Abstract
Lactic acid bacteria (LAB) belonging to the genus classically known as Lactobacillus, recently split into 25 different genera, include many relevant species for the food industry. The well-known properties of lactobacilli as probiotics make them an attractive model also for vaccines and therapeutic proteins delivery in humans. However, scarce tools are available to accomplish genetic modification of these organisms, and most are only suitable for laboratory strains. Here, we test bacterial conjugation as a new tool to introduce genetic modifications into many biotechnologically relevant laboratory and wild type lactobacilli. Using mobilizable shuttle plasmids from a donor Escherichia coli carrying either RP4 or R388 conjugative systems, we were able to get transconjugants to all tested Lactocaseibacillus casei strains, including many natural isolates, and to several other genera, including Lentilactobacillus parabuchneri, for which no transformation protocol has been reported. Transconjugants were confirmed by the presence of the oriT and 16S rRNA gene sequencing. Serendipitously, we also found transconjugants into researcher-contaminant Staphylococcus epidermidis. Conjugative DNA transfer from E. coli to S. aureus was previously described, but at very low frequencies. We have purified this recipient strain and used it in standard conjugation assays, confirming that both R388 and RP4 conjugative systems mediate mobilization of plasmids into S. epidermidis. This protocol could be assayed to introduce DNA into other Gram-positive microorganisms which are resistant to transformation.
Introduction
Lactic acid bacteria (LAB) are a heterogenic group of Gram-positive bacteria with the capacity of producing lactic acid as the main product of their sugar metabolism. Consequently, LAB are an essential microbial group in the food industry due to their use as starters in the elaboration of a great variety of fermented food and drinks, being responsible for their organoleptic properties and acting as natural preservatives (Smit and Smit, 2005; Jany and Barbier, 2008; Börner et al., 2019). Due to their extensive use during the last centuries in the food industry, some species are Generally Regarded As Safe (GRAS) by the Food and Drugs Administration (FDA) and have the status of Qualified Presumption of Safety (QPS) by the European Food Safety Authority (EFSA). The genus Lactobacillus, in addition, contains some strains that are well-known probiotics. Up to now, the genus Lactobacillus was exceptionally large and diverse, as it comprised 261 species very different at the phenotypic, ecological and genotypic level. Recently, the genus has been revisited and a new classification into 25 genera has been established (Zheng et al., 2020), which helps to reflect the great biodiversity among the species that previously were grouped as Lactobacillus. In this work, the term “lactobacilli” will remain used to designate all organisms previously classified as Lactobacillus up to 2020.
The use of lactobacilli has earned interest in human and animal biomedical applications (Wells and Mercenier, 2008; Cano-Garrido et al., 2015; Wang et al., 2016). They are crucial members of the microbiota of human mucosal surfaces, where they are involved in homeostasis processes, providing protection against pathogenic bacteria and stimulating the immune system (Isolauri and Ouwehand, 2004; Bernardeau et al., 2008). Lactobacilli have been proposed as ideal live vectors for the in situ production of therapeutic agents in the oral, nasal, genital and intestinal mucosae (Cano-Garrido et al., 2015; Wang et al., 2016; Rio et al., 2019), due to their tolerance to temperature, low pH, bile salts, or high alcohol concentrations (Bosma et al., 2017). So far, lactobacilli have been used as adjuvants or prophylactic agents against many different diseases (Reid, 2017; Mays and Nair, 2018) as well as in a range of animal husbandries (Syngai et al., 2016). Furthermore, their use in therapeutics for prevention and diagnosis (Mays and Nair, 2018) is gaining attention. However, the extended use of lactobacilli in industrial and biomedical applications is limited, since genetic tools are still underdeveloped, especially for wild-type strains (Bosma et al., 2017).
LAB were a pioneer group studied for development of genetic tools (de Vos, 2011), but these efforts were mainly focused on obtaining food-grade microorganisms rather than optimizing mutagenesis procedures (Derkx et al., 2014; Bachmann et al., 2015; Johansen, 2018; Vida and Berlec, 2020). There are several targeted genome editing methods currently available for LAB (Martin et al., 2000; Bosma et al., 2017; Hatti-Kaul et al., 2018). First studies focused on Lactococcus lactis and Lactiplantibacillus plantarum, due to their importance as starter cultures and probiotics, respectively, but several other LAB species have been found to be susceptible to genetic modification albeit with significantly lower efficiencies (de Vos, 2011; Bosma et al., 2017). The first step to accomplish targeted genetic modification is the introduction of DNA, which can be challenging in Gram-positive bacteria due to the thick peptidoglycan layer in their cell wall. The most widely used method is electroporation. Although a wide range of LAB species have been successfully transformed using generalized electroporation protocols, efficiencies varied strongly among strains and protocols need to be optimized (Landete et al., 2014; Bosma et al., 2017). In particular, transformation of lactobacilli wild-type strains has proven difficult or even not feasible. Thus the importance of exploring new approaches for efficient DNA introduction in these important LAB.
Bacterial conjugation is an efficient mechanism of horizontal gene transfer (HGT) of DNA from a donor bacterium to a recipient one which requires physical contact between them. Bacterial conjugation confers a high genomic plasticity to the prokaryotic world (de la Cruz and Davies, 2000), being the most important means of spreading resistance and virulence factors among bacteria. The conjugative machinery is composed by a Type IV secretion system (T4SS), which constitutes the physical channel for secretion of the DNA, and a number of proteins which recognize and process the DNA to be transferred (Cabezón et al., 2015). Among them, a key enzyme is the relaxase, which attaches covalently to the DNA and pilots it into the recipient cell (Guzmán-Herrador and Llosa, 2019). The DNA to be transferred must have a sequence of 100–400 bp known as origin of transfer (oriT), which is recognized by the relaxase, where it binds and cleaves the strand to be transferred.
Conjugation has been described in both Gram-negative and -positive bacteria, and even between both bacterial groups. Conjugative transfer using the RP4 transfer system from E. coli to several Gram-positive bacteria was described long ago (Trieu-Cuot et al., 1987). Dominguez and Sullivan (2013) describe a robust conjugation protocol that can be used in the transfer of genetic material from E. coli to several Bifidobacterium species. Although conjugative plasmids and transposons are very common in LAB, the details of conjugative mechanisms are still under research (Kullen and Klaenhammer, 2000; Bron et al., 2019). It has also been described that conjugative transfer happens in vivo in our microbiota, including LAB species as recipients (Aviv et al., 2016). Conjugative DNA transfer from lactobacilli has been described in a few instances to other LAB, such as Enterococcus and Lactococcus (Gevers et al., 2003). However, up to date, transfer of DNA into lactobacilli by conjugation has not been reported.
The development of genetic modification tools for Gram-positive bacteria has also focused on species with clinical relevance. An example of this is illustrated by the genus Staphylococcus, which includes many relevant strains for human health, and is also reluctant to genetic manipulation. Staphylococci are one of the main causative agents of severe nosocomial infections which require prolonged hospitalizations (Becker et al., 2014). The majority of genetic tools have been developed for Staphylococcus aureus; in other staphylococci, genetic modification is often halted by the absence of efficient transformable protocols. This is the case of coagulase-negative staphylococci (Becker et al., 2014) which include species with increasing interest in human health, such as the emerging human pathogen Staphylococcus epidermidis (Otto, 2009). Up to now, staphylococcal species are transformed via electroporation or, less frequently, by protoplast transformation (Götz and Schumacher, 1987; Augustin and Götz, 1990; Löfblom et al., 2007). However, the restriction-modification systems present in S. aureus truncate the uptake of foreign DNA (Waldron and Lindsay, 2006; Xu et al., 2011; Monk and Foster, 2012; Monk et al., 2012). The use of E. coli strains lacking dcm for production of unmethylated DNA allowed electroporation of particular strains (Monk et al., 2012; Costa et al., 2017), but it requires large amounts of DNA and is limited to specific isolates. Transformation is especially inefficient for S. epidermidis strains (Costa et al., 2017). Bacterial conjugation from E. coli to S. aureus was initially reported, albeit at low frequency (Trieu-Cuot et al., 1987), and no follow-up works are available.
