- 1Department of Histology and Embryology, Shandong First Medical University & Shandong Academy of Medical Sciences, Taian, China
- 2Department of Pathophysiology, Shandong First Medical University & Shandong Academy of Medical Sciences, Taian, China
- 3Department of Neurology, People’s Hospital of Binzhou, Binzhou, China
- 4Center for Cancer Research, National Cancer Institute, Frederick, MD, United States
Myeloid-derived suppressor cells (MDSCs) are a heterogeneous population of immature myeloid cells that accumulate in tumor-bearing hosts to reduce T cells activity and promote tumor immune escape in the tumor microenvironment (TME). The immune system in the TME can be stimulated to elicit an anti-tumor immune response through immunotherapy. The main theory of immunotherapy resides on the plasticity of the immune system and its capacity to be re-educated into a potent anti-tumor response. Thus, MDSCs within the TME became one of the major targets to improve the efficacy of tumor immunotherapy, and therapeutic strategies for tumor MDSCs were developed in the last few years. In the article, we analyzed the function of tumor MDSCs and the regulatory mechanisms of agents targeting MDSCs in tumor immunotherapy, and reviewed their therapeutic effects in MDSCs within the TME. Those data focused on discussing how to promote the differentiation and maturation of MDSCs, reduce the accumulation and expansion of MDSCs, and inhibit the function, migration and recruitment of MDSCs, further preventing the growth, invasion and metastasis of tumor. Those investigations may provide new directions for cancer therapy.
Introduction
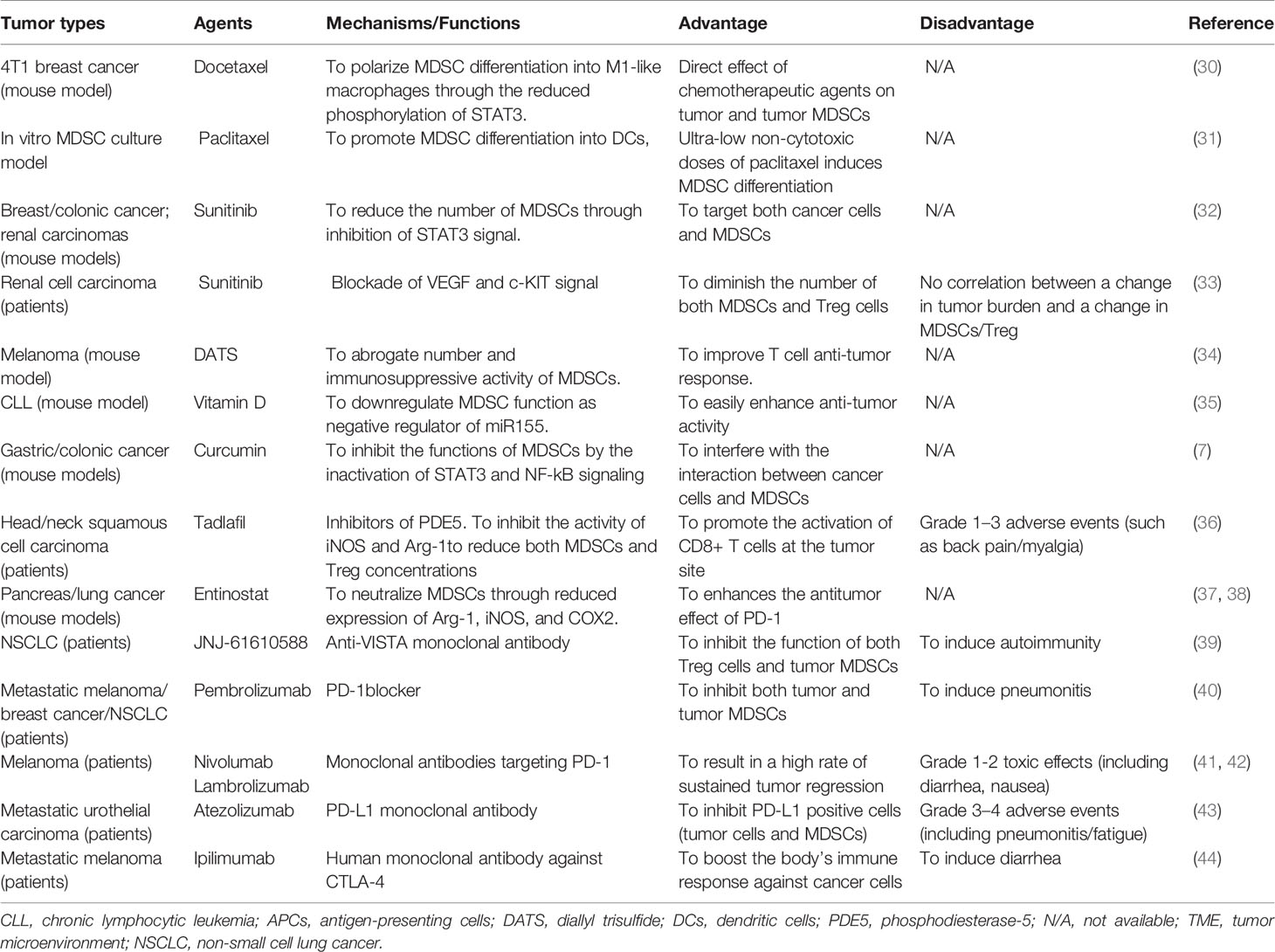
Myeloid-derived suppressor cells (MDSCs), a population of immature myeloid cells with immunosuppressive roles in tumor-bearing models or patients with tumors, have been recognized as the major suppressor of the anti-tumor response (1, 2). The underlying mechanism and function of MDSCs in the TME have been studied by our team and other scientists (3, 4). Clinical data revealed that the high-level circulating MDSCs in patients with cancers correlated with clinical stage, metastatic burden, and the resistance to both chemotherapy and immunotherapy. MDSCs were also recognized as one of the major obstacles in the treatment of cancer, especially for tumor immunotherapy. Recently, epigenetic regulation of the biologic behavior of MDSCs had emerged as a promising tool in cancer therapy (5). Multiple strategies used to target these cells were investigated to determine if the immunosuppressive effects of MDSCs can be decreased or eliminated in order to improve the efficacy of anti-cancer immunotherapy. Even though no therapeutic drugs specifically targeting MDSCs had been approved to be used in clinical treatment, the effects of some agents had been evaluated in tumor mouse models and patients with cancer (6–9). Their roles were involved in promoting the differentiation and maturation of MDSCs, reducing the accumulation and expansion of MDSCs, preventing the migration and recruitment of MDSCs into tumor sites or/and metastasis areas, and impeding the suppressive function and activity of MDSCs (9). In this review, we summarized the data about the roles and functional mechanisms of those agents targeting MDSCs, further providing novel therapeutic strategies for clinical cancer treatment.
Origin and Phenotype of MDSCs
Myeloid precursor cells, derived from hematopoietic stem cells (HSCs) in the bone marrow, developed into immature myeloid cells (IMCs) under physiological conditions. IMCs were differentiated into mature macrophages, dendritic cells (DCs), and granulocytes further. The different progenitor cells that formed the population, demonstrated a broad range of morphology and functional capacity. In contrast, in pathological conditions, such as inflammation, tumors, infections, or autoimmune diseases, there was the dramatic expansion of IMCs with the same phenotype and immune-suppressive activity, resulting in the further differentiation of IMCs into a large number of MDSCs in various tissues (10). Those MDSCs with potent immune-suppressive activity acted as negative regulators of those immune responses (11). Our data illustrated that during chronic process when inflammation transition into cancer, MDSCs downregulated anti-tumor immune responses by modulating cytokine production of macrophages and up-regulated the expression of immune-suppressive factors, such as arginase 1 (Arg-1) and inducible nitric oxide synthase (iNOS) (12, 13). Additionally, tumor MDSCs also blocked T cell anti-tumor response through the increase in the production of reactive oxygen species (ROS)/nitrogen (RNS) (14).
MDSCs lacked clear surface markers unlike monocytes, macrophages and DCs. According to surface markers, MDSCs were divided into two subsets, granulocytic/polymorphonuclear MDSCs (G-MDSCs/PMN-MDSCs) which their phenotype and morphology were similar to neutrophils, and the phenotype and morphology of monocytic MDSCs (M-MDSCs) were similar to monocyte. In tumor bearing-mice, MDSCs were characterized by co-expression of CD11b and Gr-1, which were further divided into two subtypes: CD11b+Ly6G+Ly-6Chigh monocytic MDSCs (M-MDSCs) and CD11b+Ly-6G+Ly-6Clow polymorphonuclear MDSCs (PMN-MDSCs), which were usually present in bone marrow, peripheral blood, spleen, liver, lung or various organs (14). In Human, M-MDSCs were defined as CD11b+CD14+HLA-DR−/loCD15− while PMN-MDSCs were defined as CD11b+CD14−CD15+ or CD11b+CD14−CD66b+ (4). Those phenotypes and function of MDSCs had been shown in our recent articles (3, 15). Both PMN-MDSCs and M-MDSCs had different immunosuppressive mechanisms (16).
