- 1 Department of Cell and Molecular Biology, Medical Faculty Mannheim, Heidelberg University, Mannheim, Germany
- 2 Nikon Imaging Center, Heidelberg University, Heidelberg, Germany
- 3 Max Planck Institute for Medical Research, Heidelberg, Germany
The habenular neural circuit is attracting increasing attention from researchers in fields as diverse as neuroscience, medicine, behavior, development, and evolution. Recent studies have revealed that this part of the limbic system in the dorsal diencephalon is involved in reward, addiction, and other behaviors and its impairment is associated with various neurological conditions and diseases. Since the initial description of the dorsal diencephalic conduction system (DDC) with the habenulae in its center at the end of the nineteenth century, increasingly sophisticated techniques have resolved much of its anatomy and have shown that these pathways relay information from different parts of the forebrain to the tegmentum, midbrain, and hindbrain. The first part of this review gives a brief historical overview on how the improving experimental approaches have allowed the stepwise uncovering much of the architecture of the habenula circuit as we know it today. Our brain distributes tasks differentially between left and right and it has become a paradigm that this functional lateralization is a universal feature of vertebrates. Moreover, task dependent differential brain activities have been linked to anatomical differences across the left–right axis in humans. A good way to further explore this fundamental issue will be to study the functional consequences of subtle changes in neural network formation, which requires that we fully understand DDC system development. As the habenular circuit is evolutionarily highly conserved, researchers have the option to perform such difficult experiments in more experimentally amenable vertebrate systems. Indeed, research in the last decade has shown that the zebrafish is well suited for the study of DDC system development and the phenomenon of functional lateralization. We will critically discuss the advantages of the zebrafish model, available techniques, and others that are needed to fully understand habenular circuit development.
Anatomy and Connectivity of the Dorsal Diencephalic Conduction System
The function of the brain has fascinated people for thousands of years and indeed the brain was alluded to as early as 1700 BC in an Egyptian papyrus describing cases of brain injuries and recommended treatments (Breasted, 1980). After Herophilus, the “father of anatomy,” predicted the brain as being the source of intelligence around 300 BC, analyses focused on descriptions of brain compartmentalization and anatomy (von Staden, 2007). This approach reached a peak during the renaissance when Leonardo da Vinci designed models of brain structures and introduced the general concept of asymmetry (Bell and Sons, 1897). 150 years ago, Paul Broca demonstrated that the brain is used asymmetrically across the left–right axis through analyses of patients with deficits on one or the other side of the brain (Broca, 1861). Since then it has become apparent that the phenomenon of functional brain lateralization holds true for numerous sensory and cognitive processes and is evolutionary conserved across vertebrate species (Bisazza et al., 1998; Vallortigara et al., 1999; Vallortigara and Rogers, 2005). In particular neural circuits of the limbic system are involved in processing cognitive and sensory information. The limbic system is a complex of brain structures that connects several neurotransmitter pathways involved in behavior, emotion, memory, and olfaction (Hikosaka, 2010). Two major neural pathways of this system can be distinguished: The medial forebrain bundle (MFB) interconnects the anterior olfactory areas with the lateral preoptic, lateral hypothalamic, and ventral tegmental area. In the diencephalic conduction system (DDC) system, the bilaterally formed habenulae in the epithalamus receive information through the stria medullaris from the anterior portion of the medial forebrain and relay it into the ventral midbrain via axons of the fasciculus retroflexus (Sutherland, 1982; Morgane et al., 2005).
Habenular Nuclei
The bilaterally established habenulae are often referred to as “relay station.” They are at the center of the DDC system, just dorsal to the posterior part of the thalamus with which it shares various axonal connections (Guillery, 1959; Cragg, 1961; Morgane et al., 2005). One of the first reports on anatomical differences in the habenulae across the left–right axis came from the study of bone fish in the late nineteenth century (Goronowitsch, 1883; Guglielmotti and Cristino, 2006). Thirty years later, the first functional studies were reported by Brown (1914–1915) who discovered that electrical stimulation of the habenulae evoked respiration patterns in chimpanzee. During this period the chromate staining technique was one of the earliest methods to visualize axonal connections and allowed Cajal to compile a first general description of the DDC neuronal connectivity and to uncover its main components (Cajal, 1911). In mammals, each habenula can grossly be subdivided into a medial and a lateral subnucleus, each of which is innervated by afferent axons of often different origins (Figure 1A). Injections of horseradish peroxidase in combination with autoradiographic methods identified prominent afferent projections to the lateral habenula nucleus from the globus pallidus but also from other areas of the fore- and midbrain running mostly within the stria medullaris (Herkenham and Nauta, 1977). In addition, anterograde fiber degeneration techniques and electron microscopy revealed secondary afferent projections, which have been reviewed elsewhere (Sutherland, 1982; Bianco and Wilson, 2009). These latter techniques also identified projections innervating the medial habenula subnucleus from mainly the hypothalamic area and the precommisural septum but also the lateral preoptic area, the interpeduncular nucleus (IPN), the dorsal raphe nucleus, and the superior cervical ganglion (Mitchell, 1963; Zyo et al., 1963; Powell, 1968; Smaha and Kaelber, 1973; Pierce et al., 1976).
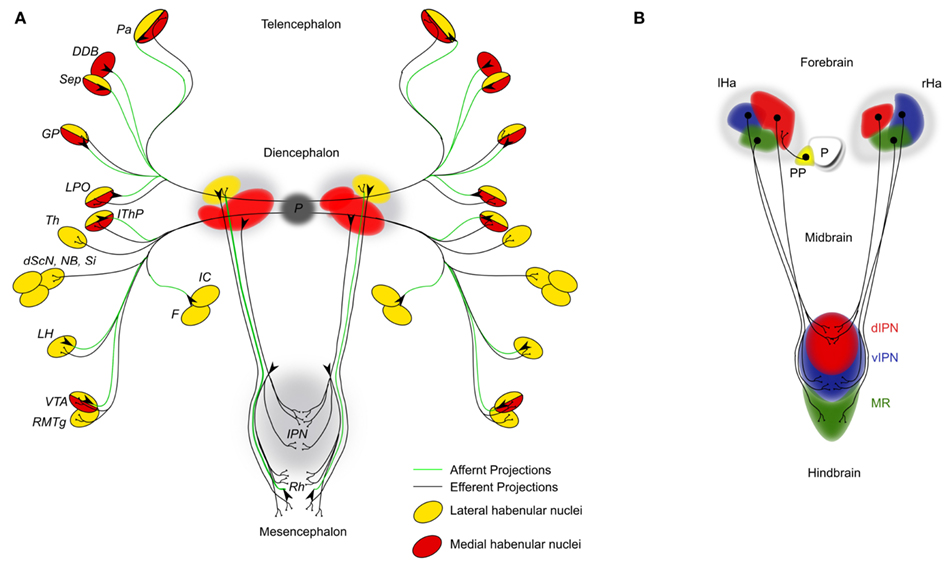
Figure 1. (A) Simplified schematic diagram of the main afferent (green lines) and efferent connections (black lines) of the dorsal diencephalic conduction system in the mammalian brain. Targets and origins of axons related to the lateral habenulae are highlighted in yellow; those of the medial habenulae in red. DDB, nucleus of diagonal band; dScN, dorsal superchiasmatic nucleus; F, fornix; GP, globus pallidus (primate homolog of the teleost entopeduncular nucleus); IC, internal capsule; IThP, inferior thalamic peduncle; LH, lateral hypothalamic area; LPO, lateral preoptic area; NB, nucleus basalis; P, pineal; Pa, pallium; RMTg, rostromedial tegmental nucleus; Sep, septum; Si, substantia innominate; Th, thalamic nuclei; VTA, ventral tegmental area of Tsai. (B) Schematic of the habenular cell composition and efferent axonal projections innervating the interpeduncular nucleus (IPN) in 4 day old zebrafish embryos. The habenular nuclei are subdivided into lateral (red), medial (blue), and ventral (green) domains. D, dorsal; Ha, habenula; IPN, interpeduncular nucleus; l, left; MR, median raphe; P, pineal organ; PP, parapineal organ; r, right; v, ventral.