The limitations of current electroporation protocols for the introduction of DNA, especially in wild-type strains, prompted us to assay conjugation as an alternative to transfer DNA into lactobacilli. Furthermore, conjugation is considered a natural mechanisms and therefore is a more accepted approach than electro-transformation (Pedersen et al., 2005). To this end, we have optimized a conjugation protocol from E. coli to lactobacilli using the promiscuous conjugative plasmids R388 and RP4. Using this protocol, we obtained transconjugants in a number of genera and species, including many wild-type strains. Serendipitously, we found that this conjugation protocol also mediates conjugal transfer from E. coli to S. epidermidis, a researcher-contaminant bacterium which normally colonizes the human skin. This conjugation protocol could be a useful approach to genetically modify other Gram-positive microorganisms which are resistant to electroporation.
Materials and Methods
Bacterial Strains and Culture Conditions
The bacterial strains used in this study are listed in Table 1. Escherichia coli strains were grown at 37°C in LB media and when necessary supplemented with 100 μg/ml ampicillin (Ap), or 50 μg/ml kanamycin (Km). Lactobacilli and S. epidermidis were grown at 37°C without aeration in MRS medium (Oxoid, Basingstoke, Hampshire, England) or on solid MRS plates supplemented with 2% agar, supplemented with 5 μg/ml erythromycin (Em) when indicated.
TABLE 1
| Bacteria | Relevant propertiesa | Reference or source |
| Escherichia coli DH5α | NxR; F- endA1 hsdR17 supE44 thi-1 recA1 gyrA96 relA1Δ(argF-lacZYA) U 169Φ80dlacΔM15 | Grant et al., 1990 |
| E. coli D1210 | SmR; recA hspR hsdM rpsl lacIq | Sadler et al., 1980 |
| E. coli S17.1 | SmR; (F-)RP4-2-Tc:Mu aph:Tn7 recA | Simon et al., 1983 |
| Furfurilactobacillus rossiae D87 | Isolated from bread dough | Del Rio et al., 2018 |
| Lacticaseibacillus casei 393 | Laboratory strain | Hansen and Lessel, 1971 |
| L. casei 12003 | Isolated from wheat dough—white bread | Alvarez-Sieiro et al., 2016 |
| L. casei 12032 | Isolated from wheat dough—Pasta | Alvarez-Sieiro et al., 2016 |
| L. casei 12042 | Isolated from wheat dough—white bread | Alvarez-Sieiro et al., 2016 |
| L. casei 13b | Dairy-derived—zamorano | Herrero-Fresno et al., 2012 |
| L. casei 41b | Dairy-derived—zamorano | Herrero-Fresno et al., 2012 |
| L. casei 5b | Dairy-derived—zamorano | Herrero-Fresno et al., 2012 |
| L. casei 61b | Dairy-derived—cabrales | Herrero-Fresno et al., 2012 |
| L. casei E2 | Dairy-derived—emmental | Herrero-Fresno et al., 2012 |
| Lacticaseibacillus paracasei 1D-CCC76 | Isolated from cheese | IPLA collection |
| Lactiplantibacillus plantarum IPLA88 | Isolated from bread dough | Laredo et al., 2013 |
| Lactobacillus crispatus HFS47 | Isolated from human feces | IPLA collection |
| Lactobacillus gasseri HFS29 | Isolated from human feces | IPLA collection |
| Latilactobacillus curvatus 1b-VPZ3 | Isolated from cheese | IPLA collection |
| Lentilactobacillus buchneri 1D-VPC30 | Isolated from cheese | IPLA collection |
| Lentilactobacillus parabuchneri 11122 | Dairy-derived—emmental | Diaz et al., 2016 |
| Levilactobacillus brevis 1D-VCC39 | Isolated from cheese | IPLA collection |
| Ligilactobacillus ruminis HFS44 | Isolated from human feces | IPLA collection |
| Limosilactobacillus reuteri IPLA11078 | Isolated from cheese | IPLA collection |
| Limosilactobacillus vaginalis IPLA11050 | Isolated from cheese | Diaz et al., 2015a |
| Loigolactobacillus coryniformis MZ25 | Isolated from cheese | IPLA collection |
| Staphylococcus epidermidis | Human skin—spontaneous isolate | This work |
Bacterial strains used in the present study.
aNxR, nalidixic acid resistance; SmR, streptomycin resistance.
The S. epidermidis strain used in this work was isolated from the researcher’s skin. The hands were placed on MRS-agar plates without antibiotics, which were then incubated at 37°C. The colonies grown were replicated on MRS-agar with and without Em 5 μg/ml. Em-sensitive colonies were selected, their 16S rRNA gene sequence was amplified by PCR, and DNA sequence was determined (STABVIDA) to confirm they were S. epidermidis.
DNA Manipulation
In order to extract genomic DNA from lactobacilli and S. epidermidis, a colony from an MRS-agar plate is punctured and resuspended in 50 μl of TE buffer. 50 μl of chloroform are added and mixed thoroughly until the mixture is homogeneous. The mixture is then centrifuged 10 min at 4°C and three phases appear. The top phase containing the genomic DNA is collected carefully and used directly for PCR analysis.
Plasmid DNA was isolated from E. coli with the GenElute Plasmid Miniprep Kit (Sigma). From lactobacilli and S. epidermidis, the protocol of Anderson and Mckay (1983) was followed with modifications to lyse the cells previous to plasmid DNA purification with the GenElute Plasmid Miniprep Kit, as follows. Lactobacillus and S. epidermidis strains were grown overnight in MRS supplemented with Em (5 μg/ml) for plasmid selection. Two milliliter cultures were centrifuged 10 min at 14,000 rpm. The pellet was resuspended in STE (sucrose 10.3%, Tris HCl 25 mM pH8, EDTA 10 mM) and centrifuged again 10 min at the same speed. The pellet was frozen at −80°C for 15 min. Then, the pellet was resuspended in 200 μl of lysis buffer (sucrose 20%, Tris HCl 10 mM pH8, EDTA 10 mM, NaCl 10 mM) with lysozyme (30 mg/ml), 2 μl of RNAse (25 mg/ml) and 20 μl of proteinase K (20 mg/ml). The sample was homogenized by vortexing, and incubated at 55°C during 30 min. Then, the lysates were applied to GenElute Plasmid Miniprep Kit (Sigma-Aldrich) to purify the plasmid DNA following the manufacturer’s protocol.
DNA and PCR products were visualized by agarose gel electrophoresis stained with SYBR Safe (Invitrogen) and visualized with a Gel Doc2000 UV system, and images were analyzed with Quantity One software (BioRad). HyperLader I (Biolabs) was used as a molecular weight marker. DNA was quantified using a Nano-Drop Spectrophotometer ND-1000. GenElute PCR Clean-Up Kit (Sigma) was used for purification of PCR products, and GenElute Agarose Spin Columns (Sigma) were used for DNA purification from agarose gels.
Plasmids and Plasmid Constructions
Plasmids used in this work and their relevant properties are listed in Table 2. Plasmid constructions were done by standard recombinant DNA techniques. Plasmid pEM110 was digested with the enzymes ClaI and SmaI (Thermo Fisher Scientific). The oriT sequences of plasmids RP4 and R388 were PCR-amplified from plasmids pLA31 and pLA32, respectively, using the oligonucleotides shown in Table 3 and high fidelity DNA polymerase PCRBIO HiFi (PCRBIOSYSTEMS). PCR fragments were digested with the same enzymes and ligated with the vector. Ligations were electroporated into E. coli (see below). The DNA sequence of the inserts was determined (STABVIDA) to verify the correct assembly of the new plasmids.
TABLE 2
| Plasmid | Relevant propertiesa | Reference or source |
| pCOR48 | pEM110- based, shuttle vector E. coli –Lactobacillus, ApR EmR; R388 oriT | This work |
| pCOR49 | pEM110-borned, shuttle vector E. coli –Lactobacillus, ApR EmR; RP4 oriT | This work |
| pEM110 | P8014-2 oriV (L. plantarum), pBR322 oriV (E. coli), EmR | Martín et al., 2004 |
| pLA31 | pSU36:RP4 oriT | Agúndez et al., 2012 |
| pLA32 | pSU36:R388 oriT | |
| pSU711 | KmR; R388 ΔoriT | Demarre et al., 2004 |
Plasmids used in the present study.
aApR, ampicillin resistance; CmR, chloramphenicol resistance; EmR, erythromicyn resistance; KmR, kanamycin resistance.
