- Department of Physiology and Medical Physics, Royal College of Surgeons in Ireland, Dublin, Ireland
Insults to the brain that are sub-threshold for damage activate endogenous protective pathways, which can temporarily protect the brain against a subsequent harmful episode. This mechanism has been named as tolerance and its protective effects have been shown in experimental models of ischemia and epilepsy. The preconditioning-stimulus can be a short period of ischemia or mild seizures induced by low doses of convulsant drugs. Gene-array profiling has shown that both ischemic and epileptic tolerance feature large-scale gene down-regulation but the mechanism are unknown. MicroRNAs are a class of small non-coding RNAs of ~20–22 nucleotides length which regulate gene expression at a post-transcriptional level via mRNA degradation or inhibition of protein translation. MicroRNAs have been shown to be regulated after non-harmful and harmful stimuli in the brain and to contribute to neuroprotective mechanisms. This review focuses on the role of microRNAs in the development of tolerance following ischemic or epileptic preconditioning.
Preconditioning
The concept of preconditioning was first described in a model of heart ischemia. Here, brief ischemic episodes protected against a subsequent ischemic insult (Murry et al., 1986), opening a new way for the treatment of heart diseases. Several preconditioned treatment paradigms have been used in the clinic (including remote preconditioning stimulus) to protect patients against an ischemic insult in heart pathologies (McCafferty et al., 2014).
Preconditioning is an adaptive response where a small dose of a harmful substance protects the brain from a subsequent damaging insult (Murry et al., 1986; Stenzel-Poore et al., 2007; Lin et al., 2008; Dirnagl et al., 2009; Assaf et al., 2011). The fact that any injury to the brain applied below the threshold of cell damage, including seizures, will induce preconditioning and neuroprotection to the brain shows the importance of understanding the mechanism underlying preconditioning and its potential as novel treatment option in brain disorders.
The preconditioning stimulus induces a transient state of protection which is called tolerance. Early work demonstrated that de novo protein synthesis was required for tolerance; this implied regulation of gene expression in a time-specific manner and having an effect only several days after the precondition stimulus (Barone et al., 1998).
Preconditioning can induce neuroprotection over two time-frames: (1) Rapid tolerance, which happens in a short period of time and is independent of protein production and is associated with synapse remodeling (Meller et al., 2008). (2) Delayed (classical) tolerance, which evolves over 1–3 days post-preconditioning and requires de novo protein production which peaks after 3 days and diminishes over the course of the following week (Stenzel-Poore et al., 2007).
Preconditioning Stimuli
Preconditioning can be induced by several stimuli in the brain, including non-injurious ischemia, cortical spreading depression, brief episodes of seizures and low doses of endotoxins (LPS, lipopolysaccharide) (Kitagawa et al., 1990; Simon et al., 1993; Kobayashi et al., 1995; Chen and Simon, 1997; Towfighi et al., 1999). A cross-tolerance phenomenon has been recognized where stimuli and challenge are from different nature. For example, a brief-seizure stimulation or LPS injection can protect against a subsequent ischemic injury (Plamondon et al., 1999; Towfighi et al., 1999).
Data has shown that the reprogramming of genes involved in cellular response to excitotoxic insults plays an important role in the preconditioning process of different tissues (Mirrione et al., 2010). Evidence from genomic studies has demonstrated that diverse stimuli which trigger neuroprotection achieved by preconditioning may share a common process which depends on a fundamental reprogramming of the response to injury, e.g., in ischemia or epileptic tolerance has been shown that 80% of genes are down-regulated. This reprogramming process can induce novel neuroprotective pathways which in turn lead to the synthesis of new proteins changing the molecular and genetic response to subsequent injury (Stenzel-Poore et al., 2004, 2007).