The fasciculus retroflexus is the most prominent efferent habenula axon bundle connecting each habenula subnuclei with the raphe nuclei and the IPN, which is differentially innervated from three different groups of medial habenular efferent projections (Kuhar et al., 1975; Bianco and Wilson, 2009). The dorsal part of the medial habenular nuclei projects to the lateral IPN, the medial part to the ventral IPN, and the lateral part to the dorsal IPN. Early work of Cajal and later electron microscopy observations from Lenn describe the unusual phenomenon of habenular fibers traversing the entire width of the IPN several times making various synaptic connections with the IPN neurons (Cajal, 1911; Lenn, 1976). Indeed, very recent DNA electroporation studies in teleosts corroborate this remarkable discovery (Bianco et al., 2008). Autoradiographic experiments identified a second group of lateral (and to a minor extend also the medial) habenular efferent projections terminating in the ventral area of Tsai and the dorsal and median raphe (Sutherland, 1982), which makes this part of the habenula being directly connected to the dopaminergic as well as the serotonergic system respectively. Additionally, the lateral habenula projects into the rostromedial tegmental nucleus, which in turn relays information into monoaminergic regions of the midbrain (Jhou et al., 2009a,b).
Interpeduncular Nucleus
The IPN was described for the first time by the early histological observation of Forel (1872) as an unpaired structure astride the midline in the posterior part of the midbrain (Forel, 1872). The first detailed description of the “nucleus interpeduncularis” was carried out by Cajal (1911) using silver chromate labeling methods in different species to show the axonal connections between the IPN and the habenular nuclei and within the IPN. In the middle of the twentieth century electron microscopy techniques allowed the identification of three different types of synapses between the habenulae and the IPN in cat (Milhaud and Pappas, 1966), a number that was corrected to four, 10 years later (Lenn, 1976). Based on anatomical features and neurotransmitter expression, at least seven different IPN subnuclei can be distinguished and these are divided into two main categories: unpaired and paired. The unpaired subnuclei are divided in three subdomains, the rostral, central, and dorsal, while the four paired subdomains are classified in intermediate, lateral, interstitial, and dorso-lateral (Hamill and Lenn, 1984).
Most axons that enter the IPN originate in the habenulae and form the fasciculus retroflexus that show characteristic high levels of acetylcholine, choline acetyltransferase, and acetylcholine esterase (Hattori et al., 1977). Measuring the levels of neuropeptides and enzymes following lesion experiments Staines and colleagues identified IPN afferent connections other than those derived from the habenula, which express additional neurotransmitters such as GABA, monoamines, and neuropeptides (Staines et al., 1980; Bianco and Wilson, 2009). These comprise axons originating in the medial frontal cortex, the nucleus of the diagonal band, the substantia innominata, the preoptic and the hypothalamic nuclei, and the supramammillary nucleus. Similar to the habenulae, the IPN is well embedded into the serotonergic and dopaminergic system as its efferent axons target the raphe nuclei and the ventral tegmental nucleus respectively. Other targets comprise the central medial region of the brain stem, the tegmental nuclei of Gudden, the hypothalamus, the septum, and the diagonal band of Broca (Hamill and Lenn, 1984).
Zebrafish as a Model to Study the Brain Asymmetry
Linking anatomical asymmetries to the lateralization of brain functions is a challenging task for neuroscientists. Starting a decade ago, the DDC system of teleosts has nourished hopes to unravel this mystery. Both the bilaterally formed habenulae and the parapineal complex exhibit distinct anatomical asymmetries that are linked to behavior (Concha and Wilson, 2001; Barth et al., 2005; Snelson et al., 2008; Bianco and Wilson, 2009; Facchin et al., 2009; Gutiérrez-Ibáñez et al., 2011). Most of the initial studies on DDC circuit development focused on unraveling the genetics underlying laterality of the asymmetries found in this structure (Halpern et al., 2003). The zebrafish is a well established model organism suitable to combine the advantages of its transparency and external development with genetics and transgenesis to follow the effect of genetic manipulations in the living animal. The most unbiased approach to uncover genes involved in developmental processes in vertebrates is the forward genetic approach of mutagenesis screens. This technique has been used extensively in zebrafish and has uncovered a vast number of genes involved in embryonic development (Driever et al., 1996). The discovery that mutants carrying mutations in genes of the Nodal signaling pathway exhibit randomized laterality of internal organs and neuroanatomical asymmetries has been a breakthrough in this field of research (Concha et al., 2000). Indeed, several Nodal pathway components such as cyclops, pitx2, and lefty1 are expressed asymmetrically in the dorsal diencephalon in a narrow time window during late somitogenesis before the appearance of neuroanatomical asymmetries. Elegant DNA electroporation approaches in transgenic zebrafish corroborated that the evolutionarily highly conserved Nodal signaling pathway is a major determinant of brain laterality in the dorsal diencephalon (Concha et al., 2003; Gamse et al., 2003). In fact, studies in numerous species ranging from sea urchins to mammals demonstrated that the function of Nodal in determining laterality in the developing embryo is a highly conserved mechanism (Hamada et al., 2002).
How does Nodal signaling confer laterality to the fish brain? Nodal influences the directionality of a group of migrating cells that detach from the medially located pineal organ (Concha et al., 2000, 2003; Gamse et al., 2003). Fate map experiments using caged fluorescein have demonstrated that these parapineal cells originate from both sides of the pineal complex and migrate exclusively to the left side of the brain, where they end up in the immediate vicinity of the left habenula (Concha et al., 2003). The analysis of mutants allowed to uncover that the t-box containing transcription factor tbx2b is essential for this migratory event as well as for the development of the correct number of parapineal cells (Snelson et al., 2008). In mammals, neither tbx2 nor the closely related tbx3 gene is expressed in the dorsal diencephalon. This led Snelson and colleagues hypothesize that the loss of tbx gene expression in the epithalamus of higher vertebrates may be the reason why mammals do not form overt parapineal structures.
