil (Kotzerke et al., 2008; Keen and Patrick, 2013; Molaei et al., 2017) (Figure 3).
There is abundant data on the impact of antibiotics on microorganisms and on soil processes mediated by bacteria and fungi. Both the effects of antibiotics on individual microbial populations as well as on the composition of entire microbial communities have been documented. A wide range of methods based on measurements of parameters that reflect the activity and abundance of total microbial communities, such as soil organic matter turnover, respiration, and microbial biomass have been used to characterize such effects. Other methods have focused on selected microbial processes such as nitrification, denitrification, sulfate, and iron reduction, methanogenesis and the activity of enzymes responsible for C, N, and P turnover (Brandt et al., 2009; Chen et al., 2013; Cui et al., 2014; Liu et al., 2015; Ma et al., 2016). A large number of studies have instead focused on assessing changes in microbial diversity, using metagenomics or 16S rRNA gene amplicon sequencing, as well as analysis of phospholipid fatty acids (PLFAs) isolated from soil (Zielezny et al., 2006; Hammesfahr et al., 2008; Reichel et al., 2015; Xu et al., 2016). In the following sections, the methods and parameters utilized to assess effects of antibiotics on the function and structure of soil microbial communities are presented and discussed (Figure 3).
Impact of Antibiotics on Soil Microbe Function
Soil Processes
Many studies have found that even low concentrations (below the MIC) of antibiotics influence various soil processes mediated by microorganisms (Table 4). A significant decrease in soil respiration (SR) was reported for soils containing sulfamethoxazole, sulfamethazine, sulfadiazine, and trimethoprim (Kotzerke et al., 2008; Liu et al., 2009), however, this effect was transient and depended on the disappearance rate of these antibiotics (DT50 2–5 days). The authors concluded that the effects of antibiotics on SR diminished due to decreased bioavailability of the antibiotic. In a study by Wepking et al. (2017), the response of microbial respiration to cephapirin, tetracycline, or erythromycin was dependent on both the type of antibiotic and the exposure of soil to dairy cattle manure. In other studies, no obvious effects of tetracycline, chlortetracycline, oxytetracycline, sulfadiazine, sulfapyridine, or tylosin on SR were observed (Thiele-Bruhn and Beck, 2005; Liu et al., 2009; Toth et al., 2011).
Table 4
| Antibiotic | Dosage (mg/kg soil) | Type of soil/origin | Main properties of soil | Experimental conditions | Effect (comparison with control, non-treated soil) | References |
|---|---|---|---|---|---|---|
| 100 | Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | Basal respiration was stimulated | Ma et al., 2014 | |
| Ciprofloxacin | 1, 5, and 50 | Ustic Cambisol (China) | Sand 12%, silt 54%, clay 34%, pH 7.9, OC 36.76 g/kg, CEC 13.82 cmol/kg | 25°C, 60% WHC, 40 days | Basal respiration was higher in treated soils at intermediate concentrations than in control at 9 and 22 days; nitrification was stimulated at 1 mg/kg and inhibited at 50 mg/kg after 9 days, nitrate and ammonium contents were not altered after antibiotic addition | Cui et al., 2014 |
| 0.2, 2, and 20 | Agricultural soil (Germany) | Sand 11%, silt 68%, clay 21%, pH 6.6, WHC 37.5%, OC 2.1% | 20°C, 60% WHC, 93 days, sludge addition at 1.8 g/kg | Soil respiration was inhibited by ~70% at all three concentrations after 2 days and as only about 35% at the end of the experiment | Girardi et al., 2011 | |
| Chlortetracycline | 1–50 | Orthic Luvisol (Germany) | Sand 3%, silt 79%, clay 18%, pH 7, OC 1.04%, WHC 48.8% | 20°C, 48 days | Basal respiration was not affected | Zielezny et al., 2006 |
| 1–300 | Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | In general, no inhibitory effect on respiration was found | Liu et al., 2009 | |
| 0.0003–0.03 | Silt loam (USA) | pH 6.5, OM 2.2%, WHC 0.313 g/g, CEC 10.6 cmol+/kg | 80% WHC, manure addition, 50 days | No effect on nitrification, Fe(III) reduction or soil respiration | Toth et al., 2011 | |
| Difloxacin | 1–100 | Loamy sand (Germany) | Sand 73.3%, silt 23.1%, clay 3.6%, pH 5.5, OC 1.7%, WHC 27% | 10°C, 60% WHC, manure addition at 40 ml/kg, 32 days | Increase rate of respiration up to 8 days; no detectable effect on ammonium and nitrate rates; higher nitrification in all treatments on day 8 and their reduction at 10 and 100 mg/kg on day 32; a significant lower potential denitrification on days 4 and 8 in all treatments | Kotzerke et al., 2011 |
| Monensin | 0.01–0.100 | Silt loam (USA) | pH 6.5, OM 2.2%, WHC 0.313 g/g, CEC 10.6 cmol+/kg | 80% WHC, manure addition, 50 days | Soil respiration was not affected; inhibition of the Fe(III) reduction was transient; low concentrations inhibited nitrogen transformation | Toth et al., 2011 |
| Norfloxacin | 5, 10, and 30 | Acidic Soil (China) | Sand 29%, silt 39%, clay 32%, pH 4.3, OM 2.4%, CEC 9.5 cmol+/kg | 25°C, 50% WHC, 42 days | No obvious inhibition on soil respiration, only slight effect on nitrogen transformation | Yang et al., 2012 |
| Oxytetracycline | 1–100 | Agricultural soil (Iran) | Sand 52.35%, silt 29.23%, clay 18.42%, pH 4.3, OC 0.95%, WHC 20% | 25°C, 50% WHC, 21 days | Impact on Fe(III) reduction at 1 and 10 mg/kg, and Fe(III) reduction was completely inhibited at concentrations above 10 mg/kg; negatively affected respiration throughout the experiment | Molaei et al., 2017 |
| 1–30 | Alfisol (China) | Sand 7.7%, silt 77.5%, clay 14.8%, pH 6.24, OM 2.4%, CEC 12.3 cmol/kg, OTC 37.3 μg/kg | 25°C, 60% WHC, 120 days | Nitrification decreased over the experiment with transient increase on day 28 | Ma et al., 2016 | |
| 10–1,000 | Luvisol (Germany) | Sand 68.4%, silt 20.4%, clay 9.9%, pH 7.1, OC 1.6%, CEC 13.1 cmol/kg | 20–25°C, 50% WHC, 14 days | No detectable effect on basal respiration; EC50 for Fe(III) reduction was 96 and 5.3 μg/g for Cambisol and Luvisol, respectively | Thiele-Bruhn and Beck, 2005 | |
| Cmabisol (Germany) | Sand 80.9%, silt 15.9%, clay 3.1%, pH 6.6, OC 0.8%, CEC 5.3 cmol/kg | |||||
| Sulfadimethoxine | 0.025–0.200 | Silt loam (USA) | pH 6.5, OM 2.2%, WHC 0.313 g/g, CEC 10.6 cmol+/kg | 80% WHC, manure addition, 50 days | Soil respiration was not affected; inhibition of soil nitrogen transformation; nearly completely blocked Fe(III) reduction throughout the 50-day experiment at higher concentrations (0.1 and 0.2 mg/kg) | Toth et al., 2011 |
| Sulfadiazine | 10 and 100 | Silt loam (Germany) | Sand 6.4%, silt 78.2%, clay 15.4%, pH 7.2, OC 2.1%, WHC 46% | 20°C, 32/61 days, manure addition at 40 g/kg soil | Only at 100 mg/kg higher ammonium and lower nitrate concentrations were detected; a reduction in CO2 production at the beginning of the experiment only in a higher treatment; increase and reduction of nitrification at 10 and 100 mg/kg, respectively after 32 days | Kotzerke et al., 2008; Schauss et al., 2009 |
| Loamy sand (Germany) | Sand 73.3%, silt 23.1%, clay 3.6%, pH 5.5, OC 1.7%, WHC 27% | Increased amounts of ammonium and reduced amounts of nitrate were determined in both treatments after 61 days; a reduction in CO2 production at the beginning of the experiment in both treatments; increase and reduction of nitrification at 10 and 100 mg/kg, respectively after 32 days | ||||
| 1–50 | Orthic Luvisol (Germany) | Sand 3%, silt 79%, clay 18%, pH 7, OC 1.04%, WHC 48.8% | 20°C, 48 days | Basal respiration was not affected | Zielezny et al., 2006 | |
| 10 and 100 | Luvisol (China) | Sand 58.4%, silt 21.7%, clay 19.9%, pH 6.24, OM 3.56%, CEC 5.38 cmol/kg | 25°C, 25% WHC, manure addition at 40 mg/kg, 28 days | Inhibition of basal respiration by both dosages on days 1 and 7; increase in basal respiration at 10 mg/kg after 28 days | Xu et al., 2016 | |
| Endogleyic Cambisol (Germany) | Sand 73.3%, silt 23.1%, clay 3.6%, pH 4.8, OC 1% | 10°C, 50% WHC, manure addition at 20, 40, and 80 g/kg, 32 days | No significant impact on the rate of SIR | Hammesfahr et al., 2011 | ||
| 10°C, 60% WHC, manure addition at 40 g/kg, 57 days | Reduction of nitrification and N mineralization; increase of ammonification | Hammesfahr et al., 2011 | ||||
| Sulfamonomethoxine | 100 | Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | Basal respiration was stimulated | Ma et al., 2014 |
| Sulfamethoxazole | 1–300 | Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | Decrease of respiration within the first 4 days; increase with increasing antibiotic concentrations after initial inhibition | Liu et al., 2009 |
| 1–100 | Agricultural soil (Iran) | Sand 52.35%, silt 29.23%, clay 18.42%, pH 4.3, OC 0.95%, WHC 20% | 25°C, 50% WHC, 21 days | Affected Fe(III) reduction at 1 and 10 mg/kg, and Fe(III) reduction was completely inhibited at concentrations above 10 mg/kg; affected respiration at different treatments over experimental period | Molaei et al., 2017 | |
| Sulfamethazine | 20 and 100 | Silt loam (Korea) | pH 6.0, OM 2.36% | 25°C, 70% WHC, 56 days, poultry manure addition (1%) | Increase of the CO2-C efflux during incubation time except for 56 day; increase of the nitrification at 28 and 56 days | Awad et al., 2016 |
| Sulfapyridine | 10–1,000 | Luvisol (Germany) | Sand 68.4%, silt 20.4%, clay 9.9%, pH 7.1, OC 1.6%, CEC 13.1 cmol/kg | 20–25°C, 50% WHC, 14 days | Basal respiration was not affected; EC50 for Fe(III) reduction was 12,400 and 310 μg/g for Cambisol and Luvisol, respectively | Thiele-Bruhn and Beck, 2005 |
| Cmabisol (Germany) | Sand 80.9%, silt 15.9%, clay 3.1%, pH 6.6, OC 0.8%, CEC 5.3 cmol/kg | |||||
| Tetracycline | 100 | Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | Basal respiration was stimulated No inhibitory effect on respiration |
Ma et al., 2014 |
| Trimethoprim | 1–300 | Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | Liu et al., 2009 | |
| Decrease of respiration within the first 4 days; increase with increasing antibiotic concentrations after initial inhibition No inhibitory effect on respiration |
||||||
| Tylosin | 2,000 | Sand (Denmark) | Sand 90.7%, silt 2.8%, clay 4.1%, pH 6.8, WHC 15%, OC 1.2% | 25°C, 60 days | No significant effect on soil respiration | Müller et al., 2002 |
Effects of the selected antibiotics on the microbial processes in soils with different characteristics.
CEC, cation exchange capacity; OC, organic carbon; OM, organic matter; SIR, substrate-induced respiration; WHC, water holding capacity.
Nitrification and/or denitrification rates were also influenced by antibiotic exposure, and the effects were strongly dependent on the type of antibiotic and the length of exposure. For example, sulfadimethoxine inhibited soil nitrification; however, this effect was only observed on some sampling days and only for a high sulfadimethoxine treatment (Toth et al., 2011). A decrease in the nitrification rate caused by a high oxytetracycline concentration (30 mg/kg) and sulfadiazine (100 mg/kg) in a single application was also observed by Ma et al. (2016) and Kotzerke et al. (2008), whereas ciprofloxacin and norfloxacin were reported to stimulate the rate of nitrification in a soil microcosm, but only at the lowest concentration of antibiotic (1 mg/kg soil) (Yang et al., 2012; Cui et al., 2014). A study by DeVries et al. (2015) revealed that low doses of sulfamethoxazole, sulfadiazine, narasin, or gentamicin (500 μg/kg soil) inhibited denitrification, but dosages < 1 μg/kg soil actually stimulated the process transiently. Toth et al. (2011) did not observe any effects of monensin and chlortetracycline, on soil nitrification at concentrations of 0.01–0.1 and 0.0003–0.3 mg/kg soil, respectively.
Antibiotics may also change the turnover rate of iron in soil. Sulfadiazine and monensin blocked Fe(III) reduction in soil over periods ranging from a few days to the end of a 50-day experiment (Toth et al., 2011). Strong inhibition of Fe(III) reduction was also observed in soil contaminated with sulfamethoxazole and oxytetracycline at concentrations >10 mg/kg soil (Molaei et al., 2017). Thiele-Bruhn and Beck (2005) calculated the EC50 value of sulfapyridine for the microbial reduction of Fe(III) in two different soils at 12.4 and 0.310 mg/kg soil. It should be noted that the lack of standardized tests hinders comparisons that would lead to general conclusions about the effects of antibiotics on biogeochemical cycles and the turnover of iron.
Enzyme Activities
Specific enzyme activity is considered to be a useful indicator of the response of microorganisms to stress caused in the soil by antibiotics (Unger et al., 2013; Liu et al., 2014, 2015; Ma et al., 2016) (Table 5). Enzyme activity indicates the potential of microbial communities to carry out biochemical processes that are essential to maintain soil quality. Any application of a toxicant that might affect the growth of soil microorganisms can induce alterations in the general activity of enzymes, such as dehydrogenases (DHAs), phosphatases (PHOSs), and urease (URE) (Gil-Sotres et al., 2005; Hammesfahr et al., 2011; Cycoń et al., 2016a). Inhibition of DHAs and URE was observed in soil amended with tetracycline at a concentration of 1 μg/kg soil, however this dosage did not affect acid phosphatase (PHOS-H) activity. Sulfamethazine applied at 53.6 μg/g soil had a significant short-term negative impact on the activities of DHAs and URE (Pinna et al., 2012). Inhibition of DHAs and arylsulphatase activities with increasing concentration of oxytetracycline at 1 to 200 mg/kg over 7 weeks was also reported by Chen et al. (2013). Benzylpenicillin, tylosin and sulfadiazine inhibited soil DHAs and PHOSs from 35 to 70% compared to the non-antibiotic amended control (Reichel et al., 2014; Akimenko et al., 2015). The results obtained by Unger et al. (2013) showed that the application of oxytetracycline or lincomycin (50 and 200 mg/kg soil) resulted in a temporary decrease in DHA activity in soil. A temporary increase, followed by a decrease, in DHA activity was found in soils containing chlortetracycline (1, 10, and 100 mg/kg soil) by Liu et al. (2015). In another study, DHA activity was unaffected by sulfapyridine or oxytetracycline, even at a dosage of 1 mg/kg soil (Thiele-Bruhn and Beck, 2005).
Table 5
| Antibiotic | Dosage (mg/kg soil) | Type of soil/origin | Main properties of soil | Experimental conditions | Effect (comparison with control, non-treated soil) | References |
|---|---|---|---|---|---|---|
| 1–300 | Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | Inhibition of acid PHOS during the 22-day experiment | Liu et al., 2009 | |
| Chlortetracycline | 10 and 100 | No data (China) | Sand 42.95%, silt 43.43%, clay 13.65%, pH 7.6, OC 20.7 g/kg | 25°C, 50% WHC, 45 days, DOM addition at 40 mg C/kg | Inhibition of DHA until day 12; decrease of acid PHOS on days 6 and 12 at 10 and 100 mg/kg, respectively; inhibition of URE by 100 mg/kg on day 45 | Liu et al., 2014 |
| 1–100 | No data (China) | 25°C, 50% WHC, 45 days | Stimulation of all enzyme activities on the first day and then inhibition of DHA and URE up to 45 days; slight effect on PHOS | Liu et al., 2015 | ||
| Lincomycin | 5–200 | Silt loam (USA) | Clay 23.4–26.2%, pH 6.7–7.3, OC 19.3–26.0 g/kg | 25°C, 35% WHC, 63 days | Low DHA at 50 and 200 mg/kg and thereafter increase in all treatments; FDA at 50 and 200 mg/kg significantly higher on day 7 | Unger et al., 2013 |
| Oxyteracycline | 1–200 | Loam (China) | Sand 52.2%, silt 38.6%, clay 9.2%, pH 8.3, OM 1.2% | 20–27°C (day) and 15–20°C (night), 50–70% WHC, manure addition at 30 mg/g, 7 weeks | DHA, ARYL, PHOS, and URE decreased with increasing concentrations of OTC | Chen et al., 2013 |
| 5–200 | Silt loam (USA) | Clay 23.4–26.2%, pH 6.7–7.3, OC 19.3–26.0 g/kg | 25°C, 35% WHC, 63 days | DHA declined at 50 and 200 mg/kg up to day 35 and recovered on day 63; FDA at 200 mg/kg significantly higher on day 7 and recovered on day 14 | Unger et al., 2013 | |
| 10–70 | Wheat rhizosphere soil (China) | No data | 30 days | Among alkaline PHOS, acid PHOS, DHA, and URE, only alkaline PHOS was higher at 10 mg/kg but further decreased at the dosage over 30 mg/kg | Yang et al., 2009 | |
| 1–30 | Alfisol (China) | Sand 7.7%, silt 77.5%, clay 14.8%, pH 6.24, OM 2.4%, CEC 12.3 cmol/kg, OTC 37.3 μg/kg | 25°C, 60% WHC, 120 days | Stimulation of DHA on day 14 and decrease by 120 days; no marked effect on PHOS and URE over the 120-day experiment | Ma et al., 2016 | |
| 10–1,000 | Luvisol (Germany) | Sand 68.4%, silt 20.4%, clay 9.9%, pH 7.1, OC 1.6%, CEC 13.1 cmol/kg | 20–25°C, 50% WHC, 14 days | No detectable effect on DHA | Thiele-Bruhn and Beck, 2005 | |
| Cmabisol (Germany) | Sand 80.9%, silt 15.9%, clay 3.1%, pH 6.6, OC 0.8%, CEC 5.3 cmol/kg | |||||
| Penicillin G | 100 and 600 | Loamy soil (Russia) | pH 7.7, humus 4.1%, Ntot 0.25%, P 28.8, Ktot 2.06% | 20–25°C, 60% WHC, 120 days | Inhibition of CAT, DHA, PHOS, and INV (20–70% of the control) | Akimenko et al., 2015 |
| Sulfadimethoxine+sulfamethoxazole+sulfamethazine | 0.09–900 | Sandy loam (USA) | Sand 85.5%, silt 8.5%, clay 6.6%, pH 6.31, OC 0.86%, CEC 8.1 cmol+/kg | 20°C, 21 days, glucose and/or manure addition | DHA and URE activities decreased with higher concentrations | Gutiérrez et al., 2010 |
| Sulfadiazine | 10 and 100 | Luvisol (China) | Sand 58.4%, silt 21.7%, clay 19.9%, pH 6.24, OM 3.56%, CEC 5.38 cmol/kg | 25°C, 25% WHC, manure addition at 40 mg/kg, 28 days | Inhibition of FDA by both dosages; DHA was significantly inhibited at 10 and 100 mg/kg within 14 days, a significant increase of DHA at 10 mg/kg after 28 days | Xu et al., 2016 |
| Endogleyic Cambisol (Germany) | Sand 73.3%, silt 23.1%, clay 3.6%, pH 4.8, OC 1% | 10°C, 50% WHC, manure addition at 20, 40, and 80 g/kg, 32 days | No significant impact on β-GLU; declined URE | Hammesfahr et al., 2011 | ||
| Sulfamethoxazole | 1–300 | Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | Decline in acid PHOS during the 22-day experiment | Liu et al., 2009 |
| 5 | Silt loam (New Zeland) | Sand 9%, silt 54%, clay 37%, pH 6.7, OC 5% | 25°C, 60% WHC, 36 days | DHA was not affected by antibiotic; an increase in the DHA | Srinivasan and Sarmah, 2014 | |
| Clay loam (New Zeland) | Sand 13.7%, silt 51%, clay 30.4%, pH 5.8, OC 4% | |||||
| Silt loam (New Zeland) | Sand 34%, silt 48%, clay 17%, pH 5.7, OC 8.2% | |||||
| Sulfamethazine | 1–300 | Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | Inhibition of acid PHOS during 22 days | Liu et al., 2009 |
| 53.6 | Sandy loam (Italy) | Sand 72.7%, silt 10.6%, clay 16.6%, pH 7.8, OC 2.8% | 7 days | A significant decrease of DHA (41%) and URE (38%) after 1 day and this effect disappeared after 7 days | Pinna et al., 2012 | |
| Sand 81.7%, silt 5.9%, clay 12.2%, pH 5.3, OC 1.7% | A significant decrease of DHA (17%) and URE (27%) after 1 day and this effect disappeared after 7 days | |||||
| Sulfapyridine | 10–1,000 | Luvisol (Germany) | Sand 68.4%, silt 20.4%, clay 9.9%, pH 7.1, OC 1.6%, CEC 13.1 cmol/kg | 20–25°C, 50% WHC, 14 days | No detectable effect on DHA | Thiele-Bruhn and Beck, 2005 |
| Cmabisol (Germany) | Sand 80.9%, silt 15.9%, clay 3.1%, pH 6.6, OC 0.8%, CEC 5.3 cmol/kg | |||||
| Tetracycline | 1–300 | Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | Inhibition of acid PHOS during 22 days | Liu et al., 2009 |
| 100 and 500 | Clay (Italy) | Sand 39.4%, silt 19.2%, clay 41.4%, pH 5.8, OM 6.9% | 20°C, 60 days | Decrease in FDA after 2 days; this effect disappeared after 7 days | Chessa et al., 2016b | |
| Sand (Italy) | Sand 72.7%, silt 10.6%, clay 16.6%, pH 7.6, OM 4.9% | |||||
| Trimethoprim | 1–300 | Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | Inhibition of acid PHOS during 22 days | Liu et al., 2009 |
| Tylosin | 1–300 | Silt loam (China) | OC 18.2 g/kg, pH 5,7 | 25°C, 60% WHC, 22 days | Decline in acid PHOS during the 22-day experiment | Liu et al., 2009 |
| 100 and 600 | Loamy soil (Russia) | pH 7.7, humus 4.1%, Ntot 0.25%, P 28.8, Ktot 2.06% | 20–25°C, 60% WHC, 120 days | Suppressing effect on CAT, DHA, PHOS, and INV (20–70% of the control) | Akimenko et al., 2015 | |
| Vancomycin | 1 and 10 | Sandy loam (Poland) | Sand 67%, silt 24%, clay 9%, pH 6.9, WHC 43%, OC 1.6%, CEC 10 cmol/kg | 22°C, 50% WHC, 90 days | A negative impact on days 1, 15, and 30 as was showed by a decrease in the values of DHA, PHOS and URE (6–32%) | Cycoń et al., 2018 |
Effects of the selected antibiotics on the enzyme activities in soils with different characteristics.
