- 1Department of Urology, The First People's Hospital of Xiushui, Jiujiang, Jiangxi, China
- 2The First Clinical College, Gannan Medical University, Ganzhou, Jiangxi, China
- 3Department of Otorhinolaryngology, The Affiliated Fengcheng Hospital of Yichun University, Fengcheng, Jiangxi, China
Prostate cancer (PCa) remains a leading cause of cancer-related mortality in men worldwide, primarily due to its propensity for therapy resistance and metastasis. Emerging evidence underscores that exosomes, nanoscale extracellular vesicles, act as critical mediators of intercellular communication within the tumor microenvironment (TME), particularly via the non-coding RNAs (ncRNAs) they transport. These molecules include microRNAs (miRNAs), circular RNAs (circRNAs), and long non-coding RNAs (lncRNAs). Exosomal ncRNAs drive tumor progression, immune evasion, and therapy resistance by reprogramming neighboring stromal cells, immune cells, and malignant cells. This review systematically examines the multifaceted roles of exosomal ncRNAs in remodeling the prostate cancer tumor microenvironment, focusing on their communication between tumor cells, tumor-stromal cells (including immune cells), and within the pre-metastatic niche preceding bone metastasis. We emphasize their mechanisms of action and clinical relevance. These findings position exosomal ncRNAs as central drivers of prostate cancer progression, revealing novel diagnostic and therapeutic opportunities. Future research must address challenges in standardizing exosome isolation techniques, resolving spatiotemporal heterogeneity, and advancing clinical translation to fully realize the potential of exosome-based strategies in precision oncology.
1 Introduction
Prostate cancer (PCa) is the second most prevalent malignant tumor in males globally, primarily affecting individuals aged 45–60, as evidenced by recent epidemiological data (1). As reported by the International Agency for Research on Cancer’s GLOBOCAN database, PCa accounted for over 1.4 million new cases and 375,000 deaths worldwide in 2020 and demonstrates a rising trend (1–3). Notably, managing advanced metastatic and castration-resistant prostate cancer (CRPC) remains a critical clinical challenge (4). While early-stage localized PCa is curable via surgery or radiotherapy (5-year survival rate approaching 100%) (5), recent research evaluating robot-assisted laparoscopic radical prostatectomy reported biochemical recurrence in approximately 10.19% of T3aN0-stage patients within 15.22 months post-surgery (6). However, the 5-year survival rate plummets to 30% once distant metastasis occurs (5). Despite advancements in surgical, radiotherapeutic, and endocrine therapies for PCa, substantial challenges persist. Tumor heterogeneity leads to marked variability in therapeutic responses, particularly to chemotherapy and endocrine interventions (7). Prolonged treatment durations exacerbate therapy-related side effects and complications, adversely affecting patients’ psychological and physiological well-being, treatment adherence, and quality of life (8). These challenges underscore the necessity for deeper mechanistic investigations into PCa pathogenesis to refine therapeutic strategies.
Cancer is widely recognized as a dynamic evolutionary process driven by intricate interactions between tumor cells and the tumor microenvironment (TME) (9, 10). Emerging studies have established that prostate cancer invasiveness is closely linked to its distinctly heterogeneous TME (11, 12). The TME comprises tumor cells, cancer-associated fibroblasts (CAFs), immune cells, vascular endothelial cells, and extracellular matrix (ECM). Non-cellular components, including growth factors, extracellular vesicles, and acellular ECM, collectively form a dynamic, pro-metastatic, and therapy-resistant ecosystem (13, 14). Exosomes, a subtype of extracellular vesicles, have emerged as pivotal mediators of intercellular communication within the TME, owing to their ability to regulate tumor progression via transport of non-coding RNAs (ncRNAs) (15, 16). These nanoscale vesicles, which are actively secreted by cells and range from 30 to 150 nm in diameter, are not merely cellular waste products but originate from multivesicular bodies (MVBs). The endosomal membrane buds inward to form an MVB, which contains numerous intraluminal vesicles; this MVB subsequently fuses with the plasma membrane, releasing these vesicles into the extracellular space as exosomes (17). Consequently, exosomes carry a diverse molecular “cargo” derived from their parent cells, including specific membrane proteins (e.g., CD9, CD63, CD81), cytoplasmic proteins, and nucleic acids (e.g., ncRNAs). Among these components, surface integrins and tissue-specific markers (e.g., PSMA) facilitate the targeted delivery of exosomes to specific tissues (18, 19). Luo et al. demonstrated that tumor-derived exosomes promote prostate cancer angiogenesis and metastasis by delivering phosphoglycerate mutase 1 (PGAM1), which binds γ-actin (ACTG1) (20). Exosomes also play a central role in CRPC progression. For instance, androgen deprivation therapy (ADT) alters exosomal miRNA profiles (e.g., upregulated miR-423-3p), driving the transition from androgen-dependent prostate cancer (ADPC) to CRPC via regulation of AR-V7 and ERG expression (21).
Evidence (22, 23) has shown that exosomes participate in the diagnosis and treatment of prostate cancer, acting as participants in pathophysiological mechanisms, biomarkers in cancer progression, and therapeutic tools. For instance, exosomes promote the progression of prostate cancer by carrying signaling molecules to act on tumor cells. However, growing recognition of ncRNAs has spurred extensive research into the functional roles of exosomal ncRNAs in prostate cancer. Table 1 summarizes the mechanisms by which exosomal ncRNAs regulate prostate cancer progression. As evidenced by the compiled data, exosomal ncRNAs drive prostate cancer progression via diverse mechanisms, including competing endogenous RNA (ceRNA) networks and epigenetic regulation. Despite considerable advances in exosomal ncRNA research, current studies remain predominantly centered on modulating prostate cancer cellular phenotypes (33). Critical gaps persist in understanding their sorting mechanisms, spatiotemporal heterogeneity, and clinical translation challenges. Two prominent examples include the paucity of longitudinal data on dynamic changes in exosomal ncRNA expression throughout disease progression, and the limited evidence for their clinical utility as interventional targets. Furthermore, the complexity of cell-cell communication networks within the TME necessitates stematic categorization to elucidate underlying principles and inform clinical strategies. This review synthesizes findings from the past five years to systematically map ncRNA-mediated cellular interaction networks within the prostate cancer TME. Specifically, it examines the role of exosomal ncRNAs in facilitating communication: (1) between tumor cells; (2) between tumor and stromal cells; (3) between tumor and immune cells; and (4) in shaping the microenvironment of prostate cancer bone metastases (Figure 1). Our objective is to identify novel therapeutic targets and early diagnostic biomarkers while advancing combinatorial treatment approaches. Deciphering the regulatory signaling logic across cell types of exosomal ncRNAs may accelerate biomarker discovery and enable ncRNA-based therapeutic interventions, advancing the long-term goal of precision oncology for advanced prostate cancer.
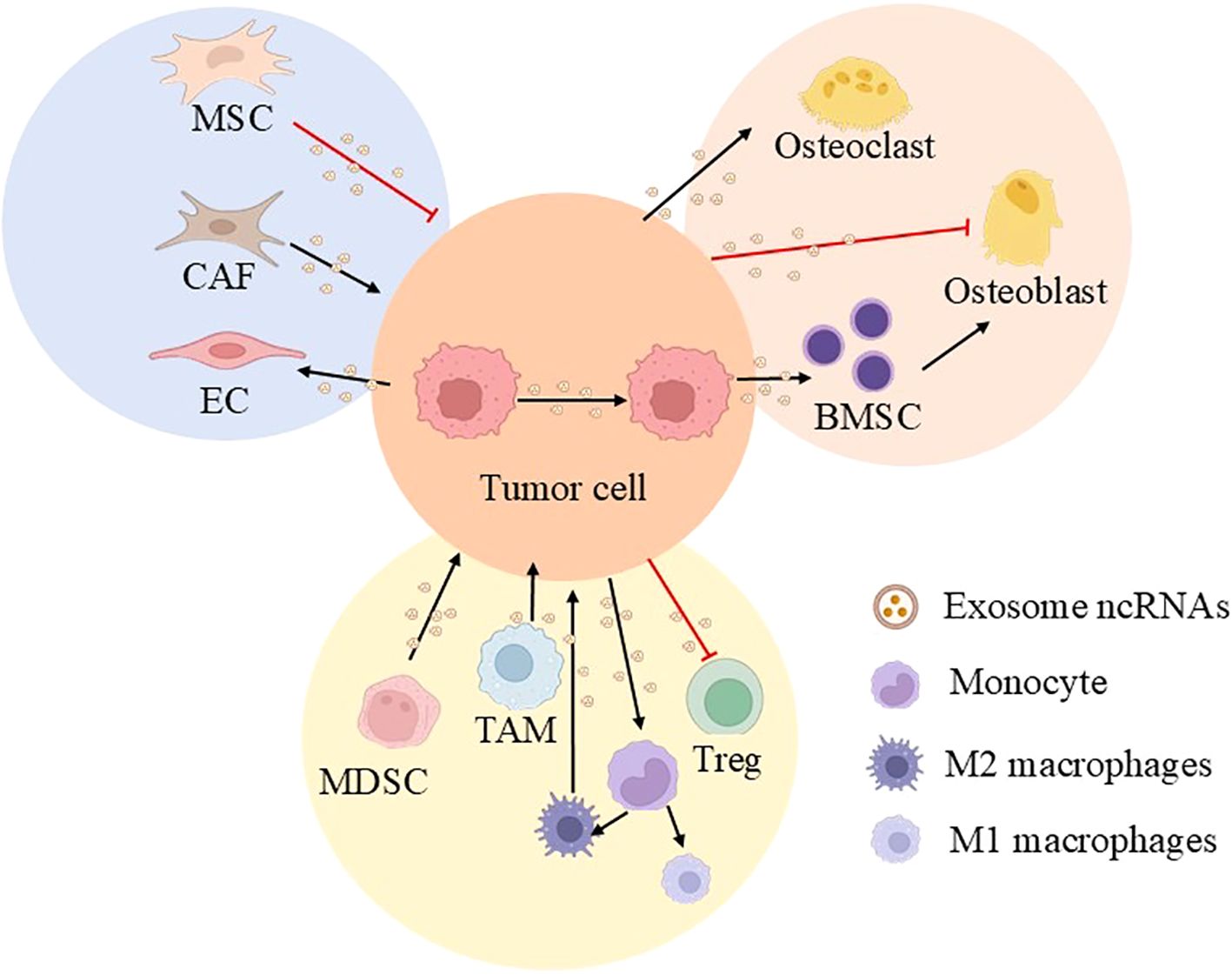
Figure 1. Outline of this review. This article takes the exosomal ncRNA communication in the tumor microenvironment of prostate cancer as the core starting point, systematically examining its functions in tumor-tumor, tumor-stromal, and tumor-immune cell crosstalk, as well as its pivotal role in shaping the bone metastatic niche. (MSC, mesenchymal stem cell; MDSC, myeloid-derived suppressor cell; BMSC, bone marrow mesenchymal stem cell; TAM, tumor-associated macrophage; CAF, cancer-associated fibroblast; Treg, regulatory T cell; EC, endothelial cell).
2 Exosomal ncRNA-mediated crosstalk between tumor cells
Acting as key messengers in intercellular communication, exosomes play a multifaceted role in prostate cancer progression. This section examines the mechanisms by which exosomal ncRNAs mediate cellular crosstalk within the tumor microenvironment, promote tumor heterogeneity, and disseminate chemoresistance. Furthermore, it provides an in-depth exploration of their pivotal role in the development of CRPC and associated therapy resistance.
2.1 Exosome-mediated tumor cell–tumor cell signaling
The TME facilitates interactions between tumor cells and neighboring non-target bystander cells, generating biological phenomena termed the paracrine-like oncogenic influence (34, 35). This confirms that in the TME, interactions exist among tumor cells, further promoting tumor maintenance and development. These intercellular interactions can be achieved through direct cell contact or indirectly via secreted factors, and ncRNAs carried by exosomes play a crucial role in this process (36). Dai et al. (37) demonstrated that exosomal miR-183, upregulated in prostate cancer, enhances proliferation, invasion, and migration of LNCaP cells. Similarly, prostate cancer-derived exosomes transport circRNAs to exert analogous oncogenic effects. For instance, exosomal circKDM4A acts as a molecular sponge for miR-338-3p, alleviating its suppression of CUL4B to activate the ubiquitin-proteasome system, thereby fostering tumor proliferation and therapy resistance (38). Mechanical cues, including fluid shear stress, induce prostate cancer cells to release exosomes enriched with miR-21-5p (39). Uptake of these exosomes by adjacent cancer cells modulates migration-associated signaling pathways, altering cellular morphology and motility. These findings underscore the role of exosomal crosstalk in the TME and elucidate a novel mechanism whereby mechanical cues regulate metastasis via exosomal miRNA signaling.
2.2 Role of exosomes in driving intratumoral heterogeneity
Even within the same tumor, cancer cells exhibit heterogeneity in morphology, invasiveness, and responses to hormones or therapies, a phenomenon termed intratumoral heterogeneity (40, 41). This heterogeneity drives tumor progression and therapeutic failure (42). However, with the continuous deepening of research, researchers have found that during the process of tumor evolution, there are interactions among these heterogeneous cells. Often, high-grade tumor cells transfer malignant phenotypes (e.g., increased invasiveness, therapy resistance) to low-grade tumor subpopulations via exosomal cargo, enhancing their abilities in proliferation, invasion, and drug resistance. For example, exosomes derived from high-Gleason-score prostate cancer cells deliver miR-153 to adjacent cells, which modulates genes involved in cell cycle regulation and migration, thereby altering recipient cell behavior (43). Exosomal miR-150-5p was also significantly correlated with high Gleason scores and implicated in metastatic progression (44). Similarly, invasive prostate cancer cells secrete elevated levels of exosomal miR-424, which promotes tumor growth and metastasis by targeting downstream cell cycle regulators (45). In CRPC, plasma exosomes contain high levels of miR-222-3p and miR-375, which activate mTORC1 and STAT3 signaling via specific gene targets, enhancing cell proliferation and anti-apoptotic potential (46, 47).