TABLE 3
| Purpose of PCR | Sequence |
| oriT for cloning | |
| ClaI-oriT R388-1 | 5′-CCGACTATCGATTCTCATTTTCTGCATCATGGTAG-3′ |
| oriT R388 401-SmaI | 5′-AGCTATCCCGGGCCGCCTCGTCCTCCAAAA-3′ |
| ClaI-oriT RP4-1 | 5′-CCGACTATCGATCCGCTTGCCCTCATCTG-3′ |
| oriT RP4-SmaI | 5′-AGCTTTCCCGGGCGCTTTTCCGCTGCATAA-3′ |
| Transconjugant confirmation | |
| oriT R388 F | 5′-CCAAGTCTACTCATTTTCTGCATCATTGT-3′ |
| oriT R388 R | 5′-CCAAGTCTACCTCTCCCGTAGTGTTACT-3′ |
| ClaI-oriT RP4-1 | 5′-CCGACTATCGATCCGCTTGCCCTCATCTG-3′ |
| oriT RP4-SmaI | 5′-AGCTTTCCCGGGCGCTTTTCCGCTGCATAA-3′ |
| 16S rRNA determination | |
| 16S 1492R | 5′-TACGGYTACCTTGTTACGACTT-3′ |
| 16S 27F | 5′-AGAGTTTGATYMTGGCTCAG-3′ |
Primers used for PCR.
Underlined sequences represent the restriction sites.
E. coli Electroporation
For preparation of electrocompetent cells, bacteria were grown to OD600 = 0.5–0.6, and pelleted by centrifugation at 4°C. Two series of washes and centrifugations (6,000 rpm on a Beckman JA-10 rotor) of 1vol milliQ water and a final wash in 1/50 volume 10% glycerol at 4°C were applied. Cells were resuspended in 1/500 vol 10% glycerol and aliquoted in 50 μl samples. Aliquots were frozen on dry ice and kept at −70°C until usage. Aliquots were mixed with < 10 ng of DNA in a 0.2 cm Gene Pulser® cuvette (BioRad) and subjected to an electric pulse (2.5 kV, 25 μF and 200 Ω) in a MicroPulserTM (BioRad). Electroporated cells were added to 1 ml LB and incubated with shaking at 37°C to allow antibiotic expression. After incubation cells were plated on antibiotic containing media.
Bacterial Conjugation
Standard conjugation assays in E. coli were performed as described in Grandoso et al. (2000). The conjugation protocol from E. coli to lactobacilli was optimized starting from the one previously described. Once the new protocol was established, conjugation to lactobacilli was performed as follows: First, donor E. coli strains were grown on LB supplemented with antibiotics overnight. Recipient lactobacilli were grown on MRS without antibiotics. The matings were performed on solid media by mixing the same volume of donor and recipient strains (100 μl of the overnight cultures) after washing with BHI media (Oxoid, Basingstoke, Hampshire, England). The bacterial mixture was then washed with BHI again, resuspended in 20 μL of BHI and transferred to a conjugation filter (0.22 μm nitrocellulose, Millipore) on a BHI-agar plate. The mating mixtures were incubated at 37°C for 24 h. Then, the filter was resuspended on BHI and appropriate dilutions were made and plated on selective media for donors, recipients and transconjugants. Donor E. coli were plated on LB agar with antibiotics for strain and plasmids selection. Recipient strains were plated on MRS. Transconjugants were plated on MRS with Em 5 μg/ml. The frequency of conjugation is expressed as the number of transconjugants per donor cell. Conjugation from E. coli into Staphylococcus epidermidis strains was performed as explained above for lactobacilli. All the manipulations of these conjugations were performed on a Faster BH-EN 2004 Class II Microbiological Safety Cabinet and using filter tips.
Analysis of Transconjugants
Transconjugants were analyzed directly from the plate for the presence of oriT. PCR reactions included an extra boiling step at the beginning to break the cells. PCRs were performed using DNA polymerase KapaTaq (KapaBiosystems) and primers indicated in Table 3. PCR products were run on agarose gels to observe the expected amplification bands.
Several transconjugants from each conjugation assay were analyzed to confirm the lactobacilli or S. epidermidis species by PCR-amplification of the 16S rRNA gene, using the universal primer pair 27F and 1492R (Table 3; Lane, 1991), and determination of the DNA sequence from the amplicon, as explained in Diaz et al. (2016).
In order to confirm the presence of the autonomous shuttle plasmid in S. epidermidis transconjugants, plasmid DNA was extracted from both the transconjugants and the strain with no plasmid, and from L. casei with and without plasmid as a control. Plasmid DNA was visualized on agarose gels. Subsequently, this plasmid DNA was electroporated into E. coli, plasmid DNA extracted again from the transformants, and analyzed by restriction digestion to test its integrity.
Results
Bacterial Conjugation From E. coli to Lacticaseibacillus casei
In order to set up a protocol for conjugative DNA transfer into lactobacilli from laboratory E. coli strains, we adapted the protocol routinely used for conjugative DNA transfer among Gram-negative bacteria on solid media (Grandoso et al., 2000). We tested two well characterized conjugative systems; those of plasmids R388 and RP4, which have been previously shown to mediate conjugative DNA transfer into a broad range of recipient cells (see section “Introduction”).
For a DNA molecule to be transferred by conjugation, the only element required in cis is the oriT. We constructed mobilizable shuttle vectors carrying replication and antibiotic resistance genes for selection in E. coli and Lactobacillus, plus the oriT of either R388 or RP4 (pCOR48 and pCOR49; Table 2). The rest of the conjugative machinery was provided in trans, either using E. coli S17.1 strain as a donor, which has the conjugative machinery of RP4 integrated into the bacterial chromosome, or using a non-mobilizable helper plasmid which provides the R388 conjugative system (pSU711; Table 2). These plasmids were tested in conjugation between E. coli strains to verify their functionality (Table 4, top rows). As negative controls, we used donors harboring the mobilizable plasmids but devoid of the rest of the conjugative machinery.
TABLE 4
| Recipient | Donor strain | Conjugative system | Conjugation frequency | Stand. Dev. | n |
| E. coli DH5α | D1210 + pSU711 + pCOR48 | R388+ | 2.5 × 10–3 | ±1.9 × 10–3 | 3 |
| E. coli DH5α | D1210 + pCOR48 | R388− | <2.6 × 10–7 | ±1.5 × 10–7 | 3 |
| E. coli DH5α | S17.1 + pCOR49 | RP4+ | 1.4 × 10–2 | ±1.3 × 10–2 | 3 |
| E. coli DH5α | D1210 + pCOR49 | RP4− | <4.6 × 10–6 | ±4.0 × 10–6 | 3 |
| L. casei 393 | D1210 + pSU711 + pCOR48 | R388+ | 2.1 × 10–4 | ±6.1 × 10–4 | 18 |
| L. casei 393 | D1210 + pCOR48 | R388− | <2.0 × 10–6 | ±2.3 × 10–6 | 18 |
| L. casei 393 | S17.1 + pCOR49 | RP4+ | 1.8 × 10–3 | ±2.0 × 10–3 | 8 |
| L. casei 393 | D1210 + pCOR49 | RP4− | <3.0 × 10–6 | ±3.1 × 10–6 | 8 |
Conjugation frequency of mobilizable shuttle plasmids into E. coli and Lacticaseibacillus casei 393.
Positive results are highlighted in bold.
In order to optimize a new protocol for conjugation from E. coli to lactobacilli, we chose the laboratory strain L. casei 393, which is easy to grow, manipulate and transform (Chassy and Flickinger, 1987). Different conditions were tested, such as the mating time, donor/recipient cell ratio, growth phases in the bacterial cultures, and culture media in the conjugation plate. After several trials, a functional protocol for conjugation between E. coli and L. casei was established. The protocol is detailed in section “Materials and Methods.” In summary, overnight cultures of both donor and recipient bacteria were mixed on BHI medium, where both donor and recipient cells can thrive, while LB and MRS allow growth only of E. coli and L. casei, respectively, allowing counter selection of donors or recipients. The mating mixtures were incubated on solid media for 24 h. Conjugation frequencies obtained are shown in Table 4 (lower rows). Transconjugants were obtained using both R388 and RP4 conjugative systems, with frequencies only 1−log lower than between E. coli strains (2.1 × 10–4 vs. 2.5 × 10–3 transconjugants/donor for R388, and 1.8 × 10–3 vs. 1.4 × 10–2 for RP4).
