- 1Biodiversity Research Cluster, Department of Biology, Faculty of Science, Mahidol University, Bangkok, Thailand
- 2Center of Excellence for Vectors and Vector-Borne Diseases, Faculty of Science, Mahidol University, Nakhon Pathom, Thailand
Ticks can transmit a wide variety of pathogens, including bacteria. Here, we report the detection of tick-associated bacteria in Chaiyaphum Province, northeastern Thailand. There have been few reports of tick-borne bacterial pathogens in the study areas, which are evergreen forests dominated by plateaus at elevations of approximately 1,000 m. In total, 94 ticks were collected from vegetation. They were screened for the presence of Coxiella, Francisella, Rickettsia, and Borrelia bacteria using PCR assays. In this study, we found ticks from two genera, Haemaphysalis and Amblyomma, that were positive for Coxiella-like bacteria (CLB) and Rickettsia. Francisella and Borrelia spp. were not detected in these two tick genera. The results revealed the evolutionary relationships of CLB in Amblyomma testudinarium, Haemaphysalis lagrangei, and Haemaphysalis obesa ticks using the 16S rRNA and rpoB markers, which clustered together with known isolates of ticks from the same genera. In contrast, the groEL marker showed different results. On the basis of the groEL phylogenetic analysis and BLAST results, three groups of CLB were found: (1) CLB from A. testudinarium grouped as a sister clade to CLB from Ixodes ricinus; (2) CLB from Haemaphysalis lagrangei was distantly related to CLB from Haemaphysalis wellingtoni; and (3) CLB from A. testudinarium grouped as sister clade to CLB from Amblyomma from French Guiana and Brazil. For Rickettsia studies, phylogenetic trees of the gltA, ompB, and sca4 genes revealed two groups of Spotted Fever Group (SFG) Rickettsiae: (1) SFG Rickettsiae that formed a sister clade with Rickettsia tamurae AT-1 (belong to the Rickettsia helvetica subgroup) in A. testudinarium and (2) SFG Rickettsiae that formed a distantly related group to Rickettsia rhipicephali 3-7-female6-CWPP (belong to the Rickettsia massiliae subgroup) in A. testudinarium. This study expanded our knowledge of the diversity of tick-borne Coxiella and Rickettsia bacteria. The pathogenic roles of these bacteria also need to be investigated further.
Introduction
Ticks are important hematophagous ectoparasites of both humans and animals. They can transmit a wide range of pathogens, such as Coxiella, Francisella, Borrelia, and Rickettsia bacteria (1–4). Hard ticks, such as the genera Amblyomma, Dermacentor, and Rhipicephalus, have been identified as the main vectors of Spotted Fever Group (SFG) Rickettsiae (5). Several tick species have been reported in Chaiyaphum Province, northeastern Thailand. These include Amblyomma testudinarium, Haemaphysalis asiatica, Haemaphysalis hystricis, Haemaphysalis semermis, Rhipicephalus haemaphysaloides, and Ixodes granulatus (6). Rickettsia spp. are gram-negative bacteria that cause SFG diseases. SFG rickettsioses have been reported in many regions of the world, including Japan, northern China, Korea (7–9), and Thailand (10). SFG rickettsioses are characterized by fever, headache, muscle pain, maculopapular rash, and developing eschar at the site of tick bites (11). In Thailand, Rickettsia spp. have been reported near the Thai-Myanmar border, such as Rickettsia sp. strain RDla420 identified in Dermacentor auratus ticks obtained from a bear and Rickettsia sp. strain RDla440 detected in Dermacentor larval ticks from a wild pig nest (12). In addition, Sumrandee et al. (13) reported the first evidence of a Rickettsia sp. that is closely related to Rickettsia tamurae in Rhipicephalus (Boophilus) microplus ticks from Thailand. In Chaiyaphum Province, Malaisri et al. (14) reported a phylogenetic analysis of new Rickettsia genotypes that were closely related to Rickettsia tamurae and Rickettsia monacensis and might be pathogenic to humans.
Q fever is a zoonotic disease caused by Coxiella burnetii; infection mainly arises through the inhalation of airborne particles contaminated with bacteria. The clinical features of Q fever include flu-like symptoms to pneumonia and granulomatous hepatitis in serious cases (15). Coxiella burnetii infections and Coxiella-like bacteria (CLB) have been found in humans and other animals in Thailand (16–19). For example, C. burnetii was found in Thai patients in Khon Kaen Province, northeastern Thailand (20). CLB can promote the reproductive fitness and development of Haemaphysalis longicornis ticks (21). In addition, CLB can interfere with the colonization and transmission of pathogens. For example, CLB can impact pathogen susceptibility in ticks, e.g., CLB can defend their Rhipicephalus haemaphysaloides tick hosts against the pathogenic microbe Babesia microti (22). Interestingly, CLB was determined to be the cause of death in a female eclectus parrot (Eclectus roratus) (23). In Thailand, CLB was also detected in Haemaphysalis ticks, such as Haemaphysalis shimoga and Haemaphysalis lagrangei (24). Moreover, Trinachartvanit et al. (25) reported CLB in Haemaphysalis wellingtoni tick-infested fowl from various parts of Thailand.
The objectives of this study were to identify the presence of bacteria in ticks collected from vegetation and to reveal the overall diversity of bacterial infections and species in ticks using PCR and phylogenetic analysis.