Regulatory Mechanism of MDSCs Within the TME
One of the main features of MDSCs was to exhibit immunosuppression activity which was involved in multiple mechanisms or/and factors including signal transducer and an activator of transcription (STATs) (such as STAT1,STAT3, and STAT6) (17), some cytokines (IFN-γ, IL-10, IL-6, GM-CSF), and special molecules (PGE2, S100 protein, and LPS, etc.) (18). MDSCs also hindered the anti-tumor roles of many immune cells in the immune system, such as Natural Killer (NK) cells, B cells and T cells. And the inhibition of T cell function was most important for evaluating the activity of MDSCs. In TME, MDSCs depleted the essential nutrients of T cells through STAT/MyD88 signaling pathway to up-regulate metabolic enzymes e.g. Arg-1, iNOS, which reduced the expression of L-arginin, a substance necessary for T cell activation and proliferation (17) (Figure 1). Thus, L-arginin metabolism played a key role in the immunosuppressive activity of MDSCs by altering the mRNA transcription (19). L-arginin was metabolized by inducible iNOS, generating citrulline and NO to suppress T cell activation and decrease MHC II molecular expression on antigen-presenting cells (APCs), and further inducing T cell apoptosis (18). The productions of ROS/RNS which MDSCs produced, were also immunosuppressive necessary factors. The action of NO and O2− produced peroxynitrite (PNT), which directly impaired T cell activation by nitrating T cell receptors and reducing the reactivity of MHC antigen complexes. PNT reduced the integration of MHC I molecules with antigenic peptides on tumor cells and nitrified T cell-specific chemokines to prevent T cell migration (17, 19–21) (Figure 1).
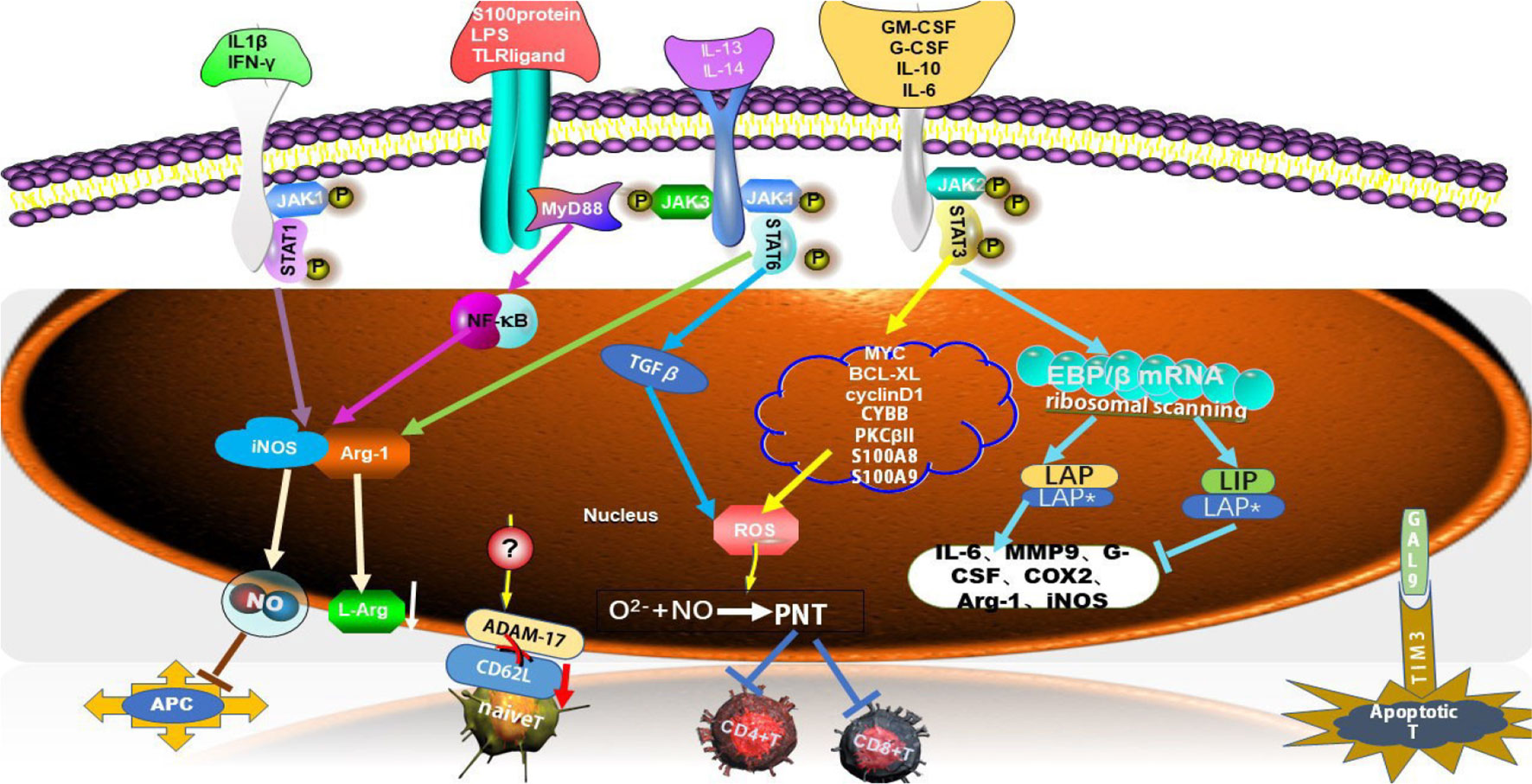
Figure 1 Multiple MDSC-mediated immunosuppressive mechanisms. MDSCs suppressed T cell function through multiple mechanisms. Several factors were involved in triggering signaling pathway, such as STAT1, STAT3, STAT6 and MyD88, which led to high expression level of immunosuppressive factors, such as Arg-1, iNOS, ROS, NO, which suppress T cells response. C/EBPβ was one of the family of CCAAT/enhancer-binding proteins (C/EBPs) which belonged to transcription factors (TFs), which had three isoforms, LAP*, LAP and LIP. S100A8 and S100A9 along with gp91phox (also known as CYBB) were part of the NADPH oxidase (NOX) complex that was responsible for the increased production of reactive oxygen species (ROS) in MDSCs. ADAM17, disintegrin and metalloproteinase domain 17; JAK, Janus kinase; STAT, signal transducer and activator of transcription; PNT, peroxynitrate; COX2, cyclooxygenase-2; APC, antigen-presenting cell; MMP9, matrix metalloproteinase 9; MyD88, myeloid differential protein-88; G-CSF, granulocyte colony-stimulating factor; GM-CSF, granulocyte-macrophage colony stimulating; BCL-XL, B-cell lymphoma XL; GAL9, galectin 9; TIM3, T cell immunoglobulin and mucin domain-containing protein 3.
Apart from the suppression roles on T cells, MDSCs also induced T cells apoptosis and blocked T cell migration which was essential for T cell responses. Galectin 9 expressed on MDSCs bond to T cell immunoglobulin and mucin domain-containing protein 3 (TIM3) on lymphocytes and induced T cell apoptosis (19) (Figure 1). MDSCs directly down-regulated the expression of CD62L on naïve T cells through the expression of TNF-α-converting enzyme (TACE/ADAM17) on MDSCs to hamper the homing of naïve T cells to lymph nodes, leading to the reduced number of those cells (22).
MDSCs and regulatory T (Treg) cells are major components of the TME. Both cell types expanded in tumor models or patients with cancer and promoted T cell dysfunction that in turn facilitated tumor progression (23). Recent studies revealed that MDSCs promoted the development and induction of Treg cells to enhance their suppressive roles on T cells (24). In Ret transgenic mouse models (melanoma model), tumor M-MDSCs also drove the recruitment of CCR5+ Treg cells through producing chemokines such as CCL3, CCL4 and CCL5, further accelerating tumor metastasis (25) (Figure 1).
Immunotherapy Through Targeting Tumor MDSCs
The suppressive roles of MDSCs on both T cells response and the functions of multiple types of cells are critical for the anti-tumor immune response. MDSCs accumulated within the TME were recognized as a major obstacle for tumor immunotherapy (3, 15). Therefore, there are increasing evidences to show that it may be novel therapy strategies for tumor to identify inhibitory factors and find therapeutic ways on tumor MDSC. In the review, we summarized recent data about therapeutic agents and methods on tumor MDSCs, based on the functional characterization of MDSCs.
Promote the Differentiation and Maturation of MDSCs
Within the TME, more IMCs were differentiated into a large number of MDSCs, whereas further differentiation of MDSCs into mature macrophage or DCs was restrained. The agents that promoted differentiation and maturation of MDSCs had been investigated as potential therapeutic strategies to reduce or eliminate MDSCs. Those agents, including all-trans retinoic acid (ATRA), IL-12, RUNX1, and CpG oligonucleotides drove MDSCs differentiation into mature myeloid cells (26–29).
Both Docetaxel and Paclitaxel treated breast cancer effectively, as the two clinical representatives of the new class of taxane drugs. Recent reports demonstrated that Docetaxel promoted MDSCs differentiation into M1-like macrophages with anti-tumor activity by reducing the phosphorylation levels of STAT3 in 4T1 breast tumor-bearing mice (30). And Paclitaxel promoted tumor MDSCs differentiation into mature DCs in a TLR4-independent manner (31) (Table 1).