In experimental models of status epilepticus, several preconditioning stimuli including bicuculline, electroshock, kindling, and low dose injection of the convulsant kainic acid, has shown to reduce brain damage after status epilepticus (Amini et al., 2014). Gene profiling has shown that down-regulation of gene expression is a general mechanism underlying the tolerance process (Borges et al., 2007; Jimenez-Mateos et al., 2008; Meller et al., 2008; Della-Morte et al., 2012), including downregulation of genes involved in ion channels, calcium signaling and excitability (Jimenez-Mateos et al., 2008).
microRNAs
The term MicroRNA, previously called small temporal RNAs (stRNAs) due to their role in the regulation of developmental timing of Caernohabditis elegans, was first introduced in 2001 (Lagos-Quintana et al., 2001; Lau et al., 2001; Lee and Ambros, 2001). Originally microRNAs were defined as small non-coding RNAs (~20–22 nucleotides) that regulate post-transcriptional gene expression in a sequence-specific manner. Since their discovery in C. elegans, microRNAs were found to be expressed in invertebrates and vertebrates, including humans, and many show highly-conservation during evolution (Lagos-Quintana et al., 2001).
MiRNAs regulate gene expression via translational inhibition, mRNA degradation or a combination of both mechanisms (O'Carroll and Schaefer, 2013). In the brain, however, miRNA targeting is frequently not associated with reduced mRNA levels of targets (Klein et al., 2007). Almost 50% of all identified miRNAs are expressed in the mammalian brain and there is significant cell- and region-specific distribution reflecting roles in gene expression that direct the functional specialization of neurons and the morphological responses that are required to adapt to their continuously changing activity state (Siegel et al., 2011; O'Carroll and Schaefer, 2013). MiRNAs and their biogenesis components display select localization within neurons, with significant enrichment in dendrites, enabling local, activity-dependent miRNA regulation of protein levels (Lugli et al., 2005, 2008; O'Carroll and Schaefer, 2013). Recent work demonstrated that certain pre-miRs have localization signals which translocate them to the synaptic sites where they are processed into mature miRNA (Bicker et al., 2013).
Most miRNA genes are expressed as a cluster within a single poly-cistronic transcript (Baskerville and Bartel, 2005) and transcribed by RNA polymerase II, resulting in the primary sequence (Cai et al., 2004; Lee et al., 2004). In the nucleus the primary sequence is recognized by the nuclear microprocessor complex containing Drosha and DGCR8 (DiGeorge syndrome critical region 8) which generate the precursor-miRNA (pre-miR) of ~60–70 nt length (Lee et al., 2003). Next, the precursor is exported into the cytoplasm by Exportin-5 in a GTP-dependent manner (Bohnsack et al., 2004). Once in the cytoplasm, microRNAs are further processed by Dicer to a single-strand RNA mature form (Lagos-Quintana et al., 2001; Lau et al., 2001; Lee and Ambros, 2001). Finally, a single strand of the mature miRNA is selected and loaded into the RISC complex and bound to the members of the Argonaute (Ago) protein family. This miRNA-bound-RISC complex is the functional and active form of the miRNA which then targets the mRNA. Binding of a 7–8 nt seed region, usually within the 3′UTR of the mRNA, results in either inhibition of the translation or mRNA degradation. This process requires the activation of de-cap enzymes and removal of the poly-A tail (Kwak and Tomari, 2012).
Several studies have shown the role of miRNAs in the brain by using mainly genetic tools in mice, including conditional or full deletion of biogenesis enzymes in the microRNA pathway. Deletion of DGCR8, which affects the production of the precursor microRNA, results in a reduction in brain size and loss of inhibitory synaptic neurotransmission (Babiarz et al., 2011; Hsu et al., 2012). Conditional deletion of Drosha in neural progenitors did not affect neurogenesis in the developing brain, but did affect differentiation and migration of neurons (Knuckles et al., 2012). Deletion of Dicer from neurons produces severe brain abnormalities, including microencephaly and defects in dendritic arborization in cortex and hippocampus (Davis et al., 2008; De Pietri Tonelli et al., 2008; Babiarz et al., 2011; Dorval et al., 2012). Mice lacking Dicer in astrocytes develop spontaneous seizures and suffer from increased premature mortality (Tao et al., 2011). These data imply that the miRNA biogenesis system is essential for brain development and function. Surprisingly, one study reported that specific deletion of Dicer in the adult mouse forebrain transiently enhanced learning and memory, although these animals later displayed degeneration of neurons in the cortex and hippocampus (Konopka et al., 2010).