2.3 Exosomal mechanisms in chemoresistance
Exosomal ncRNAs also mediate horizontal drug resistance transfer. Drug-resistant cells release FTH1P8, a pseudogene-derived RNA molecule, that binds ferritin heavy chain (FTH1) to reduce intracellular labile iron, inhibiting ferroptosis induction and spreading resistance to naive cells (48). Furthermore, researchers have found that a variety of circRNAs contribute to docetaxel resistance: circ-SFMBT2 acts as a miR-136-5p sponge to derepress TRIB1 and activate MAPK signaling (49); circ-XIAP upregulates TPD52 via miR-1182 sponging, inhibiting autophagy (50); and circSLC4A7 relieves miR-1205-mediated suppression of MAPT, enhancing microtubule stability (51). Collectively, these findings highlight exosomal communication as a critical driver of tumor heterogeneity and identify novel targets for overcoming docetaxel resistance.
2.4 Exosomal communication in ADT resistance and CRPC progression
In addition to chemotherapeutic resistance, ADT resistance in prostate cancer represents a critical challenge, as the development of CRPC signifies progression to a more aggressive and refractory stage (52). Elucidating CRPC pathogenesis and identifying predictive biomarkers remain central objectives in current research. A recent study (53) demonstrated that during ADT-induced transition from androgen-dependent to androgen-independent prostate cancer, exosome secretion and cargo composition undergo significant alterations. Androgens were further shown to modulate exosomal miRNA expression profiles and surface protein levels, thereby altering exosome targeting specificity and intercellular communication dynamics. This discovery uncovered a novel ADT resistance mechanism: exosomal ncRNA-mediated cell communication. Subsequent investigations corroborated the pivotal role of exosomal ncRNAs in CRPC progression. Both previous prospective cohort studies and plasma exosomal miRNA sequencing results have shown that miR-423-3p is significantly upregulated in CRPC patients, suggesting its important role in CRPC (54). More direct evidence (46, 55) indicated that prostate cancer-derived exosomes transfer miR-222-3p and let-7a-5p between cells. By modulating AR-V7 and ERG expression, these exosomal miRNAs facilitate the transdifferentiation of ADPC cells into androgen-independent prostate cancer (AIPC)-like cells, thereby driving androgen resistance development (summarized in Figure 2).
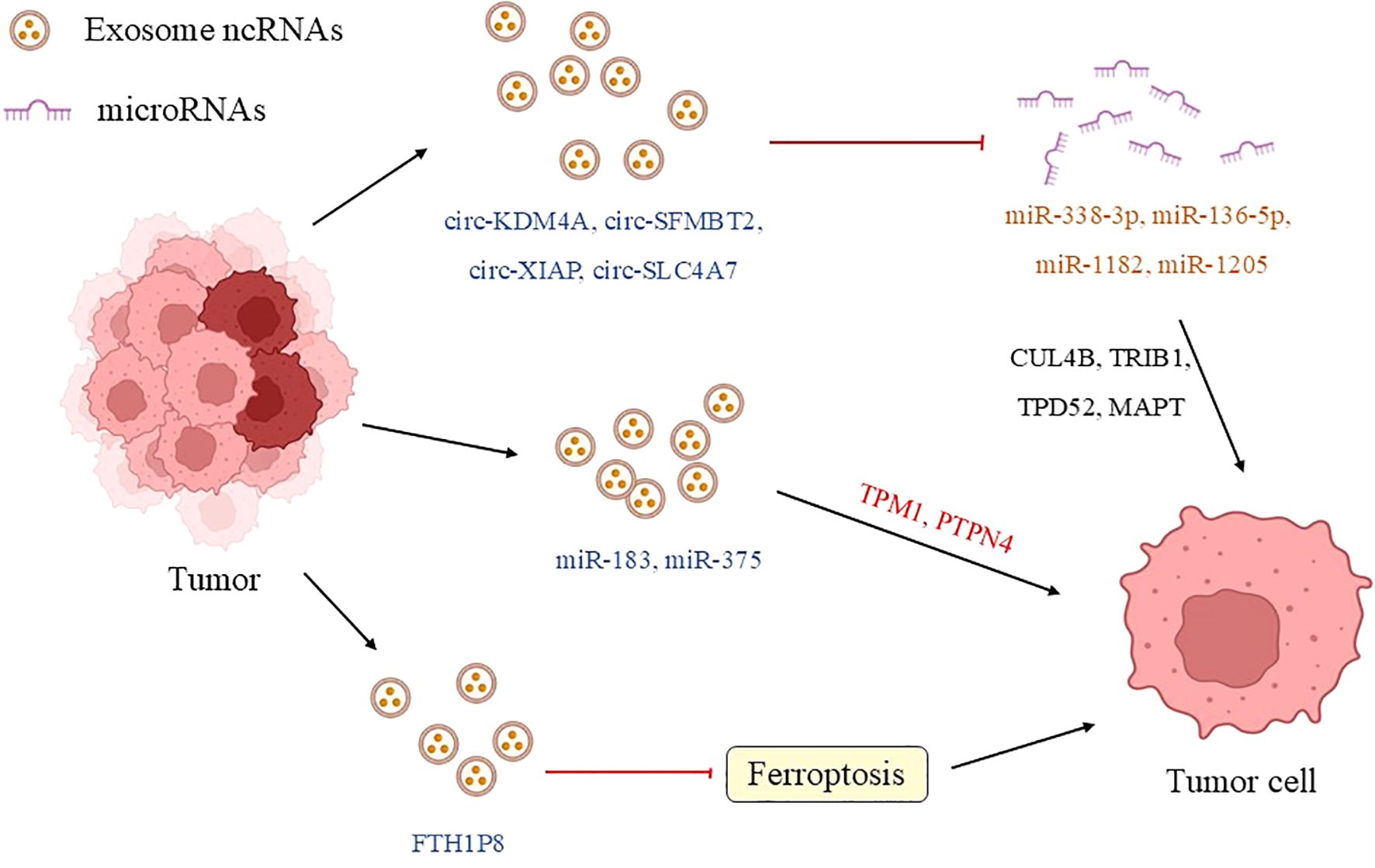
Figure 2. Exosomal ncRNA communication between tumor cells. In the tumor microenvironment of prostate cancer, tumor cells transfer ncRNAs (such as miR-183 and circKDM4A) through exosomes to regulate the biological behaviors of neighboring cells, promoting proliferation, invasion, and drug resistance, as well as tumor evolution and treatment resistance. For example, highly invasive cancer cells reprogram low-grade cells via exosomal miR-153/150-5p/424, while drug-resistant cells transmit chemotherapy resistance through exosomal circRNAs (such as circ-SFMBT2/XIAP/SLC4A7) or the pseudogene FTH1P8. These mechanisms reveal the central role of exosomal ncRNAs in the progression and drug resistance of prostate cancer. (TPM1, tropomyosin 1; PTPN4, phosphatase nonreceptor type 4; CUL4B, cullin 4B; STAT3, signal transducer and activator of transcription 3; TRIB1, tribbles homolog 1; TPD52, tumor protein D52; MAPT, microtubule-associated protein tau).
3 Exosomal ncRNA communication in tumor-stromal cell interactions
The non-malignant components within the tumor microenvironment, along with their interacting molecular networks, are collectively termed the stroma of the tumor microenvironment (16). Functionally, these stromal components not only provide support for tumor cells but also act as pivotal regulatory hubs for modulating the biological behaviors of tumors, thereby having a profound influence on tumor progression (56, 57). The stroma comprises non-cellular components with the extracellular matrix as the central element, along with cellular components including CAFs, immune cells (such as tumor-associated macrophages and lymphocytes), endothelial cells, and mesenchymal stem cells (58–60). Mounting evidence (61–63) suggests that tumor cells and the stroma within the tumor microenvironment mutually regulate the biological behaviors of tumors, including proliferation, invasion, metastasis, immune escape, and treatment resistance, via intricate bidirectional interactions. In this context, we primarily focus on the communication mediated by exosomal ncRNAs between tumor cells and stromal cells.
3.1 Exosomal ncRNA communication with CAFs
CAFs constitute a heterogeneous population of activated fibroblasts predominantly located within the tumor stroma and represent one of the most abundant stromal cell types in the tumor microenvironment (64, 65). A recent text mining study (66) analyzing CAFs revealed that the research emphasis on CAFs has transitioned from their basic biological properties in the early research phase to their functional versatility, including pro-fibrotic activity, metabolic regulation, and extracellular matrix remodeling, as well as their interactions with other constituents of the tumor microenvironment, such as immune cells and exosomes. Analogously, numerous investigations have explored the interactions among CAFs, exosomes, and tumor cells in prostate cancer. These investigations have further delineated the molecular mechanisms underlying chemotherapy and castration resistance. For instance, miR-423-5p has been shown to promote chemotherapy resistance in tumor cells by targeting the TGF-β/Smad and ferroptosis pathways (67, 68). These findings not only propose a novel therapeutic target for countering drug resistance but also imply that detecting exosomal miR-423-5p could serve as a prognostic biomarker.
Furthermore, CAFs facilitate the adaptation of prostate cancer cells to adverse conditions, including androgen deprivation and hypoxia, by downregulating exosomal ncRNA miR-146a-5p (69). Another study (70) revealed that CAFs can secrete specific miRNAs, including miR-154 and miR-376c, and transfer them to tumor cells through exosomes. These miRNAs specifically target and suppress the expression of NKX3-1, leading to the dysregulation of the AR signaling pathway and driving the transformation of prostate cancer cells towards castration resistance. Under hypoxic microenvironments, CAFs can secrete exosomal miRNAs, including miR-500a-3p. MiR-500a-3p stabilizes heat shock factor 1 (HSF1) by inhibiting the E3 ubiquitin ligase FBXW7, thereby promoting the metastasis of prostate cancer cells in response to hypoxic stress (71). These studies collectively highlight the facilitatory role of CAFs in promoting the progression of prostate cancer cells via exosomal communication. They provide novel mechanistic insights into the resistance of prostate cancer to chemotherapy and androgen deprivation therapy and offer new perspectives for developing strategies to overcome tumor drug resistance.
3.2 Exosomal ncRNA communication with MSCs
Besides CAFs, mesenchymal stem cells (MSCs) represent another crucial cell type within the tumor microenvironment stroma. MSCs are a subtype of adult stem cells exhibiting multi-directional differentiation potential. They can be sourced from diverse tissues, including bone marrow, adipose tissue, and umbilical cord (72). Additionally, they are attracted to the tumor microenvironment by chemokines, such as cytokines and growth factors, secreted by tumors (73). MSCs exhibit a dual function in relation to tumor cells. On one hand, they can facilitate tumorigenesis and progression through mechanisms such as immune suppression, angiogenesis promotion, and stromal remodeling. For instance, gastric cancer cell-derived small extracellular vesicles can induce metabolic reprogramming of bone marrow-derived MSCs (BM-MSCs) via the ERK-PPARγ-CPT1A signaling pathway, thereby enhancing lymphatic metastasis potential (74–76). On the other hand, MSCs can exert an anti-tumor effect by inducing tumor cell apoptosis, inhibiting cell proliferation, or secreting anti-angiogenic factors, such as thrombospondin-1 (TSP-1) (77).
Recent investigations have demonstrated that in prostate cancer, MSCs can exert an anti-tumor function by secreting exosomes that shuttle ncRNAs to tumor cells. For instance, exosomal miR-187 originating from bone marrow-derived mesenchymal stem cells has been verified to infiltrate prostate cancer cells. It inhibits the proliferation, migration, and invasion of prostate cancer cells and promotes apoptosis by targeting the CD276/JAK3-STAT3-Slug axis (78). Analogously, another study (79) revealed that bone marrow-derived mesenchymal stem cells can transfer miR-99b-5p to prostate cancer cells through exosomes. This action inhibits the IGF1R signaling pathway, subsequently reducing tumor growth and decreasing drug resistance. The results of these studies are highly promising. The differential expression of exosomal ncRNAs, identified through gene chip sequencing, suggests that these expression profiles hold potential as diagnostic or prognostic biomarkers. Furthermore, MSCs, which are key components of the tumor microenvironment, represent potential therapeutic targets. For instance, engineered MSCs can be utilized as drug delivery vehicles to enhance therapeutic efficacy by targeting tumor sites and releasing anti-tumor agents. As reported in the study by Kurniawati et al. (80), exosomes derived from mesenchymal stem cells were utilized as a delivery vehicle for let-7c to specifically target the MYC and AKT2 signaling pathways in castration-resistant prostate cancer cells. The results indicated that the exosomes encapsulating let-7c decreased the tumor volume by 62% and abrogated enzalutamide resistance. This study capitalizes on the natural targeting capabilities of stem cells and exosomes to offer an effective and targeted tumor treatment strategy.
3.3 Exosomal ncRNA communication with endothelial cells
It is worth noting that endothelial cells, another important cell type in the tumor stroma of prostate cancer, have also been found to have communication via exosomal ncRNAs with tumor cells. Endothelial cells within the tumor microenvironment contribute to tumor progression beyond their well-established role in supporting angiogenesis and nutrient supply. These cells engage in signal transduction with cancer cells via the secretion of various growth factors. For example, they can upregulate immune checkpoint molecules such as PD-L1, fostering an immunosuppressive microenvironment that impairs the anti-tumor immune response (81). Recent studies have further elucidated that endothelial cells function as critical hubs for the communication of extracellular vesicles, including exosomes. Through the reception and release of bioactive molecules, they significantly extend their regulatory influence over tumor biological behavior. In vitro experiments have found that exosomes secreted by PC-3 cells can promote the proliferation, migration, and lumen formation of endothelial cells. Further research (82) has confirmed that PC-3 exosomes are rich in certain pro-angiogenic proteins and microRNAs (such as miR-210). These molecules can promote angiogenesis by activating the relevant signaling pathways within endothelial cells (summarized in Figure 3). This result not only provides a new perspective for a deeper understanding of the angiogenesis mechanism of prostate cancer but also offers potential targets and strategies for the treatment of prostate cancer, suggesting that interventions targeting PC-3 exosomes and their related signaling pathways may help inhibit the angiogenesis and tumor growth of prostate cancer.
