- 1Department of Microbiology, University of Georgia, Athens, GA, United States
- 2Department of Biological Sciences, University of Alberta, Edmonton, AB, Canada
- 3Complex Carbohydrate Research Center, University of Georgia, Athens, GA, United States
Campylobacter jejuni is the leading bacterial cause of acute gastroenteritis worldwide and thus significant to public health. C. jejuni primarily lives in the gastrointestinal tracts of poultry and can contaminate meat during processing. Despite a small genome, the metabolic plasticity of C. jejuni allows proliferation in chicken ceca and mammalian host intestines, and survival in environments with a variety of temperatures, pH, osmotic conditions, and nutrient availabilities. The exact mechanism of C. jejuni infection is unknown, however, virulence requires motility. Our data suggest the C. jejuni RidA homolog, Cj1388, plays a role in flagellar biosynthesis, regulation, structure, and/or function and, as such is expected to influence virulence of the organism. Mutants lacking cj1388 have defects in motility, autoagglutination, and phage infectivity under the conditions tested. Comparison to the RidA paradigm from Salmonella enterica indicates the phenotypes of the C. jejuni cj1388 mutant are likely due to the inhibition of one or more pyridoxal 5′-phosphate-dependent enzymes by the reactive enamine 2-aminoacrylate.
Introduction
The Rid/YER057c/UK114 protein superfamily (COG0251) is broadly conserved throughout all domains of life (Kim et al., 2001; Leitner-Dagan et al., 2006; Lambrecht et al., 2012, 2013; Downs and Ernst, 2015; Niehaus et al., 2015; ElRamlawy et al., 2016). Based on phylogenetic analysis, the superfamily was divided into eight subfamilies; RidA, which includes homologs of the archetypical protein from Salmonella enterica, and Rid1-7, which are not well understood (Niehaus et al., 2015). Prokaryotic genomes can encode several members of the Rid1-7 subfamilies while also encoding one or more RidA proteins. In many genomes, the RidA homologs are not annotated with the ascribed biochemical function for these proteins. The RidA, r eactive i ntermediate d eaminase A, of S. enterica was found to be an enamine deaminase, and multiple homologs from the three domains of life have similar activity (Lambrecht et al., 2010; Lambrecht et al., 2012; Niehaus et al., 2015; ElRamlawy et al., 2016; Ernst and Downs, 2018). In some organisms, a cellular role for RidA involves quenching the reactive metabolite 2-aminoacrylate (2AA) to prevent damage to specific pyridoxal 5′-phosphate (PLP)-dependent enzymes (Schmitz and Downs, 2004; Christopherson et al., 2008; Flynn and Downs, 2013; Flynn et al., 2013; Lambrecht et al., 2013; Ernst et al., 2014, 2016; Niehaus et al., 2015; Irons et al., 2018). RidA homologs have a similar role in at least Escherichia coli, Pseudomonas aeruginosa, and Saccharomyces cerevisiae, although the phenotypic consequences of a ridA mutation depends on the specific metabolic network architecture of the organism (Borchert and Downs, 2017b; Ernst and Downs, 2018; Irons et al., 2018). Enzyme damage resulting from accumulated 2AA can impact growth, motility, biofilm formation, iron homeostasis, and potentially virulence.
Campylobacter jejuni NCTC 11168, a prominent diarrheal pathogen, encodes two members of the Rid superfamily, a RidA homolog (Cj1388), and a protein from the Rid2 subfamily (Cj0327). Data presented herein confirmed Cj1388 is a RidA protein and for clarity this locus is designated cjridA throughout. Previous data suggested that C. jejuni Cj1388 (CjRidA) plays a role in flagella-flagella interactions, possibly through regulation of flagellar glycan modification (Reuter et al., 2015). Additionally, CjRidA has been highlighted in several global -omics studies in C. jejuni strains 11168 and 81176 (strain specific gene designation cj1388 or cj1390, respectively) (Woodall et al., 2005; Reid et al., 2008; Taveirne et al., 2013; Clark et al., 2014; Flint et al., 2014; Guccione et al., 2017; Hao et al., 2017). These data sets suggest CjRidA could play a direct or indirect role in virulence, antibiotic resistance, acid adaptation, growth with bile salts, and hydrogen peroxide and oxygen stress. Finally, cjridA is a member of the HeuR (He me u tilization r egulator) regulon (Reuter et al., 2015; Johnson et al., 2016). HeuR is a PAS-domain containing regulator, and thought to be regulated in response to changing environmental cues (Reuter et al., 2015; Johnson et al., 2016). In other Campylobacter spp. including C. coli, C. upsaliensis, and C. lari, there is a co-occurrence of heuR and cjridA in the genome, suggesting cjridA may be regulated in response to differing environmental conditions in these organisms.
The CjRidA has been consistently misannotated as an endoribonuclease in C. jejuni studies, obscuring its likely connection to PLP-dependent enzymes and metabolism. In S. enterica, RidA catalyzes the hydrolysis of the 2AA intermediate formed by several PLP-dependent enzymes (Lambrecht et al., 2012; Ernst et al., 2014, 2016). In the absence of RidA, free 2AA accumulates and can covalently inactivate certain PLP-dependent enzymes such as serine hydroxymethyltransferase (SHMT) (GlyA; EC 2.1.2.1), alanine racemases (Alr/DadX; EC 5.1.1.1), and transaminase B (IlvE; EC 2.6.1.42), leading to defects in one-carbon unit metabolism, cell-wall synthesis, and isoleucine biosynthesis, respectively (Schmitz and Downs, 2004; Flynn et al., 2013; Ernst and Downs, 2016). The activity of the enzymes targeted by 2AA can be decreased by 30–50% in strains lacking RidA (Lambrecht et al., 2010; Ernst and Downs, 2016, 2018; Borchert and Downs, 2017b; Irons et al., 2018). To date, the diverse phenotypes of organisms lacking RidA suggest that there are additional and unknown targets of 2AA, possibly extending beyond PLP-dependent enzymes.
In at least S. enterica, E. coli, P. aeruginosa, S. cerevisiae, and Arabidopsis thaliana, a biosynthetic threonine/serine dehydratase (IlvA), acting on serine is the main source of 2AA in ridA mutants (Lambrecht et al., 2012; Niehaus et al., 2014; Borchert and Downs, 2017a; Ernst and Downs, 2018; Irons et al., 2018). In each of these organisms, IlvA has a regulatory domain and is allosterically inhibited by isoleucine (Gallagher et al., 1998; Schmitz and Downs, 2004). As a consequence, the presence of isoleucine eliminated generation of 2AA and suppressed the phenotypes of a ridA mutant. The specific targets of 2AA that result in detectable defects vary in different organisms. In S. enterica, 2AA accumulation causes a growth defect reversed by exogenous glycine; in E. coli 2AA accumulation-induced growth inhibition was reversed by exogenous aspartate, or purines; in P. aeruginosa, 2AA accumulation is detrimental to growth and partially reversed by exogenous proline and polyamines; and in S. cerevisiae mitochondrial accumulation of 2AA leads to loss of mitochondrial DNA and reduced heme biosynthesis (Flynn et al., 2013; Ernst and Downs, 2016, 2018; Borchert and Downs, 2017b; Irons et al., 2018) (Whitaker and Downs, unpublished). The diverse effects of 2AA emphasize the complexity of the metabolic network and our limited understanding of the integration between biochemical pathways.
This study was initiated to understand the physiological role of CjRidA (Cj1388) in C. jejuni 11168. Although the levels of cjridA were noted in multiple global studies, this gene was peripheral in those studies, and in many cases the results were not verified nor was the effect determined to be direct or indirect. The data herein suggest a role for CjRidA in flagellar biosynthesis, structure, glycosylation, and/or function. Further, we confirmed that CjRidA and the Rid2 subfamily member Cj0327, have enamine deaminase activity in vivo and in vitro. This work extends the list of organisms known to encode functional enamine deaminases of the Rid family.
Results
Cj1388 and Cj0327 Deaminate 2-Aminoacrylate in vivo
A S. enterica ridA mutant fails to grow on minimal medium with serine due to the accumulation of 2AA that is generated by the biosynthetic serine/threonine dehydratase encoded by ilvA (EC 4.3.1.19) (Schmitz and Downs, 2004; Lambrecht et al., 2013). A S. enterica ridA mutant was transformed with pBAD24 constructs harboring a gene encoding Rid proteins from C. jejuni (cj1388/cjridA or cj0327), the S. enterica ridA (seridA) under the control of an arabinose promoter, or an empty vector control. Growth was monitored in minimal glucose medium with 5 mM serine and the data are shown in Figure 1. Plasmids carrying either cjridA (pDM1577) or seridA (pDM1439) restored full growth to the S. enterica ridA mutant without inducing expression of the plasmid encoded genes. In contrast, cj0327 (pDM1588) partially restored growth, and only when its expression was induced with arabinose.
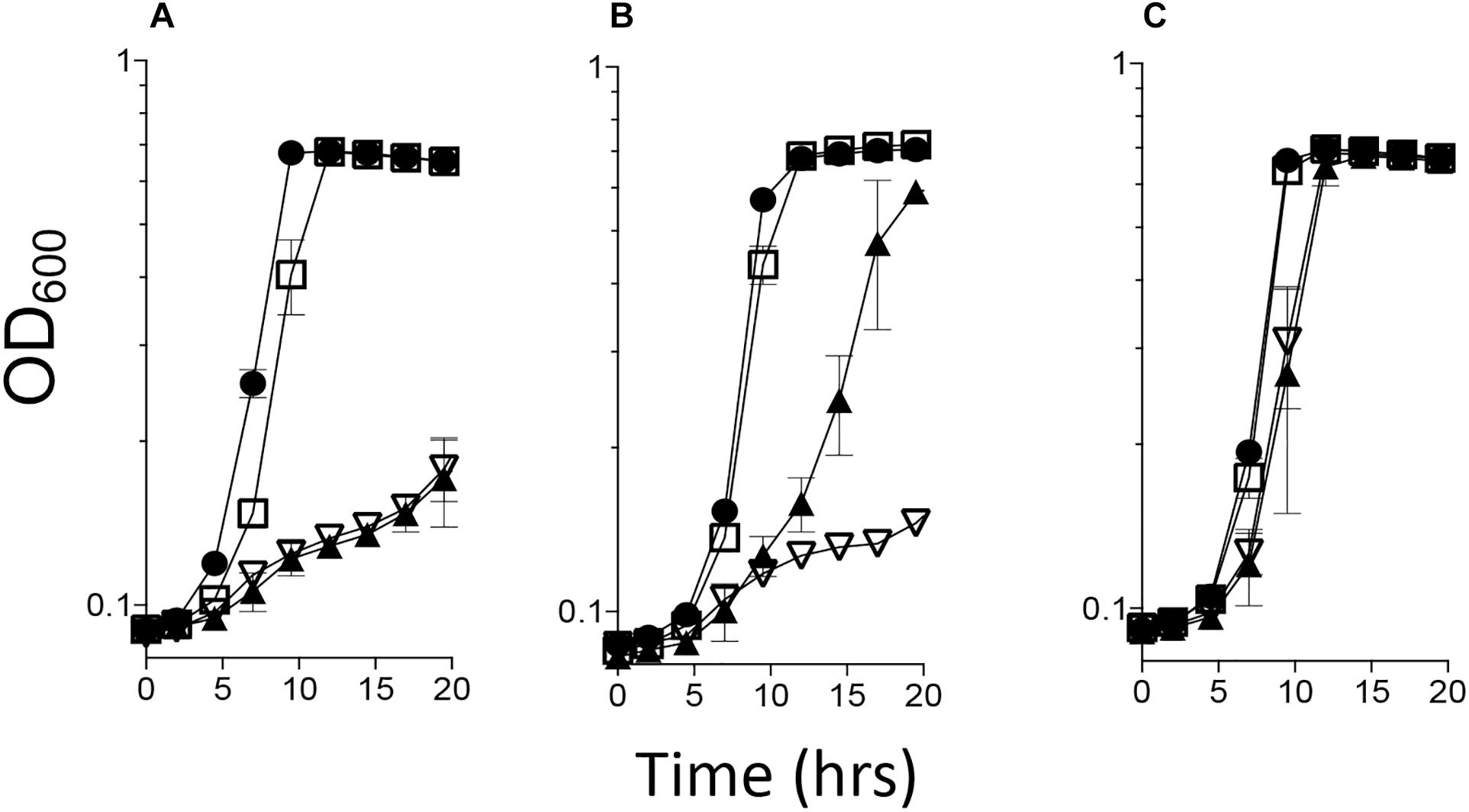
Figure 1. cjridA complements a Salmonella enterica ridA mutant. A S. enterica ridA mutant with one of four plasmids was grown in a 96-well plate at 37°C shaking in minimal glucose (11 mM) medium with: (A) serine (5 mM), (B) serine and arabinose (0.2%), or (C) serine, arabinose and isoleucine (1 mM). The S. enterica strain carried plasmids expressing S. enterica ridA (filled circles), C. jejuni ridA (cjridA, open squares), C. jejuni rid2 (cj0327, closed triangles), or empty vector (open triangles).
