Abstract
Candida auris is an emerging multi-drug resistant yeast, that causes major issues regarding patient treatment and surface disinfection in hospitals. Indeed, an important proportion of C. auris strains isolated worldwide present a decreased sensitivity to multiple and sometimes even all available antifungals. Based on recent tentative breakpoints by the CDC, it appears that in the USA about 90, 30, and < 5% of isolates have been resistant to fluconazole, amphotericin B, and echinocandins, respectively. To date, this has lead to a low therapeutic success. Furthermore, C. auris is prone to cause outbreaks, especially since it can persist for weeks in a nosocomial environment and survive high-end disinfection procedures. In this review, we describe the molecular resistance mechanisms to antifungal drugs identified so far in C. auris and compare them to those previously discovered in other Candida species. Additionally, we examine the role that biofilm formation plays in the reduced antifungal sensitivity of this organism. Finally, we summarize the few insights on how this yeast survives on hospital surfaces and discuss the challenge it presents regarding nosocomial environment disinfection.
1. Introduction
In the last decade, an emerging nosocomial Candida species has been causing severe cases of candidemia and other types of fungal infections. Outbreaks of the responsible pathogen, namely Candida auris, have been observed all around the world (see Figure 1A) (Lockhart et al., 2017).
Figure 1
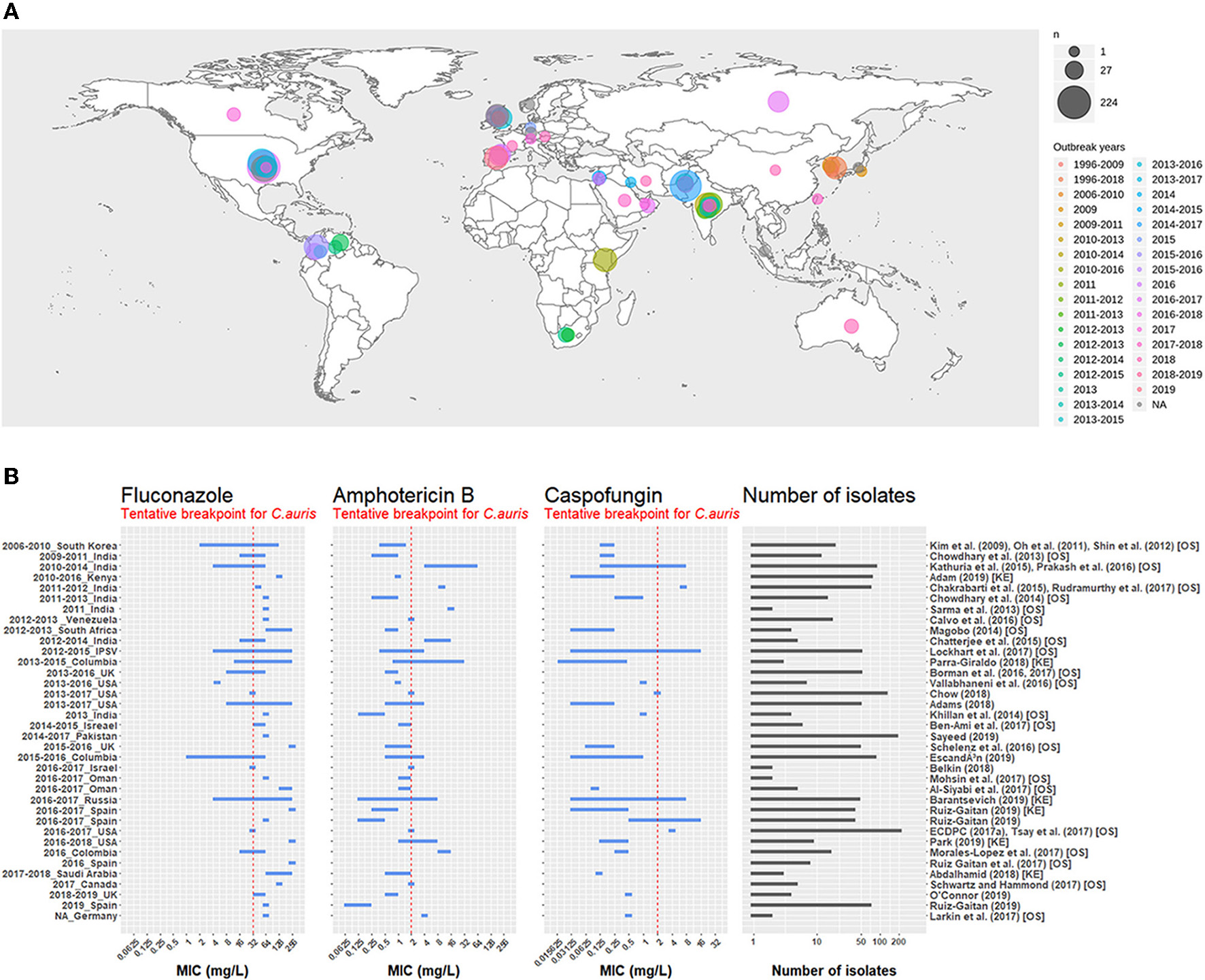
(A) Map representing the number of isolates per outbreak since 1996. The first outbreak year is brown-colored and the latest outbreak year is in pink. Circle size is proportional to the number of isolates reported for each outbreak. Outbreak survey was performed recovering data from three main reviews (Osei Sekyere, 2018; Jackson et al., 2019; Kenters et al., 2019) and complemented by a pubmed search using the query “Candida” AND “auris” AND “outbreak” as keywords. Results were limited in time, between the 21/07/2017, the latest study in the systematic review by Osei Sekyere and the 31/08/2019, when the literature search was performed. Studies not present in the review by Osei Sekyere: (Biswal et al., 2017; Abdalhamid et al., 2018; Adams et al., 2018; Belkin et al., 2018; Chow et al., 2018; Desoubeaux et al., 2018; Lesho et al., 2018; Mayor, 2018; Parra-Giraldo et al., 2018; Ruiz-Gaitán et al., 2018, 2019a,b; Sexton et al., 2018; Adam et al., 2019; Barantsevich et al., 2019; Escandón et al., 2019; Leach et al., 2019; O'Connor et al., 2019; Park et al., 2019; Sana et al., 2019; Sayeed et al., 2019). (B) MIC ranges for three antifungals measured in C. auris isolates from different outbreaks around the world. The red dotted line represents the tentative breakpoint proposed by the CDC. IPSV is an acronym for “India, Pakistan, South Africa, and Venezuela,” as strains from these countries were all analyzed in the same study. Outbreak survey was performed identically to (A). Only outbreak with >1 isolate were plotted. Studies cited by Osei Sekyere are signaled with [OS] and studies cited by Kenters et al. with [KE].