Analysis of Argonaute (Ago1–4) proteins, main components of the RISC complex, has given more diverse results (Burroughs et al., 2011). Ago-2 is the most abundant expressed member in various tissues including the brain (Liu et al., 2004) and critical for miRNA-mediated repression of mRNAs (Czech and Hannon, 2011). Deficiency in Ago-2 results in death of mice during early embryogenesis or mid-gestation (Morita et al., 2007). This reflects not only the essential role of Ago2 in embryonic development but perhaps an effect of impaired microRNA generation (Morita et al., 2007). In contrast, studies in conditional mutants showed individual deficiencies in Ago 1, 3, and 4 does not produce obvious effects in mice, suggesting a redundancy among Ago family members (O'Carroll et al., 2007).
microRNA in Ischemia Preconditioning
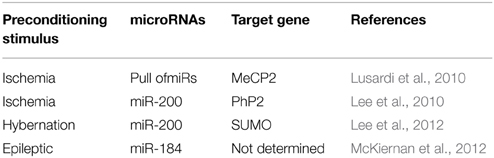
Three different publications show the regulation of microRNAs after an ischemic preconditioning stimulus in parallel (Table 1) (Dharap and Vemuganti, 2010; Lee et al., 2010; Lusardi et al., 2010).
Lusardi et al., using a brief ischemic stimulus in mice, showed that preconditioning at the same time up-regulated and down-regulated different microRNAs in the mouse cortex (Lusardi et al., 2010). A common target-gene of the down-regulated microRNAs was the transcriptional regulator MeCP2 (methyl CpG binding protein 2) (Lusardi et al., 2010). Similar results were found when hypertension was used as preconditioning stimulus in rats, where multiple down-regulated microRNAs target MeCP2 (Dharap and Vemuganti, 2010). MeCP2 was first described as a potent transcriptional repressor (Nan, 1998). Mutations in MeCP2 cause Rett syndrome, mental retardation, Angelman syndrome, and autism (Hite et al., 2009; Gonzales and LaSalle, 2010). The up-regulation of MeCP2 via microRNAs corroborated the general gene-repression in tolerance reported before (Lusardi et al., 2010), as increase levels of MeCP2 resulted in a down-regulation of the transcriptome.
In contrast to these studies, Lee et al. showed that neuroprotection achieved by brief ischemic preconditioning stimuli in mice could be due to a family member of miR-200 which targets PHD2 (prolyl hydroxylate 2), which in turn has been involved in the regulation of HIF1α after ischemia (Lee et al., 2010). Up-regulation of miR-200 shortly after the preconditioning stimulus reduced the levels of PHD2 and increased HIF1α. These results were corroborated in vitro, where the up-regulation of miR-200 using mimic decreased levels of PHD2 (Lee et al., 2010). Supporting these results, transgenic mice lacking HIF1α exposed to hypoxic preconditioning presented less neuroprotection than control mice after an hypoxia-ischemic insult (Sheldon et al., 2014).
More recently, a model of hibernation, a natural model of tolerance to ischemia, was used to study the role of microRNAs during neuroprotection in the brain (Lee et al., 2012). During torpor stages, hibernating animals lower their energy consumption, blood flow, and body temperature to an otherwise lethal level which is similar to levels during an ischemic insult (Lee et al., 2012). Microarray analysis from ground squirrel brains were performed during the active and the torpor phase and 405 microRNAs were different between both stages, with the family of miR-200 being the most representative, consistent with Lee et al. (2010). In this study the authors suggest that miR-200 family could regulate members of the SUMO family which is involved in the ubiquitin regulation and protein degradation, thereby explaining the reduction in protein levels during tolerance (Lee et al., 2012).
microRNA in Epileptic Preconditioning
Similar to ischemia, several studies analyzed the role of microRNAs after a preconditioning-seizure stimuli (Table 1 and Hatazaki et al., 2007; Kretschmann et al., 2015).
McKiernan and colleagues by using low dose of systemic KA in mice showed a de-regulation of microRNA after epileptic preconditioning. Here, a general up-regulation of microRNAs was the main response in preconditioned mice compared to control mice (McKiernan et al., 2012). Whereas 9 microRNAs were uniquely present in the preconditioning group, 39 microRNAs showed increase and 6 reduced levels in the pre-conditioning group. From the up-regulated microRNAs, 25 of the 39 were significantly regulated (McKiernan et al., 2012). Between the up-regulated microRNA group, the highest levels were found for miR-184. Inhibition of miR-184 in vivo resulted in neuronal death after a normally non-damaging preconditional stimulus (McKiernan et al., 2012) and an increase in neurodegeneration during the tolerant state after status epilepticus (damaging injury stimulation). Together, these results show a neuroprotective role of miR-184 during preconditioning (McKiernan et al., 2012).