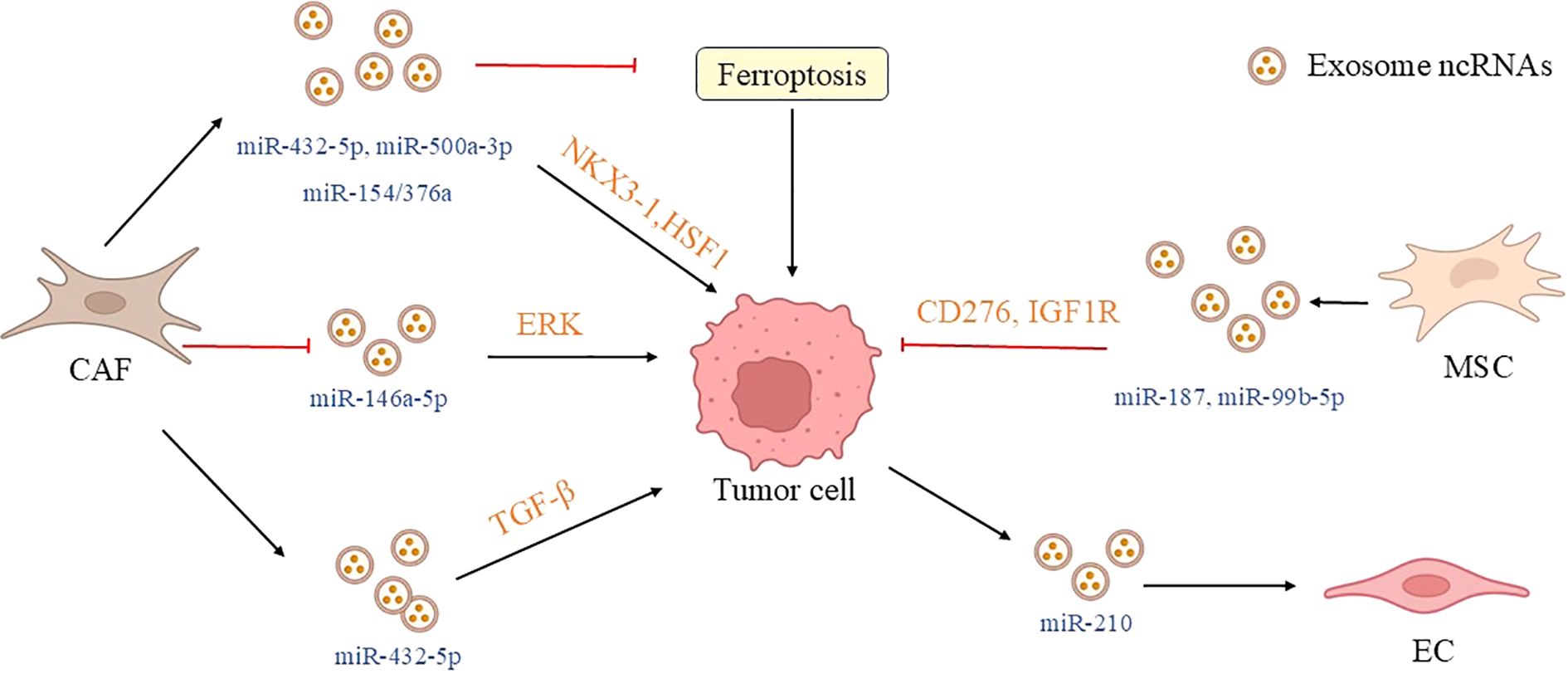
Figure 3. Exosomal ncRNA communication between tumor cells and stromal cells in the TME. Stromal components (such as CAFs, MSCs, and endothelial cells) in the tumor microenvironment form a bidirectional communication network with prostate cancer cells through exosomal ncRNAs, regulating tumor progression and treatment resistance. Exosomal miR-423-5p/miR-432-5p secreted by CAFs promotes chemoresistance and metastasis by targeting GREM2/CHAC1. The absence of CAF exosomal miR-500a-3p/miR-146a-5p induced by hypoxia or ADT enhances tumor adaptability. Exosomal miR-187/miR-99b-5p derived from MSCs can inhibit tumor growth. In addition, endothelial cells promote angiogenesis through exosomal miR-210, collectively forming a key regulatory network of the tumor microenvironment and providing a new strategy for intervening in prostate cancer by targeting stromal exosomes. (CAF, cancer-associated fibroblasts; MSC, mesenchymal stem cell; EC, endothelial cell; GREM2, gremlin-2; CHAC1, ChaC glutathione specific gamma-glutamylcyclotransferase 1; EGFR, epidermal growth factor receptor; ERK, extracellular regulated protein kinases; NKX3-1, NK3 homeobox 1; FBXW7, F-box and WD repeat domain-containing 7; HSF1, heat shock transcription factor 1; IGF1R, Insulin-like growth factor 1 receptor).
4 Exosome-derived ncRNAs modulate tumor-immune cell dialogues
The role of the immune response in tumorigenesis should not be underestimated. Consequently, the tumor immune microenvironment (TIME), a critical component of the TME, has attracted significant research interest. TIME centers on immune regulation within the TME, encompassing immune cells (e.g., T cells, myeloid-derived suppressor cells (MDSCs), tumor-associated macrophages (TAMs)), immune checkpoint molecules (e.g., PD-1/PD-L1), cytokines (e.g., IL-6, TGF-β), and their interactive networks (83, 84).
4.1 Immune cells in TIME
Immune cells within the TME constitute the fundamental components of the tumor immune ecosystem. These immune cells exhibit not only remarkable diversity in cell types but also intricate complexity in their functions, being capable of either suppressing tumor progression or enabling tumor immune escape. For example, during the early tumorigenic stages, immune cells such as CD8+ T cells and natural killer (NK) cells exert anti-tumor effects via immune surveillance (85). In contrast, MDSCs and M2-polarized macrophages secrete a repertoire of cytokines that establish an immunosuppressive microenvironment conducive to tumor progression (86, 87). Notably, a bidirectional regulatory relationship exists between tumor cells and immune cells. Tumor cells can establish an immunosuppressive microenvironment and evade immune surveillance by modulating immune cells. For instance, tumor cells can upregulate glycolysis (the Warburg effect), thereby competing for glucose in the microenvironment and impairing T cell function. Simultaneously, tumor cells can hinder the maturation of dendritic cells via lipid metabolites, such as prostaglandin E2 (88). Moreover, tumors can attract TAMs and MDSCs by secreting factors including vascular endothelial growth factor (VEGF) and colony-stimulating factor 1 (CSF-1). These recruited cells subsequently secrete enzymes, such as arginase 1 (Arg1) and inducible nitric oxide synthase (iNOS), which consume arginine crucial for T cell activation and generate immunosuppressive metabolites (89). In summary, the TIME represents a complex ecosystem comprising tumor cells, immune cells, diverse cytokines, and metabolites within tumor tissues. Its dynamic equilibrium directly influences tumor initiation, progression, metastasis, and the response to therapy, positioning it as one of the central areas of current tumor research.
4.2 Overview of the prostate cancer TIME
Unlike most tumors, the TIME of prostate cancer is predominantly immunosuppressive. Even at advanced tumor stages, the immunosuppressive microenvironment persists (90, 91). Specifically, it exhibits low PD-L1 expression, sparse immune cell infiltration (e.g., reduced CD8+ T cell abundance), and an enrichment of inhibitory immune cell populations, such as TAMs, MDSCs, and regulatory T cells (Tregs) (92). Moreover, the secretion of inhibitory cytokines, including transforming growth factor-β (TGF-β), interleukin-8 (IL-8), and interleukin-10 (IL-10), further intensifies immunosuppression (93). When compared to other solid tumors, like bladder cancer, prostate cancer represents a typical “immune-cold tumor.” In both the primary tumor site and in CRPC, immune cell infiltration is minimal. Likewise, in metastatic prostate cancer, particularly in bone metastases, the immune microenvironment exhibits enhanced immunosuppression, marked by an enrichment of M2-polarized macrophages and T-cell exhaustion (94, 95). This results in a low response rate to immune checkpoint inhibitors, such as PD-1/PD-L1 antibodies (96, 97). Additionally, recent studies have demonstrated that the androgen receptor (AR) signaling pathway plays a pivotal role in prostate cancer and engages in complex interactions with the immune microenvironment. AR signaling not only promotes the growth of prostate cancer epithelial cells but also directly regulates the establishment of the immune microenvironment by inhibiting antigen presentation (e.g., major histocompatibility complex class I (MHC-I) expression) and facilitating the recruitment of immunosuppressive cells (e.g., MDSCs) (98). Another study (99) has indicated that ADT can induce dynamic alterations in the composition of immune cells within the tumor microenvironment. Long-term ADT can activate immunosuppressive factors, such as interferon alpha 17 (IFNA17), contributing to the formation of a drug-resistant microenvironment, although the underlying mechanism remains incompletely understood. These immune features of prostate cancer present significant challenges to current immunotherapy strategies for prostate tumors.
Thus, delving deeper into the mechanisms underlying cell functions within the immune microenvironment of prostate cancer holds substantial potential for surmounting this challenge. Subsequently, we summarize the communication mediated by exosomal ncRNAs between tumor cells and immune cells within the tumor microenvironment, aiming to further elucidate the cell communication network in the immune microenvironment of prostate cancer.
4.3 Immunostimulatory exosomal ncRNA communication
As previously discussed, the tumor immune microenvironment displays temporal heterogeneity. During the early phases of tumorigenesis, the immune microenvironment predominantly exerts a tumor-suppressive effect. Nevertheless, as the tumor advances, a tumor-immunosuppressive microenvironment gradually emerges. We hypothesized that this might be partially attributable to the exosomal communication between tumor cells and immune cells. In the prostate cancer tumor microenvironment, exosomes function as critical mediators of intercellular communication, regulating complex immune responses through the delivery of bioactive molecules. Although macrophage polarization is a key aspect of this regulation, the influence of exosomes on other immune cells warrants further investigation. Research (100) demonstrates that exosomes secreted by prostate cancer cells are enriched with the chemokine CXCL14. Upon uptake by macrophages, these exosomes trigger NF-κB signaling to induce M2 polarization, thereby establishing an immunosuppressive microenvironment that facilitates tumor progression. Similarly, exosomes derived from PC3 cells promote M2 macrophage polarization via the delivery of miR-let-7b (101). Furthermore, recent studies (102) indicate that YY1 expression in M2 macrophages is upregulated by super-enhancers, promoting the secretion of exosomes containing circ-0000326. Following their uptake by prostate cancer cells, these exosomes elevate FZD7 expression by sponging miR-1258, which activates the Wnt/β-catenin signaling pathway and forms a positive feedback loop. Beyond macrophages, other immune cells significantly contribute to prostate cancer progression. For instance, exosomes from MDSCs advance CRPC through the S100A9/circMID1/miR-506-3p/MID1 axis. Mechanistically, exosomal S100A9 upregulates circMID1, which then sequesters miR-506-3p, alleviating its suppression of MID1 and subsequently activating the mTOR signaling pathway. Notably, inhibiting S100A9 significantly attenuates the tumor-promoting effects of these exosomes, reducing tumor growth by 51% (103). Meanwhile, miR-95 within TAM-derived exosomes drives tumor malignancy through multiple mechanisms. It directly targets the JUNB gene to enhance proliferation, invasion, and epithelial-mesenchymal transition in prostate cancer cells. Concurrently, miR-95 impairs endocytosis by suppressing SNX1 expression, thereby amplifying cellular sensitivity to growth factors like EGF and promoting migration and angiogenesis. Clinical analyses confirm that elevated miR-95 expression correlates with adverse clinicopathological features (104).
4.4 Immunosuppressive exosomal ncRNA communication
Regarding tumor-suppressive mechanisms, exosomal ncRNAs employ diverse regulatory pathways. For instance, the lncRNA ZNF667-AS1, derived from prostate cancer cells and delivered via exosomes, binds to the U2AF1 protein. This interaction impairs the stability of transforming growth factor-β receptor 1 (TGFBR1) mRNA, resulting in decreased TGFBR1 expression. This downregulation not only inhibits the expansion of Tregs, thereby attenuating the immunosuppressive microenvironment, but also significantly enhances tumor cell chemosensitivity to docetaxel. Animal experiments confirmed that the administration of exosomes carrying ZNF667-AS1 increased chemosensitivity by 2.3-fold (105). Separately, exosomal miR-203 secreted by prostate cancer cells induces macrophage polarization toward the M1 phenotype by suppressing SOCS3 expression and subsequently activating the STAT1 signaling pathway. The polarized M1 macrophages secrete cytokines, including tumor necrosis factor-α and interleukin-12, which directly inhibit tumor cell proliferation and migration while inducing apoptosis. In a mouse model, miR-203 overexpression reduced tumor volume by 58%, demonstrating its therapeutic potential to reverse the immunosuppressive microenvironment through macrophage reprogramming (106). Collectively, these studies reveal a complex, exosome-mediated regulatory network within the prostate cancer tumor microenvironment. On one hand, immune cells such as MDSCs and TAMs accelerate tumor progression by delivering oncogenic molecules like circMID1 and miR-95. Conversely, exosomes derived from tumor cells can also carry tumor-suppressive molecules such as ZNF667-AS1 and miR-203, which inhibit tumor growth by modulating Treg expansion, promoting anti-tumor macrophage polarization, and enhancing chemosensitivity. (summarized in Figure 4). These studies further elucidate the mechanisms underlying prostate cancer progression and enhance our understanding of the prostate cancer tumor immune microenvironment.