ARYL, arylsulfatase; CAT, catalase; CEC, cation exchange capacity; DHA, dehydrogenase; FDA, fluorescein diacetate; β-GLU, β-glucosidase; INV, invertase; OC, organic carbon; OM, organic matter; PHOS, phosphatase; URE, urease; WHC, water holding capacity.
Various effects on the activity of PHOSs from different antibiotics applied to soil have been shown. For example, six antibiotics, i.e., chlortetracycline, tetracycline, tylosin, sulfamethoxazole, sulfamethazine, and trimethoprim at dosages of 1–300 mg/kg soil inhibited the activity of PHOS-H (Liu et al., 2009, 2015). In contrast, results from Yang et al. (2009) indicated that only alkaline phosphatase (PHOS-OH) was sensitive to the application of oxytetracycline, with a 41.3% decline in enzyme activity at a concentration of 10 mg/kg soil and a further decrease of 64.3–80.8% when the dose of the antibiotic exceeded 30 mg/kg. Ma et al. (2016) found that OTC at dosages up to 30 mg/kg soil had no effect on the activity of neutral PHOS over a 120-day experimental period.
The inhibition of enzyme activity in antibiotic-treated soils may be related to inhibition of growth or death of sensitive microorganisms (Boyd and Mortland, 1990; Gianfreda et al., 1994; Alef, 1995; Marx et al., 2005). In turn, the increased activity of enzymes under antibiotics pressure may result from the ability of many bacteria to subsist on antibiotics as a carbon source (Dantas et al., 2008). We can speculate that enzymes produced by such bacteria compensate for the negative effects of antibiotics on enzyme activity by increasing the activity of the antibiotic-resistant microbial community. Similarly, an increase in the activity of some enzymes in soils treated with different antibiotics might be related to the capability of some microorganisms to make use of the antibiotics in their metabolisms, thus resulting in an increase in the abundance of some soil microorganisms and enzyme production. In addition, the presence of some antibiotics in soil may cause an overgrowth of fungi, which are generally less sensitive to antibiotics than bacteria. Fungi are a major producer of enzymes in soils and so may be responsible for observed increases in enzyme activity (Tabatabai and Bremner, 1969; Westergaard et al., 2001; Hammesfahr et al., 2011; Ding et al., 2014).
Functional Capacity Evaluation
Functional microbial diversity reflects the ability of an entire microbial community to utilize a suite of substrates (Zak et al., 1994). The response of microorganisms to the presence of antibiotics in soils, expressed as the community level physiological profile (CLPP), can be evaluated using the Biolog (EcoPlates™) or MicroResp™ methods, which are based on the utilization of 31 or 15 carbon sources, respectively, by organisms present in soil extracts (Toth et al., 2011; Ma et al., 2014: Xu et al., 2016) (Table 6). Although the exact numbers and taxonomic identities of the bacterial species responsible for the Biolog reactions remain uncertain, patterns of functional diversity indicate how biodiversity affects specific ecosystem functions (Laureto et al., 2015). For determination of CLPP, the biodiversity (Shannon H') and average well-color development (AWCD) indices have been used as indicators of changes in the catabolic potential of soil microorganisms exposed to antibiotics. For example, chlortetracycline (1 and 10 mg/kg soil) caused a decrease in AWCD values, showing low catabolic potential in the microbial community, however, this effect was seen only at the beginning of the experiment. After 35 days, irrespective of the frequency of chlortetracycline application, the AWCD values gradually recovered to the control level (Fang et al., 2014, 2016). A slight inhibitory effect on microbial activity (expressed as the H′ index) in soil was observed over a range from 1 to 300 mg oxytetracycline /kg soil. Even lower concentrations of this antibiotic (0.5 to 90 mg/kg of soil) significantly decreased the functional activity (expressed as AWCD) of the soil microbial communities (Kong et al., 2006). Sulfonamide antibiotics, such as sulfamethoxazole and sulfamethazine, can also alter the activity of microbial populations, however, they were observed to only have short-term detrimental effects (Demoling et al., 2009; Pinna et al., 2012; Pino-Otín et al., 2017). In turn, the application of chlortetracycline or sulfadimethoxine to soil did not significantly change AWCD, whereas monensin slightly increased the value of the H' index (Toth et al., 2011). In many studies, changes in the preferential degradation of groups of substrates in Eco-plates, e.g., amino acids, carbohydrates, carboxylic acids, aromatic acids, and miscellaneous, by microorganisms were observed over the course of an experimental period. Xu et al. (2016) found that a high sulfadiazine concentration decreased utilization rates of amino acids, carbohydrates, carboxylic acids, and aromatic acids by bacteria present in a soil suspension. A significant decrease in the utilization of carbohydrates and miscellaneous substrates was also observed in soil that had been treated with sulfamethoxazole (Liu et al., 2012a). However, this effect was only observed 7 days after the antibiotic application; by day 21 substrate utilization had increased compared to the first sampling day. The negative impact of antibiotics on CLPP may be mitigated by the enrichment of soil with organic matter from animal wastes, antibiotic-free manure or plant wastes (Demoling et al., 2009; Pinna et al., 2012; Liu et al., 2014). Contrarily, Wang et al. (2016) found that doxycycline application at 10 mg/kg soil increased utilization of the substrates.
Table 6
| Antibiotic | Dosage (mg/kg soil) | Type of soil/origin | Main properties of soil | Experimental conditions | Effect (comparison with control, non-treated soil) | References |
|---|---|---|---|---|---|---|
| Ciprofloxacin | 100 | Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | Functional diversity decreased | Ma et al., 2014 |
| Cefuroxime | 1 and 10 | Sandy loam (Poland) | Sand 67%, silt 24%, clay 9%, pH 6.9, WHC 43%, OC 1.6%, CEC 10 cmol/kg | 22°C, 50% WHC, 90 days | A negative impact of CEF up to 30 days | Orlewska et al., 2018a |
| 1–300 | Clay loam (China) | Sand 30.4%, silt 34.1%, clay 35.5%, pH 5.7, OC 18.2 g/kg, CEC 9.87 cmol/kg | 25°C, 60% WHC, 21 days | No effect on AWCD and Shannon indices after 7 days; a significant increase of AWCD at 300 mg CTC/kg after 21 days | Liu et al., 2012a | |
| Chlortetracycline | 1 and 100 | Inceptisol (China) | Sand 21.5%, silt 71.1%, clay 7.4%, pH 6.8, OM 3.1%, WHC 27%, CEC 10.6 cmol/kg | 25°C, 60% WHC, 35 days | AWCD and functional diversity indices decreased significantly up to 35 days | Fang et al., 2016 |
| 10 and 100 | No data (China) | Sand 42.95%, silt 43.43%, clay 13.65%, pH 7.6, OC 20.7 g/kg | 25°C, 50% WHC, 45 days, DOM addition at 40 mg C/kg | Decrease in AWCD up to 45%; a significantly detrimental effect on the diversity and richness of the microbial community with maximum decrease of 4 and 21%, respectively | Liu et al., 2014 | |
| 1–100 | No data (China) | Sand 42.95%, silt 43.43%, clay 13.65%, pH 7.6, OC 20.7 g/kg | 25°C, 50% WHC, 45 days | AWCD, richness, and Shannon indices were higher with maximum values of 56.5, 42.9, and 13.9%, respectively; no significant difference on days 6 and 12, with the exception of a larger value in the highest treatment to the end of the experiment | Liu et al., 2015 | |
| 0.0003–0.03 | Silt loam (USA) | pH 6.5, OM 2.2%, WHC 0.313 g/g, CEC 10.6 cmol+/kg | 80% WHC, manure addition, 50 days | CLPP parameters, including AWCD, Shannon diversity index, and evenness was not affected | Toth et al., 2011 | |
| Doxycycline | 10 | Silty clay loam (China) | Sand 9%, silt 63.7%, clay 27.3%, pH 7, OM 1% | 25°C, 50–65% WHC, manure addition at 10 g/kg, 56 days | Positively affected on the microbial diversity | Wang et al., 2016 |
| Monensin | 0.01–0.100 | Silt loam (USA) | pH 6.5, OM 2.2%, WHC 0.313 g/g, CEC 10.6 cmol+/kg | 80% WHC, manure addition, 50 days | 4–5% increase in Shannon diversity index in 0.025, 0.05, and 0.100 mg/kg treatments | Toth et al., 2011 |
| Oxytetracycline | 1–217 | Silt loam (Anthroposols) (China) | No data | 25°C | Functional diversity, evenness, AWCD and substrate utilization decreased significantly with increasing concentrations | Kong et al., 2006 |
| 1–200 | Loam (China) | Sand 52.2%, silt 38.6%, clay 9.2%, pH 8.3, OM 1.2% | Room temperature, 50–60% WHC, manure addition at 30 mg/g, 6 days | AWCD values increased, and the utilization of sugar and its derivatives enhanced | Liu et al., 2012b | |
| 5–200 | Silt loam (USA) | Clay 23.4–26.2%, pH 6.7–7.3, OC 19.3–26.0 g/kg | 25°C, 35% WHC, 63 days | Increase of AWCD, richness, and diversity at 50 mg/kg on day 7 | Unger et al., 2013 | |
| 1–30 | Alfisol (China) | Sand 7.7%, silt 77.5%, clay 14.8%, pH 6.24, OM 2.4%, CEC 12.3 cmol/kg, OTC 37.3 μg/kg | 25°C, 60% WHC, 120 days | Negative effect on soil microbial community metabolism, but not on functional diversity indices | Ma et al., 2016 | |
| Sulfadimethoxine | 0.025–200 | Silt loam (USA) | pH 6.5, OM 2.2%, WHC 0.313 g/g, CEC 10.6 cmol+/kg | 80% WHC, manure addition, 50 days | CLPP parameters, including AWCD, Shannon diversity index, and evenness were not affected | Toth et al., 2011 |
| Sulfadiazine | 10 and 100 | Luvisol (China) | Sand 58.4%, silt 21.7%, clay 19.9%, pH 6.24, OM 3.56%, CEC 5.38 cmol/kg | 25°C, 25% WHC, manure addition at 40 mg/kg, 28 days | High dosage decreased the utilization rates of four categories of substrates (carbohydrate, carboxylic acid, amino acid, and aromatic acid) and the values of Shannon index during the experiment, while no significant inhibition was found at lower dosage on day 28 | Xu et al., 2016 |
| Sulfamonomethoxine | 100 | Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | Negative effect on functional diversity; more than half of the CLPP substrates could not be utilized effectively in soils | Ma et al., 2014 |
| Sulfamethoxazole | 20 and 500 | Loamy sand (Netherlands) | Sand 78.9%, silt 10.4%, clay 7%, pH 4.9, OC 3.7% | 25°C, 35% WHC, 5 weeks | Negative effect on CLPP at the highest concentration | Demoling et al., 2009 |
| 10 | Silty clay loam soil (China) | Sand 9%, silt 63.7%, clay 27.3%, pH 7, OM 1% | 25°C, 50–65% WHC, manure addition at 10 g/kg, 56 days | An inhibitory effect on the microbial diversity | Wang et al., 2016 | |
| 100 and 1,000 | No data (Spain) | Sand 37.3%, silt 24.7%, clay 38.0%, pH 7.9, OM 3.9% | 22–25°C, 70% WHC, 14 days | Decrease in AWCD and physiological diversity; changes in pattern of substrate utilization (decrease in all substrates at both concentrations, except polymers, and amino acids) | Pino-Otín et al., 2017 | |
| 1–100 | Clay loam (China) | Sand 30.4%, silt 34.1%, clay 35.5%, pH 5.7, OC 18.2 g/kg, CEC 9.87 cmol/kg | 25°C, 60% WHC, 21 days | Decrease in functional diversity after 7 days; 100 mg/kg decreased AWCD and Shannon indices; enhanced soil microbial community function on day 21 | Liu et al., 2012a | |
| Sulfamethazine | 53.6 | Sandy loam (Italy) | Sand 72.7%, silt 10.6%, clay 16.6%, pH 7.8, OC 2.8% | 7 days | AWCD decreased after 1 day and increased after 7 days; richness increased during the experiment | Pinna et al., 2012 |
| Sand 81.7%, silt 5.9%, clay 12.2%, pH 5.3, OC 1.7% | AWCD decreased after 1 day and increased after 7 days; richness decreased during the experiment | |||||
| Tetracycline | 100 and 500 | Clay (Italy) | Sand 39.4%, silt 19.2%, clay 41.4%, pH 5.8, OM 6.9% | 20°C, 60 days | Microbial communities in both soils were affected in the short term 500 mg/kg soil | Chessa et al., 2016b |
| Sand (Italy) | Sand 72.7%, silt 10.6%, clay 16.6%, pH 7.6, OM 4.9% | |||||
| 100 | Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | Functional diversity decreased | Ma et al., 2014 | |
| 100 and 1,000 | No data (Spain) | Sand 37.3%, silt 24.7%, clay 38.0%, pH 7.9, OM 3.9% | 22–25oC, 70% WHC, 14 days | Decrease in physiological diversity; changes in pattern of substrate utilization (decrease in carboxylic and ketonic acids and increase in amines/amides at 100 mg/kg | Pino-Otín et al., 2017 | |
| Trimethoprim | Decrease in physiological diversity; changes in pattern of substrate utilization (decrease in carboxylic and ketonic acids and increase in polymers at 100 mg/kg) | |||||
| Tylosin | 2,000 | Sand (Denmark) | Sand 90.7%, silt 2.8%, clay 4.1%, pH 6.8, WHC 15%, OC 1.2% | 25°C, 15% WHC, 60 days | No differences in the number of substrates utilized | Westergaard et al., 2001 |
| No differences in the functional diversity patterns | Müller et al., 2002 | |||||
| Vancomycin | 1 and 10 | Sandy loam (Poland) | Sand 67%, silt 24%, clay 9%, pH 6.9, WHC 43%, OC 1.6%, CEC 10 cmol/kg | 22°C, 50% WHC, 90 days | A negative impact on days 1, 15, and 30 as was showed by a decrease in the values of the CLPP indices (10–69%) | Cycoń et al., 2018 |
Effects of the selected antibiotics on the community-level physiological profile (CLPP) in soils with different characteristics.
AWCD, average well-color development; CEC, cation exchange capacity; CLPP, community-level physiological profile; OC, organic carbon; OM, organic matter; WHC, water holding capacity.
CLPP patterns do not always show changes in the functional diversity of microorganisms in response to antibiotic. In such situations, the lack of observable effects in CLPP might be because indices expressing total microbial activity do not detect possible changes caused by low antibiotic concentrations. Moreover, repeated application of antibiotics leads to the adaptation of bacteria to these compounds and proliferation of antibiotic-resistant microorganisms. Another limitation is that the Biolog technique measures only the activity of catabolically active cells and omits non-culturable populations as well as microorganisms in a dormant state. Moreover, fast-growing microorganisms are mainly responsible for effects observed by this test (Floch et al., 2011).
Impact of Antibiotics on Microbial Community Structure
PLFA Fingerprinting
The PLFA method is a rapid tool for assessing the biomass and composition of microbial communities in soil, because various phospholipid-derived fatty acids (PLFAs) are indicative of species or microbial groups in soil. In addition, the ratios of biomass of bacteria/fungi and Gram-positive/Gram-negative bacteria are often used to evaluate the response of microorganisms to organic pollutants (Frostegård et al., 1993, 2011).
In general, application of antibiotics into soil show strong and dose-dependent effects on the microbial structure expressed as the total PLFA biomass (Table 7). For example, Chen et al. (2013) found a significant increase in bacterial and fungal PLFA biomass 7 weeks after oxytetracycline contamination of 1 and 15 mg/kg soil, but a strong decrease in the case of 200 mg antibiotic/kg treatment. A decrease in microbial PLFA biomass content was also found in soils that had been treated with penicillin G (10 and 100 mg/kg soil) and tetracycline (100 mg/kg soil) (Zhang et al., 2016a). Simultaneously, the ratios of Gram-negative/Gram-positive bacteria increased, indicating that the bacteria that were resistant to antibiotics in tested soils were more likely to be Gram-negative. A study by Unger et al. (2013) found only short-term effects (35 days) from lincomycin or oxytetracycline applications (5–200 mg/kg soil), suggesting that the resilience of soil microbial communities toward effects of these antibiotics.