The inability of Cj0327, a Rid2 subfamily member, to fully complement the S. enterica ridA mutant was consistent with the results obtained for proteins annotated as Rid1-3 from P. aeruginosa, Yersinia pestis, E. coli, Acinetobacter baylyi, and Pseudomonas syringae (Hodge-Hanson and Downs, 2017; Irons et al., 2018) (Irons et al., unpublished). The partial complementation by Cj0327 and other proteins from the Rid1, 2, and 3 subfamilies suggests that these proteins may deaminate primarily non-2AA enamines in vivo. The specific physiological role of Cj0327 was not pursued further here.
Cj1388 and Cj0327 Have Deaminase Activity in vitro
L-amino acid oxidase (LOX or LAAO)-based assays were used to assess the ability of purified Cj1388 and Cj0327 to deaminate imines in vitro (Hafner and Wellner, 1979; Niehaus et al., 2014; Niehaus et al., 2015; Hodge-Hanson and Downs, 2017; Degani et al., 2018). 2-aminobutyrate was provided as substrate, resulting in the LOX-dependent formation of 2-iminobutyrate. This imine reacts with semicarbazide to produce a semicarbazone which is monitored at 248 nm. Rid proteins can compete for the imine, converting it to the ketoacid, 2-ketobutyrate, similar to a reaction RidA catalyzes in vivo. Thus in this assay, the rate of semicarbazone formation is inversely proportional to Rid activity. The rate of semicarbazone formation (μM, min–1) with the addition of Cj1388 (CjRidA) or Cj0327 is shown in Figure 2. Consistent with the in vivo complementation data in an S. enterica ridA mutant, CjRidA has greater deaminase activity than Cj0327. When the Rid proteins are provided at higher concentrations (i.e., 10 μM), semicarbazone formation is reduced to the same extent by each protein, consistent with a saturating concentration of enzyme.
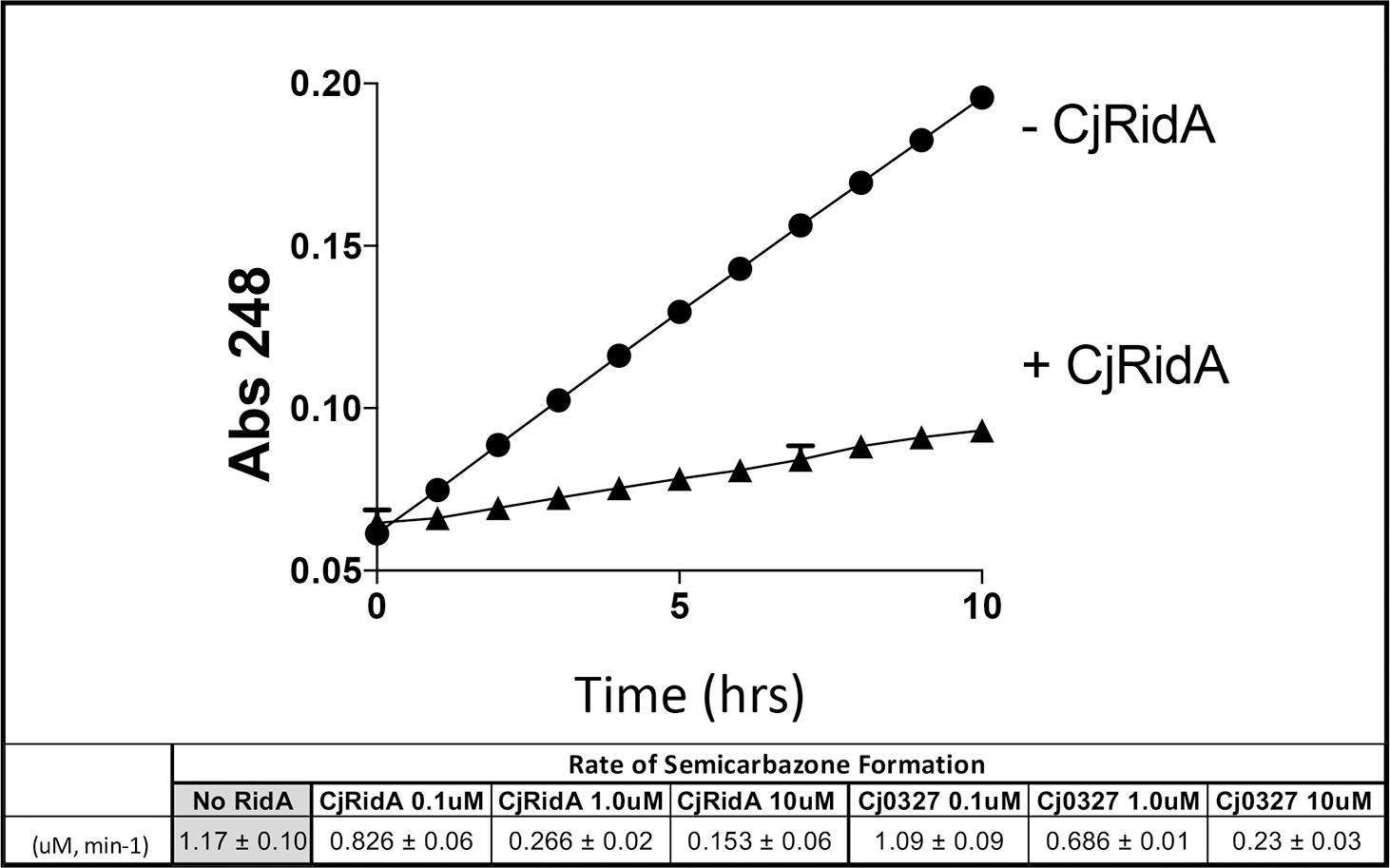
Figure 2. CjRidA and Cj0327 are imine deaminases in vitro. Each reaction mixture contained potassium pyrophosphate (50 mM, pH 8.7), neutralized semicarbazide (10 mM), bovine liver catalase (1 μg), and L-amino acid oxidase (0.5 μg) with or without the addition of CjRidA or Cj0327. 2-aminobutyrate (0.5 mM) was added to start the reaction and absorbance at 248 nm was monitored for 10 min. The graph shows the absorbance over time for reactions without CjRidA (circles) or with CjRidA at a final concentration of 1.0 μM (triangles). Error bars represent standard deviation of the mean determined from three technical triplicates by GraphPad Prism 7.0c. The molar extinction coefficient for semicarbazone (ε = 10,300 M–1 cm–1) was used to calculate the rate of semicarbazone formation (μM, min-1) in reactions without Rid proteins and with CjRidA and Cj0327 in concentrations of 0.1, 1.0, and 10 μM. Standard deviation of the mean was determined from three technical triplicates by GraphPad Prism 7.0c.
Campylobacter jejuni Mutants Lacking cjridA Have a Motility Defect
Data from several bacterial species suggested RidA is involved in flagellar biosynthesis and/or motility (Reuter et al., 2015; Borchert and Downs, 2017a; Irons et al., 2018). A variant of C. jejuni 11168 lacking cjridA was generated and assessed for swimming motility on Mueller Hinton (MH) medium with 0.4% agar and 0.01% Triphenyltetrazolium Chloride (TTC). The data (Figure 3) showed that the cjridA mutant had significantly decreased motility when compared to wild type over the course of 72 h. A non-motile, aflagellate pseC mutant was used as a control to determine the spread of the inoculum that was due to diffusion. The motility data in Figure 3 is representative of experiments that were performed more than ten times on ten different days and included three independently constructed cjridA mutants. Although there was day-to-day variation in the absolute motility measured, the difference between the mutant and control strains remained consistent at ∼2-fold. Motility was not affected by a lesion in the gene encoding the Rid2 subfamily member (cj0327) in either a wild-type background or the cjridA mutant, (data not shown). This result supported the conclusion that the CjRidA and Cj0327 proteins are not physiologically redundant in C. jejuni, consistent with what has been found in other organisms. Addition of isoleucine (1 mM) to the motility agar did not affect the motility defect of the cjridA mutant (data not shown), and motility was restored to wild-type levels when cjridA was inserted into pseudogene cj0046 and expressed with its native promoter (Figure 4).
