A commentary on
Novel pili-like surface structures of Halobacterium salinarum strain R1 are crucial for surface adhesion
by Losensky, G., Vidakovic, L., Klingl, A., Pfeifer, F., and Fröls, S. (2014). Front. Microbiol. 5:755. doi: 10.3389/fmicb.2014.00755
Type IV pili (T4P) are a group of cell surface appendages of particular interest due to broad conservation and functional versatility across the domains Bacteria and Archaea (Albers and Meyer, 2011; Giltner et al., 2012). All T4P are composed of small protein subunits known as pilins that polymerize into helical fibers through the action of assembly ATPases (Giltner et al., 2012). This core ancestral machinery has been adapted in various lineages for many cellular processes–from adhesion and biofilm formation, to motility, horizontal gene transfer (HGT) and even electricity conduction (Giltner et al., 2012; Berry and Pelicic, 2015). When T4P structures are involved in adhesion, they are known as pili, if they no longer mediate attachment, but are associated with another function, such as scavenging macromolecules (e.g., DNA uptake by Com proteins in Bacillus subtilis), or secretion of proteins through a piston-like structure (i.e., type II secretion), they are called pseudopili (Averhoff and Friedrich, 2003; Peabody et al., 2003; Chen et al., 2005). T4P appendages may also contribute to both adhesion and another function. This dual function is sometimes true for archaella: a major group of archaeal T4P appendages characterized by the ability to rotate and enable swimming motility. Archaella are functionally analogous yet structurally and genetically unrelated to bacterial flagella (Jarrell and Albers, 2012; Shahapure et al., 2014; Albers and Jarrell, 2015). T4P have been studied to a greater extent in bacteria, in part because they are often virulence factors (Giltner et al., 2012). However, recent investigations have revealed a repertoire of archaeal T4P–highlighting implications for regulatory complexity and functional diversity.
Losensky et al. (2014) demonstrated that adhesive filaments in the haloarchaeon Halobacterium salinarum R1 observed during biofilm formation (Fröls et al., 2012) are dependent on the pilus assembly ATPase gene pilB1 (Losensky et al., 2014), expanding the list of experimentally studied archaeal T4P (Table 1). Deletion of pilB1 led to a lack of pili as observed through electron microscopy and a defect in adhesion. Only 4% of a glass surface was colonized by non-piliated/non-archaellated cells (ΔflaI/ΔpilB1), relative to 36 and 44% for the parental and non-archaellated (ΔflaI) strains, respectively. The molecular composition of PilB1-dependent pili has not yet been determined, however Losensky and coauthors noted that there are over 30 candidate pilins in the Hbt. salinarum R1 genome, as indicated by the class III signal peptide prediction program FlaFind (Szabó et al., 2007b). FlaFind was used previously to show that most archaeal genomes contain many pilin/archaellin homologs (Szabó et al., 2007b; Esquivel et al., 2013). For example, Haloarcula marismortui and Haloferax volcanii have nearly 50 putative pilin/archaellin precursors (Esquivel et al., 2013).
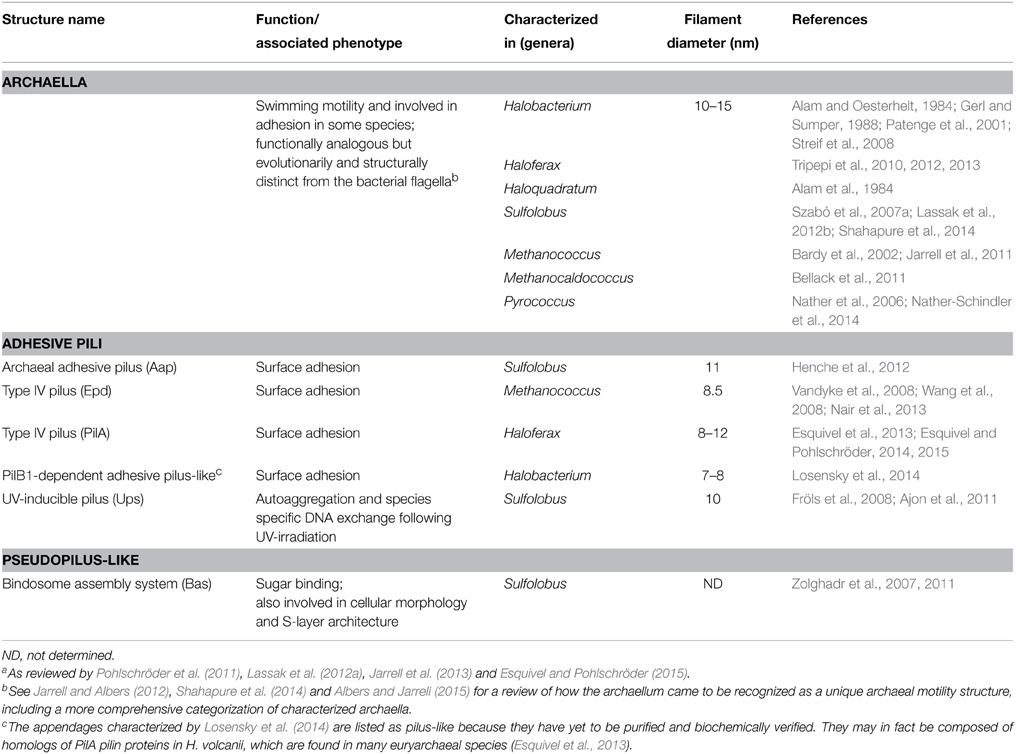
Table 1. Experimentally studied type IV pili in archaeal species: archaella, adhesive pili, and pseudopilia.