Materials and Methods
Tick Collection and Identification
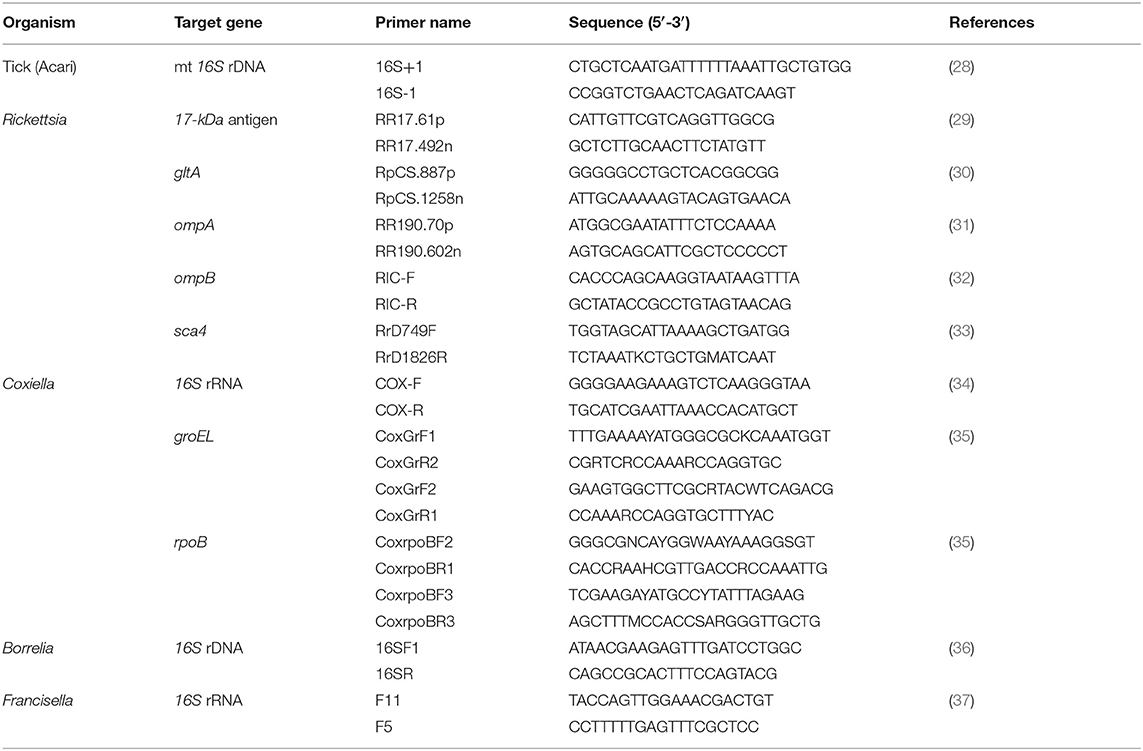
In 2014–2015, ticks (larvae, nymphs, and adults) were collected from vegetation from forests in Chaiyaphum Province, Thailand (16°16′25.2″N 101°29′02″E; 16°22′23.0″N 101°46′38.7″E; and 16°12′18.9″N 101°52′22.9″E). The areas being studied are evergreen forests dominated by plateaus at elevations of approximately 1,000 m. We placed the ticks into tubes that were held in a container with liquid nitrogen and stored them in a freezer after returning to the laboratory. Morphological identification of all tick stages was performed under a stereomicroscope (26, 27). Tick identification was performed using molecular methods on the ticks that were positive for either Coxiella or Rickettsia bacteria; previously published primers (16S+1/16S-1) for the amplification of mt 16S rDNA were used (28). There were seven pools of Haemaphysalis ticks at the immature stage, including nymphs (four pools: three pools of five and one pool of four) and larvae (three pools: one pool of six, one pool of seven, and one pool of 15). For the Amblyomma immature stage (nymphs), there were five pools of three nymphs.
DNA Extraction
Before DNA extraction, ticks were cleaned three times with 70% ethanol, 10% sodium hypochlorite, and sterile distilled water. DNA extraction was conducted using a DNeasy Blood and Tissues Kit (Qiagen) according to the manufacturer's protocol. The DNA products were stored at −20°C until use as templates for the PCR assay.
Molecular Analysis
All extracted DNA samples were used as templates for PCR assays with specific bacterial primers for detecting the presence of Coxiella, Rickettsia, Francisella, and Borrelia. Coxiella-positive bacteria were identified through the 16S rRNA, groEL (60-kDa chaperone heat shock protein B), and rpoB genes (β subunit of bacterial RNA polymerase). Rickettsia species were screened by PCR, targeting the 17-kDa antigen, citrate synthase (gltA), outer membrane protein A (ompA), outer membrane protein B (ompB), and cell surface antigen (sca4) genes. PCR primer pairs for the detection of bacterial species, primer name, target genes, and size of the amplicons (bp) are shown in Table 1. The PCR product of the expected size from each corresponding primer pair (gene) was cloned and sequenced as a positive control. A sterile distilled water negative control was also included.
DNA Purification, Sequencing, and Phylogenetic Analysis
The positive PCR products were purified with a Nucleospin Gel and PCR Clean-up Kit (Düren, Germany) and sequenced. The DNA sequence alignment of representative positive samples generated by this study was created using the CLUSTALW program. The nucleotide sequences were analyzed and blasted with the National Center for Biotechnology Information BLASTn database. Afterward, phylogenetic analyses were carried out using the maximum likelihood (ML) (38) and neighbor-joining (NJ) (39) methods (evaluated by bootstrap analysis with 1,000 replicates) for Coxiella and Rickettsia spp., respectively.
Results
Tick Collection
A total of 94 ticks were collected and identified. The species, number, and life stage of the ticks are shown in Table 2. The ticks belonged to two genera, Haemaphysalis and Amblyomma. Of the adult ticks, 13 males and 12 females belonged to H. lagrangei. The remaining ticks belonged to A. testudinarium (two female ticks). In the immature stage, Haemaphysalis sp. (22 nymphs and 28 larvae) was the most commonly collected, followed by Amblyomma sp. (17 nymphs). The species of all ticks included in the phylogenetic trees were confirmed by molecular methods, and their sequences were submitted to GenBank with the accession numbers shown in Tables 3, 4.
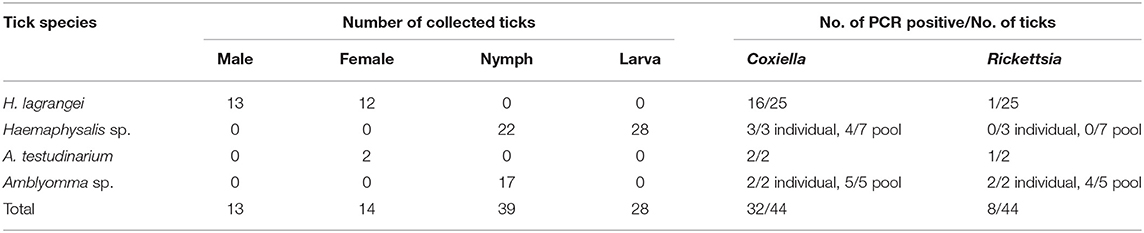
Table 2. Species, number, life stage of ticks, and results of bacterial infection in ticks analyzed by PCR (positive result of each bacterium) collected from vegetation in Chaiyaphum Province, Thailand.