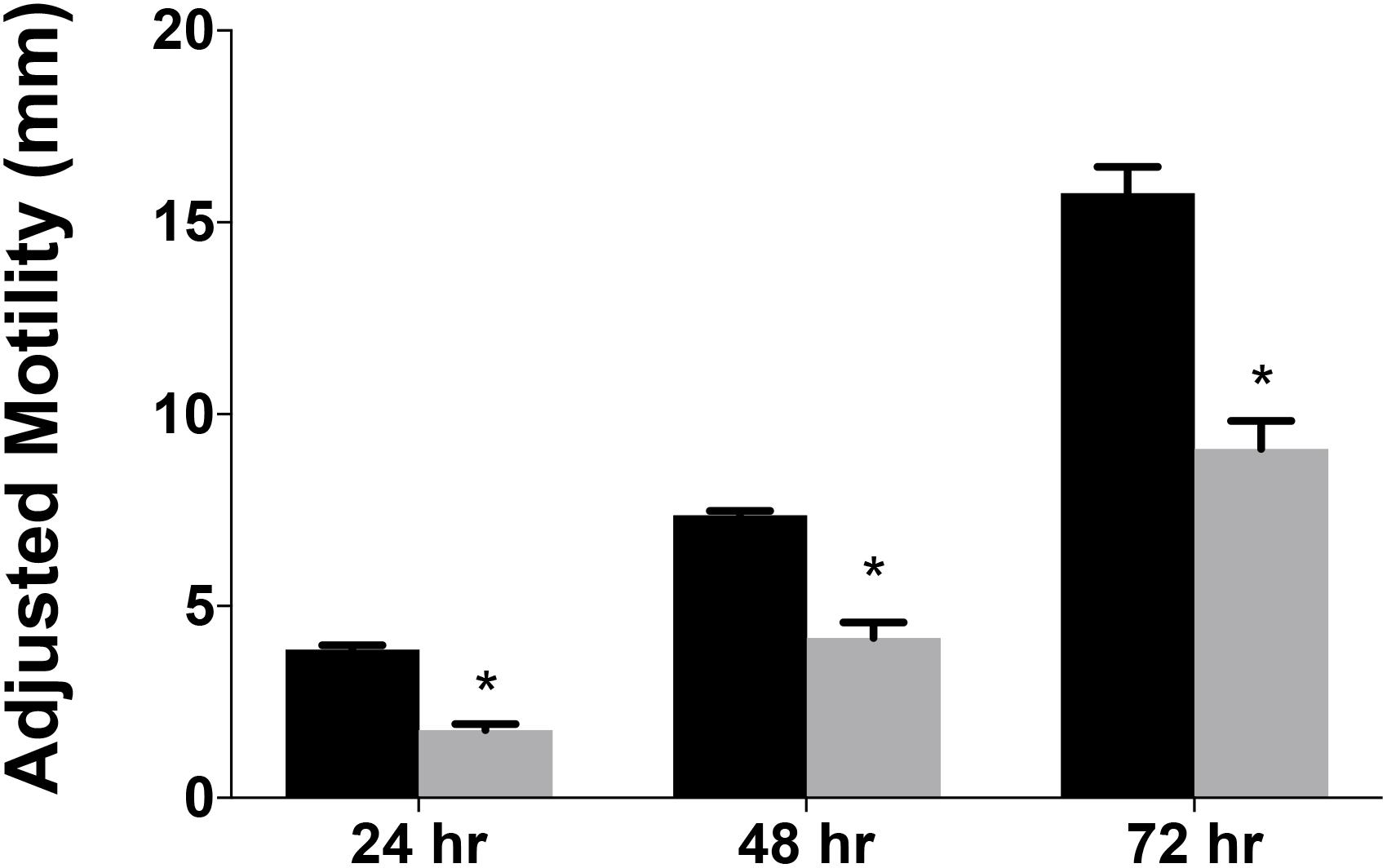
Figure 3. Campylobacter jejuni ridA mutants have a significant defect in motility. Swimming motility was determined for C. jejuni wild type (black) and a cjridA mutant (gray). Ten microliter of cell suspension was inoculated in the center of MH agar (0.4%) plate that was incubated up to 72 h in microaerophilic conditions. Motility was defined as swimming-dependent spread by subtracting the diameter of inoculum diffusion from the motility zone and dividing it by two. Error bars represent the standard errors of the mean of three technical triplicates, for wild type, and two biological replicates in technical triplicate for the cjridA mutant. Significance was determined between wild type andcjridA mutant for each time period and an asterisk denotes statistically significant (P < 0.005) variation between mutants, as determined by an unpaired Student t test performed with GraphPad Prism software, v7.0C.
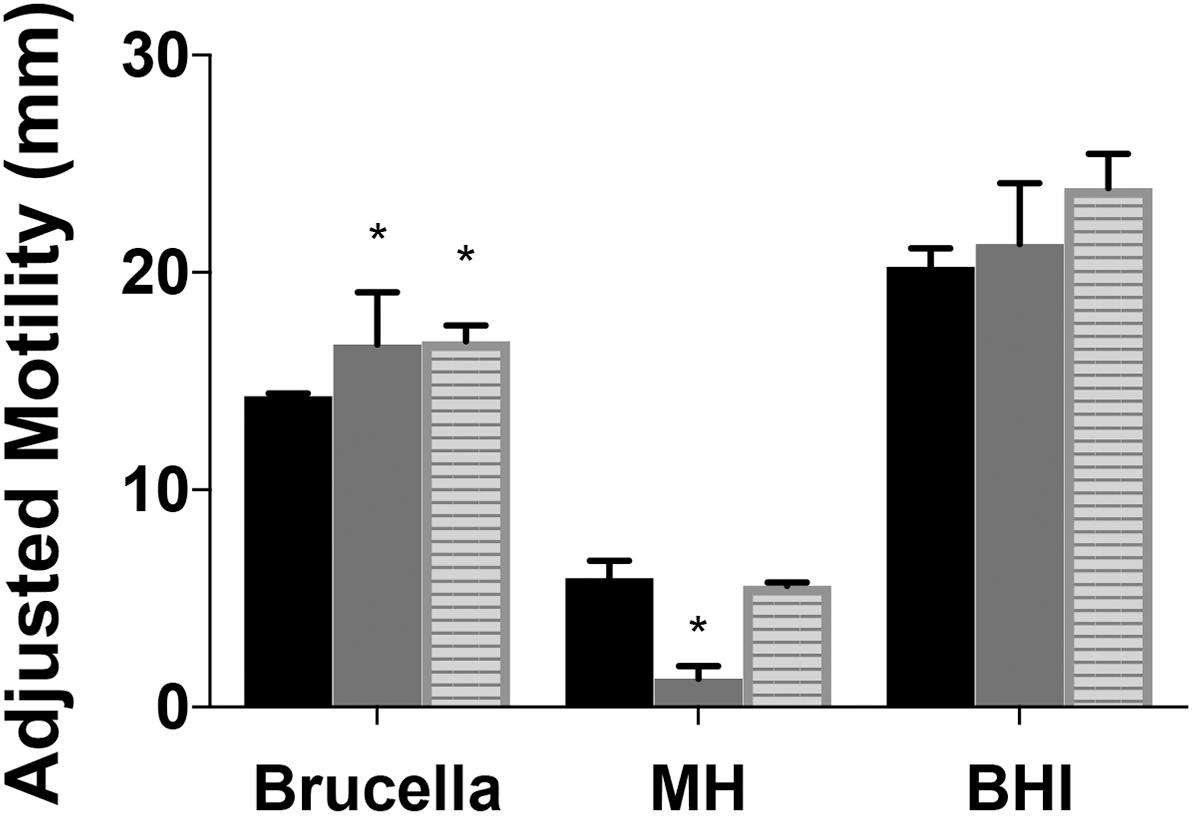
Figure 4. Motility of Campylobacter jejuni ridA mutants differs with media composition. Growth (not shown) and motility were improved by complex undefined-media. Motility of the C. jejuni wild type (black), cjridA mutant (gray), or the chromosomally complemented cjridA mutant (cj0046: cjridA-kan) (striped) was improved in Brucella or BHI as compared to MH motility agar (0.4%). Ten microliter of cell suspension was inoculated in the center of each agar (0.4%) plate that was incubated up for 36 h in microaerophilic conditions. Motility was defined as swimming-dependent spread by subtracting the diameter of inoculum diffusion from the motility zone and dividing it by two. Error bars represent the standard errors of the mean of three technical triplicates of wild-type and three biological replicates of cjridA and cj0046:cjridA-kan. Significance was determined between wild type andcjridA mutant for each time period and an asterisk denotes statistically significant (P < 0.02) variation between mutants, as determined by an unpaired Student t test performed with GraphPad Prism software, v7.0C.
A previous study reported cj1388 mutants had increased motility compared to wild-type in Brucella motility agar, a rich undefined medium (Reuter et al., 2015). Motility was assessed in Brucella, brain heart infusion (BHI), and MH motility agar for two mutants and wild type and the data are in Figure 4. Both Brucella and BHI media support increased mobility (and growth) of both mutants and wild type. Similar to what was previously reported, cjridA mutant motility was 1.1-fold and 1.4-fold higher than wild-type in BHI and Brucella motility media, respectively. Significantly, the cjridA mutant displayed a motility defect only on MH medium. The restoration of motility on the two complex media is consistent with regulation of metabolic flux in cjridA mutants limiting the production of, and/or damage by, the reactive enamine substrate of the CjRidA protein. For instance, in S. enterica and P. aeruginosa ridA mutant motility defects only arise when minimal defined media is used and the defects are eliminated by the addition of isoleucine which prevents 2AA formation. Given the complexity of metabolic systems and regulation, minimal defined media will be used in future studies to determine the impact of cjridA mutations motility.
CjRidA Is Required for Full Infection and/or Lysis by Phage NCTC 12673
The C. jejuni lytic phage, NCTC 12673 has decreased plaquing efficiency on aflagellate mutants (Javed et al., 2015a) and fails to form plaques on pseC mutants (Sacher, 2018). Plaque formation by NCTC 12673 was assessed with serially diluted aliquots of a phage lysate spotted on 0.6% agar overlays seeded with the indicated mutant or wild type. The efficiency of plating was tested on wild-type C. jejuni, a cjridA mutant (Figure 5) and a pseC mutant. As expected, no plaques were visible on the pseC mutant, which lacks the ability to synthesize pseudaminic acid, the major glycan modification of FlaA and FlaB subunits of the flagellum, and is therefore aflagellate (Javed et al., 2015b; Sacher, 2018). When plated on wild-type C. jejuni, the phage titer was 1 × 107 PFU/ml. When the same lysate was plated on a cjridA mutant, the titer was 2 × 106. This approximately 5-fold decrease in plating efficiency compared to the parental strain was consistent with the hypothesis that the cjridA mutant had a defect in flagellar biosynthesis and/or function.
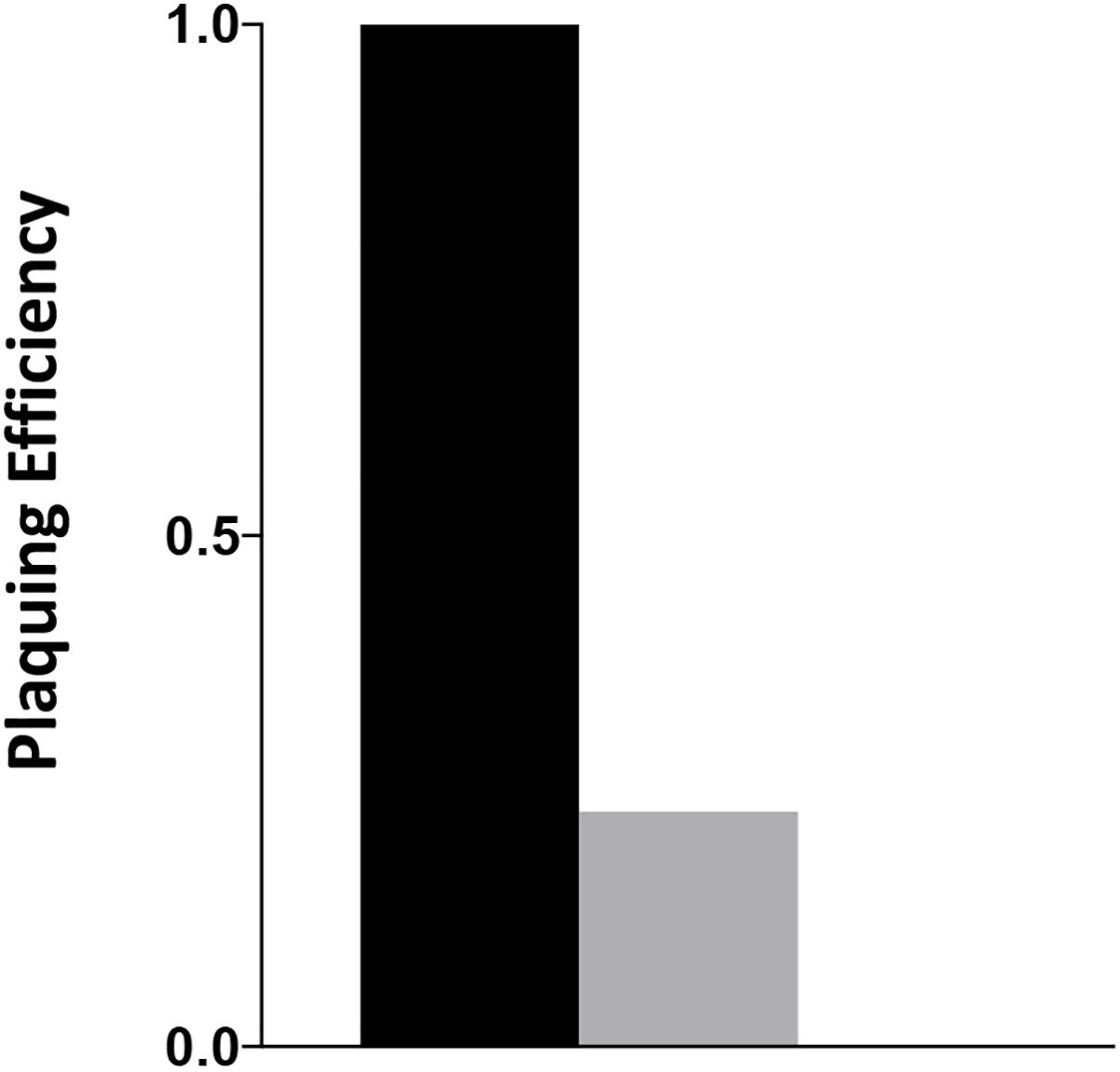
Figure 5. cjridA mutants support reduced plaque formation of phage NCTC 12673. A phage lysate was titered on C. jejuni wild type (black), a cjridA mutant (gray) and pseC mutant (plaquing efficiency of zero). Serial dilutions of the lysate were spotted on a NZCYM 0.6% agar overlay seeded with the appropriate bacterial mutant. After 1 day, plaque forming units (PFU/mL lysate) were determined with three technical triplicates of wild type, and two biological replicates in technical duplicate for the cjridA mutant. Number of plaques on wild type was defined as an efficiency of 1.
