- 1Department of Neuroscience II, Research Institute of Environmental Medicine, Nagoya University, Nagoya, Japan
- 2Department of Neural Regulation, Nagoya University Graduate School of Medicine, Nagoya, Japan
- 3Research and Education Center for Brain Science, Hokkaido University Graduate School of Medicine, Sapporo, Japan
In mammals, the central circadian clock is located in the suprachiasmatic nucleus (SCN) of the hypothalamus. Individual SCN cells exhibit intrinsic oscillations, and their circadian period and robustness are different cell by cell in the absence of cellular coupling, indicating that cellular coupling is important for coherent circadian rhythms in the SCN. Several neuropeptides such as arginine vasopressin (AVP) and vasoactive intestinal polypeptide (VIP) are expressed in the SCN, where these neuropeptides function as synchronizers and are important for entrainment to environmental light and for determining the circadian period. These neuropeptides are also related to developmental changes of the circadian system of the SCN. Transcription factors are required for the formation of neuropeptide-related neuronal networks. Although VIP is critical for synchrony of circadian rhythms in the neonatal SCN, it is not required for synchrony in the embryonic SCN. During postnatal development, the clock genes cryptochrome (Cry)1 and Cry2 are involved in the maturation of cellular networks, and AVP is involved in SCN networks. This mini-review focuses on the functional roles of neuropeptides in the SCN based on recent findings in the literature.
Introduction
Creatures on earth anticipate cyclic changes in the surrounding environment, such as day–night, seasonal, and annual changes, to adapt their own physiological functions so that they can minimize risks to survival. Among them, rhythmic changes adapting to environmental changes caused by the rotation of the earth are driven by intrinsic oscillatory mechanisms called the circadian clock. In mammals, the circadian system comprises a hierarchical structure involving the master clock in the brain and a number of peripheral clocks situated throughout the body. Almost all cells in our body possess a circadian clock in which the molecular machinery, namely, transcription–translation feedback loops involving several clock genes and their protein products, generates circadian rhythmicity (Reppert and Weaver, 2002). Circadian rhythms of cells and tissues are coordinated by the master circadian clock located in the suprachiasmatic nucleus (SCN) of the hypothalamus (Weaver, 1998). The SCN is the only clock that can entrain to the environmental light–dark (LD) time cue from the retina via the retinohypothalamic tract (RHT) (LeGates et al., 2014). Ultimately, the circadian clock in the SCN regulates rhythms within a variety of physiological output functions such as sleep/wakefulness, body temperature, and endocrine functions.
The SCN contains approximately 20,000 neurons that show circadian rhythms in clock gene expression, cytosolic Ca2+, and spontaneous firing, and the rhythms are also maintained in culture conditions, in the slice and in dispersed cells (Welsh et al., 1995; Ikeda et al., 2003; Yamaguchi et al., 2003). The distribution of the circadian period from dispersed SCN cells is larger than that from slice cultures (Herzog et al., 2004; Honma et al., 2004). Furthermore, the robustness of circadian rhythmicity in dispersed cells is weaker than that in slices (Liu et al., 2007; Ono et al., 2013). These results suggest that cellular networks are crucial for coherent circadian rhythms in the SCN.
Heterogeneous Cell Types in the Suprachiasmatic Nucleus
The SCN is, neuroanatomically and cytochemically, divided into two subregions—dorsal and ventral subregions (Figure 1). The ventral SCN receives major afferent pathways, including those from the retina, midbrain raphe nucleus, and intergeniculate leaflet (Hattar et al., 2006; McNeill et al., 2011). It contains vasoactive intestinal peptide (VIP), gastrin-releasing peptide (GRP), neurotensin (NT), and calretinin (CALR)-expressing neurons (Abrahamson and Moore, 2001). Arginine vasopressin (AVP), met-enkephalin (mENK), and angiotensin II (AII)-expressing neurons are located in the dorsal SCN.
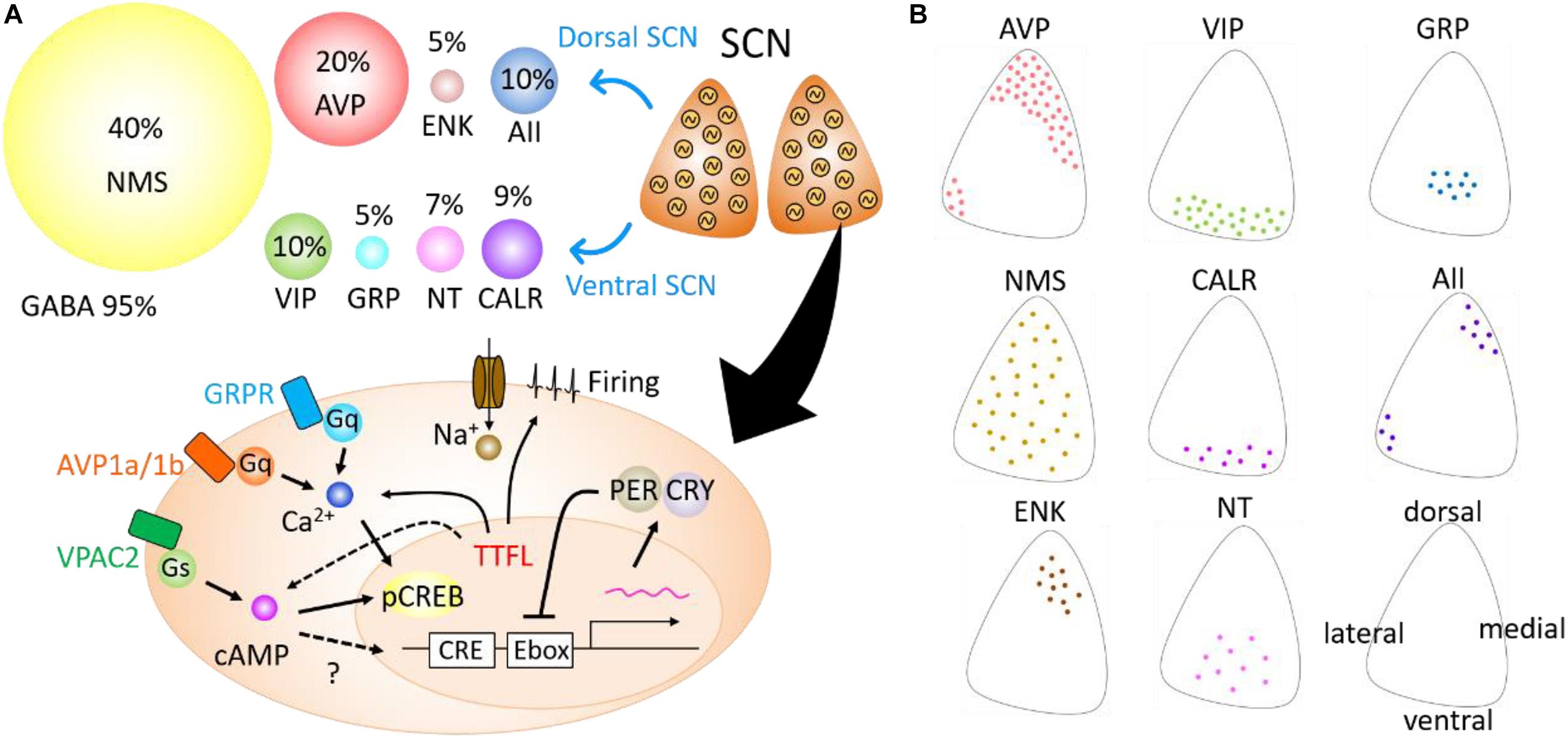
Figure 1. Neuropeptides expressed in the suprachiasmatic nucleus (SCN) and intracellular signaling. (A) A variety of neuropeptides expressed in the SCN (top). Cells with an autonomous circadian oscillator are schematically shown in the SCN in coronal plane. Circles of different sizes indicate the percentage of neurons expressed in the SCN. Arginine vasopressin (AVP), enkephalin (ENK), and angiotensin II (AII) are expressed mainly in the dorsal area of the SCN, whereas vasoactive intestinal polypeptide (VIP), gastrin-releasing peptide (GRP), neurotensin (NT), and calretinin (CALR) are mainly expressed in the ventral SCN. Neuromedin-S (NMS) is broadly expressed in the SCN, and gamma aminobutyric acid (GABA) in almost all SCN neurons. Schematic view of the intracellular pathways mediating neuropeptide signals in the SCN (bottom). Neuropeptides bind G-protein-coupled receptors (Gq, Gs) and modulate second messenger signaling such as cAMP or Ca2+. These signals facilitate the phosphorylation of CREB proteins and change the transcription of Per genes. Some signaling from transcription–translation feedback loop (TTFL) could modulate cAMP or Ca2+ rhythms. (B) Schematic drawing demonstrating the distribution of eight types of peptidergic neurons within the unilateral coronal SCN slice. Directions are shown in the lower right SCN.