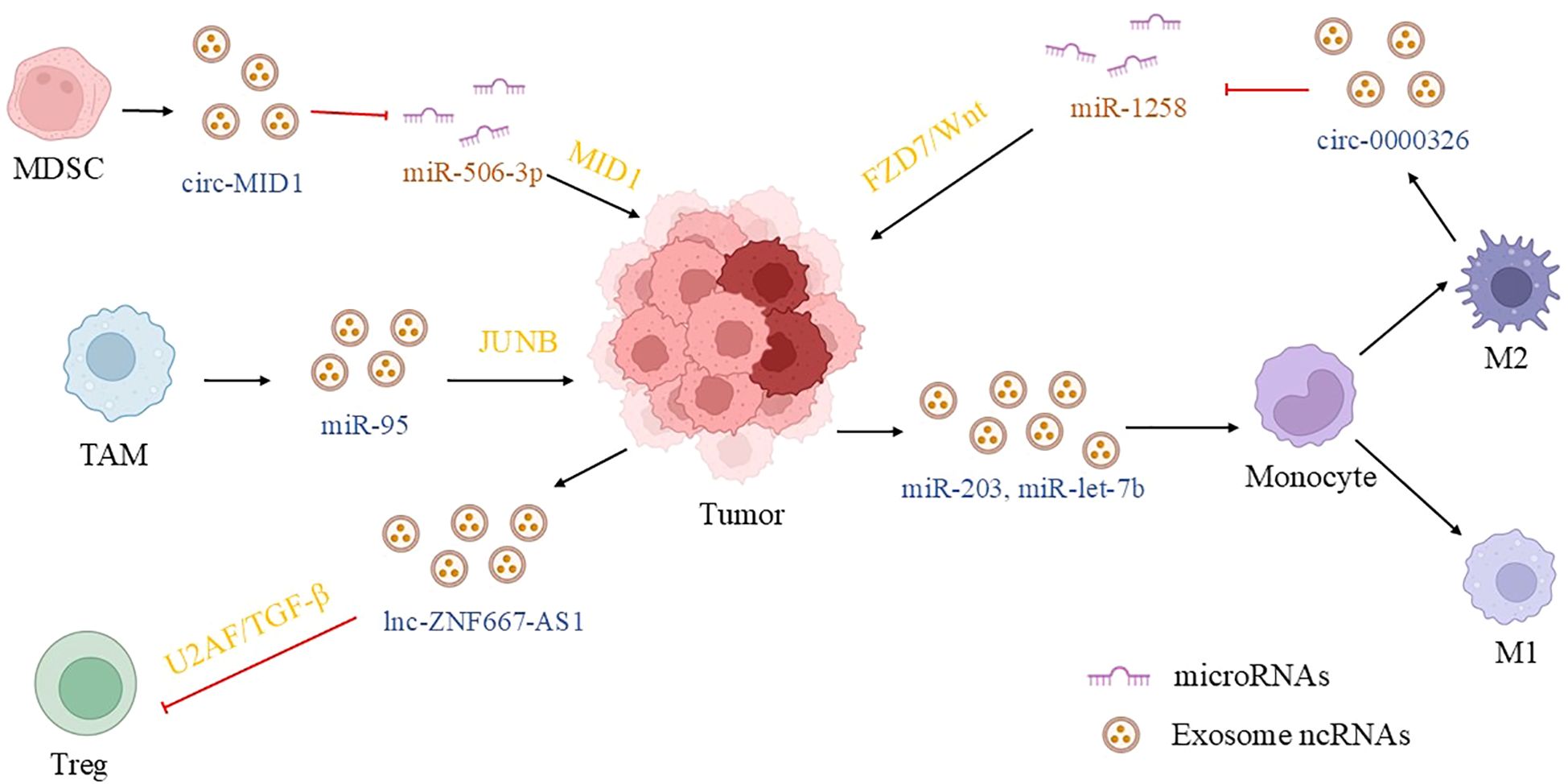
Figure 4. Exosomal ncRNA communication between prostate cancer cells and immune cells in the TME. In the immune microenvironment of prostate cancer, tumor cells and immune cells form a bidirectional regulatory network through exosomal ncRNAs. Exosomal CXCL14/miR-let-7b secreted by tumor cells induces the polarization of M2 macrophages, and M2 macrophages activate the Wnt pathway through exosomal circ-0000326 to promote tumor progression. Meanwhile, exosomal ZNF667-AS1/miR-203 of prostate cancer cells can inhibit Treg cells or induce M1 macrophages, enhancing anti-tumor immunity. In addition, immune cells (such as MDSCs/TAMs) directly promote tumor proliferation and EMT through exosomal circMID1/miR-95. This dynamic balance not only maintains the immunosuppressive microenvironment but also has the potential for immune activation, providing new strategies for targeting the immune microenvironment. (MDSC, myeloid-derived suppressor cells; TAM, tumor associated macrophage; M2, M2 macrophages; M1, M1 macrophages; MID1, midline1; JUNB, Jun B proto-oncogene; U2AF, U2 small nuclear RNA auxiliary factor 1; FZD7, frizzled class receptor 7).
5 Exosomal ncRNA signaling in remodeling the bone metastatic tumor microenvironment
5.1 Overview of PCa bone metastasis biology
Bone metastasis in advanced prostate cancer is one of the primary causes of death among patients, and this metastatic process is governed by intricate molecular mechanisms (107). Generally, this metastatic process encompasses several key steps, including primary tumor invasion, EMT, the survival and homing of circulating tumor cells, colonization within the bone microenvironment, and the bidirectional regulation between osteolysis and osteogenesis (108). Prostate cancer cells gain migratory capacity via EMT, detach from the primary tumor locus, and subsequently enter the circulatory system. Subsequently, these cells migrate through the bloodstream and lymphatic vessels. Guided by chemokines, they home in a directional manner to the bone marrow microenvironment (109, 110). Notably, certain tumor cells that have colonized within the bone metastasis microenvironment may enter a dormant state and can be reactivated subsequently upon stimulation by environmental factors, such as inflammatory factors (111, 112). Osteoblasts, osteoclasts, and mesenchymal stem cells within the bone marrow secrete growth factors, thereby creating a microenvironment conducive to tumor growth (113). Within this bone metastasis microenvironment, cancer cells activate osteoblasts and osteoclasts through the secretion of diverse factors, leading to the induction of osteoblastic and osteoclastic lesions. In prostate cancer, the lesions are predominantly osteoblastic, with accompanying osteolytic activities. Conversely, the activated osteoblasts and osteoclasts secrete growth factors, which in turn further stimulate tumor growth, thus establishing a vicious cycle (114). During the process of bone metastasis, osteoblasts secrete chemokines and growth factors, which serve as tumor homing signals and facilitate the adhesion of tumor cells to the tumor stroma (115, 116). Meanwhile, upon activation, osteoclasts induce bone resorption, releasing TGF-β, insulin-like growth factor-1 (IGF-1), calcium ions, and other substances stored in the bone matrix, thereby providing proliferation signals for tumor cells. Moreover, the cavities generated by osteoclast-mediated bone resorption offer physical space for tumor cell colonization (117). In summary, bone metastasis in prostate cancer is the outcome of dynamic interactions between cancer cells and the bone microenvironment, involving crucial processes such as EMT, chemokine-mediated homing, and the establishment of self-reinforcing feedback loops. Additionally, the bone metastasis process extensively entails interactions among diverse cell types, with tumor cells, osteoblasts, and osteoclasts being particularly significant. A more in-depth understanding of these mechanisms is instrumental in the development of targeted therapeutic strategies and the enhancement of patient prognosis. Future research endeavors should concentrate on tumor heterogeneity, the eradication of dormant cells, and the formulation of microenvironment reprogramming strategies.
As previously discussed, within the bone metastasis microenvironment, tumor cells, osteoblasts, and osteoclasts interact with one another, giving rise to a cyclic “vicious triangle”. This interaction not only perturbs bone homeostasis but also establishes a mutually reinforcing cycle through the release of diverse factors, thereby exacerbating disease progression. Consequently, an increasing number of studies are commencing to investigate the mechanisms underlying the interactions among these three cell types. In this section, we summarize the communication mediated by exosomal ncRNAs among these three cell types during the bone metastasis process of prostate cancer.
5.2 Tumor-derived exosomal ncRNAs affecting bone cells
Certain exosomal ncRNAs can activate osteogenic differentiation pathways, inducing a transition in the bone microenvironment towards an “osteoblastic” phenotype that provides structural support for tumor cell colonization. Among these, lncRNA NEAT1 and miR-375 are key regulatory factors. In vitro co-culture experiments have demonstrated that exosomes derived from prostate cancer cells are internalized by human bone marrow mesenchymal stem cells (hBMSCs). Subsequently, NEAT1 sequesters miR-205-5p, which relieves the suppression of RUNX2 and activates osteogenic differentiation signaling. This finding was further validated in nude mouse models, where exosomes overexpressing NEAT1 increased the number of mineralized nodules in bone tissue and promoted the formation of bone metastatic lesions (118). Similarly, in vitro experiments confirmed that exosomal miR-375 promotes the proliferation and migration of prostate cancer cells and induces the osteogenic differentiation of hBMSCs by targeting the DIP2C gene and consequently activating the Wnt/β-catenin pathway. Notably, serum exosomes from patients with bone metastasis exhibited significantly elevated miR-375 levels, and its expression positively correlated with bone scan positivity, suggesting its involvement in clinical bone metastasis progression (119).
In contrast to tumor-promoting ncRNAs, other exosomal ncRNAs inhibit osteoblast proliferation and differentiation by targeting key osteogenic genes, leading to diminished bone formation and even osteolytic lesions. miR-1275 and miR-940 are representative of this category, with the former’s inhibitory effect underscoring the signaling complexity within the bone microenvironment. In vitro osteoblast culture experiments revealed that upon uptake of prostate cancer cell-derived exosomal miR-1275, osteoblasts exhibit targeted silencing of the SIRT2 gene. SIRT2 downregulation reduces RUNX2 deacetylation, ultimately suppressing osteoblast proliferation and mineralization. Verification in a mouse tibial injection model confirmed that exosomes overexpressing miR-1275 reduce osteocalcin (OCN) expression in bone tissue and exacerbate osteolytic damage, providing a mechanistic explanation for the coexistence of osteolytic and osteoblastic lesions in clinical prostate cancer bone metastasis (120). Notably, exosomes isolated from a mouse orthotopic prostate cancer model were found to significantly impair osteoblast mineralization upon co-culture. This effect was mediated by exosomal miR-940, which suppresses the expression of key osteogenic differentiation genes, including ALP and RUNX2 (121). This study identifies one of the few inhibitory ncRNAs validated in in vivo models, presenting a stark contrast to the multitude of ncRNAs that promote tumor progression. This antagonistic relationship suggests that the bone microenvironment is governed by a dynamic balance between “pro-metastatic” and “anti-metastatic” signals, rather than being driven solely by promotional influences. This insight provides a new perspective for understanding the complex pathology of bone metastasis.
The disruption of osteoclast differentiation balance is a major contributor to bone damage in metastasis. Exosomal ncRNAs can influence osteoclastogenesis by modulating relevant differentiation pathways, with miR-92a-1-5p exerting a pro-osteoclast effect and miR-148a an anti-osteoclast effect. In vitro experiments on osteoclast precursor cells demonstrated that exosomal miR-92a-1-5p targets and inhibits the expression of the COL1A1 gene. This not only impairs osteogenic function but also activates osteoclastogenic transcription factors such as NFATc1, thereby promoting the fusion of precursor cells into mature, multinucleated osteoclasts (122). Conversely, following their uptake by osteoclast precursor cells, exosomes from prostate cancer PC-3 cells exert an anti-osteoclast effect via the downregulation of miR-148a. The reduction in miR-148a leads to the upregulation of its target gene, MAFB, and concurrently inhibits the PI3K/AKT/mTOR pathway, ultimately diminishing the formation of TRAP-positive multinucleated cells (123).
5.3 Bone cell-derived exosomal ncRNAs affecting tumor cells
Osteoblasts within the bone microenvironment do not passively receive tumor-derived signals; they actively contribute to tumor progression by secreting exosomes that deliver ncRNAs, which in turn enhance the proliferation, migration, and bone colonization capacity of prostate cancer cells. This process constitutes a critical link in the pathological synergy between tumors and bone. Current research has identified two core regulatory factors in this pathway: miR-140-3p and circ-DHPS, whose mechanisms have been validated through both in vitro and in vivo experiments. In vitro studies demonstrated that exosomes secreted by osteoblasts are internalized by prostate cancer cells (LNCaP, PC-3), subsequently promoting cell proliferation and migration by modulating the autophagic pathway. Subsequent validation in a nude mouse bone metastasis model confirmed that injection of osteoblast-derived exosomes significantly increased the burden of tumor cell colonization in bone tissue (124). In CRPC, a distinct clinical subtype, the role of osteoblast-derived exosomes is further modulated. A key study (125) revealed that AR inactivation alters the ncRNA cargo of osteoblast-derived exosomes, thereby influencing tumor cell chemotactic migration. Specifically, following AR knockdown in osteoblasts in vitro, the expression of circular RNA circ-DHPS in their exosomes was significantly upregulated. Upon internalization by prostate cancer cells, circ-DHPS acts as a molecular sponge for miR-214-3p. This sequestration relieves the miR-214-3p-mediated suppression of the chemokine CCL5, leading to enhanced CCL5 expression and consequently promoting tumor cell adhesion and invasion.
5.4 Reciprocal signaling via MSCs
As multipotent stem cells within the bone microenvironment, MSCs possess the capacity to differentiate into various lineages, including osteoblasts and adipocytes, and are integral to immune regulation and cytokine secretion. Exosomes derived from prostate cancer cells can indirectly influence bone metabolic balance and tumor progression by modulating MSC function. Key molecules mediating this indirect communication include lncAY927529 and miR-142-3p. In vitro experiments demonstrated that lncAY927529, carried by prostate cancer cell-derived exosomes, is internalized by ST2 cells (a model MSC line). By modulating autophagic activity in ST2 cells, lncAY927529 influences the secretion of the chemokine CXCL14, which in turn promotes the proliferation and invasion of neighboring prostate cancer cells via a paracrine mechanism (126). This pathway was validated using in vitro Transwell and co-culture models, indicating that lncRNAs can indirectly contribute to remodeling the bone metastatic niche by altering the secretory profile of MSCs. Furthermore, the regulation of MSCs by exosomal ncRNAs is susceptible to pharmacological intervention. As reported in study (127), propofol, a commonly used clinical anesthetic, can impair the osteogenic differentiation of MSCs by altering exosomal ncRNA levels. Propofol treatment significantly reduced exosomal miR-142-3p, concurrently inhibiting the osteogenic potential of MSCs. Mechanistic investigation revealed that miR-142-3p promotes osteogenesis by targeting and suppressing negative regulators of this process, such as DKK1. Propofol abrogates this pro-osteogenic effect by downregulating miR-142-3p, ultimately impeding the formation of a hospitable bone metastatic microenvironment.
5.5 Therapeutic and biomarker potential
The deepening understanding of regulatory mechanisms has revealed the considerable clinical potential of exosomal ncRNAs for the early diagnosis, prognostic assessment, and therapeutic targeting of prostate cancer bone metastasis. Current research has made preliminary progress in identifying diagnostic biomarkers and exploring therapeutic strategies. For instance, a comparative analysis of plasma exosomes from 82 patients with prostate cancer bone metastasis, 65 patients without bone metastasis, and 40 healthy controls identified a signature of four differentially expressed miRNAs (hsa-miR-125a-3p, hsa-miR-330-3p, hsa-miR-339-5p, hsa-miR-613). Among these, the expression level of hsa-miR-125a-3p was positively correlated with the stage of bone metastasis, demonstrating a diagnostic sensitivity of 78.1% and a specificity of 75.4% (128). Subsequent follow-up data indicated that patients with high hsa-miR-125a-3p expression had significantly shorter bone metastasis-free survival, suggesting its utility not only for early diagnosis but also as a prognostic indicator. Furthermore, analysis of clinical samples confirmed that serum exosomal miR-375 levels were significantly elevated in patients with bone metastasis compared to those without, and its expression positively correlated with the burden of bone metastatic lesions, thereby reinforcing the feasibility of exosomal miRNAs as diagnostic biomarkers (119).
