- Department of Microbiology and Immunology, University of Texas Health Science Center at San Antonio, San Antonio, TX, USA
The human pathogen Chlamydia trachomatis secretes numerous effectors into host cells in order to successfully establish and complete the intracellular growth cycle. Three C. trachomatis proteases [chlamydial proteasome/protease-like activity factor (CPAF), tail-specific protease (Tsp), and chlamydial high temperature requirement protein A (cHtrA)] have been localized in the cytosol of the infected cells either by direct immunofluorescence visualization or functional implication. Both CPAF and Tsp have been found to play important roles in C. trachomatis interactions with host cells although the cellular targets of cHtrA have not been identified. All three proteases contain a putative N-terminal signal sequence, suggesting that they may be secreted via a sec-dependent pathway. However, these proteases are also found in chlamydial organism-free vesicles in the lumen of the chlamydial inclusions before they are secreted into host cell cytosol, suggesting that these proteases may first be translocated into the periplasmic region via a sec-dependent pathway and then exported outside of the organisms via an outer membrane vesicles (OMVs) budding mechanism. The vesiculized proteases in the inclusion lumen can finally enter host cell cytosol via vesicle fusing with or passing through the inclusion membrane. Continuing identification and characterization of the C. trachomatis-secreted proteins (CtSPs) will not only promote our understanding of C. trachomatis pathogenic mechanisms but also allow us to gain novel insights into the OMV pathway, a well-known mechanism used by bacteria to export virulence factors although its mechanism remains elusive.
Introduction
Chlamydia trachomatis is the most frequently reported bacterial sexually transmitted infection in the US (Centers for Disease Control Prevention, 2009), which, if untreated, can lead to severe complications characterized with inflammatory pathologies, including pelvic inflammatory diseases, ectopic pregnancy, and infertility (Land et al., 2010). The chlamydial intracellular replication is thought to significantly contribute to the C. trachomatis-induced inflammatory pathologies (Stephens, 2003; Wyrick, 2010). A typical chlamydial replication cycle starts with the invasion of an epithelial cell with a chlamydial infectious elementary body (EB), which is facilitated by chlamydial injection of preexisting effectors into the epithelial cell to induce and modulate endocytosis (Clifton et al., 2004, 2005; Engel, 2004; Hower et al., 2009). Once internalized, an EB differentiates into a non-infectious but metabolically active reticulate body (RB). The RB makes new proteins not only for multiplication but also for secretion into the inclusion lumen and membrane (Rockey et al., 1995, 2002; Luo et al., 2007a,b,c; Li et al., 2008) as well as host cell cytosol (Valdivia, 2008; Zhong et al., 2009, 2011; Betts-Hampikian and Fields, 2010) via a type III secretion (T3S, Fields and Hackstadt, 2000; Valdivia, 2008; Betts-Hampikian and Fields, 2010), sec-dependent secretion (Chen et al., 2010b), or an autotransporter (Henderson and Lam, 2001; Wehrl et al., 2004; Carlson et al., 2005; Vandahl et al., 2005; Kiselev et al., 2009; Byrne, 2010) pathways. After replication, the progeny RBs differentiate back into EBs for spreading to near-by cells. It is thought that the C. trachomatis-secreted proteins (CtSPs; ref: Valdivia, 2008; Zhong, 2009, 2011; Betts-Hampikian and Fields, 2010) are not only necessary for completing the existing developmental cycle but also essential for ensuring a successful start of subsequent infection cycles. Identification and characterization of CtSPs may provide important knowledge for understanding chlamydial pathogenic mechanisms and improving diagnosis, treatment, and prevention of C. trachomatis infection. Thus, identification of CtSPs has become an intensively investigated topic for chlamydiaologists.
Chlamydia Trachomatis-Secreted Proteases in Pathogenesis
A C. trachomatis genome typically encodes more than two dozens of proteins with proteolytic activity (Stephens et al., 1998). A function-driven approach has led to the identification of a novel serine protease, designated as chlamydial proteasome/protease-like activity factor (CPAF; Zhong et al., 2001). The chlamydial proteases are usually distributed in the chlamydial cytosol, periplasmic region and/or various membranes for fulfilling their respective roles in chlamydial biology. However, some, including CPAF (encoded by ORF CT858), the tail-specific protease (Tsp, CT441), and the chlamydial high temperature requirement protein A protease (cHtrA, CT823), are also found outside of the chlamydial organisms by direct visualization and/or functional implication, suggesting that these proteases may be used by C. trachomatis organisms to target host proteins for manipulating host signaling pathways.