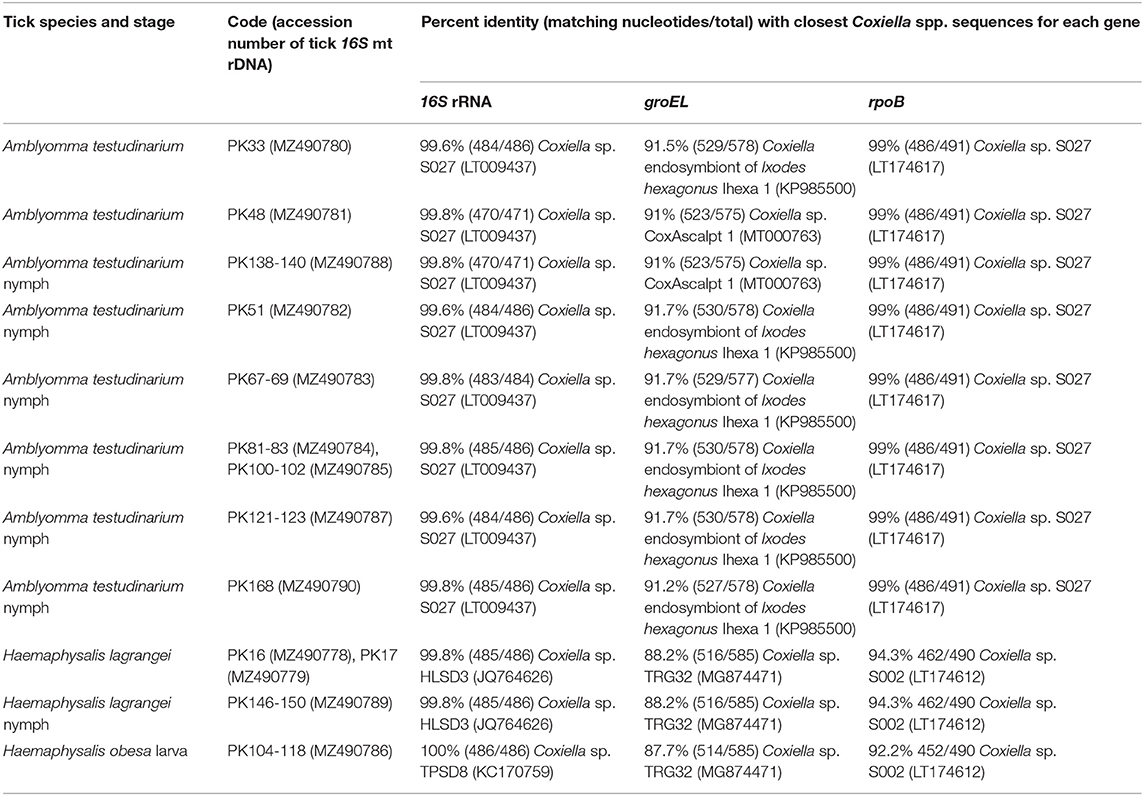
Table 3. Details of GenBank accession numbers of the Coxiella gene sequences and BLAST analysis of these sequences from tick samples collected from vegetation in Chaiyaphum Province, Thailand.
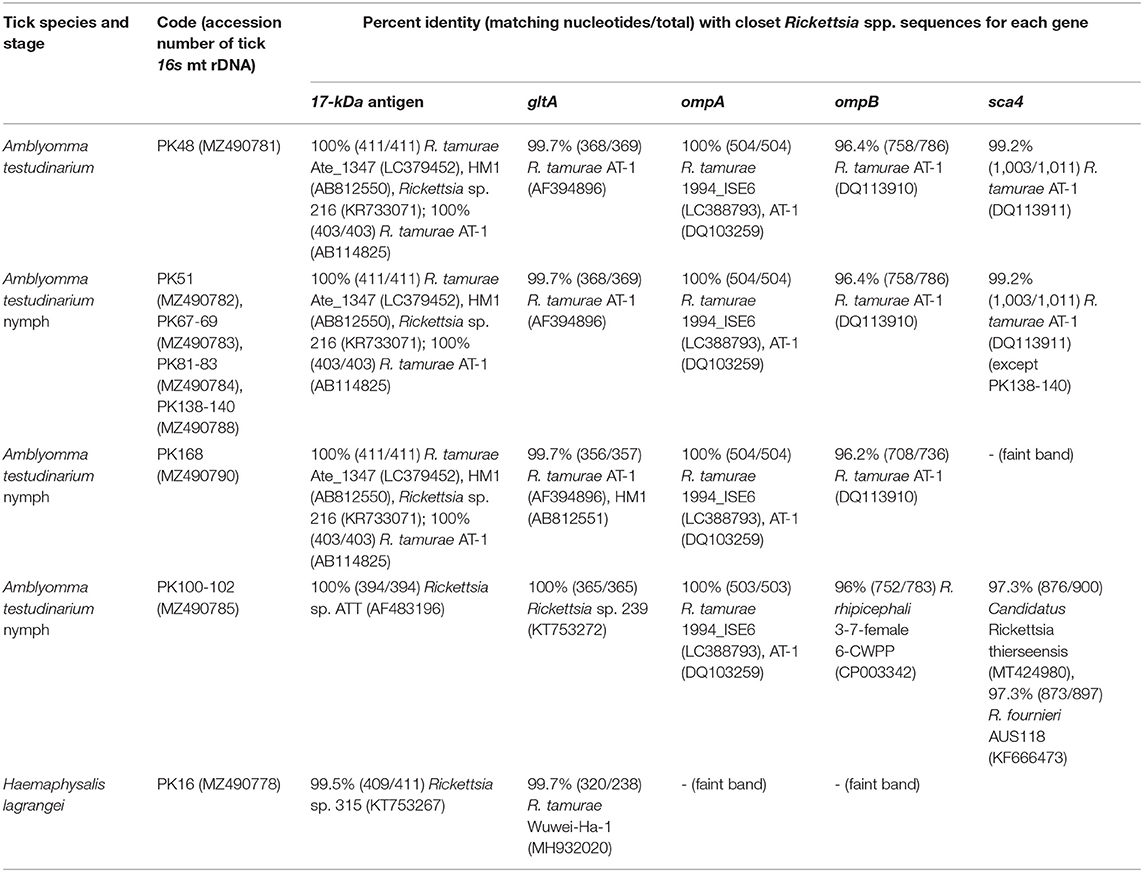
Table 4. Details of GenBank accession numbers of the Rickettsia gene sequences and BLAST analysis of these sequences from tick samples collected from vegetation in Chaiyaphum Province, Thailand.
Detection of Bacteria
Coxiella and Rickettsia bacteria were detected in two genera of ticks (Haemaphysalis and Amblyomma), whereas Francisella sp. and Borrelia sp. were not identified in this study (Table 2). Single infection with Coxiella was detected in 16 of 25 H. lagrangei adult ticks (six of 13 males and 10 of 12 females), and single infection with Rickettsia was detected in one of 25 H. lagrangei females. Adult A. testudinarium ticks were infected with Coxiella (two of two females) and Rickettsia (one of two females). Single infection with Coxiella was detected in three of three nymphs and four of seven pools of Haemaphysalis ticks (one pool of four nymphs, two pools of five nymphs, and one pool of 15 larvae). In addition, two of two nymphs and five of five pools of Amblyomma ticks were positive for Coxiella and two of two nymphs and four of five pools were positive for Rickettsia. However, infection with Rickettsia was not detected in the immature stage of Haemaphysalis. Moreover, coinfection with these two bacteria was detected in infected adults of H. lagrangei. In addition, adults of A. testudinarium were coinfected with Coxiella and Rickettsia. Coinfection with these two bacteria was also present in Amblyomma nymphal ticks.