Similar results have been found in acute seizure models not associated with neuronal death (Kretschmann et al., 2015). Kretschman and colleagues showed that 6 h after the induction of a single seizure via electrical stimulation up-regulation of microRNAs was the main response. 146 microRNAs were de-regulated, with almost 140 microRNAs being up-regulated and only few of them down-regulated (Kretschmann et al., 2015).
In a more detailed study, the expression levels of microRNAs were analyzed after electroshock stimulation, showing that microRNA levels follow three distinct patterns after non-damage neuronal activity (Eacker et al., 2011). MicroRNA regulation can be clustered in three distinct groups: Class 1, microRNAs which progressively increase their expression levels after electrical stimulation, including miR-134; Class 2, microRNAs which are strongly up-regulated in the first hours after stimulation with levels decreasing back to normal rapidly; Class 3, microRNAs that increase initially but decrease at a later time-point (12 or 24 h after the stimulation) (Eacker et al., 2011). This study shows the specific response of microRNAs after brain stimulation, and its possible role in the different tolerant-stages.
Targeting microRNAs on Brain
Several approaches for manipulating miRNA levels in neuronal cells in vitro and in vivo have been described, including recent applications of miRNA antisense oligonucleotides, miRNA gene knockout and miRNA sponges in neuronal cells. The efficiency of microRNAs inhibitors has been reported several times, inhibition of miR184 show a loss in the preconditioning neuroprotection without any toxicity (McKiernan et al., 2012). In contrast, no many approaches have shown over-expression of microRNAs in the brain, possible related with the toxic effect increase levels of RNA in the brain. The study of new drugs which target microRNAs in the central nervous system will be one of the main fields of future investigations.
Conclusions
Studying the molecular mechanism underlying the innate neuroprotection can help to understand how the brain protects itself against a damaging insult. Analysis of tolerant states has shown that a general down-regulation of genes is the main response in the brain, independently of the original insult, ischemic, or seizures. MicroRNAs are master regulators of gene-expression, its regulation after the preconditioning stimulus could be causative of the general suppression of gene-expression and being a good candidate for future target-therapy. Still several questions are remaining: Can we regulate microRNAs in the brain? Systemic administration of microRNA mimics or microRNA inhibitors will be necessary. Can we regulate microRNA expression in a time dependent manner? Tolerant is a time-dependent effect which only last for several days, a longer inhibition or over-expression of microRNAs could be toxic for the brain.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The author would like to thank Dr. Tobias Engel and Prof. David C. Henshall for critical reading of the manuscript. This work was supported by grants from the Science Foundation Ireland (13/SIRG/2114) to EJ.
References
Amini, E., Rezaei, M., Mohamed Ibrahim, N., Golpich, M., Ghasemi, R., Mohamed, Z., et al. (2014). A molecular approach to epilepsy management: from current therapeutic methods to preconditioning efforts. Mol. Neurobiol. doi: 10.1007/s12035-014-8876-5. [Epub ahead of print].