Chlamydial proteasome/protease-like activity factor is an extensively studied Chlamydia-secreted serine protease with a water molecule-mediated catalytic triad consisting of residues H105, S499, and E558. CPAF and it can undergo autoprocessing for activation (Dong et al., 2004a,b; Huang et al., 2008; Chen et al., 2009, 2010a). CPAF has a broad substrate specificity and attacks a wide spectrum of host proteins, including the transcriptional factors USF-1 (Zhong et al., 1999) and RFX5 (Zhong et al., 2000) for potentially evading immune recognition, and HIF-1 (Rupp et al., 2007) for dealing with stress responses, the BH3-only proteins (proapoptotic members of the Bcl-2 family; Fischer et al., 2004; Dong et al., 2005; Pirbhai et al., 2006) for inhibiting apoptosis (Fan et al., 1998; Greene et al., 2004; Xiao et al., 2004, 2005; Zhong et al., 2006), the DNA repairing enzyme PARP [Poly (ADP-ribose) polymerase], and cell cycling proteins (Balsara et al., 2006; Paschen et al., 2008) for altering cell cycle, cytoskeleton proteins (keratins 8 and 18, vimentin Dong et al., 2004c; Kumar and Valdivia, 2008; Savijoki et al., 2008) for promoting inclusion expansion, and even cell surface proteins CD1d (Kawana et al., 2007) and nectin-1 (Sun and Schoborg, 2009). Although the functional consequences of some of the host protein degradation by CPAF remain unknown, the host protein degradation should benefit C. trachomatis intracellular growth (Zhong, 2009).
Although Tsp was not detected outside of inclusions using standard immunofluorescence assays (data not shown), it was found to cleave host NF-κB in C. trachomatis-infected cells (Lad et al., 2007a,b), suggesting that an undetectable amount of Tsp might be secreted into host cell cytosol for interrupting NF-κB function. During C. trachomatis infection, there were conflicting observations: On one hand, C. trachomatis infection activated a wide variety of inflammatory cytokines, including IL-1, IL-6, IL-8, and TNFα (Rasmussen et al., 1997; Cheng et al., 2008); On the other, no significant NF- κB activation was detected in the infected cells (Xiao et al., 2005; Lad et al., 2007a). It turned out that the C. trachomatis organisms used the MAP kinase pathway to promote chlamydial acquisition of host lipids, during which inflammatory responses were inevitably activated (Su et al., 2004). The lack of NF- κB activation was probably due to the chlamydial ability to actively silence the NF- κB inflammatory pathway. NF- κB p65 was cleaved into two major fragments (p40 and p20) by Tsp (Lad et al., 2007a) and CPAF (Christian et al., 2010). Since the N-terminal fragment p40 maintained the ability to interact with I-κBα (a cytoplasmic inhibitor of NF-κB) and to bind to DNA but lacked transactivation capability, the p40 might be able to block the residual full length p65-mediated response via a dominant negative effect (Lad et al., 2007a). Tsp was also reported to interact with the host SRAP1 co-activator of estrogen receptor α (Borth et al., 2010). However, the biological significance of chlamydial Tsp-host SRAP-1 interaction remains unknown.
We recently detected the C. trachomatis periplasmic protease cHtrA in both the chlamydial inclusion lumen and host cell cytosol (Figure 1), suggesting that the chlamydial periplasmic protein cHtrA is also secreted into host cells. The secretion appeared to be specific since no other chlamydial periplasmic proteins including CT067 (Miller et al., 2009) were detected outside of the chlamydial inclusions (Figure 1 and data not shown). Normally, HtrA from eukaryotic and prokaryotic species exhibits both chaperone and proteolytic activities with a broad proteolytic substrate specificity (Huston et al., 2007, 2008). HtrA is a hexamer formed by staggered association of trimeric rings and the access to the proteolytic sites in central cavity is controlled by 12 PDZ domains in the sidewall (Krojer et al., 2002, 2010). In eukaryotic cells, HtrA responds to unfolded proteins in the endoplasmic reticulum (ER) by cleaving and releasing the ER membrane-anchored transcription factors ATF6 and SREBP into nucleus to activate the expression of proteins required for the unfolded protein response and cholesterol biosynthesis (Brown and Goldstein, 1999; Ye et al., 2000). In bacteria, the periplasmic HtrA, in response to the binding of C-terminal peptides from unfolded/reduced outer membrane proteins, cleaves, and releases the σE-factor to activate stress response genes (Walsh et al., 2003). Since HtrA is required for bacterial survival under high temperature, it is called High temperature requirement (Htr) protein (Missiakas et al., 1997). Although both the tertiary structure and function of HtrA are well-known, the role of cHtrA in chlamydial pathogenesis remains unclear. The finding that cHtrA was localized both in the chlamydial inclusion lumenal space and the host cell cytosol suggests that the chlamydial periplasmic cHtrA may also contribute to the chlamydial proteolysis strategies for manipulating host cell signaling pathways. However, it is still unknown how the secreted cHtrA contributes to chlamydial pathogenesis. Can the secreted cHtrA gain access to host cell ER to regulate the host unfolded protein stress responses? What are the cellular targets of the secreted cHtrA during chlamydial infection? Interestingly, HtrA from the human gastric pathogen Helicobacter pylori is also secreted outside the bacteria (Lower et al., 2008). More importantly, it can cleave E-cadherin to disrupt epithelial tight junction, which may benefit the bacterial invasion of the gastric epithelial tissues (Hoy et al., 2010). Since HtrA and other conserved proteases are known to play important roles in bacterial pathogenesis (Ingmer and Brondsted, 2009), identifying cellular targets of the Chlamydia-secreted cHtrA should provide novel insights into chlamydial pathogenic mechanisms.