The differentiation and function of MDSCs may be regulated through microRNA (miRNAs), Long non-coding RNAs (LncRNAs) and epigenetic modifying factors. Our recent data demonstrated that the differentiation and maturation of tumor MDSCs were mediated miRNAs which participated in regulating cell proliferation, differentiation, and maturation (45). The overexpression of miR-17 family members, such as miR-17-5p, miR-20a, and miR-106a in human progenitor cells repressed AML1 by binding to its promoter to down-regulate M-CSFR which induced MDSCs differentiation. MiR-223 remarkably prevented the differentiation of IMCs into MDSCs in the presence of tumor-associated factors by targeting myocyte enhancer factor 2C (MEF2C). MiR-142-3p could limit the generation of MDSCs during tumor-induced myelopoiesis by modulating STAT3 and CCAAT/enhancer-binding protein β (C/EBPβ) signal pathways (45, 46) (Figure 2).
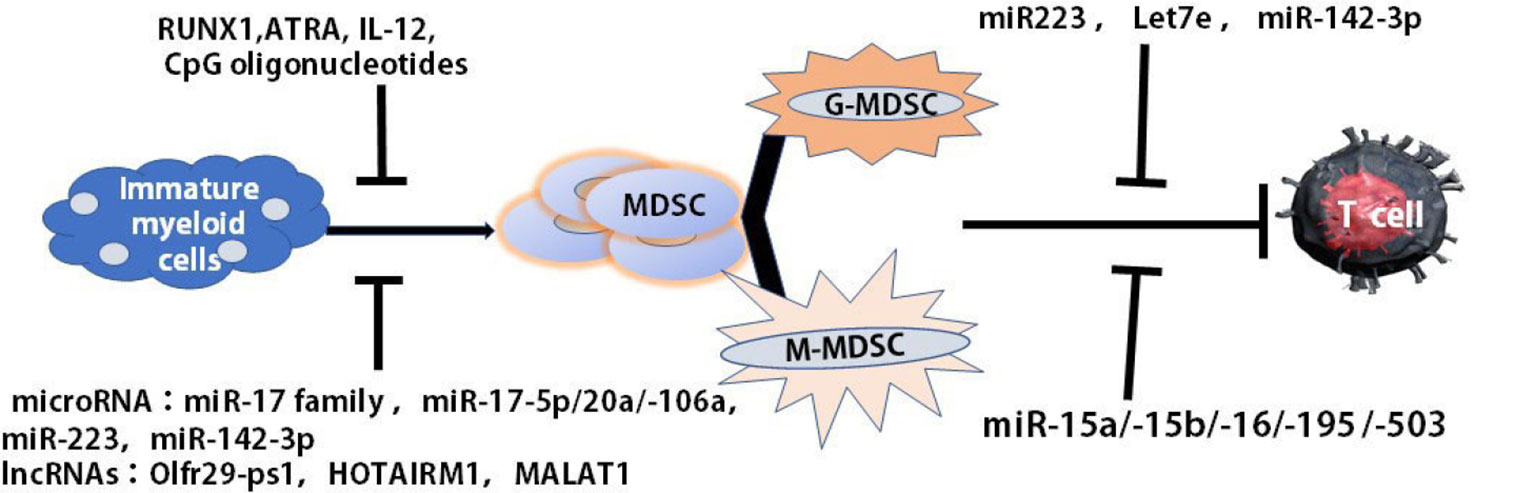
Figure 2 Multiple agents regulated the maturation and differentiation of MDSCs. The differentiation and maturation of MDSCs were regulated by multiple agents, such as RUNX1, ATRA, IL-12, CCL5, and CpG oligonucleotides as well as miRNA, lncRNA. ATRA, all-trans retinoic acid (ATRA).
LncRNAs, more than 200 introns, were intergenic, intronic and natural antisense transcripts, or transcribed from divergent enhancers and promoters (47). They modified chromatin, adjusted the networks of genetic and signal pathways in the pathogenesis of cancer and played a critical role in the regulation of the function and development of myeloid cells (48, 49). Some of the pseudogene transcripts could function as LncRNAs to regulate related gene expression by different mechanisms (50) (Figure 2). Olfr29-ps1, a LncRNA pseudogene, expressed highly in MDSCs, downregulated miR-214-3p to promote the differentiation of MO-MDSCs through IL6-mediated N6-methyladenosine (m6A) modification manner (51). Metastasis associated lung adenocarcinoma transcript 1 (MALAT1), as a nuclear intergenic of LncRNA, whose expression was involved in the differentiation of MDSC-like cells. The reduced expression of MALAT1 in the patients with lung cancer led to the increased proportions of MDSCs, indicating that MALAT1 could prevent the differentiation of tumor MDSCs (52) (Figure 2). HOXA transcript antisense RNA myeloid-specific 1 (HOTAIRM1), an intergenic lncRNA localized between homeobox (HOX)A1 and HOXA2 genes, was expressed preferentially in the myeloid lineage and was a key regulator that targets HOXA1 during myeloid cell development. HOTAIRM1 promoted the maturation of MDSCs via inducing HOXA1 expression in MDSCs to retard lung cancer growth (53). Therefore, multiple agents were found to regulate the maturation and differentiation of MDSCs (Figure 2).
Reduce the Accumulation and Expansion of MDSCs
MDSCs were not present in the circulatory system under normal physiological conditions. However, these cells accumulated in the individuals with cancer. Many agents participated in inducing the decrease of tumor MDSCs. The liver X receptor (LXR) beta-agonist could diminish the populations of both granulocytic and monocytic MDSCs in multiple mouse cancer models by inducing MDSCs apoptosis, which was mediated through the LXR target gene, ApoE, which bond to the LRP8 receptor on the surface of MDSCs (54).
AMP-activated protein kinase (AMPK), an important protein kinase, regulated energy metabolism and innated adaptive immunity by targeting the major signaling pathways. AMPK activation could diminish the expansion and activation of MDSCs through inhibiting multiple signaling pathways, such as STATs and NF-κB pathways (55, 56), and alleviated the nuclear translocation of STAT1 through the increased expression of mitogen-activated protein kinase phosphatase-1 (MKP-1) (57, 58). AMPK also decreased the expansion of MDSCs through attenuating NF-κB activation as well as oxidative and endoplasmic reticulum (ER) stresses, since NF-κB signaling played an important role in the expansion of MDSCs in tumor initiation and progression (59). As a receptor tyrosine kinase inhibitor, Sunitinib was approved by the FDA for the treatment of metastatic renal cell carcinoma (RCC). Recently, some scientists found that Sunitinib was also an immune-modulator, potently reversing tumor MDSC accumulation through STAT3 or C-Kit signal (32, 33) (Table 1).
H2S was an endogenous signaling molecule with a plenty of cellular and molecular targets, which were produced in various mammalian cells and tissues by three principal enzymes: cystathionine-β synthase (CBS), cystathionine-γ lyase (CSE), and 3-mercaptopyruvate sulfurtransferase (3-MST). In melanoma bearing mice, administration of Diallyl Trisulfide (DATS) — H2S donor and the activator of CSE, reduced the number of MDSCs in spleen and blood through the reduction of iNOS expression and NO production. The functional mechanism was involved in the endogenous production of H2S which up-regulated the expression levels of both CSE and CBS to decrease the number of MDSCs in the TME. In addition, DATS also induced MDSCs differentiating into mature APCs (34) (Table 1).
The accumulation of tumor MDSCs was also mediated by miRNA, such as miR-223 and miR-155 (60). miR-155, which was processed from the B-cell integration cluster (BIC), triggered the differentiation of CD4+ T cells into the Th1, Th2, and Th17 pathways and was required for CD8+ T cell responses to acute viral and bacterial challenges (61–63). In the TME, miR-155 promoted the accumulation of functional MDSCs through the SOCS-1 repression, facilitating tumor growth (64). In chronic lymphocytic leukemia (CLL), MDSC accumulation upregulated by exosome miR-155 was disrupted by vitamin D, one negative regulator of miR155, which enhanced anti-tumor activity (35) (Table 1).
Abrogate MDSC Immunosuppression
The inhibition of MDSC immunosuppression function is the major therapeutic target to re-establish T cell activity in cancer immunotherapy. One of the well-known pathways was the JAK-STAT3 signaling pathway which played the crucial role in the regulation of immune responses and induction of the immune escape of tumors through regulating the expression of cytokines and growth factors, further regulating the function of MDSCs (65) (Figure 3). Both STAT3 and NF-κB signaling were essential for the attenuation of Curcumin on the functions of MDSCs (7). AMPK activation phosphorylated the protein to block the activity of Janus kinase 1 (JAK1) to suppress STAT3-dependent gene expression directly, further dampening the suppressive function of MDSCs (66).