Table 7
| Antibiotic | Dosage (mg/kg soil) | Type of soil/origin | Main properties of soil | Experimental conditions | Effect (comparison with control, non-treated soil) | References |
|---|---|---|---|---|---|---|
| Ciprofoxacin | 1, 5, and 50 | Ustic Cambisol (China) | Sand 12%, silt 54%, clay 34%, pH 7.9, OC 36.76 g/kg, CEC 13.82 cmol/kg | 25°C, 60% WHC, 40 days | Decrease in the ratio of bacteria to fungi and increased in the ratio of Gram+ to Gram-bacteria, PCA of the PLFA data clearly distinguished among different concentrations | Cui et al., 2014 |
| Difloxacin | 1–100 | Loamy sand (Germany) | Sand 73.3%, silt 23.1%, clay 3.6%, pH 5.5, OC 1.7%, WHC 27% | 10°C, 60% WHC, manure addition at 40 ml/kg, 32 days | Impact on the total PLFAs only on days 1 and 8; a significant decrease in the ratio of bacteria/fungi on day 8 at 100 mg/kg; a significant decrease in the ratio of Gram+/Gram– on day 1 after application regardless of the concentration | Kotzerke et al., 2011 |
| 0.452 | Luvisol (Germany) | Sand 6%, silt 78%, clay 16%, pH 6.3, OC 1.2%, CEC 11.4 cmol/kg | 21°C, 63 days, slurry addition at 40 ml/kg | The total PLFAs increased on days 7 and 14; temporal shifts to the Gram– bacteria indicated by decrease in the ratio of Gram+/Gram– on days 14 and 63; shifts to fungi shown by lower ration of bacteria/fungi on days 42 | Reichel et al., 2013 | |
| Lincomycin | 5–200 | Silt loam (USA) | Clay 23.4–26.2%, pH 6.7–7.3, OC 19.3–26.0 g/kg | 25°C, 35% WHC, 63 days | All PLFA markers declined between 0 and 3 days; decline in total biomass, Gram+, Gram–, anaerobic bacteria, fungi, and mycorrhizae between 3 and 35 days; increase in the total biomass, bacteria/fungi ratio, Gram–, and anaerobic bacteria and protozoa between 35 and 63 days | Unger et al., 2013 |
| Oxytetracycline | 1–200 | Loam (China) | Sand 52.2%, silt 38.6%, clay 9.2%, pH 8.3, OM 1.2% | 20–27°C (day) and 15–20°C (night), 50–70% WHC, manure addition at 30 mg/g, 7 weeks | The total PLFAs increased at 1 at 15 mg/kg and decreased at 200 mg/kg; fungal PLFAs were significantly lower at 200 mg/kg; the ratios of Gram–/Gram+ at 1 and 15 mg/kg were significantly higher while at 200 mg/kg was significantly lower | Chen et al., 2013 |
| 5–200 | Silt loam (USA) | Clay 23.4–26.2%, pH 6.7–7.3, OC 19.3–26.0 g/kg | 25°C, 35% WHC, 63 days | All PLFA markers declined between 0 and 3 days; additional declines in biomass, Gram+, Gram–, anaerobic bacteria, fungi, and mycorrhizae between 3 and 35 days; decline in Gram+, fungi, and mycorrhizae and increase in biomass, ratio of bacteria/fungi, Gram–, and anaerobic bacteria and protozoa between 35 and 63 days | Unger et al., 2013 | |
| Sulfadimethoxine+ Sulfamethoxazole+ Sulfamethazine |
0.09–900 | Sandy loam (USA) | Sand 85.5%, silt 8.5%, clay 6.6%, pH 6.31, OC 0.86%, CEC 8.1 cmol+/kg | 20°C, 21 days, glucose and/or manure addition | A relative community shift toward Gram- bacteria and increase in fungal biomass | Gutiérrez et al., 2010 |
| Sulfadiazine | Luvisol (China) | Sand 58.4%, silt 21.7%, clay 19.9%, pH 6.24, OM 3.56%, CEC 5.38 cmol/kg | 25°C, 25% WHC, manure addition at 40 mg/kg, 28 days | The total PLFA, bacterial, and actinomycetes biomass was reduced; no significant effect on the ratio of Gram+/Gram– bacteria; decrease in the ratio of bacteria/fungi | Xu et al., 2016 | |
| 10 and 100 | Loamy sand (Germany) | Sand 73.3%, silt 23.1%, clay 3.6%, pH 4.8, OC 1% | 10°C, 50% WHC, manure addition at 20, 40, and 80 g/kg, 32 days | Decrease in the total PLFAs on days 1 and 8 and on day 32 for antibiotic in combination with high and low manure amendment; decrease in the ratio of bacteria/fungi for SDZ with low manure amendment on day 32; no effect on the ration of Gram+/Gram– bacteria | Hammesfahr et al., 2011 | |
| 10°C, 50% WHC, manure addition at 40 mg/g, 61 days | Decrease of the effect of manure on the total of PLFAs at all day | Hammesfahr et al., 2008 | ||||
| Silt loam (Germany) | Sand 6.4%, silt 78.2%, clay 15.4%, pH 7.2, OC 2.1% | SDZ decreased the effect of manure on the total of PLFAs at all days | ||||
| 0.256 | Silt loam (Germany) | Sand 6%, silt 78%, clay 16%, pH 6.3, OC 1.2%, CEC 11.4 cmol/kg | 21°C, 63 days, slurry addition at 40 ml/kg | The total PLFAs increased on day 63; temporal shifts to the Gram– bacteria shown by the decrease in the ratio of Gram+/Gram- on days 14 and 63; shifts to fungi characterized by lower ratio of bacteria/fungi on day 14 | Reichel et al., 2013 | |
| 1 and 10 | 21°C, 63 days, manure addition | Decrease in the total PLFAs by 14% in the rhizosphere and 3% in bulk soil in the field experiment | Reichel et al., 2014 | |||
| 20 and 500 | Loamy sand (Netherlands) | Sand 78.9%, silt 10.4%, clay 7%, pH 4.9, OC 3.7% | 25°C, 35% WHC, 5 weeks | No differences in the microbial community structure | Demoling et al., 2009 | |
| Silt loam (New Zeland) | Sand 9%, silt 54%, clay 37%, pH 6.7, OC 5% | |||||
| Sulfamethoxazole | 5 | Clay loam (New Zeland) | Sand 13.7%, silt 51%, clay 30.4%, pH 5.8, OC 4% | 25°C, 60% WHC, 36 days | A higher proportion of bacterial biomass over fungal biomass (fungal to bacterial ratio < 1) for each sampling event | Srinivasan and Sarmah, 2014 |
| Silt loam (New Zeland) | Sand 34%, silt 48%, clay 17%, pH 5.7, OC 8.2% | |||||
| Tetracycline | 100 and 500 | Clay (Italy) | Sand 39.4%, silt 19.2%, clay 41.4%, pH 5.8, OM 6.9% | 20°C, 60 days | Short-term negative effect at a higher concentration; a significant increase in the ratio of fungi: bacteria in both soil, and Gram+/Gram– in clay soil | Chessa et al., 2016b |
| Sand (Italy) | Sand 72.7%, silt 10.6%, clay 16.6%, pH 7.6, OM 4.9% | |||||
| Vancomycin | 1 and 10 | Sandy loam (Poland) | Sand 67%, silt 24%, clay 9%, pH 6.9, WHC 43%, OC 1.6%, CEC 10 cmol/kg | 22°C, 50% WHC, 90 days | A temporal shift and dominance of Gram-; a decrease in the ratio of Gram+/Gram–; increase in fungal biomass as reflected by decreased ration of bacteria/fungi | Cycoń et al., 2016a |
Effects of the selected antibiotics on the community structure of microorganisms based on the PLFA analysis in soils with different characteristics.
OC, organic carbon; OM, organic matter; PLFA, phospholipid fatty acid; WHC, water holding capacity.
An increase in total PLFA biomass was also observed in soils treated with sulfadiazine and difloxacin in slurry from pigs medicated with these antibiotics (Reichel et al., 2013). Influences from the difloxacin-slurry on PLFA biomass were observed at the beginning of the experiment (7 and 14 days), but only on the last sampling day (63 day) in the sulfadiazine-slurry. The authors hypothesized that the delayed effect of sulfadiazine was a result of continuous remobilization of antibiotic residues bound to soil particles by microorganisms. Hammesfahr et al. (2008) and Kotzerke et al. (2011), in contrast, found no changes in soil microbial communities upon addition of the above-mentioned antibiotics into soil spiked with slurry.
Several studies showed changes in the soil fungal PLFA signatures upon amendment with antibiotics. For example, the content of fungal marker fatty acids (18:1ω9 and 18:2ω6,9) significantly increased in alfalfa-amended soil and both alfalfa- and manure-amended soils with 20 and 500 mg sulfamethoxazole /kg of soil (Demoling et al., 2009). Gutiérrez et al. (2010) mixed three sulfonamides, sulfadimethoxine, sulfamethoxazole, and sulfamethazine, and observed a general increase in the proportion of fungal PLFAs among the total soil biomass PLFAs. A shift in the soil microbial community toward more fungi because of sulfapyridine application was also noted by Thiele-Bruhn and Beck (2005). The increase in the abundance of fungi may ultimately result in a decline in the productivity and quality of soils and agricultural products (Ding and He, 2010).
The PLFA method, previously used extensively for determination of microbial community composition, has been largely replaced by techniques based on the analysis of nucleic acids extracted from soil. However, PLFA-based analyses remain an adequately sensitive, efficient way to rapidly screen whether a microbial community has been affected by antibiotics.
Phylogenetic Analyses
Many authors have used PCR-dependent DNA fingerprinting techniques such as a terminal restriction fragment length polymorphism (T-RFLP) or denaturing gradient gel electrophoresis (DGGE) to evaluate the impact of antibiotics on diversity of soil microorganisms (Müller et al., 2002; Reichel et al., 2014; Chessa et al., 2016a; Orlewska et al., 2018a,b). Most of these studies target 16S rRNA genes and show altered diversity upon antibiotics applications. These changes may be explained by the disappearance of sensitive bacteria and outgrowth of resistant bacteria present in the antibiotic-contaminated soils (Table 8). For example, the application of tylosin (2 mg/kg soil) strongly modified the DGGE pattern, reducing the number of bands observed on days 15 and 22 of the experiment compared to non-exposed soil. However, on day 33 the difference was smaller, yet detectable (Westergaard et al., 2001). In a study by Müller et al. (2002), the DGGE pattern of 16S rDNA in tylosin-treated soil differed slightly to the diversity to the control soil, although colony morphology typing and potential of microbial communities to utilize selected substrates did not reveal any differences. Cycoń et al. (2016a) found that vancomycin dosed to soil at 10 mg/kg resulted in quantitative changes in microbial community patterns, suggesting selection exerted by the antibiotic. The changes in DGGE patterns suggest a decrease in populations of a species or group of species; revealing that some bacterial species were more sensitive to vancomycin than others were. This was corroborated by a decline in the values of H' and S (richness) indices. Another study showed significant effects of chlortetracycline on the microbial community that lasted for no longer than 1 week after the application of the antibiotic as the communities recovered over time until the end of the experiment (42 day) (Nelson et al., 2011). Contrarily, using PCR-DGGE Zielezny et al. (2006) showed that the application of sulfadiazine and chlortetracycline at concentrations of 1, 10, and 50 mg/kg soil had no significant effect on the structure of the soil microbial community. In another study, changes in DGGE profiles showed that sulfadiazine (10 and 100 μg/g soil) applied to soil in manure altered bacterial diversity (Hammesfahr et al., 2008). Although the DGGE profiles proved the impact of manure and sulfadiazine on the total soil bacterial communities, these effects were less distinct for pseudomonads and β-Proteobacteria. This observation may be explained by the resistance of many strains to sulfonamides, which may be only indirectly affected by sulfadiazine. Genetic changes within the β-Proteobacteria and the pseudomonads, expressed by the appearance or loss of a band in the DGGE profiles from manure, and sulfadiazine-amended soils were also reported by Reichel et al. (2014). However, statistical analysis showed that the β-Proteobacteria significantly responded to the time and moisture regimes, whereas the pseudomonads responded to the factors of time and treatment. The effects of sulfadiazine-containing manure on microbial diversity resulted in a low stability of soil bacterial communities and dominance of taxa that are known to contain human pathogens, such as Gemmatimonas, Leifsonia, Devosia, Clostridium, Shinella, and Peptostreptococcus (Ding et al., 2014). No apparent effects on microbial diversity in response to streptomycin and difloxacin were observed in similar studies (Shade et al., 2013; Jechalke et al., 2014a).
Table 8
| Antibiotic | Dosage (mg/kg soil) | Type of soil/origin | Main properties of soil | Experimental conditions | Effect (comparison with control, non-treated soil) | References |
|---|---|---|---|---|---|---|
| Amoxycillin | 10 and 100 | Silt loam (Germany) | pH 7.2, WHC 46%, OC 2.1% | 10°C, 40% WHC, manure addition at 40 ml/kg, 9 days | Transient modification in the composition of bacterial community revealed by DGGE patterns | Binh et al., 2007 |
| Loamy sand (Germany) | pH 5.5, WHC 27%, OC 1.7% | |||||
| Cefuroxime | 1 and 10 | Sandy loam (Poland) | Sand 67%, silt 24%, clay 9%, pH 6.9, WHC 43%, OC 1.6%, CEC 10 cmol/kg | 22°C, 50% WHC, 90 days | Alteration in the DGGE profiles and decline in the DGGE indices (richness and diversity) at 10 mg/kg soil, in particular on days 30, 60, and 90 | Orlewska et al., 2018c |
| Ciprofloxacin | 100 | Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | Lower abundance of six nitrogen-cycling genes including chiA, amoA, nifH, nirK, nirS, and narG in the treated soil | Ma et al., 2014 |
| 0.2, 2, and 20 | Agricultural soil (Germany) | Sand 11%, silt 68%, clay 21%, pH 6.6, WHC 37.5%, OC 2.1% | 20°C, 60% WHC, 93 days, sludge addition at 1.8 g/kg | Shift in both microbial abundance and microbial diversity based on the T-RFLP analysis | Girardi et al., 2011 | |
| Chlortetracycline | 1–50 | Orthic Luvisol (Germany) | Sand 3%, silt 79%, clay 18%, pH 7, OC 1.04%, WHC 48.8% | 20°C, 48 days | No detectable impact on the community structure showed by DGGE pattern | Zielezny et al., 2006 |
| Difloxacin | 0.452 | Luvisol (Germany) | Sand 6%, silt 78%, clay 16%, pH 6.3, OC 1.2%, CEC 11.4 cmol/kg | 21°C, 63 days, slurry addition at 40 ml/kg | Changes in DGGE profiles; additionally, clusters of the antibiotic-slurry treatment were less strongly shifted by time | Reichel et al., 2013 |
| 5 | Silt loam (Germany) | Sand 6.4%, silt 78.2%, clay 15.4%, pH 7.2, OC 2.1%, WHC 46% | Field experiment, 140 days | Significant effect of antibiotic- manure on the bacterial community composition was revealed by DGGE; quinolone resistance genes qnrB and qnrS1/qnrS2 were detected by PCR and subsequent hybridization, while qnrA was not detected; PCA of DGGE profiles suggested that effect of manure lasted till day 28; samples on days 71 and 140 were closed to untreated soil | Jechalke et al., 2014b | |
| Erythromycin | 1 and 10 | Sandy loam (Poland) | Sand 67%, silt 24%, clay 9%, pH 6.9, WHC 43%, OC 1.6%, CEC 10 cmol/kg | 22°C, 50% WHC, 90 days | Alteration in the DGGE profiles and decline in the DGGE indices (richness and diversity) at 10 mg/kg soil, in particular on days 15, 30, and 60 | Orlewska et al., 2018b |
| Lincomycin | 0.05 and 5 | Cambisol (Czech Republic) | Sand 17%, silt 57%, clay 26%, pH 7.65, OM 8.17% | 16/6°C, 40 days | Dosage of 5 mg/kg shifted bacterial diversity (16S rRNA gene—T-RFLP) in the soil with high pH, while in the soil with low pH higher percentage of actinomycetes and higher diversity of the lmrB homolog genes did not change; sequencing of 157 new clones of lmrB homologs revealed selection of lmrB homologs in both soils | Cermák et al., 2008 |
| Podzol (Czech Republic) | Sand 96%, silt 2%, clay 2%, pH 4.01, OM 9.08% | |||||
| Sulfachloropyridiazine | 1, 10, and 100 | Silt Loam (US) | Sand 19.9%, silt 56.6%, clay 23.6%, pH 7.5, OC 1.8 | 25°C, 40 days | No significant differences in banding patterns (Dice similarity coefficients above 0.9) | Accinelli et al., 2007 |
| Sand (US) | Sand 93.5%, silt 2.7%, clay 3.8%, pH 7.2, OC 0.94% | |||||
| Sulfadiazine | 20 | Silt loam (Germany) | Sand 6.4%, silt 78.2%, clay 15.4%, pH 7.2, OC 2.1%, WHC 46% | 20°C, 30 days, 50% WHC, manure addition | Contamination of the manure with antibiotic significantly reduced nifH, ammonia-oxidizing bacteria (AOB) amoA, nirK, nirS, and nosZ gene abundance patterns 20 days after application, however ammonia-oxidizing archaea (AOA), amoA gene abundance were not influenced at any sampling day | Ollivier et al., 2010 |
| 10 and 100 | Silt loam (Germany) | Sand 6.4%, silt 78.2%, clay 15.4%, pH 7.2, OC 2.1%, WHC 46% | 20°C, 61 days, manure addition at 40 g/kg soil | High prevalence of sul1 in manure and manured soils, sul2 was mainly found in the loamy sand treated with manure and high dose and sul3 was not detected; addition of manure increased nirK-harboring denitrifiers in both soils, whereas in the treatments the abundance of the nirS denitrifiers increased after the bioavailable antibiotic had declined; however, the community composition of nirS nitrite reducers investigated by DGGE did not change despite the observed alterations in abundance. | Heuer and Smalla, 2007; Schauss et al., 2009; Kleineidam et al., 2010; Heuer et al., 2011b | |
| Loamy sand (Germany) | Sand 73.3%, silt 23.1%, clay 3.6%, pH 5.5, OC 1.7%, WHC 27% | |||||
| 1–50 | Orthic Luvisol (Germany) | Sand 3%, silt 79%, clay 18%, pH 7, OC 1.04%, WHC 48.8% | 20°C, 48 days | Modification in the composition of a bacterial community shown by DGGE | Zielezny et al., 2006 | |
| 10 and 100 | Loamy sand (Germany) | Sand 73.3%, silt 23.1%, clay 3.6%, pH 5.5, OC 1% | 10°C, 50% WHC, manure addition at 40 mg/g, 61 days | Increase in the band intensity in DGGE profiles in manure treatment, while manure, and antibiotic had a more differentiated effect on bacterial populations; changes in DGGE profiles were seen even after two month of the experiment | Hammesfahr et al., 2008 | |
| Silt loam (Germany) | ||||||
| 10 and 100 | Silt loam (China) | No data | 15°C, 55% WHC, manure addition at 40 g/kg, 193 days | Repeated application at a higher concentration of 100 mg/kg soil caused visible changes in the composition of a bacterial community | Ding et al., 2014 | |
| 0.256 | Silt loam (Germany) | Sand 6%, silt 78%, clay 16%, pH 6.3, OC 1.2%, CEC 11.4 cmol/kg | 21°C, 63 days, slurry addition at 40 ml/kg | Changes in the total DGGE band profiles; the samples of control- and antibiotic-slurry treatment clustered separately mostly by time | Reichel et al., 2013 | |
| 1 and 10 | 21°C, 63 days, manure addition | DGGE pattern indicated larger structural shifts within genus of Pseudomonas in SDZ-contaminated earthworm burrows compared to bulk soils | Reichel et al., 2014 | |||
| Sulfamonmethoxine | 100 | Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | Lower abundances of six nitrogen-cycling genes including chiA, amoA, nifH, nirK, nirS, and narG were observed | Ma et al., 2014 |
| Sulfamethazine | 1, 10, and 100 | Silt Loam (US) | Sand 19.9%, silt 56.6%, clay 23.6%, pH 7.5, OC 1.8 | 25°C, 40 days | No significant t differences in banding patterns (Diece similarity coefficients above 0.9) | Accinelli et al., 2007 |
| Sand (US) | Sand 93.5%, silt 2.7%, clay 3.8%, pH 7.2, OC 0.94% | |||||
| Tetracycline | 100 | Loam (China) | Sand 42%, silt 38%, clay 20%, pH 6.31, OM 1.29%, WHC 35% | 25°C, 50% WHC, 20 days | Lower abundances of six nitrogen-cycling genes including chiA, amoA, nifH, nirK, nirS, and narG were revealed | Ma et al., 2014 |
| Tylosin | 2,000 | Sand (Denmark) | Sand 90.7%, silt 2.8%, clay 5.1%, pH 6.8, WHC 15%, OC 1.2% | 25°C, 15% WHC, 60 days | Decrease in the number of bands in DGGE profiles was detected on days 15 and 22; difference was smaller but still detectable on day 33 and at the end of experiment | Westergaard et al., 2001 |
| Small difference in the microbial diversity shown by DGGE | Müller et al., 2002 | |||||
| Vancomycin | 1 and 10 | Sandy loam (Poland) | Sand 67%, silt 24%, clay 9%, pH 6.9, WHC 43%, OC 1.6%, CEC 10 cmol/kg | 22°C, 50% WHC, 90 days | Disappearance of some bands in DGGE profiles; decrease of the overall richness and diversity of the dominant bacteria at 10 mg/kg during the 90-day experiment | Cycoń et al., 2016a |
Effects of the selected antibiotics on the community structure of microorganisms based on the genetic analyses in soils with different characteristics.