cjridA Mutants Have a Defect in Autoagglutination
The decreased motility and sensitivity to phage NCTC 12673 suggested a flagellar defect in the cjridA mutants. In both cases, the cjridA mutant phenotype fell between that of the wild type and the pseC mutant, which completely lacks flagella. Consistently, a hallmark of ridA mutants is the decreased, but not eliminated activity of the enzymes targeted by 2AA causing phenotypes that are less severe than complete lesions of the relevant enzymes. Changes in autoagglutination (AAG) can also indicate a change in flagella, specifically in flagellar glycan decoration, which correlates with a reduction in virulence (Guerry et al., 2006; Howard et al., 2009; Morrison and Imperiali, 2014). Reuter et al. reported that a cj1388 (cjridA) mutant had a slower rate of AAG compared to wild-type C. jejuni 11168, in medium supplemented with Tween-20 (0.002%) (Reuter et al., 2015). AAG was determined in our hands for wild type and cjridA mutant after suspension in several different media. Cells were harvested from MH agar plates and suspended in MH or PBS as appropriate. The cell suspension was adjusted to an OD600 of 1.0 in 5 mL of: (i) MH, (ii) MH with 0.002% Tween-20, or (iii) PBS. Consistent with previous observations, the cjridA mutant had a significant and reproducible decrease in AAG compared to wild type in MH supplemented with 0.002% Tween-20, reflected by more cells remaining in suspension (Figure 6). Each mutant was tested in triplicate. To ensure that any observed difference in phenotype was due to specific mutation, three separate clones for each mutant were tested separately and then the data were combined. The defect of a cjridA mutant appeared to reflect a slower rate of AAG, since the defect was significant after a 24-h incubation, but by 48 h the mutants were not significantly different than wild type. In our hands, other media used in reported AAG protocols (PBS or MH alone) failed to result in visible differences between the mutant and wild type. As expected, a pseC mutant showed almost complete cessation of AAG, thus another phenotype of the cjridA mutant fell between that of a wild type and the pseC mutant (Figure 6). The decreased rate of autoagglutination in a cjridA mutant supported the emerging model that CjRidA directly or indirectly affects flagellar regulation, biogenesis, glycosylation, or structure.
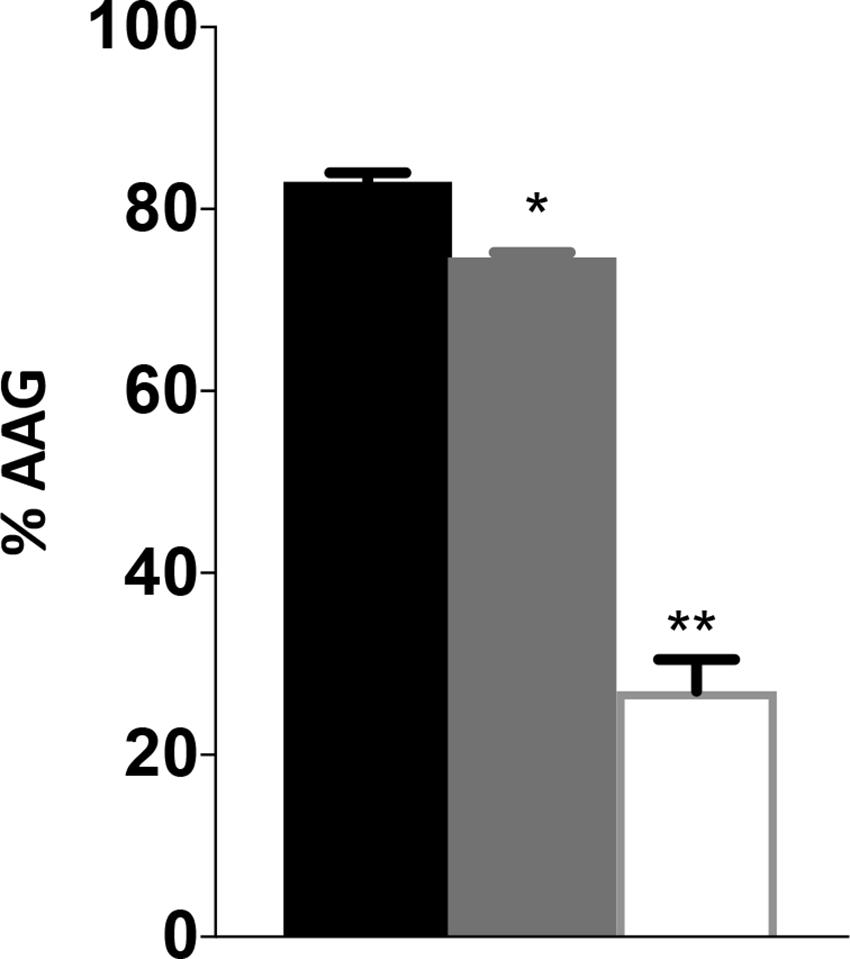
Figure 6. Campylobacter jejuni ridA mutants have an autoagglutination defect during the first 24 h. Autoagglutination was measured using C. jejuni wild type (black), a cjridA mutant (light gray) and a pseC mutant (white). % AAG represents the percentage of cells that autoagglutinated and settled in the bottom of the tube after 24 h, determined by the formula [(OD600i–OD600r)/OD600i] × 100. Error bars represent the standard errors of the mean of three technical triplicates, for wild type and pseC mutant, and two biological replicates in technical duplicate for cjridA mutants. Significance was determined between each mutant and wild type. One asterisk denotes statistically significant (P < 0.005) and two asterisks denote statistically significant (P < 0.0005) variation between mutants, as determined by an unpaired Student t test performed with GraphPad Prism software, v7.0C.
Transmission Electron Microscopy Shows CjRidA Impacts Flagella
Transmission electron microscopy (TEM) was performed on cells harvested from MH agar plates and suspended in PBS (Guerry et al., 2006). Efforts to fix cells with glutaraldehyde and formaldehyde or paraformaldehyde and stain with uranyl acetate or phosphotungstic acid failed to yield clear images and thus the cells were imaged with no fixative or stain (Figure 7B). The number of flagella were quantified using two independently constructed cjridA mutants and a wild-type strain of C. jejuni (Figure 7A). One hundred cells with two unobstructed poles from each mutant and the wild type were used for quantification. Of the one hundred wild-type cells observed, ∼ 60% had bipolar flagella, ∼20% had a single flagellum, and ∼20% had no visible flagellum. In contrast, of the 200 cjridA mutant cells observed, 20% had bipolar flagella, <40% had a single flagellum, <40% had no flagella. Beyond the number, structural anomalies of the flagella were noted in the mutant cells that were not seen in the wild-type sample (Figure 7B). First, there were “nub” structures on one or both poles of the bacterium (∼10% of mutant cells). Secondly, there were instances where flagella in the mutant were unusually long and apparently thinner than the wild type. Together these observations showed that the lack of CjRidA significantly impacted flagellar synthesis and or assembly. TEM images do not provide clarity on the specific flagellar defect caused by a cjridA mutation. Regardless, the images, in combination with the phenotypic analysis above allowed the conclusion that CjRidA is important for the full formation of a functional flagella.
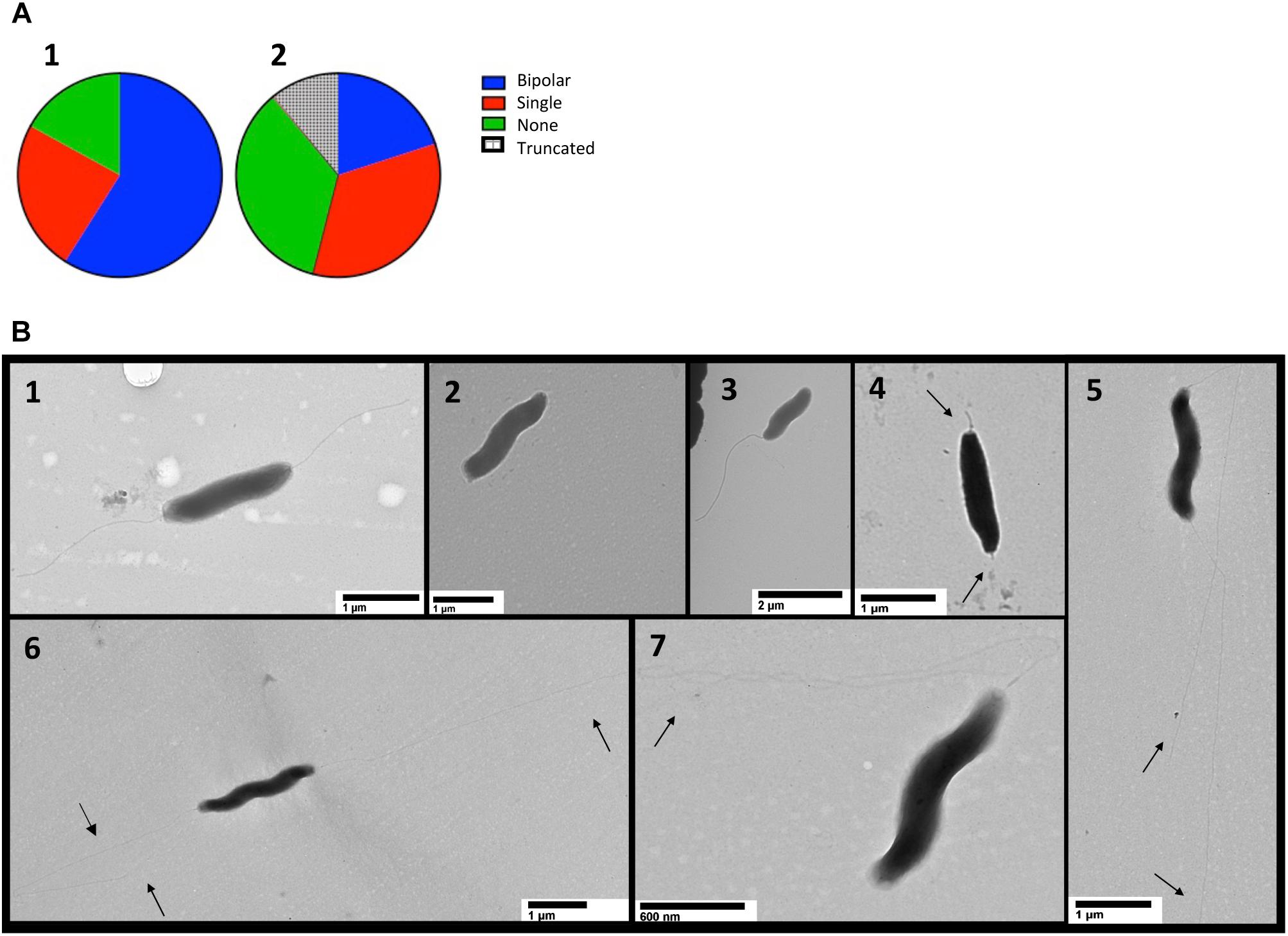
Figure 7. TEM detects flagellar differences in cjridA mutants. TEM was used to visualize the flagella of wild type and two cjridA mutants on multiple days. One hundred cells with clearly visible poles were assessed in each mutant or wild type. Pie charts in (A) represent the distribution of bipolar flagella (blue), a single polar flagellum (red) no flagellum (green) and truncated flagella (hatched). For wild type, N = 100, for cjridA, N = 200 (with 100 from each of two independent mutants) and long and thin flagella (quantified as bipolar or single) were classified by number of flagellar filaments. Lower panels (B1–B7) show representative TEM images for cells with: (B1) bipolar flagella, wild type is represented: (B2) no flagellum, cjridA mutant is represented; (B3) a single polar flagellum,cjridA mutant is represented; (B4) truncated flagella, seen only in cjridA mutants; and (B5–B7) long, potentially thin flagella, seen only in cjridA mutant.
