- 1Department of Food Science, Cornell University, Ithaca, NY, United States
- 2Department of Food Science, Penn State University, University Park, PA, United States
- 3Department of Food Science, Wageningen University, Wageningen, Netherlands
A Corrigendum on
Genes Associated With Psychrotolerant Bacillus cereus Group Isolates
by Beno, S. M., Orsi, R. H., Cheng, R. A., Kent, D. J., Kovac, J., Duncan, D. R., et al. (2019). Front. Microbiol. 10:662. doi: 10.3389/fmicb.2019.00662
In the original article, there were two minor mistakes in Figure 3 as published. The isolate FSL H8-0534 was mistakenly labeled as a clade VI isolate. We have updated the figure to demonstrate that this isolate is a clade I isolate. Additionally, isolate FSL M7-1219 had an additional “0” in the number sequence. The corrected Figure 3 appears below.
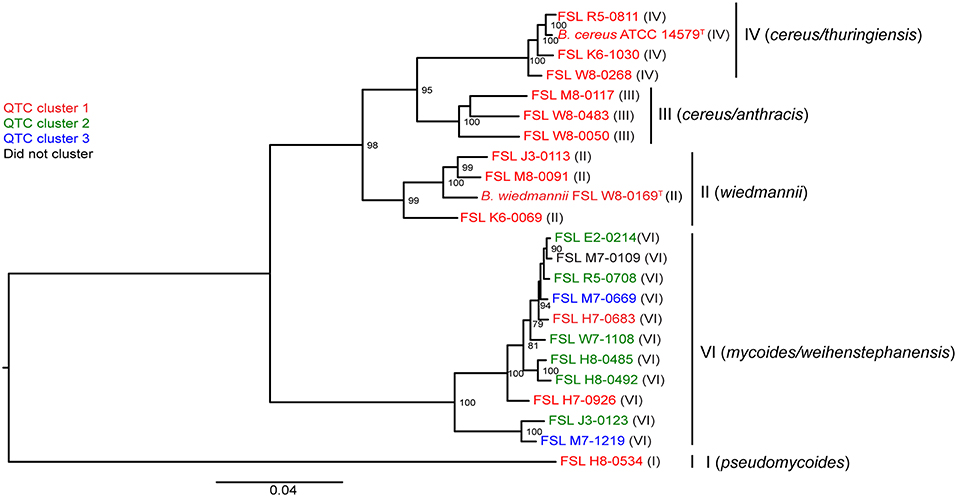
Figure 3. Phylogenetic tree constructed from the core SNPs identified in the genomes of 23 B. cereus isolates. The maximum likelihood tree was constructed using a general time-reversible (GTR) model with gamma-distributed sites and 1,000 bootstrap repetitions. Roman numerals in parentheses represent the phylogenetic clade of the isolate, as defined previously (Kovac et al., 2016). QTC clusters representing isolates with similar growth patterns in BHI and SMB at 6°C (see Figure 3A) are mapped onto the phylogenetic tree. Isolates shown in red represent QTC cluster 1 isolates (non-psychrotolerant in BHI broth or SMB). Isolates shown in green represent QTC cluster 2 isolates (psychrotolerant in BHI broth but not SMB), and isolates shown in blue represent clade 3 isolates (psychrotolerant in both BHI broth and SMB). One isolate (FSL M7-0109) did not cluster with other isolates, and is therefore shown in black font. Numbers at branch points represent bootstrap values; only bootstrap values >70 are shown.
The authors apologize for these errors and state that this does not change the scientific conclusions of the article in any way. The original article has been updated.
References
Keywords: Bacillus cereus, whole genome sequencing, psychrotolerant, spoilage, skim milk broth
Citation: Beno SM, Orsi RH, Cheng RA, Kent DJ, Kovac J, Duncan DR, Martin NH and Wiedmann M (2019) Corrigendum: Genes Associated With Psychrotolerant Bacillus cereus Group Isolates. Front. Microbiol. 10:1323. doi: 10.3389/fmicb.2019.01323
Received: 17 April 2019; Accepted: 28 May 2019;
Published: 12 June 2019.
Edited by:
Giovanna Suzzi, University of Teramo, ItalyReviewed by:
Rosanna Tofalo, University of Teramo, ItalyCopyright © 2019 Beno, Orsi, Cheng, Kent, Kovac, Duncan, Martin and Wiedmann. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Martin Wiedmann, bXcxNkBjb3JuZWxsLmVkdQ==
†Present Address: Sarah M. Beno, Department of Cell Differential and Integrative Biology, The University of Alabama, Birmingham, AB, United States
 Sarah M. Beno
Sarah M. Beno Renato H. Orsi
Renato H. Orsi Rachel A. Cheng
Rachel A. Cheng David J. Kent
David J. Kent Jasna Kovac
Jasna Kovac Diana R. Duncan
Diana R. Duncan Nicole H. Martin
Nicole H. Martin Martin Wiedmann
Martin Wiedmann