- 1Department of Biological Sciences, University of Southern California, Los Angeles, CA, USA
- 2Program in Ecology and Evolutionary Biology, Department of Biology, University of Oklahoma, Norman, OK, USA
The mixotrophic prymnesiophyte, Prymnesium parvum, is a widely distributed alga with significant ecological importance. It produces toxins and can form ecosystem disruptive blooms that result in fish kills and changes in planktonic food web structure. However, the relationship between P. parvum and its prey on the molecular level is poorly understood. In this study, we used RNA-Seq technology to study changes in gene transcription of P. parvum in three treatments with different microbial populations available as potential prey: axenic P. parvum (no prey), bacterized P. paruvm, and axenic P. parvum with ciliates added as prey. Thousands of genes were differentially expressed among the three treatments. Most notably, transcriptome data indicated that P. parvum obtained organic carbon, including fatty acids, from both bacteria and ciliate prey for energy and cellular building blocks. The data also suggested that different prey provided P. parvum with macro- and micro-nutrients, namely organic nitrogen in the form of amino acids from ciliates, and iron from bacteria. However, both transcriptomic data and growth experiments indicated that P. parvum did not grow faster in the presence of prey despite the gains in nutrients, although algal abundances attained in culture were slightly greater in the presence of prey. The relationship between phototrophy, heterotrophy and growth of P. parvum is discussed.
Introduction
Prymnesium parvum is a globally distributed prymnesiophyte alga of considerable ecological importance. P. parvum occurs in ecosystems across a wide range of salinities and environmental conditions (Guo et al., 1996; Barkoh and Fries, 2010; Johnsen et al., 2010), possesses a complex life history including haploid and diploid stages (Larsen, 1999), and exhibits mixtrophic abilities (Tillmann, 2003). The combined photosynthetic and heterotrophic capabilities of P. parvum enable the alga to form Ecosystem Disruptive Algal Blooms (EDABS) (Sunda et al., 2006). The disruptive nature of the alga to the ecosystems in which it blooms has implicated P. parvum as a cause of significant economic loss to the aquaculture industry worldwide (Barkoh and Fries, 2010).
Commonly referred to as the “golden alga,” P. parvum has appeared and attained bloom abundances in several waterways of the southwestern United States during the last two decades (Roelke et al., 2011; Hambright et al., 2015). Genetic analyses of P. parvum populations have indicated a relatively recent appearance of this species in the region, perhaps a consequence of a recent introduction of a strain from Europe, and subsequent range expansion (Lutz-Carrillo et al., 2010; Hambright et al., 2015). The alga has resulted in ecosystem disturbances ranging from changes in plankton food web structure to massive fish kills (Southard et al., 2010; Jones et al., 2013). Various reports have noted that substances produced by P. parvum can immobilize a range of co-occurring species and thereby aid in the capture of prey that are often considerably larger than the alga (Henrikson et al., 2010; Remmel and Hambright, 2012). These substances may also act as predator deterrents for potential consumers or allelopathic agents retarding the growth of competing phytoplankton (Schmidt and Hansen, 2001; Fistarol et al., 2003; Granéli and Salomon, 2010; Remmel et al., 2011).
P. parvum exhibits robust phototrophic growth in culture, a characteristic that is not always common for mixotrophic microalgae (Jones, 1997). Aside from a requirement for the vitamins thiamine (B1) and cobalamin (B12) (McLaughlin, 1958; Rahat and Reich, 1963), the alga is capable of growth on inorganic medium. Maximal growth rates of 0.6–0.8 d−1 have been reported for Prymnesium species in axenic culture (Granéli et al., 2012; Hambright et al., 2014). Conversely, growth of P. parvum in the dark is not supported by the presence of prey, and has only been achieved when the growth medium has been supplemented with high concentrations of specific organic compounds (most notably glycerol) (Rahat and Jahn, 1965). Moreover, growth rates in the dark in the presence of high concentrations of glycerol that have been reported for this species are considerably slower than growth in the light (Rahat and Jahn, 1965).
Despite a very limited ability to grow exclusively heterotrophically, P. parvum grown in the light displays clear predatory behaviors, and has been shown to be predatory on a number of types of microbes including bacteria, protists and a variety of metazoan zooplankton (Tillmann, 2003; Sopanen et al., 2006; Brooks et al., 2010). It has been suggested that up to 78% of the cellular nitrogen and 45% of the cellular phosphorus may be acquired via predation (Carvalho and Granéli, 2010). The heterotrophic capabilities of the alga include the production of a suite of toxins that exhibit hemolytic, ichthyotoxic, cytolytic, and potentially neurotoxic activities (Edvardsen and Paasche, 1998; Barkoh and Fries, 2010; Henrikson et al., 2010; Bertin et al., 2012; Granéli et al., 2012), and toxin characterization is an area of active research (Schug et al., 2010; Weissbach and Legrand, 2012; Manning and La Claire, 2013).
An alternative approach to understanding the physiological response of microbial eukaryotes to environmental factors has recently been enabled through genomic and transcriptomic analysis (Bik and Thomas, 2012; Murray et al., 2012). The latter approach has proven particularly fruitful for microbial eukaryotes in recent years (Keeling et al., 2014) because the larger genomes of many of these species relative to bacteria and archaea incur much greater sequencing costs and difficulties with genome assembly. The genome of P. parvum has not yet been sequenced, but a few studies examining gene expression in the species using a variety of methods have been conducted. La Claire (2006) established the first EST database for P. parvum and examined the most frequently detected genes. Beszteri et al. (2012) used the same approach to study P. parvum gene expression under N or P limitations. Freitag et al. (2011) correlated the expression of certain polyketide synthase genes (genes assumed to play a role in toxin production) by qPCR with toxicity of the alga when cultures were exposed to physiological shock. Koid et al. (2014) compared the gene contents of P. parvum and other three prymnesiophytes on a whole transcriptome level. These studies have provided baseline information on gene expression by the prymnesiophyte.
We conducted a transcriptomic analysis of P. parvum using RNA-Seq designed to specifically examine the effect of the presence or absence of prey on gene expression. Gene expression of the alga was analyzed when grown in laboratory culture for 2–3 weeks in three situations; (1) axenically, (2) in the presence of a mixed bacterial assemblage (potential prey and/or symbiotic interactions), and (3) grown in the presence of a ciliated protozoan that was readily consumed by the prymnesiophyte. Gene expression of P. parvum was significantly affected by the presence of other microbial taxa. Transcription data suggested that P. parvum obtained organic carbon from both the bacteria and ciliate populations. The data also suggested that the two different prey types provided different nutrients to P. parvum, namely organic nitrogen from the ciliate and iron from the bacteria. However, despite providing nutrients, the presence of prey had little positive impact on the growth rate of P. parvum, although nutrients released from prey maintained algal population growth longer than in axenic cultures.
Materials and Methods
Organisms and Cultures
Prymnesium parvum strain UOBS-LP0109 (Texoma1) was isolated from Lake Texoma, Oklahoma, USA. It was subsequently made axenic by micropipetting single cells through rinses of sterile medium, and then cultured in sterile L1 medium without silica (https://ncma.bigelow.org). The bacterivorous ciliate, Uronema marina, was isolated from Buzzards Bay, Massachusetts and maintained on its attendant bacterial flora by periodic subculturing into filtered sterile seawater containing 0.01–0.001% yeast extract.
Transcriptomics Experiment
Three treatments were performed for transcriptomic analysis: axenic P. parvum, non-axenic P. parvum with its attendant bacterial flora (hereafter called bacterized treatment), and an axenic culture of P. parvum to which an aliquot of the bacterized ciliate culture was added (hereafter called ciliate treatment). The latter treatment was prepared by inoculating an axenic P. parvum culture in mid-late exponential phase with an aliquot of the ciliate culture in early stationary phase (~50,000 ciliates· ml−1) to yield an alga:ciliate ratio of 100:1. Ciliate biomass strongly dominated the ciliate culture at the time of innoculation, although attendant bacteria were also present at ~105 · ml−1 in the combined culture. Cultures were examined using a dissecting microscope every 30–40 min following addition of the ciliates to determine when ciliates were eliminated by the alga. All ciliates were rapidly killed (<6 h) after addition of ciliates to the P. parvum culture, and the cultures were harvested as soon as no live ciliates remained.