Several transconjugants were selected for further analysis. Total DNA was extracted and used as a template for two PCR amplifications: (i) the 16S rRNA gene region, which was used for DNA sequence determination and confirmation that they were L. casei; and (ii), the corresponding oriT. An example of this analysis is shown in Figure 1. It can be observed that neither oriT is amplified from gDNA of L. casei, while the expected band for each oriT is present in DNA from the transconjugants. All these results confirm that the transconjugants obtained were bona-fide L. casei colonies which had received the pCOR shuttle plasmid by conjugation.
FIGURE 1

PCR analysis of L. casei 393 transconjugants for the amplification of R388 and RP4 oriT. The oriT amplified by PCR is indicated at the bottom (oligonucleotides indicated in Table 3). The template is indicated on top of the gel, and it was either purified DNA (R388 and RP4 as positive controls, and the recipient L. casei gDNA as negative control) or transconjugant (TC) colonies directly used for the PCR reactions. The expected plasmid in the transconjugants (R388 or RP4) is indicated. –, control reaction with no template DNA. The central lane is a molecular weight marker (GeneRuler 1 kb DNA ladder). The 500 bp band is indicated. Expected fragment size: 330 bp for R388 oriT and 280 bp for RP4 oriT.
The ability to transform lactobacilli by electroporation varies widely depending not only on the genus or species, but also on the strains. Some of the more interesting strains for biotechnological purposes are wild-type isolates, which typically show lower transformation rates than laboratory strains. We tested different strains of L. casei as conjugation recipients, isolated from natural environments. The results (Table 5) showed that we obtained transconjugants into all strains using RP4 conjugative system, whereas R388 conjugative system provided transconjugants in a subset of strains only. Transconjugants were confirmed by PCR amplification of the oriT. We confirmed that conjugation frequencies varied significantly among strains of L. casei, and frequencies were in all cases lower than that of the laboratory strain: in the case of R388, frequencies ranged around 10–7–10–5 transconjugants/donor (compared to 10–4 for L. casei 393), and in the case of RP4, we obtained between 10–7 and 10–4 transconjugants per donor (compared to 10–3 for the laboratory strain).
TABLE 5
| Conjugation frequencies | ||||
| R388 | RP4 | |||
| Recipient | ||||
| strain | + | − | + | − |
| L. casei 393 | 2.1 × 10–4 | <2.0 × 10–6 | 1.8 × 10–3 | <3.0 × 10–6 |
| L. casei 5 b | 3.5 × 10–6 | <3.6 × 10–6 | 1.9 × 10–4 | <2.5 × 10–6 |
| L. casei 13 b | 2.8 × 10–6 | <8.4 × 10–6 | 3.9 × 10–5 | <6.0 × 10–6 |
| L. casei E2 | 1.5 × 10–7 | <6.6 × 10–7 | 1.6 × 10–4 | <8.1 × 10–7 |
| L. casei 41 b | < 1.3 × 10–6 | <2.9 × 10–6 | 1.1 × 10–5 | <3.0 × 10–6 |
| L. casei 61 b | 5.0 × 10–5 | <5.8 × 10–6 | 3.8 × 10–5 | <1.2 × 10–5 |
| L. casei 12003 | 2.4 × 10–5 | <1.0 × 10–7 | 1.1 × 10–4 | <1.0 × 10–7 |
| L. casei 12032 | < 1.6 × 10–6 | <1.1 × 10–7 | 7.8 × 10–6 | <1.0 × 10–7 |
| L. casei 12042 | < 3.9 × 10–8 | <2.1 × 10–8 | 5.6 × 10–7 | <3.6 × 10–7 |
Conjugation frequencies from E. coli to different L. casei strains using R388 and RP4 systemsa.
aDonor strains as shown inTable 4. Data shown are the mean of 2 independent experiments. Positive results are highlighted in bold.
Conjugation From E. coli to Other Lactobacilli
The next step was to test conjugation to other wild-type lactobacilli, some of which are reluctant to genetic transformation by electroporation. Conjugation was performed using the same donor strains harboring R388 and RP4 conjugative systems as shown in Table 4, and using as positive control for conjugation L. casei 393 as a recipient. The conjugation frequencies obtained are shown in Table 6. Transconjugants were obtained for Lactiplantibacillus plantarum, Lentilactobacillus buchneri, Lentilactobacillus parabuchneri, Levilactobacillus brevis, and Limosilactobacillus vaginalis when using the RP4 conjugative system. No transconjugants were obtained when using the R388 conjugative system. The frequencies obtained were in all cases significantly lower than into the laboratory strain L. casei 393, ranging around 10–5–10–6 transconjugants per donor (vs. 10–3 for the laboratory strain).
TABLE 6
| Conjugative system | ||||
| R388 | RP4 | |||
| Recipient | ||||
| lactobacilli | + | − | + | − |
| L. casei 393 | 2.1 × 10–4 | < 2.0 × 10–6 | 1.8 × 10–3 | < 3.0 × 10–6 |
| L. curvatus | <2.2 × 10–9 | <5.7 × 10–9 | <1.0 × 10–7 | <1.0 × 10–8 |
| L. buchneri | <4.8 × 10–8 | <8.7 × 10–8 | 6.6 × 10–7 | <1.1 × 10–7 |
| L. brevis | <6.8 × 10–7 | <2.1 × 10–7 | 6.0 × 10–6 | <1.3 × 10–6 |
| L. paracasei | <3.2 × 10–7 | <1.4 × 10–7 | <3.1 × 10–6 | <1.4 × 10–6 |
| L. coryniformis | <8.2 × 10–7 | <1.7 × 10–7 | <1.8 × 10–6 | <1.6 × 10–7 |
| L. parabuchneri | <3.9 × 10–7 | <9.7 × 10–8 | 2.0 × 10–6 | < 2.0 × 10–6 |
| L. reuteri | <1.6 × 10–7 | <9.1 × 10–9 | <1.4 × 10–7 | <6.8 × 10–9 |
| L. vaginalis | <5.7 × 10–7 | <9.5 × 10–7 | 1.1 × 10–6 | <1.47 × 10–7 |
| L. rossiae | <1.4 × 10–6 | <2.0 × 10–6 | <5.0 × 10–6 | <1.6 × 10–6 |
| L. plantarum | <2.6 × 10–6 | <2.3 × 10–8 | 1.1 × 10–5 | <9.0 × 10–9 |
| L. crispatus | <1.0 × 10–7 | <1.8 × 10–7 | <2.0 × 10–7 | <2.7 × 10–7 |
Conjugation frequencies from E. coli to different lactobacilli using R388 and RP4 systemsa.
aDonor strains as shown inTable 4. Data shown are the mean of 2 independent experiments. Positive results are highlighted in bold.
Transconjugants were confirmed by the presence of the corresponding oriT (Figure 2). Their 16S rRNA gene sequence was amplified with primers shown in Table 3 and the DNA sequence determined, confirming in all cases the expected genera.
FIGURE 2

Amplification of RP4 oriT from transconjugants. Primers used are shown in Table 3. The template DNAs are: Lane 1: no DNA (C–, negative control). Lanes 2–4: gDNAs from recipient L. parabuchneri, L. buchneri and L. vaginalis. M, molecular marker (GeneRuler 1 kb DNA ladder). The size of the bands is 1,000, 750, 500, and 250 bp from top to bottom. The expected size of the amplicon is 280 bp. To the right, Lane 5: pCOR49 plasmid DNA (C+, positive control). Lanes 6 and 7: DNA extracts from L. parabuchneri transconjugants. Lanes 8 and 9: DNA extracts from L. buchneri transconjugants. Lane 10: gDNA extracts from L. vaginalis transconjugant.
Conjugation From E. coli to S. epidermidis
Serendipitously, we found a high number of putative transconjugants in one of the matings described above using the R388 conjugative system, which did not match the lactobacilli phenotype, although they did show amplification of the R388 oriT. Upon sequencing of the 16S rRNA gene, we found out these colonies corresponded to Staphylococcus epidermidis, a common isolate in human epidermis, and thus probably originated from a contamination from the researcher skin. Since this fact suggested that conjugation from E. coli occurred to other Gram-positives, and S. epidermidis itself is a recalcitrant organism of high biomedical interest, we decided to confirm and quantify this phenomenon. To confirm and quantify this finding, as well as to rule out that the observed phenotype was restricted to a particular isolate, S. epidermidis was isolated placing on MRS plates the hands of two other researchers, from the same and different laboratories. Colonies resembling staphylococci were obtained, cultured on MRS, checked for their sensitivity to erythromycin by replica-plating, and confirmed as S. epidermidis by 16S rRNA gene sequencing. A PCR for the R388 and RP4 oriT was performed on total DNA to verify that there was no amplification from the strains (not shown). These new isolates of S. epidermidis (isolates 1 and 2) were used as a recipient strains in conjugation assays from E. coli.