DGGE, denaturing gradient gel electrophoresis; OC, organic carbon; OM, organic matter; T-RFLP, terminal restriction fragment length polymorphisms; WHC, water holding capacity.
Diversity of Antimicrobial Resistance Genes
Studies are increasingly focusing on genes for antimicrobial resistance itself, which can be followed by metagenomic sequencing approaches. Resistance determinants present in the soil, defined as the soil resistome, include genes that confer antibiotic resistance to pathogenic and non-pathogenic species present in the environment, as well as proto-resistance genes that serve as a source of resistance elements (D'Costa et al., 2006, 2007; Perry et al., 2014). The potential transfer of antibiotic resistance among microbial populations in soil and the risk of animal and human infection are major concerns (Figure 5).
Figure 5
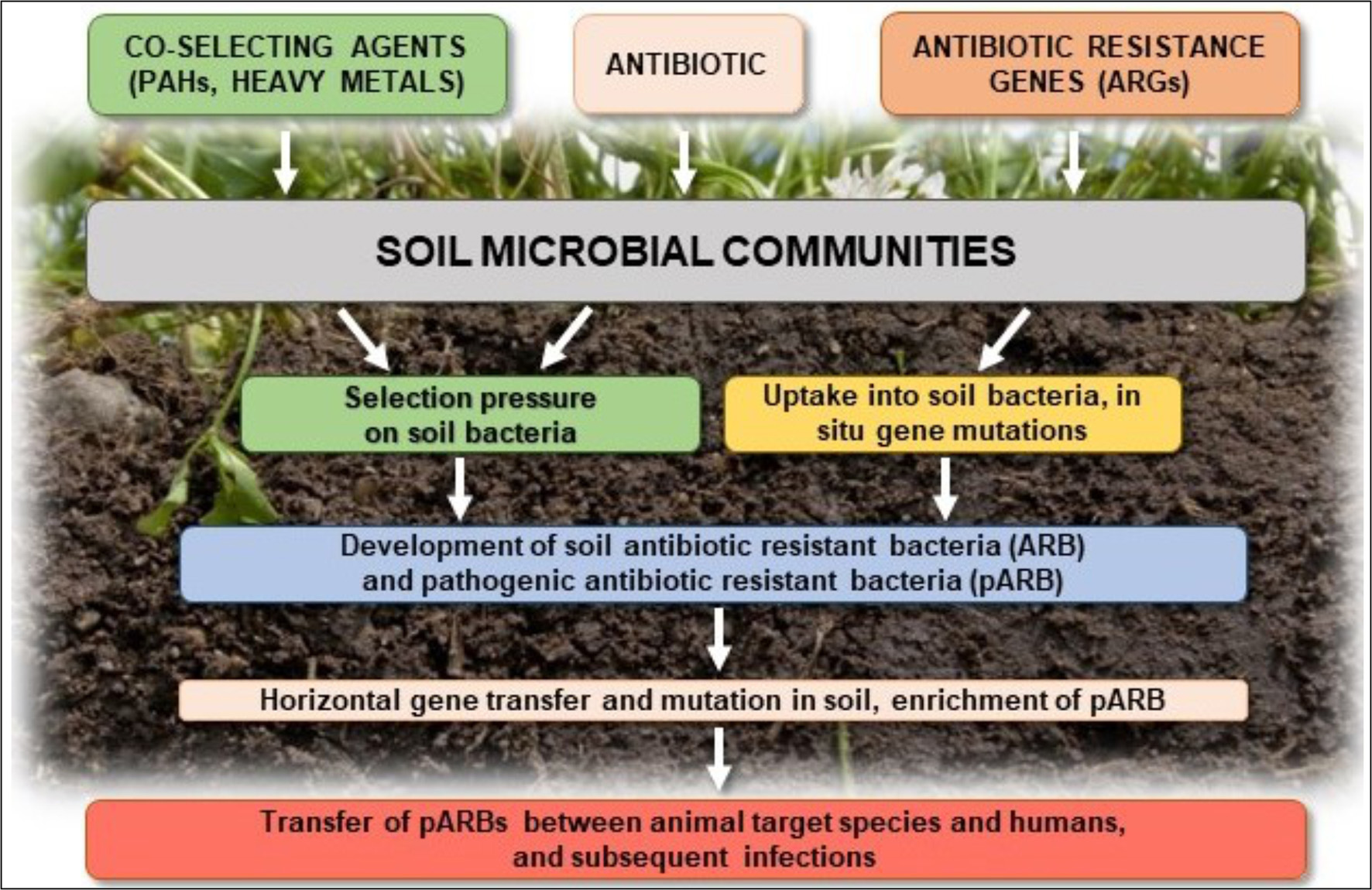
Transfer of antibiotic resistance in soil and risk of animal and human infection (based on Ashbolt et al., 2013).
Special attention is given to the detection and quantitation of ARGs, identifying resistance mechanisms, discovery of novel enzymes with unexpected activities, and the impact of pollutants and agricultural practices on the abundance of ARGs in soils. Resistance to antibiotics has been documented even in bacteria inhabiting pristine soils never exposed to these compounds, for example genes conferring resistance to common antibiotics (β-lactams, fluoroquinolones, and tetracyclines) were found in the Tibet (Chen et al., 2016). Interestingly, a tiny fraction of ARGs located on MGEs has been discovered, indicating a low, but real, potential for these ARGs to be transferred among bacteria (Chen et al., 2016). Additionally, some ARGs that have been discovered encode efflux pumps that are completely different from previously known efflux pumps. A large number (177) of ARGs encoding mostly single or multi-drug efflux pumps conferring resistance to aminoglycosides, chloramphenicol and β-lactams were reported in pristine Antarctic soils (Van Goethem et al., 2018). A low number and diversity of ARGs was detected in glacial soil and permafrost as compared to environments highly impacted by human activities in a study by Zhang et al. (2018). It has been proposed that ARGs in pristine environments most likely represent functional historical genes conferring resistance to natural antibiotics. Antibiotic resistance appeared to be transferred vertically over generations with limited to no horizontal transfer of ARGs between species (Van Goethem et al., 2018).
Diversity and abundance of ARGs is strongly influenced by agricultural practices (Wepking et al., 2017; Xiong et al., 2018). It has been documented that the spreading of manure from animals treated with different antibiotics on fields affects the abundance and diversity of ARGs (Su et al., 2014; Kyselková et al., 2015). However, it has also been documented that organic matter content, pH, and the history of soil management are important factors influencing the fate and abundance of ARGs in the soil (Cermák et al., 2008; Popowska et al., 2012).
The genes conferring resistance to minocycline, tetracycline, streptomycin, gentamycin, kanamycin, amikacin, chloramphenicol, and rifampicin in soils subjected to manure application accounted for ~70% of the total ARGs. More than half of the ARGs shared low similarity, < 60%, with the sequences of their closest proteins in GeneBank (Su et al., 2014). Among the ARGs studied (ampC, tetO, tetW, and ermB), the average abundance of ampC and tetO in the manure-treated soil was 421 and 3.3% greater in comparison with a non-treated control soil, respectively (Wepking et al., 2017). A broad-spectrum profile of ARGs has been found in 5 paddy soils of South China (Xiao et al., 2016). Based on Antibiotic Resistance Genes Database and next generation sequencing, 16 types of ARGs corresponding to 110 ARG subtypes were identified. Multi-drug resistance genes dominated in all soils (up to 47.5%), followed mainly by specific resistance genes for acriflavine, macrolide-lincosamide-streptogramin and bacitracin. Efflux pumps, antibiotic deactivation, and cellular protection were three major resistance mechanisms inferred from the resistome characterized Xiao et al. (2016).
An increase in the number and proportions of ARGs in the bacterial community was also shown by Nõlvak et al. (2016), who studied the effect of animal manure on the rate of the dissemination of ARGs (sul1, tetA, blaCTX-M, blaOXA2, and qnrS) and integron-integrase genes in soil. The abundance of genes for tetracycline resistance (tetM, tetO, and tetW) in soil that had had manure applied was significantly higher than in untreated soil (Xiong et al., 2018). It seems that an increasing abundance of ARGs resulted from inputs of ARGs already present in manure. This is because antibiotics may provide a significant selective advantage for some species or groups of antibiotic-resistant bacteria that then come to dominate the microbial community.
By contrast, many studies have shown that the dissemination of ARGs in soils is more correlated with the application of manure than with the presence of antibiotics. Though no effect on the abundance of tetG, tetO, and tetW was observed in one study, despite the presence of these genes in manure applied to the soil (Fahrenfeld et al., 2013), other studies have found an impact of manure from never antibiotic-treated animals on the number, diversity, and spread potential of ARGs (Udikovic-Kolic et al., 2014; Hu et al., 2016). This phenomenon can be explained by the stimulation of growth autochthonous bacteria and accelerated spread of ARGs by fertilizers free from antibiotics. Due to the ambiguous results of studies concerning the spread of ARGs in manure-treated soil, there is a need for future research to clearly assess the relative contribution of manure and antibiotics on microbial community structure and processes mediated by soil microorganisms.
The studies on the fate of ARGs in soil allowed not only to assess the diversity and abundance of genes of interest, but also to discover new ARGs, and described new resistance mechanisms and novel enzymes responsible for resistance of bacteria to antibiotics (Lau et al., 2017; Wang et al., 2017). Functional metagenomics for the investigation of antibiotic resistance led to the discovery of several tetracycline resistance genes from different classes in soil. Efflux pump genes, including 21 major facilitator superfamily efflux pump genes, were found to dominate the sequence libraries derived from studied soils. In addition, two new genes involved in the enzymatic inactivation of tetracycline were identified (Wang et al., 2017). Similarly, Torres-Cortés et al. (2011) described a new type of dihydrofolate reductase (protein Tm8-3; 26.8 kDa) conferring resistance to trimethoprim in soil microbiome. Using functional metagenomics they also identified three new antibiotic resistance genes conferring resistance to ampicillin, two to gentamicin, two to chloramphenicol, and four to trimethoprim in libraries generated from three different soil samples. Nine carbapenem-hydrolizing metallo-beta-lactamases (MBLs) including seven novel enzymes showing 33–59% similarity to previously described MBLs were discovered. Six originated from Proteobacteria, two from Bacteroidetes, and one from Gemmatimonadetes. MBLs were detected more frequently and exhibited higher diversity in grassland than in agricultural soil (Gudeta et al., 2016). In a study of resistome of bacterial communities in Canadian soils by functional metagenomics, 34 new ARGs with high homology to determinants conferring resistance aminoglycosides, sulfonamides and broad range of β-lactams were identified (Lau et al., 2017). High-resolution proteomics in combination with functional metagenomics resulted in the discovery of a new proline-rich peptide PPPAZI promoting resistance to various macrolides, but not to other ribosome-targeting antibiotics (Lau et al., 2017). Similarly, Donato et al. (2010) identified two novel bifunctional proteins responsible for bacteria resistance to ceftazidime and kanamycin, respectively.
Other synthetic pollutants can also influence the acquisition and maintenance of ARGs that can confer resistance not only to antibiotics but also to a number of structurally unrelated contaminants present in soil. As an example for this, ARGs were ~15 times more abundant in polycyclic aromatic hydrocarbon (PAH) contaminated soils than in similar non-contaminated soils (Chen et al., 2017). It resulted from the selection by PAHs of Proteobacteria containing ARGs encoded multi-drug efflux pumps leading to simultaneously enriching of ARGs carried by them in the soils. Moreover, most of ARGs (70%) found in the PAHs-contaminated soils were not carried by plasmids, indicating a low possibility of horizontal gene transfer (HGT) between bacteria (Chen et al., 2017).
It has recently been reported that heavy metals are also as a factor contributing to maintenance of ARGs in the environment. A study of dairy farms showed significant correlations between the abundance of ARGs, metal resistance genes and the content of Cu and Zn in cattle feces (Zhou et al., 2016). The role of heavy metals can be explained by cross-resistance, tolerance of bacteria to more than one antimicrobial agent (Chapman, 2003; Baker-Austin et al., 2006), and co-resistance, the localization of ARGs and genes encoding resistance to heavy metals and other antibacterial agents on mobile genetic elements (Chapman, 2003), mechanisms. Multi-drug efflux pumps, which mediate rapid extrusion of antibiotics and heavy metals from the cell decrease susceptibility toward these compounds, are an example of cross-resistance (Martinez et al., 2009). Co-resistance can occur due to the close arrangement of genes on a chromosome or extrachromosomal element, which increases the likelihood that genes are subject to combined transmission via HGT. Moreover, heavy metals not only trigger co-selection processes, but also increase the level of tolerance to antibiotics due to co-regulation of resistance genes (Baker-Austin et al., 2006). Such mechanisms may affect the spread of ARGs from environmental populations to bacteria of clinical importance even in the absence of direct antibiotics selection (Herrick et al., 2014).
Antibiotics concentrations in the environment are much lower than that used in therapeutic doses. However, even at sub-inhibitory concentrations (below the MIC), antibiotics may select for resistant phenotypes (Gullberg et al., 2011, 2014). It has been reported that resistant bacteria can be selected at antibiotic concentrations even 100-fold below lethal concentrations. Using data on MICs from the EUCAST database for 111 antibiotics, Bengtsson-Palme and Larsson (2016) estimated that the predicted no-effect concentrations (PNECs) range from 8 ng/L to 64 μg/L. Mutants selected by antibiotics at minimum selective concentration appeared to be more fit than those selected at high concentration and still highly resistant (Gullberg et al., 2011; Sandegren, 2014). Antibiotics at such concentrations play multifaceted roles in the soil, including influencing inter-species competition, signal communication, host–parasite interactions, virulence modulation, and biofilm formation (Sengupta et al., 2013). Moreover, they induce mutations, genetic recombination, and HGT processes (Andersson and Hughes, 2012, 2014). Sub-lethal concentrations of drugs can also increase the mutation rate and potential for enrichment of stable genetic mutants by the induction of SOS responses, increasing translation misreading, and generation of oxygen radicals. It has been reported that sub-lethal concentrations of tetracycline stimulated up to 1,000-fold higher horizontal transfer rates of mobile genetic elements in Listeria monocytogenes (Bahlm et al., 2004) and integron recombination rates in Vibrio cholerae (Guerin et al., 2009). However, the rate of spread of ARGs depends on the antibiotic concentration in the soil. Huang et al. (2016) showed that the dissipation rate of plasmid-located genes [i.e., qnrS, oqxA, aac(6′)-Ib-cr] was significantly lower in soil treated with ciprofloxacin at concentrations of 0.04 and 0.4 mg/kg soil compared to soil treated with antibiotic at 4 mg/kg and controls. Forsberg et al. (2012) also found little evidence for HGT in soil.
Because of the risks posed by the dissemination of ARGs among bacteria and plants, studies investigating new practices designed to decrease the accumulation and transport of ARGs within soil are being conducted. For example, the use of biochar as a soil amendment significantly reduced the abundance of tetracyclines (tetC, tetG, tetW, and tetX) and sulfonamides (sulI and sulII) resistance genes in soil and lettuce tissues (Ye et al., 2016; Duan et al., 2017).
Conclusion
Environmental antibiotic pollution is a problem that is expected to gain more attention in the near future since antibiotic consumption is still increasing around the world. A review of available literature shows that transformation and/or degradation are the most important processes that determine the fate of antibiotics in soils and that soil microorganisms play an important role in these processes. However, the rate of these transformation and degradation depends largely on the antibiotic structure and is affected by many abiotic and biotic factors, as is most evident in the large range of DT50 values of antibiotics in different soils. Studies have shown that within groups of similar antibiotics or even particular antibiotics, DT50 values differ significantly because of various soil properties, antibiotic dosage, and environmental conditions.
Current literature reviewed here indicates that the input of antibiotics into soil alters the structure and activity of microbial communities and the abundance of ARGs. However, results from studies in this field are often ambiguous, making environmental risk assessments related to the presence of antibiotics depend upon too many different factors to be reliable. Interactions between soil, antibiotics, and microorganisms are multifarious and many environmental factors may influence the value of tested parameters. The effect of antibiotics on the activity and diversity of microbial communities depends on the physicochemical parameters of the soil, the antimicrobial activity, and dosage of the antibiotic, as well as the time of exposure. It has become clear that microorganisms that are sensitive to different antibiotics are killed or inhibited in the presence of antibiotics, which may result in outgrowth of resistant bacteria resistant. In turn, this may alter the diversity of the soil microbial communities. Contrarily, there is some evidence that certain microorganisms can adapt to and possibly transform antibiotics. The less toxic transformation products would favor recovery of the original microbial communities from the initial disturbances caused by antibiotics exposure. Several studies found transient negative effects of antibiotics on the functional, structural and genetic diversity of soil microbial communities; a temporary loss of soil functionality with subsequent recovery.
Existing OECD and ISO ecotoxicological tests used to evaluate the toxicity of chemicals on the activity of a single species (e.g., Microtox, MARA, measurements of selected enzyme activities) and some of the processes mediated by soil microorganisms are not sufficient to gauge the effects of antibiotics on soil microbial communities. Therefore, ecotoxicological tests that combine various microbial community parameters should be developed. Moreover, future studies should be based on “omics” approaches such as genomics, transcriptomics, and proteomics, that allow for deeper examination of microbial communities than CLPP, PLFA, and PCR-DGGE methods. “Omics” methods are better tools for tracking the fate and determining dissipation rate of ARGs, as well as for recognizing new mechanisms of antibiotic resistance. Future frameworks for antibiotics should focus on the effect of soil properties on the maintenance and development of antibiotic resistance, the fate of antibiotics and ARGs in manure-fertilized soils, the rate of HGT between antibiotic-resistant and autochthonous bacteria, the link between antibiotic concentrations and genetic changes in soil resistomes, and the role of various contaminants and co-selecting agents on maintenance of ARGs in soil. Moreover, there is a need to complement biological assays with advanced analytical methods (such as LC-MS/MS, isotope dilution mass-spectrometry) and proper sample preparation, allowing assessment of antibiotic concentrations and, especially, of bioavailability in complex environmental matrices. Other problems that should be taken into consideration are variations on the procedures for estimating the limits of antibiotic detection and the lack of standard analytical methods for monitoring antibiotic in the environment. Future studies should also be focused on design and management practices that minimize the spread of ARGs from harvested crops in antibiotic-treated soils into the food chain. Finally, though it is rarely studied, the genetic potential of microorganisms involved in the degradation of antibiotics in soil warrant wider investigation.