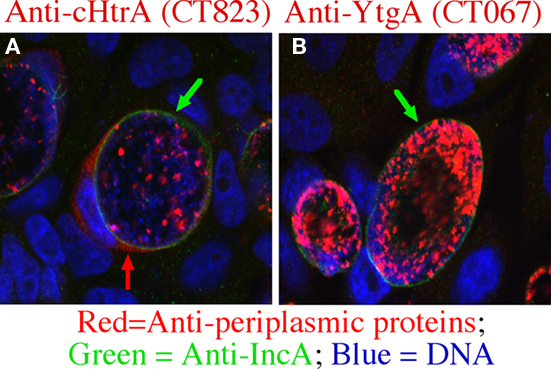
Figure 1. Chlamydial high temperature requirement protein A (cHtrA) is selectively secreted into host cell cytosol. The Chlamydia trachomatis-infected HeLa cells grown on coverslips were processed at 36 h after infection for an immunofluorescence assay as described previously (Zhong et al., 1997). The samples were triply labeled with mouse anti-cHtrA [CT823 (A)] or YtgA [CT067 (B)] antibodies visualized with a goat anti-mouse IgG conjugated with Cy3 (red), a rabbit anti-IncA antibody visualized with a goat anti-rabbit IgG conjugated with Cy2 (green), and the DNA dye Hoechst (blue). The images were acquired using an Olympus confocal microscope. Green arrows indicate inclusion membrane while the red arrow indicates cHtrA secreted into host cell cytosol. Note that only cHtrA but not YtgA was detected outside of the inclusion membrane although both are considered chlamydial periplasmic proteins (Miller et al., 2009).
Potential Pathways Required for c. Trachomatis Secretion of Proteases into Host Cell Cytosol
All three proteases localized in the host cell cytosol contain an N-terminal signal sequence, suggesting that they may be translocated into the periplasmic region via a sec-dependent secretion pathway. Among the many known secretion pathways, the sec-dependent or twin-arginine translocon (Tat) pathway is used by bacteria to deliver proteins into the periplasmic space. The Tat translocase, consisting of the TatA/E, B and C proteins, is responsible for transporting folded proteins across the inner membrane (Dilks et al., 2003; Lee et al., 2006). However, C. trachomatis genome does not encode any homolog of the Tat translocases (Stephens et al., 1998) but encodes all essential components required for a functional sec-dependent pathway (Stephens et al., 1998), suggesting that C. trachomatis organisms can use the universally conserved sec-dependent pathway to translocate proteins with an N-terminal signal sequence from cytoplasm into periplasmic regions. Indeed, we have previously demonstrated that a sec-dependent pathway is required for exporting CPAF to host cell cytosol (Chen et al., 2010b). The N-terminal signal sequence of CPAF (CPAFss) was cleaved from mature CPAF and CPAFss directed translocation of PhoA into bacterial periplasm.
However, the sec-dependent pathway alone can only deliver its cargoes into the periplasmic region. The periplasmic proteins are further exported outside of the bacterial organisms using the chaperone/usher, autotransporter, or Type II outer membrane GspD pore complex pathways. Although chlamydial genome encodes homologs of both autotransporters and GspD, these pathways deliver periplasmic proteins out of the organisms in free form. It is hard to imagine how free proteins secreted into the inclusion lumen can pass through the inclusion membrane and enter host cell cytosol in a regulated/controlled manner. Interestingly, both CPAF (Figure 2) and cHtrA (data not shown) were detected in organism-free granules in the lumen of inclusions before their secretion into host cell cytosol. It is thus hypothesized that the chlamydial proteases that are translocated into the periplasmic regions may be further exported to the lumen of inclusions via an outer membrane vesicles (OMVs) budding mechanism. This hypothesis is supported by various previous observations that the chlamydial RB outer membrane was induced to undergo vesiculation (Matsumoto and Manire, 1970) and chlamydial organism-free vesicles were detected both inside (Jorgensen and Valdivia, 2008) and outside of inclusion membrane (Giles et al., 2006). The vesiculized proteases may further enter host cell cytosol by vesicle fusing with or passing through the inclusion membrane. Although OMVs have been recognized as an essential means for gram-negative bacteria to secrete virulence factors (Bomberger et al., 2009; Ellis and Kuehn, 2010; Parker et al., 2010; Unal et al., 2010), the precise mechanisms on how OMVs are regulated remain unknown (Haurat et al., 2011). That’s why some remain skeptical about whether OMVs can represent a specific mechanism for protein secretion. The C. trachomatis-infected cells by allowing multiple vesiculized “effectors” to be studied simultaneously in a confined space may provide a unique opportunity for us to gain novel insights into the mechanisms of OMVs.