Conjugation frequencies obtained are summarized in Table 7. We obtained S. epidermidis transconjugants for both recipient strains, using both R388 and RP4 conjugative systems, confirming that plasmids can be mobilized by conjugation from E. coli to S. epidermidis. It is interesting to note that in this case, the R388 system was similar or even more efficient in rendering transconjugants than RP4. Transconjugants were confirmed by PCR for amplification of the corresponding oriT, and their 16S rRNA gene sequence was amplified by PCR and the DNA sequence determined, to verify that they were S. epidermidis.
TABLE 7
| Conjugation frequencies | ||||
| R388 | RP4 | |||
| Recipient | ||||
| bacteria | + | − | + | − |
| S. epidermidis isolate 1 | 2.5 × 10–6 | <1.4 × 10–6 | 8.1 × 10–8 | <1.1 × 10–8 |
| S. epidermidis isolate 2 | 3.5 × 10–7 | <7.8 × 10–7 | 1.7 × 10–7 | <2.1 × 10–7 |
Conjugation frequencies from E. coli to S. epidermidis using conjugative systems R388 and RP4a.
aDonor strains as shown inTable 4. Data shown are the mean of 3–4 independent experiments. Positive results are highlighted in bold.
The mobilizable shuttle plasmids used for our mating assays (pCOR48 and pCOR49; Table 2) carry origins of replication for E. coli and Lactobacillus, but it is not known if the plasmid can replicate in staphylococci. In order to determine if the S. epidermidis transconjugants harbored the shuttle plasmid as an episome, or they were the result of integration of the plasmid into the S. epidermidis chromosome, plasmid DNA was extracted from several transconjugants. In parallel, plasmid DNA was extracted from the strain without plasmids as a negative control, and from L. casei 393 transconjugants obtained in previous conjugation assays, as positive controls. The DNA samples were run on agarose gels, where the plasmid DNA was visible in the S. epidermidis transconjugants (not shown). For further confirmation, these plasmid DNA samples were used to electroporate E. coli. Ampicillin-resistant colonies were obtained and their plasmid DNA was extracted and analyzed by restriction digestion in parallel with the original plasmid DNA, to confirm the presence of the pCOR shuttle plasmid (Figure 3). It can be observed that the plasmid recovered from the E. coli cells transformed with plasmid DNA extracted from the L. casei or S. epidermidis transconjugants maintains the same restriction pattern as the original shuttle plasmid present in the donor E. coli strain. Thus, the shuttle plasmid is able to replicate in S. epidermidis.
FIGURE 3
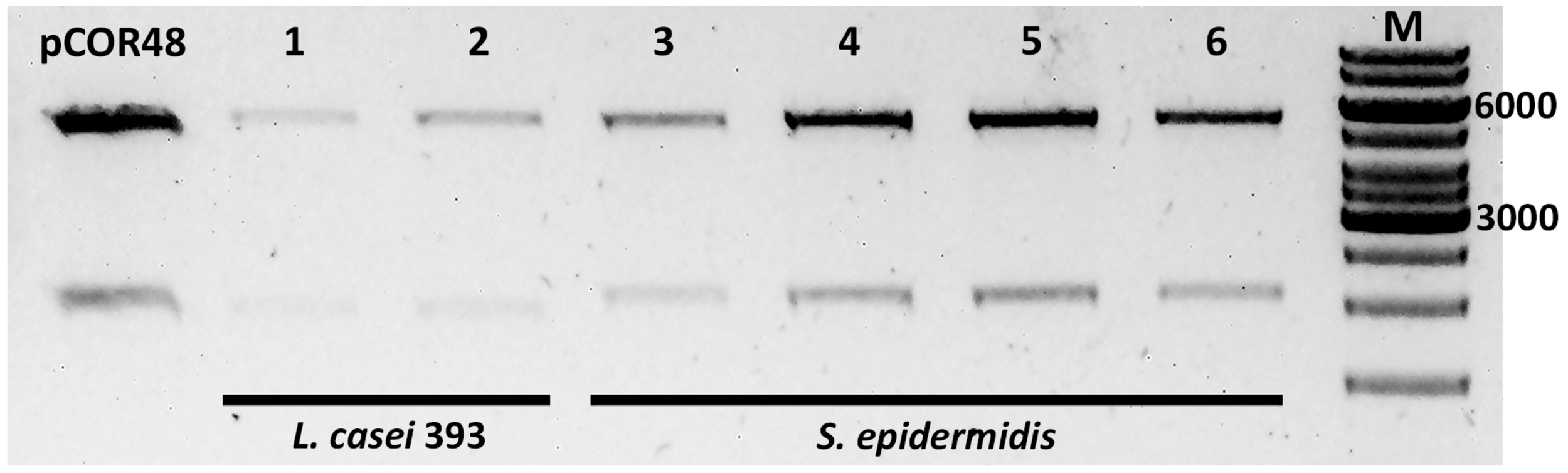
Analysis of plasmids extracted from E. coli previously transformed with plasmid DNA from transconjugants. Plasmid DNA samples were digested with enzyme EcoRI (Fast Digest, Thermo Fisher Scientific), purified, and run on an agarose gel. pCOR48, plasmid DNA (positive control). The transconjugants from which the electroporated DNA was extracted were L. casei 393 (pCOR48) (1–2), and S. epidermidis (pCOR48) from two different colonies (3–4 and 5–6). M, molecular weight marker (GeneRuler 1 kb DNA ladder). The 6,000 and 3,000 bp are indicated. Expected fragment sizes: 5,436 and 2,000 bp.
Discussion
Targeted genetic modification of bacteria with biotechnological and biomedical potential is a prerequisite for most processes of genetic improvement, whether it is to introduce a plasmid contributing to increase production of the desired substance, or to introduce scarless genetic modifications in strains to be applied to human consumption or for medical uses. These processes all have a first requisite consisting on the introduction of DNA into the target cell. A number of protocols exist using bacterial transformation, conjugation, phage transduction, or even protoplast fusions, which are available for most laboratory strains. However, the need for novel substances to use as antimicrobials, food additives, probiotics, or therapeutic substances, has propelled the search for wild-type strains providing the desired properties, which require subsequent optimization steps. Introducing foreign DNA into these microorganisms often proves challenging and even impossible. Among LAB, electroporation is the most widely used method, due to its simplicity, efficiency and wide applicability; however, efficiencies vary strongly among species and even strains, and protocols need to be optimized for each of them (Wang et al., 2020). In particular, transformation of wild-type strains has proven difficult or even not feasible. This is the case for many lactobacilli which, in addition of including some of the most relevant species in the food industry, has an increasingly important biomedical interest, due to both its potential as human live delivery vector and to the existence of emerging human pathogens. Thus, there is an open niche for new DNA introduction protocols.
Bacterial conjugation is a naturally efficient and promiscuous mechanism of horizontal gene transfer, which operates among all main bacterial types. Conjugative DNA transfer from E. coli to several LAB has been reported (see section “Introduction”), but to our knowledge, there are no reports of conjugation from E. coli into lactobacilli. In this work, we prove that it is possible to introduce DNA by conjugation into lactobacilli from E. coli, not only to the model laboratory strain L. casei 393, but to a number of other genera, species and natural isolates, typically reluctant to transformation. In particular, we successfully obtained transconjugants in L. plantarum, L. buchneri, L. parabuchneri, L. brevis, and L. vaginalis (Table 6). Since no transconjugants were obtained in the negative controls lacking the conjugative machinery, we conclude that DNA transfer is happening through conjugation, and not through other mechanisms such as natural transformation, nanotubes, or extracellular vesicles. Dedicated electroporation protocols have been published for each of these species, reflecting the inherent difficulty of transforming them (Stephenson et al., 2011; Spath et al., 2012; Zhang et al., 2012). In the case of L. parabuchneri, to our knowledge there are no reports of transformation, which makes this result especially significative. L. parabuchneri is a member of cheese flora, contributing to its organoleptic properties and ripening process (Fröhlich-Wyder et al., 2013). Moreover, some species have been characterized as potential probiotics (Agostini et al., 2018). On the other hand, some strains of L. parabuchneri are mainly responsible for the undesirable accumulation of the biogenic amine histamine in cheese (Diaz et al., 2015b). Thus, the ability to manipulate genetically this species has high scientific interest, as well as both biotechnological and biomedical potential.