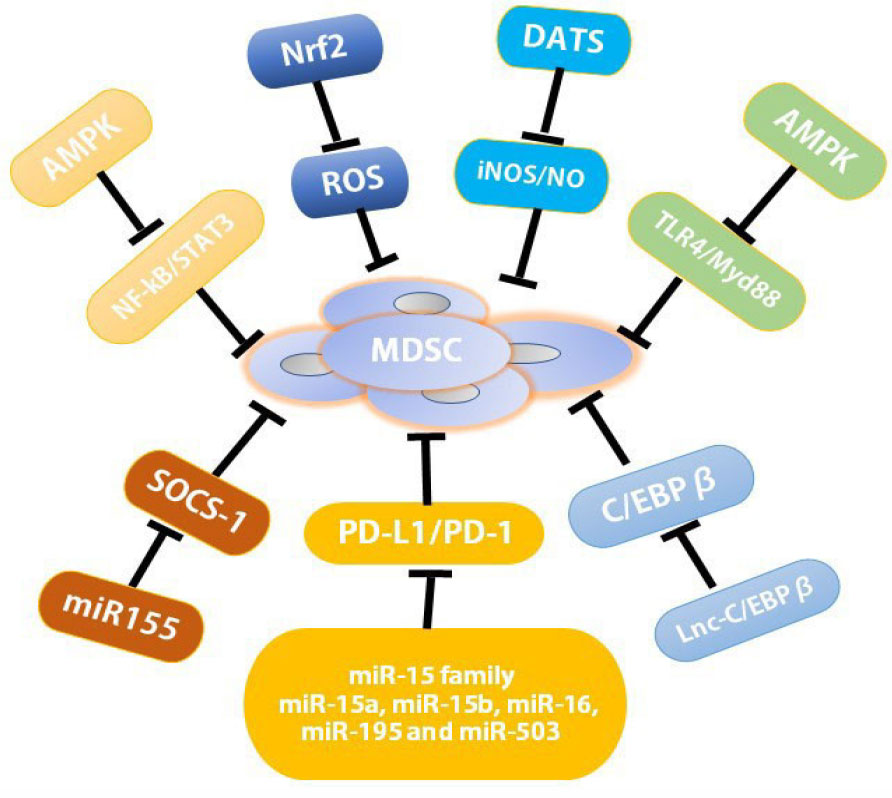
Figure 3 Resistance mechanisms to MDSC immunosuppression. AMPK activation inhibited the TLR4/Myd88 pathway as well as STATs and NF-κB pathways to attenuate the immunosuppressive function of tumor MDSCs; miR-15 family including miR-15a, miR-15b, miR-16, miR-195, and miR-503, down-regulated the suppressive function of MDSCs and/or Tregs within the TME through blocking PD-L1/PD-1 signaling pathway; miR-155 regulated tumor MDSC via the repression of SOCS-1 in MDSCs; Nrf2 activation reduced intracellular ROS production to abrogate MDSC immunosuppression; DATS, as H2S donors and an activator of CSE reduced the number of MDSCs in spleen and blood through the inhibition of iNOS expression and NO production; Lnc-C/EBPβ mastered suppressive functions and differentiation of MDSCs by binding to C/EBPβ (specifically to the LIP isoform). AMPK, AMP-activated protein kinase; C/EBPs, CCAAT/enhancer-binding proteins; miRNA, MicroRNA; LncRNAs, long non-coding RNAs; MALAT1, metastasis associated lung adenocarcinoma transcript 1; DATS, diallyl trisulfide.
Besides reducing the accumulation and expansion of MDSCs, AMPK activation also attenuated the immunosuppressive function of tumor MDSCs through TLR4/Myd88 pathway (67–69). Metformin, one anti-diabetes drug broadly used for the treatment of type 2 diabetes, might act as a new anti-cancer drug since it reduced the immunosuppressive activity of MDSCs by triggering the activation of AMPK (70, 71) (Figure 3). Metformin down-regulated the function of G-MDSCs through AMPK/STAT3 pathways, delaying tumor progression in CT-26 cell colon cancer mouse model (71). In patients with ovarian cancer (OC), Metformin suppressed hypoxia-inducible factor-α (HIF-1α) which blocked CD39/CD73 expression in MDSC to impair MDSC function (72) (Figure 3).
MDSCs upregulated the expression of immune suppressive factors such as ROS, iNOS and Arg-1 to reduce T cells anti-tumor activity. Thus, those factors above became important therapeutic targets. Nuclear factor erythroid 2-related factor 2 (Nrf2) modulated the expression of antioxidant enzymes, including NADPH, NQO1, Hem oxygenase to confer cryoprotection against oxidative stress (73). Selective activation of Nrf2 reduced intracellular ROS production to abrogate MDSC immunosuppression, further reducing tumor metastasis. The promising anticancer drugs had been tested in Phase 1 clinical trials of patients with tumors (73–75). Moreover, Tadalafil was found to reduce activity of iNOS and Arg-1 on both MDSCs and Tregs to disrupt their roles in patients with head/neck squamous cell carcinoma (HNSCC), even though there were low grade of adverse events (such as back pain/myalgia) (36). Entinostat, as class I histone deacetylase inhibitor (HDAC), neutralized MDSCs through reducing the expression of both Arg-1 and iNOS in mouse models of pancreas/lung cancer (37, 38) (Table 1).
Checkpoints were divided into stimulatory and inhibitory forms, which precisely regulated T cell activation. Its balance-maintained self-tolerance and prevented autoimmunity. The upregulation of inhibitory checkpoints led to T cell exhaustion by inhibiting TCR and interactions between co-stimulatory molecules and ligands present on APCs (39). The inhibitory checkpoints included cytotoxic T lymphocyte–associated antigen 4 (CTLA-4), programmed death 1 (PD-1), and V-domain Ig suppressor of T cell activation (VISTA) etc. CTLA-4 were expressed on T cells and bond to CD80/CD86 to mediate the inhibition of Tregs.) (76). PD-1 regulated T cell activation through binding to its ligands, programmed death ligands 1 (PD-L1, B7-H1, or CD274) or 2 (PDL-2, B7-DC, or CD273). The immunotherapy of PD-L1/PD-1 on tumor has been broadly applied. There were increasing evidence that immunosuppression function of MDSCs was mediated by PD-L1 (77). PD-L1 was constitutively expressed and inducible in tumor cells and tumor MDSCs, and bond to PD-1 to suppress T cell activation as an inhibitory ligand. In mouse bladder cancers, PD-L1 expression on tumor-associated MDSCs was associated with the expression of cyclooxygenase-2 (COX2), microsomal prostaglandin E (mPGES1), and prostaglandin E2 (PGE2), as well as their capacity to induce apoptosis of CD8+ T cells (78). The expression of PD-L1 on MDSCs was upregulated in response to multiple microenvironment signals, including hypoxia via HIF1α, and IFN-γ via the STAT1/IRF1 axis (79). In addition, the interaction between PD-L1/PD-1 and miRNAs was required for the function of tumor MDSCs (77). Our recent report revealed that five members of the miR-15 family, including miR-15a, miR-15b, miR-16, miR-195, and miR-503, down-regulated the suppressive function of MDSCs and/or Tregs in the TME through blocking PD-L1/PD-1 signaling pathway (45). The expression of PD-L1 on tumor MDSCs was modulated by the miR-93/106b miRNA cluster of miR-17 family through the STAT3 pathway. Those PD-L1 expression levels on MDSCs could be reduced significantly after the treatment of miR-93 mimics (9, 45). Thus, these PD-1/miRNA/STAT3 pathways provided a new idea of treatment idea for hindering MDSC-associated tumor metastasis (80) (Table 1).
VISTA, another negative checkpoint regulator in the B7 family, enhanced Treg maturation and inhibited T cell activation and hence contributed to the TME (81–83). Its blockade led to the decreases of both Treg cells and MDSCs in the TME, activated DCs, inducing tumor regression in AML mouse models (84–86) and human oral squamous cell carcinoma (OSCC) (87, 88). Recently, the drugs targeting CTLA-4, PD-1, PD-L1, and VISTA were approved to inhibit tumor growth and metastasis, even though these drugs had adverse events for patients with cancer during the treatment (40–44) (Table 1).
LncRNAs also participated in regulating the function of MDSCs through mediating gene transcription. Multiple LncRNAs have been described in myeloid derive cells. Recently, Lnc-C/EBPβ was identified to master suppressive functions of MDSCs by binding to C/EBPβ (specifically to the LIP isoform). C/EBPβ was one of the families of C/EBPs—a group of transcription factors (TFs), which had three isoforms, LAP*, LAP, and LIP, with the former one mainly functioning as transcriptional activator of the expression of immunosuppressive genes such as Arg-1, NOS2, NOX2, or COX2 (89). After exogenous Lnc-C/EBPβ treatment, these immunosuppressive factors above were decreased in quantity, indicating that Lnc-c/EBPβ may play a negative regulatory role in the immunosuppressive function of MDSCs (90) (Figure 3). LncRNA Pvt1 expression in G-MDSCs was upregulated under hypoxia which was a typical feature within the TME. LncRNA Pvt1 knockdown led to the decreased suppression of G-MDSCs, partially restoring T cell antitumor responses (91). In humans, the overexpression of LncRNA PVT1 was tightly associated with a variety of cancer types, including hepatocellular carcinoma, gastric cancer, esophageal cancer, and acute myeloid leukemia (92–97) (Figure 3).