Whole genome sequencing determined the presence of four specific clades within the C. auris species: East Asia, South Asia, South Africa, and South America. All clades are characterized by distinct single nucleotide polymorphisms (SNPs), which highlights the independent and worldwide emergence of this pathogen (Lockhart et al., 2017).
Comparative studies in different animal models (murine and Galleria mellonella larvae) showed strain-dependant virulence levels for C. auris. In most cases, C.auris strains proved to be less virulent than C. albicans (Borman et al., 2016; Ben-Ami et al., 2017; Fakhim et al., 2018; Wang et al., 2018). However, reported cases of C. auris outbreaks were linked with a high mortality rate, primarily in immunocompromised patients. Other conditions such as diabetes, cardiovascular and/or pulmonary diseases, sepsis or a prior treatment with antibiotics seemed also to be important risk factors (Osei Sekyere, 2018).
Exceptionally for a yeast, C. auris has successfully spread in nosocomial environments. For example, in a UK hospital, a single patient colonized with C. auris led to subsequent identifications of other cases among other hospitalized patients. The pathogen was also detected on the healthcare staff even several months after its first isolation, suggesting an efficient human to human transmission. Additionally, traces of C. auris had been found on bed sides, windowsills, monitors, and other equipment throughout the hospital, showing that C. auris is capable to persist on nosocomial surfaces (Schelenz et al., 2016), in contrast to other Candida species. Investigation into an outbreak in another UK hospital reported that the use of reusable equipment such as axillary temperature probes was the primary cause of transmission to patients (Eyre et al., 2018), further illustrating its capability to survive on surfaces.
The capacity to survive in nosocomial environments could be due to the ability of C. auris to form biofilms; a growth form in which cells are arranged together in microcolonies protected by a matrix of glucan. By convention, cells enclosed in the matrix are called sessile, and cells outside of it are called planktonic. C. auris cells protected by this structure were proven to be difficult to eliminate, even with high-levels of disinfection procedures (Kean et al., 2018b).
In addition to potentially allowing an increased survival on inert surfaces, the ability of C. auris to form biofilms may also be an important factor of its resistance to systemic antifungals used in patient treatment. Indeed, the fact that biofilm forming cells were isolated from wounds and catheter tips indicates that some C. auris cells could survive in biofilm-form in patients (Borman et al., 2016) and it was shown that sessile cells presented reduced susceptibility for several antifungals. (Sherry et al., 2017). Finally, biofilm has also been shown to be an important virulence factor, as biofilm forming C. auris strains are linked to increased morbidity and mortality (Borman et al., 2016).
With this review we aim to highlight molecular resistance mechanisms acquired by C. auris, which led to its extraordinary survival in nosocomial environment and in hospitalized patients.
2. C. auris Antifungal Susceptibility
After 10 years of recording minimum inhibitory concentration (MIC) for available antifungals, the CDC recently published a tentative of breakpoints to better define C. auris antifungal susceptibility1. It clearly appeared that C. auris has exceptionally high MICs for the four main classes of antifungal drugs: azoles, echinocandins, polyenes (Figure 1B), and nucleoside analogs. In this chapter, we will examine in detail two factors that proved to play a role in this decreased susceptibility to antifungal treatments: molecular resistance mechanisms and biofilm-associated resistance mechanisms.
2.1. Molecular Resistance Mechanisms
We define here molecular resistance as the ways individual cells can decrease their sensitivity to antifungals due to genetic modifications leading to drug target modification, efflux pumps overexpression, or metabolism modifications (see for review: Pemán et al., 2009; Sanglard et al., 2009; Krishnasamy et al., 2018). This section compiles, for each class of antifungal, the main molecular resistance mechanisms found in C. auris up to now. Additionally, Table 1 summarizes for each drug, the mode of action, the related resistance mechanisms described in non-auris Candida and the resistance mechanisms discovered in C. auris, as of yet.
Table 1
| Antifungal class | Target | Effect | Resistance mechanism(s) in non-auris Candida spp. | Resistance mechanism(s) in Candida auris |
|---|---|---|---|---|
| Azoles | Inhibits Lanosterol 14-alpha -demethylase (LD), the protein catalyzing the conversion of lanosterol to ergosterol | Weaker cell membrane, inhibiting growth and sensitizing to host immune system Accumulation of toxic intermediary compound | MSF efflux pumps overexpression due to mutation in Mrr1 transcription factor (TF). [a] CDR genes overexpression, encoding ABC efflux pumps, due to mutation in Tac1 TF. [a] Point mutation in gene encoding LD (ERG11) Upregulation of ERG11 due to mutation in Upc2 TF. [a] ERG3 mutation:Bypass ergosterol synthesis pathway to prevent toxic compound accumulation | Presence of MFS efflux pumps genes [b]. Higher ABC efflux pumps activity compared to C.glabrata. [c] Significant contribution of CDR1 to resistance. [d] ERG11 point mutation : Y132F ; [f] K143R [f] Presence of ERG3 [e] but no mutation associated with increased resistance reported |
| Echinocandin | Inhibits beta(1,3)D-glucan synthase | Defective cell wall formation | Mutation in FKS1 and FKS2 gene, encoding the target enzyme | Presence of FKS1 and FKS2 In FKS1 three mutations linked to increased MIC: S639Y; [g] S639P; [g] S639F. [h] |
| Polyene | Binds ergosterol molecules in the cytoplasmic membrane | Affects membrane permeability by forming multimeric pores and causes oxidative damage | Membrane composition alteration to reduce target exposition Increase catalase activity to counter oxidative damage | 5 SNPs in different genomic loci associated with reduced sensitivity. [i] |
| Nucleoside analogs | Transformed in the fungal cell in two products that respectively inhibit thymidine synthase and get incorporated in RNA | DNA and RNA synthesis inhibition | Modification of enzymes involved in 5-FC uptake or metabolism Competitive inhibition of 5-FC effect by increased pyrimidine synthesis | Amino-acid substitution (F211I) in FUR1, a gene involved in 5-FC metabolism found in a single flucytosine- resistant isolate [g] |
Pharmacodynamics of the four main antifungal classes and linked resistance mechanisms in Candida spp. in general and C. auris specifically.