Taking advantage of bioluminescence reporters, circadian rhythms can be recorded in individual SCN cells. In dispersed cell culture, individual SCN cells show autonomous circadian rhythms, but not all cells maintain rhythmicity. Webb et al. (2009) measured PER2:LUC bioluminescence from dispersed SCN cells and performed post hoc immunostaining to identify which neurons (i.e., those expressing AVP or VIP) are responsible for rhythm generation. They found that the circadian period and percentage of rhythmic cells did not differ between AVP and VIP neurons and concluded that intrinsic circadian oscillation was not restricted to a single class of neuropeptidergic neurons in the SCN. They also treated SCN slices with tetrodotoxin (TTX) for 6 days twice at 6-day intervals and examined PER2:LUC rhythmicity. Since TTX suppresses neural communication, neurons exhibiting circadian rhythms via neural inputs no longer exhibit circadian rhythms following TTX treatment, while neurons with a cell-autonomous circadian clock continue to show circadian PER2:rhythms even under TTX treatment. If circadian peacemaking neurons are unique to certain cell types, repeated TTX treatments would reveal circadian rhythms in the same cells. However, the results showed that the cells exhibiting circadian rhythms under TTX treatment were not the same between the two trials. Furthermore, the location of neurons that maintained circadian rhythms in both TTX treatments was not regionally specific; rather, these neurons were scattered throughout the SCN. These results suggest that neurons throughout the SCN are intrinsic but unstable circadian oscillators that are capable of showing robust oscillation by neural networks. A recent study by Hirata et al. (2019) demonstrated that by culturing dispersed SCN cells on microfabricated islands, a substantial number of solitary SCN neurons with no contact with other cells exhibited robust circadian rhythms with regard to clock gene expression and intracellular Ca2+ levels. Since TTX treatment suppresses action potential, neuronal firing may itself affect the circadian molecular rhythms in solitary SCN neurons.
Roles of Vasoactive Intestinal Polypeptide Neurons in the Suprachiasmatic Nucleus
VIP is known as a synchronizer of neuronal networks in the SCN, and approximately 10% of the SCN consists of VIP-positive neurons (Abrahamson and Moore, 2001). Mice lacking VIP or VIP receptor 2 (VPAC2) show desynchronized circadian rhythms in SCN neurons in culture as well as attenuated behavioral rhythms (Harmar et al., 2002; Colwell et al., 2003; Aton et al., 2005; Maywood et al., 2006). Similarly, the application of an excessive amount of VIP onto cultured SCN slices also attenuates the synchrony of cellular circadian rhythms (An et al., 2013). The rhythmic release of VIP in cultured SCN slices has been previously reported (Shinohara et al., 1995; Shinohara et al., 1999). These results suggest that rhythmic release of VIP is important for synchrony of circadian rhythms in SCN neurons. Chemogenetic activation of VIP neurons also changes the spatiotemporal patterns of PER2:LUC rhythms, with lengthening of the circadian period and attenuated amplitude of circadian rhythms (Brancaccio et al., 2013). Maywood et al. (2011) demonstrated that paracrine signaling in the SCN is critical for the synchronization of circadian rhythms. The authors cultured VIP or VPAC2 knockout (KO) SCN containing a PER2:LUC reporter with a wild-type (WT) graft SCN, and they placed a semipermeable membrane that only passed small molecules, such as neuropeptides, between two SCN slices. Importantly, the circadian rhythms of VIP KO SCN were restored by coculture of WT SCN, indicating that small diffusible molecules such as neuropeptides from the WT SCN are critical for rhythm synchronization. However, it took longer to restore the circadian rhythms of VPAC2 KO SCN as compared to VIP KO SCN. These results indicate that VIP and other molecules, such as AVP and GRP, are crucial for the synchronization of circadian rhythms in the SCN.
VIP and gamma aminobutyric acid (GABA) signaling work synergistically to sustain circadian phase differences in the SCN. For example, a long-day photoperiod shortens the activity time and changes cellular coupling in the SCN (Inagaki et al., 2007). Interestingly, under a long-day photoperiod, the circadian phase of PER2:LUC rhythms between the dorsal and ventral SCN is decoupled at the beginning of culture, which gradually resynchronizes under culture conditions (Evans et al., 2013; Myung et al., 2015). This resynchronization is blocked by a VIP or GABAA receptor antagonist. VIP receptor antagonists attenuate both the advance and delay portions of the coupling response curve (i.e., the resetting responses of SCN core neurons as compared to the initial phase relationship with SCN shell neurons). However, in this study, the GABAA receptor antagonist mainly attenuated the advance portion of the curve. Furthermore, the effects of GABA receptor antagonists are not evident for circadian rhythms in the cultured WT SCN (Aton et al., 2006; Ono et al., 2019), though GABA dramatically restores the synchrony of circadian rhythms in the VIP KO cultured SCN (Freeman et al., 2013).