All three treatments were grown in 1–2 L volumes in 2800 ml Pyrex glass Fernbach flasks in L1 medium minus silica at 18 ppt salinity and 18°C. Cultures were incubated at an irradiance of ~300 μE·m−2s−1, measured using a QSL-100 sensor with QSP-170 deckbox (Biospherical Instruments Inc.). Cultures were incubated on a 12:12 h light:dark cycle.
Growth Experiment
The effect of the presence or absence of prey on the growth of P. parvum was established in batch cultures of the alga. All cultures used in growth experiments were grown under the same conditions as the transcriptome experiment except that they were in ~100 mL volumes. The same three treatments of P. parvum that were used for transcriptomic analysis were examined. All treatments were conducted in duplicate. On day 16, cells from a ciliate culture were added to an axenic P. parvum culture at a ratio of 100:1 (alga:ciliate). Growth of P. parvum was monitored by performing direct microscopic cell counts on samples periodically removed from each flask over a 28-day period. Growth rates in different treatments were compared using One-Way ANOVA test.
RNA Extraction and cDNA Production
All samples were harvested during mid-exponential growth phase in the middle of the 12 h light cycle. Cultures were spun down in an Eppendorf 5810R centrifuge using the A-4-62 rotor at 3200 rcf for 15 min at 15°C. The supernatant was carefully decanted, and 1–2 ml of TRI reagent (Ambion) was added to the pellet and vortexed until the pellet fully dissolved. Homogenates were then either processed immediately using Ribopure kit (Ambion), or stored at −80°C for later processing. Eluted RNA was treated with DNase (Sigma) to remove DNA contamination. Samples were cleaned and concentrated using RNA Clean and Concentrator-25 (Zymo Research). RNA was quantified using a Qubit 2.0 Fluorometer (Invitrogen) and run on an E-gel iBase with E-gel Gel EX 1% (Invitrogen) to check for nucleic acid quality.
RNA samples were sent to the National Center for Genome Resources for processing and sequencing. RNA was quantified again at the sequencing center using Invitrogen Qubit Q32855, and RNA quality was assessed using the Agilent 2100 Bioanalyzer. cDNA libraries were made from 2 μg RNA using Illumina's TruSeq RNA Sample Preparation Kit. The average insert size of each library ranged from 250 to 350 bp. Libraries were sequenced on an Illumina HiSeq 2000 which generated 2 × 50 bp (paired-end) reads. The original sequence data are publicly available from NCBI Sequence Read Archive under accession number SRA166613 and sample IDs MMETSP0006_2, MMETSP0815, and MMETSP0008_2, for axenic, bacterized and ciliate treatment, respectively.
Sequencing and Bioinformatic Analyses
Bioinformatic analysis procedures were adapted and developed from guidelines established by the Marine Microbial Eukaryote Transcriptome Sequencing Project (https://www.marinemicroeukaryotes.org) (Keeling et al., 2014). Sequences of all treatments were first checked for quality using the FASTX toolkit (http://hannonlab.cshl.edu/fastx_toolkit/index.html) with options “-p 80 -q 20.”
An assembled transcriptome of P. parvum was then generated as a reference for downstream alignment processes. All sequences of the ciliate treatment were excluded from the assembly to ensure that the P. parvum assembled transcriptome was not contaminated with ciliate sequences that might have remained after ciliates were killed by the alga. Although the bacterized treatment and ciliate treatment also contained bacteria, the sequence data from those treatments should have been reasonably free of bacterial sequences because RNA was polyA-selected in our protocol for all treatments. Sequences from the axenic and bacterized treatments were combined and assembled de novo using a hybrid approach containing both a de Bruijn graph and an overlap-based algorithm method. Sequences were first assembled using ABySS v. 1.3.2 (Simpson et al., 2009) at four different k-mer settings of 19, 25, 31, and 37. The resulting four assemblies were merged using Trans-ABySS v. 1.4.4 (Robertson et al., 2010). Redundant contigs were removed from the assembly using CD-Hit-EST v. 4.5.7 (Li and Godzik, 2006) with option “-c 0.99.” The remaining contigs were then further assembled using the overlap-based program CAP3 (Huang and Madan, 1999) with options “-p 99 -o 50 −k 0.” Scaffolding of the resulting contigs was inferred using ABySS with default parameters (Simpson et al., 2009). The GapCloser v. 1.12 application of the SOAPdenovo (Luo et al., 2012) was employed to try to fill in gaps in the assembly. Scaffolds were broken into contigs where gaps remained unfilled. Contigs shorter than 150 bp were discarded. CD-Hit-EST was used again to remove redundant contigs.
All contigs were searched against SILVA database (Quast et al., 2013) using BLAST to identify and remove rRNA sequences. Protein coding genes with at least 150 bp were predicted from the remaining contigs using ESTscan v. 3.0.3 (Iseli et al., 1999). Genes were annotated using an evalue cutoff of 1e-5 based on a variety of database searches including HMMER3 v. 3.1b1 (http://hmmer.janelia.org) searches against Pfam (http://pfam.sanger.ac.uk) and Tigrfam (http://www.jcvi.org/cgi-bin/tigrfams/index.cgi) databases. A BLAST search against NCBI nr database was also carried out using the same e-value cutoff. Genes without hits to Pfam or Tigrfam were annotated based on their best hits in the NCBI nr database. KEGG and Gene Ontology annotations were also obtained using KEGG Automatic Annotation Server (http://www.genome.jp/tools/kaas/) and Blast2GO v. 2.5 (Conesa et al., 2005), respectively, to provide additional functional understanding of the genes. Annotations of some genes of interest were edited after manual inspection of all the information available.
Sequences from each of the three datasets were aligned back to the assembled transcriptome using BWA v. 0.6.1-r104 (Li and Durbin, 2009). Read pairs that were aligned to genes correctly (i.e., paired reads aligned to the same gene, on the opposite strand, with fragment size of ~200 bp) were counted using a custom PERL script. Statistical analyses of the read counts of each gene were carried out using the “exact test” function of edgeR v. 3.6.2 (Robinson et al., 2010) with common dispersion set at 0.1. Pairwise comparisons between all three treatments were carried out. P-values were adjusted to false discovery rate using p.adjust in R software v. 3.1.0 (Benjamini and Hochberg, 1995). Only genes with adjusted p-values smaller than 0.05 were deemed as having significantly different expression levels between treatments. Normalized expression levels in the form of FPKM (Fragments per kilobase exon per million fragments aligned) values were calculated. Expression levels in bacterized and ciliate treatments relative to those in axenic treatment were calculated and plotted in figures.
Results
Growth Experiment
A growth experiment was conducted to determine whether the presence of prey (bacteria or ciliates) had a significant effect on the growth rate or overall yield of P. parvum (Figure 1). Growth rates of the alga were not significantly different (p = 0.20, ANOVA) among the three treatments in early exponential phase (0.36 ± 0.02, 0.42 ± 0.01, 0.38 ± 0.04 d−1 during the first 9 days of the experiment for axenic, bacterized, and ciliate treatment, respectively. Axenic and ciliate treatments did not differ during that period). The addition of ciliates at late exponential phase of the alga on day 16 allowed P. parvum to continue growing (0.12 ± 0.05 d−1) while cultures without ciliates exhibited negligible growth (0.03 ± 0.01 d−1). P. parvum in the bacterized treatment reached the maximum abundance of 4.28 ± 0.03 × 106 cells mL−1, which is slightly higher than that of the axenic treatment (3.80 ± 0.17 × 106 cells mL−1). The addition of ciliates also increased the maximum abundance of P. parvum to (4.95 ± 0.25 × 106 cells mL−1).
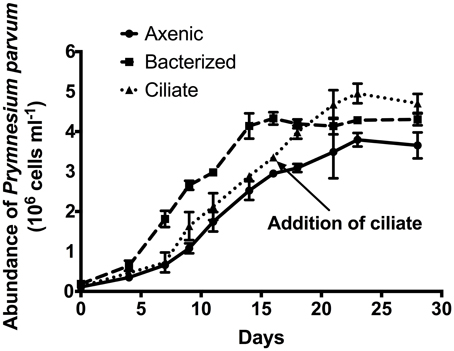
Figure 1. Growth of the prymnesiophyte, Prymnesium parvum, in axenic culture (circles, solid line), in non-axenic culture with attendant bacterial flora (squares, dashed line), and in axenic culture with the addition of ciliates at late exponential phase (triangles, dotted line). Arrow indicates Day 16 when ciliates were added to the axenic P. parvum culture. In the transcriptome experiment, cells were harvested from axenic and bacterized treatment just prior to the addition of ciliates, while cells were harvested from ciliate treatment ~6 h after the addition of ciliates. Symbols and error bars represent mean ± standard deviation of two replicates.