Adapted from Krishnasamy et al. (2018). With additional information from: [a] Vandeputte et al. (2012), [b] Sharma et al. (2016), [c] Ben-Ami et al. (2017), [d] Chowdhary et al. (2018), [e] Rybak et al. (2019), [f] Healey et al. (2018), [g] Rhodes et al. (2018), [h] Kordalewska et al. (2018), [i] Escandón et al. (2019).
2.1.1. Mechanisms of Resistance to Azoles
2.1.1.1. MFS and ABC efflux pumps overexpression
Efflux pumps are proteins that transport components across the cell membrane. Some of them can pump drugs outside of the cell, lowering their concentration and effect on the cell. There are two major families of efflux pumps involved in antifungal resistance: ATP Binding Cassette (ABC) and Major Facilitator Superfamily (MFS) transporters. Enhanced overexpression of efflux pumps is one of the major resistance mechanism to azoles in pathogenic Candida species (Schuetzer-Muehlbauer et al., 2003; Morschhäuser et al., 2007).
Indian strains of C. auris sequenced in two different studies exhibit numerous genes orthologous to ABC and MFS transporters of C. albicans (Chatterjee et al., 2015; Sharma et al., 2016). Interestingly, C. auris strains that were isolated in Israel showed an intrinsicly high ABC transporter activity compared to C. glabrata and C. haemulonii, by measuring efflux of a fluorescent substrate (Rhodamine 6G) (Ben-Ami et al., 2017).
In C. albicans, CDR1 is a gene encoding for an ABC efflux pump, known for its role in azole resistance (Sanglard et al., 1995). A gene homologous to CDR1 was found in C. auris. (Rybak et al., 2019). The same study also showed that the deletion of this gene could increase susceptibility of resistant strains from 64- to 128-fold (Rybak et al., 2019).
2.1.1.2. Point mutations in ERG11
Ergosterol is a key membrane component in fungi. In Candida spp., its biosynthesis is mediated, among others, by the enzyme Lanosterol 14-alpha-demethylase (LD), encoded by the gene ERG11 which converts lanosterol to ergosterol. The LD is the primary target of azoles, antifungals that inhibit the function of the enzyme and effectively shut down ergosterol biosynthesis, thus impairing membrane integrity (Sanglard et al., 1998).
Point mutations in ERG11 have been shown to reduce azole sensitivity in Candida spp., particularly in three “hot-spot” regions located between amino-acids 105–165, 266–287, and 405–488 (Vandeputte et al., 2012).
Interestingly, after having retrieved the amino acid sequences of 44 C. auris isolates from India, 15 missense mutations were found when aligned to the C. albicans wild-type ERG11 sequence (Chowdhary et al., 2018). Five of these mutations were already associated with azole resistance in C. albicans. Among these variants two were found in every resistant strain tested in this study, namely Y132F or K143R (Chowdhary et al., 2018). It can also be noted that these mutations are in the first “hot spot” discussed earlier.
A screen for variants in the ERG11 gene of C. auris strains from Columbia identified the same Y132F and K143R substitutions. Upon heterologous expression of these two mutations in S. cerivisiae, MICs for azoles presented a two-fold increase as compared to S. cerevisiae strains expressing the wild-type C. auris ERG11 gene (Healey et al., 2018).
2.1.1.3. ERG11 overexpression
In C. albicans, an overexpression of ERG11 has been linked to a resistance to azole treatment as well.The increased production of LD overwhelms the capacity of the antifungal to inhibit the activity of the protein, resulting in an active protein despite drug treatment (Lopez-Ribot et al., 1998).
In C. auris, real time PCR experiments showed that in absence of fluconazole, there was no difference in ERG11 expression between fluconazole-susceptible and fluconazole-resistant strains (Chowdhary et al., 2018). In presence of the antifungal, ERG11 expression rises in resistant strains when compared to a control without fluconazole (Chowdhary et al., 2018). However, the effect of this drug on ERG11 expression was not tested on susceptible strains. Therefore, conditionally increased expression of ERG11 in presence of fluconazole has not been demonstrated to be specific to resistant strains, until now.
2.1.2. Mechanisms of Resistance to Echinocandins
Beta(1,3)D-glucan is a key component of the fungal cell wall and is encoded by the FKS1 and FKS2 genes, two subunits of the Beta(1,3)D-glucan synthase. Echinocandins inhibit this enzyme, decreasing the amount of glucans in the cell wall (Martins et al., 2011).
In C. albicans and other non-auris Candida species, several mutations leading to echinocandin resistance were found in the same two regions of FKS1 and FKS2. So, they were named “hot-spots” 1 and 2 (HS1 and HS2). In the C. albicans FKS1 gene these “hot-spots” are located between amino acids 641–649 and 1,345–1,365 (Park et al., 2005). Sequencing of the corresponding hot-spot regions of 38 C. auris strains lead to the discovery of an S639F amino acid substitution that is correlated with pan-echinocandin resistance: it was present in all four resistant strains and absent in the 34 susceptible ones. This position in C. auris FKS1 is aligned to the position 645 (also a Serine) in C. albicans (Chowdhary et al., 2018). Interestingly, this mutation is in the region aligning to the HS1 of C. albicans FKS1. Other studies observed different mutations at the same location in echinocandin-resistant C. auris strains: S639Y and S639P (Rhodes et al., 2018). The latter was confirmed to cause echinocandin resistance in vivo in a mouse model (Kordalewska et al., 2018).