Statements
Author contributions
MC, AM, and ZP-S conceived and designed content of the paper and wrote the paper. MC prepared the figures and tables.
Acknowledgments
This work was financed by the National Science Center, Poland (No. DEC-2014/15B/NZ9/04414).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The reviewer UNR and handling Editor declared their shared affiliation.
References
1
Accinelli C. Koskinen W. C. Becker J. M. Sadowsky M. J. (2007). Environmental fate of two sulfonamide antimicrobial agents in soil. J. Agric. Food Chem. 55, 2677–2682. 10.1021/jf063709j
2
Aga D. S. Lenczewski M. Snow D. Muurinen J. Sallach J. B. Wallace J. S. (2016). Challenges in the measurement of antibiotics and in evaluating their impacts in agroecosystems: a critical review. J. Environ. Qual. 45, 407–419. 10.2134/jeq2015.07039
3
Akimenko Y. V. Kazeev K. S. Kolesnikov S. I. (2015). Impact assessment of soil contamination with antibiotics (For example, an ordinary chernozem). Am. J. Appl. Sci. 12, 80–88. 10.3844/ajassp.2015.80.88
4
Alef K. (1995). Dehydrogenase activity, in Methods in Applied Soil Microbiology and Biochemistry, eds AlefK.NannipieriP. (London: Academic), 228–231.
5
Amorim C. L. Moreira I. S. Maia A. S. Tiritan M. E. Castro P. M. L. (2014). Biodegradation of ofloxacin, norfloxacin, and ciprofloxacin as single and mixed substrates by Labrys portucalensis F11. Appl. Microbiol. Biotechnol. 98, 3181–3190. 10.1007/s00253-013-5333-8
6
Andersson D. I. Hughes D. (2012). Evolution of antibiotic resistance at non-lethal drug concentration. Drug Resist. Updat. 15, 162–172. 10.1016/j.drup.2012.03.005
7
Andersson D. I. Hughes D. (2014). Microbiological effects of sublethal levels of antibiotics. Nat. Rev. Microbiol. 12, 465–478. 10.1038/nrmicro3270
8
Ashbolt N. J. Amézquita A. Backhaus T. Borriello P. Brandt K. K. Collignon P. et al . (2013). Human health risk assessment (HHRA) for environmental development and transfer of antibiotic resistance. Environ. Health Perspect. 121, 993–1001. 10.1289/ehp.1206316
9
Atashgahi S. Sánchez-Andrea I. Heipieper H. J. van der Meer J. R. Stams A. J.M. Smidt H. (2018). Prospects for harnessing biocide resistance for bioremediation and detoxification. Science360, 743–746. 10.1126/science.aar3778
10
Aust M.-O. Godlinski F. Travis G. R. Hao X. McAllister T. A. Leinweber P. et al . (2008). Distribution of sulfamethazine, chlortetracycline and tylosin in manure and soil of Canadian feedlots after subtherapeutic use in cattle. Environ. Pollut. 156, 1243–1251. 10.1016/j.envpol.2008.03.011
11
Aust M.-O. Thiele-Bruhn S. Seeger J. Godlinski F. Meissner R. Leinweber P. (2010). Sulfonamides leach from sandy loam soils under common agricultural practice. Water. Air. Soil Pollut. 211, 143–156. 10.1007/s11270-009-0288-1
12
Awad Y. M. Kim S.-C. Abd El-Azeem S. A. M. Kim K.-H. Kim K.-R. Kim K. et al . (2014). Veterinary antibiotics contamination in water, sediment, and soil near a swine manure composting facility. Environ. Earth Sci. 71, 1433–1440. 10.1007/s12665-013-2548-z
13
Awad Y. M. Ok Y. S. Igalavithana A. D. Lee Y. H. Sonn Y.-K. Usman A. R. A. et al . (2016). Sulphamethazine in poultry manure changes carbon and nitrogen mineralisation in soils. Chem. Ecol. 32, 899–918. 10.1080/02757540.2016.1216104
14
Bahlm M. I. Sørensen S. J. Hansen L. H. Licht T. R. (2004). Effect of tetracycline on transfer and establishment of the tetracycline-inducible conjugative transposon Tn916 in the guts of gnotobiotic rats. Appl. Environ. Microbiol. 70, 758–764. 10.1128/AEM.70.2.758-764.2004
15
Bak S. A. Hansen M. Krogh K. A. Brandt A. Halling-Sørensen B. Björklund E. (2013). Development and validation of an SPE methodology combined with LC-MS/MS for the determination of four ionophores in aqueous environmental matrices. Int. J. Environ. Anal. Chem. 93, 1500–1512. 10.1080/03067319.2013.763250
16
Baker-Austin C. Wright M. S. Stepanauskas R. McArthur J. V. (2006). Co-selection of antibiotic and metal resistance. Trends Microbiol.14, 176–182. 10.1016/j.tim.2006.02.006
17
Barra Caracciolo A. Topp E. Grenni P. (2015). Pharmaceuticals in the environment: biodegradation and effects on natural microbial communities. A review. J. Pharm. Biomed. Anal.106, 25–36. 10.1016/j.jpba.2014.11.040
18
Bengtsson-Palme J. Larsson D. G. J. (2016). Concentrations of antibiotics predicted to select for resistant bacteria: proposed limits for environmental regulation. Environ. Int.86, 140–149. 10.1016/j.envint.2015.10.015
19
Berendsen B. J. A. Wegh R. S. Memelink J. Zuidema T. Stolker A. A. M. (2015). The analysis of animal feces as a tool to monitor antibiotic usage. Talanta132, 258–268. 10.1016/j.talanta.2014.09.022
20
Binh C. T. T. Heuer H. Gomes N. C. M. Kotzerke A. Fulle M. Wilke B. M. et al . (2007). Reichel et aShort-term effects of amoxicillin on bacterial communities in manured soil. FEMS Microbiol. Ecol. 62, 290–302. 10.1111/j.1574-6941.2007.00393.x
21
Blackwell P. A. Kay P. Boxall A. B. A. (2007). The dissipation and transport of veterinary antibiotics in a sandy loam soil. Chemosphere67, 292–299. 10.1016/j.chemosphere.2006.09.095
22
Bouki C. Venieri D. Diamadopoulos E. (2013). Detection and fate of antibiotic resistant bacteria in wastewater treatment plants: a review. Ecotoxicol. Environ. Saf. 91, 1–9. 10.1016/j.ecoenv.2013.01.016
23
Boxall A. B. A. Blackwell P. Cavallo R. Kay P. Tolls J. (2002). The sorption and transport of a sulphonamide antibiotic in soil systems. Toxicol. Lett. 131, 19–28. 10.1016/S0378-4274(02)00063-2
24
Boxall A. B. A. Johnson P. Smith E. J. Sinclair C. J. Stutt E. Levy L. S. (2006). Uptake of veterinary medicines from soils into plants. J. Agric. Food Chem. 54, 2288–2297. 10.1021/jf053041t
25
Boyd S. A. Mortland M. M. (1990). Enzyme interactions with clays and clay-organic matter complexes, in Soil BiochemistryVol. 6, eds BollagJ. M.StotzkyG. (New York, NY: Marcel Dekker Inc.), 1–28.
26
Brandt K. K. Amézquita A. Backhaus T. Boxall A. Coors A. Heberer T. et al . (2015). Ecotoxicological assessment of antibiotics: a call for improved consideration of microorganisms. Environ. Int. 85, 189–205. 10.1016/j.envint.2015.09.013
27
Brandt K. K. Sjøholm O. R. Krogh K. A. Halling-Sørensen B. Nybroe O. (2009). Increased pollution-induced bacterial community tolerance to sulfadiazine in soil hotspots amended with artificial root exudates. Environ. Sci. Technol. 43, 2963–2968. 10.1021/es803546y
28
Braschi I. Blasioli S. Fellet C. Lorenzini R. Garelli A. Pori M. et al . (2013). Persistence and degradation of new β-lactam antibiotics in the soil and water environment. Chemosphere93, 152–159. 10.1016/j.chemosphere.2013.05.016
29
Carlson J. C. Mabury S. A. (2006). Dissipation kinetics and mobility of chlortetracycline, tylosin, and monensin in an agricultural soil in Northumberland County, Ontario, Canada. Environ. Toxicol. Chem. 25, 1–10. 10.1897/04-657R.1
30
Carter L. J. Harris E. Williams M. Ryan J. J. Kookana R. S. Boxall A. B. A. (2014). Fate and uptake of pharmaceuticals in soil-plant systems. J. Agric. Food Chem. 62, 5955–5963. 10.1021/jf404282y,
31
CDC Centers for Disease (2015). Control and Prevention. Outpatient Antibiotic Prescription–United States, (Atlanta).
32
Cermák L. Kopecký J. Novotná J. Omelka M. Parkhomenko N. Plháčková K. et al . (2008). Bacterial communities of two contrasting soils reacted differently to lincomycin treatment. Appl. Soil Ecol. 40, 348–358. 10.1016/j.apsoil.2008.06.001
33
Chapman J. S. (2003). Disinfectant resistance mechanisms, cross-resistance, and co-resistance. Int. Biodeterior. Biodegr. 51, 271–276. 10.1016/S0964-8305(03)00044-1
34
Chen B. He R. Yuan K. Chen E. Lin L. Chen X. et al . (2017). Polycyclic aromatic hydrocarbons (PAHs) enriching antibiotic resistance genes (ARGs) in the soils. Environ. Pollut.220, 1005–1013. 10.1016/j.envpol.2016.11.047
35
Chen B. Yuan K. Chen X. Yang Y. Zhang T. Wang Y. et al . (2016). Metagenomic analysis revealing antibiotic resistance genes (ARGs) and their genetic compartments in the Tibetan Environment. Environ. Sci. Technol. 50, 6670–6679. 10.1021/acs.est.6b00619
36
Chen C. Li J. Chen P. Ding R. Zhang P. Li X. (2014). Occurrence of antibiotics and antibiotic resistances in soils from wastewater irrigation areas in Beijing and Tianjin, China. Environ. Pollut. 193, 94–101. 10.1016/j.envpol.2014.06.005
37
Chen W. Liu W. Pan N. Jiao W. Wang M. (2013). Oxytetracycline on functions and structure of soil microbial community. J. Soil Sci. Plant Nutr. 13, 967–975. 10.4067/S0718-95162013005000076
38
Chen Y. S. Zhang H. B. Luo Y. M. Song J. (2012). Occurrence and assessment of veterinary antibiotics in swine manures: a case study in East China. Chinese Sci. Bull. 57, 606–614. 10.1007/s11434-011-4830-3
39
Cheng M. Wu L. Huang Y. Luo Y. Christie P. (2014). Total concentrations of heavy metals and occurrence of antibiotics in sewage sludges from cities throughout China. J. Soils Sediments14, 1123–1135. 10.1007/s11368-014-0850-3
40
Chessa L. Jechalke S. Ding G.-C. Pusino A. Mangia N. P. Smalla K. (2016a). The presence of tetracycline in cow manure changes the impact of repeated manure application on soil bacterial communities. Biol. Fertil. Soils52, 1121–1134. 10.1007/s00374-016-1150-4
41
Chessa L. Pusino A. Garau G. Mangia N. P. Pinna M. V. (2016b). Soil microbial response to tetracycline in two different soils amended with cow manure. Environ. Sci. Pollut. Res. 23, 5807–5817. 10.1007/s11356-015-5789-4
42
Conkle J. L. Lattao C. White J. R. Cook R. L. (2010). Competitive sorption and desorption behavior for three fluoroquinolone antibiotics in a wastewater treatment wetland soil. Chemosphere80, 1353–1359. 10.1016/j.chemosphere.2010.06.012
43
Crane M. Boxall A. B. A. Barret K. (2010). Veterinary Medicines in the Environment. Boca Raton, FL: CRC Press.
44
Cui H. Wang S.-P. Fu J. Zhou Z.-Q. Zhang N. Guo L. (2014). Influence of ciprofloxacin on microbial community structure and function in soils. Biol. Fertil. Soils50, 939–947. 10.1007/s00374-014-0914-y
45
Cycoń M. Borymski S. Orlewska K. Wasik T. J. Piotrowska-Seget Z. (2016b). An analysis of the effects of vancomycin and/or vancomycin-resistant Citrobacter freundii exposure on the microbial community structure in soil. Front. Microbiol. 7:1015. 10.3389/fmicb.2016.01015
46
Cycoń M. Borymski S. Zołnierczyk B. Piotrowska-Seget Z. (2016a). Variable effects of non-steroidal anti-inflammatory drugs (NSAIDs) on selected biochemical processes mediated by soil microorganisms. Front. Microbiol. 7:1969. 10.3389/fmicb.2016.01969
47
Cycoń M. Orlewska K. Markowicz A. Zmijowska A. Smolen-Dzirba J. Bratosiewicz-Wasik J. et al . (2018). Vancomycin and/or multidrug-resistant Citrobacter freundii altered the metabolic pattern of soil microbial community. Front. Microbiol.9:1047. 10.3389/fmicb.2018.01047
48
Cycoń M. Piotrowska-Seget Z. (2015). Biochemical and microbial soil functioning after application of the insecticide imidacloprid. J. Environ. Sci.27, 147–158. 10.1016/j.jes.2014.05.034
49
Cycoń M. Piotrowska-Seget Z. (2016). Pyrethroid-degrading microorganisms and their potential for the bioremediation of contaminated soils: a review. Front. Microbiol.7:1463. 10.3389/fmicb.2016.01463
50
Cycoń M. Zmijowska A. Piotrowska-Seget Z. (2011). Biodegradation kinetics of 2,4-D by bacterial strains isolated from soil. Cent. Eur. J. Biol.6, 188–198. 10.2478/s11535-011-0005-0
51
Daghrir R. Drogui P. (2013). Tetracycline antibiotics in the environment: a review. Environ. Chem. Lett. 11, 209–227. 10.1007/s10311-013-0404-8
52
Dantas G. Sommer M. O. A. Oluwasegun R. D. Church G. M. (2008). Bacteria subsisting on antibiotics. Science320, 100–103. 10.1126/science.1155157
53
Davis J. G. Truman C. C. Kim S. C. Ascoughd J. C. Carlsone K. (2006). Antibiotic transport via runoff and soil loss. J. Environ. Qual. 35, 2250–2260. 10.2134/jeq2005.0348
54
D'Costa V. M. Griffith E. Wright G. D. (2007). Expanding the soil antibiotic resistome: exploring environmental diversity. Curr. Opin. Microbiol. 10, 481–489. 10.1016/j.mib.2007.08.009
55
D'Costa V. M. McGrann K. M. Hughes D. W. Wright G. D. (2006). Sampling the antibiotic resistome. Science311, 374–377. 10.1126/science.1120800
56
De la Torre A. Iglesias I. Carballo M. Ramírez P. Muñoz M. J. (2012). An approach for mapping the vulnerability of European Union soils to antibiotic contamination. Sci. Total Environ. 414, 672–679. 10.1016/j.scitotenv.2011.10.032
57
Demoling L. A. Bååth E. Greve G. Wouterse M. Schmitt H. (2009). Effects of sulfamethoxazole on soil microbial communities after adding substrate. Soil Biol. Biochem. 41, 840–848. 10.1016/j.soilbio.2009.02.001
58
DeVries S. L. Loving M. Li X. Zhang P. (2015). The effect of ultralow-dose antibiotics exposure on soil nitrate and N2O flux. Sci. Rep. 5:16818. 10.1038/srep16818
59
DeVries S. L. Zhang P. (2016). Antibiotics and the terrestrial nitrogen cycle: a review. Curr. Pollut. Rep.2, 51–67. 10.1007/s40726-016-0027-3
60
Ding C. He J. (2010). Effect of antibiotics in the environment on microbial populations. Appl. Microbiol. Biotechnol. 87, 925–941. 10.1007/s00253-010-2649-5
61
Ding G. C. Radl V. Schloter-Hai B. Jechalke S. Heuer H. Smalla K. et al . (2014). Dynamics of soil bacterial communities in response to repeated application of manure containing sulfadiazine. PLoS ONE9:e92958. 10.1371/journal.pone.0092958
62
Ding Y. Zhang W. Gu C. Xagoraraki I. Li H. (2011). Determination of pharmaceuticals in biosolids using accelerated solvent extraction and liquid chromatography/tandem mass spectrometry. J. Chromatogr. A1218, 10–16. 10.1016/j.chroma.2010.10.112
63
Dolliver H. Gupta S. Noll S. (2008). Antibiotic degradation during manure composting. J. Environ. Qual. 37:1245. 10.2134/jeq2007.0399
64
Dolliver H. Kumar K. Gupta S. (2007). Sulfamethazine uptake by plants from manure-amended soil. J. Environ. Qual. 36:1224. 10.2134/jeq2006.0266
65
Donato J. J. Moe L. A. Converse B. J. Smart K. D. Berklein E. C. McManus P. S. et al . (2010). Metagenomic analysis of apple orchard soil reveals antibiotic resistance genes encoding predicted bifunctional proteins. Appl. Environ. Microbiol.76, 4396–4401. 10.1128/AEM.01763-09
66
Du L. Liu W. (2012). Occurrence, fate, and ecotoxicity of antibiotics in agro-ecosystems. A review. Agron. Sustain. Dev. 32, 309–327. 10.1007/s13593-011-0062-9
67
Duan M. Li H. Gu J. Tuo X. Sun W. Qian X. et al . (2017). Effects of biochar on reducing the abundance of oxytetracycline, antibiotic resistance genes, and human pathogenic bacteria in soil and lettuce. Environ. Pollut. 224, 787–795. 10.1016/j.envpol.2017.01.021
68
ECDC (2018). ESAC-Net surveillance data (2017). Summary of the latest data on antibiotic consumption in the European Union. European Centre for Disease Prevention and Control. Antimicrobial consumption, in ECDC Annual Epidemiological Report for 2016. (Stockholm: ECDC).
69
Erickson B. D. Elkins C. A. Mullis L. B. Heinze T. M. Wagner R. D. Cerniglia C. E. (2014). A metallo-β-lactamase is responsible for the degradation of ceftiofur by the bovine intestinal bacterium Bacillus cereus P41. Vet. Microbiol. 172, 499–504. 10.1016/j.vetmic.2014.05.032
70
Fahrenfeld N. Ma Y. O'Brien M. Pruden A. (2013). Reclaimed water as a reservoir of antibiotic resistance genes: distribution system and irrigation implications. Front. Microbiol. 4:130. 10.3389/fmicb.2013.00130
71
Fan Z. Casey F. X. M. Hakk H. Larsen G. L. Khan E. (2011). Sorption, fate, and mobility of sulfonamides in soils. Water Air. Soil Pollut. 218, 49–61. 10.1007/s11270-010-0623-6
72
Fang H. Han L. Cui Y. Xue Y. Cai L. Yu Y. (2016). Changes in soil microbial community structure and function associated with degradation and resistance of carbendazim and chlortetracycline during repeated treatments. Sci. Total Environ. 572, 1203–1212. 10.1016/j.scitotenv.2016.08.038
73
Fang H. Han Y. Yin Y. Pan X. Yu Y. (2014). Variations in dissipation rate, microbial function and antibiotic resistance due to repeated introductions of manure containing sulfadiazine and chlortetracycline to soil. Chemosphere96, 51–56. 10.1016/j.chemosphere.2013.07.016
74
Fatta-Kassinos D. Kalavrouziotis I. K. Koukoulakis P. H. Vasquez M. I. (2011). The risks associated with wastewater reuse and xenobiotics in the agroecological environment. Sci. Total Environ. 409, 3555–3563. 10.1016/j.scitotenv.2010.03.036
75
Fernández-Calviño D. Bermúdez-Couso A. Arias-Estévez M. Nóvoa-Muñoz J. C. Fernández-Sanjurjo M. J. Álvarez-Rodríguez E. et al . (2015). Competitive adsorption/desorption of tetracycline, oxytetracycline and chlortetracycline on two acid soils: stirred flow chamber experiments. Chemosphere134, 361–366. 10.1016/j.chemosphere.2015.04.098
76
Floch C. Chevremont A.-C. Joanico K. Capowiez Y. Criquet S. (2011). Indicators of pesticide contamination: soil enzyme compared to functional diversity of bacterial communities via Biolog® Ecoplates. Eur. J. Soil Biol. 47, 256–263. 10.1016/j.ejsobi.2011.05.007
77
Forsberg K. J. Reyes A. Wang B. Selleck E. M. Sommer M. O. A. Dantas G. (2012). The shared antibiotic resistome of soil bacteria and human pathogens. Science337, 1107–1111. 10.1126/science.1220761
78
Förster M. Laabs V. Lamshöft M. Groeneweg J. Zühlke S. Spiteller M. et al . (2009). Sequestration of manure-applied sulfadiazine residues in soils. Environ. Sci. Technol. 43, 1824–1830. 10.1021/es8026538
79
Frostegård Å. Tunlid A. Bååth E. (2011). Use and misuse of PLFA measurements in soils. Soil Biol. Biochem. 43, 1621–1625. 10.1016/j.soilbio.2010.11.021
80
Frostegård Å. Tunlid A. Baath E. (1993). Phospholipid fatty acid composition, biomass, and activity of microbial communities from two soil types experimentally exposed to different heavy metals. Appl. Environ. Microbiol. 59, 3605–3617.