Overview of the Transcriptome
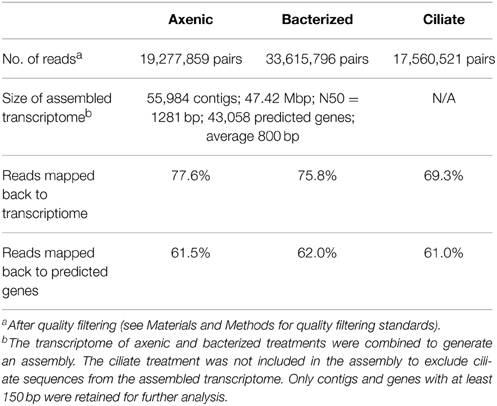
The assembled transcriptome of P. parvum contained 55,984 contigs and 43,058 genes or partial genes, totaling 47.42 million base pairs (Table 1). Most of the physiologically necessary genes were found in the transcriptome (Table S1). For example, the transcriptome contained all genes necessary for glycolysis, the TCA cycle, nitrogen metabolism, nucleotide biosynthesis and almost all genes required for the biosynthesis of all 20 amino acids. Based on this information, it is reasonable to assume that the transcriptome had enough sequencing depth to cover most of the transcribed genes. Consistent with previous experiments (McLaughlin, 1958; Rahat and Reich, 1963), the P. parvum transcriptome contained no genes for the synthesis of thiamine and cobalamin but did contain most of the genes required for the synthesis of other vitamins.
Among the 43,058 genes identified in this study, only a small percentage (9.2% or 3972 genes) had significantly different expression levels between at least a pair of treatments (Figure 2). Similar numbers of genes (1600~1900) had higher expression levels in each of the three treatments (1612, 1861, and 1645 genes in the axenic, bacteria and ciliate treatments, respectively; Figure 2). The overlap among those sets of genes, however, was quite interesting. A total of 618 genes, or 33 and 38% of all genes with elevated expression levels in the bacterized and ciliate treatments, respectively, had higher expression levels in both treatments compared to the axenic treatment. In comparison, 426 genes were shared between the axenic and bacterized treatments, and only 102 genes were shared between the axenic and ciliate treatments, in terms of genes with higher expression levels compared to the third treatment (Figure 2).
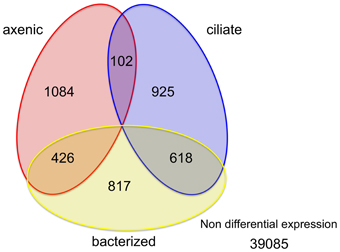
Figure 2. Distribution of genes according to changes in their expression levels among three treatments. Numbers in each area indicate numbers of genes whose expression levels were higher in that treatment compared to the other two or in two treatments compared to the third.
A more detailed evaluation of differential gene expression and function with respect to roles in P. parvum showed that most (>50%) of its genes had no matches or functional annotations to existing databases, owing to the relative scarcity of protistan sequence data (Caron et al., 2009). Nevertheless, we were able to infer interesting patterns from changes in the transcriptional patterns of those genes with putative functions (Figure 3). Genes belonging to the same pathway or metabolic function had very similar transcriptional patterns among the treatments. For example, all differential expressed genes associated with fatty acid metabolism had higher expression levels in the presence of either prey type and formed a cluster in the upper right quadrant of Figure 3, while genes associated with iron uptake clustered together in the lower left quadrant (i.e., the latter genes had lower expression levels in the presence of both prey). Figure 3 also illustrates that genes associated with different metabolic functions often demonstrated different transcriptional patterns (as noted above). While changes in transcription do not necessarily equate with protein concentrations or enzymatic activities, the transcriptional patterns observed in this study provided insights into the relationship between P. parvum and the presence of potential prey, and different prey types.
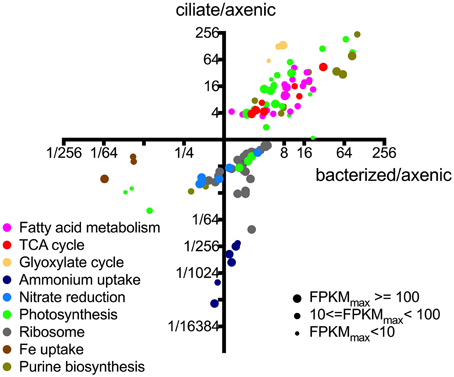
Figure 3. Relative expression levels of individual genes of P. parvum within different metabolic pathways or functions. Each dot represents an individual gene. X-axis values indicate expression levels in bacterized treatment relative to those in axenic treatment (FPKMbacterized/FPKMaxenic). Y-axis values indicate expression levels in ciliate treatment relative to those in axenic treatment (FPKMciliate/FPKMaxenic). Sizes of the dots are proportional to the number of reads assigned to the genes. Not all genes within certain pathways or functions are shown, only those that have differential expression between at least a pair of treatments. These functions and pathways with significantly differential gene expression were plotted in Figures 3–5 to further show consistent patterns within each function/pathway that are highlighted in our discussion of core metabolism of P. parvum. For list of genes and their read counts, FPKM values, and relative expression levels, refer to Table S2.
Carbon Metabolism
Genes involved in several carbon metabolism pathways had higher expression levels in both treatments with prey, compared to the axenic treatment. Several copies of genes of every step in the fatty acid oxidation pathway, including acyl-CoA dehydrogenase, enoyl-CoA hydratase, 3-hydroxyacyl-CoA dehydrogenase, and acetyl-CoA C-acetyltransferase, had ~9-fold higher expression levels when prey, either bacteria or ciliates, were present (Figures 3, 4A). Two genes encoding an electron transfer flavoprotein gene and an electron transfer flavoprotein ubiquinone oxidoreductase that also play a role in fatty acid oxidation (Watmough and Frerman, 2010) had similar transcription patterns (Table S2).
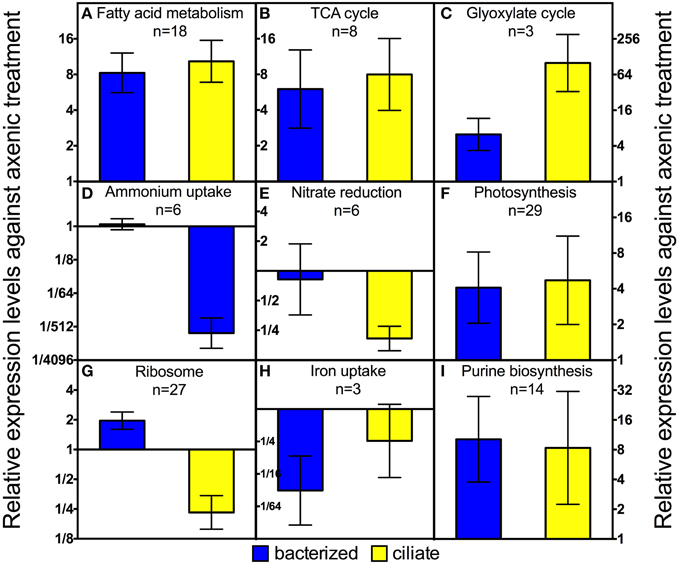
Figure 4. Average expression levels among different groups of genes of P. parvum in bacterized and ciliate treatment relative to those in axenic treatment. Geometric means ± 95% confidence intervals are shown for relative expression levels against the axenic treatment of genes of each pathway or function. For list of genes and their read counts, FPKM values, and relative expression levels, please refer to Table S2.
Fatty acid oxidation produces a large quantity of acetyl-CoA, which can be used in two different pathways in P. parvum mitochondria: complete oxidation through the TCA cycle or transformation to succinate through the glyoxylate cycle, with the former pathway being catabolic and the latter being anabolic. As expected, genes involved in both pathways had higher expression levels as a result of increased influx of acetyl-CoA when prey were available (Figures 4B,C). Genes responsible for almost every step of the TCA cycle were regulated in strikingly similar patterns as those of fatty acid oxidation (Figure 4B). Expression levels of two genes specifically involved in the glyoxylate cycle, malate synthase and isocitrate lyase (the rest of the pathway overlaps with the TCA cycle) increased ~6-fold in the bacterized treatment relative to the axenic one, and increased ~100-fold when ciliates were present (Figure 4C).