FKS2 was also found in a single copy in the C. auris genome (Sharma et al., 2016), but no mutation associated with echinocandin resistance were found in this gene to our knowledge.
2.1.3. Mechanisms of Resistance to Polyenes
As shown in the Figure 1B, several C. auris strains are resistant to Amphotericin B (AMB). In the Candida species, modifications of the sterol composition of the membrane have been highlighted as a resistance mechanism (Haynes et al., 1996; Nolte et al., 1997). In C. albicans, mutations in ERG 2, 3, 5, 6 or 11 have been shown to have this effect (Arendrup and Patterson, 2017). Rhodes et al. screened 27 C. auris isolates from the UK for SNPs in these genes in strains displaying reduced sensitivity to AMB. However, no variants explaining these differences in drug susceptibility were found (Rhodes et al., 2018).
2.1.4. Mechanisms of Resistance to Flucytosine (5-Fluorocytosine)
Flucytosine is a nucleoside analog that inhibits nucleic acid synthesis. After cell entry, flucytosine has to be activated to have an antifungal effect. This activation requires, among others, the protein encoded by the gene FUR1 Waldorf and Polak (1983). In non-auris Candida species mutations in FUR1 were demonstrated to be linked with flucytosine resistance (Vandeputte et al., 2012).
Rhodes et al. sequenced a flucytosine resistant C. auris strain and observed a F211I amino acid substitution in the FUR1 gene (Rhodes et al., 2018). This specific missense mutation has no known equivalent in other Candida species, therefore additional studies are required to determine if this mutation is the cause of the resistance to flucytosine in the tested C. auris strain (Rhodes et al., 2018). However, this drug is less employed than other antifungals. Because of that, fewer studies have been performed to better understand the resistance of C. auris to this compound.
To summarize, some resistance mechanisms previously discovered in Candida species like C. glabrata and C. albicans, were also showcased by C. auris. They are listed in the last column of the Table 1. However, C. auris antifungal resistance is only partially explained by these previously described resistance mechanisms. In order to eventually design new therapeutic agents and more generally to improve patient care, further studies are needed to better understand the underlying mechanisms leading to the exceptionally high MIC ranges presented by some C. auris strains (see Figure 1B).
2.2. Biofilm: An Antifungal Resistant Way of Life
It was shown that MICs for several antifungals are higher in C. auris sessile cells (up to 4-, 20-, 60-fold for voriconazole, Amphotericin B, and Micafungin, respectively) compared to planktonic cells (Sherry et al., 2017). Similarly, minimal biofilm eradication concentrations were shown to be 512-fold higher than MICs for echinocandins and azoles (Romera et al., 2019). This phenomenon was previously shown for C. albicans (Hawser and Douglas, 1995). The molecular mechanisms responsible for these elevated MIC are still largely unknown in C. auris but some studies have already provided some insights.
MSF and ABC efflux pump-encoding genes were found to be upregulated (2- to 4-fold) in sessile cells compared to planktonic cells (Kean et al., 2018a). Concordantly, the activity of corresponding proteins was 2-fold increased. However, when treated with efflux pump inhibitors the antifungal sensitivity of sessile cells was increased 4- to 16-fold after 12 h. This suggests that efflux pumps play an important role in the resistance to antifungals displayed by this type of cells (Kean et al., 2018a).
In C. albicans, the exopolymer matrix is known to non-specifically bind to all classes of antifungals and sequestrate them outside the cells (Nett et al., 2010). As Candida spp. share a core polysaccharide profile, it was suggested that this mechanism might be conserved in C. auris as well (Kean et al., 2018a). This was later confirmed in in vivo and in vitro assays (Dominguez et al., 2019).
3. Nosocomial Persistence
The capacity of C. auris to colonize and persist on surfaces is exceptional: it is able to persist longer on moist surfaces in comparison to C. albicans (Piedrahita et al., 2017). It also has a prolonged metabolic activity on surfaces, akin to C. parapsilosis, a known skin and plastic colonizer (Piedrahita et al., 2017; Welsh et al., 2017).
The increased survival of C. auris on surfaces was investigated by two studies. The first claimed it could be linked to an enhanced environmental stress resistance to temperature and other stressors (Kean et al., 2018b). The second hypothesizes that it could be due to the formation of biofilm. However, this remains to be proven as, so far, no biofilm forming isolate was recovered from environmental surfaces (Ku et al., 2018). It is to be noted that these two possibilities are not mutually exclusive, and that biofilm formation might even be the cause of the increased stress resistance (Ku et al., 2018).
The persistence of C. auris in nosocomial environments is heightened by the resistance of this pathogen to disinfection procedures. The efficacy of NaOCl and peracetic acid on stainless steel, polymer (polyester coverslips) and cellulose surfaces were tested. Both disinfectants showed a significant efficacy to kill C. auris cells on all surfaces. However, some viable cells remained after applying NaOCl on non-porous surfaces (stainless steel and polyester coverslips). In fact, higher concentrations of this disinfectant coupled to longer exposure times were required to lower regrowth, but even then, they were not able to completely eradicate the pathogen. For example, a 10,000 parts per million (ppm) NaOCl concentration coupled to a 5 min exposure on stainless steel, was enough to observe a significant reduction of colonies. However, substantial regrowth was observed after re-inoculation onto a rich medium (Kean et al., 2018b).
For peracetic acid, which is a high-level disinfectant, this regrowth was surface dependent, as no regrowth was observed for polymer as opposed to stainless steel. They performed the same experiments on C. glabrata and C. albicans and obtained similar results (Kean et al., 2018b).