Prevent the Migration and Recruitment of MDSCs
MDSCs exhibited their immunosuppressive activity mainly within the TME. Therefore, the intensive investigations were also conducted to block the migration of MDSC to the tumor sites. PGE2 was involved in tumor angiogenesis and progression via the recruitment of MDSCs (98–100). The inactivation of COX‐2/PGE2 signaling succeeds in reducing MDSC recruitment to retard tumor growth (101). CSF‐1R, a tyrosine kinase receptor, was also involved in the migration of MDSCs (8). Treatments targeting the receptor or its ligand CSF‐1R/CSF‐1 were found to prevent MDSC recruitment to the tumor site to improve T‐cell activity. In addition, anti-glycan antibodies designed to target the receptor for advanced glycation end products (RAGE) were revealed to prevent the recruitment of MDSCs to cancer areas through the S100A8/A9 feedback loop (8).
The chemokines and chemokine receptors played important roles in the migration of tumor MDSCs. The chemokine receptors on MDSCs as therapeutic targets have been used to prevent the recruitment of MDSCs to the tumor sites or metastatic areas. MDSCs were driven into the TME through the chemokines receptor CCR5 expressed on MDSCs with the help of the ligands CCL3, CCL4, and CCL5 (102). In melanoma mouse models and human patients, MDSCs that expressed CCR5 were found to have more potent immunosuppressive effect compared to those that did not express CCR5. Blattner et al. demonstrated that the blockade of CCR5 dampened the recruitment and immunosuppressive activity of MDSCs and improved survival rate (103). Elevated level of CCL2 and CCL5 were present in the TME to recruit MDSCs through chemokine receptor CXCR2 (104, 105). CXCR2+ MDSCs promoted tumor expansion and metastasis in breast cancer (106). By targeting CXCR2, MDSCs were diminished to block tumor metastasis, promoting T‐cell infiltration into the tumors and extending survival in pancreatic cancer (107). Metformin reduced CXCL1 secretion in esophageal squamous cell carcinoma (ESCC) cells through enhancing AMPK phosphorylation and inducing Dachshund homolog 1 (DACH1) expression. Knockdown of both AMPK and DACH1 blocked the effect of metformin on MDSC chemotaxis, indicating that AMPK-DACH1-CXCL1 axis played a significant role in Metformin-regulated migration of MDSCs (70). In summary, targeting chemokine receptors on MDSCs could be applied to prevent the migration and accumulation of MDSCs in the TME.
Conclusion and Perspectives
Tumor MDSCs were recognized as a major obstacle for tumor immunotherapy, As those results, many scientists continue looking for the inhibitory products targeting tumor MDSCs and evaluating their effects to develop new therapeutics that improve the efficacy of cancer immunotherapy strategies. While many of the cancer immunotherapies focus on the manipulation of T cells, the therapy targeting MDSCs may provide another idea for anti-tumor treatment.
MDSCs are major functional cells in the TME through multiple mechanisms to facilitate the harnessing of anti-tumor response. The radiolabeled MDSCs imaging approach to visualize MDSCs migration and tumor homing in vivo is used to further analyze the effects of cancer therapies and immune-therapeutics on MDSC migration and their capacity to infiltrate tumors (108). In addition, MDSCs from patients with tumor are quantified as one marker to differentiate patients with active vs inactive cancer-including radiation necrosis, using liquid biopsies (109). Those results indicated that MDSCs may also one useful clinical biomarker for evaluating radiotherapy effect and identifying tumor metastasis.
Author Contributions
XDG wrote, reviewed, and revised manuscript, figures, and tables. HS prepared the figures and tables. SH reviewed and revised the manuscript. YS revised the manuscript. PQ wrote, reviewed, and revised manuscript, figures, and tables. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by the National Natural Science Foundation of China (grant 81572868); Science Foundation of Shandong (grant ZR2018LC012).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Talmadge JE, Gabrilovich DI. History of myeloid-derived suppressor cells. Nat Rev Cancer (2013) 13:739–52. doi: 10.1038/nrc3581
2. Gabrilovich DI, Bronte V, Chen SH, Colombo MP, Ochoa A, Ostrand-Rosenberg S, et al. The terminology issue for myeloid-derived suppressor cells. Cancer Res (2007) 67:425. doi: 10.1158/0008-5472.CAN-06-3037
3. Qu P, Wang LZ, Lin PC. Expansion and functions of myeloid-derived suppressor cells in the tumor microenvironment. Cancer Lett (2016) 380:253–6. doi: 10.1016/j.canlet.2015.10.022
4. Gabrilovich DI. Myeloid-Derived Suppressor Cells. Cancer Immunol Res (2017) 5:3–8. doi: 10.1158/2326-6066.CIR-16-0297
5. Zhang C, Wang S, Liu Y, Yang C. Epigenetics in myeloid derived suppressor cells: a sheathed sword towards cancer. Oncotarget (2016) 7:57452–63. doi: 10.18632/oncotarget.10767
6. Salminen A, Kauppinen A, Kaarniranta K. AMPK activation inhibits the functions of myeloid-derived suppressor cells (MDSC): impact on cancer and aging. J Mol Med (Berl) (2019) 97:1049–64. doi: 10.1007/s00109-019-01795-9
7. Salminen A, Kaarniranta K, Kauppinen A. Phytochemicals inhibit the immunosuppressive functions of myeloid-derived suppressor cells (MDSC): Impact on cancer and age-related chronic inflammatory disorders. Int Immunopharmacol (2018) 61:231–40. doi: 10.1016/j.intimp.2018.06.005
8. Law AMK, Valdes-Mora F, Gallego-Ortega D. Myeloid-Derived Suppressor Cells as a Therapeutic Target for Cancer. Cells (2020) 9(3):561. doi: 10.3390/cells9030561
9. Consonni FM, Porta C, Marino A, Pandolfo C, Mola S, Bleve A, et al. Myeloid-Derived Suppressor Cells: Ductile Targets in Disease. Front Immunol (2019) 10:949. doi: 10.3389/fimmu.2019.00949
10. Veglia F, Perego M, Gabrilovich D. Myeloid-derived suppressor cells coming of age. Nat Immunol (2018) 19:108–19. doi: 10.1038/s41590-017-0022-x
11. Zhang Y, Qiu Z, Qiu Y, Su T, Qu P, Jia A. Functional Regulation of Ginsenosides on Myeloid Immunosuppressive Cells in the Tumor Microenvironment. Integr Cancer Ther (2019) 18:1534735419886655. doi: 10.1177/1534735419886655
12. Qu P, Yan C, Du H. Matrix metalloproteinase 12 overexpression in myeloid lineage cells plays a key role in modulating myelopoiesis, immune suppression, and lung tumorigenesis. Blood (2011) 117:4476–89. doi: 10.1182/blood-2010-07-298380
13. Qu P, Du H, Wang X, Yan C. Matrix metalloproteinase 12 overexpression in lung epithelial cells plays a key role in emphysema to lung bronchioalveolar adenocarcinoma transition. Cancer Res (2009) 69:7252–61. doi: 10.1158/0008-5472.CAN-09-0577
14. Sawanobori Y, Ueha S, Kurachi M, Shimaoka T, Talmadge JE, Abe J, et al. Chemokine-mediated rapid turnover of myeloid-derived suppressor cells in tumor-bearing mice. Blood (2008) 111:5457–66. doi: 10.1182/blood-2008-01-136895
15. Qu P, Boelte KC, Lin PC. Negative regulation of myeloid-derived suppressor cells in cancer. Immunol Invest (2012) 41:562–80. doi: 10.3109/08820139.2012.685538
16. Thomas AC, Mattila JT. “Of mice and men”: arginine metabolism in macrophages. Front Immunol (2014) 5:479. doi: 10.3389/fimmu.2014.00479
17. Gabrilovich DI, Nagaraj S. Myeloid-derived suppressor cells as regulators of the immune system. Nat Rev Immunol (2009) 9:162–74. doi: 10.1038/nri2506