Leveraging these elucidated regulatory mechanisms, targeting exosomal ncRNAs or their downstream pathways has emerged as a promising direction for treating bone metastasis. In vitro experiments demonstrated that inhibition of miR-1275 significantly restored SIRT2/RUNX2 signaling in osteoblasts and increased mineralized nodule formation (120). A separate study confirmed that knockdown of miR-375 suppressed the proliferation and migration of prostate cancer cells by attenuating Wnt pathway activity (119). Moreover, a combination strategy that concurrently targets key ncRNAs and critical pathway components within the tumor-bone microenvironment may yield synergistic therapeutic effects. For example, one investigation found that the combined application of a miR-140-3p inhibitor and an AKT inhibitor more potently suppressed the proliferation and bone colonization of prostate cancer cells (124). This combinatorial approach demonstrated robust anti-tumor efficacy in nude mouse models, providing a strong experimental foundation for future clinical translation. (summarized in Figure 5).
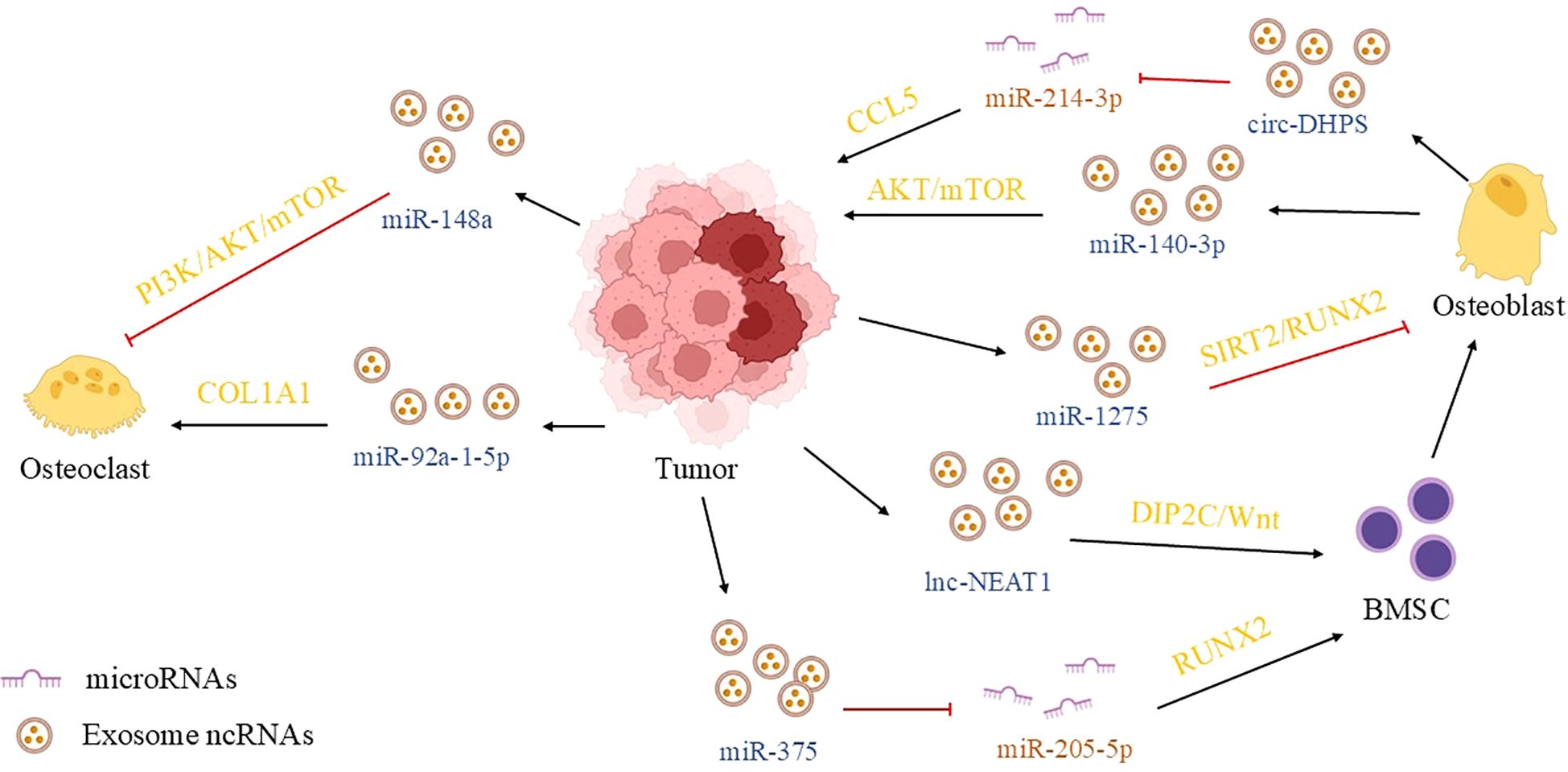
Figure 5. The role of exosomal ncRNA communication in shaping the bone metastasis microenvironment of prostate cancer. In the bone metastasis microenvironment of prostate cancer, tumor cells, osteoblasts, and osteoclasts form a “vicious triangle” through exosomal ncRNAs: PCa exosomes carry NEAT1/miR-375 to induce the osteogenic differentiation of mesenchymal stem cells, while miR-1275/miR-92a-1-5p inhibits osteogenic activity or activates osteoclasts, disrupting bone homeostasis. Osteoblasts promote tumor progression through exosomal miR-140-3p/circ-DHPS, and ncRNAs related to osteoclast activation (such as miR-92a-1-5p) exacerbate osteolytic lesions. This suggests that targeting exosomal communication in the bone microenvironment may become a new anti-metastasis strategy. (BMSC, bone marrow-derived mesenchymal stem cells; ACER2, alkaline ceramidase 2; CCL5, C-C chemokine ligand 5; SIRT2, sirtuin 2; RUNX2, runt-related transcription factor 2; DIP2C, disco interacting protein 2 homolog C).
6 Conclusion and prospect
PCa, which ranks among the leading causes of cancer-related fatalities in men globally, continues to present significant clinical challenges regarding treatment resistance and metastatic spread. In recent years, exosomes, serving as crucial mediators of intercellular communication within the TME, have garnered substantial attention. This is because of the distinctive roles played by the ncRNAs they carry in regulating tumor progression, facilitating immune escape, and contributing to drug resistance.
Exosomal ncRNAs act as “molecular messengers” in the TME of PCa by mediating intercellular communication. In the intercellular communication between tumor cells, exosomal ncRNAs serve as molecular messengers, promoting tumor progression and enhancing drug resistance. For instance, prostate cancer cells with a high Gleason score secrete exosomal microRNAs (miR-153, miR-150-5p, and miR-424), which reprogram neighboring low-grade cells and enhance their invasive phenotypes (43–45). In addition, the bidirectional exosomal communication between tumor cells and stromal cells (such as CAFs and MSCs) further shapes the pro-tumor microenvironment (67, 68, 70, 71, 78, 79). On the other hand, the immune characteristics of the prostate cancer TIME are closely associated with the regulation of exosomal ncRNAs. Tumor cells induce the polarization of macrophages towards the M2 phenotype via exosomal CXCL14 or microRNA (miR-let-7b), suppressing the anti-tumor immune response (100, 101). Meanwhile, circular RNA (circMID1) or microRNA (miR-95) secreted by MDSCs or TAMs directly drives the progression of CRPC (103, 104). Intriguingly, some exosomal ncRNAs (such as ZNF667-AS1 or miR-203) can also counteract immunosuppression, augmenting the anti-tumor effect by inhibiting the proliferation of Tregs or inducing the polarization of M1 macrophages (105, 106). This bidirectional regulation unveils the intricate role of exosomal ncRNAs in the prostate cancer TIME, suggesting their potential as targets for enhancing immunotherapy sensitivity. Similarly, in the bone metastasis microenvironment, exosomal ncRNAs exacerbate the imbalance between osteogenesis and osteolysis by mediating the “vicious triangle” interaction (119, 122, 124, 125, 129). To summarize, exosomal ncRNAs collectively drive the malignant progression of prostate cancer by mediating complex intercellular crosstalk within TME. Then, how does this intricate communication network evolve as the disease progresses from local advancement to castration resistance, and eventually colonizes the bone? During tumor proliferation and early metastasis, exosomal miR-183 promotes cancer cell invasion and bone metastasis by targeting TPM1 (37), whereas circKDM4A upregulates CUL4B by sponging miR-338-3p to drive malignant progression (38). The progression to CRPC marks a critical turning point, characterized by distinct alterations in exosomal ncRNA expression: For instance, miR-222-3p and miR-375 enhance treatment resistance by activating the mTOR signaling pathway and the PTPN4/STAT3 axis, respectively (46, 47). Similarly, let-7a-5p and LINC01213 facilitate the transition to androgen-independent growth, thereby promoting the development of CRPC (31, 55). Furthermore, circRNAs including circ-SFMBT2, circSLC4A7, and circ-XIAP confer docetaxel resistance via ceRNA mechanisms, collectively forming a robust cross-resistance network (49–51). The role of exosomal ncRNAs in remodeling the TME to drive distant metastasis is also significant. For example, CAF-derived miR-500a-3p and miR-423-5p remodel the microenvironment to promote metastasis and drug resistance under hypoxic or post-treatment conditions (50, 68). Additionally, macrophage-associated circ-0000326 accelerates disease progression via the Wnt signaling pathway (102). Ultimately, in the terminal stage of bone metastasis, exosomal molecules such as miR-375 and lncRNA NEAT1 orchestrate osteoblastic and osteoclastic activity, thereby generating a pre-metastatic niche that supports tumor colonization (118, 119). Collectively, these findings delineate a detailed regulatory network of exosomal ncRNAs throughout the dynamic evolution of prostate cancer, thereby enhancing our understanding of its underlying mechanisms.
In addition, exosomal ncRNAs have increasingly been applied in clinical translational research. In the context of diagnosis, plasma exosomal microRNA (miR-423-3p) or microRNA combinations (such as miR-125a-3p, miR-613) have demonstrated potential as early diagnostic markers for CRPC or bone metastasis (21, 130). The stability and accessibility of ncRNAs render them ideal biomarkers for early diagnosis. For example, in lung cancer, miRNA panels derived from liquid biopsies can differentiate benign from malignant lesions (131), while in prostate cancer, lncRNAs such as PCA3 have been established as clinical diagnostic markers (132). Furthermore, circRNAs exhibit significant potential as diagnostic and prognostic markers for central nervous system tumors, a property attributed to their high stability and tissue specificity (133). Similarly, the expression profiles of exosomal ncRNAs in hematological malignancies facilitate disease classification and monitoring of therapeutic efficacy, underscoring their broad potential in precision medicine (134). In the realm of treatment, strategies such as engineering exosomes derived from MSCs to be loaded with let-7c or targeting and inhibiting miR-222-3p have significantly reversed drug resistance in preclinical models (80). Furthermore, significant progress has been made in applying exosomal ncRNAs in adjuvant immunotherapy. As previously discussed, exosomal ncRNAs can modulate immune cells within the tumor microenvironment to foster an immunosuppressive state. Targeting these specific ncRNAs presents a promising strategy to reverse immune escape and enhance the efficacy of checkpoint blockade therapy (135). Additionally, certain exosomal miRNAs, including miR-192 and miR-21, have been shown to regulate post-vaccination immune responses, suggesting their potential utility as vaccine adjuvants or predictive biomarkers for vaccine efficacy (136). Despite these promising findings, the clinical translation of exosomal ncRNAs faces several challenges. Significant inter-patient heterogeneity presents a primary obstacle; for instance, in hepatocellular carcinoma, individual variations in exosomal miRNA expression profiles compromise their consistency as universal biomarkers (137). A second major challenge is the technical difficulty in detecting these ncRNAs due to their exceptionally low abundance in bodily fluids, which often leads to insufficient analytical sensitivity (138). The lack of standardized protocols for exosome isolation represents another prominent issue, as inconsistencies across different platforms hinder the comparability of research findings and clinical results. This is particularly problematic for central nervous system tumors, where the blood-brain barrier limits the concentration of exosomal nucleic acids in the periphery, and variability in detection sensitivity among methods further obstructs clinical application (139). Finally, the clinical utility of exosomal biomarkers requires validation in large-scale, prospective cohorts. When considering exosomes as drug delivery systems, additional complexities arise, including the need to standardize isolation, purification, targeting efficiency, and large-scale production, all of which complicate the regulatory approval pathway (140, 141). In the future, it will be essential to establish large-scale, multi-center cohorts to validate the diagnostic value of exosomal ncRNAs. Additionally, developing more sensitive and specific detection technologies (such as nano-flow cytometry) and enriching tumor-specific exosomes in conjunction with surface markers (such as prostate-specific membrane antigen, PSMA) will enhance the sensitivity and specificity of detection. Simultaneously, spatial transcriptomics or CRISPR-labeling tracking technologies can be employed to analyze the dynamic changes of exosomal ncRNAs during tumor progression, dormancy, and recurrence. This will contribute to uncovering the spatiotemporal patterns of their regulatory networks. Second, treatment strategies targeting exosomes must strike a balance between efficiency and safety.
From the perspective of mechanism research, the regulatory network of exosomal ncRNAs extends far beyond the solitary ceRNA model. In the future, it will be imperative to integrate multi-omics data (such as the circRNA-miRNA-mRNA co-expression network) to analyze their synergistic or antagonistic effects and explore their interactions with epigenetic modifications. For instance, exosomal lncA1BG-AS1 influences m6A modification by regulating ZC3H13, indicating that epigenetic regulation could be a crucial pathway for exosomal ncRNAs to exert their functions. Moreover, the role of exosomes in tumor cell quiescence remains ambiguous. Quiescent cells in the bone metastasis microenvironment sustain a quiescent state by transferring molecules such as miR-34a via exosomes. Targeting these “quiescent signals” could potentially prevent metastasis recurrence, offering a direction for the development of novel maintenance therapies. Furthermore, regarding the clinical challenge of bone metastasis in PCa, the “vicious cycle” mediated by exosomal ncRNAs offers multiple targets for intervention. On the one hand, utilizing exosomes derived from BMSCs to deliver anti-resorptive drugs or ncRNA antagonists (such as miR-375) can directly disrupt the osteogenesis-tumor interaction. On the other hand, modulating exosomal circ-DHPS or lncAY927529 could reverse the pro-metastatic microenvironment. Additionally, inhibitors of exosomal microRNA (miR-92a-1-5p) that target osteoclast activation could potentially restore bone homeostasis and enhance the quality of life of patients. These certainly offer potential directions, yet there remains a significant distance to traverse before these goals can be truly accomplished.