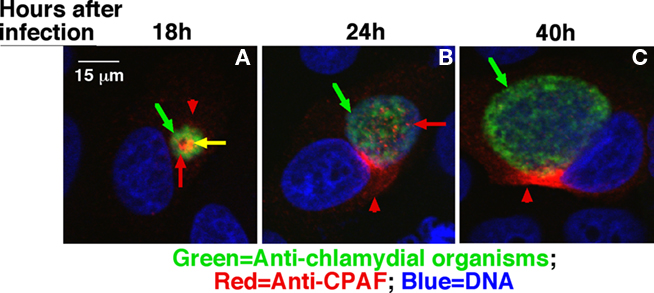
Figure 2. Detection of CPAF in chlamydial organism-free vesicles in the lumen of chlamydial inclusions. The Chlamydia trachomatis-infected HeLa cells grown on coverslips were processed at 18 h (A), 24 h (B), or 40 h (C) after infection as indicated on top of the figure for an immunofluorescence assay as described in Figure 1 legend except that the monoclonal antibody 100a against CPAF was used to replace the mouse antibodies and a rabbit anti-chlamydial organism antibody was used to replace the anti-IncA antibody. The images were also acquired using an Olympus confocal microscope. Green arrows indicate chlamydial organisms. Red arrows indicate CPAF-laden granules that are free of chlamydial organisms while the yellow arrow indicates CPAF that overlaps with chlamydial organisms. Red arrowheads indicate CPAF molecules secreted into host cell cytosol. Please note that at the early time points, CPAF was detected in granules in the inclusions and many of the CPAF-positive granules were free of chlamydial organisms in the 24 h sample.
Conflict of Interest Statement
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgment
This work was supported in part by grants from the US National Institutes of Health.
References
Balsara, Z. R., Misaghi, S., Lafave, J. N., and Starnbach, M. N. (2006). Chlamydia trachomatis infection induces cleavage of the mitotic cyclin B1. Infect. Immun. 74, 5602–5608.
Betts-Hampikian, H., and Fields, K. (2010). The chlamydial type III secretion mechanism: revealing cracks in a tough nut. Front. Microbiol. 1:114. doi: 10.3389/fmicb.2010.00114
Bomberger, J. M., Maceachran, D. P., Coutermarsh, B. A., Ye, S., O’Toole, G. A., and Stanton, B. A. (2009). Long-distance delivery of bacterial virulence factors by Pseudomonas aeruginosa outer membrane vesicles. PLoS Pathog. 5, e1000382. doi: 10.1371/journal.ppat.1000382
Borth, N., Massier, J., Franke, C., Sachse, K., Saluz, H. P., and Hanel, F. (2010). Chlamydial protease CT441 interacts with SRAP1 co-activator of estrogen receptor alpha and partially alleviates its co-activation activity. J. Steroid Biochem. Mol. Biol. 119, 89–95.
Brown, M. S., and Goldstein, J. L. (1999). A proteolytic pathway that controls the cholesterol content of membranes, cells, and blood. Proc. Natl. Acad. Sci. U.S.A. 96, 11041–11048.
Byrne, G. I. (2010). Chlamydia trachomatis strains and virulence: rethinking links to infection prevalence and disease severity. J. Infect. Dis. 201(Suppl. 2), S126–S133.
Carlson, J. H., Porcella, S. F., McClarty, G., and Caldwell, H. D. (2005). Comparative genomic analysis of Chlamydia trachomatis oculotropic and genitotropic strains. Infect. Immun. 73, 6407–6418.
Centers for Disease Control and Prevention. (2009). “Sexually transmitted disease surveillance, 2008,” in U. S. Department of Health and Human Services, Atlanta, GA. Available at: http://www.cdc.gov/std/stats08/toc.htm
Chen, D., Chai, J., Hart, P. J., and Zhong, G. (2009). Identifying catalytic residues in CPAF, a Chlamydia-secreted protease. Arch. Biochem. Biophys. 485, 16–23.
Chen, D., Lei, L., Flores, R., Huang, Z., Wu, Z., Chai, J., and Zhong, G. (2010a). Autoprocessing and self-activation of the secreted protease CPAF in Chlamydia-infected cells. Microb. Pathog. 49, 164–173.
Chen, D., Lei, L., Lu, C., Flores, R., DeLisa, M. P., Roberts, T. C., Romesberg, F. E., and Zhong, G. (2010b). Secretion of the chlamydial virulence factor CPAF requires the Sec-dependent pathway. Microbiology 156, 3031–3040.