81
Gao L. Shi Y. Li W. Liu J. Cai Y. (2015). Occurrence and distribution of antibiotics in urban soil in Beijing and Shanghai, China. Environ. Sci. Pollut. Res. 22, 11360–11371. 10.1007/s11356-015-4230-3
82
Gavalchin J. Katz S. E. (1994). The persistence of fecal-borne antibiotics in soil. J. Assoc. Anal. Chem. Int. 77, 481–485.
83
Gelband H. Miller-Petrie M. Pant S. Gandra S. Levinson J. Barter D. et al . (2015). State of the world's antibiotics, in Center for Disease Dynamics, Economics & Policy, eds CDDEP (Washington, DC: CDDEP), 1–84.
84
Ghosh S. LaPara T. M. (2007). The effects of subtherapeutic antibiotic use in farm animals on the proliferation and persistence of antibiotic resistance among soil bacteria. ISME J. 1, 191–203. 10.1038/ismej.2007.31
85
Ghosh S. Sadowsky M. J. Roberts M. C. Gralnick J. A. LaPara T. M. (2009). Sphingobacterium sp. strain PM2-P1-29 harbours a functional tet(X) gene encoding for the degradation of tetracycline. J. Appl. Microbiol.106, 1336–1342. 10.1111/j.1365-2672.2008.04101.x
86
Gianfreda L. Sannino F. Ortega N. Nannipieri P. (1994). Activity of free and immobilized urease in soil: effects of pesticides. Soil Biol. Biochem. 26, 777–784. 10.1016/0038-0717(94)90273-9
87
Gil-Sotres F. Trasar-Cepeda C. Leirós M. C. Seoane S. (2005). Different approaches to evaluating soil quality using biochemical properties. Soil Biol. Biochem. 37, 877–887. 10.1016/j.soilbio.2004.10.003
88
Girardi C. Greve J. Lamshöft M. Fetzer I. Miltner A. Schäffer A. et al . (2011). Biodegradation of ciprofloxacin in water and soil and its effects on the microbial communities. J. Hazard. Mater. 198, 22–30. 10.1016/j.jhazmat.2011.10.004
89
Göbel A Thomsen A. McArdell C. S. Alder A. C Giger W. Theiß N. Löffler D. et al . (2005). Extraction and determination of sulfonamides, macrolides, and trimethoprim in sewage sludge. J. Chromatogr. A1085, 179–189. 10.1016/j.chroma.2005.05.051
90
Grenni P. Ancona V. Barra Caracciolo A. (2018). Ecological effects of antibiotics on natural ecosystems: a review. Microchem. J. 136, 25–39. 10.1016/j.microc.2017.02.006
91
Grote M. Schwake-Anduschus C. Michel R. Stevens H. Heyser W. Langenkämper G. et al . (2007). Incorporation of veterinary antibiotics into crops from manured soil. Landbauforsch. Volkenrode57, 25–32.
92
Gudeta D. D. Bortolaia V. Pollini S. Docquier J.-D. Rossolini G. M. Amos A. et al . (2016). Expanding the repertoire of carbapenem-hydrolyzing metallo-β-lactamases by functional metagenomic analysis of soil microbiota. Front. Microbiol.7:1985. 10.3389/fmicb.2016.01985
93
Guerin E. Cambray G. Sanchez-Alberola N. Campoy S. Erill I. Da Re S. et al . (2009). The SOS response controls integron recombination. Science324:1034. 10.1126/science.1172914
94
Gulkowska A. Leung H. W. So M. K. Taniyasu S. Yamashita N. Yeung L. W. Y. et al . (2008). Removal of antibiotics from wastewater by sewage treatment facilities in Hong Kong and Shenzhen, China. Water Res.42, 395–403. 10.1016/j.watres.2007.07.031
95
Gullberg E. Albrecht L. M. Karlsson C. Sandegren L. Andersson D. I. (2014). Selection of a multidrug resistance plasmid by sublethal levels of antibiotics and heavy metals. MBio5, e01918–e01914. 10.1128/mBio.01918-14
96
Gullberg E. Cao S. Berg O. G. Ilbäck C. Sandegren L. Hughes D. et al . (2011). Selection of resistant bacteria at very low antibiotic concentrations. PLoS Pathog. 7:e1002158. 10.1371/journal.ppat.1002158
97
Gutiérrez I. R. Watanabe N. Harter T. Glaser B. Radke M. (2010). Effect of sulfonamide antibiotics on microbial diversity and activity in a Californian Mollic Haploxeralf. J. Soils Sediments10, 537–544. 10.1007/s11368-009-0168-8
98
Halling-Sørensen B. Jacobsen A.-M. Jensen J. Sengeløv G. Vaclavik E. Ingerslev F. (2005). Dissipation and effects of chlortetracycline and tylosin in two agricultural soils: a field-scale study in southern Denmark. Environ. Toxicol. Chem. 24, 802–810. 10.1897/03-576.1
99
Hammesfahr U. Bierl R. Thiele-Bruhn S. (2011). Combined effects of the antibiotic sulfadiazine and liquid manure on the soil microbial-community structure and functions. J. Plant Nutr. Soil Sci. 174, 614–623. 10.1002/jpln.201000322
100
Hammesfahr U. Heuer H. Manzke B. Smalla K. Thiele-Bruhn S. (2008). Impact of the antibiotic sulfadiazine and pig manure on the microbial community structure in agricultural soils. Soil Biol. Biochem. 40, 1583–1591. 10.1016/j.soilbio.2008.01.010
101
Hamscher G. Pawelzick H. T. Höper H. Nau H. (2005). Different behavior of tetracyclines and sulfonamides in sandy soils after repeated fertilization with liquid manure. Environ. Toxicol. Chem. 24, 861–868. 10.1897/04-182R.1
102
Hamscher G. Sczesny S. Höper H. Nau H. (2002). Determination of persistent tetracycline residues in soil fertilized with liquid manure by high-performance liquid chromatography with electrospray ionization tandem mass spectrometry. Anal. Chem. 74, 1509–1518. 10.1021/ac015588m
103
Herrick J. B. Haynes R. Heringa S. Brooks J. M. Sobota L. T. (2014). Coselection for resistance to multiple late-generation human therapeutic antibiotics encoded on tetracycline resistance plasmids captured from uncultivated stream and soil bacteria. J. Appl. Microbiol. 117, 380–389. 10.1111/jam.12538
104
Heuer H. Schmitt H. Smalla K. (2011a). Antibiotic resistance gene spread due to manure application on agricultural fields. Curr. Opin. Microbiol. 14, 236–243. 10.1016/j.mib.2011.04.009
105
Heuer H. Smalla K. (2007). Manure and sulfadiazine synergistically increased bacterial antibiotic resistance in soil over at least two months. Environ. Microbiol. 9, 657–666. 10.1111/j.1462-2920.2006.01185.x
106
Heuer H. Solehati Q. Zimmerling U. Kleineidam K. Schloter M. Müller T. et al . (2011b). Accumulation of sulfonamide resistance genes in arable soils due to repeated application of manure containing sulfadiazine. Appl. Environ. Microbiol. 77, 2527–2530. 10.1128/AEM.02577-10
107
Hirth N. Topp E. Dörfler U. Stupperich E. Munch J. C. Schroll R. (2016). An effective bioremediation approach for enhanced microbial degradation of the veterinary antibiotic sulfamethazine in an agricultural soil. Chem. Biol. Technol. Agric. 3:29. 10.1186/s40538-016-0080-6
108
Ho Y. B. Zakaria M. P. Latif P. A. Saari N. (2014). Occurrence of veterinary antibiotics and progesterone in broiler manure and agricultural soil in Malaysia. Sci. Total Environ.488–489, 261–267. 10.1016/j.scitotenv.2014.04.109
109
Hou J. Wan W. Mao D. Wang C. Mu Q. Qin S. et al . (2015). Occurrence and distribution of sulfonamides, tetracyclines, quinolones, macrolides, and nitrofurans in livestock manure and amended soils of Northern China. Environ. Sci. Pollut. Res. 22, 4545–4554. 10.1007/s11356-014-3632-y
110
Hu D. Coats J. R. (2007). Aerobic degradation and photolysis of tylosin in water and soil. Environ. Toxicol. Chem. 26, 884–889. 10.1897/06-197R.1
111
Hu H.-W. Han X.-M. Shi X.-Z. Wang J.-T. Han L.-L. Chen D. et al . (2016). Temporal changes of antibiotic-resistance genes and bacterial communities in two contrasting soils treated with cattle manure. FEMS Microbiol. Ecol. 92, 1–13. 10.1093/femsec/fiv169
112
Hu X. Zhou Q. Luo Y. (2010). Occurrence and source analysis of typical veterinary antibiotics in manure, soil, vegetables and groundwater from organic vegetable bases, northern China. Environ. Pollut. 158, 2992–2998. 10.1016/j.envpol.2010.05.023
113
Huang T. Xu Y. Zeng J. Zhao D.-H. Li L. Liao X.-P. et al . (2016). Low-concentration ciprofloxacin selects plasmid-mediated quinolone resistance encoding genes and affects bacterial taxa in soil containing manure. Front. Microbiol. 7:1730. 10.3389/fmicb.2016.01730
114
Huang X. Liu C. Li K. Liu F. Liao D. Liu L. et al . (2013). Occurrence and distribution of veterinary antibiotics and tetracycline resistance genes in farmland soils around swine feedlots in Fujian Province, China. Environ. Sci. Pollut. Res. 20, 9066–9074. 10.1007/s11356-013-1905-5
115
Ingerslev F. Halling-Sørensen B. (2001). Biodegradability of metronidazole, olaquindox, and tylosin and formation of tylosin degradation products in aerobic soil-manure slurries. Ecotoxicol. Environ. Saf. 48, 311–320. 10.1006/eesa.2000.2026
116
Jacobsen A. M. Halling-Sørensen B. Ingerslev F. Hansen S. H. (2004). Simultaneous extraction of tetracycline, macrolide and sulfonamide antibiotics from agricultural soils using pressurised liquid extraction, followed by solid-phase extraction and liquid chromatography-tandem mass spectrometry. J. Chromatogr. A1038, 157–170. 10.1016/j.chroma.2004.03.034
117
Jechalke S. Focks A. Rosendahl I. Groeneweg J. Siemens J. Heuer H. et al . (2014a). Structural and functional response of the soil bacterial community to application of manure from difloxacin-treated pigs. FEMS Microbiol. Ecol. 87, 78–88. 10.1111/1574-6941.12191
118
Jechalke S. Heuer H. Siemens J. Amelung W. Smalla K. (2014b). Fate and effects of veterinary antibiotics in soil. Trends Microbiol. 22, 536–545. 10.1016/j.tim.2014.05.005
119
Jones-Lepp T. L. Stevens R. (2007). Pharmaceuticals and personal care products in biosolids/sewage sludge: the interface between analytical chemistry and regulation. Anal. Bioanal. Chem. 387, 1173–1183. 10.1007/s00216-006-0942-z
120
Kang D. H. Gupta S. Rosen C. Fritz V. Singh A. Chander Y. et al . (2013). Antibiotic uptake by vegetable crops from manure-applied soils. J. Agric. Food Chem. 61, 9992–10001. 10.1021/jf404045m
121
Karci A. Balcioglu I. A. (2009). Investigation of the tetracycline, sulfonamide, and fluoroquinolone antimicrobial compounds in animal manure and agricultural soils in Turkey. Sci. Total Environ. 407, 4652–4664. 10.1016/j.scitotenv.2009.04.047
122
Kay P. Blackwell P. A. Boxall A. B. A. (2004). Fate of veterinary antibiotics in a macroporous tile drained clay soil. Environ. Toxicol. Chem. 23, 1136–1143. 10.1897/03-374
123
Kay P. Blackwell P. A. Boxall A. B. A. (2005). A lysimeter experiment to investigate the leaching of veterinary antibiotics through a clay soil and comparison with field data. Environ. Pollut. 134, 333–341. 10.1016/j.envpol.2004.07.021
124
Keen P. L. Patrick D. M. (2013). Tracking change: a look at the ecological footprint of antibiotics and antimicrobial resistance. Antibiotics2, 191–205. 10.3390/antibiotics2020191
125
Kim D.-W. Heinze T. M. Kim B.-S. Schnackenberg L. K. Woodling K. A. Sutherland J. B. (2011). Modification of norfloxacin by a Microbacterium sp. strain isolated from a wastewater treatment plant. Appl. Environ. Microbiol. 77, 6100–6108. 10.1128/AEM.00545-11
126
Kinney C. A. Furlong E. T. Zaugg S. D. Burkhardt M. R. Werner S. L. Cahill J. D. et al . (2006). Survey of organic wastewater contaminants in biosolids destined for land application. Environ. Sci. Technol. 40, 7207–7215. 10.1021/es0603406
127
Klein E,. Y. Van Boeckel T. P. Martinez E. M. Pant S. Gandra S. Levin S. A. et al . (2018). Global increase and geographic convergence in antibiotic consumption between 2000 and 2015. Proc. Natl. Acad. Sci. U.S.A.115, E3463–E3479. 10.1073/pnas.1717295115
128
Kleineidam K. Sharma S. Kotzerke A. Heuer H. Thiele-Bruhn S. Smalla K. et al . (2010). Effect of sulfadiazine on abundance and diversity of denitrifying bacteria by determining nirK and nirS genes in two arable soils. Microb. Ecol. 60, 703–707. 10.1007/s00248-010-9691-9
129
Koba O. Golovko O. Kodešová R. Fér M. Grabic R. (2017). Antibiotics degradation in soil: a case of clindamycin, trimethoprim, sulfamethoxazole and their transformation products. Environ. Pollut. 220, 1251–1263. 10.1016/j.envpol.2016.11.007
130
Kong W. Li C. Dolhi J. M. Li S. He J. Qiao M. (2012). Characteristics of oxytetracycline sorption and potential bioavailability in soils with various physical-chemical properties. Chemosphere87, 542–548. 10.1016/j.chemosphere.2011.12.062
131
Kong W.-D. Zhu Y.-G. Fu B.-J. Marschner P. He J.-Z. (2006). The veterinary antibiotic oxytetracycline and Cu influence functional diversity of the soil microbial community. Environ. Pollut. 143, 129–137. 10.1016/j.envpol.2005.11.003
132
Kotzerke A. Hammesfahr U. Kleineidam K. Lamshöft M. Thiele-Bruhn S. Schloter M. et al . (2011). Influence of difloxacin-contaminated manure on microbial community structure and function in soils. Biol. Fertil. Soils47, 177–186. 10.1007/s00374-010-0517-1
133
Kotzerke A. Sharma S. Schauss K. Heuer H. Thiele-Bruhn S. Smalla K. et al . (2008). Alterations in soil microbial activity and N-transformation processes due to sulfadiazine loads in pig-manure. Environ. Pollut. 153, 315–322. 10.1016/j.envpol.2007.08.020
134
Kuchta S. L. Cessna A. J. Elliott J. A. Peru K. M. Headley J. V. (2009). Transport of lincomycin to surface and ground water from manure-amended cropland. J. Environ. Qual. 38, 1719–1729. 10.2134/jeq2008.0365
135
Kulkarni P. Olson N. D. Raspanti G. A. Goldstein R. E. R. Gibbs S. G. Sapkota A. et al . (2017). Antibiotic concentrations decrease during wastewater treatment but persist at low levels in reclaimed water. Int. J. Environ. Res. Public Health14:668. 10.3390/ijerph14060668
136
Kumar K. Gupta C. SChander Y. Singh A. K. (2005a). Antibiotic use in agriculture and its impact on the terrestrial environment. Adv. Agron. 87, 1–54. 10.1016/S0065-2113(05)87001-4
137
Kumar K. Gupta S. C. Baidoo S. K. Chander Y. Rosen C. J. (2005b). Antibiotic uptake by plants from soil fertilized with animal manure. J. Environ. Qual. 34, 2082–2085. 10.2134/jeq2005.0026
138
Kumar R. R. Lee J. T. Cho J. Y. (2012). Fate, occurrence, and toxicity of veterinary antibiotics in environment. J. Korean Soc. Appl. Biol. Chem. 55, 701–709. 10.1007/s13765-012-2220-4
139
Kümmerer K. (2001). Drugs in the environment: Emission of drugs, diagnostic aids and disinfectants into wastewater by hospitals in relation to other sources - A review. Chemosphere45, 957–969. 10.1016/S0045-6535(01)00144-8
140
Kümmerer K. (2009). Antibiotics in the aquatic environment–a review–Part, I. Chemosphere75, 417–434. 10.1016/j.chemosphere.2008.11.086
141
Kyselková M. Jirout J. Chronáková A. Vrchotová N. Bradley R. Schmitt H. et al . (2013). Cow excrements enhance the occurrence of tetracycline resistance genes in soil regardless of their oxytetracycline content. Chemosphere93, 2413–2418. 10.1016/j.chemosphere.2013.08.058
142
Kyselková M. Kotrbová L. Bhumibhamon G. Chronáková A. Jirout J. Vrchotová N. et al . (2015). Tetracycline resistance genes persist in soil amended with cattle feces independently from chlortetracycline selection pressure. Soil Biol. Biochem. 81, 259–265. 10.1016/j.soilbio.2014.11.018
143
Larsson D. G.J. (2014). Antibiotics in the environment. Ups. J. Med. Sci. 119, 108–112. 10.3109/03009734.2014.896438
144
Lau C. H.-F. van Engelen K. Gordon S. Renaud J. Topp E. (2017). Novel antibiotic resistance determinants from agricultural soil exposed to antibiotics widely used in human medicine and animal farming. Appl. Environ. Microbiol. 83, e00989–e00917. 10.1128/AEM.00989-17
145
Laureto L. M. O. Cianciaruso M. V. Samia D. S. M. (2015). Functional diversity: an overview of its history and applicability. Braz. J. Natur. Conserv. 13, 112–116. 10.1016/j.ncon.2015.11.001
146
Leal R. M. P. Alleoni L. R. F. Tornisielo V. L. Regitano J. B. (2013). Sorption of fluoroquinolones and sulfonamides in 13 Brazilian soils. Chemosphere92, 979–985. 10.1016/j.chemosphere.2013.03.018
147
Leal R. M. P. Figueira R. F. Tornisielo V. L. Regitano J. B. (2012). Occurrence and sorption of fluoroquinolones in poultry litters and soils from São Paulo State, Brazil. Sci. Total Environ. 432, 344–349. 10.1016/j.scitotenv.2012.06.002
148
Leng Y. Bao J. Chang G. Zheng H. Li X. Du J. et al . (2016). Biotransformation of tetracycline by a novel bacterial strain Stenotrophomonas maltophilia DT1. J. Hazard. Mater. 318, 125–133. 10.1016/j.jhazmat.2016.06.053
149
Li C. Chen J. Wang J. Ma Z. Han P. Luan Y. et al . (2015). Occurrence of antibiotics in soils and manures from greenhouse vegetable production bases of Beijing, China and an associated risk assessment. Sci. Total Environ.521–522, 101–107. 10.1016/j.scitotenv.2015.03.070
150
Li L.-L. Huang L.-D. Chung R.-S. Fok K.-H. Zhang Y.-S. (2010a). Sorption and dissipation of tetracyclines in soils and compost. Pedosphere20, 807–816. 10.1016/S1002-0160(10)60071-9
151
Li W. Shi Y. Gao L. Liu J. Cai Y. (2013). Occurrence, distribution and potential affecting factors of antibiotics in sewage sludge of wastewater treatment plants in China. Sci. Tot. Environ.445–446, 306–313. 10.1016/j.scitotenv.2012.12.050
152
Li Y.-W. Mo C.-H. Zhao N. Tai Y.-P. Bao Y.-P. Wang J.-Y. et al . (2009). Investigation of sulfonamides and tetracyclines antibiotics in soils from various vegetable fields. Huanjing Kexue/Environmental Sci. 30, 1762–1766.