In summary, exosomal ncRNAs serving as the “molecular bridge” within the TME of PCa, drive disease progression by facilitating complex intercellular communications and offer unparalleled opportunities for accurate diagnosis and targeted treatment. Despite the challenges they still encounter in exosome isolation techniques, heterogeneity analysis, and clinical translation, through interdisciplinary collaboration and technological innovation, exosomal ncRNAs are hold potential to be a key contributor to overcoming the treatment predicament of PCa. Future research endeavors should concentrate on unveiling their dynamic regulatory networks, formulating effective targeted strategies, and facilitating the implementation of personalized treatment regimens. By decoding the molecular language of exosomal ncRNAs, we can refine diagnostic precision, guide individualized therapies, and ultimately improve survival and quality of life for patients with advanced PCa.
Author contributions
HD: Writing – review & editing, Writing – original draft, Investigation, Conceptualization, Supervision, Project administration. BX: Methodology, Validation, Visualization, Writing – original draft, Writing – review & editing. PL: Writing – original draft, Resources, Visualization, Data curation, Methodology, Investigation, Writing – review & editing. TC: Formal Analysis, Project administration, Writing – review & editing, Conceptualization, Investigation. JZ: Validation, Methodology, Investigation, Writing – review & editing, Supervision.
Funding
The author(s) declare that no financial support was received for the research, and/or publication of this article.
Acknowledgments
All figures are created with BioRender.com.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Generative AI statement
The author(s) declare that no Generative AI was used in the creation of this manuscript.
Any alternative text (alt text) provided alongside figures in this article has been generated by Frontiers with the support of artificial intelligence and reasonable efforts have been made to ensure accuracy, including review by the authors wherever possible. If you identify any issues, please contact us.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
1. Al-Ghazawi M, Salameh H, Amo-Afful S, Khasawneh S, and Ghanem R. An in-depth look into the epidemiological and etiological aspects of prostate cancer: A literature review. Cureus. (2023) 15:e48252. doi: 10.7759/cureus.48252
2. Luo LS, Jiang JF, Luan HH, Zi H, Zhu C, Li BH, et al. Spatial and temporal patterns of prostate cancer burden and their association with Socio-Demographic Index in Asia, 1990-2019. Prostate. (2022) 82:193–202. doi: 10.1002/pros.24258
3. Serdà-Ferrer BC, Sanvisens A, Fuentes-Raspall R, Puigdemont M, Farré X, Vidal-Vila A, et al. Significantly reduced incidence and improved survival from prostate cancer over 25 years. BMC Public Health. (2023) 23:2552. doi: 10.1186/s12889-023-17440-7
4. Bergengren O, Pekala KR, Matsoukas K, Fainberg J, Mungovan SF, Bratt O, et al. 2022 Update on prostate cancer epidemiology and risk factors-A systematic review. Eur Urol. (2023) 84:191–206. doi: 10.1016/j.eururo.2023.04.021
5. Yechiel Y, Chicheportiche A, Keidar Z, and Ben-Haim S. Prostate cancer radioligand therapy: beta-labeled radiopharmaceuticals. PET Clin. (2024) 19:389–99. doi: 10.1016/j.cpet.2024.03.011
6. Kim JS, Chung JH, Song W, Kang M, Sung HH, Jeon HG, et al. Risk factors for prostate-specific antigen persistence in pT3aN0 prostate cancer after robot-assisted laparoscopic radical prostatectomy: a retrospective study. J Yeungnam Med Sci. (2023) 40:412–8. doi: 10.12701/jyms.2023.00234
7. Dathathri E, Isebia KT, Abali F, Lolkema MP, Martens JWM, Terstappen L, et al. Liquid biopsy based circulating biomarkers in metastatic prostate cancer. Front Oncol. (2022) 12:863472. doi: 10.3389/fonc.2022.863472
8. Zhou R and Xu CL. Application effect of case management nursing based on patient safety in patients with prostate cancer. World J Clin cases. (2024) 12:6070–6. doi: 10.12998/wjcc.v12.i27.6070
9. Shi L, Sheng J, Wang M, Luo H, Zhu J, Zhang B, et al. Combination therapy of TGF-β Blockade and commensal-derived probiotics provides enhanced antitumor immune response and tumor suppression. Theranostics. (2019) 9:4115–29. doi: 10.7150/thno.35131
10. Merlo LM, Pepper JW, Reid BJ, and Maley CC. Cancer as an evolutionary and ecological process. Nat Rev Cancer. (2006) 6:924–35. doi: 10.1038/nrc2013
11. Bian X, Wang W, Abudurexiti M, Zhang X, Ma W, Shi G, et al. Integration analysis of single-cell multi-omics reveals prostate cancer heterogeneity. Adv Sci (Weinh). (2024) 11:e2305724. doi: 10.1002/advs.202305724
12. Xin S, Liu X, Li Z, Sun X, Wang R, Zhang Z, et al. ScRNA-seq revealed an immunosuppression state and tumor microenvironment heterogeneity related to lymph node metastasis in prostate cancer. Exp Hematol Oncol. (2023) 12:49. doi: 10.1186/s40164-023-00407-0
13. Peng C, Xu Y, Wu J, Wu D, Zhou L, and Xia X. TME-related biomimetic strategies against cancer. Int J Nanomed. (2024) 19:109–35. doi: 10.2147/IJN.S441135
14. Xiao Y and Yu D. Tumor microenvironment as a therapeutic target in cancer. Pharmacol Ther. (2021) 221:107753. doi: 10.1016/j.pharmthera.2020.107753
15. Pathania AS and Challagundla KB. Exosomal long non-coding RNAs: emerging players in the tumor microenvironment. Mol Ther Nucleic Acids. (2021) 23:1371–83. doi: 10.1016/j.omtn.2020.09.039
16. Wang D, Zhang W, Zhang C, Wang L, Chen H, and Xu J. Exosomal non-coding RNAs have a significant effect on tumor metastasis. Mol Ther Nucleic Acids. (2022) 29:16–35. doi: 10.1016/j.omtn.2022.05.034
17. Liu Q-W, He Y, and Xu WW. Molecular functions and therapeutic applications of exosomal noncoding RNAs in cancer. Exp Mol Med. (2022) 54:216–25. doi: 10.1038/s12276-022-00744-w
18. Rubinstein E, Théry C, and Zimmermann P. Tetraspanins affect membrane structures and the trafficking of molecular partners: what impact on extracellular vesicles? Biochem Soc Trans. (2025) 0:371–82. doi: 10.1042/BST20240523
19. Bei Y, Das S, Rodosthenous RS, Holvoet P, Vanhaverbeke M, Monteiro MC, et al. Extracellular vesicles in cardiovascular theranostics. Theranostics. (2017) 7:4168–82. doi: 10.7150/thno.21274
20. Luo JQ, Yang TW, Wu J, Lai HH, Zou LB, Chen WB, et al. Exosomal PGAM1 promotes prostate cancer angiogenesis and metastasis by interacting with ACTG1. Cell Death Dis. (2023) 14:502. doi: 10.1038/s41419-023-06007-4
21. Ma S, Wang Z, Xiong Z, Ge Y, Xu MY, Zhang J, et al. Enhancer transcription profiling reveals an enhancer RNA-driven ferroptosis and new therapeutic opportunities in prostate cancer. Signal Transduct Target Ther. (2025) 10:87. doi: 10.1038/s41392-025-02170-6
22. Tang S, Cheng H, Zang X, Tian J, Ling Z, Wang L, et al. Small extracellular vesicles: crucial mediators for prostate cancer. J Nanobiotechnol. (2025) 23:230. doi: 10.1186/s12951-025-03326-w
23. Vlaeminck-Guillem V. Exosomes and prostate cancer management. Semin Cancer Biol. (2021) 86:101–11. doi: 10.1016/j.semcancer.2021.08.004
24. Ding L, Zheng Q, Lin Y, Wang R, Wang H, Luo W, et al. Exosome-derived circTFDP2 promotes prostate cancer progression by preventing PARP1 from caspase-3-dependent cleavage. Clin Transl Med. (2023) 13:e1156. doi: 10.1002/ctm2.1156
25. Jiang Y, Zhao H, Chen Y, Li K, Li T, Chen J, et al. Exosomal long noncoding RNA HOXD-AS1 promotes prostate cancer metastasis via miR-361-5p/FOXM1 axis. Cell Death Dis. (2021) 12:1129. doi: 10.1038/s41419-021-04421-0
26. Yang Z, Luo Y, Zhang F, and Ma L. Exosome-derived lncRNA A1BG-AS1 attenuates the progression of prostate cancer depending on ZC3H13-mediated m6A modification. Cell Div. (2024) 19:5. doi: 10.1186/s13008-024-00110-4
27. Wei Y, Chen Z, Zhang R, Wu B, Lin L, Zhu Q, et al. Blood circulating exosomes carrying microRNA-423-5p regulates cell progression in prostate cancer via targeting FRMD3. J Cancer. (2022) 13:2970–81. doi: 10.7150/jca.71706
28. Zhang G, Liu Y, Yang J, Wang H, and Xing Z. Inhibition of circ_0081234 reduces prostate cancer tumor growth and metastasis via the miR-1/MAP 3 K1 axis. J Gene Med. (2022) 24:e3376. doi: 10.1002/jgm.3376
29. Tang Y, Liu J, Li X, and Wang W. Exosomal circRNA HIPK3 knockdown inhibited cell proliferation and metastasis in prostate cancer by regulating miR-212/BMI-1 pathway. J Biosci. (2021) 46:69. doi: 10.1007/s12038-021-00190-2
30. Jiang X, Xu Y, Liu R, and Guo S. Exosomal lincROR Promotes Docetaxel Resistance in Prostate Cancer through a β-catenin/HIF1α Positive Feedback Loop. Mol Cancer Res. (2023) 21:472–82. doi: 10.1158/1541-7786.MCR-22-0458
31. Guo Z, Lu X, Yang F, He C, Qin L, Yang N, et al. Exosomal LINC01213 plays a role in the transition of androgen-dependent prostate cancer cells into androgen-independent manners. J Oncol. (2022) 2022:8058770. doi: 10.1155/2022/8058770
32. Wang C, Zhou C, Wang D, Zhang YF, Lv HX, He H, et al. Proangiogenic potential of plasma exosomes from prostate cancer patients. Cell Signal. (2024) 124:111398. doi: 10.1016/j.cellsig.2024.111398
33. Zeng Q, Liu CH, Ampuero J, Wu D, Jiang W, Zhou L, et al. Circular RNAs in non-alcoholic fatty liver disease: Functions and clinical significance. RNA Biol. (2024) 21:1–15. doi: 10.1080/15476286.2023.2290769
34. Rezaei M, Esmailzadeh A, and Shanei A. Bystander effect of therapeutic ultrasound in the presence of cisplatin: an in vitro study on human melanoma cells. J BioMed Phys Eng. (2023) 13:433–42. doi: 10.31661/jbpe.v0i0.2105-1337
35. Giugliano F, Corti C, Tarantino P, Michelini F, and Curigliano G. Bystander effect of antibody-drug conjugates: fact or fiction? Curr Oncol Rep. (2022) 24:809–17. doi: 10.1007/s11912-022-01266-4
36. Xu Z, Chen Y, Ma L, Chen Y, Liu J, Guo Y, et al. Role of exosomal non-coding RNAs from tumor cells and tumor-associated macrophages in the tumor microenvironment. Mol Ther. (2022) 30:3133–54. doi: 10.1016/j.ymthe.2022.01.046
37. Dai Y and Gao X. Inhibition of cancer cell-derived exosomal microRNA-183 suppresses cell growth and metastasis in prostate cancer by upregulating TPM1. Cancer Cell Int. (2021) 21:145. doi: 10.1186/s12935-020-01686-x
38. Huang G, Jiang Z, Zhu W, and Wu Z. Exosomal circKDM4A Induces CUL4B to Promote Prostate Cancer Cell Malignancy in a miR-338-3p-Dependent Manner. Biochem Genet. (2023) 61:390–409. doi: 10.1007/s10528-022-10251-2
39. Liu Z, Jin Q, Yan T, Wo Y, Liu H, and Wang Y. Exosome-mediated transduction of mechanical force regulates prostate cancer migration via microRNA. Biochem Biophys Rep. (2022) 31:101299. doi: 10.1016/j.bbrep.2022.101299
40. Angerilli V, Fontana E, Lonardi S, Sbaraglia M, Borelli B, Munari G, et al. Intratumor morphologic and transcriptomic heterogeneity in (V600E)BRAF-mutated metastatic colorectal adenocarcinomas. ESMO Open. (2021) 6:100211. doi: 10.1016/j.esmoop.2021.100211
41. Shirshin EA, Shirmanova MV, Gayer AV, Lukina MM, Nikonova EE, Yakimov BP, et al. Label-free sensing of cells with fluorescence lifetime imaging: The quest for metabolic heterogeneity. Proc Natl Acad Sci U.S.A. (2022) 119:e2118241119. doi: 10.1073/pnas.2118241119
42. Li Y, Van Alsten SC, Lee DN, Kim T, Calhoun BC, Perou CM, et al. Visual intratumor heterogeneity and breast tumor progression. Cancers (Basel). (2024) 16:2294. doi: 10.3390/cancers16132294
43. Bertoli G, Panio A, Cava C, Gallivanone F, Alini M, Strano G, et al. Secreted miR-153 controls proliferation and invasion of higher gleason score prostate cancer. Int J Mol Sci. (2022) 23:6339. doi: 10.3390/ijms23116339
44. Cruz-Burgos M, Cortés-Ramírez SA, Losada-García A, Morales-Pacheco M, Martínez-Martínez E, Morales-Montor JG, et al. Unraveling the role of EV-derived miR-150-5p in prostate cancer metastasis and its association with high-grade gleason scores: implications for diagnosis. Cancers (Basel). (2023) 15:4148. doi: 10.3390/cancers15164148