Cj0828 Is the Biosynthetic Serine/Threonine Dehydratase in C. jejuni
In five organisms previously characterized, the phenotypic effects of eliminating the RidA homolog were due to the accumulation of 2AA, generated by a PLP-dependent serine threonine dehydratase enzyme (EC 4.3.1.19). As a consequence, a hallmark of the paradigm thus far has been the suppression of all defects by exogenous isoleucine, which allosterically inhibits the dehydratase enzyme(s). Within this context, it was striking that phenotypes associated with a cjridA mutation in C. jejuni were apparent in nutrient (MH) medium that contained abundant isoleucine. C. jejuni encodes a single gene annotated as a PLP-dependent serine/threonine dehydratase, (Cj0828, EC 4.3.1.19). Cj0828 shares 32% identity to S. enterica IlvA (Figure 8) but it lacks the C-terminal domain that contains the allosteric site for inhibition by isoleucine (Gallagher et al., 1998; Chen et al., 2013). These data suggested that if cj0828 encoded the legitimate biosynthetic threonine dehydratase, the presence of isoleucine would not prevent generation of 2AA by this enzyme. An insertion deletion was introduced into cj0828 and growth was tested on a defined minimal medium (MCLMAN). In minimal medium, the cj0828 mutant required isoleucine for full growth, indicating this gene product was the biosynthetic serine/threonine dehydratase in vivo (data not shown). To reflect this result, the gene was renamed cjilvA. The identification of cjilvA suggested three possible scenarios to explain the RidA paradigm in C. jejuni: 1) cjilvA is constitutively expressed and thus generates 2AA even on nutrient medium, 2) there are other enzyme(s) in the cell that generate 2AA, or 3) 2AA is not responsible for the phenotypes of the cjridA mutant. The latter would suggest there was another reactive metabolite produced in the cell that is quenched by CjRidA.

Figure 8. Campylobacter jejuni Cj0828 is an IlvA homolog. Salmonella enterica IlvA and Campylobacter jejuni Cj0828 protein sequences share 32% identity. Cj0828 (cjIlvA) lacks the C-terminal domain that contain the site of allosteric regulation by isoleucine.
C. jejuni Expands Features of the RidA Paradigm
The contribution of CjIlvA to the phenotypes of a cjridA mutant was tested by constructing a double mutant. A cjilvA loss of function mutation was introduced into a cjridA mutant background and the resulting double mutant was assessed for motility. Motility assays were performed with the cjridA cjilvA double mutant on MH with 0.4% agar (Figure 9). The motility of the cjilvA mutant was indistinguishable from the parental wild type. Similarly, the motility of the cjridA cjilvA double mutant was no different than the single cjridA mutant. Importantly, both mutants carrying a cjridA mutation had a >2-fold decrease in motility compared to their respective parental strain. These data showed that CjIlvA was not the source of sufficient 2AA to result in the phenotypes detected for the cjridA mutant on MH motility agar. Thus, the source of the reactive enamine presumed to be responsible for the flagellar defects of the cjridA mutants remains to be determined.
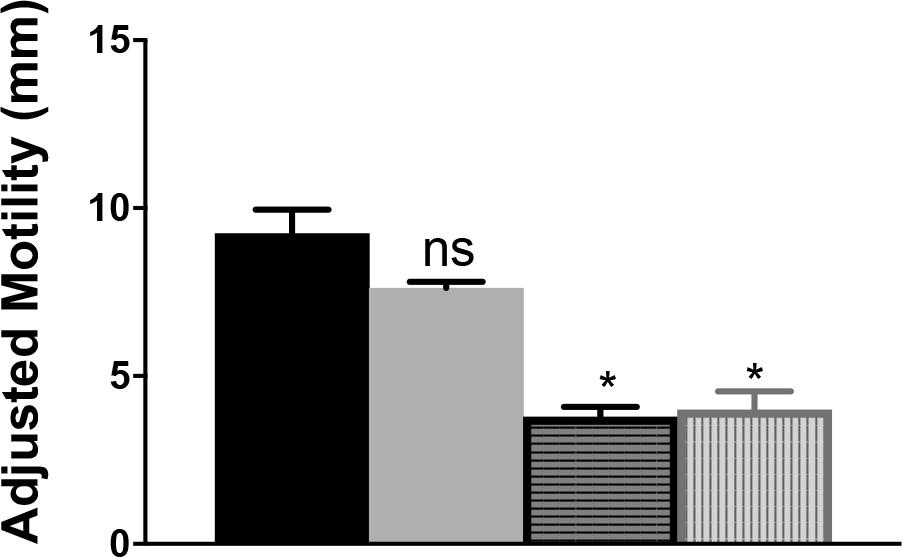
Figure 9. A cjilvA mutation does not eliminate the phenotype of a cjridA mutant. Mutants were grown on BHI overnight. Then cells were scraped from agar surface, resuspended in 1 mL of PBS and the OD600 was set to 1.0. Then, 10 μL of cell suspension was used to inoculate the center of a MH 0.4% agar plate. After 48 h the diameter of motility was measured and the values adjusted to account for the swimming-independent spread of the inoculum, as determined by the non-motile pseC mutant. The wild-type parental strain (black) and the ilvA:cat mutant (gray) had no defect in motility. The cjridA mutant (horizontal stripe) had a >2-fold decrease in motility that was not restored in acjridA ilvA double mutant (vertical stripe). Error bars represent the standard errors of the mean of three technical triplicates, for wild type, and two biological replicates in technical triplicate for cjridA and cjridA cj0828 mutants. Significance was determined between each mutant and wild type; asterisk denotes statistically significant (P < 0.0001) variation between mutants, as determined by an unpaired Student t test performed with GraphPad Prism software, v7.0C.
Discussion
The data herein demonstrate that the gene designated cj1388 in Campylobacter jejuni 11168 is a RidA with 2AA deaminase activity in vivo. C. jejuni is the first organism to date where the major phenotypic consequences of lacking RidA are not caused by the activity of a serine threonine dehydratase. Thus C. jejuni provides an opportunity to identify additional generators of reactive enamine(s) like 2AA, that can impact the physiology of different organisms in the absence of RidA. One of the two additional 2AA generators found in S. enterica, cysteine desulfhydrase (CdsH; EC 2.5.1.47), appears to be present in C. jejuni and additional work will determine if this enzyme has a role in generating the phenotypes of a cjridA mutant.
Results presented herein, which used three independent cjridA mutants, suggest C. jejuni 11168 lacking ridA has a defect in flagellar biosynthesis, regulation, or structure. C. jejuni mutants lacking cjridA have defects in motility, AAG, and phage infectivity, all of which require or are enhanced by flagella (Guerry et al., 1991; Golden and Acheson, 2002; Javed et al., 2015b). Motility is essential for C. jejuni to move through the viscous mucosal environment to colonize a human host, and protein glycosylation is essential for flagellar biosynthesis and function. Flagellum (FlaA and FlaB) subunits are modified by O-linked pseudaminic and legionaminic acids and their derivatives at up to 19 Ser/Thr sites before export and assembly of the flagellar apparatus (Thibault et al., 2001; Logan et al., 2002; Schirm et al., 2005; Logan, 2006; Ewing et al., 2009). Importantly, thus far the only defined targets of accumulated 2AA are PLP-dependent enzymes. Given the importance of glycosylation of the flagellar subunits, it is possible that the UDP-4-amino-4,6-dideoxy-N-acetyl-ß–L-altrosamine transaminase (Cj1294/PseC; EC 2.6.1.92), a fold-type II PLP-dependent enzyme, could be a critical target of 2AA and thus be damaged in a cjridA mutant.
Our favored model suggests that 2AA accumulates in a cjridA mutant and damages PLP-dependent enzyme, PseC, leading to a decrease in pseudaminic acid modification on FlaA. Consistent with this model, changes in FlaA glycosylation affect AAG, motility, and virulence (Misawa and Blaser, 2000; Thibault et al., 2001; Logan et al., 2002; Schirm et al., 2005; Guerry et al., 2006; Ewing et al., 2009). Based on other examples, damage by 2AA is expected to reduce the activity of PseC 30–50% (Lambrecht et al., 2010; Ernst and Downs, 2016, 2018; Borchert and Downs, 2017b; Irons et al., 2018). In this case, the phenotypes resulting from PseC damage could vary among the cell population and be similar to the range of phenotypes previously shown from flaA point mutations (Ewing et al., 2009; Ulasi et al., 2015; Zebian et al., 2016).