Combinations of forward and reverse genetics and transgenesis allowed to reveal another pathway involved in the migration of parapineal cells (Regan et al., 2009). Implantations of FGF8 protein coated microbeads into mutant brains being otherwise devoid of FGF8 mediated signaling identified FGF signaling being crucial in this process. Moreover, transient inactivation of the FGF signaling pathway using drug (SU5402) treatments of embryos revealed that FGF signaling is required at the onset of parapineal cell migration. The FGF pathway appears to be necessary for the parapineal cells to migrate and acts in concert with Nodal signaling to allow their migration into the correct location. As tbx mediated signaling and FGF signaling are known to interact during developmental processes (Rodriguez-Esteban et al., 1999) it is tempting to speculate that also in the dorsal diencephalon tbx2/3 and FGF signaling are epistatically related to facilitate the migration of parapineal cells.
In fish, amphibians, reptiles, birds, and possibly other vertebrates, the bilaterally formed habenulae display side typical characteristics both on anatomical and morphological level (Concha and Wilson, 2001; Figure 1B). The laterality of these asymmetric characteristics and the location of parapineal cells are always concordantly established (Concha et al., 2000). For instance, in wild type embryos with left sided Nodal signaling the parapineal cells will migrate to the left and the left and right habenula will show left and right specific characteristics respectively. This scenario is inverted in manipulated embryos, in which Nodal signaling on the right side of the diencephalon results in a right sided migration of the parapineal cells, leading to the right habenula exhibiting left specific features and vice versa.
The parapineal organ and the habenulae are physically connected through ipsilateral axonal projections from the parapineal cells into the left habenula. Evidence is accumulating that both of these asymmetrically formed structures communicate with one another. Transgenic embryos with labeled pineal complex allow the targeting and photoablation of parapineal cells just prior to the onset of migration (Concha et al., 2003; Gamse et al., 2003; Bianco et al., 2008). In the absence of parapineal cells, the habenulae develop largely symmetric suggesting that signals derived from these cells are important for asymmetric development of the habenulae. In line with this, tbx2b mutants, in which parapineal cell development and migration are impaired, exhibit a loss of habenula asymmetry (Snelson et al., 2008). Conversely, ablations of cells forming the left habenula result in parapineal cell migration defects (Concha et al., 2003). These findings suggest that so far unknown signals derived from either structure establish the communication required for asymmetric DDC system development.
A first clue as to the signaling pathways involved in this communication comes from our previous work on Wnt/beta-catenin signaling (Carl et al., 2007). The upregulation of this pathway in axin1/masterblind mutant embryos (Heisenberg et al., 2001) results in a loss of habenula asymmetry similar to the phenotype of wild type embryos, in which parapineal cells have been removed. Intriguingly, the parapineal cells in most axin1 mutants do migrate normally to the left side of the brain suggesting that Wnt/beta-catenin signaling is involved in the communication between the parapineal cells and the habenula. However, it remains to be determined by which mechanism and in which brain structure Wnt signaling acts in this process.
Besides uncovering these communication signals it will be important to investigate whether parapineal cells misplaced to the right side of the brain have an effect on the development of the right habenula. This would provide an answer to the question whether permissive factors present only on the left side of the brain are required to establish the communication. However, transplantation of cells or tissues at advanced stages of zebrafish development is technically challenging and has so far hampered progress to resolve this issue.
The current availability of five teleost genome sequences facilitates comparative studies between evolutionary distant related species such as zebrafish and medaka to find out about conserved genetic elements underlying DDC network formation (Carl et al., 2007). This comparative approach can also be used to identify conserved and species specific axonal pathways as well as their targeting within the teleost lineage (Signore et al., 2009). It is however equally important to unravel such homologies between higher and lower vertebrates (Aizawa et al., 2011). This will allow us to relate functional consequences of network alterations in lower vertebrates to mechanisms in higher vertebrates. For instance, the asymmetries in the habenulae of mammals are rather subtle when compared to the clear neuroanatomical asymmetries in lower vertebrates (Wree et al., 1981). The habenulae in teleosts can grossly be divided into a dorsal and a ventral domain. The dorsal habenula consists of a lateral subnucleus being larger on the left side and the medial subnucleus being larger on the right, while the ventral domains show no signs of asymmetries (Figure 1B). Recently, significant progress regarding the identification of homologies has been made through the analysis of cell population specific genes and by using anterograde and retrograde dye-labelings of axonal habenula tracts (Amo et al., 2010). The dorsal habenulae in fish were found homologs to the medial habenula of mammals, while the ventral habenula in fish shows homology to the mammalian lateral habenula.
Much of the work mentioned to understand the DDC neural network is based on techniques applied to fixed embryos. These studies are starting to be successfully complemented by the use of live embryos. Combinations of transgenic techniques, DNA electroporation and the use of photoconvertible proteins have helped to identify mainly conserved elements of the DDC system such as afferent projections from different areas of the forebrain like the eminentia thalami, posterior tuberculum, and pallium (Hendricks and Jesuthasan, 2007; Miyasakan et al., 2009). Other findings were rather surprising as for example unlike in mammals, olfactory bulb neurons as well as pallium derived axons were discovered to send their axons asymmetrically into the habenulae. One theory now is that such asymmetric connections may be involved in the habenula laterality dependent differential eye use of zebrafish (Barth et al., 2005; Facchin et al., 2009) as no direct connections between the visual system and the habenulae have been discovered to date.
Like in all vertebrates analyzed to date the fasciculus retroflexus in fish connects the habenulae with the IPN and median raphe. A number of studies used lipophilic dyes to label efferent habenula axons and show that the differential targeting of left versus right sided habenula axons convert the left–right asymmetry in the dorsal diencephalon into a dorso-ventral asymmetry in the ventral midbrain (Gamse et al., 2005; Aizawa et al., 2006; Carl et al., 2007; Bianco et al., 2008). Focal DNA electroporation was used to fluorescently label single habenula efferent axons (Bianco et al., 2008). These experiments have shown that axons not only segregate laterotopically in the IPN but differ significantly in their morphology and arborization. It remains a challenging task to understand the functional significance of these results as well as the mechanism allowing these axons to cross the midline multiple times.
It has become apparent that at least habenula efferent axons receive much of their targeting information within the habenula, while information within their targets appears important for their fine tuning (Bianco et al., 2008; Roussigne et al., 2011). Axons derived from the lateral habenula cell subpopulation will target preferentially the dorsal IPN, while medial habenula cell derived axons predominantly innervate the ventral side (Roussigne et al., 2011). Axons exiting the ventral habenula domain target the median raphe nuclei (Figure 1B). However, also intermediate targets innervated by axons derived from the medial habenula subpopulation have been proposed. This suggests the presence of habenula cells or cell clusters within the three subpopulations with divergent characteristics (Aizawa et al., 2006). It appears that the currently favored subdivision into three habenula cell populations is by far oversimplified. In fact, only a very few genes found to be expressed asymmetrically across the left–right axis seem to be specifically localized to one particular habenula cell population. The kctd12.1/left over gene is expressed exclusively in lateral habenula cells (Gamse et al., 2005). Conversely kctd12.2/right on gene as well as the tg(brn3a:GFP) transgene are markers for the medial cell population (Aizawa et al., 2007). Other asymmetrically expressed genes are mostly so called right sided markers with greater expression on the right side such as kctd8/dexter and tag1 (Gamse et al., 2005; Carl et al., 2007). However, these genes are not restricted to only one subpopulation of habenula cells. This indicates that different groups of cells express different sets or combinations of genes at a given time point. Thus, the different content of information of cells or cell clusters within one habenula cell subpopulation may allow the generation of far more distinct habenula cell populations across the anterior–posterior (A–P) and the dorso-ventral (D-V) axes. These may provide different cues to efferent and possibly afferent axons and allow the establishment of stereotype axonal projection pattern important for behavior.