DNA Sequencing and Phylogenetic Analysis
A phylogenetic tree based on the partial sequences of 16S rRNA revealed that all Coxiella sequences detected in this study belonged to two endosymbiotic groups (Figure 1A). Coxiella sequences detected in H. lagrangei and Haemaphysalis obesa were in the first group. PK16, PK17, and PK146-150 clustered with CLB of HLSD3 found in H. lagrangei, whereas PK104-118 grouped with CLB of TPSD8 detected in H. obesa. These sequences were closely related to those of CLB in H. hystricis S002 (LT009432), H. longicornis 47 (AY342035), and H. longicornis A (AB001519). Coxiella sequences in A. testudinarium and Amblyomma sp. formed a monophyletic clade and clustered together with CLB found in Amblyomma sp. S027 and A. testudinarium AMTKK2.1 from Malaysia and Thailand, respectively (Figure 1A). BLAST analysis of the Coxiella 16S rRNA and groEL and rpoB gene sequences from Haemaphysalis and Amblyomma ticks is shown in Table 3.
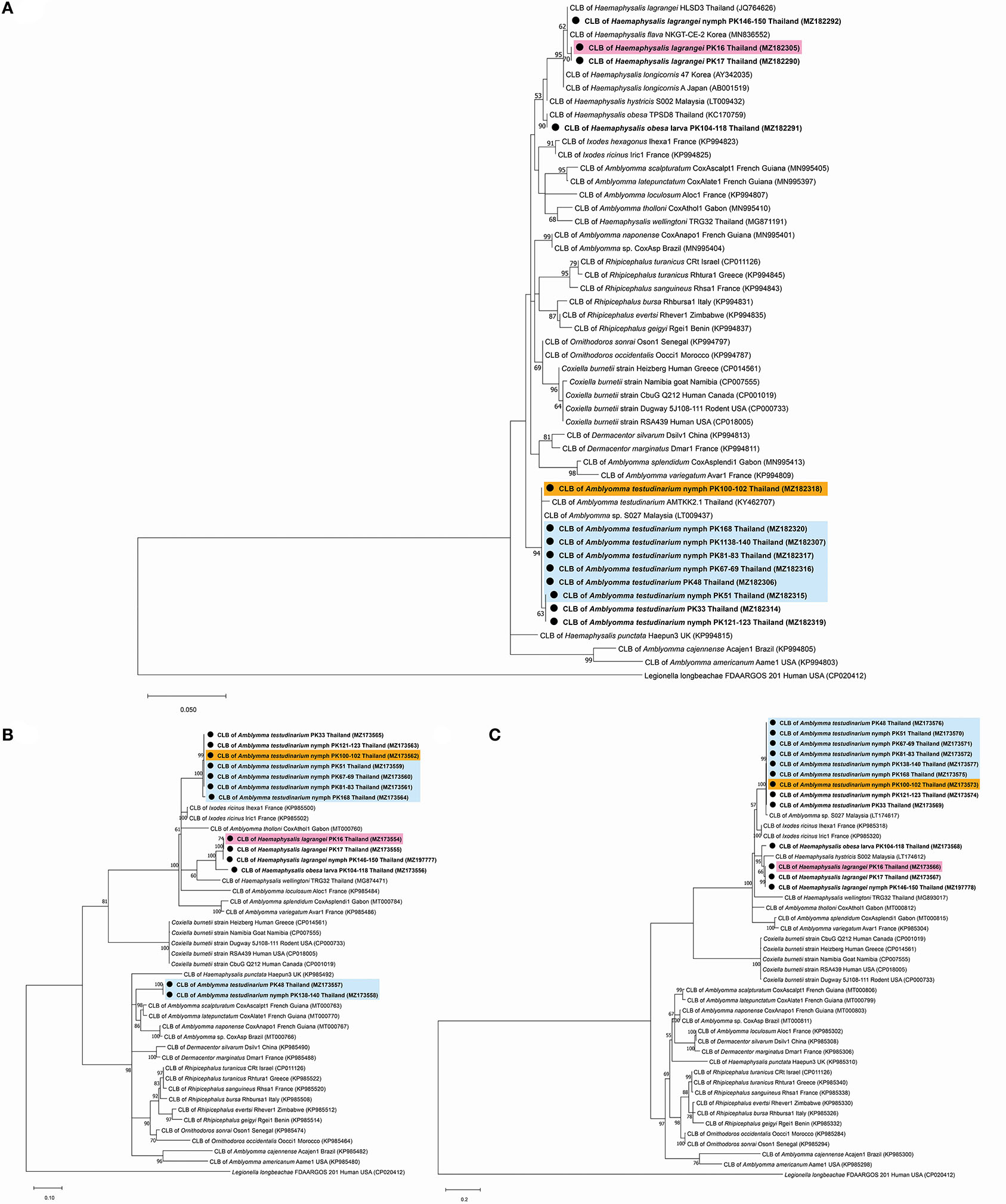
Figure 1. Phylogenetic tree of Coxiella species gene sequences using the maximum likelihood method with 1,000 bootstrap replicates (bootstrap values < 50% are not shown). (A) 16S rRNA gene: Legionella longbeachae was used as the outgroup. (B) groEL gene: Legionella longbeachae was used as the outgroup. (C) rpoB gene: Legionella longbeachae was used as the outgroup. DNA from Coxiella spp. amplified from ticks identified in this study is indicated by black dots and bold font. The scale bar indicates nucleotide substitutions per site.
Interestingly, the BLAST results of the groEL gene sequences from this work showed DNA sequence identities of <92% compared to the existing sequences. Phylogenetic trees based on the partial sequences of groEL revealed that CLB in this study were clustered into three groups (Figure 1B). All sequences of Coxiella spp. in Amblyomma (except PK48 and PK138-140) grouped together as sister clades to the CLB of I. ricinus. CLB in Haemaphysalis grouped as sister clades with CLB detected in H. wellingtoni TRG32 (MG874471). Interestingly, the groEL sequences of PK48 and PK138-140 clearly formed separate clades from the other Coxiella spp. in A. testudinarium obtained in the first group, although they still formed a sister clade with the Amblyomma genus from other geographical regions. CLB in Haemaphysalis (PK16, PK17, PK104-118, and PK146-150) from this study (Figure 1B) formed a distinct clade from those of CLB in Amblyomma. Importantly, the BLAST results of the rpoB gene sequences from this work exhibited DNA sequence identities <95% compared to the existing sequences for Haemaphysalis (PK16, PK17, PK146-150, and PK104-118). Phylogenetic trees based on the partial sequences of rpoB revealed that all CLB sequences detected in this study were also divided into two groups. The first group included Coxiella spp. detected in Amblyomma ticks that formed a monophyletic clade and grouped together with CLB in Amblyomma sp. S027 (LT174617) from Malaysia. The second group included Coxiella sequences found in Haemaphysalis that formed an independent clade and grouped with CLB in H. hystricis S002 (LT174612) from Malaysia (Figure 1C).