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Assaf, F., Fishbein, M., Gafni, M., Keren, O., and Sarne, Y. (2011). Pre- and post-conditioning treatment with an ultra-low dose of Delta9-tetrahydrocannabinol (THC) protects against pentylenetetrazole (PTZ)-induced cognitive damage. Behav. Brain Res. 220, 194–201. doi: 10.1016/j.bbr.2011.02.005
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Babiarz, J. E., Hsu, R., Melton, C., Thomas, M., Ullian, E. M., and Blelloch, R. (2011). A role for noncanonical microRNAs in the mammalian brain revealed by phenotypic differences in Dgcr8 versus Dicer1 knockouts and small RNA sequencing. RNA 17, 1489–1501. doi: 10.1261/rna.2442211
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Barone, F. C., White, R. F., Spera, P. A., Ellison, J., Currie, R. W., Wang, X., et al. (1998). Ischemic preconditioning and brain tolerance: temporal histological and functional outcomes, protein synthesis requirement, and interleukin-1 receptor antagonist and early gene expression. Stroke 29, 1937–1950. discussion: 1950–1931. doi: 10.1161/01.STR.29.9.1937
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Baskerville, S., and Bartel, D. P. (2005). Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA 11, 241–247. doi: 10.1261/rna.7240905
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bicker, S., Khudayberdiev, S., Weiß, K., Zocher, K., Baumeister, S., and Schratt, G. (2013). The DEAH-box helicase DHX36 mediates dendritic localization of the neuronal precursor-microRNA-134. Genes Dev. 27, 991–996. doi: 10.1101/gad.211243.112
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bohnsack, M. T., Czaplinski, K., and Gorlich, D. (2004). Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 10, 185–191. doi: 10.1261/rna.5167604
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Borges, K., Shaw, R., and Dingledine, R. (2007). Gene expression changes after seizure preconditioning in the three major hippocampal cell layers. Neurobiol. Dis. 26, 66–77. doi: 10.1016/j.nbd.2006.12.001
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Burroughs, A. M., Ando, Y., de Hoon, M. J., Tomaru, Y., Suzuki, H., Hayashizaki, Y., et al. (2011). Deep-sequencing of human Argonaute-associated small RNAs provides insight into miRNA sorting and reveals Argonaute association with RNA fragments of diverse origin. RNA Biol. 8, 158–177. doi: 10.4161/rna.8.1.14300
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Cai, X., Hagedorn, C. H., and Cullen, B. R. (2004). Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 10, 1957–1966. doi: 10.1126/science.1102513
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Chen, J., and Simon, R. (1997). Ischemic tolerance in the brain. Neurology 48, 306–311. doi: 10.1212/WNL.48.2.306
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Czech, B., and Hannon, G. J. (2011). Small RNA sorting: matchmaking for Argonautes. Nat. Rev. Genet. 12, 19–31. doi: 10.1038/nrg2916
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Davis, T. H., Cuellar, T. L., Koch, S. M., Barker, A. J., Harfe, B. D., McManus, M. T., et al. (2008). Conditional loss of Dicer disrupts cellular and tissue morphogenesis in the cortex and hippocampus. J. Neurosci. 28, 4322–4330. doi: 10.1523/JNEUROSCI.4815-07.2008
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Della-Morte, D., Guadagni, F., Palmirotta, R., Ferroni, P., Testa, G., Cacciatore, F., et al. (2012). Genetics and genomics of ischemic tolerance: focus on cardiac and cerebral ischemic preconditioning. Pharmacogenomics 13, 1741–1757. doi: 10.2217/pgs.12.157
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
De Pietri Tonelli, D., Pulvers, J. N., Haffner, C., Murchison, E. P., Hannon, G. J., and Huttner, W. B. (2008). miRNAs are essential for survival and differentiation of newborn neurons but not for expansion of neural progenitors during early neurogenesis in the mouse embryonic neocortex. Development 135, 3911–3921. doi: 10.1242/dev.025080
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Dharap, A., and Vemuganti, R. (2010). Ischemic pre-conditioning alters cerebral microRNAs that are upstream to neuroprotective signaling pathways. J. Neurochem. 113, 1685–1691. doi: 10.1111/j.1471-4159.2010.06735.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Dirnagl, U., Becker, K., and Meisel, A. (2009). Preconditioning and tolerance against cerebral ischaemia: from experimental strategies to clinical use. Lancet Neurol. 8, 398–412. doi: 10.1016/S1474-4422(09)70054-7