This shows that high-level disinfection procedures may be insufficient for a complete surface cleansing and that the pre-established guidelines for surface disinfection have to be adapted to these Candida species (Kean et al., 2018b). Factors like the type of surfaces and exposure times should be considered when implementing these disinfection procedures (Kean et al., 2018b; Ku et al., 2018) not only cost and ease of use, as is it often the case (Ku et al., 2018).
Based on the results of two studies (Cadnum et al., 2017; Rutala et al., 2019), the CDC recommends using disinfectants effective against Clostridioides difficile spores. If not possible, they suggest alternatives such as hydrogen peroxyde 0.5–1.4% or quaternary ammonium compounds supplemented with isopropyl alcohol and/or ethyl alcohol2. Finally, ultraviolet-C light could also be an interesting candidate for surfaces disinfection. It proved highly effective at killing C. auris colonies, given sufficient exposure time and distance. However, further studies would be necessary before implementing such methods in hospital settings (de Groot et al., 2019).
4. Conclusion
The pathogenic and virulent capacities of C. auris are profoundly worrying. Especially since outbreaks occurred independently in different regions of the world and that the associated isolates showed decreased susceptibility to the most commonly used antifungal drugs used for patient treatment (see Figure 1).
To explain the reduced antifungal susceptibility in C. auris, researchers made use of the extended knowledge already available on other Candida species, particularly C. albicans. As a result, some of the resistance mechanisms of C. auris were highlighted (see Table 1). More precisely, two point mutations in ERG11 and the overexpression of the ABC transporter Cdr1 were proven to decrease fluconazole sensitivity. Furthermore, an amino acid substitution in FKS1 was shown to reduce the sensitivity of C. auris to echinocandins, which is normally the drug of choice for treatment against this pathogen (Bidaud et al., 2018). Finally, a mutation in FUR1 was identified in a flucytosine resistant C. auris strain, but remains to be demonstrated as the cause of this reduced sensitivity. Biofilm is thought to be a resistance mechanism per se as it was shown to decrease drug susceptibility. However, further research is still necessary to describe the whole variety of processes that take place in this growth form. It is important to note that biofilm is a complex structure that remains poorly understood and difficult to investigate, even in well-known species such as C. albicans.
The resistance properties of C. auris are coupled to a propensity to cause nosocomial infections. Its ability to adhere to a variety of nosocomial surfaces and survive disinfection are critical propagation factors. The presence of multiple types of surfaces poses a challenge for clinical institutions, as standard disinfection procedures are inefficient and unadapted for each type of them. This problem, also arises from the insufficient research on this subject. In addition, a better understanding of the mechanisms underlying C. auris disinfectant resistance would help to improve hygiene protocols and avoid future nosocomial outbreaks.
Statements
Author contributions
FC, AG, and LJ participated equally in the redaction of this review. AC supervised and participated in the redaction of this review.
Acknowledgments
We would like to thank Clara Heiman as a native English speaker to have revised our manuscript.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Footnotes
1.^CDC (2019). Antifungal Susceptibility Testing and Interpretation. Available online at: https://www.cdc.gov/fungal/candida-auris/c-auris-antifungal.html (accessed April 29, 2019).
2.^CDC (2018). Infection Prevention and Control for Candida auris. Available online at: https://www.cdc.gov/fungal/candida-auris/c-auris-infection-control.html (accessed November 12, 2019).
References
1
AbdalhamidB.AlmaghrabiR.AlthawadiS.OmraniA. (2018). First report of Candida auris infections from Saudi Arabia. J. Infect. Public Health11, 598–599. 10.1016/j.jiph.2018.05.010
2
AdamR. D.RevathiG.OkindaN.FontaineM.ShahJ.KagothoE.et al. (2019). Analysis of Candida auris fungemia at a single facility in Kenya. Int. J. Infect. Dis.85, 182–187. 10.1016/j.ijid.2019.06.001
3
AdamsE.QuinnM.TsayS.PoirotE.ChaturvediS.SouthwickK.et al. (2018). Candida auris in healthcare facilities, New York, USA, 2013-2017. Emerg. Infect. Dis.24, 1816–1824. 10.3201/eid2410.180649
4
ArendrupM. C.PattersonT. F. (2017). Multidrug-resistant candida: epidemiology, molecular mechanisms, and treatment. J. Infect. Dis.216, S445–S451. 10.1093/infdis/jix131
5
BarantsevichN. E.OrlovaO. E.ShlyakhtoE. V.JohnsonE. M.WoodfordN.Lass-FloerlC.et al. (2019). Emergence of Candida auris in Russia. J. Hosp. Infect.102, 445–448. 10.1016/j.jhin.2019.02.021
6
BelkinA.GazitZ.KellerN.Ben-AmiR.Wieder-FinesodA.NovikovA.et al. (2018). Candida auris infection leading to nosocomial transmission, Israel, 2017. Emerg. Infect. Dis.24, 801–804. 10.3201/eid2404.171715