The efficiency of conjugation into the model laboratory strain L. casei 393 was around 10–3 or 10–4 transconjugants per donor, depending on the conjugative system used (RP4 or R388), which is higher than in early reports of conjugative transfer between distantly related bacteria (Trieu-Cuot et al., 1987). Comparable rates were obtained in conjugation experiments from E. coli to Bifidobacterium (Dominguez and Sullivan, 2013) where differences between strains were apparent. The efficiency of the different lactobacilli species as recipients also varied widely, and was always lower than that of the laboratory strain (Table 6). Some species were not transformed. There is no taxonomic explanation for this difference: according to the recent reclassification of the genus Lactobacillus (Zheng et al., 2020), L. casei shares the genus with its closest relative L. paracasei, for which we obtained no transconjugants. In fact, a survey of different L. casei natural isolates (Table 5) showed also ample variation within the species. The difference in conjugation frequencies could have multiple causes, such as the existence of different restriction-modification systems, and very likely, the presence of other plasmids in the wild-type strains; further studies would be necessary to determine the factors interfering with conjugation, which could lead to increased efficiencies and a wider range of potential recipients. Moreover, as for some species the frequency of conjugation obtained in some cases is just at the limit of detection of the mating assays, we think that an optimization of the protocol will probably extend the range of recipient species.
Serendipitously, we found that our conjugation assay also mediated DNA transfer into a researcher-contaminant S. epidermidis, so we isolated this species from different researchers and quantified the DNA transfer, confirming the presence of transconjugants. We have also confirmed the episomal presence of the shuttle plasmid in the transconjugants, meaning that one of the origins of replication present in this plasmid is functional in staphylococci. The mobilizable shuttle plasmids pCOR48 and pCOR49 contain the pBR322 origin of replication, which is functional in E. coli but not in Gram-positive organisms, and the replicon of P8014-2, a plasmid isolated from L. plantarum (Leer et al., 1992). There are a number of broad-host-range plasmids of Gram-positive bacteria which can replicate in both Lactobacillus and Staphylococcus spp. (Jain and Srivastava, 2013). Our results indicate that the Lactobacillus plasmid P8014-2 replicon is also functional in both lactobacilli and staphylococci. In fact, this replicon includes a sequence at position 2051 (5′-TTCTTATCTTGATA-3′) which is identical to the plus origin of replication of plasmid pC194 (Gros et al., 1987), capable of replication in S. aureus and Bacillus subtilis (Horinouchi and Weisblum, 1982).
Our finding that bacterial conjugation can be used to introduce DNA into S. epidermidis is significant. As stated in the Introduction, staphylococci are difficult to transform, and conjugative DNA transfer from E. coli has only been reported for S. aureus (Trieu-Cuot et al., 1987) using an IncP plasmid. S. epidermidis is a component of the human microbiota and also an emerging pathogen (Otto, 2009), leading to an increasing interest in its genetic manipulation. Up to now, few reports have addressed electroporation and transduction, respectively, of specific S. epidermidis strains (Monk et al., 2012; Winstel et al., 2015; Costa et al., 2017). The strategies used to increase the transformation efficiency of S. aureus have little efficiency on S. epidermidis (Monk et al., 2012; Costa et al., 2017). Thus, adding bacterial conjugation to this scarce toolbox will undoubtedly facilitate the generation of genetically modified strains. Conjugation into the laboratory strain L. casei 393 worked efficiently using both RP4 and R388 conjugative systems, although the efficiency was higher with the RP4 system. This result was expected, since the RP4 transfer system has been widely used to transfer DNA into distantly related bacteria and even eukaryotic cells, due to its intrinsic promiscuity (Bates et al., 1998; Luzhetskyy et al., 2006). In contrast, this is, to our knowledge, the first report of conjugative transfer to any Gram-positive bacteria mediated by R388. Moreover, conjugation into S. epidermidis was more efficient using R388 than RP4 conjugative system (Table 7). These results underscore the importance of assaying different conjugative systems, and point to the R388 conjugative system as a suitable candidate to explore other recalcitrant microorganisms as recipients of bacterial conjugation assays.
With this work, we show that a single conjugation protocol allows the introduction of foreign DNA into many different genera, species, and wild-type strains. The result obtained accidentally with S. epidermidis suggests that the range of Gram-positive bacteria which can act as recipients of conjugative DNA transfer from E. coli may be wider than suspected. Using E. coli as a donor laboratory strain implies access to almost unlimited genetic tools to generate the desired DNA to be transferred. Bacterial conjugation is a simple assay, which allows the transfer of DNA molecules of any size, even whole genomes (Isaacs et al., 2011). In addition, bacterial conjugation is considered a natural process, as opposed to electroporation; conjugation has been exploited to introduce natural plasmids into LAB strains, which can be considered non-genetically modified when this technology is used instead of electroporation (Pedersen et al., 2005; Bron et al., 2019). These features are relevant for the genetic manipulation of LAB, for their use in food fermentation as probiotics or as live vector for mucosal delivery of therapeutic proteins (Wells and Mercenier, 2008).
Statements
Data availability statement
The original contributions presented in the study are included in the article/supplementary material, further inquiries can be directed to the corresponding author/s.
Author contributions
MÁ and ML conceived the work. SS, DG-H, RM-C, and MM performed the experiments. SS, DG-H, RM-C, MÁ, and ML analyzed the results. SS, MÁ, and ML wrote the manuscript. All authors contributed to the article and approved the submitted version.
Funding
Work in ML lab was supported by the grant BIO2017-87190-R from the Spanish Ministry of Science and Innovation. Work in MÁ lab was funded by the Spanish State Research Agency (AEI) and the European Regional Development Fund (FEDER) (AGL2016-78708-R, AEI/FEDER, EU). DG-H was a recipient of a predoctoral appointment from the University of Cantabria. RM-C received an Erasmus+ traineeship grant.
Acknowledgments
We are grateful to Matilde Cabezas for technical help, and to Mapi Garcillán for helpful suggestions.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Dedication
This article is dedicated to the memory of María Cruz Martín.
References
1
AgostiniC.EckertC.VincenziA.MachadoB. L.JordonB. C.KipperJ. P.et al (2018). Characterization of technological and probiotic properties of indigenous Lactobacillus spp. from south Brazil.3 Biotech.8:451. 10.1007/s13205-018-1469-7
2
AgúndezL.González-PrietoC.MachónC.LlosaM. (2012). Site-specific integration of foreign dna into minimal bacterial and human target sequences mediated by a conjugative relaxase.PLoS One7:e31047. 10.1371/journal.pone.0031047
3
Alvarez-SieiroP.Montalbán-LópezM.MuD.KuipersO. P. (2016). Bacteriocins of lactic acid bacteria: extending the family.Appl. Microbiol. Biotechnol.100, 2939–2951. 10.1007/s00253-016-7343-9
4
AndersonD. G.MckayL. L. (1983). Simple and rapid method for isolating large plasmid DNA from lactic streptococcit.Appl. Environ. Microbiol.46549–552.