153
Li Y.-W. Wu X.-L. Mo C.-H. Tai Y.-P. Huang X.-P. Xiang L. (2011). Investigation of sulfonamide, tetracycline, and quinolone antibiotics in vegetable farmland soil in the pearl river delta area, Southern China. J. Agric. Food Chem. 59, 7268–7276. 10.1021/jf1047578
154
Li Z. Chang P.-H. Jean J.-S. Jiang W.-T. Wang C.-J. (2010b). Interaction between tetracycline and smectite in aqueous solution. J. Colloid Interface Sci. 341, 311–319. 10.1016/j.jcis.2009.09.054
155
Lillenberg M. Yurchenko S. Kipper K. Herodes K. Pihl V. Lõhmus R. et al . (2010). Presence of fluoroquinolones and sulfonamides in urban sewage sludge and their degradation as a result of composting. Int. J. Environ. Sci. Technol. 7, 307–312. 10.1007/BF03326140
156
Lin K. Gan J. (2011). Sorption and degradation of wastewater-associated non-steroidal anti-inflammatory drugs and antibiotics in soils. Chemosphere83, 240–246. 10.1016/j.chemosphere.2010.12.083
157
Liu B. Li Y. Zhang X. Wang J. Gao M. (2014). Combined effects of chlortetracycline and dissolved organic matter extracted from pig manure on the functional diversity of soil microbial community. Soil Biol. Biochem. 74, 148–155. 10.1016/j.soilbio.2014.03.005
158
Liu B. Li Y. Zhang X. Wang J. Gao M. (2015). Effects of chlortetracycline on soil microbial communities: comparisons of enzyme activities to the functional diversity via Biolog EcoPlates™. Eur. J. Soil Biol. 68, 69–76. 10.1016/j.ejsobi.2015.01.002
159
Liu F. Wu J. Ying G.-G. Luo Z. Feng H. (2012a). Changes in functional diversity of soil microbial community with addition of antibiotics sulfamethoxazole and chlortetracycline. Appl. Microbiol. Biotechnol. 95, 1615–1623. 10.1007/s00253-011-3831-0
160
Liu F. Ying G.-G. Tao R. Zhao J.-L. Yang J.-F. Zhao L.-F. (2009). Effects of six selected antibiotics on plant growth and soil microbial and enzymatic activities. Environ. Pollut. 157, 1636–1642. 10.1016/j.envpol.2008.12.021
161
Liu L. Liu Y-H Liu C-X. Huang X. (2016). Accumulation of antibiotics and tet resistance genes from swine wastewater in wetland soils. Environ. Eng. Manage. J. 15, 2137–2145. 10.30638/eemj.2016.231
162
Liu W. Pan N. Chen W. Jiao W. Wang M. (2012b). Effect of veterinary oxytetracycline on functional diversity of soil microbial community. Plant Soil Environ. 58, 295–301. 10.1016/j.ibiod.2012.06.010
163
Liu Z. Han Y. Jing M. Chen J. (2017). Sorption and transport of sulfonamides in soils amended with wheat straw-derived biochar: effects of water pH, coexistence copper ion, and dissolved organic matter. J. Soils Sediments17, 771–779. 10.1007/s11368-015-1319-8
164
Łukaszewicz P. Białk-Bielinska A. Dołzonek J. Kumirska J. Caban M. Stepnowski P. (2018). A new approach for the extraction of tetracyclines from soil matrices: application of the microwave-extraction technique. Anal. Bioanal. Chem. 410, 1697–1707. 10.1007/s00216-017-0815-7
165
Ma J. Lin H. Sun W. Wang Q. Yu Q. Zhao Y. et al . (2014). Soil microbial systems respond differentially to tetracycline, sulfamonomethoxine, and ciprofloxacin entering soil under pot experimental conditions alone and in combination. Environ. Sci. Pollut. Res. 21, 7436–7448. 10.1007/s11356-014-2685-2
166
Ma T. Pan X. Chen L. Liu W. Christie P. Luo Y. et al . (2016). Effects of different concentrations and application frequencies of oxytetracycline on soil enzyme activities and microbial community diversity. Eur. J. Soil Biol. 76, 53–60. 10.1016/j.ejsobi.2016.07.004
167
Manzetti S. Ghisi R. (2014). The environmental release and fate of antibiotics. Mar. Pollut. Bull. 79, 7–15. 10.1016/j.marpolbul.2014.01.005
168
Martens R. Wetzstein H.-G. Zadrazil F. Capelari M. Hoffmann P. Schmeer N. (1996). Degradation of the fluoroquinolone enrofloxacin by wood-rotting fungi. Appl. Environ. Microbiol. 62, 4206–4209.
169
Martinez J. L. Sánchez M. B. Martínez-Solano L. Hernandez A. Garmendia L. Fajardo A. et al . (2009). Functional role of bacterial multidrug efflux pumps in microbial natural ecosystems. FEMS Microbiol. Rev. 33, 430–449. 10.1111/j.1574-976.2008.00157.x
170
Martínez-Carballo E. González-Barreiro C. Scharf S. Gans O. (2007). Environmental monitoring study of selected veterinary antibiotics in animal manure and soils in Austria. Environ. Pollut. 148, 570–579. 10.1016/j.envpol.2006.11.035
171
Martínez-Hernández V. Meffe R. Herrera López S. de Bustamante I. (2016). The role of sorption and biodegradation in the removal of acetaminophen, carbamazepine, caffeine, naproxen and sulfamethoxazole during soil contact: a kinetics study. Sci. Total Environ. 559, 232–241. 10.1016/j.scitotenv.2016.03.131
172
Marx M.-C. Kandeler E. Wood M. Wermbter N. Jarvis S. C. (2005). Exploring the enzymatic landscape: distribution and kinetics of hydrolytic enzymes in soil particle-size fractions. Soil Biol. Biochem. 37, 35–48. 10.1016/j.soilbio.2004.05.024
173
Massé D. I. Saady N. M. C. Gilbert Y. (2014). Potential of biological processes to eliminate antibiotics in livestock manure: an overview. Animals4, 146–163. 10.3390/ani4020146
174
McFarland J. W. Berger C. M. Froshauer S. A. Hayashi S. F. Hecker S. J. Jaynes B. H. et al . (1997). Quantitative structure-activity relationships among macrolide antibacterial agents: in vitro and in vivo potency against Pasteurella multocida. J. Med. Chem. 40, 1340–1346. 10.1021/jm960436i
175
Michael I. Rizzo L. McArdell C. S. Manaia C. M. Merlin C. Schwartz T. et al . (2013). Urban wastewater treatment plants as hotspots for the release of antibiotics in the environment: a review. Water Res.47, 957–995. 10.1016/j.watres.2012.11.027
176
Mitchell S. M. Ullman J. L. Teel A. L. Watts R. J. (2015). Hydrolysis of amphenicol and macrolide antibiotics: chloramphenicol, florfenicol, spiramycin, and tylosin. Chemosphere134, 504–511. 10.1016/j.chemosphere.2014.08.050
177
Molaei A. Lakzian A. Haghnia G. Astaraei A. Rasouli-Sadaghiani M. Ceccherini M. T. et al . (2017). Assessment of some cultural experimental methods to study the effects of antibiotics on microbial activities in a soil: an incubation study. PLoS ONE12:e0180663. 10.1371/journal.pone.0180663
178
Mulla S. I. Hu A. Sun Q. Li J. Suanon F. Ashfaq M. et al . (2018). Biodegradation of sulfamethoxazole in bacteria from three different origins. J. Environ. Manage. 206, 93–102. 10.1016/1.jenvman.2017.10.029
179
Müller A. K. Westergaard K. Christensen S. Sørensen S. J. (2002). The diversity and function of soil microbial communities exposed to different disturbances. Microb. Ecol. 44, 49–58. 10.1007/s00248-001-0042-8
180
Muñoz I. Gómez-Ramos M. J. Agüera A. Fernández-Alba A. R. García-Reyes J. F. Molina-Díaz A. (2009). Chemical evaluation of contaminants in wastewater effluents and the environmental risk of reusing effluents in agriculture. Trends Anal. Chem. 28, 676–694. 10.1016/j.trac.2009.03.007
181
Nelson K. L. Brözel V. S. Gibson S. A. Thaler R. Clay S. A. (2011). Influence of manure from pigs fed chlortetracycline as growth promotant on soil microbial community structure. World, J. Microbiol. Biotechnol. 27, 659–668. 10.1007/s11274-010-0504-6
182
Nõlvak H. Truu M. Kanger K. Tampere M. Espenberg M. Loit E. et al . (2016). Inorganic and organic fertilizers impact the abundance and proportion of antibiotic resistance and integron-integrase genes in agricultural grassland soil. Sci. Total Environ. 562, 678–689. 10.1016/j.scitotenv.2016.04.035
183
Nowara A. Burhenne J. Spiteller M. (1997). Binding of fluoroquinolone carboxylic acid derivatives to clay minerals. J. Agric. Food Chem. 45, 1459–1463.
184
Ollivier J. Kleineidam K. Reichel R. Thiele-Bruhn S. Kotzerke A. Kindler R. et al . (2010). Effect of sulfadiazine-contaminated pig manure on the abundances of genes and transcripts involved in nitrogen transformation in the root-rhizosphere complexes of maize and clover. Appl. Environ. Microbiol. 76, 7903–7909. 10.1128/AEM.01252-10
185
Orlewska K. Markowicz A. Piotrowska-Seget Z. Smolen-Dzirba J. Cycoń M. (2018a). Functional diversity of soil microbial communities in response to the application of cefuroxime and/or antibiotic-resistant Pseudomonas putida strain MC1. Sustain10:3549. 10.3390/su10103549
186
Orlewska K. Piotrowska-Seget Z. Bratosiewicz-Wasik J. Cycoń M. (2018b). Characterization of bacterial diversity in soil contaminated with the macrolide antibiotic erythromycin and/or inoculated with a multidrug-resistant Raoultella sp. strain using the PCR-DGGE approach. Appl. Soil Ecol. 126, 57–64. 10.1016/j.apsoil.2018.02.019
187
Orlewska K. Piotrowska-Seget Z. Cycoń M. (2018c). Use of the PCR-DGGE method for the analysis of the bacterial community structure in soil treated with the cephalosporin antibiotic cefuroxime and/or inoculated with a multidrug-resistant Pseudomonas putida strain MC1. Front. Microbiol.9:1387. 10.3389/fmicb.2018.01387
188
Pan M. Chu L. M. (2016). Adsorption and degradation of five selected antibiotics in agricultural soil. Sci. Total Environ.545–546, 48–56. 10.1016/j.scitotenv.2015.12.040
189
Pan M. Chu L. M. (2017a). Leaching behavior of veterinary antibiotics in animal manure-applied soils. Sci. Total Environ. 579, 466–473. 10.1016/j.scitotenv.2016.11.072
190
Pan M. Chu L. M. (2017b). Fate of antibiotics in soil and their uptake by edible crops. Sci. Total Environ.599–600, 500–512. 10.1016/j.scitotenv.2017.04.214
191
Pan M. Wong C. K. C. Chu L. M. (2014). Distribution of antibiotics in wastewater-irrigated soils and their accumulation in vegetable crops in the Pearl River Delta, Southern China. J. Agric. Food Chem. 62, 11062–11069. 10.1021/jf503850v
192
Pan X. Qiang Z. Ben W. Chen M. (2011). Residual veterinary antibiotics in swine manure from concentrated animal feeding operations in Shandong Province, China. Chemosphere84, 695–700. 10.1016/j.chemosphere.2011.03.022
193
Park J. Y. Huwe B. (2016). Effect of pH and soil structure on transport of sulfonamide antibiotics in agricultural soils. Environ. Pollut. 213, 561–570. 10.1016/j.envpol.2016.01.089
194
Park S. Choi K. (2008). Hazard assessment of commonly used agricultural antibiotics on aquatic ecosystems. Ecotoxicology17, 526–538. 10.1007/s10646-008-0209-x
195
Perry J. A. Westman E. L. Wright G. D. (2014). The antibiotic resistome: what's new?Curr. Opin. Microbiol. 21, 45–50. 10.1016/j.mib.2014.09.002
196
Pinna M. V. Castaldi P. Deiana P. Pusino A. Garau G. (2012). Sorption behavior of sulfamethazine on unamended and manure-amended soils and short-term impact on soil microbial community. Ecotoxicol. Environ. Saf. 84, 234–242. 10.1016/j.ecoenv.2012.07.006
197
Pino-Otín M. R. Muñiz S. Val J. Navarro E. (2017). Effects of 18 pharmaceuticals on the physiological diversity of edaphic microorganisms. Sci. Total Environ. 595, 441–450. 10.1016/j.scitotenv.2017.04.002
198
Popowska M. Rzeczycka M. Miernik A. Krawczyk-Balska A. Walsh F. Duffy B. (2012). Influence of soil use on prevalence of tetracyclinestreptomycin, and erythromycin resistance and associated resistance genes. Antimicrob. Agents Chemother. 56, 1434–1443. 10.1128/AAC.05766-11
199
Rabølle M. Spliid N. H. (2000). Sorption and mobility of metronidazole, olaquindox, oxytetracycline and tylosin in soil. Chemosphere40, 715–722. 10.1016/S0045-6535(99)00442-7
200
Rafii F. Williams A. J. Park M. Sims L. M. Heinze T. M. Cerniglia C. E. et al . (2009). Isolation of bacterial strains from bovine fecal microflora capable of degradation of ceftiofur. Vet. Microbiol. 139, 89–96. 10.1016/j.vetmic.2009.04.023
201
Reichel R. Michelini L. Ghisi R. Thiele-Bruhn S. (2015). Soil bacterial community response to sulfadiazine in the soil-root zone. J. Plant Nutr. Soil Sci. 178, 499–506. 10.1002/jpln.201400352
202
Reichel R. Radl V. Rosendahl I. Albert A. Amelung W. Schloter M. et al . (2014). Soil microbial community responses to antibiotic-contaminated manure under different soil moisture regimes. Appl. Microbiol. Biotechnol. 98, 6487–6495. 10.1007/s00253-014-5717-4
203
Reichel R. Rosendahl I. Peeters E. T. H. M. Focks A. Groeneweg J. Bierl R. et al . (2013). Effects of slurry from sulfadiazine- (SDZ) and difloxacin- (DIF) medicated pigs on the structural diversity of microorganisms in bulk and rhizosphere soil. Soil Biol. Biochem. 62, 82–91. 10.1016/j.soilbio.2013.03.007
204
Rosendahl I. Siemens J. Groeneweg J. Linzbach E. Laabs V. Herrmann C. et al . (2011). Dissipation and sequestration of the veterinary antibiotic sulfadiazine and its metabolites under field conditions. Environ. Sci. Technol. 45, 5216–5222. 10.1021/es200326t
205
Rosendahl I. Siemens J. Kindler R. Groeneweg J. Zimmermann J. Czerwinski S. et al . (2012). Persistence of the fluoroquinolone antibiotic difloxacin in soil and lacking effects on nitrogen turnover. J. Environ. Qual. 41, 1275–1283. 10.2134/jeq2011.0459
206
Rutgersson C. Fick J. Marathe N. Kristiansson E. Janzon A. Angelin M. et al . (2014). Fluoroquinolones and qnr genes in sediment, water, soil, and human fecal flora in an environment polluted by manufacturing discharges. Environ. Sci. Technol. 48, 7825–7832. 10.1021/es501452a
207
Sandegren L. (2014). Selection of antibiotic resistance at very low antibiotic concentrations. Upsala J. Med. Sci. 119, 103–110. 10.3109/03009734.2014.90445
208
Sarmah A. K. Meyer M. T. Boxall A. B. A. (2006). A global perspective on the use, sales, exposure pathways, occurrence, fate and effects of veterinary antibiotics (VAs) in the environment. Chemosphere65, 725–759. 10.1016/j.chemosphere.2006.03.026
209
Sassman S. A. Lee L. S. (2007). Sorption and degradation in soils of veterinary ionophore antibiotics: Monensin and lasalocid. Environ. Toxicol. Chem. 26, 1614–1621. 10.1897/07-073R.1
210
Schauss K. Focks A. Heuer H. Kotzerke A. Schmitt H. Thiele-Bruhn S. et al . (2009). Analysis, fate and effects of the antibiotic sulfadiazine in soil ecosystems. TrAC - Trends Anal. Chem. 28, 612–618. 10.1016/j.trac.2009.02.009
211
Schlüsener M. P. Von Arb M. A. Bester K. (2006). Elimination of macrolides, tiamulin, and salinomycin during manure storage. Arch. Environ. Contam. Toxicol. 51, 21–28. 10.1007/s00244-004-0240-8
212
Schmitt H. Haapakangas H. Van Beelen P. (2005). Effects of antibiotics on soil microorganisms: time and nutrients influence pollution-induced community tolerance. Soil Biol. Biochem. 37, 1882–1892. 10.1016/j.soilbio.2005.02.022
213
Sengupta S. Chattopadhyay M. K. Grossart H-P. (2013). The multifaceted roles of antibiotics and antibiotic resistance in nature. Front. Microbiol. 4:47. 10.3389/fmicb.2013.00047
214
Shade A. Klimowicz A. K. Spear R. N. Linske M. Donato J. J. Hogan C. S. et al . (2013). Streptomycin application has no detectable effect on bacterial community structure in apple orchard soil. Appl. Environ. Microbiol. 79, 6617–6625. 10.1128/AEM.02017-13
215
Shi Y. Gao L. Li W. Liu J. Cai Y. (2012). Investigation of fluoroquinolones, sulfonamides and macrolides in long-term wastewater irrigation soil in Tianjin, China. Bull. Environ. Contam. Toxicol. 89, 857–861. 10.1007/s00128-012-0761-1
216
Singer H. P. Wossner A. E. McArdell C. S. Fenner K. (2016). Rapid screening for exposure to “non-target” pharmaceuticals from wastewater effluents by combining HRMS-based suspect screening and exposure modeling. Environ. Sci. Technol. 50, 6698–6707. 10.1021/acs.est.5b03332
217
Sittig S. Kasteel R. Groeneweg J. Hofmann D. Thiele B. Köppchen S. et al . (2014). Dynamics of transformation of the veterinary antibiotic sulfadiazine in two soils. Chemosphere95, 470–477. 10.1016/j.chemosphere.2013.09.100
218
Srinivasan P. Sarmah A. K. (2014). Dissipation of sulfamethoxazole in pasture soils as affected by soil and environmental factors. Sci. Total Environ.479–480, 284–291. 10.1016/j.scitotenv.2014.02.014
219
Stoob K. Singer H. P. Mueller S. R. Schwarzenbach R. P. Stamm C. H. (2007). Dissipation and transport of veterinary sulfonamide antibiotics after manure application to grassland in a small catchment. Environ. Sci. Technol. 41, 7349–7355. 10.1021/es070840e
220
Su J. Q. Wei B. Xu C. Y. Qiao M. Zhu Y. G. (2014). Functional metagenomic characterization of antibiotic resistance genes in agricultural soils from China. Environ. Int. 65, 9–15. 10.1016/j.envint.2013.12.010
221
Sukul P. Lamshöft M. Zühlke S. Spiteller M. (2008). Sorption and desorption of sulfadiazine in soil and soil-manure systems. Chemosphere73, 1344–1350. 10.1016/j.chemosphere.2008.06.066
222
Sułowicz S. Piotrowska-Seget Z. (2016). Response of microbial communities from an apple orchard and grassland soils to the first-time application of the fungicide tetraconazole. Ecotoxicol. Environ. Saf. 124, 193–201. 10.1016/j.ecoenv.2015.10.025
223
Tabatabai M. A. Bremner J. M. (1969). Use of p-nitrophenyl phosphate for assay of soil phosphatase activity. Soil Biol. Biochem. 1, 301–307.