7
Ben-AmiR.BermanJ.NovikovA.BashE.Shachor-MeyouhasY.ZakinS.et al. (2017). Multidrug-resistant Candida haemulonii and C. auris, Tel Aviv, Israel. Emerg. Infect. Dis.23, 195–203. 10.3201/eid2302.161486
8
BidaudA. L.ChowdharyA.DannaouiE. (2018). Candida auris: an emerging drug resistant yeast – A mini-review. J. Mycol. Méd.28, 568–573. 10.1016/j.mycmed.2018.06.007
9
BiswalM.RudramurthyS. M.JainN.ShamanthA. S.SharmaD.JainK.et al. (2017). Controlling a possible outbreak of Candida auris infection: lessons learnt from multiple interventions. J. Hosp. Infect.97, 363–370. 10.1016/j.jhin.2017.09.009
10
BormanA. M.SzekelyA.JohnsonE. M. (2016). Comparative pathogenicity of United Kingdom isolates of the emerging pathogen Candida auris and other key pathogenic candida species. mSphere1:e00189–16. 10.1128/mSphere.00189-16
11
CadnumJ. L.ShaikhA. A.PiedrahitaC. T.SankarT.JencsonA. L.LarkinE. L.et al. (2017). Effectiveness of disinfectants against Candida auris and other candida species. Infect. Control Hosp. Epidemiol.38, 1240–1243. 10.1017/ice.2017.162
12
ChatterjeeS.AlampalliS. V.NageshanR. K.ChettiarS. T.JoshiS.TatuU. S. (2015). Draft genome of a commonly misdiagnosed multidrug resistant pathogen Candida auris. BMC Genomics16:686. 10.1186/s12864-015-1863-z
13
ChowN. A.GadeL.TsayS. V.ForsbergK.GreenkoJ. A.SouthwickK. L.et al. (2018). Multiple introductions and subsequent transmission of multidrug-resistant Candida auris in the USA: a molecular epidemiological survey. Lancet Infect. Dis.18, 1377–1384. 10.1016/S1473-3099(18)30597-8
14
ChowdharyA.PrakashA.SharmaC.KordalewskaM.KumarA.SarmaS.et al. (2018). A multicentre study of antifungal susceptibility patterns among 350 Candida auris isolates (2009–17) in India: role of the ERG11 and FKS1 genes in azole and echinocandin resistance. J. Antimicrob. Chemother.73, 891–899. 10.1093/jac/dkx480
15
de GrootT.ChowdharyA.MeisJ. F.VossA. (2019). Killing of Candida auris by UV-C: importance of exposure time and distance. Mycoses62, 408–412. 10.1111/myc.12903
16
DesoubeauxG.BaillyE.GuillaumeC.De KyvonM.-A.TellierA.-C.MorangeV.et al. (2018). Candida auris in contemporary mycology labs: a few practical tricks to identify it reliably according to one recent French experience. J. Mycol. Med.28, 407–410. 10.1016/j.mycmed.2018.02.011
17
DominguezE. G.ZarnowskiR.ChoyH. L.ZhaoM.SanchezH.NettJ. E.et al. (2019). Conserved role for biofilm matrix polysaccharides in Candida auris drug resistance. mSphere4:e00680–18. 10.1128/mSphereDirect.00680-18
18
EscandónP.ChowN. A.CaceresD. H.GadeL.BerkowE. L.ArmstrongP.et al. (2019). Molecular epidemiology of Candida auris in colombia reveals a highly related, countrywide colonization with regional patterns in amphotericin B resistance. Clin. Infect. Dis.68, 15–21. 10.1093/cid/ciy411
19
EyreD. W.SheppardA. E.MadderH.MoirI.MoroneyR.QuanT. P.et al. (2018). A Candida auris outbreak and its control in an intensive care setting. N. Engl. J. Med.379, 1322–1331. 10.1056/NEJMoa1714373
20
FakhimH.VaeziA.DannaouiE.ChowdharyA.NasiryD.FaeliL.et al. (2018). Comparative virulence of Candida auris with Candida haemulonii, Candida glabrata and Candida albicans in a murine model. Mycoses61, 377–382. 10.1111/myc.12754
21
HawserS. P.DouglasL. J. (1995). Resistance of Candida albicans biofilms to antifungal agents in vitro. Antimicrob. Agents Chemother.39, 2128–2131. 10.1128/AAC.39.9.2128
22
HaynesM. P.ChongP. L.-G.BuckleyH. R.PieringerR. A. (1996). Fluorescence studies on the molecular action of amphotericin B on susceptible and resistant fungal cells. Biochemistry35, 7983–7992. 10.1021/bi952910c
23
HealeyK. R.KordalewskaM.OrtigosaC. J.SinghA.BerríoI.ChowdharyA.et al. (2018). Limited ERG11 mutations identified in isolates of Candida auris directly contribute to reduced azole susceptibility. Antimicrob. Agents Chemother.62, e01427–18. 10.1128/AAC.01427-18
24
JacksonB. R.ChowN.ForsbergK.LitvintsevaA. P.LockhartS. R.WelshR.et al. (2019). On the origins of a species: what might explain the rise of Candida auris?J. Fungi5:E58. 10.3390/jof5030058
25
KeanR.DelaneyC.SherryL.BormanA.JohnsonE. M.RichardsonM. D.et al. (2018a). Transcriptome assembly and profiling of Candida auris reveals novel insights into biofilm-mediated resistance. mSphere3:e00334–18. 10.1128/mSphere.00334-18
26
KeanR.SherryL.TownsendE.McKloudE.ShortB.AkinbobolaA.et al. (2018b). Surface disinfection challenges for Candida auris: an in-vitro study. J. Hosp. Infect.98, 433–436. 10.1016/j.jhin.2017.11.015
27
KentersN.KiernanM.ChowdharyA.DenningD. W.PemánJ.SarisK.et al. (2019). Control of Candida auris in healthcare institutions. Outcome of an ISAC expert meeting. Int. J. Antimicrob. Agents54, 400–406. 10.1016/j.ijantimicag.2019.08.013
28
KordalewskaM.LeeA.ParkS.BerrioI.ChowdharyA.ZhaoY.et al. (2018). Understanding echinocandin resistance in the emerging pathogen Candida auris. Antimicrob. Agents Chemother.62:e00238–18. 10.1128/AAC.00238-18
29