Materials and Methods
Bacterial Strains, Plasmids and Media
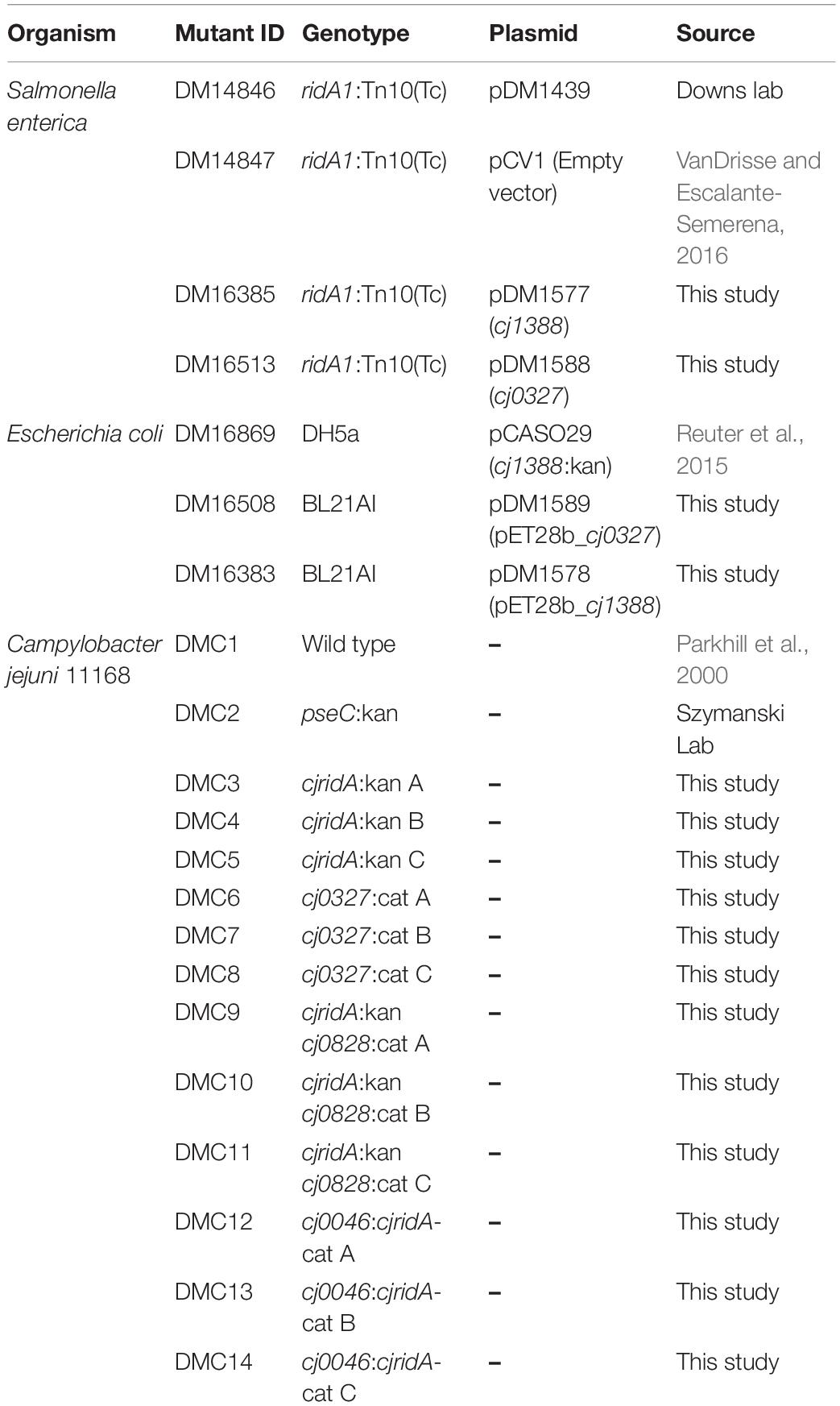
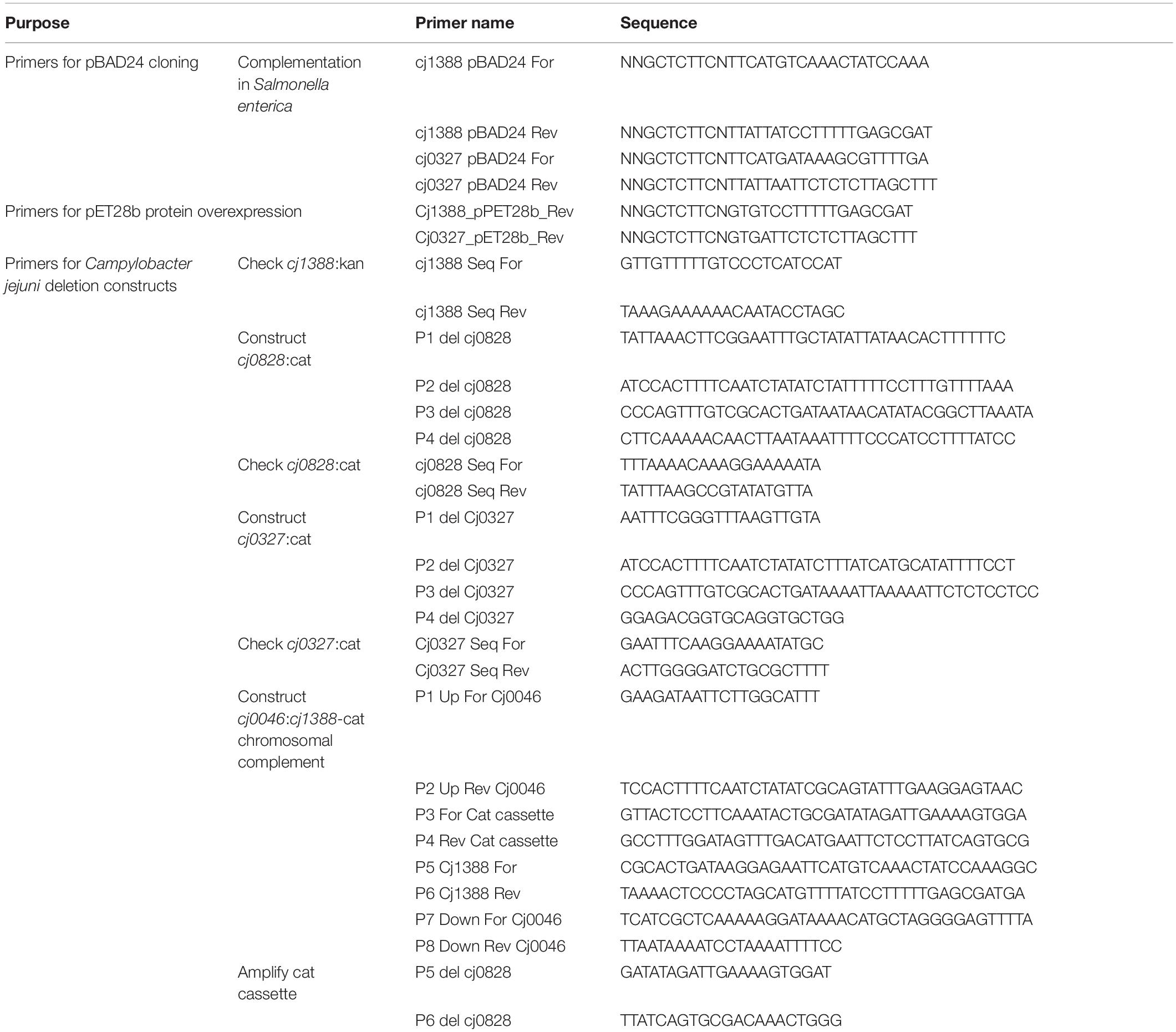
The strains, plasmids and primers used in this study are listed in Tables 1–3 with their sources. C. jejuni human isolate NCTC 11168 (Parkhill et al., 2000) was used as the parental strain. Derivatives of S. enterica serovar Typhimurium LT2 (S. enterica) were used for in vivo complementation studies.
Derivatives of C. jejuni 11168 were grown on Mueller Hinton (MH, 21 g/liter), Brain heart infusion (BHI, 37 g/L), Brucella agar (28.1 g/L) or NZCYM (22 g/L) at 37°C under microaerobic conditions (85% N2, 10% CO2, 5% O2) (Blattner et al., 1997). S. enterica and E. coli strains were grown in Difco Nutrient Broth (8 g/l) with NaCl (5 g/l) at 37°C. Minimal medium was NCE salts with MgSO4 (Vogel and Bonner, 1956), trace minerals (Balch and Wolfe, 1976), and 11 mM glucose. Additions, isoleucine (1 mM), and serine (5 mM) were added as indicated. Antibiotic concentrations were as follows; 150 μg/mL ampicillin or 50 μg/mL kanamycin were used for S. enterica and 15 μg/mL chloramphenicol or 30 μg/mL kanamycin were used for C. jejuni. When needed to induce expression of genes in relevant plasmids, L-arabinose was added (0.2%). Chemicals were purchased from MilliporeSigma (Sigma-Aldrich, St. Louis, MO).
Growth Quantification
Growth of S. enterica in liquid culture was assessed using a BioTek Elx808 microtiter plate reader following optical density at 650 nm at 37°C with slow shaking speed. Overnight cultures of S. enterica in biological triplicate were grown in rich medium at 37°C, pelleted and resuspended in an equal volume of sterile NaCl (8.5 g/L). The resulting cell suspension was used to inoculate growth medium (2% inoculum) and growth was monitored for 24 h. The resulting data were plotted using GraphPad Prism 7.0, generating curves in log10-format that display the mean of three replicates and standard deviation of the mean. Specific growth rates (μ) were calculated according to the following equation: ln(X/X0)/T, where X is OD650, X0 is the starting OD650 of the exponential growth period monitored, and T is time in hours.
Molecular Biology
A plasmid (pCASO29) with a deletion/kanamycin insertion construct in cj1388 was used to construct a cjridA:kan mutant (DMC3, DMC4, and DMC5) (Reuter et al., 2015). A pseC:kan mutant was obtained from the Szymanski laboratory collection. Additional mutants were constructed using standard methods (Hansen et al., 2007; Tan and Berg, 2004). Briefly, to generate an insertion/deletion in a gene of interest, homology both up- and down-stream to the gene of interest was joined to a drug resistance cassette by overlap extension PCR. PCR products were purified using Qiagen gel extraction kit (ID 28506). The natural competence of C. jejuni was exploited to transform the PCR product into cells grown on nutrient rich medium, BHI with 2% yeast extract. After 24 h growth, cells were streaked on selective medium and colonies formed after 3-5 days. Colonies were streaked for isolation and culture stocks were frozen in glycerol. Insertion deletions of relevant genes was confirmed by PCR. The complete protocol for Natural Transformation can be found on Protocols.io at https://dx.doi.org/10.17504/protocols.io.magc2bw.
Derivatives of plasmid pBAD24 and pET28b were created using a BspQI restriction cloning method as previously described by Galloway et al. (2013) with a modified vector that contained the BspQI site (pCV1) (VanDrisse and Escalante-Semerena, 2016). S. enterica or E. coli competent cells were prepared and transformed using standard method. Transformants were recovered in nutrient broth, plated to selective medium at 37°C before isolating and confirming the plasmid structure.
For pseC:km mutant construction, pseC was amplified from 11168 using the pseC-F and pseC-R primers (Table 3). This insert was purified, digested with BamHI and XhoI, inserted into PCRscript (Stratagene) digested with the same enzymes, and transformed into E. coli DH5α. The extracted plasmids were digested with XhoI and BamHI to confirm insert presence and one plasmid subsequently digested with NcoI, purified, blunted and treated with alkaline phosphatase to prevent re-ligation. The pseC gene was interrupted by ligating a kanamycin resistance cassette (km). The mutant was confirmed using the pseC-F/mid KmR primer pair (Table 3) and sequenced.
Protein Production and Purification
Proteins CjRidA and Cj0327 were purified from E. coli strain BL21AI harboring pET vector constructs. The polyhistidine-tagged proteins were purified by nickel-affinity chromatography as previously described (Lambrecht et al., 2010). Overnight cultures in LB (10 mL) were used to inoculate flasks containing super broth (1.5 L) supplemented with kanamycin (50 μg/mL). Cultures were grown at 37°C with shaking until an OD650 between 0.7 and 1.0 was reached. Arabinose (0.2%) was added to induce T7 RNA polymerase for 18 h. Cells were harvested by centrifugation at 7,000 × g and the cell pellets were stored at −80°C until use. Binding buffer (50 mM potassium phosphate pH 7.5, 100 mM NaCl, 5 mM imidazole, and 10% glycerol) was added to thaw cells (2 mL per gram wet cell weight) along with lysozyme (1 mg/mL) and DNase (20 Units/mL), and the cells were lysed with a One Shot Cell Disruptor at 18,000 psi (Constant Systems). The lysate was clarified by centrifugation (40,000 × g for 45 minutes) and passed through a 0.45 μm syringe filter (Argos Technologies) prior to being loaded onto 5 mL HisTrapTM HP column and purified using the manufacturer’s protocol (GE Healthcare). Protein was eluted with a 0–100% gradient of elution buffer (50 mM potassium phosphate pH 7.5, 100 mM NaCl, 500 mM imidazole and 10% glycerol). The fractions were assessed for purity, pooled, and concentrated using a 7,000 molecular weight cut-off protein concentrator (Millipore). The protein preparations were dialyzed into storage buffer (50 mM potassium phosphate pH 7.5, 10% glycerol) using a PD-10 desalting column (GE Healthcare). Proteins were subjected to SDS/PAGE and purity was assessed using a Foto/Analyst FX (Fotodyne) imager and TotalLab Quant v11 densitometry software. Protein concentration was quantified using BCA Protein Assay (Thermo Scientific), and the samples were frozen in liquid nitrogen and stored at −80°C.