BLAST analysis of the Rickettsia 17-kDa antigen, gltA, ompA, ompB, and sca4 gene sequences from Amblyomma and Haemaphysalis ticks is shown in Table 4. The bands for ompA and ompB from PK16 were faint and could not be sequenced. Interestingly, the BLAST search results of all ompB genes studied herein showed <97% DNA sequence similarity compared to the existing ompB genes in Rickettsia spp. In addition, the BLAST search results of PK100-102 showed that the Rickettsia ompB gene sequences from A. testudinarium nymphal ticks had 96% sequence similarity to Rickettsia rhipicephali 3-7-female 6-CWPP (CP003342). Moreover, the BLAST search results of the sca4 gene of PK100-102 showed 97.3% identity to Candidatus Rickettsia thierseensis (MT424980) and Rickettsia fournieri AUS118 (KF666473). Phylogenetic trees based on the partial sequences of the 17-kDa, gltA, ompB, and sca4 genes from Rickettsia sp. are shown in Figures 2A–D. Phylogenetic trees based on the partial sequences of the 17-kDa (Figure 2A) and gltA (Figure 2B) genes indicated that the Rickettsia spp. detected in this study formed three groups. The first group (PK16) was clustered with Rickettsia sp. HOT2 and Rickettsia sp. 315 (17-kDa gene), and this group was within the Rickettsia massiliae subgroup of Rickettsia. Phylogenetically, the gltA PK16 sequence grouped with the Rickettsia raoultii clade (R. massiliae subgroup) and was closely related to the clades containing Rickettsia japonica and Rickettsia heilongjiangensis. The second group included PK100-102, which clustered together with Rickettsia sp. in A. testudinarium from Laos. The third group (PK48, PK51, PK67-69, PK81-83, PK138-140, and PK168) formed a sister clade with R. tamurae based on their 17-kDa and gltA gene sequences.
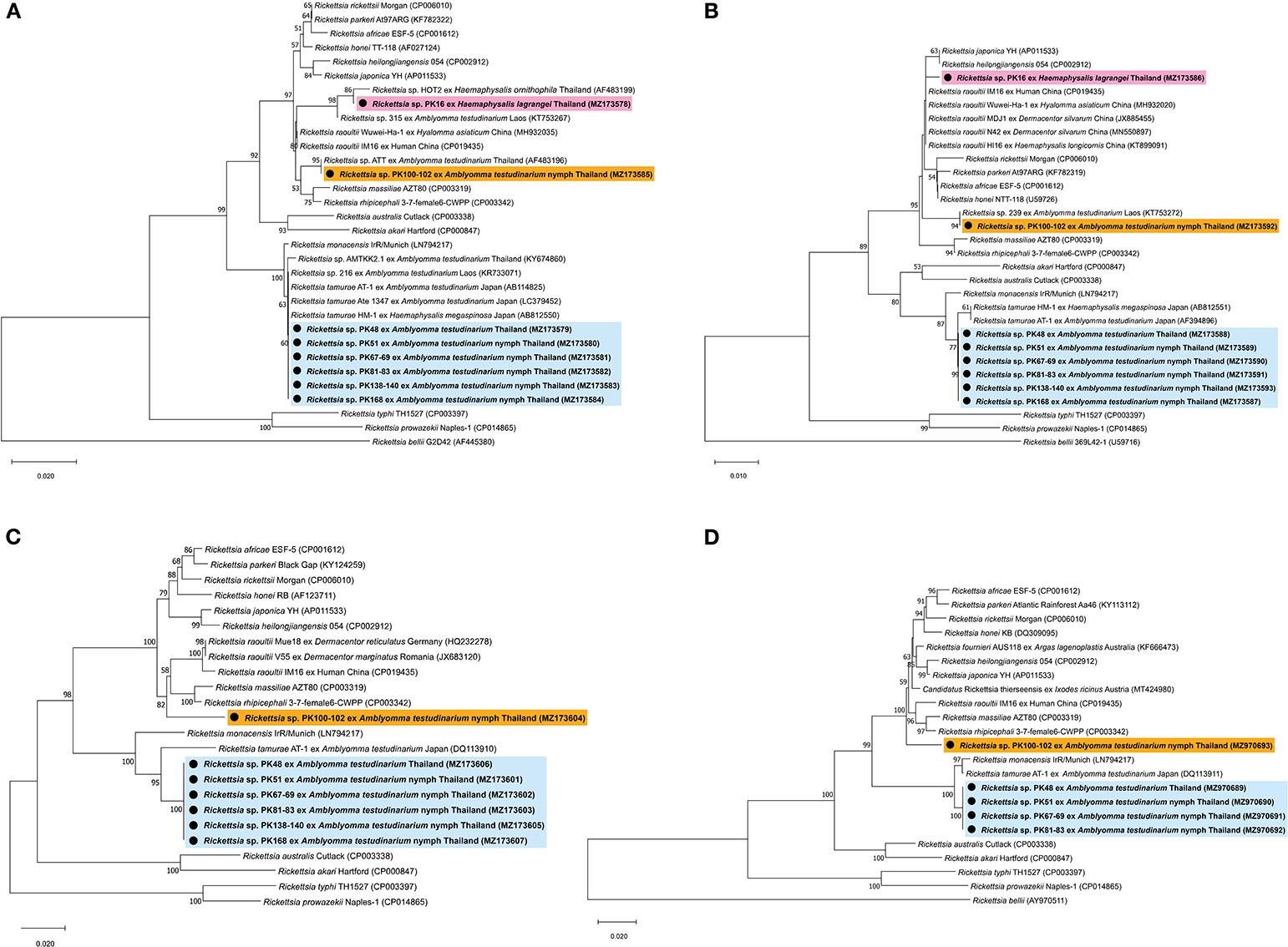
Figure 2. Phylogenetic tree of Rickettsia species using the neighbor-joining method with 1,000 bootstrap replicates (bootstrap values < 50% are not shown). (A) 17-kDa antigen gene: Rickettsia bellii was used as the outgroup. (B) gltA gene: Rickettsia bellii was used as the outgroup. (C) ompB gene: Rickettsia typhi and Rickettsia prowazekii were used as outgroups. (D) sca4 gene: Rickettsia bellii was used as the outgroup. DNA from Rickettsia spp. amplified from ticks identified in this study is indicated by black dots and bold font. The scale bar indicates nucleotide substitutions per site.
However, phylogenetic analysis based on the partial sequence of the ompA gene was not included in this study because the ompA amplicons might be the products of contaminated reactions. In addition, phylogenetic trees based on the partial sequences of the ompB gene revealed that Rickettsia spp. formed two groups: the first group (including PK100-102) formed a sister clade to R. raoultii, R. rhipicephali, and R. massiliae (R. massiliae subgroup), and the second group (including PK48, PK51, PK67-69, PK81-83, PK138-140, and PK168) grouped within the clade containing R. tamurae (Rickettsia helvetica subgroup) (Figure 2C). A phylogenetic tree of the sca4 gene (not including sequence PK138-140) showed the presence of two groups within SFG Rickettsia (Figure 2D), similar to the results for the ompB gene.