Expression patterns of glycolysis/gluconeogenesis genes were inconsistent among themselves. However, genes involved in the connection between glycolysis/gluconeogenesis and other carbon pathways had interesting expression patterns. Genes encoding pyruvate carboxylase and PEP carboxykinase had slightly higher expression levels in the bacterized treatment, and even higher levels in the ciliate treatment. These two proteins convert pyruvate and TCA cycle metabolites through oxaloacetate to PEP, and are generally regarded as gluconeogenic enzymes. Conversely, PEP carboxylase, which operates in the opposite direction of PEP carboxykinase, decreased significantly in its transcription level in the ciliate treatment (Figure 5, Table S2).
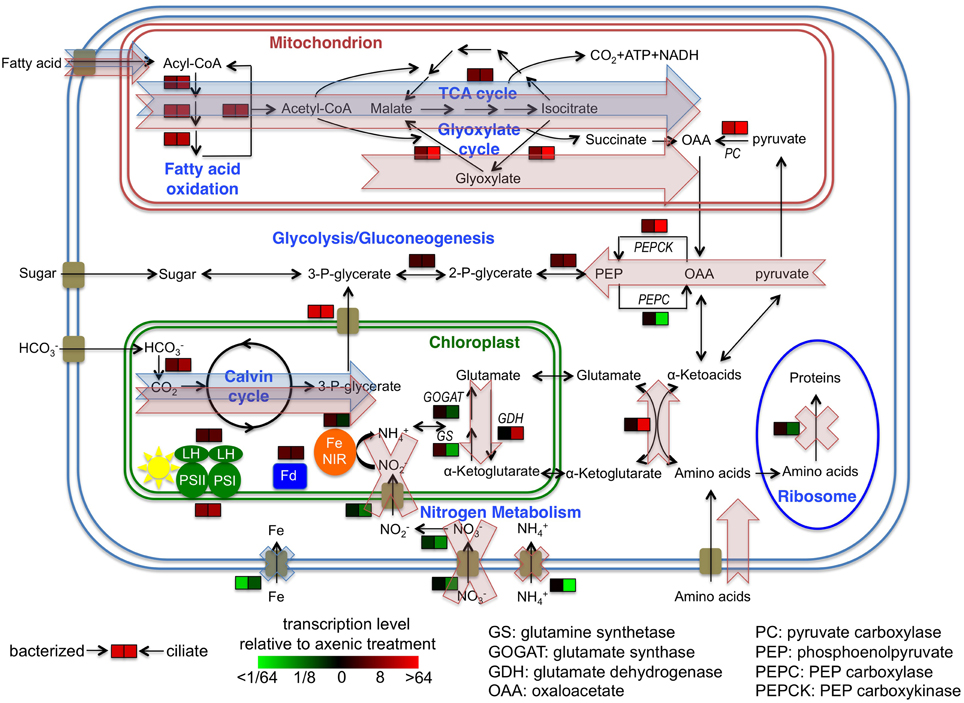
Figure 5. Cellular overview of gene transcription of Prymnesium parvum under three treatments and hypothetical model of metabolic activities. Transcription levels of select genes, pathways or functions in bacterized and ciliate treatments relative to axenic treatment are shown. Blue shaded arrows and crosses indicate proposed model of metabolic flows and suppression of functions of P. parvum, respectively, in response to the presence of bacteria in the bacterized treatment. Red shaded arrows and crosses indicate the same flows and suppressions in response to the presence of ciliates in the ciliate treatment. Cellular localizations of proteins are based on knowledge of homologs in other protists or in silico predictions.
Nitrogen Metabolism
Transcription data suggested dramatic changes in modes of nitrogen acquisition in response to the addition of prey and particularly to the presence of the ciliates. First, expression levels of ammonium uptake genes drastically decreased in the ciliate treatment, compared to the other two treatments (Figure 4D). Those genes include several copies of an ammonium transporter and two periplasmic L-amino acid oxidases. The latter two genes were similar to a homolog in the green algal species, Chlamydomonas reinhardtii, that has been shown to oxidize extracellular amino acids to release ammonium (Vallon et al., 1993). Similar although less dramatic transcriptional changes were observed for nitrate reduction genes. Nitrate transporter, nitrate reductase, ferredoxin-nitrite reductase and several nitrite transporters had ~6-fold lower expression levels in the ciliate treatment relative to the axenic or bacterized treatment (Figure 4E).
The balance and interconversion between α-ketoglutarate and glutamate is central to nitrogen metabolism, especially amino acid metabolism, as the pair provide the amino group to most if not all amino acids during their biosynthesis, and accept the amino group when they are metabolized. Expression levels of glutamine synthetase (GS) and glutamine oxoglutarate aminotransferase (GOGAT), which converts α-ketoglutarate to glutamate, decreased significantly in the ciliate treatment. On the other hand, expression levels of glutamate dehydrogenase (GDH), which catalyzes the opposite reaction, increased significantly (Figure 5, Table S2). Meanwhile, an aminotransferase gene, which catalyzes the exchange of amino groups between α-ketoglutarate/glutamate and other amino acids/α-ketoacids, had significantly higher expression levels in the ciliate treatment (Figure 5, Table S2).
Photosynthesis
A large number of genes involved in photosynthesis, such as most photosynthetic reaction center genes and ribulose-bisphosphate carboxylase genes are encoded by the chloroplast genome. Their mRNAs were not retained in the polyA selection process in our protocol, and their sequences were largely undetected in our datasets. However, we were still able to obtain transcriptional patterns of many chromosomal genes involved in photosynthesis or CO2 fixation (Figure 4F). Four genes encoding photosystem I or II proteins were detected in the transcriptome, and all of them had higher expression levels when prey were present. The transcriptome also included many copies of chlorophyll a/b binding proteins, some of which were among the most actively transcribed genes. Of these, 20 genes were differentially expressed, and 17 of those had higher expression levels in the presence of prey. Two genes encoding plant type ferredoxin, the major electron acceptor from photosystem I, increased their expression levels in the presence of prey. Three genes encoding carbonic anhydrases, which supply CO2 to the Calvin cycle, had similar expression patterns. Two other genes encoding phosphoglycerate transporters that presumably transport the end products of the Calvin cycle outside of chloroplast (Neuhaus and Wagner, 2000) had similar expression patterns (Figure 5, Table S2).
Other Notable Pathways and Functions
Twenty-seven genes encoding ribosomal proteins exhibited similar transcriptional patterns. On average, their expression levels decreased ~4-fold in the ciliate treatment compared to the axenic treatment (Figure 4G). The same genes had slightly higher expression levels (~2 fold) in the bacterized treatment compared to the axenic treatment.
Genes encoding three proteins involved in iron uptake were regulated similarly. They include the low iron inducible periplasmic protein FEA1 and two high affinity iron permeases FTR1. FEA1 has been shown to be one of the major proteins excreted by iron deficient Chlamydomonas reinhardtii (Allen et al., 2007). FEA1 may also facilitate iron uptake when expressed heterologously in yeast and plants (Narayanan et al., 2011). FTR1 is part of a high affinity iron uptake system consisting of a permease and an oxidase (Stearman et al., 1996). The transcriptional levels of these three genes were ~30-fold lower in the bacterized treatment than in the axenic treatment. Those levels also decreased, although to a lesser degree (~8-fold) in the ciliate treatment (Figure 4H).
Expression levels of 14 genes involving in purine biosynthesis increased in both bacterized and ciliate treatments compared to axenic treatment (Figure 4I). Most of the 15 genes encode proteins involved in the early, generic part of the purine biosynthesis pathway. Therefore, it was unclear whether there was a targeted increase in adenine or guanine biosynthesis. In comparison, pyrimidine biosynthesis genes did not change significantly among the three treatments.
Genes responsible for the biosynthesis of prymnesins are not yet fully characterized. However, since the two identified prymnesins are polyketides (Igarashi et al., 1996), it has been speculated that polyketide synthases play an important role in the synthesis of prymnesins (Manning and La Claire, 2010). Eleven different genes annotated as polyketide synthase were found in the transcriptome. Among them, only one had differential expression (slightly higher in the ciliate treatment), the other 10 were not differentially expressed in any of the treatments. Homologs of the genes previously studied by qPCR (Freitag et al., 2011) were among those not differentially expressed.