Cheng, W., Shivshankar, P., Zhong, Y., Chen, D., Li, Z., and Zhong, G. (2008). Intracellular interleukin-1alpha mediates interleukin-8 production induced by Chlamydia trachomatis infection via a mechanism independent of type I interleukin-1 receptor. Infect. Immun. 76, 942–951.
Christian, J., Vier, J., Paschen, S. A., and Hacker, G. (2010). Cleavage of the NF-{kappa}B family protein p65/RelA by the chlamydial protease-like activity factor (CPAF) impairs proinflammatory signaling in cells infected with chlamydiae. J. Biol. Chem. 285, 41320–41327.
Clifton, D. R., Dooley, C. A., Grieshaber, S. S., Carabeo, R. A., Fields, K. A., and Hackstadt, T. (2005). Tyrosine phosphorylation of the chlamydial effector protein Tarp is species specific and not required for recruitment of actin. Infect. Immun. 73, 3860–3868.
Clifton, D. R., Fields, K. A., Grieshaber, S. S., Dooley, C. A., Fischer, E. R., Mead, D. J., Carabeo, R. A., and Hackstadt, T. (2004). A chlamydial type III translocated protein is tyrosine-phosphorylated at the site of entry and associated with recruitment of actin. Proc. Natl. Acad. Sci. U.S.A. 101, 10166–10171.
Dilks, K., Rose, R. W., Hartmann, E., and Pohlschroder, M. (2003). Prokaryotic utilization of the twin-arginine translocation pathway: a genomic survey. J. Bacteriol. 185, 1478–1483.
Dong, F., Pirbhai, M., Xiao, Y., Zhong, Y., Wu, Y., and Zhong, G. (2005). Degradation of the proapoptotic proteins Bik, Puma, and Bim with Bcl-2 domain 3 homology in Chlamydia trachomatis-infected cells. Infect. Immun. 73, 1861–1864.
Dong, F., Pirbhai, M., Zhong, Y., and Zhong, G. (2004a). Cleavage-dependent activation of a Chlamydia-secreted protease. Mol. Microbiol. 52, 1487–1494.
Dong, F., Sharma, J., Xiao, Y., Zhong, Y., and Zhong, G. (2004b). Intramolecular dimerization is required for the Chlamydia-secreted protease CPAF to degrade host transcriptional factors. Infect. Immun. 72, 3869–3875.
Dong, F., Su, H., Huang, Y., Zhong, Y., and Zhong, G. (2004c). Cleavage of host keratin 8 by a Chlamydia-secreted protease. Infect. Immun. 72, 3863–3868.
Ellis, T. N., and Kuehn, M. J. (2010). Virulence and immunomodulatory roles of bacterial outer membrane vesicles. Microbiol. Mol. Biol. Rev. 74, 81–94.
Engel, J. (2004). Tarp and Arp: how Chlamydia induces its own entry. Proc. Natl. Acad. Sci. U.S.A. 101, 9947–9948.
Fan, T., Lu, H., Hu, H., Shi, L., McClarty, G. A., Nance, D. M., Greenberg, A. H., and Zhong, G. (1998). Inhibition of apoptosis in Chlamydia-infected cells: blockade of mitochondrial cytochrome c release and caspase activation. J. Exp. Med. 187, 487–496.
Fields, K. A., and Hackstadt, T. (2000). Evidence for the secretion of Chlamydia trachomatis CopN by a type III secretion mechanism. Mol. Microbiol. 38, 1048–1060.
Fischer, S. F., Vier, J., Kirschnek, S., Klos, A., Hess, S., Ying, S., and Hacker, G. (2004). Chlamydia inhibit host cell apoptosis by degradation of proapoptotic BH3-only proteins. J. Exp. Med. 200, 905–916.
Giles, D. K., Whittimore, J. D., LaRue, R. W., Raulston, J. E., and Wyrick, P. B. (2006). Ultrastructural analysis of chlamydial antigen-containing vesicles everting from the Chlamydia trachomatis inclusion. Microbes Infect. 8, 1579–1591.
Greene, W., Xiao, Y., Huang, Y., McClarty, G., and Zhong, G. (2004). Chlamydia-infected cells continue to undergo mitosis and resist induction of apoptosis. Infect. Immun. 72, 451–460.
Haurat, M. F., Aduse-Opoku, J., Rangarajan, M., Dorobantu, L., Gray, M. R., Curtis, M. A., and Feldman, M. F. (2011). Selective sorting of cargo proteins into bacterial membrane vesicles. J. Biol. Chem. 286, 1269–1276.
Henderson, I. R., and Lam, A. C. (2001). Polymorphic proteins of Chlamydia spp.-autotransporters beyond the Proteobacteria. Trends Microbiol. 9, 573–578.