Application of VIP into the culture medium induces a phase-dependent phase delay shift in the circadian PER2:LUC rhythm of the SCN slice (An et al., 2011). The VIP-mediated phase shift could be due to cAMP signaling because VIP application increases intracellular cAMP and activity of the cAMP response element (CRE) using cAMP FRET sensor and CRE-luc reporter, respectively (An et al., 2011; Hamnett et al., 2019). In addition, extracellular signal-regulated kinases (ERKs) 1/2 and dual specificity phosphatase (DUSP) 4 signaling may also be involved in the VIP-induced responses of circadian rhythms in the SCN, which is independent of the cAMP response element-binding protein (CREB) pathway (Hamnett et al., 2019). Similarly, optogenetic activation of VIP neurons causes a phase delay shift in circadian PER2:LUC rhythms in the SCN slice as well as behavioral rhythms (Mazuski et al., 2018; Patton et al., 2020). Intriguingly, Mazuski et al. (2018) found that VIP neurons showed tonic or irregular firing patterns and optogenetically mimicked irregular firing induced large phase shifts in both PER2:LUC rhythms in the SCN and behavioral rhythms. This result shows the physiological significance of irregular firing in SCN VIP neurons for the entrainment of circadian rhythms.
VIP neurons exhibit circadian rhythms during spontaneous firing in cultured SCN slices (Hermanstyne et al., 2016). Taking advantage of the in vivo fiber photometry method, Jones et al. (2018) demonstrated that VIP neurons exhibit circadian rhythms in spontaneous calcium activity in vivo under LD and constant dark (DD) conditions, but not under constant light (LL) conditions. In addition, calcium activity was evoked by light pulses with daily peaks occurring at approximately CT 12, which is consistent with light-induced circadian phase shifts. Chemogenetic suppression of neuronal activity of VIP neurons attenuates light-induced phase shifts. These results indicate that VIP neurons are important for light-mediated resetting of circadian rhythms.
VIP neurons may play an important role in the circadian output rhythms. Optogenetic activation of VIP neurons in the SCN suppresses paraventricular nucleus (PVN) neurons via GABA release, and chemogenetic suppression of these neurons in turn increases corticosterone concentration (Paul et al., 2020). This regulation could be mediated by corticotropin-releasing factor (CRF) neurons in the PVN because activation of SCN neurons suppresses CRF neuronal activity (Ono et al., 2020). The SCN to PVN neuronal pathways also regulate sleep and wakefulness (Ono et al., 2020). VIP neurons regulate nighttime sleep called “siesta” in mice, which is due to a specific group of VIP neurons that is active during nighttime in the SCN (Collins et al., 2020). Recently, single-cell RNA sequencing analysis revealed two types of VIP neurons in the SCN: pacemaker and non-pacemaker cells. Such techniques help categorize cell types in the SCN that regulate a variety of physiological functions (Todd et al., 2020; Wen et al., 2020).
Roles of Arginine Vasopressin Neurons in the Suprachiasmatic Nucleus
Approximately 20% of SCN consists of AVP-positive neurons, which is the second largest peptide population in the SCN (Abrahamson and Moore, 2001). The Avp promoter contains an E-box element, which is the target of CLOCK/BMAL1 transcription factors, and Avp mRNA levels exhibit a robust daily rhythm with daytime peak (Uhl and Reppert, 1986; Jin et al., 1999). Avp-Eluc reporter mice also show circadian Avp expression rhythms in SCN slice (Yoshikawa et al., 2015). Since the Avp promoter contains an E-box, Avp mRNA levels in the SCN are low throughout the day in Clock mutant mice (Jin et al., 1999). CRY1 and CRY2 are negative elements of E-box-related transcription. Thus, Avp expression is expected to increase without CRYs. However, Avp expression was suppressed and arrhythmic in the SCN of Cry1 and Cry2 double-deficient (Cry1/Cry2 KO) mice, suggesting that suppression of E-box-related transcription is not only due to Cry genes.
Although VIP is a strong synchronizer in the SCN, the AVP functions as a weak synchronizer. For example, the circadian rhythms of VPAC2 KO SCN are restored by the coculture of WT SCN, but this restoration is inhibited by the application of AVP receptor antagonists (Maywood et al., 2011). Edwards et al. (2016) demonstrated that the arrhythmicity of Cry1/Cry2 KO SCN was restored by AAV-mediated transduction of Cry1. Interestingly, treatment of Cry1/Cry2 KO SCN with AVP receptor antagonists only induced low-amplitude circadian rhythms when applied simultaneously with Cry1 gene transduction, but it did not attenuate restored rhythms when treated after circadian rhythms were restored by Cry1 transduction, suggesting that AVP signaling is required for the induction, but not maintenance, of CRY-dependent circadian rhythms in Cry1/Cry2 KO SCN (Edwards et al., 2016). Loss of the AVP receptors V1a and V1b in the SCN can modify neuronal networks in the SCN. The clock gene, Per1, expression rhythm in the SCN is known to show a phase gradient with phase-leading at the dorsomedial SCN and -lagging at the ventrolateral area (Yamaguchi et al., 2003). This spatiotemporal pattern was perturbed by the application of cycloheximide (CHX), a translation inhibitor, and restored after washout in WT mice, but not in V1a and V1b KO mice (Yamaguchi et al., 2013). These mice show immediate re-entrainment to a new LD environment after an abrupt shift in LD cycles, suggesting that V1a and V1b confer SCN resistance to external perturbation.
The functional roles of the circadian clock of AVP neurons in the SCN were evaluated by genetically manipulating AVP neurons using the Cre-loxP system. AVP neuron-specific Bmal1 KO (Avp-Bmal1–/–) mice showed longer activity time with lengthening of the free-running period than WT mice (Mieda et al., 2015), but VIP neuron-specific Bmal1 KO did not affect circadian behavioral rhythms (Lee et al., 2015). PER2:LUC rhythms in the dorsal SCN of Avp-Bmal1–/– mice exhibited a lengthened circadian period and attenuated amplitude (Mieda et al., 2015) and failed to resynchronize circadian rhythms in the SCN after the washing out of TTX (Shan et al., 2020). Lee et al. (2015) demonstrated similar results in neuromedin-S (NMS) Cre mice. Overexpression of PER2 or deletion of Bmal1 in NMS neurons in the SCN abolished circadian wheel-running activity rhythms under constant darkness (DD). NMS KO, per se, did not change circadian behavioral rhythms, indicating that NMS itself has no role in the SCN circadian clock. Approximately 40% of the SCN consists of NMS-positive neurons, and VIP and AVP are coexpressed in NMS neurons. These results suggest that AVP neurons may modulate the coupling of the SCN network for morning and evening behavioral rhythms.