Discussion
CLB have been identified in several tick genera, including Haemaphysalis and Amblyomma, and in at least two-thirds of tick species (15, 40, 41). Moreover, the tissue distribution of this symbiont within ticks showed that CLB specifically colonized the ovaries of female Amblyomma cajennense (42) and H. longicornis ticks (43), which also indicated that CLB is associated with the regulation of tick reproductive fitness (42, 44, 45). By using specific fluorescent foci, CLB were also observed in several tick tissues, including Malpighian tubules, salivary glands, and the midgut (46). In Thailand, the presence of CLB in the Haemaphysalis genus has also been documented (13, 24, 25). Our results added information on CLB in H. lagrangei, H. obesa, and A. testudinarium ticks from Chaiyaphum Province.
The phylogenetic tree based on ML analysis using the 16S rRNA and rpoB genes showed that the detected CLB from both Haemaphysalis and Amblyomma ticks obtained in this study were clustered in the same clade as CLB gene sequences from similar genera. However, some groEL gene sequences of CLB in A. testudinarium ticks (accession numbers MZ173557 and MZ173558) detected in this study formed two separate clades. The groEL gene sequence of CLB in A. testudinarium (accession numbers MZ173557 and MZ173558) clustered with Amblyomma from other countries. Another group clustered with the Ixodes genus. This clade of groEL genes was also close to CLB associated with H. wellingtoni collected from domestic fowl from Trang Province, Thailand (25). The reason that the infection pattern shown by the groEL gene of CLB found in A. testudinarium was grouped with other previously published Coxiella sequences from the other tick genera may be horizontal gene transfer. A few examples have shown that accidental horizontal transmission occurs among host individuals, including during cofeeding. For example, the highly efficient exchange of the rompA gene of Rickettsia conorii israelensis was demonstrated between infected and uninfected Rhipicephalus sanguineus ticks feeding nearby each other on a dog that was not formerly infected with these bacteria (47). Interestingly, the results for the 16S rRNA and rpoB markers in this study revealed that the CLB in A. testudinarium, H. lagrangei, and H. obesa ticks clustered together with known isolates, in contrast to the results found with the groEL marker. On the basis of the groEL phylogenetic analysis and BLAST results in this work, we found three groups of CLB: (1) CLB from A. testudinarium grouped as a sister clade with CLB from I. ricinus; (2) CLB from H. lagrangei was distantly related to CLB from H. wellingtoni; and (3) CLB from A. testudinarium grouped as a sister clade with CLB from Amblyomma ticks from French Guiana and Brazil.
Rickettsia spp. detected in ticks from this study grouped with SFG Rickettsiae, which are pathogenic bacteria. On the basis of the phylogenetic analysis, we showed that Rickettsia sp. detected from H. lagrangei tick (PK16) obtained in this study were clustered in different clades containing the rickettsial genes of Amblyomma ticks. Moreover, Rickettsia sp. detected from the H. lagrangei tick (PK16-MZ490778) grouped with Rickettsia sp. HOT2, which clustered with the R. raoultii clade based on the 17-kDa gene. It has been reported that the presence of HOT2 Rickettsia has been detected in Haemaphysalis ornithophila ticks from Khao Yai National Park, Thailand (48). On the basis of the gltA gene sequence, PK16 grouped within the R. raoultii clade (R. massiliae subgroup). However, the ompA and ompB genes of PK16 could not be sequenced due to the presence of faint bands.
The BLAST analysis showed that the ompA genes of Rickettsia spp. collected from Amblyomma ticks were identical (100% DNA sequence identity) to those from R. tamurae AT-1 in all Rickettsia samples detected in this study. In this scenario, it is possible that the detection of the ompA sequences may have arisen due to PCR bias, resulting in this very surprising finding. An alternative explanation could be that the ompA amplicons were the product of a contaminated reaction. We did not include the phylogenetic analysis of the ompA gene in this study.
On the basis of the gltA, ompB, and sca4 phylogenetic analyses and BLAST results from this work, we found two groups of SFG Rickettsiae: (1) SFG Rickettsiae that grouped as a sister clade with R. tamurae AT-1 (belonging to the R. helvetica subgroup) in A. testudinarium and (2) SFG Rickettsiae that was distantly related to R. rhipicephali 3-7-female 6-CWPP (belonging to the R. massiliae subgroup) in A. testudinarium. The pathogenic roles of these bacteria need to be studied further.
Conclusions
From the results of the groEL phylogenetic analysis, CLB clades were found to group as a sister clades to CLB from I. ricinus, CLB from H. wellingtoni, and CLB from Amblyomma ticks from French Guiana and Brazil. In addition, on the basis of the gltA, ompB, and sca4 phylogenetic analyses, SFG Rickettsiae formed two groups: a sister clade to R. tamurae AT-1 (belonging to the R. helvetica subgroup) and a clade distantly related to R. rhipicephali 3-7-female 6-CWPP (belonging to the R. massiliae subgroup). This study demonstrates the diversity of CLB and Rickettsia bacteria with their host ticks, which may act as potential vectors.
Data Availability Statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/supplementary material.
Author Contributions
AA wrote and edited manuscript and planned and designed the research with advice from VB and WT. PU wrote the original draft. VB and WT edited the manuscript. WK analyzed the data of the phylogenetic tree and writing. RS took care of the experiments. All authors contributed to the article and approved the submitted version.
Funding
This research was supported by grants from Mahidol University, the Center of Excellence on Biodiversity (BDC) and the Office of Higher Education Commission (BDC-PG3-163005). This research was also supported by the Thailand Research Fund through the Royal Golden Jubilee PhD Program (Grant No. PHD 0175/2560) to PU.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher's Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Acknowledgments
The authors would like to thank Dr. Surachai Chansri for ticks from his field collections.