Hower, S., Wolf, K., and Fields, K. A. (2009). Evidence that CT694 is a novel Chlamydia trachomatis T3S substrate capable of functioning during invasion or early cycle development. Mol. Microbiol. 72, 1423–1437.
Hoy, B., Lower, M., Weydig, C., Carra, G., Tegtmeyer, N., Geppert, T., Schroder, P., Sewald, N., Backert, S., Schneider, G., and Wessler, S. (2010). Helicobacter pylori HtrA is a new secreted virulence factor that cleaves E-cadherin to disrupt intercellular adhesion. EMBO Rep. 11, 798–804.
Huang, Z., Feng, Y., Chen, D., Wu, X., Huang, S., Wang, X., Xiao, X., Li, W., Huang, N., Gu, L., Zhong, G., and Chai, J. (2008). Structural basis for activation and inhibition of the secreted Chlamydia protease CPAF. Cell Host Microbe 4, 529–542.
Huston, W. M., Swedberg, J. E., Harris, J. M., Walsh, T. P., Mathews, S. A., and Timms, P. (2007). The temperature activated HtrA protease from pathogen Chlamydia trachomatis acts as both a chaperone and protease at 37 degrees C. FEBS Lett. 581, 3382–3386.
Huston, W. M., Theodoropoulos, C., Mathews, S. A., and Timms, P. (2008). Chlamydia trachomatis responds to heat shock, penicillin induced persistence, and IFN-gamma persistence by altering levels of the extracytoplasmic stress response protease HtrA. BMC Microbiol. 8, 190. doi: 10.1186/1471-2180-8-190
Ingmer, H., and Brondsted, L. (2009). Proteases in bacterial pathogenesis. Res. Microbiol. 160, 704–710.
Jorgensen, I., and Valdivia, R. H. (2008). Pmp-like proteins Pls1 and Pls2 are secreted into the lumen of the Chlamydia trachomatis inclusion. Infect. Immun. 76, 3940–3950.
Kawana, K., Quayle, A. J., Ficarra, M., Ibana, J. A., Shen, L., Kawana, Y., Yang, H., Marrero, L., Yavagal, S., Greene, S. J., Zhang, Y. X., Pyles, R. B., Blumberg, R. S., and Schust, D. J. (2007). CD1d degradation in Chlamydia trachomatis-infected epithelial cells is the result of both cellular and chlamydial proteasomal activity. J. Biol. Chem. 282, 7368–7375.
Kiselev, A. O., Skinner, M. C., and Lampe, M. F. (2009). Analysis of pmpD expression and PmpD post-translational processing during the life cycle of Chlamydia trachomatis serovars A, D, and L2. PLoS ONE 4, e5191. doi:10.1371/journal.pone.0005191
Krojer, T., Garrido-Franco, M., Huber, R., Ehrmann, M., and Clausen, T. (2002). Crystal structure of DegP (HtrA) reveals a new protease-chaperone machine. Nature 416, 455–459.
Krojer, T., Sawa, J., Huber, R., and Clausen, T. (2010). HtrA proteases have a conserved activation mechanism that can be triggered by distinct molecular cues. Nat. Struct. Mol. Biol. 17, 844–852.
Kumar, Y., and Valdivia, R. H. (2008). Actin and intermediate filaments stabilize the Chlamydia trachomatis vacuole by forming dynamic structural scaffolds. Cell Host Microbe 4, 159–169.
Lad, S. P., Li, J., da Silva Correia, J., Pan, Q., Gadwal, S., Ulevitch, R. J., and Li, E. (2007a). Cleavage of p65/RelA of the NF-kappaB pathway by Chlamydia. Proc. Natl. Acad. Sci. U.S.A. 104, 2933–2938.
Lad, S. P., Yang, G., Scott, D. A., Wang, G., Nair, P., Mathison, J., Reddy, V. S., and Li, E. (2007b). Chlamydial CT441 is a PDZ domain-containing tail-specific protease that interferes with the NF-kappaB pathway of immune response. J. Bacteriol. 189, 6619–6625.
Land, J. A., Van Bergen, J. E., Morre, S. A., and Postma, M. J. (2010). Epidemiology of Chlamydia trachomatis infection in women and the cost-effectiveness of screening. Hum. Reprod. Update 16, 189–204.
Lee, P. A., Tullman-Ercek, D., and Georgiou, G. (2006). The bacterial twin-arginine translocation pathway. Annu. Rev. Microbiol. 60, 373–395.
Li, Z., Chen, C., Chen, D., Wu, Y., Zhong, Y., and Zhong, G. (2008). Characterization of fifty putative inclusion membrane proteins encoded in the Chlamydia trachomatis genome. Infect. Immun. 76, 2746–2757.