But how is habenula asymmetry established in the first place? Recent evidences suggest that Nodal signaling is not only required for brain laterality decisions but also for differences in neurogenesis across the midline (Roussigne et al., 2009, 2011). Furthermore, Aizawa et al. (2007) identified a time sequence of early habenula asymmetry using mutant, transgenic, and BrdU-labeling approaches on fixed embryos. Cells of the initially equally large pools of precursors start to proliferate earlier on the left side. During subsequent differentiation steps influenced by light induced melatonin production (de Borsetti et al., 2011) more cells of the early born lateral habenula cell subpopulation differentiate on the left side, possibly under the influence of Nodal signaling (Roussigne et al., 2009), while habenula cells on the right side increase their proliferation rate resulting in similar total numbers of habenula cells. Later during development, the remaining habenula precursor cells differentiate and give rise to the medial habenula cell subtype, which is consequently larger on the right side. A pathway regulating the differentiation events is the Notch signaling pathway (Aizawa et al., 2007). Using heat shock inducible constructs to transiently influence Notch signaling activity and mutant analyses Aizawa and colleagues show that Notch signaling delays habenula cell differentiation resulting in equal numbers of the later born medial cell subtype.
Mutant analyses and drug treatment experiments implicated also the Wnt/beta-catenin pathway in the establishment of habenula asymmetry (Carl et al., 2007). Wnt signaling as well as transcription factors such as Six3 (Inbal et al., 2007) are crucial regulators of asymmetric Nodal gene expression and act as early as gastrulation stages. At subsequent mid-somitogenesis stages Wnt signaling is again involved in Nodel gene expression and embryos with increased Wnt signaling develop less lateral habenula cells on the left side resulting in symmetric habenulae. Still, nothing is known as to the timing or the mechanism, by which Wnt signaling is involved in this process. However, the striking resemblance of habenula phenotypes resulting from inappropriate upregulation of either the Notch or the Wnt pathway may give a hint regarding a possible interaction during the establishment of habenula asymmetry.
Remaining Challenges and How We Can Address Them
Following DDC Network Formation
Much of the current understanding of morphological changes during embryonic brain development is based on data collected from labelings in fixed samples at different developmental stages. Thus, it is rather difficult to draw conclusions regarding the highly dynamic processes of brain development and neural network formation, because continuity is lacking when considering the course of events. The zebrafish has proven to be an excellent model to follow neural network formation during embryonic development (Aramaki and Hatta, 2006). Although the DDC system is well accessible in the zebrafish, it spans several hundred micrometers in both the A–P and the D-V directions, and following the entire DDC system development over time requires time-lapse imaging for at least 72 h.
Conventional confocal laser scan microscopy (CLSM) is commonly used for time-lapse imaging. However, the effect of photobleaching of the fluorescence requires increasing laser power over time, which in turn results in phototoxicity (Squirrell et al., 1999). In fact, our attempts to perform time-lapse recordings of GFP-transgenic embryos for more than 40 h resulted in the death of the embryo.
Two-photon microscopy uses a very restricted excitation volume and longer excitation wavelengths, which circumvents the problem of photobleaching and photodamage (Helmchen and Denk, 2005; Svoboda and Yasuda, 2006). We used this technique to image the novel zebrafish enhancer trap line Et(-1.0otpa:mmGFP)hd1, which expresses GFP in possibly all cells of the habenulae and its efferent projections starting as early as 32 hours post fertilization (hpf; Figure 2 and data not shown). Applying a similar approach to that previously used to image developing blood vessels (Kamei and Weinstein, 2005), no detectable harm to the animal or a decrease in GFP intensity was observed. The pictures show GFP expressing habenula cells in the dorsal diencephalon at 38 and 47 hpf and habenula afferent projections entering the midbrain at 57 hpf, eventually innervating the IPN about 300 μm away from their source in the ventral midbrain (Figures 2A–D).
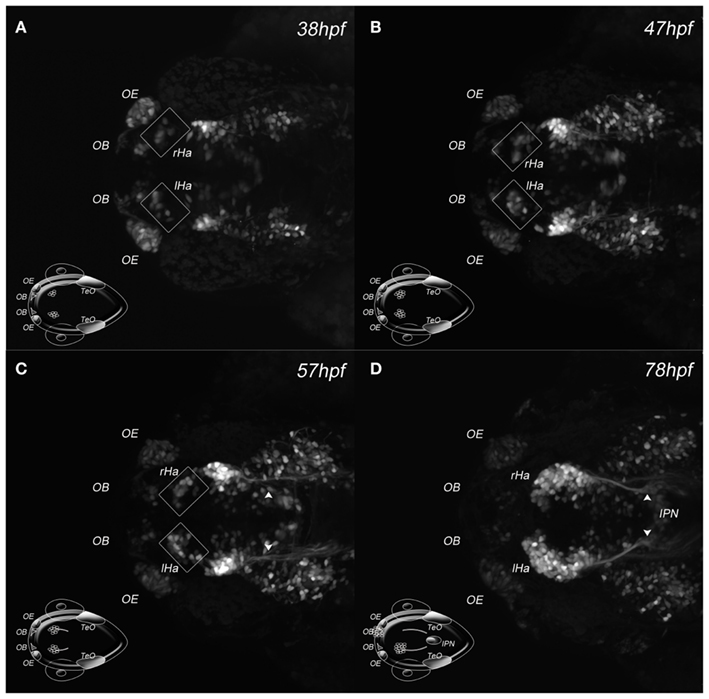
Figure 2. Two-photon excitation of DDC system architecture. (A–D) Maximum intensity projections of four developmental stages acquired by in vivo 2PM of a Et(-1.0otpa:mmGFP)hd1 transgenic zebrafish embryo; dorsal views, anterior left; developmental stages are indicated. The cartoon inserts show regions of GFP expressing cells for orientation. (A–C) Squares frame GFP expressing habenular cells. (C) The arrowheads mark habenula efferent axons. (D) The arrowheads mark habenula efferent axons entering the IPN. Ha, habenula; IPN, interpeduncular nucleus; l, left; OB, olfactory bulb; OE, olfactory epithelium; r, right; TeO, optic tectum.
We next compared the efficacy of 2PM versus CLSM to image deep into the brain and record the axonal innervation of the IPN (not shown). CLSM was never able to reach the depth attained by 2PM, being limited to a penetration of ∼100–120 μm in comparison to >400 μm achieved in two-photon excitation (2PE), where the final limitation was due to the absence of tagged cells. Figure 3 shows the comparison between images of the IPN of embryos recorded by CLSM (Figures 3A–D) and 2PM (Figures 3E–H) at different depths using partial maximum intensity projections of the whole Z-stack. Note that IPN structures are still visible reaching the maximum imaging depth using the CLSM (Figure 3D).