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Dorval, V., Smith, P. Y., Delay, C., Calvo, E., Planel, E., Zommer, N., et al. (2012). Gene network and pathway analysis of mice with conditional ablation of Dicer in post-mitotic neurons. PLoS ONE 7:e44060. doi: 10.1371/journal.pone.0044060
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Eacker, S. M., Keuss, M. J., Berezikov, E., Dawson, V. L., and Dawson, T. M. (2011). Neuronal activity regulates hippocampal miRNA expression. PLoS ONE 6:e25068. doi: 10.1371/journal.pone.0025068
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gonzales, M. L., and LaSalle, J. M. (2010). The role of MeCP2 in brain development and neurodevelopmental disorders. Curr. Psychiatry Rep. 12, 127–134. doi: 10.1007/s11920-010-0097-7
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hatazaki, S., Bellver-Estelles, C., Jimenez-Mateos, E. M., Meller, R., Bonner, C., Murphy, N., et al. (2007). Microarray profile of seizure damage-refractory hippocampal CA3 in a mouse model of epileptic preconditioning. Neuroscience 150, 467–477. doi: 10.1016/j.neuroscience.2007.09.020
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hite, K. C., Adams, V. H., and Hansen, J. C. (2009). Recent advances in MeCP2 structure and function. Biochem. Cell Biol. 87, 219–227. doi: 10.1139/O08-115
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hsu, R., Schofield, C. M., Dela Cruz, C. G., Jones-Davis, D. M., Blelloch, R., and Ullian, E. M. (2012). Loss of microRNAs in pyramidal neurons leads to specific changes in inhibitory synaptic transmission in the prefrontal cortex. Mol. Cell. Neurosci. 50, 283–292. doi: 10.1016/j.mcn.2012.06.002
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jimenez-Mateos, E. M., Hatazaki, S., Johnson, M. B., Bellver-Estelles, C., Mouri, G., Bonner, C., et al. (2008). Hippocampal transcriptome after status epilepticus in mice rendered seizure damage-tolerant by epileptic preconditioning features suppressed calcium and neuronal excitability pathways. Neurobiol. Dis. 32, 442–453. doi: 10.1016/j.nbd.2008.08.008
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kitagawa, K., Matsumoto, M., Kuwabara, K., Tagaya, M., Ohtsuki, T., Hata, R., et al. (1990). “Ischemic tolerance” phenomenon found in the brain. Brain Res. 528, 21–24. doi: 10.1016/0006-8993(90)90189-I
Klein, M. E., Lioy, D. T., Ma, L., Impey, S., Mandel, G., and Goodman, R. H. (2007). Homeostatic regulation of MeCP2 expression by a CREB-induced microRNA. Nat. Neurosci. 10, 1513–1514. doi: 10.1038/nn2010
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Knuckles, P., Vogt, M. A., Lugert, S., Milo, M., Chong, M. M., Hautbergue, G. M., et al. (2012). Drosha regulates neurogenesis by controlling neurogenin 2 expression independent of microRNAs. Nat. Neurosci. 15, 962–969. doi: 10.1038/nn.3139
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kobayashi, S., Harris, V. A., and Welsh, F. A. (1995). Spreading depression induces tolerance of cortical neurons to ischemia in rat brain. J. Cereb. Blood Flow Metab. 15, 721–727. doi: 10.1038/jcbfm.1995.92
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Konopka, W., Kiryk, A., Novak, M., Herwerth, M., Parkitna, J. R., Wawrzyniak, M., et al. (2010). MicroRNA loss enhances learning and memory in mice. J. Neurosci. 30, 14835–14842. doi: 10.1523/JNEUROSCI.3030-10.2010
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kretschmann, A., Danis, B., Andonovic, L., Abnaof, K., van Rikxoort, M., Siegel, F., et al. (2015). Different microRNA profiles in chronic epilepsy versus acute seizure mouse models. J. Mol. Neurosci. 55, 466–479. doi: 10.1007/s12031-014-0368-6
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Kwak, P. B., and Tomari, Y. (2012). The N domain of Argonaute drives duplex unwinding during RISC assembly. Nat. Struct. Mol. Biol. 19, 145–151. doi: 10.1038/nsmb.2232
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lagos-Quintana, M., Rauhut, R., Lendeckel, W., and Tuschl, T. (2001). Identification of novel genes coding for small expressed RNAs. Science 294, 853–858. doi: 10.1126/science.1064921
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lugli, G. I., Larson, J., Martone, M. E., Jones, Y., and Smalheiser, N. R. (2005). Dicer and eIF2c are enriched at postsynaptic densities in adult mouse brain and are modified by neuronal activity in a calpain-dependent manner. J. Neurochem. 94, 896–905. doi: 10.1111/j.1471-4159.2005.03224.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lau, N. C., Lim, L. P., Weinstein, E. G., and Bartel, D. P. (2001). An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294, 858–862. doi: 10.1126/science.1065062