Lower, M., Weydig, C., Metzler, D., Reuter, A., Starzinski-Powitz, A., Wessler, S., and Schneider, G. (2008). Prediction of extracellular proteases of the human pathogen Helicobacter pylori reveals proteolytic activity of the Hp1018/19 protein HtrA. PLoS ONE 3, e3510. doi: 10.1371/journal.pone.0003510
Luo, J., Jia, T., Flores, R., Chen, D., and Zhong, G. (2007a). Hypothetical protein Cpn0308 is localized in the Chlamydia pneumoniae inclusion membrane. Infect. Immun. 75, 497–503.
Luo, J., Jia, T., Zhong, Y., Chen, D., Flores, R., and Zhong, G. (2007b). Localization of the hypothetical protein Cpn0585 in the inclusion membrane of Chlamydia pneumoniae-infected cells. Microb. Pathog. 42, 111–116.
Luo, J., Liu, G., Zhong, Y., Jia, T., Liu, K., Chen, D., and Zhong, G. (2007c). Characterization of hypothetical proteins Cpn0146, 0147, 0284 and 0285 that are predicted to be in the Chlamydia pneumoniae inclusion membrane. BMC Microbiol. 7, 38. doi: 10.1186/1471-2180-7-38
Matsumoto, A., and Manire, G. P. (1970). Electron microscopic observations on the effects of penicillin on the morphology of Chlamydia psittaci. J. Bacteriol. 101, 278–285.
Miller, J. D., Sal, M. S., Schell, M., Whittimore, J. D., and Raulston, J. E. (2009). Chlamydia trachomatis YtgA is an iron-binding periplasmic protein induced by iron restriction. Microbiology 155, 2884–2894.
Missiakas, D., Mayer, M. P., Lemaire, M., Georgopoulos, C., and Raina, S. (1997). Modulation of the Escherichia coli sigmaE (RpoE) heat-shock transcription-factor activity by the RseA, RseB and RseC proteins. Mol. Microbiol. 24, 355–371.
Parker, H., Chitcholtan, K., Hampton, M. B., and Keenan, J. I. (2010). Uptake of Helicobacter pylori outer membrane vesicles by gastric epithelial cells. Infect. Immun. 78, 5054–5061.
Paschen, S. A., Christian, J. G., Vier, J., Schmidt, F., Walch, A., Ojcius, D. M., and Hacker, G. (2008). Cytopathicity of Chlamydia is largely reproduced by expression of a single chlamydial protease. J. Cell Biol. 182, 117–127.
Pirbhai, M., Dong, F., Zhong, Y., Pan, K. Z., and Zhong, G. (2006). The secreted protease factor CPAF is responsible for degrading pro-apoptotic BH3-only proteins in Chlamydia trachomatis-infected cells. J. Biol. Chem. 281, 31495–31501.
Rasmussen, S. J., Eckmann, L., Quayle, A. J., Shen, L., Zhang, Y. X., Anderson, D. J., Fierer, J., Stephens, R. S., and Kagnoff, M. F. (1997). Secretion of proinflammatory cytokines by epithelial cells in response to Chlamydia infection suggests a central role for epithelial cells in chlamydial pathogenesis. J. Clin. Invest. 99, 77–87.
Rockey, D. D., Heinzen, R. A., and Hackstadt, T. (1995). Cloning and characterization of a Chlamydia psittaci gene coding for a protein localized in the inclusion membrane of infected cells. Mol. Microbiol. 15, 617–626.
Rockey, D. D., Scidmore, M. A., Bannantine, J. P., and Brown, W. J. (2002). Proteins in the chlamydial inclusion membrane. Microbes Infect. 4, 333–340.
Rupp, J., Gieffers, J., Klinger, M., van Zandbergen, G., Wrase, R., Maass, M., Solbach, W., Deiwick, J., and Hellwig-Burgel, T. (2007). Chlamydia pneumoniae directly interferes with HIF-1alpha stabilization in human host cells. Cell. Microbiol. 9, 2181–2191.
Savijoki, K., Alvesalo, J., Vuorela, P., Leinonen, M., and Kalkkinen, N. (2008). Proteomic analysis of Chlamydia pneumoniae-infected HL cells reveals extensive degradation of cytoskeletal proteins. FEMS Immunol. Med. Microbiol. 54, 375–384.
Stephens, R. S. (2003). The cellular paradigm of chlamydial pathogenesis. Trends Microbiol. 11, 44–51.
Stephens, R. S., Kalman, S., Lammel, C., Fan, J., Marathe, R., Aravind, L., Mitchell, W., Olinger, L., Tatusov, R. L., Zhao, Q., Koonin, E. V., and Davis, R. W. (1998). Genome sequence of an obligate intracellular pathogen of humans: Chlamydia trachomatis. Science 282, 754–759.
Su, H., McClarty, G., Dong, F., Hatch, G. M., Pan, Z. K., and Zhong, G. (2004). Activation of Raf/MEK/ERK/cPLA2 signaling pathway is essential for chlamydial acquisition of host glycerophospholipids. J. Biol. Chem. 279, 9409–9416.