KrishnasamyL.KrishnakumarS.KumaramanickavelG.SaikumarC. (2018). Molecular mechanisms of antifungal drug resistance in Candida species. J. Clin. Diagn. Res. 12, DE01–DE06. 10.7860/JCDR/2018/36218.11961
30
KuT. S. N.WalravenC. J.LeeS. A. (2018). Candida auris: disinfectants and implications for infection control. Front. Microbiol.9:726. 10.3389/fmicb.2018.00726
31
LeachL.RussellA.ZhuY.ChaturvediS.ChaturvediV. (2019). A rapid and automated sample-to-result Candida auris real-time PCR assay for high-throughput testing of surveillance samples with BD MAX™ open system. J. Clin. Microbiol. 57:e00630–19. 10.1128/JCM.00630-19
32
LeshoE. P.BronsteinM. Z.McGannP.StamJ.KwakY.MaybankR.et al. (2018). Importation, mitigation, and genomic epidemiology of Candida auris at a large teaching hospital. Infect. Control Hosp. Epidemiol.39, 53–57. 10.1017/ice.2017.231
33
LockhartS. R.EtienneK. A.VallabhaneniS.FarooqiJ.ChowdharyA.GovenderN. P.et al. (2017). Simultaneous emergence of multidrug-resistant Candida auris on 3 continents confirmed by whole-genome sequencing and epidemiological analyses. Clin. Infect. Dis.64, 134–140. 10.1093/cid/ciw691
34
Lopez-RibotJ. L.McAteeR. K.LeeL. N.KirkpatrickW. R.WhiteT. C.SanglardD.et al. (1998). Distinct patterns of gene expression associated with development of fluconazole resistance in serial Candida albicans isolates from human immunodeficiency virus-infected patients with oropharyngeal candidiasis. Antimicrob. Agents Chemother.42, 2932–2937. 10.1128/AAC.42.11.2932
35
MartinsI. M.CortésJ. C. G.MuñozJ.MorenoM. B.RamosM.Clemente-RamosJ. A.et al. (2011). Differential activities of three families of specific beta(1,3)glucan synthase inhibitors in wild-type and resistant strains of fission yeast. J. Biol. Chem.286, 3484–3496. 10.1074/jbc.M110.174300
36
MayorS. (2018). Candida auris outbreak was linked to reusable axillary temperature probes, study finds. BMJ363:k4133. 10.1136/bmj.k4133
37
MorschhäuserJ.BarkerK. S.LiuT. T.Bla-WarmuthJ.HomayouniR.RogersP. D. (2007). The transcription factor Mrr1p controls expression of the MDR1 efflux pump and mediates multidrug resistance in Candida albicans. PLoS Pathog.3:e164. 10.1371/journal.ppat.0030164
38
NettJ. E.CrawfordK.MarchilloK.AndesD. R. (2010). Role of Fks1p and matrix glucan in Candida albicans biofilm resistance to an echinocandin, pyrimidine, and polyene. Antimicrob. Agents Chemother.54, 3505–3508. 10.1128/AAC.00227-10
39
NolteF. S.ParkinsonT.FalconerD. J.DixS.WilliamsJ.GilmoreC.et al. (1997). Isolation and characterization of fluconazole- and amphotericin B-resistant Candida albicans from blood of two patients with leukemia. Antimicrob. Agents Chemother.41, 196–199. 10.1128/AAC.41.1.196
40
O'ConnorC.BicanicT.DaveJ.EvansT. J.MoxeyP.AdamuU.et al. (2019). Candida auris outbreak on a vascular ward - the unexpected arrival of an anticipated pathogen. J. Hosp. Infect. 103, 106–108. 10.1016/j.jhin.2019.06.002
41
Osei SekyereJ. (2018). Candida auris: a systematic review and meta-analysis of current updates on an emerging multidrug-resistant pathogen. MicrobiologyOpen7:e00578. 10.1002/mbo3.578
42
ParkJ. Y.BradleyN.BrooksS.BurneyS.WassnerC. (2019). Management of patients with Candida auris fungemia at community hospital, Brooklyn, New York, USA, 2016–20181. Emerg. Infect. Dis.25, 601–602. 10.3201/eid2503.180927
43
ParkS.KellyR.KahnJ. N.RoblesJ.HsuM.-J.RegisterE.et al. (2005). Specific substitutions in the echinocandin target Fks1p account for reduced susceptibility of rare laboratory and clinical Candida sp. isolates. Antimicrob. Agents Chemother.49, 3264–3273. 10.1128/AAC.49.8.3264-3273.2005
44
Parra-GiraldoC. M.ValderramaS. L.Cortes-FraileG.GarzónJ. R.ArizaB. E.MorioF.et al. (2018). First report of sporadic cases of Candida auris in Colombia. Int. J. Infect. Dis.69, 63–67. 10.1016/j.ijid.2018.01.034
45
PemánJ.CantónE.Espinel-IngroffA. (2009). Antifungal drug resistance mechanisms. Expert Rev. Anti Infect. Ther.7, 453–460. 10.1586/eri.09.18
46
PiedrahitaC. T.CadnumJ. L.JencsonA. L.ShaikhA. A.GhannoumM. A.DonskeyC. J. (2017). Environmental surfaces in healthcare facilities are a potential source for transmission of Candida auris and other candida species. Infect. Control Hosp. Epidemiol.38, 1107–1109. 10.1017/ice.2017.127
47
RhodesJ.AbdolrasouliA.FarrerR. A.CuomoC. A.AanensenD. M.Armstrong-JamesD.et al. (2018). Genomic epidemiology of the UK outbreak of the emerging human fungal pathogen Candida auris. Emerg. Microbes Infect.7:43. 10.1101/201343
48
RomeraD.Aguilera-CorreaJ. J.GadeaI.Viñuela-SandovalL.García-RodríguezJ.EstebanJ. (2019). Candida auris: a comparison between planktonic and biofilm susceptibility to antifungal drugs. J. Med. Microbiol.68, 1353–1358. 10.1099/jmm.0.001036
49
Ruiz-GaitánA.MartínezH.MoretA. M.CalabuigE.TasiasM.Alastruey-IzquierdoA.et al. (2019a). Detection and treatment of Candida auris in an outbreak situation: risk factors for developing colonization and candidemia by this new species in critically ill patients. Expert Rev. Anti Infect. Ther.17, 295–305. 10.1080/14787210.2019.1592675
50