18. Melero-Jerez C, Ortega MC, Moline-Velazquez V, Clemente D. Myeloid derived suppressor cells in inflammatory conditions of the central nervous system. Biochim Biophys Acta (2016) 1862:368–80. doi: 10.1016/j.bbadis.2015.10.015
19. Gabrilovich DI, Ostrand-Rosenberg S, Bronte V. Coordinated regulation of myeloid cells by tumours. Nat Rev Immunol (2012) 12:253–68. doi: 10.1038/nri3175
20. Sergei Kusmartsev YN, Yoder D. Antigen-Specific Inhibition of CD8+ T Cell Response by Immature Myeloid Cells in Cancer Is Mediated by Reactive Oxygen Species. J Immunol (2004) 172:989–99. doi: 10.4049/jimmunol.172.2.989
21. Gupta K, Nagaraj S, Pisarev V, Kinarsky L, Sherman S, Kang L, et al. Altered recognition of antigen is a novel mechanism of CD8+ tolerance in cancer. Nat Med (2007) 13:828–35. doi: 10.1038/nm1609
22. Hanson EM, Clements VK, Sinha P, Ilkovitch D, Ostrand-Rosenberg S. Myeloid-Derived Suppressor Cells Down-Regulate L-Selectin Expression on CD4+ and CD8+ T Cells. J Immunol (2009) 183:937–44. doi: 10.4049/jimmunol.0804253
23. Lindau D, Gielen P, Kroesen M, Wesseling P, Adema GJ. The immunosuppressive tumour network: myeloid-derived suppressor cells, regulatory T cells and natural killer T cells. Immunology (2013) 138:105–15. doi: 10.1111/imm.12036
24. Fujimura T, Kambayashi Y, Aiba S. Crosstalk between regulatory T cells (Tregs) and myeloid derived suppressor cells (MDSCs) during melanoma growth. Oncoimmunology (2012) 1:1433–4. doi: 10.4161/onci.21176
25. Bruno A, Mortara L, Baci D, Noonan DM, Albini A. Myeloid Derived Suppressor Cells Interactions With Natural Killer Cells and Pro-angiogenic Activities: Roles in Tumor Progression. Front Immunol (2019) 10:771. doi: 10.3389/fimmu.2019.00771
26. Mirza N, Fishman M, Fricke I, Dunn M, Neuger AM, Frost TJ, et al. All-trans-retinoic acid improves differentiation of myeloid cells and immune response in cancer patients. Cancer Res (2006) 66:9299–307. doi: 10.1158/0008-5472.CAN-06-1690
27. Iclozan C, Antonia S, Chiappori A, Chen DT, Gabrilovich D. Therapeutic regulation of myeloid-derived suppressor cells and immune response to cancer vaccine in patients with extensive stage small cell lung cancer. Cancer Immunol Immunother (2013) 62:909–18. doi: 10.1007/s00262-013-1396-8
28. Kusmartsev S, Su Z, Heiser A, Dannull J, Eruslanov E, Kubler H, et al. Reversal of myeloid cell-mediated immunosuppression in patients with metastatic renal cell carcinoma. Clin Cancer Res (2008) 14:8270–8. doi: 10.1158/1078-0432.CCR-08-0165
29. Wang H, Li W, Guo R, Sun J, Cui J, Wang G, et al. An intragenic long noncoding RNA interacts epigenetically with the RUNX1 promoter and enhancer chromatin DNA in hematopoietic malignancies. Int J Cancer (2014) 135:2783–94. doi: 10.1002/ijc.28922
30. Kodumudi KN, Woan K, Gilvary DL, Sahakian E, Wei S, Djeu JY. A novel chemoimmunomodulating property of docetaxel: suppression of myeloid-derived suppressor cells in tumor bearers. Clin Cancer Res (2010) 16:4583–94. doi: 10.1158/1078-0432.CCR-10-0733
31. Michels T, Shurin GV, Naiditch H, Sevko A, Umansky V, Shurin MR. Paclitaxel promotes differentiation of myeloid-derived suppressor cells into dendritic cells in vitro in a TLR4-independent manner. J Immunotoxicol (2012) 9:292–300. doi: 10.3109/1547691X.2011.642418
32. Ko JS, Rayman P, Ireland J, Swaidani S, Li G, Bunting KD, et al. Direct and differential suppression of myeloid-derived suppressor cell subsets by sunitinib is compartmentally constrained. Cancer Res (2010) 70:3526–36. doi: 10.1158/0008-5472.CAN-09-3278
33. Ko JS, Zea AH, Rini BI, Ireland JL, Elson P, Cohen P, et al. Sunitinib mediates reversal of myeloid-derived suppressor cell accumulation in renal cell carcinoma patients. Clin Cancer Res (2009) 15:2148–57. doi: 10.1158/1078-0432.CCR-08-1332
34. De Cicco P, Ercolano G, Rubino V, Terrazzano G, Ruggiero G, Cirino G, et al. Modulation of the functions of myeloid-derived suppressor cells : a new strategy of hydrogen sulfide anti-cancer effects. Br J Pharmacol (2020) 177:884–97. doi: 10.1111/bph.14824
35. Bruns H, Bottcher M, Qorraj M, Fabri M, Jitschin S, Dindorf J, et al. CLL-cell-mediated MDSC induction by exosomal miR-155 transfer is disrupted by vitamin D. Leukemia (2017) 31:985–8. doi: 10.1038/leu.2016.378
36. Weed DT, Vella JL, Reis IM, De la Fuente AC, Gomez C, Sargi Z, et al. Tadalafil reduces myeloid-derived suppressor cells and regulatory T cells and promotes tumor immunity in patients with head and neck squamous cell carcinoma. Clin Cancer Res (2015) 21:39–48. doi: 10.1158/1078-0432.CCR-14-1711
37. Orillion A, Hashimoto A, Damayanti N, Shen L, Adelaiye-Ogala R, Arisa S, et al. Entinostat Neutralizes Myeloid-Derived Suppressor Cells and Enhances the Antitumor Effect of PD-1 Inhibition in Murine Models of Lung and Renal Cell Carcinoma. Clin Cancer Res (2017) 23:5187–201. doi: 10.1158/1078-0432.CCR-17-0741
38. Christmas BJ, Rafie CI, Hopkins AC, Scott BA, Ma HS, Cruz KA, et al. Entinostat Converts Immune-Resistant Breast and Pancreatic Cancers into Checkpoint-Responsive Tumors by Reprogramming Tumor-Infiltrating MDSCs. Cancer Immunol Res (2018) 6:1561–77. doi: 10.1158/2326-6066.CIR-18-0070
39. Popovic A, Jaffee EM, Zaidi N. Emerging strategies for combination checkpoint modulators in cancer immunotherapy. J Clin Invest (2018) 128:3209–18. doi: 10.1172/JCI120775
40. Chen L, Han X. Anti-PD-1/PD-L1 therapy of human cancer: past, present, and future. J Clin Invest (2015) 125:3384–91. doi: 10.1172/JCI80011
41. Weber J, Gibney G, Kudchadkar R, Yu B, Cheng P, Martinez AJ, et al. Phase I/II Study of Metastatic Melanoma Patients Treated with Nivolumab Who Had Progressed after Ipilimumab. Cancer Immunol Res (2016) 4:345–53. doi: 10.1158/2326-6066.CIR-15-0193
42. Hamid O, Robert C, Daud A, Hodi FS, Hwu WJ, Kefford R, et al. Safety and tumor responses with lambrolizumab (anti-PD-1) in melanoma. N Engl J Med (2013) 369:134–44. doi: 10.1056/NEJMoa1305133
43. Rosenberg JE, Hoffman-Censits J, Powles T, van der Heijden MS, Balar AV, Necchi A, et al. Atezolizumab in patients with locally advanced and metastatic urothelial carcinoma who have progressed following treatment with platinum-based chemotherapy: a single-arm, multicentre, phase 2 trial. Lancet (2016) 387:1909–20. doi: 10.1016/S0140-6736(16)00561-4
44. McDermott DF, Hodi FS, O’Day SJ, Weber RW, Sosman JA, Haanen JB, et al. Improved Survival with Ipilimumab in Patients with Metastatic Melanoma. New Engl J Med (2010) 363:711–23. doi: 10.1056/NEJMx100063
45. Su Y, Qiu Y, Qiu Z, Qu P. MicroRNA networks regulate the differentiation, expansion and suppression function of myeloid-derived suppressor cells in tumor microenvironment. J Cancer (2019) 10:4350–6. doi: 10.7150/jca.35205
46. Holtschke T, Lohler J, Kanno Y, Fehr T, Giese N, Rosenbauer F, et al. Immunodeficiency and chronic myelogenous leukemia-like syndrome in mice with a targeted mutation of the ICSBP gene. Cell (1996) 87:307–17. doi: 10.1016/S0092-8674(00)81348-3
47. Leija Montoya G, Gonzalez Ramirez J, Sandoval Basilio J, Serafin Higuera I, Isiordia Espinoza M, Gonzalez Gonzalez R, et al. Long Non-coding RNAs: Regulators of the Activity of Myeloid-Derived Suppressor Cells. Front Immunol (2019) 10:1734. doi: 10.3389/fimmu.2019.01734
48. Chen MT, Lin HS, Shen C, Ma YN, Wang F, Zhao HL, et al. PU.1-Regulated Long Noncoding RNA lnc-MC Controls Human Monocyte/Macrophage Differentiation through Interaction with MicroRNA 199a-5p. Mol Cell Biol (2015) 35:3212–24. doi: 10.1128/MCB.00429-15