5
AugustinJ.GötzF. (1990). Transformation of Staphylococcus epidermidis and other Staphylococcal species with plasmid DNA by electroporation.FEMS Microbiol. Lett.66203–207. 10.1111/j.1574-6968.1990.tb03997.x
6
AvivG.RahavG.Gal-morO. (2016). Horizontal transfer of the Salmonella enterica serovar infantis resistance and virulence plasmid pESI to the gut microbiota of. mBio.Am. Soc. Microbiol.7:e01395-16. 10.1128/mBio.01395-16
7
BachmannH.PronkJ. T.KleerebezemM.TeusinkB. (2015). Evolutionary engineering to enhance starter culture performance in food fermentations.Curr. Opin. Biotechnol.321–7. 10.1016/j.copbio.2014.09.003
8
BatesS.CashmoreA. M.WilkinsB. M. (1998). IncP plasmids are unusually effective in mediating conjugation of Escherichia coli and Saccharomyces cerevisiae: involvement of the Tra2 mating system.J. Bacteriol.1806538–6543. 10.1128/jb.180.24.6538-6543.1998
9
BeckerK.HeilmannC.PetersG. (2014). Coagulase-negative staphylococci.Clin. Microbiol. Rev.27870–926. 10.1128/CMR.00109-13
10
BernardeauM.PaulJ.Henri-dubernetS.GuéguenM. (2008). Safety assessment of dairy microorganisms: the Lactobacillus genus ⋆.Int. J. Food Microbiol.126278–285. 10.1016/j.ijfoodmicro.2007.08.015
11
BörnerR. A.KandasamyV.AxelsenA. M.NielsenA. T.BosmaE. F. (2019). Genome editing of lactic acid bacteria: opportunities for food, feed, pharma and biotech.FEMS Microbiol. Lett366:fny291. 10.1093/femsle/fny291
12
BosmaE. F.ForsterJ.NielsenA. T. (2017). Lactobacilli and pediococci as versatile cell factories – evaluation of strain properties and genetic tools.Biotechnol. Adv.35419–442. 10.1016/j.biotechadv.2017.04.002
13
BronP. A.MarcelliB.MulderJ.Van Der ElsS.MorawskaL. P.KuipersO. P.et al (2019). Renaissance of traditional DNA transfer strategies for improvement of industrial lactic acid bacteria.Curr. Opin. Biotechnol.5661–68. 10.1016/j.copbio.2018.09.004
14
CabezónE.Ripoll-RozadaJ.PeñaA.de la CruzF.ArechagaI. (2015). Towards an integrated model of bacterial conjugation.FEMS Microbiol. Rev.3981–95. 10.1111/1574-6976.12085
15
Cano-GarridoO. C.Seras-FranzosoJ. S.Garcia-FruitósE. G. (2015). Lactic acid bacteria: reviewing the potential of a promising delivery live vector for biomedical purposes.Microb. Cell Fact.14:137. 10.1186/s12934-015-0313-6
16
ChassyB. M.FlickingerJ. L. (1987). Transformation of Lactobacillus casei by electroporation.FEMS Microbiol. Lett.44173–177.
17
CostaS. K.DoneganN. P.CorvagliaA. R.FrançoisP.CheungA. L. (2017). Bypassing the restriction system to improve transformation of Staphylococcus epidermidis.J. Bacteriol.199:e00271-17. 10.1128/JB.00271-17
18
de la CruzF.DaviesJ. (2000). Horizontal gene transfer and the origin of species: lessons from bacteria.Trends Microbiol.8128–133.
19
de VosW. M. (2011). Systems solutions by lactic acid bacteria: from paradigms to practice.Microb. Cell Fact.10:S2.
20
Del RioB.Alvarez-SieiroP.RedruelloB.MartinM. C.FernandezM.LaderoV.et al (2018). Lactobacillus rossiae strain isolated from sourdough produces putrescine from arginine.Sci. Rep.8:3989. 10.1038/s41598-018-22309-6
21
DemarreG.GuéroutA.-M.Matsumoto-MashimoC.Rowe-MagnusD. A.MarlièreP.MazelD. (2004). A new family of mobilizable suicide plasmids based on broad host range R388 plasmid (IncW) and RP4 plasmid (IncPα) conjugative machineries and their cognate Escherichia coli host strains.Res. Microbiol.156245–255. 10.1016/j.resmic.2004.09.007
22
DerkxP. M. F.JanzenT.SørensenK. I.ChristensenJ. E.Stuer-lauridsenB.JohansenE. (2014). The art of strain improvement of industrial lactic acid bacteria without the use of recombinant DNA technology.Microb. Cell Fact.131–13.
23
DiazM.del RioB.LaderoV.RedruelloB.FernándezM.MartinM. C.et al (2015a). Isolation and typification of histamine-producing Lactobacillus vaginalis strains from cheese.Int. J. Food Microbiol.215117–123. 10.1016/j.ijfoodmicro.2015.08.026
24
DiazM.LaderoV.RedruelloB.Sanchez-llanaE.RioB.FernandezM.et al (2015b). A PCR-DGGE method for the identification of histamine-producing bacteria in cheese.Food Control63216–223. 10.1016/j.foodcont.2015.11.035
25
DiazM.RioB.Sanchez-llanaE.LaderoV.MartinM. C.AlvarezM. A.et al (2016). Histamine-producing Lactobacillus parabuchneri strains isolated from grated cheese can form bio films on stainless steel.Food Microbiol.5985–91. 10.1016/j.fm.2016.05.012
26
DominguezW.SullivanD. J. O. (2013). Developing an efficient and reproducible conjugation-based gene transfer system for bifidobacteria.Microbiology159328–338. 10.1099/mic.0.061408-0
27
Fröhlich-WyderM. T.GuggisbergD.BadertscherR.WechslerD.WittwerA.IrmlerS. (2013). The effect of Lactobacillus buchneri and Lactobacillus parabuchneri on the eye formation of semi-hard cheese.Int. Dairy J.33120–128. 10.1016/j.idairyj.2013.03.004
28
GeversD.HuysG.SwingsJ. (2003). In vitro conjugal transfer of tetracycline resistance from Lactobacillus isolates to other gram-positive bacteria.FEMS Microbiol. Lett.225125–130. 10.1016/S0378-1097(03)00505-6
29
GötzF.SchumacherB. (1987). Improvements of protoplast transformation in Staphylococcus carnosus.FEMS Microbiol. Lett.40285–288. 10.1111/j.1574-6968.1987.tb02040.x
30
GrandosoG.AvilaP.CayónA.HernandoM. A.LlosaM.de la CruzF. (2000). Two active-site tyrosyl residues of protein TrwC act sequentially at the origin of transfer during plasmid R388 conjugation.J. Mol. Biol.2951163–1172.
31
GrantS. G. N.JesseeJ.BloomF. R.HanahanD. (1990). Differential plasmid rescue from transgenic mouse DNAs into Escherichia coli methylation-restriction mutants.Proc. Natl. Acad. Sci. U.S.A.874645–4649.
32
GrosM. F.Te RieleH.EhrlichS. D. (1987). Rolling circle replication of single-stranded DNA plasmid pC194.EMBO J.63863–3869. 10.1002/j.1460-2075.1987.tb02724.x
33
Guzmán-HerradorD. L.LlosaM. (2019). The secret life of conjugative relaxases.Plasmid104:102415. 10.1016/j.plasmid.2019.102415
34
HansenP. A.LesselE. F. (1971). Lactobacillus casei (Orla-Jensen) comb. nov.Int. J. Syst. Bacteriol.21, 69–71.
35
Hatti-KaulR.ChenL.DishishaT.ElH. (2018). Lactic acid bacteria: from starter cultures to producers of chemicals.FEMS Microbiol. Lett.365:fny213. 10.1093/femsle/fny213
36
Herrero-FresnoA.MartínezN.Sánchez-LlanaE.DíazM.FernándezM.MartinM. C.et al (2012). Lactobacillus casei strains isolated from cheese reduce biogenic amine accumulation in an experimental model.Int. J. Food Microbiol.157, 297–304. 10.1016/j.ijfoodmicro.2012.06.002
37
HorinouchiS.WeisblumB. (1982). Nucleotide sequence and functional map of pC194, a plasmid that specifies inducible chloramphenicol resistance.J. Bacteriol.150815–825. 10.1128/jb.150.2.815-825.1982
38
IsaacsF. J.CarrP. A.WangH. H.LajoieM. J.SterlingB.KraalL.et al (2011). Precise manipulation of chromosomes in vivo enables genome- wide codon replacement farren.Science333348–353. 10.1016/j.physbeh.2017.03.040
39
IsolauriE.OuwehandA. C. (2004). Probiotics.Best Pract. Res. Clin. Gastroenterol.18299–313. 10.1053/ybega.2004.443
40
JainA.SrivastavaP. (2013). Broad host range plasmids.FEMS Microbiol. Lett.34887–96. 10.1111/1574-6968.12241
41
JanyJ.BarbierG. (2008). Culture-independent methods for identifying microbial communities in cheese.Food Microbiol.25839–848. 10.1016/j.fm.2008.06.003
42
JohansenE. (2018). Use of natural selection and evolution to develop new starter cultures for fermented foods.Annu. Rev. Food Sci. Technol.9411–428.
43
KullenM. J.KlaenhammerT. R. (2000). Genetic modification of intestinal Lactobacilli and Bifidobacteria.Curr. Issues Mol. Biol.241–50.