Three different phosphate transporters were differentially expressed. However, the pattern of their expression among the three treatments was not consistent. An ABC-type and a Pho4 family phosphate transporter both had higher expression levels (~4-fold) in the ciliate treatment. A sodium-dependent phosphate transporter had higher expression levels in the bacterized treatment (~4-fold).
Discussion
Comparison with Previous Gene Expression Studies
The size of the P. parvum transcriptome is comparable to other protistan transcriptomes in the Marine Microbial Eukaryote Transcriptome Sequencing Project (https://www.marinemicroeukaryotes.org), the transcriptomes of other prymnesiophytes (Koid et al., 2014), and protistan genomes in general (Caron et al., 2009). The transcriptome was also compared to EST data previously obtained for P. parvum. Out of 6286 EST sequences generated for P. parvum strain UTEX no. 2797 (La Claire, 2006), 5626 (89.5%) had a homolog in the transcriptome of clone UOBS-LP0109 obtained in this study, with average nucleotide sequence identity of 98.9%. Out of 17,153 EST sequences generated for P. parvum strain RL10 (Beszteri et al., 2012), 14,271 (83.2%) had a homolog in this transcriptome, with average nucleotide sequence identity of 95.8%. The numbers of shared genes were probably slight underestimates because of the limitations of the de novo short read assembly. P. parvum strain UOBS-LP0109 shares a large numbers of genes with both other strains, but was more closely related to strain UTEX no. 2797 than strain RL10 in terms of both percentage of shared genes and sequence similarity. This finding is not surprising considering the proximity of the sources of strain UOBS-LP0109 (Lake Texoma, on the border of Oklahoma and Texas) and strain UTEX no. 2797 (Texas). Many of the most frequently-encountered ESTs in the study of strain UTEX no. 2797 (La Claire, 2006), such as multiple chlorophyll-binding proteins, ABC-type phosphate transporter, and phosphoenolpyruvate carboxykinase, were also among the genes with the most read pairs.
Nutrient Uptake from Prey
Numerous studies have documented the ability of P. parvum to kill or prey on a variety organisms, ranging from bacteria to phototrophic and heterotrophic protists, zooplankton and even fish (Tillmann, 2003; Southard et al., 2010; Remmel and Hambright, 2012). It has been reported that heterotrophic nutrition by this toxic, mixotrophic alga may provide a substantial percentage of the macronutrients and carbon required for growth (Carvalho and Granéli, 2010). However, the specific contribution to algal nutrition provided by consumed prey remains unclear, and there is virtually no information on how P. parvum might respond at the cellular or molecular level to the availability of prey. We evaluated changes in gene transcription of the alga in response to two different prey assemblages: a mixed bacterial assemblage, or one consisting of largely a ciliated protist with a background assemblage of bacteria. We realize the lack of replication in our transcriptome study and therefore have been purposefully conservative in our interpretation of the data. In many cases, different genes involved in the same pathway had similar transcription patterns (Figure 3), thereby providing a level of confidence to their collective transcription patterns. Inference from the data was only drawn in those cases, and was never based on the transcription pattern of a single gene. We also recognize and acknowledge the complexity of the ciliate treatment in that many factors could have affected gene transcription of P. parvum leading to difficulties in attributing P. parvum response to a specific factor. Those factors include ciliate cells, bacterial cells, nutrients from the ciliate culture, and nutrients released as a consequence of ciliates consuming bacteria. We assume that P. parvum gene transcription changes in the ciliate treatment were likely the result of its predation on ciliates because ciliate biomass greatly outweighed bacterial biomass in that treatment, but dissolved substances released by the ciliates may also have elicited responses from the alga. Regardless, our results clearly indicated that P. parvum killed prey and acquired nutrients in their presence, and also provided insights into the response of specific metabolic processes affected by the presence of other microorganisms.
The transcriptomic data obtained in this study indicate some shared changes in gene expression to the presence of bacteria or ciliates (relative to the alga grown axenically), as well as some changes that were specific to these two types of prey assemblages. For carbon metabolism, expression levels of genes involved in fatty acid metabolism and the TCA cycle were higher in both treatments with prey (Figures 4A,B). This finding indicates that P. parvum probably used fatty acids from both prey types. Changes in expression levels of genes involved in the glyoxylate cycle, pyruvate carboxylase, PEP carboxykinase, and PEP carboxylase were much more dramatic in the ciliate treatment than in the bacterized treatment (Figure 5). This implies that organic carbon from prey was used in different ways in those two treatments. Specifically, it seemed that more fatty acids were used for biomass production and less for energy production in the ciliate treatment (Figure 5).
In nitrogen metabolism, significant changes in gene expression were only observed in the ciliate treatment. The presence of ciliates led to dramatic decreases in expression levels of genes involved in inorganic nitrogen uptake, especially ammonium uptake (Figures 3, 4D). On the other hand, the transcription patterns of GS, GOGAT, GDH and an aminotransferase gene indicate increased catabolism of amino acids in the ciliate treatment (Figure 5). Collectively, these data suggests that amino acids, presumably obtained either directly from the ciliate or mediated by the ciliate (i.e., perhaps as a consequence of ciliate predation on co-occurring bacteria), were metabolized in P. parvum. Nitrogen needs of P. parvum were likely satisfied by the influx of organic nitrogen, which led to the suppression of inorganic nitrogen uptake. In all, these data imply that ciliates might be a very effective nitrogen source for P. parvum in nature. Ciliates have been shown to be an important source of nitrogen for other mixotrophic protists in nitrogen-limited environments (Bockstahler and Coats, 1993; Smalley and Coats, 2002). This hypothesis is also consistent with previous findings that have reported greater toxicity of P. parvum under nitrogen or phosphate limitation (Johansson and Granéli, 1999; Granéli and Johansson, 2003; Hambright et al., 2014), and suppression of toxicity when high doses of ammonium are provided (Grover et al., 2007, 2013).
Expression levels of genes involved in iron uptake decreased sharply in the presence of bacteria, and to a lesser degree in the ciliate treatment (Figure 4H). Both prey assemblages were likely sources of iron for P. parvum in our experiment, although it would appear that bacteria alone were more effective in making iron accessible than the ciliate. This finding is consistent with the varied and complex relationships known to occur between phytoplankton and co-occurring bacteria (Amin et al., 2012). Maranger et al. (1998) experimentally demonstrated that the ingestion of bacteria by a different mixotrophic phytoflagellate, Ochromonas sp., provided iron for cellular growth of the alga. We speculate that the digestion of ingested prey by P. parvum may meet the iron requirement for growth of the alga and thereby reduce the need for iron uptake and transport. Alternatively, iron availability may be increased in the presence of other microbes, obviating a requirement for phagotrophy by P. parvum. For example, predation on bacteria by heterotrophic protists has been shown to relieve iron stress of co-occurring algae (Barbeau et al., 1996). Most previous studies of P. parvum have focused on the importance of nitrogen and phosphate contained in prey biomass as a factor in the toxicity of the alga, since N:P ratio has a direct impact on the toxicity of P. parvum (Granéli et al., 2012). Our results broaden these findings to include the possible importance of iron to the metabolism of P. parvum. The bacterial assemblage in our culture was part of the native bacterial assemblage occurring during a P. parvum bloom. It would be interesting to determine whether P. parvum has a preference among different bacteria as a means of acquiring iron.
Contribution of Predation to the Growth of P. parvum
The ciliate employed in the study, Uronema marina, was readily killed when introduced into cultures of P. parvum at high abundance. The process involved immobilization and attachment of several algal cells to each ciliate, as has been described previously (Tillmann, 2003). Bacterivory has also been reported for P. parvum (Nygaard and Tobiesen, 1993), although ingestion of bacteria was not confirmed in our study. It has been speculated that prey capture and digestion by P. parvum provide substantial sources of major nutrients (N, P) or organic carbon for energy and growth (Carvalho and Granéli, 2010; Granéli et al., 2012). Our transcriptional data supported this hypothesis, for nitrogen and carbon.