Ruiz-GaitánA.MoretA. M.Tasias-PitarchM.Aleixandre-LópezA. I.Martínez-MorelH.CalabuigE.et al. (2018). An outbreak due to Candida auris with prolonged colonisation and candidaemia in a tertiary care European hospital. Mycoses61, 498–505. 10.1111/myc.12781
51
Ruiz-GaitánA. C.CantónE.Fernández-RiveroM. E.RamírezP.PemánJ. (2019b). Outbreak of Candida auris in Spain: a comparison of antifungal activity by three methods with published data. Int. J. Antimicrob. Agents53, 541–546. 10.1016/j.ijantimicag.2019.02.005
52
RutalaW. A.KanamoriH.GergenM. F.Sickbert-BennettE. E.WeberD. J. (2019). Susceptibility of Candida auris and Candida albicans to 21 germicides used in healthcare facilities. Infect. Control Hosp. Epidemiol.40, 380–382. 10.1017/ice.2019.1
53
RybakJ. M.DoorleyL. A.NishimotoA. T.BarkerK. S.PalmerG. E.RogersP. D. (2019). Abrogation of triazole resistance upon deletion of CDR1 in a clinical isolate of Candida auris. Antimicrob. Agents Chemother.63, e00057–19. 10.1128/AAC.00057-19
54
SanaF.HussainW.ZamanG.SattiL.KhurshidU.KhadimM. T. (2019). Candida auris outbreak report from Pakistan: a success story of infection control in ICUs of a tertiary care hospital. J. Hosp. Infect. 103, 108–110. 10.1016/j.jhin.2019.06.011
55
SanglardD.CosteA.FerrariS. (2009). Antifungal drug resistance mechanisms in fungal pathogens from the perspective of transcriptional gene regulation. FEMS Yeast Res.9, 1029–1050. 10.1111/j.1567-1364.2009.00578.x
56
SanglardD.IscherF.KoymansL.BilleJ. (1998). Amino acid substitutions in the cytochrome P-450 lanosterol 14-demethylase (CYP51a1) from azole-resistant Candida albicans clinical isolates contribute to resistance to azole antifungal agents. Antimicrob. Agents Chemother.42, 241–253.
57
SanglardD.KuchlerK.IscherF.PaganiJ. L.MonodM.BilleJ. (1995). Mechanisms of resistance to azole antifungal agents in Candida albicans isolates from AIDS patients involve specific multidrug transporters. Antimicrob. Agents Chemother.39, 2378–2386. 10.1128/AAC.39.11.2378
58
SayeedM. A.FarooqiJ.JabeenK.AwanS.MahmoodS. F. (2019). Clinical spectrum and factors impacting outcome of Candida auris: a single center study from Pakistan. BMC Infect. Dis.19:384. 10.1186/s12879-019-3999-y
59
SchelenzS.HagenF.RhodesJ. L.AbdolrasouliA.ChowdharyA.HallA.et al. (2016). First hospital outbreak of the globally emerging Candida auris in a European hospital. Antimicrob. Resist. Infect. Control5:35. 10.1186/s13756-016-0132-5
60
Schuetzer-MuehlbauerM.WillingerB.EgnerR.EckerG.KuchlerK. (2003). Reversal of antifungal resistance mediated by ABC efflux pumps from Candida albicans functionally expressed in yeast. Int. J. Antimicrob. Agents22, 291–300. 10.1016/S0924-8579(03)00213-9
61
SextonD. J.KordalewskaM.BentzM. L.WelshR. M.PerlinD. S.LitvintsevaA. P. (2018). Direct detection of emergent fungal pathogen Candida auris in clinical skin swabs by SYBR green-based quantitative PCR assay. J. Clin. Microbiol.56:e01337–18. 10.1128/JCM.01337-18
62
SharmaC.KumarN.PandeyR.MeisJ. F.ChowdharyA. (2016). Whole genome sequencing of emerging multidrug resistant Candida auris isolates in India demonstrates low genetic variation. New Microbes New Infect.13, 77–82. 10.1016/j.nmni.2016.07.003
63
SherryL.RamageG.KeanR.BormanA.JohnsonE. M.RichardsonM. D.et al. (2017). Biofilm-forming capability of highly virulent, multidrug-resistant Candida auris. Emerg. Infect. Dis.23, 328–331. 10.3201/eid2302.161320
64
VandeputteP.FerrariS.CosteA. T. (2012). Antifungal resistance and new strategies to control fungal infections. Int. J. Microbiol.2012:713687. 10.1155/2012/713687
65
WaldorfA. R.PolakA. (1983). Mechanisms of action of 5-fluorocytosine. Antimicrob. Agents Chemother.23, 79–85. 10.1128/AAC.23.1.79
66
WangX.BingJ.ZhengQ.ZhangF.LiuJ.YueH.et al. (2018). The first isolate of Candida auris in China: clinical and biological aspects. Emerging Microbes & Infections7, 1–9. 10.1038/s41426-018-0095-0
67
WelshR. M.BentzM. L.ShamsA.HoustonH.LyonsA.RoseL. J.et al. (2017). Survival, persistence, and isolation of the emerging multidrug-resistant pathogenic yeast Candida auris on a plastic health care surface. J. Clin. Microbiol.55, 2996–3005. 10.1128/JCM.00921-17
Summary
Keywords
Candida auris, antifungal resistance, biofilm, emerging pathogen, MIC, outbreaks
Citation
Chaabane F, Graf A, Jequier L and Coste AT (2019) Review on Antifungal Resistance Mechanisms in the Emerging Pathogen Candida auris. Front. Microbiol. 10:2788. doi: 10.3389/fmicb.2019.02788
Received
19 September 2019
Accepted
15 November 2019
Published
29 November 2019
Volume
10 - 2019
Edited by
Ana Traven, Monash University, Australia
Reviewed by
Jeniel E. Nett, University of Wisconsin-Madison, United States; Delma S. Childers, University of Aberdeen, United Kingdom
Updates
Copyright
© 2019 Chaabane, Graf, Jequier and Coste.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Alix T. Coste alix.coste@chuv.ch
This article was submitted to Fungi and Their Interactions, a section of the journal Frontiers in Microbiology
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.