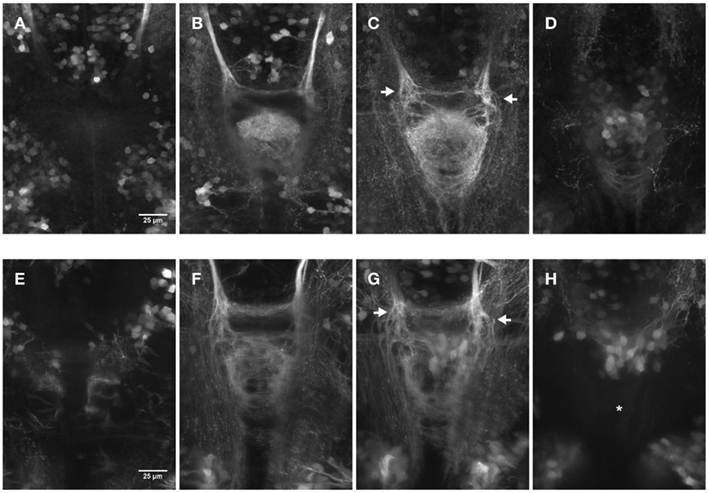
Figure 3. Comparison of dorsal views (150 × 200 μm) of two 4 day old embryos focused on the IPN. The pictures show partial maximum intensity projections for CLSM (A–D) and 2PM (E–H). (A–D) Maximum intensity projections of a range of 15 μm each for a total Z-height of 60 μm. (E–H) Maximum intensity projections of a range of 25 μm each for a total Z-height of 100 μm. Different focal planes are pictured due to the differences in range. Laser power correction was used in both cases to compensate for increasing depth. The display range is kept constant within the series and the gamma was corrected to 0.5 using FiJi for display purpose. (C–G) The arrows highlight the axons entering the IPN area more dorsal on the left and more ventral on the right side. Asterisk in (H): less structures can be observed compared to (D) due to the fact that (H) shows a deeper range past the IPN that view (D) does (9–34 μm versus 2–17 μm after the “end” of the IPN).
These experiments show the obvious advantages of 2PM over CLSM when working in vivo on whole animals: besides the possibility of long-term time-lapse imaging without harming the specimens, the imaging depth of 2PM in thick samples or tissues is far superior to that of CLSM.
The number of transgenic zebrafish lines that label cells of the DDC system has increased exponentially during the last years. Using 2PM time-lapse imaging we can now analyze the spatio-temporal formation of the entire asymmetric DDC system. We have the exciting possibility to learn all about the origins of habenula cells, timing of events during DDC system development and axonal targeting and shaping. These are the essential prerequisites if we are to analyze the consequences of genetic or laser manipulations in their full spectrum.
Future Goals
Progress has been made as to the genetic network underlying the asymmetric formation of the DDC system. Several pathways including Wnt/beta-catenin, Notch, Fgf, and Nodal have been identified but their epistatic relationship remains unknown. Which factors are involved in the communication of parapineal cells and the habenulae? Which are the molecules important for guidance of habenula projections, their targeting and shaping? Some of these questions can readily be addressed while others need creativity and the development of new resources and techniques. A variety of tools to manipulate genetic pathways in time are already available ranging from signal cascade specific drugs to transgenic animals harboring heat shock inducible constructs. For instance, the established infrared laser based “laser evoked gene operator (LEGO)” for focal activation of such constructs offers the exciting possibility to study the consequences of manipulating genes in one particular region of the brain at a given time point (Deguchi et al., 2009; Kamei et al., 2009). Also the application of the Gal4–VP16 system in combination with tetanus toxin or nitroreductase to specifically ablate particular structures of the DDC system have been used successfully (Agetsuma et al., 2010; Lee et al., 2010). Other tools are still missing such as for instance promoters that drive gene expression in defined areas of the circuit to not only visualize or ablate these structures but as well to misexpress genes of interest and analyze the consequences on DDC system development. Also techniques for simultaneous timely and spatially controlled activation of genes are rather limited in the zebrafish.
The targeting of axonal projections exiting the habenulae is a good read out as to the nature of the habenula cell types. Based on this targeting and on the expression of marker genes the habenulae have been divided into lateral, medial, and ventral cell subpopulations. However, as outlined above the situation appears much more complex and one can easily imagine that many more cell subtypes could be distinguished. For instance, what makes some cells of the medial habenula cell subpopulation send their axons into an intermediate area of the IPN rather than into the ventral part (Aizawa et al., 2006)? Focal electroporation techniques have been applied to follow single axons innervating the IPN. Although this being a very informative method, it is very difficult and laborious to apply. The same holds true for the tracking of single axons in networks of such complexity. With the development of reliable multicolor labeling techniques on fixed as well as living animals we may be able to approach this complex task. For instance, one promising tool to map habenula axonal projections that has been used to study retino-tectal projections in mammals is the brainbow technique (Livet et al., 2007). The unique, randomly generated combinatorial expression of various fluorescent proteins allows researchers to label cells and their axons in various colors (over 100) to discriminate between them. This technique (Pan et al., 2011) requires the identification of genes with appropriate promoters, which drive gene expression in habenula precursor cells and their descendants such as the cxcr4b gene (Roussigne et al., 2009). Moreover, this labeling technique will have to be combined with super-resolution light microscopy to allow the unique identification of neurites (Helmstaedter et al., 2008).
The use of in vivo long-term time-lapse 2PM to study DDC system development in the living embryo will give us many clues as to the timing of neural circuit developmental processes. Moreover, we will be able to compare DDC system development between normal and mutant embryos and study the impact of the mutated gene in great detail. Although progress has been made, the lack of appropriate transgenic lines or vital dyes to mark specific embryonic structures to be used as landmarks is currently hampering the rapid detailed description of axonal targeting. It also remains difficult to actually trace single axons in embryos in which the entire circuit is labeled. Promoters are needed that drive for instance the photoconvertible Kaede protein (Ando et al., 2002) or the recently reported photoswitchable monomeric orange (PSmOrange) protein in habenula precursor cells (Subach et al., 2011). Photoconversion of the Kaede protein switches its fluorescence from green to red while the PSmOrange is converted from orange into far-red fluorescence. Photoconversion of single cells or cell clusters in such a line followed by long-term multiphoton time-lapse microscopy would be a promising tool to learn more about the development of the DDC system architecture.
We have now acquired the knowledge, tools, and resources to visualize the DDC system in the living animal. Moreover we can manipulate habenula laterality and asymmetry either genetically or by leaving the embryo genetically unchanged. We even have the means to introduce rather subtle changes in neural activities using a variety of optogenetic tools that have been reviewed elsewhere (Del Bene and Wyart, 2011). But before we manipulate neural activity we need to visualize it for comparison between normal and manipulated embryos. Increasing numbers of genetically encoded reporters of neural activity have been generated to study the locomotor, olfactory, and visual systems. The usefulness of genetically encoded calcium indicators (GECIs) for visualizing neural activity upon electrical stimulation or touch escape responses in the living zebrafish was first demonstrated in 2003 (Higashijima et al., 2003). Since then, a multitude of reporters have been developed which can also be stimulated by odors (Li et al., 2005) or light (Sumbre et al., 2008; Del Bene et al., 2010; Naumann et al., 2010). Although these reporters are promising tools, they only indirectly measure neural activity as they visualize changes in Ca2+-levels. Recently, the improved voltage sensor Mermaid has been reported to allow recordings of spikes in the fish similar to action potentials (Tsutsui et al., 2010). It will be exciting to generate transgenic animals, which express such reporters under the control of habenula cell population specific promoters to investigate neural activity upon stimulation. This will allow to unravel the functional properties of the DDC system.