Many mixotrophic protists grow faster when prey are available compared to axenic growth (Sanders, 2011). Surprisingly, our transcriptional data and our direct measurements of growth of the alga indicated that P. parvum did not grow faster despite its demonstrable predatory behavior. Protein synthesis at ribosomes is a huge component of cell growth, therefore ribosome synthesis could at least in part reflect cell growth. It has been proposed that ribosome synthesis levels could be a good measure for growth (Rudra and Warner, 2004). The expression levels of genes encoding P. parvum ribosomal proteins increased only slightly in the bacterized treatment compared to levels observed for the axenic treatment (Figure 4G). However, P. parvum did not grow significantly faster in the presence of bacteria (Figure 1). On the other hand, the expression levels of these genes decreased in the ciliate treatment (Figure 4G), implying P. parvum synthesized fewer ribosomes. Although the addition of ciliates in the late exponential phase maintained growth of P. parvum, that growth rate was slower than that of an axenic culture in the early-mid exponential phase (p < 0.001, ANOVA, Figure 1). The treatments with prey did support slightly higher yields of the alga relative to the axenic P. parvum culture. This outcome is presumably a result of the additional nutrients available from the prey or the culture medium of the prey. Overall, our findings are consistent with a previous study in which P. parvum fed Rhodomonas salina did not grow faster than a monoculture of P. parvum, even under nutrient deficient conditions (Carvalho and Granéli, 2010).
One possible explanation for the lack of increased growth rate in the presence of prey is the cost of making heterotrophic cellular machinery when prey become available. However, we did not observe an initial decrease followed by later recovery in growth rate that would be expected if such hypothesis were true. Additionally, killing of the ciliate cells started almost instantaneously when they were added to the P. parvum culture, suggesting P. parvum was already toxic and predatory when growing axenically. Therefore, a more likely explanation is that P. parvum is critically dependent on some aspect of phototrophy, and that phototrophy presents a bottleneck controlling growth rate, at least under the specific growth conditions employed in this study. A photosynthetic bottleneck is implied by the observation that P. parvum cannot grow or survive in the dark even when prey are available at high abundances (Carvalho and Granéli, 2010; Brutemark and Granéli, 2011). The only known exception to this rule is that P. parvum can grow very slowly in the dark when supplied with high concentrations of certain organic compounds (Rahat and Jahn, 1965). However, the concentrations of organic compounds required for its growth in the dark are much higher than concentrations that would be experienced in nature. The mechanism and metabolic details of such growth are unknown.
Carvalho and Granéli (2010) suggested that photosynthesis and phagotrophy in P. parvum might supplement each other, employing “co-metabolism” for growth. Our results do not directly support this hypothesis, since the presence of prey in our study did not enhance growth rate per se beyond that enabled by phototrophic growth. However, one observation in our study was at least partially consistent with their hypothesis. A number of genes involved in photosynthesis, although far from complete and conclusive, increased their expression levels in response to prey despite the fact that there was no difference in light conditions during the experiment. The presence of prey may have relieved some of the cellular processes required to obtain major nutrient elements (e.g., N), as evidenced by changes in gene expression among our experimental treatments. As a consequence, more resources might have been made available for making the photosynthetic apparatus. Alternatively, prey may have provided some micronutrient(s) that support photosynthesis. For example, increased availability of iron may have had a positive effect on photosynthesis since the photosynthetic apparatus includes many proteins and cofactors that contain Fe-S clusters.
It was unclear from our transcriptomic data where or how nutrients acquired by P. parvum from its heterotrophic activities were used, since growth rate was not noticeably enhanced. One possibility is storage, possibly in carbohydrates. Some of our data supported this hypothesis as expression levels of some gluconeogenic genes increased in treatments with prey, especially in the ciliate treatment (Figure 5). However, genes involved in starch biosynthesis were not significantly regulated. Other forms of storage are also possible, although there were no transcriptional data indicating which form. If this hypothesis is true, it means that P. parvum may be acquiring nutrients from prey and storing them for growth when conditions are amenable for photosynthesis. This is consistent with the observation that P. parvum took up nitrogen and phosphorus from Myrionecta rubra but did not grow in the dark (Brutemark and Granéli, 2011).
Our study investigated expression of a wide range of genes in the mixotrophic protist P. parvum in the presence or absence of two different prey assemblages. Our gene expression data clearly indicated that P. parvum obtained a variety of nutrients from its prey including organic carbon in the form of fatty acids from both the bacteria and the ciliate, organic nitrogen in the form of amino acids from the ciliate, and iron from the bacteria. Despite the nutrient gains, P. parvum did not grow faster, implying that its growth is ultimately controlled by its photosynthetic activity, at least in the cases of tested prey types. Our study revealed transcriptional behaviors of P. parvum in response to prey availability, and has led to several testable hypotheses relating to interactions between P. parvum and its prey. These results provide the necessary foundation for future molecular studies regarding the contribution of prey for growth and maintenance of this ecologically and economically important organism.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This research is funded by the Gordon and Betty Moore Foundation through Grant GBMF3299 to DAC and KBH. This sequencing effort was funded in part by the Gordon and Betty Moore Foundation through Grant GBMF2637 to the National Center for Genome Resources.
Supplementary Material
The Supplementary Material for this article can be found online at: http://www.frontiersin.org/journal/10.3389/fmicb.2015.00319/abstract
References
Allen, M. D., del Campo, J. A., Kropat, J., and Merchant, S. S. (2007). FEA1, FEA2, and FRE1, encoding two homologous secreted proteins and a candidate ferrireductase, are expressed coordinately with FOX1 and FTR1 in iron-deficient Chlamydomonas reinhardtii. Eukaryot. Cell 6, 1841–1852. doi: 10.1128/EC.00205-07
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Amin, S. A., Parker, M. S., and Armbrust, E. V. (2012). Interactions between diatoms and bacteria. Microbiol. Mol. Biol. Rev. 76, 667–684. doi: 10.1128/MMBR.00007-12
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Barbeau, K., Moffett, J. W., Caron, D. A., Croot, P. L., and Erdner, D. L. (1996). Role of protozoan grazing in relieving iron limitation of phytoplankton. Nature 380, 61–64. doi: 10.1038/380061a0
Barkoh, A., and Fries, L. T. (2010). Aspects of the origins, ecology, and control of Golden Alga Prymnesium parvum: introduction to the featured collection. J. Am. Water Resour. Assoc. 46, 1–5. doi: 10.1111/j.1752-1688.2009.00394.x
Benjamini, Y., and Hochberg, Y. (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300.
Bertin, M. J., Zimba, P. V., Beauchesne, K. R., Huncik, K. M., and Moeller, P. D. R. (2012). The contribution of fatty acid amides to Prymnesium parvum carter toxicity. Harmful Algae 20, 117–125. doi: 10.1016/j.hal.2012.08.004
Beszteri, S., Yang, I., Jaeckisch, N., Tillman, U., Frickenhaus, S., Glöckner, G., et al. (2012). Transcriptomic response of the toxic prymnesiophyte Prymnesium parvum (N. Carter) to phosphorus and nitrogen starvation. Harmful Algae 18, 1–15. doi: 10.1016/j.hal.2012.03.003
Bik, H. M., and Thomas, W. K. (2012). Metagenomics will highlight and drive links to taxonomic data: reply to Murray. Trends Ecol. Evol. 27, 652–653. doi: 10.1016/j.tree.2012.09.005
Bockstahler, K. R., and Coats, D. W. (1993). Grazing of the mixotrophic dinoflagellate Gymnodinium sanguineum on ciliate populations of Chesapeake Bay. Mar. Biol. 116, 477–487. doi: 10.1007/BF00350065
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Brooks, B. W., James, S. V., Valenti, J. T. W., Urena-Boeck, F., Serrano, C., Berninger, J. P., et al. (2010). Comparative toxicity of Prymnesium parvum in Inland waters. J. Am. Water Resour. Assoc. 46, 45–62. doi: 10.1111/j.1752-1688.2009.00390.x
Brutemark, A., and Granéli, E. (2011). Role of mixotrophy and light for growth and survival of the toxic haptophyte Prymnesium parvum. Harmful Algae 10, 388–394. doi: 10.1016/j.hal.2011.01.005
Caron, D. A., Worden, A. Z., Countway, P. D., Demir, E., and Heidelberg, K. B. (2009). Protists are microbes too: a perspective. ISME J. 3, 4–12. doi: 10.1038/ismej.2008.101
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Carvalho, W. F., and Granéli, E. (2010). Contribution of phagotrophy versus autotrophy to Prymnesium parvum growth under nitrogen and phosphorus sufficiency and deficiency. Harmful Algae 9, 105–115. doi: 10.1016/j.hal.2009.08.007
Conesa, A., Götz, S., Garcia-Gomez, J. M., Terol, J., Talon, M., and Robles, M. (2005). Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21, 3674–3676. doi: 10.1093/bioinformatics/bti610
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Edvardsen, B., and Paasche, E. (1998). “Bloom dynamics and physiology of Prymnesium and Chrysochromulina,” in Physiological Ecology of Harmful Algal Blooms, Vol. G 41, eds D. M. Anderson, A. D. Cembella, and G. M. Hallegraef (Berlin: Springer-Verlag), 193–208.