45. Albino D, Falcione M, Uboldi V, Temilola DO, Sandrini G, Merulla J, et al. Circulating extracellular vesicles release oncogenic miR-424 in experimental models and patients with aggressive prostate cancer. Commun Biol. (2021) 4:119. doi: 10.1038/s42003-020-01642-5
46. Wang W, Kong P, Feng K, Liu C, Gong X, Sun T, et al. Exosomal miR-222-3p contributes to castration-resistant prostate cancer by activating mTOR signaling. Cancer Sci. (2023) 114:4252–69. doi: 10.1111/cas.15948
47. Gan J, Liu S, Zhang Y, He L, Bai L, Liao R, et al. MicroRNA-375 is a therapeutic target for castration-resistant prostate cancer through the PTPN4/STAT3 axis. Exp Mol Med. (2022) 54:1290–305. doi: 10.1038/s12276-022-00837-6
48. Lu J, Zou Q, Li Y, Xiong C, Tao L, Wu J, et al. FTH1P8 induces and transmits docetaxel resistance by inhibiting ferroptosis in prostate cancer. BioMed Pharmacother. (2024) 180:117472. doi: 10.1016/j.biopha.2024.117472
49. Tan X, Song X, Fan B, Li M, Zhang A, and Pei L. Exosomal circRNA Scm-like with four Malignant brain tumor domains 2 (circ-SFMBT2) enhances the docetaxel resistance of prostate cancer via the microRNA-136-5p/tribbles homolog 1 pathway. Anticancer Drugs. (2022) 33:871–82. doi: 10.1097/CAD.0000000000001365
50. Zhang H, Li M, Zhang J, Shen Y, and Gui Q. Exosomal circ-XIAP promotes docetaxel resistance in prostate cancer by regulating miR-1182/TPD52 axis. Drug Des Devel Ther. (2021) 15:1835–49. doi: 10.2147/DDDT.S300376
51. Lin A, Li J, and He W. CircSLC4A7 in resistant-cells-derived exosomes promotes docetaxel resistance via the miR-1205/MAPT axis in prostate cancer. IUBMB Life. (2024) 76:1342–55. doi: 10.1002/iub.2915
52. Zhang Y, Chen B, Xu N, Xu P, Lin W, Liu C, et al. Exosomes promote the transition of androgen-dependent prostate cancer cells into androgen-independent manner through up-regulating the heme oxygenase-1. Int J Nanomed. (2021) 16:315–27. doi: 10.2147/IJN.S281710
53. Martens-Uzunova ES, Kusuma GD, Crucitta S, Lim HK, Cooper C, Riches JE, et al. Androgens alter the heterogeneity of small extracellular vesicles and the small RNA cargo in prostate cancer. J Extracell Vesicles. (2021) 10:e12136. doi: 10.1002/jev2.12136
54. Guo T, Wang Y, Jia J, Mao X, Stankiewicz E, Scandura G, et al. The Identification of Plasma Exosomal miR-423-3p as a Potential Predictive Biomarker for Prostate Cancer Castration-Resistance Development by Plasma Exosomal miRNA Sequencing. Front Cell Dev Biol. (2020) 8:602493. doi: 10.3389/fcell.2020.602493
55. Lei L, Yu L, Fan W, and Hao X. Prostate cancer small extracellular vesicles participate in androgen-independent transformation of prostate cancer by transferring let-7a-5p. Heliyon. (2022) 8:e12114. doi: 10.1016/j.heliyon.2022.e12114
56. Zhang N, Hao J, Cai Y, and Wang M. Research advances of secretory proteins in Malignant tumors. Chin J Cancer Res. (2021) 33:115–32. doi: 10.21147/j.issn.1000-9604.2021.01.12
57. Sleeboom JJF, van Tienderen GS, Schenke-Layland K, van der Laan LJW, Khalil AA, and Verstegen MMA. The extracellular matrix as hallmark of cancer and metastasis: From biomechanics to therapeutic targets. Sci Transl Med. (2024) 16:eadg3840. doi: 10.1126/scitranslmed.adg3840
58. Kazakova A, Sudarskikh T, Kovalev O, Kzhyshkowska J, and Larionova I. Interaction of tumor−associated macrophages with stromal and immune components in solid tumors: Research progress (Review). Int J Oncol. (2023) 62:32. doi: 10.3892/ijo.2023.5480
59. Zhang W, Liu M, Li W, and Song Y. Immune cells in the B-cell lymphoma microenvironment: From basic research to clinical applications. Chin Med J (Engl). (2024) 137:776–90. doi: 10.1097/CM9.0000000000002919
60. Gao T, Yuan D, He B, Gao Y, Liu C, Sun H, et al. Identification of autophagy related genes in predicting the prognosis and aiding 5- fluorouracil therapy of colorectal cancer. Heliyon. (2022) 8:e09033. doi: 10.1016/j.heliyon.2022.e09033
61. Chen QY, Gao B, Tong D, and Huang C. Crosstalk between extracellular vesicles and tumor-associated macrophage in the tumor microenvironment. Cancer Lett. (2023) 552:215979. doi: 10.1016/j.canlet.2022.215979
62. Li T, Chen D, Liu H, Tao Y, He X, Zang S, et al. Spatially targeting and regulating tumor-associated macrophages using a raspberry-like micellar system sensitizes pancreatic cancer chemoimmunotherapy. Nanoscale. (2022) 14:13098–112. doi: 10.1039/D2NR03053E
63. Naser R, Fakhoury I, El-Fouani A, Abi-Habib R, and El-Sibai M. Role of the tumor microenvironment in cancer hallmarks and targeted therapy (Review). Int J Oncol. (2023) 62:23. doi: 10.3892/ijo.2022.5471
64. Zhang X, Shi X, Xie F, Liu Y, Wei X, Cai Y, et al. Dissecting pulmonary fibroblasts heterogeneity in lung development, health and diseases. Heliyon. (2023) 9:e19428. doi: 10.1016/j.heliyon.2023.e19428
65. Xu Y, Li W, Lin S, Liu B, Wu P, and Li L. Fibroblast diversity and plasticity in the tumor microenvironment: roles in immunity and relevant therapies. Cell Commun Signal. (2023) 21:234. doi: 10.1186/s12964-023-01204-2
66. Cheng K, Li W, Wu H, and Li C. Mapping knowledge structure and themes trends of cancer-associated fibroblasts: a text-mining study. Front Mol Biosci. (2023) 10:1302016. doi: 10.3389/fmolb.2023.1302016
67. Zhao J, Shen J, Mao L, Yang T, Liu J, and Hongbin S. Cancer associated fibroblast secreted miR-432-5p targets CHAC1 to inhibit ferroptosis and promote acquired chemoresistance in prostate cancer. Oncogene. (2024) 43:2104–14. doi: 10.1038/s41388-024-03057-6
68. Shan G, Gu J, Zhou D, Li L, Cheng W, Wang Y, et al. Cancer-associated fibroblast-secreted exosomal miR-423-5p promotes chemotherapy resistance in prostate cancer by targeting GREM2 through the TGF-β signaling pathway. Exp Mol Med. (2020) 52:1809–22. doi: 10.1038/s12276-020-0431-z
69. Zhang Y, Zhao J, Ding M, Su Y, Cui D, Jiang C, et al. Loss of exosomal miR-146a-5p from cancer-associated fibroblasts after androgen deprivation therapy contributes to prostate cancer metastasis. J Exp Clin Cancer Res. (2020) 39:282. doi: 10.1186/s13046-020-01761-1
70. Matsuda C, Ishii K, Nakagawa Y, Shirai T, Sasaki T, Hirokawa YS, et al. Fibroblast-derived exosomal microRNA regulates NKX3–1 expression in androgen-sensitive, androgen receptor-dependent prostate cancer cells. J Cell Biochem. (2023) 124:1135–44. doi: 10.1002/jcb.30435
71. Liu Z, Lin Z, Jiang M, Zhu G, Xiong T, Cao F, et al. Cancer-associated fibroblast exosomes promote prostate cancer metastasis through miR-500a-3p/FBXW7/HSF1 axis under hypoxic microenvironment. Cancer Gene Ther. (2024) 31:698–709. doi: 10.1038/s41417-024-00742-2
72. Wang P, Zhang S, Meng Q, Zhu P, and Yuan W. Treatment and application of stem cells from different sources for cartilage injury: a literature review. Ann Transl Med. (2022) 10:610. doi: 10.21037/atm-22-1715
73. Park A, Choi S, Do J, Kim Y, Kim KS, Koh E, et al. ZO-1 regulates the migration of mesenchymal stem cells in cooperation with α-catenin in response to breast tumor cells. Cell Death Discov. (2024) 10:19. doi: 10.1038/s41420-023-01793-4
74. Huang J, Wang X, Wen J, Zhao X, Wu C, Wang L, et al. Gastric cancer cell-originated small extracellular vesicle induces metabolic reprogramming of BM-MSCs through ERK-PPARγ-CPT1A signaling to potentiate lymphatic metastasis. Cancer Cell Int. (2023) 23:87. doi: 10.1186/s12935-023-02935-5
75. Zhang L. The role of mesenchymal stem cells in modulating the breast cancer microenvironment. Cell Transplant. (2023) 32:9636897231220073. doi: 10.1177/09636897231220073
76. Zhang L, Xiang J, Zhang F, Liu L, and Hu C. MSCs can be a double-edged sword in tumorigenesis. Front Oncol. (2022) 12:1047907. doi: 10.3389/fonc.2022.1047907
77. Yao J, Sun L, Gao F, and Zhu W. Mesenchymal stem/stromal cells: dedicator to maintain tumor homeostasis. Hum Cell. (2024) 38:21. doi: 10.1007/s13577-024-01154-y
78. Li C, Sun Z, Song Y, and Zhang Y. Suppressive function of bone marrow-derived mesenchymal stem cell-derived exosomal microRNA-187 in prostate cancer. Cancer Biol Ther. (2022) 23:1–14. doi: 10.1080/15384047.2022.2123675
79. Jiang S, Chen H, He K, and Wang J. Human bone marrow mesenchymal stem cells-derived exosomes attenuated prostate cancer progression via the miR-99b-5p/IGF1R axis. Bioengineered. (2022) 13:2004–16. doi: 10.1080/21655979.2021.2009416
80. Kurniawati I, Liu MC, Hsieh CL, Do AD, and Sung SY. Targeting castration-resistant prostate cancer using mesenchymal stem cell exosomes for therapeutic microRNA-let-7c delivery. Front Biosci (Landmark Ed). (2022) 27:256. doi: 10.31083/j.fbl2709256
81. Aliazis K, Christofides A, Shah R, Yeo YY, Jiang S, Charest A, et al. The tumor microenvironment's role in the response to immune checkpoint blockade. Nat Cancer. (2025) 6:924–37. doi: 10.1038/s43018-025-00986-3
82. Prigol AN, Rode MP, Silva AH, Cisilotto J, and Creczynski-Pasa TB. Pro-angiogenic effect of PC-3 exosomes in endothelial cells in vitro. Cell Signal. (2021) 87:110126. doi: 10.1016/j.cellsig.2021.110126
83. Sun W, Hu S, and Wang X. Advances and clinical applications of immune checkpoint inhibitors in hematological Malignancies. Cancer Commun (Lond). (2024) 44:1071–97. doi: 10.1002/cac2.12587
84. Wang L, Guo W, Guo Z, Yu J, Tan J, Simons DL, et al. PD-L1-expressing tumor-associated macrophages are immunostimulatory and associate with good clinical outcome in human breast cancer. Cell Rep Med. (2024) 5:101420. doi: 10.1016/j.xcrm.2024.101420
85. Zhu S, Wang Y, Tang J, and Cao M. Radiotherapy induced immunogenic cell death by remodeling tumor immune microenvironment. Front Immunol. (2022) 13:1074477. doi: 10.3389/fimmu.2022.1074477
86. Bai R and Cui J. Development of immunotherapy strategies targeting tumor microenvironment is fiercely ongoing. Front Immunol. (2022) 13:890166. doi: 10.3389/fimmu.2022.890166
87. Kim H, Park HJ, Chang HW, Back JH, Lee SJ, Park YE, et al. Exosome-guided direct reprogramming of tumor-associated macrophages from protumorigenic to antitumorigenic to fight cancer. Bioact Mater. (2023) 25:527–40. doi: 10.1016/j.bioactmat.2022.07.021
88. Jin Y, Huang Y, Ren H, Huang H, Lai C, Wang W, et al. Nano-enhanced immunotherapy: Targeting the immunosuppressive tumor microenvironment. Biomaterials. (2024) 305:122463. doi: 10.1016/j.biomaterials.2023.122463
89. Kalkusova K, Smite S, Darras E, Taborska P, Stakheev D, Vannucci L, et al. Mast cells and dendritic cells as cellular immune checkpoints in immunotherapy of solid tumors. Int J Mol Sci. (2022) 23:11080. doi: 10.3390/ijms231911080
90. Xu P, Yang JC, Chen B, Nip C, Van Dyke JE, Zhang X, et al. Androgen receptor blockade resistance with enzalutamide in prostate cancer results in immunosuppressive alterations in the tumor immune microenvironment. J Immunother Cancer. (2023) 11:e006581. doi: 10.1136/jitc-2022-006581
91. Koerner J, Horvath D, Oliveri F, Li J, and Basler M. Suppression of prostate cancer and amelioration of the immunosuppressive tumor microenvironment through selective immunoproteasome inhibition. Oncoimmunology. (2023) 12:2156091. doi: 10.1080/2162402X.2022.2156091
92. Wang D, Xing C, Liang Y, Wang C, Zhao P, Liang X, et al. Ultrasound imaging of tumor vascular CD93 with MMRN2 modified microbubbles for immune microenvironment prediction. Adv Mater. (2024) 36:e2310421. doi: 10.1002/adma.202310421
93. Hadimani SM, Das S, and Harish KG. An immunohistochemical evaluation of tumor-associated macrophages (M1 and M2) in carcinoma prostate - An institutional study. J Cancer Res Ther. (2023) 19:S300–s305. doi: 10.4103/jcrt.jcrt_497_22