The next step then is to use readily available and constantly improving tools to selectively inhibit or activate neural activity of habenula neurons (Del Bene and Wyart, 2011) and to investigate the effect on larval behavior. This approach can be refined by for instance making use of the differing excitations spectra of microbial opsins used to modulate neural activity and to perform a combinatorial analysis of the effect of activating neurons of different habenula subpopulations in the same animal. One caveat of using zebrafish is the existence of only a few reliable behavior assays that correlate habenula laterality or asymmetry with behavior and in these cases there has been no direct demonstration that habenular circuits are involved in the described behaviors (Barth et al., 2005; Facchin et al., 2009). The habenulae have been reported to be involved in syndromes such as depression and schizophrenia but we are far from having the possibility to use the fish as a disease model for such syndromes. One successfully applied test perhaps closest to a human depression syndrome is an escape assay, which implicated that afferent habenula projections derived from the telencephalon are important in this process (Lee et al., 2010). The usefulness for automated behavior and drug testing makes the zebrafish an exquisite model for large scale screens (Gerlai, 2010; Zhong and Lin, 2011). Furthermore, the vast variety of techniques and resources available for the zebrafish together with the enormous potential of a research field such as neurological diseases should be encouraging enough for scientists to think about novel readout systems.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Steve Wilson for thoughts and comments on the manuscript, Peter Bankhead for the MIP script and Ulrike Engel for support. This work was supported by the Behrens-Weise Stiftung and Max Planck Society to Soojin Ryu and DFG-SFB488-A18, DFG CA298/3-1, and the Wnt-Forschergruppe 1036 (AP-2) to Matthias Carl.
References
Agetsuma, M., Aizawa, H., Aoki, T., Nakayama, R., Takahoko, M., Goto, M., Sassa, T., Amo, R., Shiraki, T., Kawakami, K., Hosoya, T., Higashijima, S., and Okamoto, H. (2010). The habenula is crucial for experience-dependent modification of fear responses in zebrafish. Nat. Neurosci. 13, 1354–1356.
Aizawa, H., Bianco, I. H., Hamaoka, T., Miyashita, T., Uemura, O., Concha, M. L., Russell, C., Wilson, S. W., and Okamoto, H. (2006). Laterotopic representation of left-right information onto the dorso-ventral axis of a zebrafish midbrain target nucleus. Curr. Biol. 15, 238–243.
Aizawa, H., Goto, M., Sato, T., and Okamoto, H. (2007). Temporally regulated asymmetric neurogenesis causes left-right difference in the zebrafish habenular structures. Dev. Cell 12, 87–98.
Aizawa, H., Amo, R., and Okamoto, H. (2011). Phylogeny and ontogeny of the habenular structure. Front. Neurosci. 5, 138.
Amo, R., Aizawa, H., Takahoko, M., Kobayashi, M., Takahashi, R., Aoki, T., and Okamoto, H. (2010). Identification of the zebrafish ventral habenula as a homolog of the mammalian lateral habenula. J. Neurosci. 30, 1566–1574.
Ando, R., Hama, H., Yamamoto-Hino, M., Mizuno, H., and Miyawaki, A. (2002). An optical marker based on the UV-induced green-to-red photoconversion of a fluorescent protein. Proc. Natl. Acad. Sci. U.S.A. 99, 12651–12656.
Aramaki, S., and Hatta, K. (2006). Visualizing neurons one-by-one in vivo: optical dissection and reconstruction of neural networks with reversible fluorescent proteins. Dev. Dyn. 235, 2192–2199.
Barth, K. A., Miklosi, A., Watkins, J., Bianco, I. H., Wilson, S. W., and Andrew, R. J. (2005). fsi zebrafish show concordant reversal of laterality of viscera, neuroanatomy, and a subset of behavioral responses. Curr. Biol. 15, 844–850.
Bianco, I. H., Carl, M., Russell, C., Clarke, J. D., and Wilson, S. W. (2008). Brain asymmetry is encoded at the level of axon terminal morphology. Neural Dev. 3, 9.
Bianco, I. H., and Wilson, S. W. (2009). The habenular nuclei: a conserved asymmetric relay station in the vertebrate brain. Phil. Trans. R. Soc. B Biol. Sci. 364, 1005–1020.
Bisazza, A., Rogers, L. J., and Vallortigara, G. (1998). The origins of cerebral asymmetry: a review of evidence of behavioural and brain lateralization in fishes, reptiles and amphibians. Neurosci. Biobehav. Rev. 22, 411–426.
Broca, P. P. (1861). Remarques sur la siège de la faculté du langage ariticulè, suivies d’une observation d’apèmie. Bull. Soc. Anatomique 2, 330–357.
Brown, T. G. (1914–1915). Note on the physiology of the basal ganglia and midbrain of the anthropoid ape, especially in reference to the act of laughter. J. Physiol. 47, 195–207.
Cajal, S. R. Y. (1911). Histologie du Systeme Nerveux de l’homme et des Vertebres, Vol. 2. Parie: Maloine, 270–275.
Carl, M., Bianco, I. H., Bajoghli, B., Aghaallaei, N., Czerny, T., and Wilson, S. W. (2007). Wnt/Axin1/beta-catenin signaling regulates asymmetric nodal activation, elaboration, and concordance of cns asymmetries. Neuron 55, 393–405.
Concha, M. L., Burdine, R. D., Russell, C., Schier, A. F., and Wilson, S. W. (2000). A nodal signaling pathway regulates the laterality of neuroanatomical asymmetries in the zebrafish forebrain. Neuron 28, 399–409.
Concha, M. L., Russell, C., Regan, J. C., Tawk, M., Sidi, S., Gilmour, D. T., Kapsimali, M., Sumoy, L., Goldstone, K., Amaya, E., Kimelman, D., Nicolson, T., Gründer, S., Gomperts, M., Clarke, J. D., and Wilson, S. W. (2003). Local tissue interactions across the dorsal midline of the forebrain establish CNS laterality. Neuron 39, 423–438.
Concha, M. L., and Wilson, S. W. (2001). Asymmetry in the epithalamus of vertebrates. J. Anat. 199, 63–84.
de Borsetti, N. H., Dean, B. J., Bain, E. J., Clanton, J. A., Taylor, R. W., and Gamse, J. T. (2011). Light and melatonin schedule neuronal differentiation in the habenular nuclei. Dev. Biol. 358, 251–261.
Deguchi, T., Itoh, M., Urawa, H., Matsumoto, T., Nakayama, S., Kawasaki, T., Kitano, T., Oda, S., Mitani, H., Takahashi, T., Todo, T., Sato, J., Okada, K., Hatta, K., Yuba, S., and Kamei, Y. (2009). Infrared laser-mediated local gene induction in medaka, zebrafish and Arabidopsis thaliana. Dev. Growth Differ. 51, 769–775.