AVP neurons are also important for anticipatory thirst prior to sleep (Gizowski et al., 2016). Optogenetic activation of AVP neurons in the SCN increased water intake around ZT23 via AVP release in the organum vasculosum laminae terminalis (OVLT), an important brain area for water intake. In contrast, optogenetic suppression of neuronal activity in AVP neurons decreased water intake. These results suggest that the anticipatory thirst prior to sleep is driven by excitatory peptidergic neurotransmission mediated by AVP release in the SCN.
Neuropeptide-Related Neuronal Networks in the Suprachiasmatic Nucleus During Development
During development, dramatic changes in genetic, cellular, and circuit levels were observed in the SCN. Neurogenesis mainly occurs in the SCN around embryonic days 11–15 in mice (Shimada and Nakamura, 1973; Okamura et al., 1983; Kabrita and Davis, 2008; Shimogori et al., 2010), and synaptogenesis occurs rapidly around postnatal days 4–10 (Moore and Bernstein, 1989). Transcription factors such as Six3, Six6, Lhx1, and Fzd5 are expressed during the embryonic period in the SCN (Shimogori et al., 2010; VanDunk et al., 2011). AVP and VIP mRNA are observed from around embryonic days 17–18 (Romero and Silver, 1990; Isobe and Muramatsu, 1995; Ban et al., 1997; VanDunk et al., 2011). Interestingly, Six6-deficient mice show weak wheel-running rhythms and no clear AVP and VIP expression in the SCN (Clark et al., 2013). Six6-Cre-dependent Lhx1 deficiency (Six3-Cre;Lhx1loxp/loxp) attenuates neuropeptide expression in the SCN, including Avp, Vip, Grp, Prok2, Enk, and Nms (Bedont et al., 2014). In a study of Six3-Cre;Lhx1loxp/loxp mice, synchrony of circadian PER2:LUC rhythms in the SCN was suppressed and the circadian rhythmicity of wheel-running rhythms was attenuated; these results are phenotypically similar to Vip or VPAC2-deficient mice (Harmar et al., 2002; Colwell et al., 2003). Rorα-Cre;Lhx1loxP/loxP mice also exhibited similar circadian phenotypes (Hatori et al., 2014). These results suggest that transcription factors are required for the maturation of neuronal networks in the SCN.
Circadian rhythms in the SCN have been observed before birth. For example, day/night differences in 2-deoxyglucose uptake were observed at embryonic day 19 in vivo (Reppert and Schwartz, 1983). Per1 expression or PER2:LUC rhythms in the SCN are detectable around embryonic days 15–18 (Shimomura et al., 2001; Ansari et al., 2009; Wreschnig et al., 2014; Landgraf et al., 2015; Carmona-Alcocer et al., 2018). Notably, in the fetal SCN, synchronization of circadian PER2:LUC rhythms was not affected by the sodium channel inhibitor, TTX, VPAC2 receptor antagonist, GABAA receptor antagonist, or gap junction inhibitor, suggesting that VIP, GABA, gap junction, and neuronal firing are not required for synchronization of circadian rhythms in the fetal SCN (Carmona-Alcocer et al., 2018).
VIP neurons are suggested to be important for the maturation of neuronal networks in the SCN during development. Ablation of VIP neurons in the adult SCN using AAV-mediated expression of caspase3 induced shortening of circadian behavioral rhythms (Mazuski et al., 2020), but only 15% of mice became behaviorally arrhythmic. In contrast, 50% of the mice were arrhythmic when VIP or VPAC2 was deleted globally. In addition, ablation of VIP neurons in the neonatal SCN significantly dampens the circadian PER2:LUC rhythm (Mazuski et al., 2020). These results suggest that VIP neurons are important for synchrony in the SCN during early development but not in adulthood.
On the other hand, Todd et al. (2020) reported that even in adult mice, ablation of VIP neurons in the SCN by AAV-mediated diphtheria toxin A subunit (DTA) expression profoundly disrupted circadian locomotor activity recorded by a telemetry transmitter and running wheel. The difference in behavioral phenotypes in the two studies might be related to the number of VIP neurons spared from deletion and recording methods. The role of VIP neurons in circadian behavioral rhythms is still controversial and remains to be studied.
The importance of neuropeptides and clock gene, Cry, for coherent circadian rhythms in the SCN during postnatal development has also been reported. Cry1 and Cry2 are thought to be essential genes for circadian rhythms (Okamura et al., 1999; van der Horst et al., 1999; Albus et al., 2002; Yamaguchi et al., 2003). However, circadian Per1-luc, PER2:LUC, and spontaneous firing rhythms have been observed in single SCN cells in Cry1/Cry2-deficient mice (Maywood et al., 2011; Ono et al., 2013; Figure 2). Importantly, the cellular circadian rhythms of Cry1/Cry2-deficient SCN were synchronized during the neonatal period but became desynchronized around postnatal day 21, where circadian behavioral rhythms could be measured. Furthermore, synchronized circadian rhythms in neonatal Cry1/Cry2-deficient SCN were desynchronized without VIP signaling. Avp expression was also attenuated in Cry1/Cry2-deficient SCN in both neonatal and adult SCN (Ono et al., 2016; Figure 2B). These results indicate that VIP and AVP are responsible for CRY-independent and -dependent cellular coupling, respectively, and that these neuropeptides are differentially involved in the maturation of circadian networks in the SCN.