224
Tang X. Lou C. Wang S. Lu Y. Liu M. Hashmi M. Z. et al . (2015). Effects of long-term manure applications on the occurrence of antibiotics and antibiotic resistance genes (ARGs) in paddy soils: evidence from four field experiments in south of China. Soil Biol. Biochem. 90, 179–187. 10.1016/j.soilbio.2015.07.027
225
Tasho R. P. Cho J. Y. (2016). Veterinary antibiotics in animal waste, its distribution in soil and uptake by plants: a review. Sci. Total Environ.563–564, 366–376. 10.1016/j.scitotenv.2016.04.140
226
Teixidó M. Granados M. Prat M. D. Beltrán J. L. (2012). Sorption of tetracyclines onto natural soils: data analysis and prediction. Environ. Sci. Pollut. Res. 19, 3087–3095. 10.1007/s11356-012-0954-5
227
Thiele S. (2000). Adsorption of the antibiotic pharmaceutical compound sulfapyridine by a long-term differently fertilized loess Chernozem. J. Plant Nutr. Soil Sci. 163, 589–594. 10.1002/1522-2624(200012)163:6<589::AID-JPLN589>3.0.CO;2-5
228
Thiele-Bruhn S. (2003). Pharmaceutical antibiotic compounds in soils - a review. J. Plant Nutr. Soil Sci. 166, 145–167. 10.1002/jpln.200390023
229
Thiele-Bruhn S. (2005). Microbial inhibition by pharmaceutical antibiotics in different soils - Dose-response relations determined with the iron(III) reduction test. Environ. Toxicol. Chem. 24, 869–876. 10.1897/04-166R.1
230
Thiele-Bruhn S. Beck I.-C. (2005). Effects of sulfonamide and tetracycline antibiotics on soil microbial activity and microbial biomass. Chemosphere59, 457–465. 10.1016/j.chemosphere.2005.01.023
231
Thiele-Bruhn S. Peters D. (2007). Photodegradation of pharmaceutical antibiotics on slurry and soil surfaces. Landbauforsch. Volkenrode57, 13–23.
232
Thiele-Bruhn S. Seibicke T. Schulten H.-R. Leinweber P. (2004). Sorption of sulfonamide pharmaceutical antibiotics on whole soils and particle-size fractions. J. Environ. Qual. 33, 1331–1342.
233
Tolls J. (2001). Sorption of veterinary pharmaceuticals in soils: a review. Environ. Sci. Technol. 35, 3397–3406. 10.1021/es0003021
234
Topp E. Chapman R. Devers-Lamrani M. Hartmann A. Marti R. Martin-Laurent F. et al . (2013). Accelerated biodegradation of veterinary antibiotics in agricultural soil following long-term exposure, and isolation of a sulfamethazinedegrading microbacterium sp. J. Environ. Qual. 42, 173–178. 10.2134/jeq2012.0162
235
Topp E. Renaud J. Sumarah M. Sabourin L. (2016). Reduced persistence of the macrolide antibiotics erythromycin, clarithromycin and azithromycin in agricultural soil following several years of exposure in the field. Sci. Total Environ. 562, 136–144. 10.1016/j.scitotenv.2016.03.210
236
Torres-Cortés G. Millán V. Ramírez-Saad H. C. Nisa-Martínez R. Toro N. Martínez-Abarca F. (2011). Characterization of novel antibiotic resistance genes identified by functional metagenomics on soil samples. Environ. Microbiol.13, 1101–1114. 10.1111/j.1462-2920.2010.02422.x
237
Toth J. D. Feng Y. Dou Z. (2011). Veterinary antibiotics at environmentally relevant concentrations inhibit soil iron reduction and nitrification. Soil Biol. Biochem. 43, 2470–2472. 10.1016/j.soilbio.2011.09.004
238
Udikovic-Kolic N. Wichmann F. Broderick N. A. Handelsman J. (2014). Bloom of resident antibiotic-resistant bacteria in soil following manure fertilization. Proc. Natl. Acad. Sci. U.S.A. 111, 15202–15207. 10.1073/pnas.1409836111
239
Unger I. M. Goyne K. W. Kennedy A. C. Kremer R. J. McLain J. E. T. Williams C. F. (2013). Antibiotic effects on microbial community characteristics in soils under conservation management practices. Soil Sci. Soc. Am. J. 77, 100–112. 10.2136/sssaj2012.0099
240
US EPA (2009). Targeted National Sewage Sludge Survey. EPA-822-R-08-018. Washington, DC: U.S. Environmental Protection Agency.
241
Van Doorslaer X. Dewulf J. Van Langenhove H. Demeestere K. (2014). Fluoroquinolone antibiotics: an emerging class of environmental micropollutants. Sci. Total Environ.500–501, 250–269. 10.1016/j.scitotenv.2014.08.075
242
Van Goethem M. W. Pierneef R. Bezuidt O. K. I. Van De Peer Y. Cowan D. A. Makhalanyane T. P. (2018). A reservoir of “historical” antibiotic resistance genes in remote pristine Antarctic soils. Microbiome6:40. 10.1186/s40168-018-0424-5
243
Varma A. Buscot F. (2005). Microorganisms in Soils: Roles in Genesis and Functions. Berlin Heidelberg: Springer-Verlag.
244
Vaz S. Lopes W. T. Martin-Neto L. (2015). Study of molecular interactions between humic acid from Brazilian soil and the antibiotic oxytetracycline. Environ. Technol. Innov. 4, 260–267. 10.1016/j.eti.2015.09.004
245
Vazquez-Roig P. Segarra R. Blasco C. Andreu V. Picó Y. (2010). Determination of pharmaceuticals in soils and sediments by pressurized liquid extraction and liquid chromatography tandem mass spectrometry. J. Chromatogr. A1217, 2471–2483. 10.1016/j.chroma.2009.11.033
246
Walters E. McClellan K. Halden R. U. (2010). Occurrence and loss over three years of 72 pharmaceuticals and personal care products from biosolids-soil mixtures in outdoor mesocosms. Water Res. 44, 6011–6020. 10.1016/j.watres.2010.07.051
247
Wang J. Lin H. Sun W. Xia Y. Ma J. Fu J. et al . (2016). Variations in the fate and biological effects of sulfamethoxazole, norfloxacin and doxycycline in different vegetable-soil systems following manure application. J. Hazard. Mater. 304, 49–57. 10.1016/j.jhazmat.2015.10.038
248
Wang Q. Guo M. Yates S. R. (2006). Degradation kinetics of manure-derived sulfadimethoxine in amended soil. J. Agric. Food Chem. 54, 157–163. 10.1021/jf052216w
249
Wang Q. Yates S. R. (2008). Laboratory study of oxytetracycline degradation kinetics in animal manure and soil. J. Agric. Food Chem. 56, 1683–1688. 10.1021/jf072927p
250
Wang S. Gao X. Gao Y. Li Y. Cao M. Xi Z. et al . (2017). Tetracycline resistance genes identified from distinct soil environments in China by functional metagenomics. Front. Microbiol. 8:1406. 10.3389/fmicb.2017.01406
251
Wang S. Wang H. (2015). Adsorption behavior of antibiotic in soil environment: a critical review. Front. Environ. Sci. Eng. 9, 565–574. 10.1007/s11783-015-0801-2
252
Watanabe N. Bergamaschi B. A. Loftin K. A. Meyer M. T. Harter T. (2010). Use and environmental occurrence of antibiotics in freestall dairy farms with manured forage fields. Environ. Sci. Technol. 44, 6591–6600. 10.1021/es100834s
253
Weerasinghe C. A. Towner D. (1997). Aerobic biodegradation of virginiamycin in soil. Environ. Toxicol. Chem. 16, 1873–1876. 10.1002/etc.5620160916
254
Wegst-Uhrich S. R. Navarro D. A. G. Zimmerman L. Aga D. S. (2014). Assessing antibiotic sorption in soil: a literature review and new case studies on sulfonamides and macrolides. Chem. Cent. J. 8:5. 10.1186/1752-153X-8-5
255
Wei X. Wu S. C. Nie X. P. Yediler A. Wong M. H. (2009). The effects of residual tetracycline on soil enzymatic activities and plant growth. J. Environ. Sci. Heal.44, 461–471. 10.1080/03601230902935139
256
Wellington E. M. H. Boxall A. B. A. Cross P. Feil E. J. Gaze W. H. Hawkey P. M. et al . (2013). The role of the natural environment in the emergence of antibiotic resistance in Gram-negative bacteria. Lancet Infect. Dis. 13, 155–165. 10.1016/S1473-3099(12)70317-1
257
Wen X. Wang Y. Zou Y. Ma B. Wu Y. (2018). No evidential correlation between veterinary antibiotic degradation ability and resistance genes in microorganisms during the biodegradation of doxycycline. Ecotoxicol. Environ. Saf. 147, 759–766. 10.1016/j.ecoenv.2017.09.025
258
Wepking C. Avera B. Badgley B. Barrett J. E. Franklin J. Knowlton K. F. et al . (2017). Exposure to dairy manure leads to greater antibiotic resistance and increased mass-specific respiration in soil microbial communities. Proc. R. Soc. B Biol. Sci. 284, 20162233. 10.1098/rspb.2016.2233
259
Westergaard K. Müller A. K. Christensen S. Bloem J. Sørensen S. J. (2001). Effects of tylosin as a disturbance on the soil microbial community. Soil Biol. Biochem. 33, 2061–2071. 10.1016/S0038-0717(01)00134-1
260
Wu L. Pan X. Chen L. Huang Y. Teng Y. Luo Y. et al . (2013). Occurrence and distribution of heavy metals and tetracyclines in agricultural soils after typical land use change in east China. Environ. Sci. Pollut. Res. 20, 8342–8354. 10.1007/s11356-013-1532-1
261
Wu X.-L. Xiang L. Yan Q.-Y. Jiang Y.-N. Li Y.-W. Huang X.-P. et al . (2014). Distribution and risk assessment of quinolone antibiotics in the soils from organic vegetable farms of a subtropical city, Southern China. Sci. Total Environ. 487, 399–406. 10.1016/j.scitotenv.2014.04.015
262
Xiao K.-Q. Li B. Ma L. Bao P. Zhou X. Zhang T. et al . (2016). Metagenomic profiles of antibiotic resistance genes in paddy soils from South China. FEMS Microbiol. Ecol.92:fiw023. 10.1093/femsec/fiw023
263
Xin Z. Fengwei T. Gang W. Xiaoming L. Qiuxiang Z. Hao Z. et al . (2012). Isolation, identification and characterization of human intestinal bacteria with the ability to utilize chloramphenicol as the sole source of carbon and energy. FEMS Microbiol. Ecol. 82, 703–712. 10.1111/j.1574-6941.2012.01440.x
264
Xiong W. Wang M. Dai J. Sun Y. Zeng Z. (2018). Application of manure containing tetracyclines slowed down the dissipation of tet resistance genes and caused changes in the composition of soil bacteria. Ecotoxicol. Environ. Saf. 147, 455–460. 10.1016/j.ecoenv.2017.08.061
265
Xu Y. Yu W. Ma Q. Wang J. Zhou H. Jiang C. (2016). The combined effect of sulfadiazine and copper on soil microbial activity and community structure. Ecotoxicol. Environ. Saf. 134, 43–52. 10.1016/j.ecoenv.2016.06.041
266
Yang J.-F. Ying G.-G. Liu S. Zhou L.-J. Zhao J.-L. Tao R. et al . (2012). Biological degradation and microbial function effect of norfloxacin in a soil under different conditions. J. Environ. Sci. Heal.47, 288–295. 10.1080/03601234.2012.638886
267
Yang J.-F. Ying G.-G. Zhao J.-L. Tao R. Su H.-C. Chen F. (2010). Simultaneous determination of four classes of antibiotics in sediments of the Pearl Rivers using RRLC-MS/MS. Sci. Total Environ. 408, 3424–3432. 10.1016/j.scitotenv.2010.03.049
268
Yang J.-F. Ying G.-G. Zhou L.-J. Liu S. Zhao J.-L. (2009). Dissipation of oxytetracycline in soils under different redox conditions. Environ. Pollut. 157, 2704–2709. 10.1016/j.envpol.2009.04.031
269
Yang Y. Owino A. A. Gao Y. Yan X. Xu C. Wang J. (2016). Occurrence, composition and risk assessment of antibiotics in soils from Kenya, Africa. Ecotoxicology25, 1194–1201. 10.1007/s10646-016-1673-3
270
Ye M. Sun M. Feng Y. Wan J. Xie S. Tian D. et al . (2016). Effect of biochar amendment on the control of soil sulfonamides, antibiotic-resistant bacteria, and gene enrichment in lettuce tissues. J. Hazard. Mater. 309, 219–227. 10.1016/j.jhazmat.2015.10.074
271
Zak J. C. Willing M. R. Moorhead D. L. Wildman H. G. (1994). Functional diversity of microbial communities: a quantitative approach. Soil Biol. Biochem. 26, 1101–1108. 10.1016/0038-0717(94)90131-7
272
Zhang H. Zhou Y. Huang Y. Wu L. Liu X. Luo Y. (2016a). Residues and risks of veterinary antibiotics in protected vegetable soils following application of different manures. Chemosphere152, 229–237. 10.1016/j.chemosphere.2016.02.111
273
Zhang J. Wei Y. Chen M. Tong J. Xiong J. Ao Z. (2015). Occurrence and fate of antibiotic and heavy metal resistance genes in the total process of biological treatment and land application of animal manure: a review. Huanjing Kexue Xuebao/Acta Sci. Circum.35, 935–946. 10.13671/j.hjkxxb.2014.0843
274
Zhang J. Q. Dong Y. H. (2008). Effect of low-molecular-weight organic acids on the adsorption of norfloxacin in typical variable charge soils of China. J. Hazard. Mater. 151, 833–839. 10.1016/j.jhazmat.2007.11.046
275
Zhang Q. Dick W. A. (2014). Growth of soil bacteria, on penicillin and neomycin, not previously exposed to these antibiotics. Sci. Total Environ. 493, 445–453. 10.1016/j.scitotenv.2014.05.114
276
Zhang Q. Shamsi I. H. Zhou H. Wei L. Zhu Z. Adil M. F. et al . (2016b). Soil microbial communities and mineralization responses to penicillin and tetracycline loads. Res. Rev. J. Microbiol. Biotechnol. 5, 7–15.
277
Zhang S. Yang G. Hou S. Zhang T. Li Z. Liang F. (2018). Distribution of ARGs and MGEs among glacial soil, permafrost, and sediment using metagenomic analysis. Environ. Pollut. 234, 339–346. 10.1016/j.envpol.2017.11.031
278
Zhang W. Qiu L. Gong A. Yuan X. (2017b). Isolation and characterization of a high-efficiency erythromycin A-degrading Ochrobactrum sp. strain. Mar. Pollut. Bull. 114, 896–902. 10.1016/j.marpolbul.2016.10.076
279
Zhang W.-W. Wen Y.-Y. Niu Z.-L. Yin K. Xu D.-X. Chen L.-X. (2012). Isolation and characterization of sulfonamide-degrading bacteria Escherichia sp. HS21 and Acinetobacter sp. HS51. World J. Microbiol. Biotechnol.28, 447–452. 10.1007/s11274-011-0834-z
280
Zhang Y. Hu S. Zhang H. Shen G. Yuan Z. Zhang W. (2017a). Degradation kinetics and mechanism of sulfadiazine and sulfamethoxazole in an agricultural soil system with manure application. Sci. Total Environ.607–608, 1348–1356. 10.1016/j.scitotenv.2017.07.083
281
Zhao L. Dong Y. H. Wang H. (2010). Residues of veterinary antibiotics in manures from feedlot livestock in eight provinces of China. Sci. Total Environ. 408, 1069–1075. 10.1016/j.scitotenv.2009.11.014
282
Zhou B. Wang C. Zhao Q. Wang Y. Huo M. Wang J. et al . (2016). Prevalence and dissemination of antibiotic resistance genes and coselection of heavy metals in Chinese dairy farms. J. Hazard. Mater.320, 10–17. 10.1016/j.jhazmat.2016.08.007
283
Zhou L.-J. Ying G.-G. Liu S. Zhang R.-Q. Lai H.-J. Chen Z.-F. et al . (2013a). Excretion masses and environmental occurrence of antibiotics in typical swine and dairy cattle farms in China. Sci. Total Environ. 444, 183–195. 10.1016/j.scitotenv.2012.11.087
284
Zhou L.-J. Ying G.-G. Liu S. Zhao J.-L. Yang B. Chen Z.-F. et al . (2013b). Occurrence and fate of eleven classes of antibiotics in two typical wastewater treatment plants in South China. Sci. Total Environ.452–453, 365–376. 10.1016/j.scitotenv.2013.03.010
285
Zhou L.-J. Ying G.-G. Zhao J.-L. Yang J.-F. Wang L. Yang B. et al . (2011). Trends in the occurrence of human and veterinary antibiotics in the sediments of the Yellow River, Hai River and Liao River in northern China. Environ. Pollut. 159, 1877–1885. 10.1016/j.envpol.2011.03.034
286
Zhou Y. Niu L. Zhu S. Lu H. Liu W. (2017). Occurrence, abundance, and distribution of sulfonamide and tetracycline resistance genes in agricultural soils across China. Sci. Total Environ. 599–600, 1977–1983. 10.1016/j.scitotenv.2017.05.152
287
Zielezny Y. Groeneweg J. Vereecken H. Tappe W. (2006). Impact of sulfadiazine and chlorotetracycline on soil bacterial community structure and respiratory activity. Soil Biol. Biochem. 38, 2372–2380. 10.1016/j.soilbio.2006.01.031
288
ŽiŽek S. Dobeic M. Pintarič Š. Zidar P. Kobal S. Vidrih M. (2015). Degradation and dissipation of the veterinary ionophore lasalocid in manure and soil. Chemosphere138, 947–951. 10.1016/j.chemosphere.2014.12.032
Summary
Keywords
antibiotics, degradation, DT50, microbial activities, microbial community structure, antibiotic resistance genes, metagenomics, soil
Citation
Cycoń M, Mrozik A and Piotrowska-Seget Z (2019) Antibiotics in the Soil Environment—Degradation and Their Impact on Microbial Activity and Diversity. Front. Microbiol. 10:338. doi: 10.3389/fmicb.2019.00338
Received
05 July 2018
Accepted
08 February 2019
Published
08 March 2019
Volume
10 - 2019
Edited by
Matthias E. Kaestner, Helmholtz Centre for Environmental Research (UFZ), Germany
Reviewed by
Ulisses Nunes da Rocha, Helmholtz Centre for Environmental Research (UFZ), Germany; Jan Roelof Van Der Meer, Université de Lausanne, Switzerland
Updates
Copyright
© 2019 Cycoń, Mrozik and Piotrowska-Seget.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mariusz Cycoń mcycon@sum.edu.pl
This article was submitted to Microbiotechnology, Ecotoxicology and Bioremediation, a section of the journal Frontiers in Microbiology
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.