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lee, R. C., and Ambros, V. (2001). An extensive class of small RNAs in Caenorhabditis elegans. Science 294, 862–864. doi: 10.1126/science.1065329
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lee, S. T., Chu, K., Jung, K. H., Yoon, H. J., Jeon, D., Kang, K. M., et al. (2010). MicroRNAs induced during ischemic preconditioning. Stroke 41, 1646–1651. doi: 10.1161/STROKEAHA.110.579649
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lee, Y., Ahn, C., Han, J., Choi, H., Kim, J., Yim, J., et al. (2003). The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419. doi: 10.1038/nature01957
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lee, Y. J., Johnson, K. R., and Hallenbeck, J. M. (2012). Global protein conjugation by ubiquitin-like-modifiers during ischemic stress is regulated by microRNAs and confers robust tolerance to ischemia. PLoS ONE 7:e47787. doi: 10.1371/journal.pone.0047787
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lee, Y., Kim, M., Han, J., Yeom, K. H., Lee, S., Baek, S. H., et al. (2004). MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 23, 4051–4060. doi: 10.1038/sj.emboj.7600385
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lin, J. H., Lou, N., Kang, N., Takano, T., Hu, F., Han, X., et al. (2008). A central role of connexin 43 in hypoxic preconditioning. J. Neurosci. 28, 681–695. doi: 10.1523/JNEUROSCI.3827-07.2008
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Liu, J., Carmell, M. A., Rivas, F. V., Marsden, C. G., Thomson, J. M., Song, J. J., et al. (2004). Argonaute2 is the catalytic engine of mammalian RNAi. Science 305, 1437–1441. doi: 10.1126/science.1102513
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lugli, G., Torvik, V. I., Larson, J., and Smalheiser, N. R. (2008). Expression of microRNAs and their precursors in synaptic fractions of adult mouse forebrain. J. Neurochem. 106, 650–661. doi: 10.1111/j.1471-4159.2008.05413.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lusardi, T. A., Farr, C. D., Faulkner, C. L., Pignataro, G., Yang, T., Lan, J., et al. (2010). Ischemic preconditioning regulates expression of microRNAs and a predicted target, MeCP2, in mouse cortex. J. Cereb. Blood Flow Metab. 30, 744–756. doi: 10.1038/jcbfm.2009.253
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
McCafferty, K., Forbes, S., Thiemermann, C., and Yaqoob, M. M. (2014). The challenge of translating ischemic conditioning from animal models to humans: the role of comorbidities. Dis. Model. Mech. 7, 1321–1333. doi: 10.1242/dmm.016741
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
McKiernan, R. C., Jimenez-Mateos, E. M., Sano, T., Bray, I., Stallings, R. L., Simon, R. P., et al. (2012). Expression profiling the microRNA response to epileptic preconditioning identifies miR-184 as a modulator of seizure-induced neuronal death. Exp. Neurol. 237, 346–354. doi: 10.1016/j.expneurol.2012.06.029
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Meller, R., Thompson, S. J., Lusardi, T. A., Ordonez, A. N., Ashley, M. D., Jessick, V., et al. (2008). Ubiquitin proteasome-mediated synaptic reorganization: a novel mechanism underlying rapid ischemic tolerance. J. Neurosci. 28, 50–59. doi: 10.1523/JNEUROSCI.3474-07.2008
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Mirrione, M. M., Konomos, D. K., Gravanis, I., Dewey, S. L., Aguzzi, A., Heppner, F. L., et al. (2010). Microglial ablation and lipopolysaccharide preconditioning affects pilocarpine-induced seizures in mice. Neurobiol. Dis. 39, 85–97. doi: 10.1016/j.nbd.2010.04.001
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Morita, S., Horii, T., Kimura, M., Goto, Y., Ochiya, T., and Hatada, I. (2007). One Argonaute family member, Eif2c2 (Ago2), is essential for development and appears not to be involved in DNA methylation. Genomics 89, 687–696. doi: 10.1016/j.ygeno.2007.01.004
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Murry, C. E., Jennings, R. B., and Reimer, K. A. (1986). Preconditioning with ischemia: a delay of lethal cell injury in ischemic myocardium. Circulation 74, 1124–1136. doi: 10.1161/01.CIR.74.5.1124
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Nan, X. (1998). Gene silencing by methyl-CpG-binding proteins. Novartis Found. Symp. 214, 6–16. discussion: 16–21, 46–50.