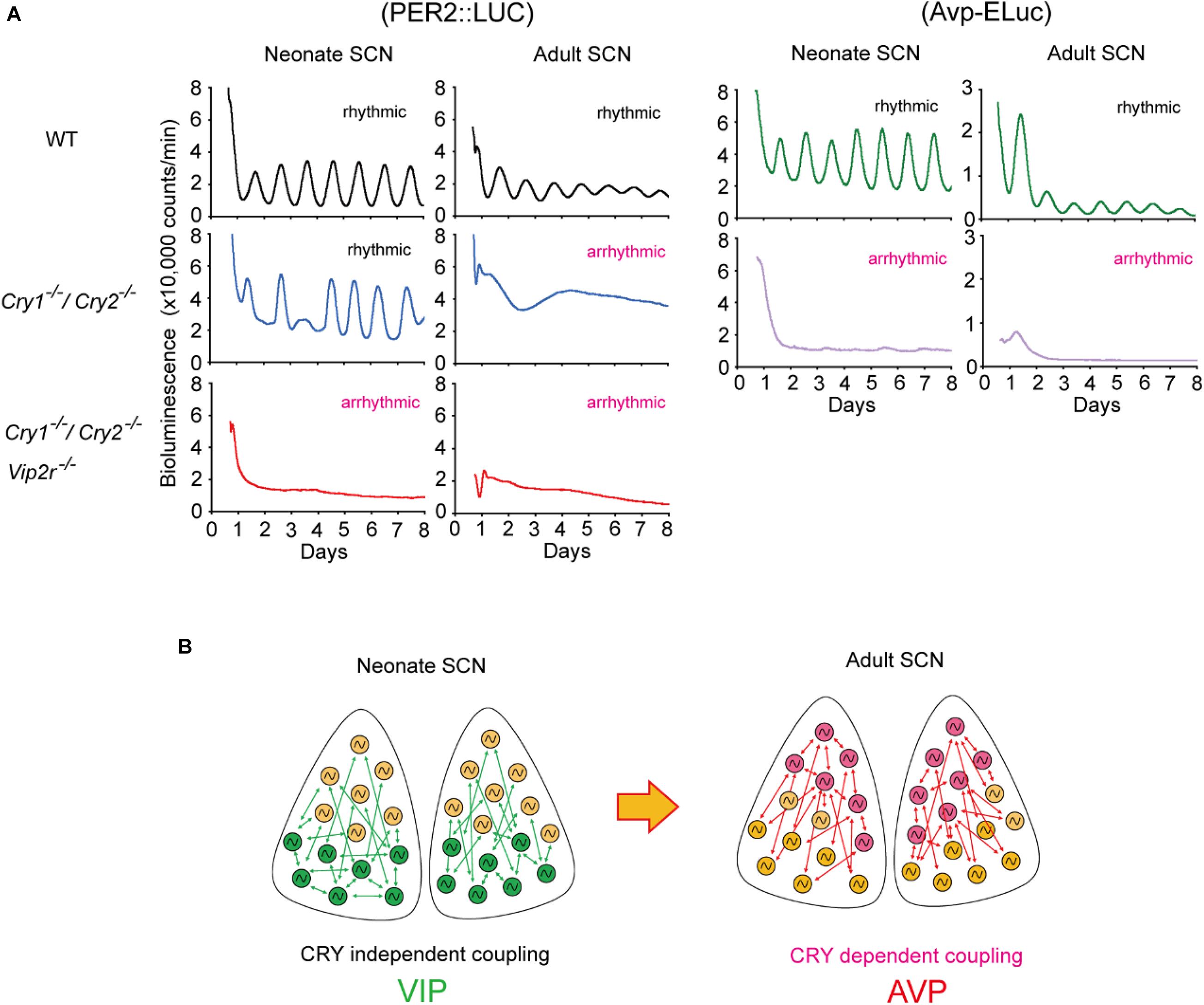
Figure 2. Developmental changes in cellular coupling mechanisms in the suprachiasmatic nucleus (SCN). (A) Tissue-level PER2:LUC bioluminescence from the cultured SCN in neonate or adult mice (left). Wild-type (WT) SCNs show circadian rhythms in both neonatal and adult SCNs. Cryptochrome (Cry)1– /–/Cry2– /– SCNs show robust circadian rhythms in the neonate SCN, but not in the adult SCN. Whereas Cry1– /–/Cry2– /–/vasoactive intestinal polypeptide (Vip)2r– /– SCN is arrhythmic both in the neonate and adult, which is due to desynchronization of cellular circadian rhythms. Arginine vasopressin (Avp)-ELuc bioluminescence from the cultured SCN in neonate or adult mice (right). Circadian Avp expression rhythms are observed in the WT SCN, but they are arrhythmic and expression is attenuated in Cry1– /–/Cry2– /– SCN both at neonatal and adult periods. (B) Schematic view of a model of cellular coupling in the SCN during postnatal development. CRY1 and CRY2 are involved in the developmental shift from CRY-independent (via VIP) to CRY-dependent (via AVP) networks in the SCN. Green and pink circles indicate VIP and AVP neurons, respectively. Green and pink arrows indicate signaling via VIP and AVP, respectively. This shift is observed around postnatal days 14–21. Modified from Ono et al., 2016.
Discussion
Since the first report demonstrating the importance of the SCN for circadian rhythms in behavior (Moore and Eichler, 1972; Stephan and Zucker, 1972), SCN circadian rhythms have been recorded in slice culture as well as in situ. Since culturing neonatal SCN is easier than culturing adult SCN, researchers have used neonatal SCN to explain phenotypes of behavior of mice lacking specific genes. However, recent studies have demonstrated that neuronal networks in the SCN are modulated during development. CRY1 and CRY2 are involved in the maturation of neuronal networks in the SCN during postnatal development. Although the functions of neuropeptides in the SCN cellular network have been investigated, their roles as output signals regulating peripheral circadian oscillators have not been clearly identified. Recent advances in various methodologies allow us to manipulate cell type- and developmental stage-specific gene expression, cellular signaling, and neuronal activity, which can help identify mechanisms for the developmental aspects of circadian rhythms in the SCN.
Author Contributions
DO and SH wrote the manuscript with support of KH. All authors contributed to the article and approved the submitted version.
Funding
This work was supported in part by the Uehara Memorial Foundation, Kowa Life Science Foundation, Takeda Science Foundation, Kato Memorial Bioscience Foundation, DAIKO FOUNDATION, SECOM Science and Technology Foundation, Research Foundation for Opto-Science and Technology, The Nakatani Foundation for Advancement of Measuring Technologies in Biomedical Engineering, and the JSPS KAKENHI (18H02477, 15H04679, and 15K12763).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Abrahamson, E. E., and Moore, R. Y. (2001). Suprachiasmatic nucleus in the mouse: retinal innervation, intrinsic organization and efferent projections. Brain Res. 916, 172–191. doi: 10.1016/s0006-8993(01)02890-6
Albus, H., Bonnefont, X., Chaves, I., Yasui, A., Doczy, J., van der Horst, G. T. J., et al. (2002). Cryptochrome-deficient mice lack circadian electrical activity in the suprachiasmatic nuclei. Curr. Biol. 12, 1130–1133. doi: 10.1016/s0960-9822(02)00923-5
An, S. W., Irwin, R. P., Allen, C. N., Tsai, C., and Herzog, E. D. (2011). Vasoactive intestinal polypeptide requires parallel changes in adenylate cyclase and phospholipase C to entrain circadian rhythms to a predictable phase. J. Neurophysiol. 105, 2289–2296. doi: 10.1152/jn.00966.2010
An, S., Harang, R., Meeker, K., Granados-Fuentes, D., Tsai, C. A., Mazuski, C., et al. (2013). A neuropeptide speeds circadian entrainment by reducing intercellular synchrony. Proc. Natl. Acad. Sci. USA 110, E4355–E4361.