Del Bene, F., and Wyart, C. (2011). Optogenetics: a new enlightenment age for zebrafish neurobiology. Dev. Neurobiol. 72, 404–414.
Del Bene, F., Wyart, C., Robles, E., Tran, A., Looger, L., Scott, E. K., Isacoff, E. Y., and Baier, H. (2010). Filtering of visual information in the tectum by an identified neural circuit. Science 330, 669–673.
Driever, W., Solnica-Krezel, L., Shier, A. F., Neuhaus, S. C. F., Malicki, J., Stemple, D. L., Stanier, D. Y. R., Zwartkruis, F., Abdelilah, S., Rangini, Z., Belak, J., and Boggs, C. (1996). A genetic screen for mutations affecting embryogenesis in zebrafish. Development 123, 37–46.
Facchin, L., Burgess, H. A., Siddiqi, M., Granato, M., and Halpern, M. E. (2009). Determining the function of zebrafish epithalamic asymmetry. Philos. Trans. R. Soc. Lond. B Biol. Sci. 364, 1021–1032.
Forel, A. (1872). Beitrage zur kenntniss des thalamus opticus und der ihn umgebenden gebilde bei den saugethieren. Sitzber. k. Akad. Wiss. Wien. 65–66, 25–58.
Gamse, J. T., Kuan, Y. S., Macurak, M., Brosamle, C., Thisse, B., Thisse, C., and Halpern, M. E. (2005). Directional asymmetry of the zebrafish epithalamus guides dorsoventral innervation of the midbrain target. Development 132, 4869–4881.
Gamse, J. T., Thisse, C., Thisse, B., and Halpern, M. E. (2003). The parapineal mediates left-right asymmetry in the zebrafish diencephalon. Development 130, 1059–1068.
Gerlai, R. (2010). High-throughput behavioral screens: the first step towards finding genes involved in vertebrate brain function using zebrafish. Molecules 15, 2609–2622.
Goronowitsch, N. (1883). Das gehirn und die cranialnerven von Acipenser ruthenus. Morphol. Jahrb. 13, 427–574.
Guglielmotti, V., and Cristino, L. (2006). The interplay between the pineal complex and the habenular nuclei in lower vertebrates in the context of the evolution of cerebral asymmetry. Brain Res. Bull. 69, 475–488.
Guillery, R. W. (1959). Afferent fibers to the dorso-medial thalamic nucleus in the cat. J. Anat. 93, 403–419.
Gutiérrez-Ibáñez, C., Reddon, A. R., Kreuzer, M. B., Wylie, D. R., and Hurd, P. L. (2011). Variation in asymmetry of the habenular nucleus correlates with behavioural asymmetry in a cichlid fish. Behav. Brain Res. 221, 189–196.
Halpern, M. E., Liang, J. O., and Gamse, J. T. (2003). Leaning to the left: laterality in the zebrafish forebrain. Trends Neurosci. 26, 308–313.
Hamada, H., Meno, C., Watanabe, D., and Saijoh, Y. (2002). Establishment of vertebrate left-right asymmetry. Nat. Rev. Genet. 3, 103–113.
Hamill, G. S., and Lenn, N. J. (1984). The subnuclear organization of the rat interpeduncular nucleus: a light and electron microscopic study. J. Comp. Neurol. 222, 396–408.
Hattori, T., McGeer, E. G., Singh, V. K., and McGeer, P. L. (1977). Cholinergic synapse of the interpeduncular nucleus. Exp. Neurol. 55, 666–679.
Heisenberg, C. P., Houart, C., Take-uchi, M., Rauch, G. J., Young, N., Coutinho, P., Masai, I., Caneparo, L., Concha, M. L., Geisler, R., Dale, T. C., Wilson, S. W., and Stemple, D. L. (2001). A mutation in the Gsk3-binding domain of zebrafish masterblind/axin1 leads to a fate transformation of telencephalon and eyes to diencephalon. Genes Dev. 15, 1427–1434.
Helmstaedter, M., Briggman, K. L., and Denk, W. (2008). 3D structural imaging of the brain with photons and electrons. Curr. Opin. Neurobiol. 18, 633–641.
Hendricks, M., and Jesuthasan, S. (2007). Asymmetric innervation of the habenula in zebrafish. J. Comp. Neurol. 502, 611–619.
Herkenham, M., and Nauta, W. J. H. (1977). Afferent connections of the habenular nuclei in the rat. A horseradish peroxidase study, with a note on the fiber-of-passage problem. J. Comp. Neurol. 173, 123–146.
Higashijima, S., Masino, M. A., Mandel, G., and Fetcho, J. R. (2003). Imaging neuronal activity during zebrafish behavior with a genetically encoded calcium indicator. J. Neurophysiol. 90, 3986–3997.
Hikosaka, O. (2010). The habenula: from stress evasion to value-based decision-making. Nat. Neurosci. 11, 503–513.
Inbal, A., Kim, S. H., Shin, J., and Solnica-Krezel, L. (2007). Six3 represses nodal activity to establish early brain asymmetry in zebrafish. Neuron 55, 407–415.
Jhou, T. C., Fields, H. L., Baxter, M. G., Saper, C. B., and Holland, P. C. (2009a). The rostromedial tegmental nucleus (RMTg), a GABAergic afferent to midbrain dopamine neurons, encodes aversive stimuli and inhibits motor responses. Neuron 61, 786–800.
Jhou, T. C., Geisler, S., Marinelli, M., DeGarmo, B., and Zahm, D. S. (2009b). The mesopontine rostromedial tegmental nucleus: a structure targeted by the lateral habenula that projects to the ventral tegmental area of Tsai and substantia nigra compacta. J. Comp. Neurol. 513, 566–596.
Kamei, M., and Weinstein, B. M. (2005). Long-term time-lapse fluorescence imaging of developing zebrafish. Zebrafish 2, 113–123.
Kamei, Y., Suzuki, M., Watanabe, K., Fujimori, K., Kawasaki, T., Deguchi, T., Yoneda, Y., Todo, T., Takagi, S., Funatsu, T., and Yuba, S. (2009). Infrared laser-mediated gene induction in targeted single cells in vivo. Nat. Methods 6, 79–81.
Kuhar, M. J., DeHaven, R. N., Yamamura, H. I., Rommel-Spacher, H., and Simon, J. R. (1975). Further evidence for cholinergic habenulo-interpeduncular neurons: pharmacologic and functional characteristics. Brain Res. 97, 265–275.
Lee, A., Mathuru, A. S., Teh, C., Kibat, C., Korzh, V., Penney, T. B., and Jesuthasan, S. (2010). The habenula prevents helpless behavior in larval zebrafish. Curr. Biol. 20, 1–6.
Lenn, N. J. (1976). Synapses in the interpeduncular nucleus: Electron microscopy of normal and habenula lesioned rats. J. Comp. Neurol. 166, 73–100.