L-Amino Acid Oxidase Assays
The LOX-based assay for Rid activity was adapted from a previously described assay and has been used to assess activity of Rid proteins from several organisms (Hafner and Wellner, 1979; Niehaus et al., 2015; Hodge-Hanson and Downs, 2017; Degani et al., 2018). The 2-iminobutyrate intermediate from 2-aminobutyrate was derivatized with semicarbazide resulting in semicarbazone detected by absorbance at 248 nm. The assay mixture (100 μL total volume) contained potassium pyrophosphate (50 mM, pH 8.7), neutralized semicarbazide (10 mM), bovine liver catalase (1 μg), L-amino acid oxidase (0.5 μg) and 0.1, 1.0 or 10 μM Rid protein. Reactions were started in a 96-well quartz plate with the addition of 2-aminobutyrate to the final concentration of 0.5 mM. Following the addition of substrate, the path length for each well was measured and used along with the molar extinction coefficient for semicarbazone (ε = 10,300 M–1 cm–1) to calculate the rate of semicarbazone formation. Standard deviation of the mean was determined from three technical triplicates by GraphPad Prism 7.0c.
Motility
Assays for swimming motility were done by modifying previously described methods (Guerry et al., 1991; Palyada et al., 2009; Neal-McKinney and Konkel, 2012; Vorwerk et al., 2014). Briefly, bacteria were harvested from overnight growth on BHI or MH agar plates into PBS and the OD600 was adjusted to 1.0. Ten microliter of the bacteria suspension was inoculated on individual plates by gently piercing the soft agar before expelling the cell suspension into 0.4% agar Brucella, BHI, or MH. Agar plates were incubated at 37°C for 24–72 h. The diameter of each swimming halo was measured and recorded in millimeter (mm). A non-motile pseC mutant served as a negative control; the spread of the pseC inoculum was subtracted from the motility zone diameter of the experimental strains and the number divided by 2 to get the motility distances as reported in mm. The data shown represent the mean of three technical replicates. For each mutation of interest, three independently isolated mutants were tested to ensure phase variability did not contribute to motility defects. Error bars represent the standard error of the mean. Statistical significance (P < 0.02) was determined by unpaired Student’s test (t test) using GraphPad Prism 7.0c.
Phage NCTC 12673 Plaque Assay
Plaque formation by NCTC 12673 phage was tested by spotting dilutions of a lysate onto a freshly inoculated bacterial suspension using a standard agar overlay method (Javed et al., 2015a). Briefly, 100 μL of a bacterial suspension (OD600 of ∼0.35) was mixed with 5 mL sterile 0.6% molten NZCYM agar (Sigma-Aldrich, St. Louis, MO) and poured onto the surface of a NZCYM plate. After 15 min, 10 μL aliquots of serial dilutions of a phage lysate were spotted onto the agar surface and allowed to completely dry before incubation at 37°C under microaerobic conditions. After 24 h, plaques were counted and the apparent number of PFU/mL was determined.
Autoagglutination Assays
Published protocols for autoagglutination were adapted for use (Misawa and Blaser, 2000; Guerry et al., 2006; Reuter et al., 2015). Simply, cells were harvested from overnight growth on MH agar plates and resuspended in MH broth. The OD600 was measured and adjusted to 1.0 in 5 mL of MH broth with 0.002% Tween-20 in a glass test tube. The top 1 mL was removed and OD600 measured (OD600i). The remaining 4 mL sat without shaking at room temperature in air. At 24, and 48 h, a 1 mL aliquot of the liquid was removed and the absorbance was measured to obtain the recorded OD600 (OD600r). The percent of autoagglutination (%AAG) reported was calculated as [(OD600i−OD600r)/OD600i] × 100.
Data Availability
All datasets generated for this study are included in the manuscript and/or the supplementary files.
Author Contributions
DD and JI conceived the project, designed the experiments, analyzed the data, and wrote the manuscript. JS and CS provided advice on the experimental design and edits in manuscript writing. JI performed the experiments. JI, JS, DD, and CS contributed reagents, materials, and analysis tools.
Funding
This work was supported by a competitive grant GM095837 from the National Institutes of Health (DD). JS is a recipient of an NSERC Alexander Graham Bell Canada Graduate Student Scholarship. CS is an Alberta Innovates Strategic Chair in Bacterial Glycomics.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors thank the Robert J. Maier laboratory in the Department of Microbiology, University of Georgia for providing space, supplies, and we especially thank Stéphane Benoit for technical guidance. The authors also thank Stephen Andersen and Muhammad Afzal Javed for construction of the pseC mutant, Bruce Pearson at the Quadram Institute for supplying plasmids used to construct cjridA:kan deletion mutants, and John Shields and Mary Ard at the Georgia Electron Microscopy core for training and technical assistance.
References
Balch, W. E., and Wolfe, R. S. (1976). New approach to the cultivation of methanogenic bacteria: 2-mercaptoethanesulfonic acid (HS-CoM)-dependent growth of Methanobacterium ruminantium in a pressurized atmosphere. Appl. Environ. Microb. 32, 781–791.
Blattner, F. R., Plunkett, G. III, Bloch, C. A., Perna, N. T., Burland, V., Riley, M., et al. (1997). The complete genome sequence of Escherichia coli K-12. Science 277, 1453–1474.
Borchert, A. J., and Downs, D. M. (2017a). Endogenously generated 2-aminoacrylate inhibits motility in Salmonella enterica. Sci. Rep. 7:12971. doi: 10.1038/s41598-017-13030-x
Borchert, A. J., and Downs, D. M. (2017b). The response to 2-aminoacrylate differs in Escherichia coli and Salmonella enterica, despite shared metabolic components. J. Bacteriol. 199, e140–17. doi: 10.1128/JB.00140-17
Chen, L., Chen, Z., Zheng, P., Sun, J., and Zeng, A. P. (2013). Study and reengineering of the binding sites and allosteric regulation of biosynthetic threonine deaminase by isoleucine and valine in Escherichia coli. Appl. Microbiol. Biotechnol. 97, 2939–2949. doi: 10.1007/s00253-012-4176-z
Christopherson, M. R., Schmitz, G. E., and Downs, D. M. (2008). YjgF is required for isoleucine biosynthesis when Salmonella enterica is grown on pyruvate medium. J. Bacteriol. 190, 3057–3062. doi: 10.1128/JB.01700-07
Clark, C. G., Chong, P. M., McCorrister, S. J., Simon, P., Walker, M., Lee, D. M., et al. (2014). The CJIE1 prophage of Campylobacter jejuni affects protein expression in growth media with and without bile salts. BMC Microbiol. 14:70. doi: 10.1186/1471-2180-14-70
Degani, G., Barbiroli, A., Regazzoni, L., Popolo, L., and Vanoni, M. A. (2018). Imine deaminase activity and conformational stability of UK114, the mammalian member of the rid protein family active in amino acid metabolism. Int. J. Mol. Sci. 19:945. doi: 10.3390/ijms19040945
Downs, D. M., and Ernst, D. C. (2015). From microbiology to cancer biology: the Rid protein family prevents cellular damage caused by endogenously generated reactive nitrogen species. Mol. Microbiol. 96, 211–219. doi: 10.1111/mmi.12945
ElRamlawy, K. G., Fujimura, T., Baba, K., Kim, J. W., Kawamoto, C., Isobe, T., et al. (2016). Der f 34, a novel major house dust mite allergen belonging to a highly conserved Rid/YjgF/YER057c/UK114 family of imine deaminases. J. Biol. Chem. 291, 21607–21615. doi: 10.1074/jbc.m116.728006
Ernst, D. C., Anderson, M. E., and Downs, D. M. (2016). L-2,3-diaminopropionate generates diverse metabolic stresses in Salmonella enterica. Mol. Microbiol. 101, 210–223. doi: 10.1111/mmi.13384
Ernst, D. C., and Downs, D. M. (2016). 2-Aminoacrylate stress induces a context-dependent glycine requirement inridAStrains of Salmonella enterica. J. Bacteriol. 198, 536–543. doi: 10.1128/JB.00804-15
Ernst, D. C., and Downs, D. M. (2018). Mmf1p couples amino acid metabolism to mitochondrial DNA maintenance in Saccharomyces cerevisiae. mBio 9, e84–18. doi: 10.1128/mBio.00084-18
Ernst, D. C., Lambrecht, J. A., Schomer, R. A., and Downs, D. M. (2014). Endogenous synthesis of 2-aminoacrylate contributes to cysteine sensitivity in Salmonella enterica. J. Bacteriol. 196, 3335–3342. doi: 10.1128/JB.01960-14
Ewing, C. P., Andreishcheva, E., and Guerry, P. (2009). Functional characterization of flagellin glycosylation in Campylobacter jejuni 81-176. J. Bacteriol. 191, 7086–7093. doi: 10.1128/JB.00378-09
Flint, A., Sun, Y. Q., Butcher, J., Stahl, M., Huang, H., and Stintzi, A. (2014). Phenotypic screening of a targeted mutant library reveals Campylobacter jejuni defenses against oxidative stress. Infect. Immun. 82, 2266–2275. doi: 10.1128/IAI.01528-13
Flynn, J. M., Christopherson, M. R., and Downs, D. M. (2013). Decreased coenzyme A levels in ridA mutant strains of Salmonella enterica result from inactivated serine hydroxymethyltransferase. Mol. Microbiol. 89, 751–759. doi: 10.1111/mmi.12313
Flynn, J. M., and Downs, D. M. (2013). In the absence of RidA, endogenous 2-aminoacrylate inactivates alanine racemases by modifying the pyridoxal 5′-phosphate cofactor. J. Bacteriol. 195, 3603–3609. doi: 10.1128/JB.00463-13
Gallagher, D. T., Gilliland, G. L., Xiao, G., Zondlo, J., Fisher, K. E., Chinchilla, D., et al. (1998). Structure and control of pyridoxal phosphate dependent allosteric threonine deaminase. Structure 6, 465–475. doi: 10.1016/s0969-2126(98)00048-3
Galloway, N. R., Toutkoushian, H., Nune, M., Bose, N., and Momany, C. (2013). Rapid cloning for protein crystallography using type IIS restriction enzymes. Cryst. Growth Des. 13, 2833–2839. doi: 10.1021/cg400171z
Golden, N. J., and Acheson, D. W. (2002). Identification of motility and autoagglutination Campylobacter jejuni mutants by random transposon mutagenesis. Infect. Immun. 70, 1761–1771. doi: 10.1128/iai.70.4.1761-1771.2002
Guccione, E. J., Kendall, J. J., Hitchcock, A., Garg, N., White, M. A., Mulholland, F., et al. (2017). Transcriptome and proteome dynamics in chemostat culture reveal how Campylobacter jejuni modulates metabolism, stress responses and virulence factors upon changes in oxygen availability. Environ. Microbiol. 19, 4326–4348. doi: 10.1111/1462-2920.13930
Guerry, P., Alm, R. A., Power, M. E., Logan, S. M., and Trust, T. J. (1991). Role of two flagellin genes in Campylobacter motility. J. Bacteriol. 173, 4757–4764. doi: 10.1128/jb.173.15.4757-4764.1991
Guerry, P., Ewing, C. P., Schirm, M., Lorenzo, M., Kelly, J., Pattarini, D., et al. (2006). Changes in flagellin glycosylation affect Campylobacter autoagglutination and virulence. Mol. Microbiol. 60, 299–311. doi: 10.1111/j.1365-2958.2006.05100.x
Hafner, E. W., and Wellner, D. (1979). Reactivity of the imino acids formed in the amino acid oxidase reaction. Biochemistry 18, 411–417. doi: 10.1021/bi00570a004
Hansen, C. R., Khatiwara, A., Ziprin, R., and Kwon, Y. M. (2007). Rapid construction of Campylobacter jejuni deletion mutants. Lett. Appl. Microbiol. 45, 599–603. doi: 10.1111/j.1472-765x.2007.02232.x
Hao, H., Li, F., Han, J., Foley, S. L., Dai, M., Wang, X., et al. (2017). Cj1199 Affect the development of erythromycin resistance in campylobacter jejuni through regulation of leucine biosynthesis. Front. microbiol. 8:16. doi: 10.3389/fmicb.2017.00016
Hodge-Hanson, K. M., and Downs, D. M. (2017). Members of the Rid protein family have broad imine deaminase activity and can accelerate the Pseudomonas aeruginosa D-arginine dehydrogenase (DauA) reaction in vitro. PLoS One 12:e0185544. doi: 10.1371/journal.pone.0185544
Howard, S. L., Jagannathan, A., Soo, E. C., Hui, J. P., Aubry, A. J., Ahmed, I., et al. (2009). Campylobacter jejuni glycosylation island important in cell charge, legionaminic acid biosynthesis, and colonization of chickens. Infect. Immun. 77, 2544–2556. doi: 10.1128/IAI.01425-08
Irons, J., Hodge-Hanson, K. M., and Downs, D. M. (2018). PA5339, a RidA homolog, is required for full growth in Pseudomonas aeruginosa. J. Bacteriol. 200, e434–18.