Ansari, N., Agathagelidis, M., Lee, C., Korf, H. W., and von Gall, C. (2009). Differential maturation of circadian rhythms in clock gene proteins in the suprachiasmatic nucleus and the pars tuberalis during mouse ontogeny. Eur. J. Neurosci. 29, 477–489. doi: 10.1111/j.1460-9568.2008.06605.x
Aton, S. J., Colwell, C. S., Harmar, A. J., Waschek, J., and Herzog, E. D. (2005). Vasoactive intestinal polypeptide mediates circadian rhythmicity and synchrony in mammalian clock neurons. Nat. Neurosci. 8, 476–483. doi: 10.1038/nn1419
Aton, S. J., Huettner, J. E., Straume, M., and Herzog, E. D. (2006). GABA and G(i/o) differentially control circadian rhythms and synchrony in clock neurons. Proc. Natl. Acad. Sci. USA 103, 19188–19193. doi: 10.1073/pnas.0607466103
Ban, Y., Shigeyoshi, Y., and Okamura, H. (1997). Development of vasoactive intestinal peptide mRNA rhythm in the rat suprachiasmatic nucleus. J. Neurosci. 17, 3920–3931. doi: 10.1523/jneurosci.17-10-03920.1997
Bedont, J. L., LeGates, T. A., Slat, E. A., Byerly, M. S., Wang, H., Hu, J. F., et al. (2014). Lhx1 controls terminal differentiation and circadian function of the suprachiasmatic nucleus. Cell Rep. 7, 609–622. doi: 10.1016/j.celrep.2014.03.060
Brancaccio, M., Maywood, E. S., Chesham, J. E., Loudon, A. S. I., and Hastings, M. H. (2013). A Gq-Ca2+ Axis controls circuit-level encoding of circadian time in the suprachiasmatic nucleus. Neuron 78, 714–728. doi: 10.1016/j.neuron.2013.03.011
Carmona-Alcocer, V., Abel, J. H., Sun, T. C., Petzold, L. R., Doyle, F. J., Simms, C. L., et al. (2018). Ontogeny of circadian rhythms and synchrony in the suprachiasmatic nucleus. J. Neurosci. 38, 1326–1334. doi: 10.1523/jneurosci.2006-17.2017
Clark, D. D., Gorman, M. R., Hatori, M., Meadows, J. D., Panda, S., and Mellon, P. L. (2013). Aberrant development of the suprachiasmatic nucleus and circadian rhythms in mice lacking the homeodomain protein Six6. J. Biol. Rhythm 28, 15–25. doi: 10.1177/0748730412468084
Collins, B., Pierre-Ferrer, S., Muheim, C., Lukacsovich, D., Cai, Y. C., Spinnler, A., et al. (2020). Circadian VIPergic neurons of the suprachiasmatic nuclei sculpt the sleep-wake cycle. Neuron 108, 486–499.e5.
Colwell, C. S., Michel, S., Itri, J., Rodriguez, W., Tam, J., Lelievre, V., et al. (2003). Disrupted circadian rhythms in VIP- and PHI-deficient mice. Am. J. Physiol. Reg. I 285, R939–R949.
Edwards, M. D., Brancaccio, M., Chesham, J. E., Maywood, E. S., and Hastings, M. H. (2016). Rhythmic expression of cryptochrome induces the circadian clock of arrhythmic suprachiasmatic nuclei through arginine vasopressin signaling. Proc. Natl. Acad. Sci. U S A, 113, 2732–2737.
Evans, J. A., Leise, T. L., Castanon-Cervantes, O., and Davidson, A. J. (2013). Dynamic interactions mediated by nonredundant signaling mechanisms couple circadian clock neurons. Neuron 80, 973–983. doi: 10.1016/j.neuron.2013.08.022
Freeman, G. M. Jr., Krock, R. M., Aton, S. J., Thaben, P., and Herzog, E. D. (2013). GABA networks destabilize genetic oscillations in the circadian pacemaker. Neuron 78, 799–806. doi: 10.1016/j.neuron.2013.04.003
Gizowski, C., Zaelzer, C., and Bourque, C. W. (2016). Clock-driven vasopressin neurotransmission mediates anticipatory thirst prior to sleep. Nature 537, 685–688. doi: 10.1038/nature19756
Hamnett, R., Crosby, P., Chesham, J. E., and Hastings, M. H. (2019). Vasoactive intestinal peptide controls the suprachiasmatic circadian clock network via ERK1/2 and DUSP4 signalling. Nat. Commun. 10:542.
Harmar, A. J., Marston, H. M., Shen, S. B., Spratt, C., West, K. M., Sheward, W. J., et al. (2002). The VPAC(2) receptor is essential for circadian function in the mouse suprachiasmatic nuclei. Cell 109, 497–508. doi: 10.1016/s0092-8674(02)00736-5
Hatori, M., Gill, S., Mure, L. S., Goulding, M., O’Leary, D. D. M., and Panda, S. (2014). Lhx1 maintains synchrony among circadian oscillator neurons of the SCN. Elife 3:e03357.
Hattar, S., Kumar, M., Park, A., Tong, P., Tung, J., Yau, K.W., and Berson, D.M. (2006). Central projections of melanopsin-expressing retinal ganglion cells in the mouse. J. Comp. Neurol. 497, 326–349. doi: 10.1002/cne.20970
Hermanstyne, T. O., Simms, C. L., Carrasquillo, Y., Herzog, E. D., and Nerbonne, J. M. (2016). Distinct firing properties of vasoactive intestinal peptide-expressing neurons in the suprachiasmatic nucleus. J. Biol. Rhythm 31, 57–67. doi: 10.1177/0748730415619745
Herzog, E. D., Aton, S. J., Numano, R., Sakaki, Y., and Tei, H. (2004). Temporal precision in the mammalian circadian system: a reliable clock from less reliable neurons. J. Biol. Rhythm 19, 35–46. doi: 10.1177/0748730403260776
Hirata, Y., Enoki, R., Kuribayashi-Shigetomi, K., Oda, Y., Honma, S., and Honma, K. (2019). Circadian rhythms in Per1, PER2 and Ca2+ of a solitary SCN neuron cultured on a microisland. Sci. Rep. 9:18271.