References
1. Gehringer H, Schacht E, Maylaender N, Zeman E, Kaysser P, Oehme R, et al. Presence of an emerging subclone of Francisella tularensis holarctica in Ixodes ricinus ticks from South-Western Germany. Ticks Tick Borne Dis. (2013) 4:93–100. doi: 10.1016/j.ttbdis.2012.09.001
2. Cicuttin GL, De Salvo MN, Venzal JM, Nava S. Borrelia spp. in ticks and birds from a protected urban area in Buenos Aires city, Argentina. Ticks Tick Borne Dis. (2019) 10:101282. doi: 10.1016/j.ttbdis.2019.101282
3. Knap N, Zele D, Glinsek Biskup U, Avsic-Zupanc T, Vengust G. The prevalence of Coxiella burnetii in ticks and animals in Slovenia. BMC Vet Res. (2019) 15:15368. doi: 10.1186/s12917-019-2130-3
4. Seo MG, Kwon OD, Kwak D. Molecular detection of Rickettsia raoultii, Rickettsia tamurae, and associated pathogens from ticks parasitizing water deer (Hydropotes inermis argyropus) in South Korea. Ticks Tick Borne Dis. (2021) 12:101712. doi: 10.1016/j.ttbdis.2021.101712
5. Parola P, Paddock CD, Socolovschi C, Labruna MB, Mediannikov O, Kernif T, et al. Update on tick-borne rickettsioses around the world: a geographic approach. Clin Microbiol Rev. (2013) 26:657–702. doi: 10.1128/CMR.00032-13
6. Grassman LI Jr, Sarataphan N, Tewes ME, Silvy NJ, Nakanakrat T. Ticks (Acari: Ixodidae) parasitizing wild carnivores in Phu Khieo wildlife sanctuary, Thailand. J Parasitol. (2004) 90:657–9. doi: 10.1645/GE-3327RN
7. Lee JH, Park HS, Jung KD, Jang WJ, Koh SE, Kang SS, et al. Identification of the spotted fever group rickettsiae detected from Haemaphysalis longicornis in Korea. Microbiol Immunol. (2003) 47:301–4. doi: 10.1111/j.1348-0421.2003.tb03399.x
8. Thu MJ, Qiu Y, Matsuno K, Kajihara M, Mori-Kajihara A, Omori R, et al. Diversity of spotted fever group rickettsiae and their association with host ticks in Japan. Sci Rep. (2019) 9:1500. doi: 10.1038/s41598-018-37836-5
9. Guo WP, Wang YH, Lu Q, Xu G, Luo Y, Ni X, et al. Molecular detection of spotted fever group rickettsiae in hard ticks, northern China. Transbound Emerg Dis. (2019) 66:1587–96. doi: 10.1111/tbed.13184
10. Bhengsri S, Baggett HC, Edouard S, Dowell SF, Dasch GA, Fisk TL, et al. Sennetsu Neorickettsiosis, spotted fever group, and typhus group rickettsioses in three provinces in Thailand. Am J Trop Med Hyg. (2016) 95:43–9. doi: 10.4269/ajtmh.15-0752
11. Wood H, Artsob H. Spotted fever group rickettsiae: a brief review and a Canadian perspective. Zoonoses Public Health. (2012) 59:65–79. doi: 10.1111/j.1863-2378.2012.01472.x
12. Parola P, Cornet JP, Sanogo YO, Miller RS, Thien HV, Gonzalez JP, et al. Detection of Ehrlichia spp, Anaplasma spp, Rickettsia spp, and other eubacteria in ticks from the Thai-Myanmar border and Vietnam. J Clin Microbiol. (2003) 41:1600–8. doi: 10.1128/JCM.41.4.1600-1608.2003
13. Sumrandee C, Baimai V, Trinachartvanit W, Ahantarig A. Molecular detection of Rickettsia, Anaplasma, Coxiella and Francisella bacteria in ticks collected from Artiodactyla in Thailand. Ticks Tick Borne Dis. (2016) 7:678–89. doi: 10.1016/j.ttbdis.2016.02.015
14. Malaisri P, Hirunkanokpun S, Baimai V, Trinachartvanit W, Ahantarig A. Detection of Rickettsia and Anaplasma from hard ticks in Thailand. J Vector Ecol. (2015) 40:262–8. doi: 10.1111/jvec.12163
15. Duron O, Sidi-Boumedine K, Rousset E, Moutailler S, Jourdain E. The importance of ticks in Q fever transmission: what has (and has not) been demonstrated? Trends Parasitol. (2015) 31:536–52. doi: 10.1016/j.pt.2015.06.014
16. Porntip L, Pranee S, Ratchadawan A, Thidarut B, Nadchanan W, Sirintip B, et al. Molecular detection and treatment of tick-borne pathogens in domestic dogs in Khon Kaen, Northeastern Thailand. Southeast Asian J Trop Med Public Health. (2014) 45:1157–66.
17. Pachirat O, Fournier PE, Pussadhamma B, Taksinachanekij S, Lulitanond V, Baggett HC, et al. The first reported cases of Q fever endocarditis in Thailand. Infect Dis Rep. (2012) 4:17–8. doi: 10.4081/idr.2012.e7
18. Suputtamongkol Y, Rolain JM, Losuwanaruk K, Niwatayakul K, Suttinont C, Chierakul W, et al. Q fever in Thailand. Emerg Infect Dis. (2003) 9:1186–88. doi: 10.3201/eid0909.030086
19. Yingst SL, Opaschaitat P, Kanitpun R, Thammasart S, Ekgatat M, Jirathanawat V, et al. Q fever surveillance in ruminants, Thailand, 2012. Emerg Infect Dis. (2013) 19:2056–58. doi: 10.3201/eid1912.130624
20. Doung-Ngern P, Chuxnum T, Pangjai D, Opaschaitat P, Kittiwan N, Rodtian P, et al. Seroprevalence of Coxiella burnetii antibodies among ruminants and occupationally exposed people in Thailand, 2012–2013. Am J Trop Med Hyg. (2017) 96:786–90. doi: 10.4269/ajtmh.16-0336
21. Zhang CM, Li NX, Zhang TT, Qiu ZX, Li Y, Li LW, et al. Endosymbiont CLS-HI plays a role in reproduction and development of Haemaphysalis longicornis. Exp Appl Acarol. (2017) 73:429–38. doi: 10.1007/s10493-017-0194-y
22. Li LH, Zhang Y, Zhu D, Zhou XN. Endosymbionts alter larva-to-nymph transstadial transmission of Babesia microti in Rhipicephalus haemaphysaloides ticks. Front Microbiol. (2018) 9:1415. doi: 10.3389/fmicb.2018.01415
23. Vapniarsky N, Barr BC, Murphy B. Systemic Coxiella-like infection with myocarditis and hepatitis in an eclectus parrot (Eclectus roratus). Vet Pathol. (2012) 49:717–22. doi: 10.1177/0300985811409251
24. Arthan W, Sumrandee C, Hirunkanokpun S, Kitthawee S, Baimai V, Trinachartvanit W, et al. Detection of Coxiella-like endosymbiont in Haemaphysalis tick in Thailand. Ticks Tick Borne Dis. (2015) 6:63–8. doi: 10.1016/j.ttbdis.2014.09.005
25. Trinachartvanit W, Maneewong S, Kaenkan W, Usananan P, Baimai V, Ahantarig A. Coxiella-like bacteria in fowl ticks from Thailand. Parasit Vectors. (2018) 11:670. doi: 10.1186/s13071-018-3259-9
26. Tanskull P, Inlao I. Keys to the adult ticks of Haemaphysalis Koch, 1844, in Thailand with notes on changes in taxonomy (Acari: Ixodoidea: Ixodidae). J Med Entomol. (1989) 26:573–601. doi: 10.1093/jmedent/26.6.573