Sun, J., and Schoborg, R. V. (2009). The host adherens junction molecule nectin-1 is degraded by chlamydial protease-like activity factor (CPAF) in Chlamydia trachomatis-infected genital epithelial cells. Microbes Infect. 11, 12–19.
Unal, C. M., Schaar, V., and Riesbeck, K. (2010). Bacterial outer membrane vesicles in disease and preventive medicine. Semin. Immunopathol. [Epub ahead of print].
Valdivia, R. H. (2008). Chlamydia effector proteins and new insights into chlamydial cellular microbiology. Curr. Opin. Microbiol. 11, 53–59.
Vandahl, B. B., Stensballe, A., Roepstorff, P., Christiansen, G., and Birkelund, S. (2005). Secretion of Cpn0796 from Chlamydia pneumoniae into the host cell cytoplasm by an autotransporter mechanism. Cell. Microbiol. 7, 825–836.
Walsh, N. P., Alba, B. M., Bose, B., Gross, C. A., and Sauer, R. T. (2003). OMP peptide signals initiate the envelope-stress response by activating DegS protease via relief of inhibition mediated by its PDZ domain. Cell 113, 61–71.
Wehrl, W., Brinkmann, V., Jungblut, P. R., Meyer, T. F., and Szczepek, A. J. (2004). From the inside out-processing of the chlamydial autotransporter PmpD and its role in bacterial adhesion and activation of human host cells. Mol. Microbiol. 51, 319–334.
Wyrick, P. B. (2010). Chlamydia trachomatis persistence in vitro: an overview. J. Infect. Dis. 201(Suppl. 2), S88–S95.
Xiao, Y., Zhong, Y., Greene, W., Dong, F., and Zhong, G. (2004). Chlamydia trachomatis infection inhibits both Bax and Bak activation induced by staurosporine. Infect. Immun. 72, 5470–5474.
Xiao, Y., Zhong, Y., Su, H., Zhou, Z., Chiao, P., and Zhong, G. (2005). NF-kappa B activation is not required for Chlamydia trachomatis inhibition of host epithelial cell apoptosis. J. Immunol. 174, 1701–1708.
Ye, J., Rawson, R. B., Komuro, R., Chen, X., Dave, U. P., Prywes, R., Brown, M. S., and Goldstein, J. L. (2000). ER stress induces cleavage of membrane-bound ATF6 by the same proteases that process SREBPs. Mol. Cell 6, 1355–1364.
Zhong, G. (2009). Killing me softly: chlamydial use of proteolysis for evading host defenses. Trends Microbiol. 17, 467–474.
Zhong, G., Fan, P., Ji, H., Dong, F., and Huang, Y. (2001). Identification of a chlamydial protease-like activity factor responsible for the degradation of host transcription factors. J. Exp. Med. 193, 935–942.
Zhong, G., Fan, T., and Liu, L. (1999). Chlamydia inhibits interferon gamma-inducible major histocompatibility complex class II expression by degradation of upstream stimulatory factor 1. J. Exp. Med. 189, 1931–1938.
Zhong, G., Lei, L., Gong, S., Lu, C., Qi, M., and Chen, D. (2011). Chlamydia-secreted proteins in chlamydial interactions with host cells. Curr. Chem. Biol. 5, 29–37.
Zhong, G., Liu, L., Fan, T., Fan, P., and Ji, H. (2000). Degradation of transcription factor RFX5 during the inhibition of both constitutive and interferon gamma-inducible major histocompatibility complex class I expression in Chlamydia-infected cells. J. Exp. Med. 191, 1525–1534.
Zhong, G., Reis e Sousa, C., and Germain, R. N. (1997). Production, specificity, and functionality of monoclonal antibodies to specific peptide-major histocompatibility complex class II complexes formed by processing of exogenous protein. Proc. Natl. Acad. Sci. U.S.A. 94, 13856–13861.
Keywords: Chlamydia trachomatis, proteases, secretion, pathogenesis
Citation: Zhong G (2011) Chlamydia trachomatis secretion of proteases for manipulating host signaling pathways. Front. Microbio. 2:14. doi: 10.3389/fmicb.2011.00014
Received: 08 January 2011;
Paper pending published: 17 January 2011;
Accepted: 19 January 2011;
Published online: 08 February 2011.
Edited by:
Robert Heinzen, NIH/NIAID-RML, USACopyright: © 2011 Zhong. This is an open-access article subject to an exclusive license agreement between the authors and Frontiers Media SA, which permits unrestricted use, distribution, and reproduction in any medium, provided the original authors and source are credited.
*Correspondence: Guangming Zhong, Department of Microbiology and Immunology, University of Texas Health Science Center at San Antonio, 7703 Floyd Curl Drive, San Antonio, TX 78229, USA. e-mail:emhvbmdnQHV0aHNjc2EuZWR1