Javed, M. A., Sacher, J. C., van Alphen, L. B., Patry, R. T., and Szymanski, C. M. (2015a). A flagellar glycan-specific protein encoded by campylobacter phages inhibits host cell growth. Viruses 7, 6661–6674. doi: 10.3390/v7122964
Javed, M. A., van Alphen, L. B., Sacher, J., Ding, W., Kelly, J., Nargang, C., et al. (2015b). A receptor-binding protein of Campylobacter jejuni bacteriophage NCTC 12673 recognizes flagellin glycosylated with acetamidino-modified pseudaminic acid. Mol. Microbiol. 95, 101–115. doi: 10.1111/mmi.12849
Johnson, J. G., Gaddy, J. A., and DiRita, V. J. (2016). The PAS domain-containing protein HeuR regulates heme uptake in Campylobacter jejuni. mBio 7, e1691–16. doi: 10.1128/mBio.01691-16
Kim, J.-M., Yoshikawa, H., and Shirahige, K. (2001). A member of the YER057c/yjgf/Uk114 family links isoleucine biosynthesis to intact mitochondria maintenance in Saccharomyces cerevisiae. Genes Cells 6, 507–517. doi: 10.1046/j.1365-2443.2001.00443.x
Lambrecht, J. A., Browne, B. A., and Downs, D. M. (2010). Members of the YjgF/YER057c/UK114 family of proteins inhibit phosphoribosylamine synthesis in vitro. J. Biol. Chem. 285, 34401–34407. doi: 10.1074/jbc.M110.160515
Lambrecht, J. A., Flynn, J. M., and Downs, D. M. (2012). Conserved YjgF protein family deaminates reactive enamine/imine intermediates of pyridoxal 5′-phosphate (PLP)-dependent enzyme reactions. J. Biol. Chem. 287, 3454–3461. doi: 10.1074/jbc.M111.304477
Lambrecht, J. A., Schmitz, G. E., and Downs, D. M. (2013). RidA proteins prevent metabolic damage inflicted by PLP-dependent dehydratases in all domains of life. mBio 4, e33–13. doi: 10.1128/mBio.00033-13
Leitner-Dagan, Y., Ovadis, M., Zuker, A., Shklarman, E., Ohad, I., Tzfira, T., et al. (2006). CHRD, a plant member of the evolutionarily conserved YjgF family, influences photosynthesis and chromoplastogenesis. Planta 225, 89–102. doi: 10.1007/s00425-006-0332-y
Logan, S. M. (2006). Flagellar glycosylation - a new component of the motility repertoire? Microbiology 152, 1249–1262. doi: 10.1099/mic.0.28735-0
Logan, S. M., Kelly, J. F., Thibault, P., Ewing, C. P., and Guerry, P. (2002). Structural heterogeneity of carbohydrate modifications affects serospecificity of Campylobacter flagellins. Mol. Microbiol. 46, 587–597. doi: 10.1046/j.1365-2958.2002.03185.x
Misawa, N., and Blaser, M. J. (2000). Detection and characterization of autoagglutination activity by Campylobacter jejuni. Infect. Immun. 68, 6168–6175. doi: 10.1128/iai.68.11.6168-6175.2000
Morrison, M. J., and Imperiali, B. (2014). The renaissance of bacillosamine and its derivatives: pathway characterization and implications in pathogenicity. Biochemistry. 53, 624–638. doi: 10.1021/bi401546r
Neal-McKinney, J. M., and Konkel, M. E. (2012). The Campylobacter jejuni CiaC virulence protein is secreted from the flagellum and delivered to the cytosol of host cells. Front. Cell Infect. Microbiol. 2:31. doi: 10.3389/fcimb.2012.00031
Niehaus, T. D., Gerdes, S., Hodge-Hanson, K., Zhukov, A., Cooper, A. J., ElBadawi-Sidhu, M., et al. (2015). Genomic and experimental evidence for multiple metabolic functions in the RidA/YjgF/YER057c/UK114 (Rid) protein family. BMC Genomics 16:382. doi: 10.1186/s12864-015-1584-3
Niehaus, T. D., Nguyen, T. N., Gidda, S. K., ElBadawi-Sidhu, M., Lambrecht, J. A., McCarty, D. R., et al. (2014). Arabidopsis and maize RidA proteins preempt reactive enamine/imine damage to branched-chain amino acid biosynthesis in plastids. Plant cell 26, 3010–3022. doi: 10.1105/tpc.114.126854
Palyada, K., Sun, Y. Q., Flint, A., Butcher, J., Naikare, H., and Stintzi, A. (2009). Characterization of the oxidative stress stimulon and PerR regulon of Campylobacter jejuni. BMC Genomics 10:481. doi: 10.1186/1471-2164-10-481
Parkhill, J., Wren, B. W., Mungall, K., Ketley, J. M., Churcher, C., Basham, D., et al. (2000). The genome sequence of the food-borne pathogen Campylobacter jejuni reveals hypervariable sequences. Nature 403, 665–668. doi: 10.1038/35001088
Reid, A. N., Pandey, R., Palyada, K., Whitworth, L., Doukhanine, E., and Stintzi, A. (2008). Identification of Campylobacter jejuni genes contributing to acid adaptation by transcriptional profiling and genome-wide mutagenesis. Appl. Environ. Microbiol. 74, 1598–1612. doi: 10.1128/AEM.01508-07
Reuter, M., Periago, P. M., Mulholland, F., Brown, H. L., and van Vliet, A. H. (2015). A PAS domain-containing regulator controls flagella-flagella interactions in Campylobacter jejuni. Front. Microbiol. 6:770. doi: 10.3389/fmicb.2015.00770
Sacher, J. C. (2018). Insights into the Role of the Flagellar Glycosylation System in Campylobacter jejuni Phage-Host Interactions. Ph.D thesis, University of Alberta, Edmonton, AB.
Schirm, M., Schoenhofenm, I., Logan, S. M., Wldron, K., and Thibault, P. (2005). Identification of unusual bacterial glycosylatin by tandem mass spectrometry analysis of intact proteins. Anal. Chem. 77, 7774–7782. doi: 10.1021/ac051316y
Schmitz, G., and Downs, D. M. (2004). Reduced Transaminase B (IlvE) activity caused by the lack of yjgF is dependent on the status of threonine deaminase (IlvA) in Salmonella enterica serovar typhimurium. J. Bacteriol. 186, 803–810. doi: 10.1128/jb.186.3.803-810.2004
Tan, S., and Berg, D. E. (2004). Motility of urease-deficient derivatives of Helicobacter pylori. J. Bacteriol. 186, 885–888. doi: 10.1128/jb.186.3.885-888.2004
Taveirne, M. E., Theriot, C. M., Livny, J., and DiRita, V. J. (2013). The complete Campylobacter jejuni transcriptome during colonization of a natural host determined by RNAseq. PLoS One 8:e73586. doi: 10.1371/journal.pone.0073586
Thibault, P., Logan, S. M., Kelly, J. F., Brisson, J. R., Ewing, C. P., Trust, T. J., et al. (2001). Identification of the carbohydrate moieties and glycosylation motifs in Campylobacter jejuni flagellin. J. Biol. Chem. 276, 34862–34870.
Ulasi, G. N., Creese, A. J., Hui, S. X., Penn, C. W., and Cooper, H. J. (2015). Comprehensive mapping of O-glycosylation in flagellin from Campylobacter jejuni 11168: a multienzyme differential ion mobility mass spectrometry approach. Proteomics 15, 2733–2745. doi: 10.1002/pmic.201400533
VanDrisse, C. M., and Escalante-Semerena, J. C. (2016). New high-cloning-efficiency vectors for complementation studies and recombinant protein overproduction in Escherichia coli and Salmonella enterica. Plasmid 86, 1–6. doi: 10.1016/j.plasmid.2016.05.001
Vogel, H. J., and Bonner, D. M. (1956). Acetylornithase of Escherichia coli: partial purification and some properties. J. Biol. Chem. 218, 97–106.
Vorwerk, H., Mohr, J., Huber, C., Wensel, O., Schmidt-Hohagen, K., Gripp, E., et al. (2014). Utilization of host-derived cysteine-containing peptides overcomes the restricted sulphur metabolism of Campylobacter jejuni. Mol. Microbiol. 93, 1224–1245. doi: 10.1111/mmi.12732
Woodall, C. A., Jones, M. A., Barrow, P. A., Hinds, J., Marsden, G. L., Kelly, D. J., et al. (2005). Campylobacter jejuni gene expression in the chick cecum: evidence for adaptation to a low-oxygen environment. Infect. Immun. 73, 5278–5285. doi: 10.1128/iai.73.8.5278-5285.2005
Zebian, N., Merkx-Jacques, A., Pittock, P. P., Houle, S., Dozois, C. M., Lajoie, G. A., et al. (2016). Comprehensive analysis of flagellin glycosylation in Campylobacter jejuni NCTC 11168 reveals incorporation of legionaminic acid and its importance for host colonization. Glycobiology 26, 386–397. doi: 10.1093/glycob/cwv104
Keywords: RidA, Cj1388, motility, autoagglutination, flagella, 2-aminoacrylate, Campylobacter jejuni
Citation: Irons J, Sacher JC, Szymanski CM and Downs DM (2019) Cj1388 Is a RidA Homolog and Is Required for Flagella Biosynthesis and/or Function in Campylobacter jejuni. Front. Microbiol. 10:2058. doi: 10.3389/fmicb.2019.02058
Received: 08 April 2019; Accepted: 20 August 2019;
Published: 06 September 2019.
Edited by:
Hua Xiang, Institute of Microbiology (CAS), ChinaReviewed by:
Ximin Zeng, The University of Tennessee, Knoxville, United StatesBeile Gao, South China Sea Institute of Oceanology (CAS), China
Copyright © 2019 Irons, Sacher, Szymanski and Downs. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Diana M. Downs, ZG1kb3duc0B1Z2EuZWR1
 Jessica Irons
Jessica Irons Jessica C. Sacher
Jessica C. Sacher Christine M. Szymanski1,2,3
Christine M. Szymanski1,2,3 Diana M. Downs
Diana M. Downs