27. Voltzit OV, Keirans JE. A review of Asian Amblyomma species (Acari, Ixodida, Ixodidae). Acarina. (2002) 10:95–136.
28. Black WC, Piesman J. Phylogeny of hard- and soft-tick taxa (Acari: Ixodida) based on mitochondrial 16S rDNA sequences. Proc Natl Acad Sci U S A. (1994) 91:10034–8. doi: 10.1073/pnas.91.21.10034
29. Williams SG, Sacci JB Jr, Schriefer ME, Andersen EM, Fujioka KK, Sorvillo FJ, et al. Typhus and typhuslike rickettsiae associated with opossums and their fleas in Los Angeles County, California. J Clin Microbiol. (1992) 30:1758–62. doi: 10.1128/jcm.30.7.1758-1762.1992
30. Roux V, Rydkina E, Eremeeva M, Raoult D. Citrate synthase gene comparison, a new tool for phylogenetic analysis, and its application for the rickettsiae. Int J Syst Bacteriol. (1997) 47:252–61. doi: 10.1099/00207713-47-2-252
31. Regnery RL, Spruill CL, Plikaytis BD. Genotypic identification of rickettsiae and estimation of intraspecies sequence divergence for portions of two rickettsial genes. J Bacteriol. (1991) 173:1576–89. doi: 10.1128/jb.173.5.1576-1589.1991
32. Hirunkanokpun S, Ahantarig A, Baimai V, Pramual P, Trinachartvanit W. A new record of Rickettsia japonica in ticks infesting a Burmese ferret-badger in Thailand. Trop. Biomed. (in press).
33. Jiang J, Sangkasuwan V, Lerdthusnee K, Sukwit S, Chuenchitra T, Rozmajzl PJ, et al. Human infection with Rickettsia honei, Thailand. Emerg Infect Dis. (2005) 11:1473–5. doi: 10.3201/eid1109.050011
34. Almeida AP, Marcili A, Leite RC, Nieri-Bastos FA, Domingues LN, Martins JR, et al. Coxiella symbiont in the tick Ornithodoros rostratus (Acari: Argasidae). Ticks Tick Borne Dis. (2012) 3:203–6. doi: 10.1016/j.ttbdis.2012.02.003
35. Duron O, Noel V, McCoy KD, Bonazzi M, Sidi-Boumedine K, Morel O, et al. The recent evolution of a maternally-inherited endosymbiont of ticks led to the emergence of the Q fever pathogen, Coxiella burnetii. PLoS Pathog. (2015) 11:e1004892. doi: 10.1371/journal.ppat.1004892
36. Masuzawa T, Fukui T, Miyake M, Oh HB, Cho MK, Chang WH, et al. Determination of members of a Borrelia afzelii-related group isolated from Ixodes nipponensis in Korea as Borrelia valaisiana. Int J Syst Bacteriol. (1999) 49:1409–15. doi: 10.1099/00207713-49-4-1409
37. Forsman M, Sandstrom G, Sjostedt A. Analysis of 16S ribosomal DNA sequences of Francisella strains and utilization for determination of the phylogeny of the genus and for identification of strains by PCR. Int J Syst Bacteriol. (1994) 44:38–46. doi: 10.1099/00207713-44-1-38
38. Felsenstein J. Evolutionary trees from DNA sequences: a maximum likelihood approach. J Mol Evol. (1981) 17:368–76. doi: 10.1007/BF01734359
39. Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. (1987) 4:406–25.
40. Mediannikov O, Ivanov L, Nishikawa M, Saito R, Sidelnikov YN, Zdanovskaya NI, et al. Molecular evidence of Coxiella-like microorganism harbored by Haemaphysalis concinnae ticks in the Russian Far East. Ann N Y Acad Sci. (2003) 990:226–8. doi: 10.1111/j.1749-6632.2003.tb07367.x
41. Jasinskas A, Zhong J, Barbour AG. Highly prevalent Coxiella sp. bacterium in the tick vector Amblyomma americanum. Appl Environ Microbiol. (2007) 73:334–6. doi: 10.1128/AEM.02009-06
42. Machado-Ferreira E, Dietrich G, Hojgaard A, Levin M, Piesman J, Zeidner NS, et al. Coxiella symbionts in the Cayenne tick Amblyomma cajennense. Microb Ecol. (2011) 62:134–42. doi: 10.1007/s00248-011-9868-x
43. Wang M, Zhu D, Dai J, Zhong Z, Zhang Y, Wang J. Tissue localization and variation of major symbionts in Haemaphysalis longicornis, Rhipicephalus haemaphysaloides, and Dermacentor silvarum in China. Appl Environ Microbiol. (2018) 84:e00029–18. doi: 10.1128/AEM.00029-18
44. Lalzar I, Friedmann Y, Gottlieb Y. Tissue tropism and vertical transmission of Coxiella in Rhipicephalus sanguineus and Rhipicephalus turanicus ticks. Environ Microbiol. (2014) 16:3657–68. doi: 10.1111/1462-2920.12455
45. Guizzo MG, Parizi LF, Nunes RD, Schama R, Albano RM, Tirloni L, et al. A Coxiella mutualist symbiont is essential to the development of Rhipicephalus microplus. Sci Rep. (2017) 7:17554. doi: 10.1038/s41598-017-17309-x
46. Buysse M, Plantard O, McCoy KD, Duron O, Menard C. Tissue localization of Coxiella-like endosymbionts in three European tick species through fluorescence in situ hybridization. Ticks Tick Borne Dis. (2019) 10:798–804. doi: 10.1016/j.ttbdis.2019.03.014
47. Zemtsova G, Killmaster LF, Mumcuoglu KY, Levin ML. Co-feeding as a route for transmission of Rickettsia conorii israelensis between Rhipicephalus sanguineus ticks. Exp Appl Acarol. (2010) 52:383–92. doi: 10.1007/s10493-010-9375-7
Keywords: Coxiella-like bacteria, Rickettsia, Haemaphysalis, Amblyomma, tick
Citation: Usananan P, Kaenkan W, Sudsangiem R, Baimai V, Trinachartvanit W and Ahantarig A (2022) Phylogenetic Studies of Coxiella-Like Bacteria and Spotted Fever Group Rickettsiae in Ticks Collected From Vegetation in Chaiyaphum Province, Thailand. Front. Vet. Sci. 9:849893. doi: 10.3389/fvets.2022.849893
Received: 06 January 2022; Accepted: 17 February 2022;
Published: 06 April 2022.
Edited by:
Nicola Pugliese, University of Bari Aldo Moro, ItalyReviewed by:
Donato Antonio Raele, Experimental Zooprophylactic Institute of Puglia and Basilicata (IZSPB), ItalyZhijun Yu, Hebei Normal University, China
Copyright © 2022 Usananan, Kaenkan, Sudsangiem, Baimai, Trinachartvanit and Ahantarig. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Arunee Ahantarig, YXJ1bmVlLmFoYUBtYWhpZG9sLmFjLnRo
 Pawiga Usananan
Pawiga Usananan Warissara Kaenkan
Warissara Kaenkan Ronnayuth Sudsangiem
Ronnayuth Sudsangiem Visut Baimai
Visut Baimai Wachareeporn Trinachartvanit
Wachareeporn Trinachartvanit Arunee Ahantarig
Arunee Ahantarig