Honma, S., Nakamura, W., Shirakawa, T., and Honma, K. (2004). Diversity in the circadian periods of single neurons of the rat suprachiasmatic nucleus depends on nuclear structure and intrinsic period. Neurosci. Lett. 358, 173–176. doi: 10.1016/j.neulet.2004.01.022
Ikeda, M., Sugiyama, T., Wallace, C. S., Gompf, H. S., Yoshioka, T., Miyawaki, A., et al. (2003). Circadian dynamics of cytosolic and nuclear Ca2+ in single suprachiasmatic nucleus neurons. Neuron 38, 253–263. doi: 10.1016/s0896-6273(03)00164-8
Inagaki, N., Honma, S., Ono, D., Tanahashi, Y., and Honma, K. (2007). Separate oscillating cell groups in mouse suprachiasmatic nucleus couple photoperiodically to the onset and end of daily activity. Proc. Natl. Acad. Sci. USA 104, 7664–7669. doi: 10.1073/pnas.0607713104
Isobe, Y., and Muramatsu, K. (1995). Day-night differences in the contents of vasoactive-intestinal-peptide, gastrin-releasing peptide and arg-vasopressin in the suprachiasmatic nucleus of rat pups during postnatal-development. Neurosci. Lett. 188, 45–48. doi: 10.1016/0304-3940(95)11391-9
Jin, X. W., Shearman, L. P., Weaver, D. R., Zylka, M. J., De Vries, G. J., and Reppert, S. M. (1999). A molecular mechanism regulating rhythmic output from the suprachiasmatic circadian clock. Cell 96, 57–68. doi: 10.1016/s0092-8674(00)80959-9
Jones, J. R., Simon, T., Lones, L., and Herzog, E. D. (2018). SCN VIP neurons are essential for normal light-mediated resetting of the circadian system. J. Neurosci. 38, 7986–7995. doi: 10.1523/jneurosci.1322-18.2018
Kabrita, C. S., and Davis, F. C. (2008). Development of the mouse suprachiasmatic nucleus: determination of time of cell origin and spatial arrangements within the nucleus. Brain Res. 1195, 20–27. doi: 10.1016/j.brainres.2007.12.020
Landgraf, D., Achten, C., Dallmann, F., and Oster, H. (2015). Embryonic development and maternal regulation of murine circadian clock function. Chronobiol. Int. 32, 416–427. doi: 10.3109/07420528.2014.986576
Lee, I. T., Chang, A. S., Manandhar, M., Shan, Y. L., Fan, J. M., Izumo, M., et al. (2015). Neuromedin S-Producing neurons act as essential pacemakers in the suprachiasmatic nucleus to couple clock neurons and dictate circadian rhythms. Neuron 85, 1086–1102. doi: 10.1016/j.neuron.2015.02.006
LeGates, T. A., Fernandez, D. C., and Hattar, S. (2014). Light as a central modulator of circadian rhythms, sleep and affect. Nat. Rev. Neurosci. 15, 443–454. doi: 10.1038/nrn3743
Liu, A. C., Welsh, D. K., Ko, C. H., Tran, H. G., Zhang, E. E., Priest, A. A., et al. (2007). Intercellular coupling confers robustness against mutations in the SCN circadian clock network. Cell 129, 605–616. doi: 10.1016/j.cell.2007.02.047
Maywood, E. S., Chesham, J. E., O’Brien, J. A., and Hastings, M. H. (2011). A diversity of paracrine signals sustains molecular circadian cycling in suprachiasmatic nucleus circuits. Proc. Natl. Acad. Sci. U. S. A. 108, 14306–14311.
Maywood, E. S., Reddy, A. B., Wong, G. K. Y., O’Neill, J. S., O’Brien, J. A., McMahon, D. G., et al. (2006). Synchronization and maintenance of timekeeping in suprachiasmatic circadian clock cells by neuropeptidergic signaling. Curr. Biol. 16, 599–605. doi: 10.1016/j.cub.2006.02.023
Mazuski, C., Abel, J. H., Chen, S. P., Hermanstyne, T. O., Jones, J. R., Simon, T., et al. (2018). Entrainment of circadian rhythms depends on firing rates and neuropeptide release of VIP SCN neurons. Neuron 99, 555–563.e5.
Mazuski, C., Chen, S. P., and Herzog, E. D. (2020). Different roles for VIP neurons in the neonatal and adult suprachiasmatic nucleus. J. Biol. Rhythm 35, 465–475. doi: 10.1177/0748730420932073
McNeill, D. S., Sheely, C. J., Ecker, J. L., Badea, T. C., Morhardt, D., Guido, W., and Hattar, S. (2011). Development of melanopsin-based irradiance detecting circuitry. Neural Dev. 6:8. doi: 10.1186/1749-8104-6-8
Mieda, M., Ono, D., Hasegawa, E., Okamoto, H., Honma, K., Honma, S., et al. (2015). Cellular clocks in AVP neurons of the SCN are critical for interneuronal coupling regulating circadian behavior rhythm. Neuron 85, 1103–1116. doi: 10.1016/j.neuron.2015.02.005
Moore, R. Y., and Bernstein, M. E. (1989). Synaptogenesis in the rat suprachiasmatic nucleus demonstrated by electron-microscopy and Synapsin-1 immunoreactivity. J. Neurosci. 9, 2151–2162. doi: 10.1523/jneurosci.09-06-02151.1989
Moore, R. Y., and Eichler, V. B. (1972). Loss of a circadian adrenal corticosterone rhythm following suprachiasmatic lesions in rat. Brain Res. 42, 201–206. doi: 10.1016/0006-8993(72)90054-6
Myung, J., Hong, S., DeWoskin, D., De Schutter, E., Forger, D. B., and Takumi, T. (2015). GABA-mediated repulsive coupling between circadian clock neurons in the SCN encodes seasonal time. Proc. Natl. Acad. Sci. USA 112, E3920–E3929.
Okamura, H., Fukui, K., Koyama, E., Tsutou, H. L. O., Tsutou, T., Terubayashi, H., et al. (1983). Time of vasopressin neuron origin in the mouse hypothalamus - examination by combined technique of immunocytochemistry and [H-3]thymidine autoradiography. Dev. Brain Res. 9, 223–226.
Okamura, H., Miyake, S., Sumi, Y., Yamaguchi, S., Yasui, A., Muijtjens, M., et al. (1999). Photic induction of mPer1 and mPer2 in Cry-deficient mice lacking a biological clock. Science 286, 2531–2534. doi: 10.1126/science.286.5449.2531
Ono, D., Honma, K., Yanagawa, Y., Yamanaka, A., and Honma, S. (2019). GABA in the suprachiasmatic nucleus refines circadian output rhythms in mice. Commun. Biol. 2:232.
Ono, D., Honma, S., and Honma, K. (2013). Cryptochromes are critical for the development of coherent circadian rhythms in the mouse suprachiasmatic nucleus. Nat. Commun. 4:1666.
Ono, D., Honma, S., and Honma, K. (2016). Differential roles of AVP and VIP signaling in the postnatal changes of neural networks for coherent circadian rhythms in the SCN. Sci. Adv. 2:e1600960. doi: 10.1126/sciadv.1600960
Ono, D., Mukai, Y., Hung, C. J., Chowdhury, S., Sugiyama, T., and Yamanaka, A. (2020). The mammalian circadian pacemaker regulates wakefulness via CRF neurons in the paraventricular nucleus of the hypothalamus. Sci. Adv. 6:eabd0384. doi: 10.1126/sciadv.abd0384
Patton, A. P., Edwards, M. D., Smyllie, N. J., Hamnett, R., Chesham, J. E., Brancaccio, M., et al. (2020). The VIP-VPAC2 neuropeptidergic axis is a cellular pacemaking hub of the suprachiasmatic nucleus circadian circuit. Nat. Commun. 11:3394.