O'Carroll, D., Mecklenbrauker, I., Das, P. P., Santana, A., Koenig, U., Enright, A. J., et al. (2007). A Slicer-independent role for Argonaute 2 in hematopoiesis and the microRNA pathway. Genes Dev. 21, 1999–2004. doi: 10.1101/gad.1565607
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
O'Carroll, D., and Schaefer, A. (2013). General principals of miRNA biogenesis and regulation in the brain. Neuropsychopharmacology 38, 39–54. doi: 10.1038/npp.2012.87
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Plamondon, H., Blondeau, N., Heurteaux, C., and Lazdunski, M. (1999). Mutually protective actions of kainic acid epileptic preconditioning and sublethal global ischemia on hippocampal neuronal death: involvement of adenosine A1 receptors and K(ATP) channels. J. Cereb. Blood Flow Metab. 19, 1296–1308. doi: 10.1097/00004647-199912000-00002
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sheldon, R. A., Lee, C. L., Jiang, X., Knox, R. N., and Ferriero, D. M. (2014). Hypoxic preconditioning protection is eliminated in HIF-1alpha knockout mice subjected to neonatal hypoxia-ischemia. Pediatr. Res. 76, 46–53. doi: 10.1038/pr.2014.53
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Siegel, G., Saba, R., and Schratt, G. (2011). microRNAs in neurons: manifold regulatory roles at the synapse. Curr. Opin. Genet. Dev. 21, 491–497. doi: 10.1016/j.gde.2011.04.008
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Simon, R. P., Niiro, M., and Gwinn, R. (1993). Prior ischemic stress protects against experimental stroke. Neurosci. Lett. 163, 135–137. doi: 10.1016/0304-3940(93)90364-Q
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Stenzel-Poore, M. P., Stevens, S. L., King, J. S., and Simon, R. P. (2007). Preconditioning reprograms the response to ischemic injury and primes the emergence of unique endogenous neuroprotective phenotypes: a speculative synthesis. Stroke 38, 680–685. doi: 10.1161/01.STR.0000251444.56487.4c
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Stenzel-Poore, M. P., Stevens, S. L., and Simon, R. P. (2004). Genomics of preconditioning. Stroke 35(11 Suppl. 1), 2683–2686. doi: 10.1161/01.STR.0000143735.89281.bb
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Tao, J., Wu, H., Lin, Q., Wei, W., Lu, X. H., Cantle, J. P., et al. (2011). Deletion of astroglial Dicer causes non-cell-autonomous neuronal dysfunction and degeneration. J. Neurosci. 31, 8306–8319. doi: 10.1523/JNEUROSCI.0567-11.2011
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Towfighi, J., Housman, C., Mauger, D., and Vannucci, R. C. (1999). Effect of seizures on cerebral hypoxic-ischemic lesions in immature rats. Brain Res. Dev. Brain Res. 113, 83–95. doi: 10.1016/S0165-3806(99)00004-8
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Keywords: preconditioning, microRNAs, ischemia, epilepsy, neuroprotection
Citation: Jimenez-Mateos EM (2015) Role of MicroRNAs in innate neuroprotection mechanisms due to preconditioning of the brain. Front. Neurosci. 9:118. doi: 10.3389/fnins.2015.00118
Received: 31 December 2014; Accepted: 23 March 2015;
Published: 21 April 2015.
Edited by:
Giuseppe Pignataro, Federico II University of Naples, ItalyReviewed by:
Varun Kesherwani, University of Nebraska Medical Center, USAAntonio Vinciguerra, Federico II University of Naples, Italy
Copyright © 2015 Jimenez-Mateos. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Eva M. Jimenez-Mateos, Department of Physiology and Medical Physics, Royal College of Surgeons in Ireland, 123 St Stephen's Green, Dublin 2, IrelandZXZhamltZW5lekByY3NpLmll
 Eva M. Jimenez-Mateos
Eva M. Jimenez-Mateos