44
LandeteJ. M.ArquésJ. L.PeiroténÁLangaS.MedinaM. (2014). An improved method for the electrotransformation of lactic acid bacteria: a comparative survey.J. Microbiol. Methods105130–133. 10.1016/j.mimet.2014.07.022
45
LaneD. J. (1991). “16S/23S rRNA sequencing,” in Nucleic Acid Techniques in Bacterial Systematic, edsStackebrandtE.GoodfellowM. (New York, NY: John Wiley and Sons), 115–175.
46
LaredoV.Alvarez-SieiroP.RedruelloB.Del RioB.LinaresD. M.MartínM. C.et al (2013). Draft genome sequence of Lactobacillus plantarum strain IPLA 88.Genome Announc.1:e00524-13. 10.1038/nbt.2491
47
LeerR. J.van LuijkN.PosnoM.PouwelsP. H. (1992). Structural and functional analysis of two cryptic plasmids from Lactobacillus pentosus MD353 and Lactobacillus plantarum ATCC 8014.Mol. Gen. Genet.234265–274. 10.1007/BF00283847
48
LöfblomJ.KronqvistN.UhlénM.StåhlS.WernérusH. (2007). Optimization of electroporation-mediated transformation: Staphylococcus carnosus as model organism.J. Appl. Microbiol.102736–747. 10.1111/j.1365-2672.2006.03127.x
49
LuzhetskyyA.FedoryshynM.GromykoO.OstashB.RebetsY.BechtholdA.et al (2006). IncP plasmids are most effective in mediating conjugation between Escherichia coli and Streptomycetes.Russ. J. Genet.42476–481. 10.1134/S1022795406050036
50
MartinM. C.AlonsoJ. C.SuaJ. E.AlvarezM. A.LaP. (2000). Generation of food-grade recombinant lactic acid bacterium strains by site-specific recombination.Appl. Environ. Microbiol.662599–2604.
51
MartínM. C.FernándezM.Martín-AlonsoJ. M.ParraF.BogaJ. A.AlvarezM. A. (2004). Nisin-controlled expression of Norwalk virus VP60 protein in Lactobacillus casei à Ferna.FEMS Microbiol. Lett.237385–391. 10.1016/j.femsle.2004.07.002
52
MaysZ. J. S.NairN. U. (2018). Synthetic biology in probiotic lactic acid bacteria: at the frontier of living therapeutics.Curr. Opin. Biotechnol.53224–231. 10.1016/j.copbio.2018.01.028
53
MonkI. R.FosterT. J. (2012). Genetic manipulation of Staphylococci-breaking through the barrier.Front. Cell. Infect. Microbiol.2:49. 10.3389/fcimb.2012.00049
54
MonkI. R.ShahI. M.XuM. (2012). Transforming the untransformable: application of direct transformation to manipulate genetically Staphylococcus aureus and Staphylococcus epidermidis.MBio3e277–e211. 10.1128/mBio.00277-11
55
OttoM. (2009). Staphylococcus epidermidis – the “accidental” pathogen.Nat. Rev. Microbiol.7555–567. 10.1038/nrmicro2182
56
PedersenM. B.IversenS. L.SørensenK. I.JohansenE. (2005). The long and winding road from the research laboratory to industrial applications of lactic acid bacteria.FEMS Microbiol. Rev.29611–624. 10.1016/j.femsre.2005.04.001
57
ReidG. (2017). The development of probiotics for women ’ s health.Can. J. Microbiol.63269–277.
58
RioB.RedruelloB.FernandezM.MartinM. C.LaderoV.AlvarezM. A. (2019). Lactic acid bacteria as a live delivery system for the in situ production of nanobodies in the human gastrointestinal tract.Front. Microbiol.9:3179. 10.3389/fmicb.2018.03179
59
SadlerJ. R.TecklenburgM.BetzJ. L. (1980). Plasmids containing many tandem copies of a synthetic lactose operator.Gene8279–300.
60
SimonR.PrieferU.PühlerA. (1983). A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in gram negative bacteria.Nat. Biotechnol.1784–791.
61
SmitG.SmitB. A. (2005). Flavour formation by lactic acid bacteria and biochemical flavour profiling of cheese products.FEMS Microbiol. Rev.29591–610. 10.1016/j.femsre.2005.04.002
62
SpathK.HeinlS.EggerE.GrabherrR. (2012). Lactobacillus plantarum and Lactobacillus buchneri as expression systems: evaluation of different origins of replication for the design of suitable shuttle vectors.Mol. Biotechnol.5240–48. 10.1007/s12033-011-9471-x
63
StephensonD.MooreR. J.AllisonG. E. (2011). Transformation of, and heterologous protein expression in, Lactobacillus agilis and Lactobacillus vaginalis isolates from the chicken gastrointestinal tract.Appl. Environ. Microbiol.77220–228. 10.1128/AEM.02006-10
64
SyngaiG. G.GopiR.BharaliR.DeyS.LakshmananG. M. A.AhmedG. (2016). Probiotics – the versatile functional food ingredients.J. Food Sci. Technol.53921–933. 10.1007/s13197-015-2011-0
65
Trieu-CuotP.CarlierC.MartinP.CourvalinP. (1987). Plasmid transfer by conjugation from Escherichia coli to gram-positive bacteria.FEMS Microbiol. Lett.48289–294.
66
VidaT.BerlecA. (2020). Safety aspects of genetically modified lactic acid bacteria.Microorganisms8:297.
67
WaldronD. E.LindsayJ. A. (2006). Sau1: a novel lineage-specific type I restriction-modification system that blocks horizontal gene transfer into Staphylococcus aureus and between S. aureus isolates of different lineages.J. Bacteriol.1885578–5585. 10.1128/JB.00418-06
68
WangC.CuiY.QuX. (2020). Optimization of electrotransformation (ETF) conditions in lactic acid bacteria (LAB).J. Microbiol. Methods174:105944. 10.1016/j.mimet.2020.105944
69
WangM.GaoZ.ZhangY.PanL. (2016). Lactic acid bacteria as mucosal delivery vehicles: a realistic therapeutic option.Appl. Microbiol. Biotechnol.1005691–5701. 10.1007/s00253-016-7557-x
70
WellsJ. M.MercenierA. (2008). Mucosal delivery of therapeutic and prophylactic molecules using lactic acid bacteria.Nat. Rev. Microbiol.6349–362. 10.1038/nrmicro1840
71
WinstelV.KühnerP.KrismerB.PeschelA.RohdeH. (2015). Transfer of plasmid DNA to clinical coagulase-negative Staphylococcal pathogens by using a unique bacteriophage.Appl. Environ. Microbiol.812481–2488. 10.1128/AEM.04190-14
72
XuS. Y.CorvagliaA. R.ChanS. H.ZhengY.LinderP. (2011). A type IV modification-dependent restriction enzyme SauUSI from Staphylococcus aureus subsp. aureus USA300.Nucleic Acids Res.395597–5610. 10.1093/nar/gkr098
73
ZhangY.ChengK.GaoQ.GaoN. F. (2012). Elaboration of an electroporation protocol for Lactobacillus brevis.Adv. Mater. Res.393–395729–732. 10.4028/www.scientific.net/AMR.393-395.729
74
ZhengJ.WittouckS.SalvettiE.FranzC. M. A. P.HarrisH. M. B.MattarelliP.et al (2020). A taxonomic note on the genus Lactobacillus: description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae.Int. J. Syst. Evol. Microbiol.702782–2858. 10.1099/ijsem.0.004107
Summary
Keywords
bacterial conjugation, lactic acid bacteria, Lactobacillus, Staphylococcus epidermidis, plasmid RP4, plasmid R388
Citation
Samperio S, Guzmán-Herrador DL, May-Cuz R, Martín MC, Álvarez MA and Llosa M (2021) Conjugative DNA Transfer From E. coli to Transformation-Resistant Lactobacilli. Front. Microbiol. 12:606629. doi: 10.3389/fmicb.2021.606629
Received
15 September 2020
Accepted
22 January 2021
Published
11 February 2021
Volume
12 - 2021
Edited by
Eva M. Top, University of Idaho, United States
Reviewed by
Elisabeth Grohmann, Beuth Hochschule für Technik Berlin, Germany; Sarah Lebeer, University of Antwerp, Belgium
Updates
Copyright
© 2021 Samperio, Guzmán-Herrador, May-Cuz, Martín, Álvarez and Llosa.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Matxalen Llosa, llosam@unican.esMiguel A. Álvarez, maag@ipla.csic.es
†Deceased in April 2020
This article was submitted to Evolutionary and Genomic Microbiology, a section of the journal Frontiers in Microbiology
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.