94. JiaWei Z, ChunXia D, CunDong L, Yang L, JianKun Y, HaiFeng D, et al. M2 subtype tumor associated macrophages (M2-TAMs) infiltration predicts poor response rate of immune checkpoint inhibitors treatment for prostate cancer. Ann Med. (2021) 53:730–40. doi: 10.1080/07853890.2021.1924396
95. Rathi N, McFarland TR, Nussenzveig R, Agarwal N, and Swami U. Evolving role of immunotherapy in metastatic castration refractory prostate cancer. Drugs. (2021) 81:191–206. doi: 10.1007/s40265-020-01456-z
96. Anton A, Hutchinson R, Hovens CM, Christie M, Ryan A, Gibbs P, et al. An immune suppressive tumor microenvironment in primary prostate cancer promotes tumor immune escape. PloS One. (2024) 19:e0301943. doi: 10.1371/journal.pone.0301943
97. Zhang J, Ding X, Peng K, Jia Z, and Yang J. Identification of biomarkers for immunotherapy response in prostate cancer and potential drugs to alleviate immunosuppression. Aging (Albany NY). (2022) 14:4839–57. doi: 10.18632/aging.204115
98. Nourbakhsh NS, Naeimi S, Moghanibashi M, and Baradaran B. Bicalutamide reveals immunomodulatory effects in prostate cancer by regulating immunogenic dendritic cell maturation. Tissue Cell. (2024) 91:102530. doi: 10.1016/j.tice.2024.102530
99. Chetta P, Sriram R, and Zadra G. Lactate as key metabolite in prostate cancer progression: what are the clinical implications? Cancers (Basel). (2023) 15:3473. doi: 10.3390/cancers15133473
100. Tian HY, Liang Q, Shi Z, and Zhao H. Exosomal CXCL14 contributes to M2 macrophage polarization through NF-κB signaling in prostate cancer. Oxid Med Cell Longev. (2022) 2022:7616696. doi: 10.1155/2022/7616696
101. Costanzi E, Romani R, Scarpelli P, and Bellezza I. Extracellular vesicles-mediated transfer of miRNA let-7b from PC3 cells to macrophages. Genes (Basel). (2020) 11:1495. doi: 10.3390/genes11121495
102. Guan H, Tao H, Luo J, Wan L, Hu H, Chen L, et al. Upregulation of YY1 in M2 macrophages promotes secretion of exosomes containing hsa-circ-0000326 via super-enhancers to facilitate prostate cancer progression. Mol Cell Biochem. (2025) 480:3873–88. doi: 10.1007/s11010-025-05222-1
103. Gao F, Xu Q, Tang Z, Zhang N, Huang Y, Li Z, et al. Exosomes derived from myeloid-derived suppressor cells facilitate castration-resistant prostate cancer progression via S100A9/circMID1/miR-506-3p/MID1. J Transl Med. (2022) 20:346. doi: 10.1186/s12967-022-03494-5
104. Guan H, Peng R, Fang F, Mao L, Chen Z, Yang S, et al. Tumor-associated macrophages promote prostate cancer progression via exosome-mediated miR-95 transfer. J Cell Physiol. (2020) 235:9729–42. doi: 10.1002/jcp.29784
105. Shi Z, Pu W, Li M, Aihemaitijiang M, Li S, Zhang X, et al. Prostate cancer cell-derived exosomes ZNF667-AS1 reduces TGFBR1 mRNA stability to inhibit Treg expansion and DTX resistance by binding to U2AF1. Mol Med. (2024) 30:179. doi: 10.1186/s10020-024-00947-z
106. Zhang LS, Chen QC, Zong HT, and Xia Q. Exosome miRNA-203 promotes M1 macrophage polarization and inhibits prostate cancer tumor progression. Mol Cell Biochem. (2024) 479:2459–70. doi: 10.1007/s11010-023-04854-5
107. Archer Goode E, Wang N, and Munkley J. Prostate cancer bone metastases biology and clinical management (Review). Oncol Lett. (2023) 25:163. doi: 10.3892/ol.2023.13749
108. Verbruggen SW. Role of the osteocyte in bone metastasis - The importance of networking. J Bone Oncol. (2024) 44:100526. doi: 10.1016/j.jbo.2024.100526
109. Li Y, Liu F, Cai Q, Deng L, Ouyang Q, Zhang XH, et al. Invasion and metastasis in cancer: molecular insights and therapeutic targets. Signal Transduct Target Ther. (2025) 10:57. doi: 10.1038/s41392-025-02148-4
110. Mohammed SI, Torres-Luquis O, Zhou W, Lanman NA, Espina V, and Liotta L. Tumor-draining lymph secretome en route to the regional lymph node in breast cancer metastasis. Breast Cancer (Dove Med Press). (2020) 12:57–67. doi: 10.2147/BCTT.S236168
111. Byrne NM, Summers MA, and McDonald MM. Tumor cell dormancy and reactivation in bone: skeletal biology and therapeutic opportunities. JBMR Plus. (2019) 3:e10125. doi: 10.1002/jbm4.10125
112. Dai R, Liu M, Xiang X, Xi Z, and Xu H. Osteoblasts and osteoclasts: an important switch of tumour cell dormancy during bone metastasis. J Exp Clin Cancer Res. (2022) 41:316. doi: 10.1186/s13046-022-02520-0
113. Ahamad N and Singh BB. Calcium channels and their role in regenerative medicine. World J Stem Cells. (2021) 13:260–80. doi: 10.4252/wjsc.v13.i4.260
114. Tae JH and Chang IH. Animal models of bone metastatic prostate cancer. Investig Clin Urol. (2023) 64:219–28. doi: 10.4111/icu.20230026
115. Joseph GJ, Johnson DB, and Johnson RW. Immune checkpoint inhibitors in bone metastasis: Clinical challenges, toxicities, and mechanisms. J Bone Oncol. (2023) 43:100505. doi: 10.1016/j.jbo.2023.100505
116. Kang Y. Dissecting tumor-stromal interactions in breast cancer bone metastasis. Endocrinol Metab (Seoul). (2016) 31:206–12. doi: 10.3803/EnM.2016.31.2.206
117. Juárez P and Guise TA. TGF-β in cancer and bone: implications for treatment of bone metastases. Bone. (2011) 48:23–9. doi: 10.1016/j.bone.2010.08.004
118. Mo C, Huang B, Zhuang J, Jiang S, Guo S, and Mao X. LncRNA nuclear-enriched abundant transcript 1 shuttled by prostate cancer cells-secreted exosomes initiates osteoblastic phenotypes in the bone metastatic microenvironment via miR-205-5p/runt-related transcription factor 2/splicing factor proline- and glutamine-rich/polypyrimidine tract-binding protein 2 axis. Clin Transl Med. (2021) 11:e493. doi: 10.1002/ctm2.493
119. Liu Y, Yang C, Chen S, Liu W, Liang J, He S, et al. Cancer-derived exosomal miR-375 targets DIP2C and promotes osteoblastic metastasis and prostate cancer progression by regulating the Wnt signaling pathway. Cancer Gene Ther. (2023) 30:437–49. doi: 10.1038/s41417-022-00563-1
120. Zou Z, Dai R, Deng N, Su W, and Liu P. Exosomal miR-1275 secreted by prostate cancer cells modulates osteoblast proliferation and activity by targeting the SIRT2/RUNX2 cascade. Cell Transplant. (2021) 30:9636897211052977. doi: 10.1177/09636897211052977
121. Furesi G, de Jesus Domingues AM, Alexopoulou D, Dahl A, Hackl M, Schmidt JR, et al. Exosomal miRNAs from prostate cancer impair osteoblast function in mice. Int J Mol Sci. (2022) 23:1285. doi: 10.3390/ijms23031285
122. Yu L, Sui B, Fan W, Lei L, Zhou L, Yang L, et al. Exosomes derived from osteogenic tumor activate osteoclast differentiation and concurrently inhibit osteogenesis by transferring COL1A1-targeting miRNA-92a-1-5p. J Extracell Vesicles. (2021) 10:e12056. doi: 10.1002/jev2.12056
123. Tian G, Hu K, Qiu S, Xie Y, Cao Y, Ni S, et al. Exosomes derived from PC-3 cells suppress osteoclast differentiation by downregulating miR-148a and blocking the PI3K/AKT/mTOR pathway. Exp Ther Med. (2021) 22:1304. doi: 10.3892/etm.2021.10739
124. Liu Y, Chen S, Guo K, Ma S, Wang X, Liu Q, et al. Osteoblast-derived exosomal miR-140-3p targets ACER2 and increases the progression of prostate cancer via the AKT/mTOR pathway-mediated inhibition of autophagy. FASEB J. (2024) 38:e70206. doi: 10.1096/fj.202401480R
125. Yang Z, Chen JQ, Liu TJ, Chen YL, Ma ZK, Fan YZ, et al. Knocking down AR promotes osteoblasts to recruit prostate cancer cells by altering exosomal circ-DHPS/miR-214-3p/CCL5 pathway. Asian J Androl. (2024) 26:195–204. doi: 10.4103/aja202351
126. Li Q, Hu J, Shi Y, Xiao M, Bi T, Wang C, et al. Exosomal lncAY927529 enhances prostate cancer cell proliferation and invasion through regulating bone microenvironment. Cell Cycle. (2021) 20:2531–46. doi: 10.1080/15384101.2021.1992853
127. Chen N, Qian J, Wu D, Chen W, Mao J, Cai Y, et al. Propofol mediates bone metastasis by regulating PC-derived exosomal miR-142-3p. Bull Cancer. (2023) 110:265–74. doi: 10.1016/j.bulcan.2023.01.008
128. Lu Z, Hou J, Li X, Zhou J, Luo B, Liang S, et al. Exosome-derived miRNAs as potential biomarkers for prostate bone metastasis. Int J Gen Med. (2022) 15:5369–83. doi: 10.2147/IJGM.S361981
129. Yin X, Zheng X, Liu M, Wang D, Sun H, Qiu Y, et al. Exosomal miR-663b targets Ets2-repressor factor to promote proliferation and the epithelial-mesenchymal transition of bladder cancer cells. Cell Biol Int. (2020) 44:958–65. doi: 10.1002/cbin.11292
130. Ninio-Many L, Grossman H, Levi M, Zilber S, Tsarfaty I, Shomron N, et al. MicroRNA miR-125a-3p modulates molecular pathway of motility and migration in prostate cancer cells. Oncoscience. (2014) 1:250–61. doi: 10.18632/oncoscience.30
131. Iqbal MA, Arora S, Prakasam G, Calin GA, and Syed MA. MicroRNA in lung cancer: role, mechanisms, pathways and therapeutic relevance. Mol Aspects Med. (2018) 70:3–20. doi: 10.1016/j.mam.2018.07.003
132. Mirzaei S, Paskeh MDA, Okina E, Gholami MH, Hushmandi K, Hashemi M, et al. Molecular Landscape of LncRNAs in Prostate Cancer: A focus on pathways and therapeutic targets for intervention. J Exp Clin Cancer Res: CR. (2022) 41:214. doi: 10.1186/s13046-022-02406-1
133. Zhang B, Zhang H, Wang Z, Cao H, Zhang N, Dai Z, et al. The regulatory role and clinical application prospects of circRNA in the occurrence and development of CNS tumors. CNS Neurosci Ther. (2023) 30:e14500. doi: 10.1111/cns.14500
134. Jahromi FNA, Dowran R, and Jafari R. Recent advances in the roles of exosomal microRNAs (exomiRs) in hematologic neoplasms: pathogenesis, diagnosis, and treatment. Cell Commun Signaling: CCS. (2023) 21:88. doi: 10.1186/s12964-023-01102-7
135. Huang Q, Zhong X, Li J, Hu R, Yi J, Sun J, et al. Exosomal ncRNAs: Multifunctional contributors to the immunosuppressive tumor microenvironment of hepatocellular carcinoma. Biomed Pharmacother = Biomed Pharmacother. (2024) 173:116409. doi: 10.1016/j.biopha.2024.116409
136. Oshiumi H. Circulating extracellular vesicles carry immune regulatory miRNAs and regulate vaccine efficacy and local inflammatory response after vaccination. Front In Immunol. (2021) 12:685344. doi: 10.3389/fimmu.2021.685344
137. Pan J-H, Zhou H, Zhao X-X, Ding H, Li W, Qin L, et al. Role of exosomes and exosomal microRNAs in hepatocellular carcinoma: Potential in diagnosis and antitumour treatments (Review). Int J Mol Med. (2018) 41:1809–16. doi: 10.3892/ijmm.2018.3383
138. Rijavec E, Coco S, Genova C, Rossi G, Longo L, and Grossi F. Liquid biopsy in non-small cell lung cancer: highlights and challenges. Cancers. (2019) 12:17. doi: 10.3390/cancers12010017
139. Zanin L, Sachkova A, Panciani PP, Rohde V, Fontanella MM, and Schatlo B. Liquid biopsy in low-grade glioma: A systematic review and a proposal for a clinical utility score. Cell Mol Neurobiol. (2023) 43:3833–45. doi: 10.1007/s10571-023-01406-9
140. Khawar MB, Abbasi MH, Siddique Z, Arif A, and Sheikh N. An update on novel therapeutic warfronts of extracellular vesicles (EVs) in cancer treatment: where we are standing right now and where to go in the future. Oxid Med Cell Longevity. (2019) 2019:9702562. doi: 10.1155/2019/9702562
Keywords: exosomes, non-coding RNA, prostate cancer, tumor microenvironment, immune microenvironment
Citation: Duan H, Xu B, Luo P, Chen T and Zou J (2025) Exosomal non-coding RNAs: orchestrators of intercellular crosstalk in the prostate cancer tumor microenvironment. Front. Immunol. 16:1644861. doi: 10.3389/fimmu.2025.1644861
Received: 11 June 2025; Accepted: 04 November 2025; Revised: 26 October 2025;
Published: 21 November 2025.
Edited by:
Xudong Zhu, Cancer Hospital of Dalian University of Technology (Liaoning Cancer Institute and Hospital), ChinaReviewed by:
Rahul Shivahare, The Ohio State University, United StatesAniket Mali, Bharati Vidyapeeth Deemed University, India
Copyright © 2025 Duan, Xu, Luo, Chen and Zou. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Huanglin Duan, MTU3Nzk4MzExNzlAMTYzLmNvbQ==
†These authors have contributed equally to this work and share first authorship
 Huanglin Duan
Huanglin Duan Baisheng Xu1†
Baisheng Xu1† Peiyue Luo
Peiyue Luo Tao Chen
Tao Chen Jun Zou
Jun Zou