Li, J., Mack, J. A., Souren, M., Yaksi, E., Higashijima, S., Mione, M., Fetcho, J. R., and Friedrich, R. W. (2005). Early development of functional spatial maps in the zebrafish olfactory bulb. J. Neurosci. 25, 5784–5795.
Livet, J., Weissman, T. A., Kang, H., Draft, R. W., Lu, J., Bennis, R. A., Sanes, J. R., and Lichtman, J. W. (2007). Transgenic strategies for combinatorial expression of fluorescent proteins in the nervous system. Nature 450, 56–62.
Milhaud, M., and Pappas, G. D. (1966). The fine structure of neurons and synapses of the habenula of the cat with special reference to subjunctional bodies. Brain Res. 3, 158–173.
Mitchell, R. (1963). Connections of the habenula and of the interpeduncular nucleus in the cat. J. Comp. Neurol. 121, 441–457.
Miyasakan, N., Morimoto, K., Tsubokawa, T., Higashijima, S., Okamoto, H., and Yoshihara, Y. (2009). From the olfactory bulb to higher brain centers: genetic visualization of secondary olfactory pathways in zebrafish. J. Neurosci. 30, 1566–1574.
Morgane, P. J., Galler, J. R., and Mokler, D. J. (2005). A review of systems and networks of the limbic forebrain/limbic midbrain. Prog. Neurobiol. 75, 143–160.
Naumann, E. A., Kampff, A. R., Prober, D. A., Schier, A. F., and Engert, F. (2010). Monitoring neural activity with bioluminescence during natural behavior. Nat. Neurosci. 13, 513–520.
Pan, Y. A., Livet, J., Sanes, J. R., Lichtman, J. W., and Schier, A. F. (2011). “Multicolor brainbow imaging in zebrafish,” in Imaging in Developmental Biology: A Laboratory Manual, eds R. Wong, and J. Sharpe (Cold Spring Harbor: Cold Spring Harbor Laboratory Press), 191–198.
Pierce, E. T., Foote, W. E., and Hobson, J. A. (1976). The efferent connection of the nucleus raphe dorsalis. Brain Res. 107, 137–144.
Powell, E. W. (1968). Septohabenular connections in the rat, cat and monkey. J. Comp. Neurol. 134, 145–150.
Regan, J. C., Concha, M. L., Roussigne, M., Russell, C., and Wilson, S. W. (2009). An Fgf8-dependent bistable cell migratory event establishes CNS asymmetry. Neuron 61, 27–34.
Rodriguez-Esteban, C., Tsukui, T., Yonei, S., Magallon, J., Tamura, K., and Belmonte, J. C. I. (1999). The T-box genes Tbx4 and Tbx5 regulate limb outgrowth and identity. Nature 398, 814–818.
Roussigne, M., Bianco, I. H., Wilson, S. W., and Blader, P. (2009). Nodal signalling imposes left-right asymmetry upon neurogenesis in the habenular nuclei. Development 136, 1549–1557.
Roussigne, M., Blader, P., and Wilson, S. W. (2011). The zebrafish epithalamus clears a path through the complexity of brain lateralization. Dev. Neurobiol. 72, 269–281.
Signore, I. A., Guerrero, N., Loosli, F., Colombo, A., Villalón, A., Wittbrodt, J., and Concha, M. L. (2009). Zebrafish and medaka: model organisms for a comparative developmental approach of brain asymmetry. Philos. Trans. R. Soc. Lond. B Biol. Sci. 364, 991–1003.
Smaha, L. A., and Kaelber, W. W. (1973). Efferent fiber projections of the habenula and the interpeduncular nucleus. An experimental study in the opossum and cat. Exp. Brain Res. 16, 291–308.
Snelson, C. D., Santhakumar, K., Halpern, M. E., and Gamse, J. T. (2008). Tbx2b is required for the development of the parapineal organ. Development 135, 1693–1702.
Squirrell, J. M., Wokosin, D. L., White, J. G., and Bavister, B. D. (1999). Long-term two-photon fluorescence imaging of mammalian embryos without compromising viability. Nat. Biotechnol. 17, 763–767.
Staines, W. A., Nagy, J. I., Vincent, S. R., and Fibiger, H. C. (1980). Neurotransmitters contained in the efferents of the striatum. Brain Res. 194, 391–402.
Subach, O. M., Patterson, G. H., Ting, L.-M., Wang, Y., Condeelis, J. S., and Verkhusha, V. V. (2011). A photoswitchable orange-to-far-red fluorescent protein, PSmOrange. Nat. Methods 8, 771–777.
Sumbre, G., Muto, A., Baier, H., and Poo, M. M. (2008). Entrained rhythmic activities of neuronal ensembles as perceptual memory of time interval. Nature 456, 102–106.
Sutherland, R. J. (1982). The dorsal diencephalic conduction system: a review of the anatomy and functions of the habenular complex. Neurosci. Biobehav. Rev. 6, 1–13.
Svoboda, K., and Yasuda, R. (2006). Principles of two-photon excitation microscopy and its applications to neuroscience. Neuron 50, 823–839.
Tsutsui, H., Higashijima, S. I., Miyawaki, A., and Okamura, Y. (2010). Visualizing voltage dynamics in zebrafish heart. J. Physiol. (Lond.) 588, 2017–2021.
Vallortigara, G., and Rogers, L. J. (2005). Survival with an asymmetrical brain: advantages and disadvantages of cerebral lateralization. Behav. Brain Sci. 28, 575–633.
Vallortigara, G., Rogers, L. J., and Bisazza, A. (1999). Possible evolutionary origins of cognitive brain lateralization. Brain Res. Brain Res. Rev. 30, 164–175.
von Staden, H. (2007). Herophilus: The Art of Medicine in Early Alexandria. Cambridge: University Press.
Wree, A., Zilles, K., and Schleicher, A. (1981). Growth of fresh volumes and spontaneous cell death in the nuclei habenulae of albino rats during ontogenesis. Anat. Embryol. (Berl.) 161, 419–431.
Zhong, H., and Lin, S. (2011). Chemical screening with zebrafish embryos. Methods Mol. Biol. 716, 193–205.
Keywords: habenula, asymmetry, zebrafish, DDC, epithalamus, neural circuit, 2PM imaging
Citation: Beretta CA, Dross N, Guiterrez-Triana JA, Ryu S and Carl M (2012) Habenula circuit development: past, present, and future. Front. Neurosci. 6:51. doi: 10.3389/fnins.2012.00051
Received: 31 January 2012; Paper pending published: 14 February 2012;
Accepted: 27 March 2012; Published online: 23 April 2012.
Edited by:
Steffen Scholpp, Karlsruhe Institute of Technology, GermanyReviewed by:
Martin Meyer, King’s College London, UKHidenori Aizawa, Tokyo Medical and Dental University, Japan
Copyright: © 2012 Beretta, Dross, Guiterrez-Triana, Ryu and Carl. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: Matthias Carl, Department of Cell and Molecular Biology, Medical Faculty Mannheim, Heidelberg University, Ludolf-Krehl Strasse 13-17, 68167 Mannheim, Germany. e-mail:bWF0dGhpYXMuY2FybEBtZWRtYS51bmktaGVpZGVsYmVyZy5kZQ==
 Carlo A. Beretta1
Carlo A. Beretta1