Fistarol, G. O., Legrand, C., and Granéli, E. (2003). Allelopathic effect of Prymnesium parvum on a natural plankton community. Mar. Ecol. Prog. Ser. 255, 115–125. doi: 10.3354/meps255115
Freitag, M., Beszteri, B., Vogel, H., and John, U. (2011). Effects of physiological shock treatments on toxicity and polyketide synthase gene expression in Prymnesium parvum (Prymnesiophyceae). Eur. J. Phycol. 46, 193–201. doi: 10.1080/09670262.2011.591438
Granéli, E., Edvardsen, B., Roelke, D. L., and Hagström, J. A. (2012). The ecophysiology and bloom dynamics of Prymnesium spp. Harmful Algae 14, 260–270. doi: 10.1016/j.hal.2011.10.024
Granéli, E., and Johansson, N. (2003). Increase in the production of allelopathic substances by Prymnesium parvum cells grown under N- or P-deficient conditions. Harmful Algae 2, 135–145. doi: 10.1016/S1568-9883(03)00006-4
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Granéli, E., and Salomon, P. S. (2010). Factors influencing allelopathy and toxicity in Prymnesium parvum. J. Am. Water Resour. Assoc. 46, 108–120. doi: 10.1111/j.1752-1688.2009.00395.x
Grover, J. P., Baker, J. W., Ureña-Boeck, F., Brooks, B. W., Errera, R. M., Roelke, D. L., et al. (2007). Laboratory tests of ammonium and barley straw extract as agents to suppress abundance of the harmful alga Prymnesium parvum and its toxicity to fish. Water Res. 41, 2503–2512. doi: 10.1016/j.watres.2007.03.025
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Grover, J. P., Roelke, D. L., Brooks, B. W., Gable, G. M., Neisch, M. T., Hayden, N. J., et al. (2013). Ammonium treatments to suppress toxic blooms of Prymnesium parvum in a subtropical lake of semi-arid climate: results from in situ mesocosm experiments. Water Res. 47, 4274–4285. doi: 10.1016/j.watres.2013.05.001
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Guo, M., Harrison, P. J., and Taylor, F. J. R. (1996). Fish kills related to Prymnesium parvum N. Carter (Haptophyta) in the People's Republic of China. J. Appl. Phycol. 8, 111–117. doi: 10.1007/BF02186313
Hambright, K. D., Beyer, J. E., Easton, J. D., Zamor, R. M., Easton, A. C., and Hallidayschult, T. C. (2015). The niche of an invasive marine microbe in a subtropical freshwater impoundment. ISME J. 9, 256–264. doi: 10.1038/ismej.2014.103
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hambright, K. D., Easton, J. D., Zamor, R. M., Beyer, J., Easton, A. C., and Allison, B. (2014). Regulation of growth and toxicity of a mixotrophic microbe: implications for understanding range expansion in Prymnesium parvum. Freshwater Sci. 33, 745–754. doi: 10.1086/677198
Henrikson, J. C., Gharfeh, M. S., Easton, A. C., Easton, J. D., Glenn, K. L., Shadfan, M, et al. (2010). Reassessing the ichthyotoxin profile of cultured Prymnesium parvum (golden algae) and comparing it to samples collected from recent freshwater bloom and fish kill events in North America. Toxicon 55, 1396–1404. doi: 10.1016/j.toxicon.2010.02.017
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Huang, X., and Madan, A. (1999). CAP3: A DNA sequence assembly program. Genome Res. 9, 868–877. doi: 10.1101/gr.9.9.868
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Igarashi, T., Satake, M., and Yasumoto, T. (1996). Prymnesin-2: a potent ichthyotoxic and hemolytic glycoside isolated from the red tide alga Prymnesium parvum. J. Am. Chem. Soc. 118, 479–480. doi: 10.1021/ja9534112
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Iseli, C., Jongeneel, C. V., and Bucher, P. (1999). ESTScan: a program for detecting, evaluating, and reconstructing potential coding regions in EST sequences. Proc. Int. Conf. Intell. Syst. Mol. Biol. 138–148.
Johansson, N., and Granéli, E. (1999). Influence of different nutrient conditions on cell density, chemical composition and toxicity of Prymnesium parvum (Haptophyta) in semi-continuous cultures. J. Exp. Mar. Biol. Ecol. 239, 243–258. doi: 10.1016/S0022-0981(99)00048-9
Johnsen, T. M., Eikrem, W., Olseng, C. D., Tollefsen, K. E., and Bjerknes, V. (2010). Prymnesium parvum: the Norwegian experience. J. Am. Water Resour. Assoc. 46, 6–13. doi: 10.1111/j.1752-1688.2009.00386.x
Jones, A. C., Liao, T. S. V., Najar, F. Z., Roe, B. A., Hambright, K. D., and Caron, D. A. (2013). Seasonality and disturbance: annual pattern and response of the bacterial and microbial eukaryotic assemblages in a freshwater ecosystem. Environ. Microbiol. 15, 2557–2572. doi: 10.1111/1462-2920.12151
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jones, H. (1997). A classification of mixotrophic protists based on their behaviour. Freshwater Biol. 37, 35–43. doi: 10.1046/j.1365-2427.1997.00138.x
Keeling, P. J., Burki, F., Wilcox, H. M., Allam, B., Allen, E. E., Amaral-Zettler, L. A., et al. (2014). The Marine Microbial Eukaryote Transcriptome Sequencing Project (MMETSP): Illuminating the functional diversity of eukaryotic life in the oceans through transcriptome sequencing. PLoS Biol. 12:e1001889. doi: 10.1371/journal.pbio.1001889
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Koid, A. E., Liu, Z., Terrado, R., Jones, A. C., Caron, D. A., and Heidelberg, K. B. (2014). Comparative transcriptome analyses of four Prymnesiophyte algae. PLoS ONE 9:e97801. doi: 10.1371/journal.pone.0097801
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
La Claire, J. W. II. (2006). Analysis of expressed sequence tags from the harmful alga, Prymnesium parvum (Prymnesiophyceae, Haptophyceae). Mar. Biotechnol. 8, 534–546. doi: 10.1007/s10126-005-5182-2
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Larsen, A. (1999). Prymnesium parvum and P. patelliferum (Haptophyta) – one species. Phycologia 38, 541–543. doi: 10.2216/i0031-8884-38-6-541.1
Li, H., and Durbin, R. (2009). Fast and accurate short read alignment with Burrows-Wheeler Transform. Bioinformatics 25, 1754–1760. doi: 10.1093/bioinformatics/btp324
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Li, W., and Godzik, A. (2006). Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22, 1658–1659. doi: 10.1093/bioinformatics/btl158
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Luo, R., Liu, B., Xie, Y., Li, Z., Huang, W., Yuan, J., et al. (2012). SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1:18. doi: 10.1186/2047-217X-1-18
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lutz-Carrillo, D. J., Southard, G. M., and Fries, L. T. (2010). Global genetic relationships among isolates of Golden Alga (Prymnesium parvum). J. Am. Water Resour. Assoc. 46, 24–32. doi: 10.1111/j.1752-1688.2009.00388.x
Manning, S. R., and La Claire, J. W. II. (2010). Prymnesins: toxic metabolites of the Golden Alga, Prymnesium parvum Carter (Haptophyta). Mar. Drugs 8, 678–704. doi: 10.3390/md8030678
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Manning, S. R., and La Claire, J. W. II. (2013). Isolation of polyketides from Prymnesium parvum (Haptophyta) and their detection by liquid chromatography/mass spectrometry metabolic fingerprint analysis. Anal. Biochem. 442, 189–195. doi: 10.1016/j.ab.2013.07.034
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Maranger, R., Bird, D. F., and Price, N. M. (1998). Iron acquisition by photosynthetic marine phytoplankton from ingested bacteria. Nature 396, 248–251. doi: 10.1038/24352