49. Zhang X, Lian Z, Padden C, Gerstein MB, Rozowsky J, Snyder M, et al. A myelopoiesis-associated regulatory intergenic noncoding RNA transcript within the human HOXA cluster. Blood (2009) 113:2526–34. doi: 10.1182/blood-2008-06-162164
50. Rapicavoli NA, Qu K, Zhang J, Mikhail M, Laberge RM, Chang HY. A mammalian pseudogene lncRNA at the interface of inflammation and anti-inflammatory therapeutics. Elife (2013) 2:e00762. doi: 10.7554/eLife.00762
51. Shang W, Gao Y, Tang Z, Zhang Y, Yang R. The Pseudogene Olfr29-ps1 Promotes the Suppressive Function and Differentiation of Monocytic MDSCs. Cancer Immunol Res (2019) 7:813–27. doi: 10.1158/2326-6066.CIR-18-0443
52. Zhou Q, Tang X, Tian X, Tian J, Zhang Y, Ma J, et al. LncRNA MALAT1 negatively regulates MDSCs in patients with lung cancer. J Cancer (2018) 9:2436–42. doi: 10.7150/jca.24796
53. Tian X, Ma J, Wang T, Tian J, Zhang Y, Mao L, et al. Long Non-Coding RNA HOXA Transcript Antisense RNA Myeloid-Specific 1-HOXA1 Axis Downregulates the Immunosuppressive Activity of Myeloid-Derived Suppressor Cells in Lung Cancer. Front Immunol (2018) 9:473. doi: 10.3389/fimmu.2018.00473
54. Tavazoie MF, Pollack I, Tanqueco R, Ostendorf BN, Reis BS, Gonsalves FC, et al. LXR/ApoE Activation Restricts Innate Immune Suppression in Cancer. Cell (2018) 172:825–40.e18. doi: 10.1016/j.cell.2017.12.026
55. Millrud CR, Bergenfelz C, Leandersson K. On the origin of myeloid-derived suppressor cells. Oncotarget (2017) 8:3649–65. doi: 10.18632/oncotarget.12278
56. Salminen A, Hyttinen JM, Kaarniranta K. AMP-activated protein kinase inhibits NF-kappaB signaling and inflammation: impact on healthspan and lifespan. J Mol Med (Berl) (2011) 89:667–76. doi: 10.1007/s00109-011-0748-0
57. Meares GP, Qin H, Liu Y, Holdbrooks AT, Benveniste EN. AMP-activated protein kinase restricts IFN-gamma signaling. J Immunol (2013) 190:372–80. doi: 10.4049/jimmunol.1202390
58. Li H, He C, Viollet B, Zou M-H, Xie Z. AMP-activated Protein Kinase Suppresses Vascular Inflammation in vivo by Inhibiting Signal Transducer and Activator of Transcription-1. Diabetes (2015) 64:4285–97. doi: 10.2337/db15-0107
59. Karin M, Greten FR. NF-kappaB: linking inflammation and immunity to cancer development and progression. Nat Rev Immunol (2005) 5:749–59. doi: 10.1038/nri1703
60. Liu Q, Zhang M, Jiang X, Zhang Z, Dai L, Min S, et al. miR-223 suppresses differentiation of tumor-induced CD11b(+) Gr1(+) myeloid-derived suppressor cells from bone marrow cells. Int J Cancer (2011) 129:2662–73. doi: 10.1002/ijc.25921
61. Dudda JC, Salaun B, Ji Y, Palmer DC, Monnot GC, Merck E, et al. MicroRNA-155 is required for effector CD8+ T cell responses to virus infection and cancer. Immunity (2013) 38:742–53. doi: 10.1016/j.immuni.2012.12.006
62. Gallina G, Dolcetti L, Serafini P, De Santo C, Marigo I, Colombo MP, et al. Tumors induce a subset of inflammatory monocytes with immunosuppressive activity on CD8+ T cells. J Clin Invest (2006) 116:2777–90. doi: 10.1172/JCI28828
63. Gracias DT, Stelekati E, Hope JL, Boesteanu AC, Doering TA, Norton J, et al. The microRNA miR-155 controls CD8(+) T cell responses by regulating interferon signaling. Nat Immunol (2013) 14:593–602. doi: 10.1038/ni.2576
64. Chen S, Wang L, Fan J, Ye C, Dominguez D, Zhang Y, et al. Host miR155 promotes tumor growth through a myeloid-derived suppressor cell-dependent mechanism. Cancer Res (2015) 75:519–31. doi: 10.1158/0008-5472.CAN-14-2331
65. Hillmer EJ, Zhang H, Li HS, Watowich SS. STAT3 signaling in immunity. Cytokine Growth Factor Rev (2016) 31:1–15. doi: 10.1016/j.cytogfr.2016.05.001
66. Rutherford C, Speirs C, Williams JJ, Ewart MA, Mancini SJ, Hawley SA, et al. Phosphorylation of Janus kinase 1 (JAK1) by AMP-activated protein kinase (AMPK) links energy sensing to anti-inflammatory signaling. Sci Signal (2016) 9:ra109. doi: 10.1126/scisignal.aaf8566
67. Vaez H, Najafi M, Rameshrad M, Toutounchi NS, Garjani M, Barar J, et al. AMPK activation by metformin inhibits local innate immune responses in the isolated rat heart by suppression of TLR 4-related pathway. Int Immunopharmacol (2016) 40:501–7. doi: 10.1016/j.intimp.2016.10.002
68. Rameshrad M, Soraya H, Maleki-Dizaji N, Vaez H, Garjani A. A-769662, a direct AMPK activator, attenuates lipopolysaccharide-induced acute heart and lung inflammation in rats. Mol Med Rep (2016) 13:2843–9. doi: 10.3892/mmr.2016.4821
69. Hong EH, Chang SY, Lee BR, Kim YS, Lee JM, Kang CY, et al. Blockade of Myd88 signaling induces antitumor effects by skewing the immunosuppressive function of myeloid-derived suppressor cells. Int J Cancer (2013) 132:2839–48. doi: 10.1002/ijc.27974
70. Qin G, Lian J, Huang L, Zhao Q, Liu S, Zhang Z, et al. Metformin blocks myeloid-derived suppressor cell accumulation through AMPK-DACH1-CXCL1 axis. Oncoimmunology (2018) 7:e1442167. doi: 10.1080/2162402X.2018.1442167
71. Xu P, Yin K, Tang X, Tian J, Zhang Y, Ma J, et al. Metformin inhibits the function of granulocytic myeloid-derived suppressor cells in tumor-bearing mice. BioMed Pharmacother (2019) 120:109458. doi: 10.1016/j.biopha.2019.109458
72. Li L, Wang L, Li J, Fan Z, Yang L, Zhang Z, et al. Metformin-Induced Reduction of CD39 and CD73 Blocks Myeloid-Derived Suppressor Cell Activity in Patients with Ovarian Cancer. Cancer Res (2018) 78:1779–91. doi: 10.1158/0008-5472.CAN-17-2460
73. Ohl K, Tenbrock K. Reactive Oxygen Species as Regulators of MDSC-Mediated Immune Suppression. Front Immunol (2018) 9:2499. doi: 10.3389/fimmu.2018.02499
74. Wang YY, Yang YX, Zhe H, He ZX, Zhou SF. Bardoxolone methyl (CDDO-Me) as a therapeutic agent: an update on its pharmacokinetic and pharmacodynamic properties. Drug Des Devel Ther (2014) 8:2075–88. doi: 10.2147/DDDT.S68872
75. Nagaraj S, Youn JI, Weber H, Iclozan C, Lu L, Cotter MJ, et al. Anti-inflammatory triterpenoid blocks immune suppressive function of MDSCs and improves immune response in cancer. Clin Cancer Res (2010) 16:1812–23. doi: 10.1158/1078-0432.CCR-09-3272
76. Sasidharan Nair V, Elkord E. Immune checkpoint inhibitors in cancer therapy: a focus on T-regulatory cells. Immunol Cell Biol (2018) 96:21–33. doi: 10.1111/imcb.1003
77. Lu C, Redd PS, Lee JR, Savage N, Liu K. The expression profiles and regulation of PD-L1 in tumor-induced myeloid-derived suppressor cells. Oncoimmunology (2016) 5:e1247135. doi: 10.1080/2162402X.2016.1247135
78. Prima V, Kaliberova LN, Kaliberov S, Curiel DT, Kusmartsev S. COX2/mPGES1/PGE2 pathway regulates PD-L1 expression in tumor-associated macrophages and myeloid-derived suppressor cells. Proc Natl Acad Sci U S A (2017) 114:1117–22. doi: 10.1073/pnas.1612920114
79. Noman MZ, Desantis G, Janji B, Hasmim M, Karray S, Dessen P, et al. PD-L1 is a novel direct target of HIF-1alpha, and its blockade under hypoxia enhanced MDSC-mediated T cell activation. J Exp Med (2014) 211:781–90. doi: 10.1084/jem.20131916
80. Goze C, Reynes C, Forestier L, Sabatier R, Duffau H. Pilot Study of Whole Blood MicroRNAs as Potential Tools for Diffuse Low-Grade Gliomas Detection. Cell Mol Neurobiol (2018) 38:715–25. doi: 10.1007/s10571-017-0536-7
81. Wang L, Rubinstein R, Lines JL, Wasiuk A, Ahonen C, Guo Y, et al. VISTA, a novel mouse Ig superfamily ligand that negatively regulates T cell responses. J Exp Med (2011) 208:577–92. doi: 10.1084/jem.20100619