Paul, S., Hanna, L., Harding, C., Hayter, E. A., Walmsley, L., Bechtold, D. A., et al. (2020). Output from VIP cells of the mammalian central clock regulates daily physiological rhythms. Nat. Commun. 11:1453.
Reppert, S. M., and Schwartz, W. J. (1983). Maternal coordination of the fetal biological clock inutero. Science 220, 969–971. doi: 10.1126/science.6844923
Reppert, S. M., and Weaver, D. R. (2002). Coordination of circadian timing in mammals. Nature 418, 935–941. doi: 10.1038/nature00965
Romero, M. T., and Silver, R. (1990). Time course of peptidergic expression in fetal suprachiasmatic nucleus transplanted into adult hamster. Dev. Brain Res. 57, 1–6. doi: 10.1016/0165-3806(90)90177-z
Shan, Y., Abel, J. H., Li, Y., Izumo, M., Cox, K. H., Jeong, B., et al. (2020). Dual-color single-cell imaging of the suprachiasmatic nucleus reveals a circadian role in network synchrony. Neuron 108, 164–179 e167.
Shimada, M., and Nakamura, T. (1973). Time of neuron origin in mouse hypothalamic nuclei. Exp. Neurol. 41, 163–173. doi: 10.1016/0014-4886(73)90187-8
Shimogori, T., Lee, D. A., Miranda-Angulo, A., Yang, Y. Q., Wang, H., Jiang, L. Z., et al. (2010). A genomic atlas of mouse hypothalamic development. Nat. Neurosci. 13, U767–U153.
Shimomura, H., Moriya, T., Sudo, M., Wakamatsu, H., Akiyama, M., Miyake, Y., et al. (2001). Differential daily expression of Per1 and Per2 mRNA in the suprachiasmatic nucleus of fetal and early postnatal mice. Eur. J. Neurosci. 13, 687–693. doi: 10.1046/j.0953-816x.2000.01438.x
Shinohara, K., Honma, S., Katsuno, Y., Abe, H., and Honma, K. (1995). 2 Distinct Oscillators in the rat suprachiasmatic nucleus in-Vitro. Proc. Natl. Acad. Sci. USA 92, 7396–7400. doi: 10.1073/pnas.92.16.7396
Shinohara, K., Tominaga, K., and Inouye, S. I. T. (1999). Phase dependent response of vasoactive intestinal polypeptide to light and darkness in the suprachiasmatic nucleus. Neurosci. Res. 33, 105–110. doi: 10.1016/s0168-0102(98)00122-9
Stephan, F. K., and Zucker, I. (1972). Circadian-rhythms in drinking behavior and locomotor activity of rats are eliminated by hypothalamic-lesions. Proc. Natl. Acad. Sci. USA 69, 1583–1586. doi: 10.1073/pnas.69.6.1583
Todd, W. D., Venner, A., Anaclet, C., Broadhurst, R. Y., De Luca, R., Bandaru, S. S., et al. (2020). Suprachiasmatic VIP neurons are required for normal circadian rhythmicity and comprised of molecularly distinct subpopulations. Nat. Commun. 11:4410.
Uhl, G. R., and Reppert, S. M. (1986). Suprachiasmatic nucleus vasopressin messenger-rna - circadian variation in normal and brattleboro rats. Science 232, 390–393. doi: 10.1126/science.3961487
van der Horst, G. T. J., Muijtjens, M., Kobayashi, K., Takano, R., Kanno, S., Takao, M., et al. (1999). Mammalian Cry1 and Cry2 are essential for maintenance of circadian rhythms. Nature 398, 627–630. doi: 10.1038/19323
VanDunk, C., Hunter, L. A., and Gray, P. A. (2011). Development, maturation, and necessity of transcription factors in the mouse suprachiasmatic nucleus. J. Neurosci. 31, 6457–6467. doi: 10.1523/jneurosci.5385-10.2011
Weaver, D. R. (1998). The suprachiasmatic nucleus: a 25-year retrospective. J. Biol. Rhythm 13, 100–112. doi: 10.1177/074873098128999952
Webb, A. B., Angelo, N., Huettner, J. E., and Herzog, E. D. (2009). Intrinsic, nondeterministic circadian rhythm generation in identified mammalian neurons. Proc. Natl. Acad. Sci. USA 106, 16493–16498. doi: 10.1073/pnas.0902768106
Welsh, D. K., Logothetis, D. E., Meister, M., and Reppert, S. M. (1995). Individual Neurons dissociated from rat suprachiasmatic nucleus express independently phased circadian firing rhythms. Neuron 14, 697–706. doi: 10.1016/0896-6273(95)90214-7
Wen, S., Ma, D., Zhao, M., Xie, L., Wu, Q., Gou, L., et al. (2020). Spatiotemporal single-cell analysis of gene expression in the mouse suprachiasmatic nucleus. Nat. Neurosci. 23, 456–467. doi: 10.1038/s41593-020-0586-x
Wreschnig, D., Dolatshad, H., and Davis, F. C. (2014). Embryonic development of circadian oscillations in the mouse hypothalamus. J. Biol. Rhythm 29, 299–310. doi: 10.1177/0748730414545086
Yamaguchi, S., Isejima, H., Matsuo, T., Okura, R., Yagita, K., Kobayashi, M., et al. (2003). Synchronization of cellular clocks in the suprachiasmatic nucleus. Science 302, 1408–1412. doi: 10.1126/science.1089287
Yamaguchi, Y., Suzuki, T., Mizoro, Y., Kori, H., Okada, K., Chen, Y. L., et al. (2013). Mice genetically deficient in vasopressin V1a and V1b receptors are resistant to jet lag. Science 342, 85–90. doi: 10.1126/science.1238599
Keywords: circadian rhythm, suprachiasmatic nucleus, AVP, VIP, neuronal coupling, synchronization, entrainment
Citation: Ono D, Honma K and Honma S (2021) Roles of Neuropeptides, VIP and AVP, in the Mammalian Central Circadian Clock. Front. Neurosci. 15:650154. doi: 10.3389/fnins.2021.650154
Received: 06 January 2021; Accepted: 10 March 2021;
Published: 15 April 2021.
Edited by:
Masayuki Ikeda, University of Toyama, JapanReviewed by:
Elizabeth S. Maywood, MRC Laboratory of Molecular Biology (LMB), United KingdomWilliam David Todd, University of Wyoming, United States
Copyright © 2021 Ono, Honma and Honma. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Daisuke Ono, ZGFpLW9ub0ByaWVtLm5hZ295YS11LmFjLmpw; Sato Honma, c2F0aG9ubWFAbWVkLmhva3VkYWkuYWMuanA=
 Daisuke Ono
Daisuke Ono Ken-ichi Honma
Ken-ichi Honma Sato Honma
Sato Honma