McLaughlin, J. J. A. (1958). Euryhaline chrysomonads: nutrition and toxigenesis in Prymnesium parvum, with notes on Isochrysis galbana and Monochrysis lutheri. J. Protozool. 5, 75–81. doi: 10.1111/j.1550-7408.1958.tb02529.x
Murray, S. A., Patterson, D. J., and Thessen, A. E. (2012). Transcriptomics and microbial eukaryote diversity: a way forward. Trends. Ecol. Evol. 27, 651–652. doi: 10.1016/j.tree.2012.08.011
Narayanan, N. N., Ihemere, U., Chiu, W. T., Siritunga, D., Rajamani, S., Singh, S., et al. (2011). The iron assimilatory protein, FEA1, from Chlamydomonas reinhardtii facilitates iron-specific metal uptake in yeast and plants. Front. Plant, Sci. 2:67. doi: 10.3389/fpls.2011.00067
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Neuhaus, H. E., and Wagner, R. (2000). Solute pores, ion channels, and metabolite transporters in the outer and inner envelope membranes of higher plant plastids. Biochim. Biophys. Acta. 1465, 307–323. doi: 10.1016/S0005-2736(00)00146-2
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Nygaard, K., and Tobiesen, A. (1993). Bacterivory in algae: a survival strategy during nutrient limitation. Limnol. Oceanogr. 38, 273–279. doi: 10.4319/lo.1993.38.2.0273
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., et al. (2013). The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucl. Acids Res. 41, D590–D596. doi: 10.1093/nar/gks1219
Rahat, M., and Jahn, T. L. (1965). Growth of Prymnesium parvum in the dark: note on ichthyotoxin formation. J. Protozool. 12, 246–250. doi: 10.1111/j.1550-7408.1965.tb01845.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Rahat, M., and Reich, K. (1963). The B12 vitamins and growth of the flagellate Prymnesium parvum. J. Gen. Microbiol. 31, 195–202. doi: 10.1099/00221287-31-2-195
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Remmel, E. J., and Hambright, K. D. (2012). Toxin-assisted micropredation: experimental evidence shows that contact micropredation rather than exotoxicity is the role of Prymnesium toxins. Ecol. Lett. 15, 126–132. doi: 10.1111/j.1461-0248.2011.01718.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Remmel, E. J., Kohmescher, N., Larson, J. H., and Hambright, K. D. (2011). An experimental analysis of harmful algae-zooplankton interactions and the ultimate defense. Limnol. Oceanogr. 56, 461–470. doi: 10.4319/lo.2011.56.2.0461
Robertson, G., Schein, J., Chiu, R., Corbett, R., Field, M., Jackman, S. D., et al. (2010). De novo assembly and analysis of RNA-seq data. Nat. Methods 7, 909–912. doi: 10.1038/nmeth.1517
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Robinson, M. D., McCarthy, D. J., and Smyth, G. K. (2010). edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140. doi: 10.1093/bioinformatics/btp616
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Roelke, D. L., Grover, J. P., Brooks, B. W., Glass, J., Buzan, D., Southard, G. M., et al. (2011). A decade of fish-killing Prymnesium parvum blooms in Texas: roles of inflow and salinity. J. Plankton Res. 33, 243–253. doi: 10.1093/plankt/fbq079
Rudra, D., and Warner, J. R. (2004). What better measure than ribosome synthesis? Genes Dev. 18, 2431–2436. doi: 10.1101/gad.1256704
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sanders, R. W. (2011). Alternative nutritional strategies in protists: symposium introduction and a review of freshwater protists that combine photosynthesis and heterotrophy. J. Eukaryot. Microbiol. 58, 181–184. doi: 10.1111/j.1550-7408.2011.00543.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Schmidt, L. E, and Hansen, P. J. (2001). Allelopathy in the prymnesiophyte Chrysochromulina polylepis: effect of cell concentration, growth phase and pH. Mar. Ecol. Prog. Ser. 216, 67–81. doi: 10.3354/meps216067
Schug, K. A., Skingel, T. R., Spencer, S. E., Serrano, C. A., Le, C. Q., Schug, C. A., et al. (2010). Hemolysis, fish mortality, and LC-ESI-MS of cultured crude and fractionated Golden Alga (Prymnesium parvum). J. Am. Water Resour. Assoc. 46, 33–44. doi: 10.1111/j.1752-1688.2009.00389.x
Simpson, J. T., Wong, K., Jackman, S. D., Schein, J. E., Jones, S. J., and Birol, I. (2009). ABySS: a parallel assembler for short read sequence data. Genome Res. 19, 1117–1123. doi: 10.1101/gr.089532.108
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Smalley, G. W., and Coats, D. W. (2002). Ecology of the red-tide dinoflagellate Ceratium furca: distribution, mixotrophy, and grazing impact on ciliate populations of Chesapeake Bay. J. Eukaryot. Microbiol. 49, 63–73. doi: 10.1111/j.1550-7408.2002.tb00343.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sopanen, S., Koski, M., Kuuppo, P., Uronen, P., Legrand, C., and Tamminen, T. (2006). Toxic haptophyte Prymnesium parvum affects grazing, survival, egestion and egg production of the calanoid copepods Eurytemora affinis and Acartia bifilosa. Mar. Ecol. Progr. Ser. 327, 223–232. doi: 10.3354/meps327223
Southard, G. M., Fries, L. T., and Barkoh, A. (2010). Prymnesium parvum: the Texas experience. J. Am. Water Resour. Assoc. 46, 14–23. doi: 10.1111/j.1752-1688.2009.00387.x
Stearman, R., Yuan, D. S., Yamaguchi-Iwai, Y., Klausner, R. D., and Dancis, A. (1996). A permease-oxidase complex involved in high-affinity iron uptake in yeast. Science 271, 1552–1557. doi: 10.1126/science.271.5255.1552
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Sunda, W. G., Granéli, E., and Gobler, C. J. (2006). Positive feedback and the development and persistence of ecosystem disruptive algal blooms. J. Phycol. 42, 963–974. doi: 10.1111/j.1529-8817.2006.00261.x
Tillmann, U. (2003). Kill and eat your predator: a winning strategy of the planktonic flagellate Prymnesium parvum. Aq. Microb. Ecol. 32, 73–84. doi: 10.3354/ame032073
Vallon, O., Bulté, L., Kuras, R., Olive, J., and Wollman, F. A. (1993). Extensive accumulation of an extracellular L-amino-acid oxidase during gametogenesis of Chlamydomonas reinhardtii. Eur. J. Biochem. 215, 351–360. doi: 10.1111/j.1432-1033.1993.tb18041.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Watmough, N. J., and Frerman, F. E. (2010). The electron transfer flavoprotein: ubiquinone oxidoreductaases. Biochim. Biophys. Acta 1797, 1910–1916. doi: 10.1016/j.bbabio.2010.10.007
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Keywords: Prymnesium parvum, transcriptomics, gene expression, mixotrophic protist, golden algae, harmful algae
Citation: Liu Z, Jones AC, Campbell V, Hambright KD, Heidelberg KB and Caron DA (2015) Gene expression in the mixotrophic prymnesiophyte, Prymnesium parvum, responds to prey availability. Front. Microbiol. 6:319. doi: 10.3389/fmicb.2015.00319
Received: 05 December 2014; Accepted: 30 March 2015;
Published: 20 April 2015.
Edited by:
Senjie Lin, University of Connecticut, USAReviewed by:
Tatiana Rynearson, University of Rhode Island, USATom O. Delmont, Marine Biological Lab, USA
Copyright © 2015 Liu, Jones, Campbell, Hambright, Heidelberg and Caron. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zhenfeng Liu, Department of Biological Sciences, University of Southern California, 3616 Trousdale Parkway, AHF 231, Los Angeles, CA 90089-0371, USAemhlbmZlbmcubGl1MUBnbWFpbC5jb20=
 Zhenfeng Liu
Zhenfeng Liu Adriane C. Jones1
Adriane C. Jones1 Victoria Campbell
Victoria Campbell K. David Hambright
K. David Hambright Karla B. Heidelberg
Karla B. Heidelberg David A. Caron
David A. Caron