82. Lines JL, Pantazi E, Mak J, Sempere LF, Wang L, O’Connell S, et al. VISTA is an immune checkpoint molecule for human T cells. Cancer Res (2014) 74:1924–32. doi: 10.1158/0008-5472.CAN-13-1504
83. Toor SM, Elkord E. Therapeutic prospects of targeting myeloid-derived suppressor cells and immune checkpoints in cancer. Immunol Cell Biol (2018) 96:888–97. doi: 10.1111/imcb.12054
84. Le Mercier I, Chen W, Lines JL, Day M, Li J, Sergent P, et al. VISTA Regulates the Development of Protective Antitumor Immunity. Cancer Res (2014) 74:1933–44. doi: 10.1158/0008-5472.CAN-13-1506
85. Lines JL, Sempere LF, Broughton T, Wang L, Noelle R. VISTA is a novel broad-spectrum negative checkpoint regulator for cancer immunotherapy. Cancer Immunol Res (2014) 2:510–7. doi: 10.1158/2326-6066.CIR-14-0072
86. Wang L, Le Mercier I, Putra J, Chen W, Liu J, Schenk AD, et al. Disruption of the immune-checkpoint VISTA gene imparts a proinflammatory phenotype with predisposition to the development of autoimmunity. Proc Natl Acad Sci U S A (2014) 111:14846–51. doi: 10.1073/pnas.1407447111
87. Wu L, Deng WW, Huang CF, Bu LL, Yu GT, Mao L, et al. Expression of VISTA correlated with immunosuppression and synergized with CD8 to predict survival in human oral squamous cell carcinoma. Cancer Immunol Immunother (2017) 66:627–36. doi: 10.1007/s00262-017-1968-0
88. Wang L, Jia B, Claxton DF, Ehmann WC, Rybka WB, Mineishi S, et al. VISTA is highly expressed on MDSCs and mediates an inhibition of T cell response in patients with AML. Oncoimmunology (2018) 7:e1469594. doi: 10.1080/2162402X.2018.1469594
89. Wang W, Xia X, Mao L, Wang S. The CCAAT/Enhancer-Binding Protein Family: Its Roles in MDSC Expansion and Function. Front Immunol (2019) 10:1804. doi: 10.3389/fimmu.2019.01804
90. Gao Y, Sun W, Shang W, Li Y, Zhang D, Wang T, et al. Lnc-C/EBPbeta Negatively Regulates the Suppressive Function of Myeloid-Derived Suppressor Cells. Cancer Immunol Res (2018) 6:1352–63. doi: 10.1158/2326-6066.CIR-18-0108
91. Zheng Y, Tian X, Wang T, Xia X, Cao F, Tian J, et al. Long noncoding RNA Pvt1 regulates the immunosuppression activity of granulocytic myeloid-derived suppressor cells in tumor-bearing mice. Mol Cancer (2019) 18:61. doi: 10.1186/s12943-019-0978-2
92. Zheng X, Hu H, Li S. High expression of lncRNA PVT1 promotes invasion by inducing epithelial-to-mesenchymal transition in esophageal cancer. Oncol Lett (2016) 12:2357–62. doi: 10.3892/ol.2016.5026
93. Zhao J, Du P, Cui P, Qin Y, Hu C, Wu J, et al. LncRNA PVT1 promotes angiogenesis via activating the STAT3/VEGFA axis in gastric cancer. Oncogene (2018) 37:4094–109. doi: 10.1038/s41388-018-0250-z
94. Zhang S, Zhang G, Liu J. Long noncoding RNA PVT1 promotes cervical cancer progression through epigenetically silencing miR-200b. Apmis (2016) 124:649–58. doi: 10.1111/apm.12555
95. Xu Y, Luo X, He W, Chen G, Li Y, Li W, et al. Long Non-Coding RNA PVT1/miR-150/ HIG2 Axis Regulates the Proliferation, Invasion and the Balance of Iron Metabolism of Hepatocellular Carcinoma. Cell Physiol Biochem (2018) 49:1403–19. doi: 10.1159/000493445
96. Tian Z, Cao S, Li C, Xu M, Wei H, Yang H, et al. LncRNA PVT1 regulates growth, migration, and invasion of bladder cancer by miR-31/ CDK1. J Cell Physiol (2019) 234:4799–811. doi: 10.1002/jcp.27279
97. Cui M, You L, Ren X, Zhao W, Liao Q, Zhao Y. Long non-coding RNA PVT1 and cancer. Biochem Biophys Res Commun (2016) 471:10–4. doi: 10.1016/j.bbrc.2015.12.101
98. Eruslanov E, Daurkin I, Ortiz J, Vieweg J, Kusmartsev S. Pivotal Advance: Tumor-mediated induction of myeloid-derived suppressor cells and M2-polarized macrophages by altering intracellular PGE(2) catabolism in myeloid cells. J Leukoc Biol (2010) 88:839–48. doi: 10.1189/jlb.1209821
99. Ochoa AC, Zea AH, Hernandez C, Rodriguez PC. Arginase, prostaglandins, and myeloid-derived suppressor cells in renal cell carcinoma. Clin Cancer Res (2007) 13:721s–6s. doi: 10.1158/1078-0432.CCR-06-2197
100. Sinha P, Clements VK, Fulton AM, Ostrand-Rosenberg S. Prostaglandin E2 promotes tumor progression by inducing myeloid-derived suppressor cells. Cancer Res (2007) 67:4507–13. doi: 10.1158/0008-5472.CAN-06-4174
101. Veltman JD, Lambers ME, van Nimwegen M, Hendriks RW, Hoogsteden HC, Aerts JG, et al. COX-2 inhibition improves immunotherapy and is associated with decreased numbers of myeloid-derived suppressor cells in mesothelioma. Celecoxib influences MDSC function. BMC Cancer (2010) 10:464. doi: 10.1186/1471-2407-10-464
102. Weber R, Fleming V, Hu X, Nagibin V, Groth C, Altevogt P, et al. Myeloid-Derived Suppressor Cells Hinder the Anti-Cancer Activity of Immune Checkpoint Inhibitors. Front Immunol (2018) 9:1310. doi: 10.3389/fimmu.2018.01310
103. Blattner C, Fleming V, Weber R, Himmelhan B, Altevogt P, Gebhardt C, et al. CCR5(+) Myeloid-Derived Suppressor Cells Are Enriched and Activated in Melanoma Lesions. Cancer Res (2018) 78:157–67. doi: 10.1158/0008-5472.CAN-17-0348
104. Toh B, Wang X, Keeble J, Sim WJ, Khoo K, Wong WC, et al. Mesenchymal transition and dissemination of cancer cells is driven by myeloid-derived suppressor cells infiltrating the primary tumor. PloS Biol (2011) 9:e1001162. doi: 10.1371/journal.pbio.1001162
105. Katoh H, Wang D, Daikoku T, Sun H, Dey SK, Dubois RN. CXCR2-expressing myeloid-derived suppressor cells are essential to promote colitis-associated tumorigenesis. Cancer Cell (2013) 24:631–44. doi: 10.1016/j.ccr.2013.10.009
106. Zhu H, Gu Y, Xue Y, Yuan M, Cao X, Liu Q. CXCR2(+) MDSCs promote breast cancer progression by inducing EMT and activated T cell exhaustion. Oncotarget (2017) 8:114554–67. doi: 10.18632/oncotarget.23020
107. Steele CW, Karim SA, Leach JDG, Bailey P, Upstill-Goddard R, Rishi L, et al. CXCR2 Inhibition Profoundly Suppresses Metastases and Augments Immunotherapy in Pancreatic Ductal Adenocarcinoma. Cancer Cell (2016) 29:832–45. doi: 10.1016/j.ccell.2016.04.014
108. Hoffmann SHL, Reck DI, Maurer A, Fehrenbacher B, Sceneay JE, Poxleitner M, et al. Visualization and quantification of in vivo homing kinetics of myeloid-derived suppressor cells in primary and metastatic cancer. Theranostics (2019) 9:5869–85. doi: 10.7150/thno.33275
109. Soler DC, Kerstetter-Fogle A, Elder T, Raghavan A, Barnholtz-Sloan JS, Cooper KD, et al. A Liquid Biopsy to Assess Brain Tumor Recurrence: Presence of Circulating Mo-MDSC and CD14+ VNN2+ Myeloid Cells as Biomarkers That Distinguish Brain Metastasis From Radiation Necrosis Following Stereotactic Radiosurgery. Neurosurgery (2020) 88:E67–72. doi: 10.1093/neuros/nyaa334
Keywords: cancer, tumor immunotherapy, tumor microenvironment, myeloid-derived suppressor cells, inhibitory factors
Citation: Gao X, Sui H, Zhao S, Gao X, Su Y and Qu P (2021) Immunotherapy Targeting Myeloid-Derived Suppressor Cells (MDSCs) in Tumor Microenvironment. Front. Immunol. 11:585214. doi: 10.3389/fimmu.2020.585214
Received: 20 July 2020; Accepted: 17 December 2020;
Published: 04 February 2021.
Edited by:
Nicola Giuliani, University of Parma, ItalyReviewed by:
Wei Chen, Stanford University, United StatesMichael Hader, University Hospital Erlangen, Germany
Copyright © 2021 Gao, Sui, Zhao, Gao, Su and Qu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yanping Su, c3UteWFucGluZ0AxNjMuY29t; Peng Qu, cGVuZ3F1amkyMDAwQGdtYWlsLmNvbQ==
 Xidan Gao1
Xidan Gao1 Hongshu Sui
Hongshu Sui Yanping Su
Yanping Su Peng Qu
Peng Qu