Some of these pilins could be associated with additional functions. Hfx. volcanii has an ability for social motility in static liquid (Chimileski et al., 2014) and T4P could be involved in this activity (Esquivel and Pohlschröder, 2015), whereby they may attach to extracellular matrix along the substratum, similar to the S-motility system that pulls Myxococcus xanthus cells forward (Hodgkin and Kaiser, 1979; Zusman et al., 2007). There could be more archaeal T4P-related surface structures that scavenge macromolecules as well, like the bindosome of Sulfolobus solfataricus (Zolghadr et al., 2007, 2011).
Investigations of archaeal T4P leave open the possibility for undiscovered mechanisms for contacting abiotic surfaces or other cells. For instance, even in the non-piliated/non-archaellated Hbt. salinarum strain, adhesion was not completely abolished (Losensky et al., 2014). Similar residual adhesion has been observed in Hfx. volcanii (Tripepi et al., 2010, 2013). In both cases, pilins that remain present in the membrane but cannot be assembled into pili without the assembly ATPase(s) likely explain low levels of adhesion (Esquivel and Pohlschröder, 2014). There are two other cell-to-cell contact phenomena in Hfx. volcanii that do not require archaella or pili: Ca2+ dependent autoaggregation (Tripepi et al., 2010), and an HGT mechanism known as mating (Rosenshine et al., 1989; Tripepi et al., 2010; Naor et al., 2012). Additional types of extracellular polymers or fibers found in bacterial species could be present in archaea, such as amyloid protein (Chimileski et al., 2014). Unusual, genetically ambiguous non-T4P structures have already been observed in other archaeal species, including the hamus of the SM1 euryarchaeon (Moissl et al., 2005; Perras et al., 2014) and the cannulae of Pyrodictium cells (Nickell et al., 2003).
A plausible explanation for having a wide array of appendages is a capacity for differential regulation (Jarrell, 2012; Lassak et al., 2012a; Jarrell et al., 2013). Indeed, a number of studies point to dynamic regulatory systems controlling archaeal T4P. In Hbt. salinarum, pilB1 expression was upregulated relative to flaI in adherent cells (Losensky et al., 2014), suggesting archaella and pili have antagonistically regulated functions in motility (when a planktonic state is favorable) and for adhesion (during biofilm formation), as in Hfx. volcanii (Tripepi et al., 2010; Esquivel and Pohlschröder, 2014, 2015). Haloarcula marismortui has two archaellins that are expressed under different temperatures and salinities (Syutkin et al., 2014), termed ecoparalogs. Likewise, the six Hfx. volcanii pilA paralogs, any one of which can restore adhesion when expressed in a null mutant [ΔpilA(1–6)] (Esquivel et al., 2013), may be ecoparalogs as well. Intriguingly, deleting flgA2, one of two archaellin genes in Hfx. volcanii, produced a hypermotile phenotype, rather than a motility defect (Tripepi et al., 2013). Archaellins are also regulated through N-glycosylation (Guan et al., 2012; Tripepi et al., 2012) and regulatory proteins controlling adhesive pili and archaella have been identified in Sulfolobus acidocaldarius (Reimann et al., 2012; Orell et al., 2013; Vassart et al., 2013).
As more T4P are described in archaeal groups, a common theme is appearing. A multitude of individual pilins/archaellins from one or more loci may appear to be redundant–contributing to appendages that are difficult to differentiate through electron microscopy and often depend on the same assembly ATPase. However, to the contrary, the maintenance of more than one pilus and archaellum subunit gene is likely due to a complex regulatory network and the corresponding advantages of functional versatility. Subsets of pilins may be expressed in different combinations as a response to a variety of specific environmental conditions and/or cellular functions.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
RTP was supported by the National Science Foundation (award numbers DEB-0910290 and DEB-0830024) and the NASA Astrobiology: Exobiology and Evolutionary Biology Program (grant number NNX12AD70G).
References
Ajon, M., Fröls, S., Van Wolferen, M., Stoecker, K., Teichmann, D., Driessen, A. J., et al. (2011). UV-inducible DNA exchange in hyperthermophilic archaea mediated by type IV pili. Mol. Microbiol. 82, 807–817. doi: 10.1111/j.1365-2958.2011.07861.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Alam, M., Claviez, M., Oesterhelt, D., and Kessel, M. (1984). Flagella and motility behavior of square bacteria. EMBO J. 3, 2899–2903.
Alam, M., and Oesterhelt, D. (1984). Morphology, function and isolation of halobacterial flagella. J. Mol. Biol. 176, 459–475.
Albers, S.-V., and Jarrell, K. F. (2015). The archaellum: how archaea swim. Front. Microbiol. 6:23. doi: 10.3389/fmicb.2015.00023
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Albers, S. V., and Meyer, B. H. (2011). The archaeal cell envelope. Nat. Rev. Microbiol. 9, 414–426. doi: 10.1038/nrmicro2576
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Averhoff, B., and Friedrich, A. (2003). Type IV pili-related natural transformation systems: DNA transport in mesophilic and thermophilic bacteria. Arch. Microbiol. 180, 385–393. doi: 10.1007/s00203-003-0616-6
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bardy, S. L., Mori, T., Komoriya, K., Aizawa, S. I., and Jarrell, K. F. (2002). Identification and localization of flagellins FlaA and FlaB3 within flagella of Methanococcus voltae. J. Bacteriol. 184, 5223–5233. doi: 10.1128/jb.184.19.5223-5233.2002
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Bellack, A., Huber, H., Rachel, R., Wanner, G., and Wirth, R. (2011). Methanocaldococcus villosus sp. nov., a heavily flagellated archaeon that adheres to surfaces and forms cell-cell contacts. Int. J. Syst. Evol. Microbiol. 61, 1239–1245. doi: 10.1099/ijs.0.023663-0
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Berry, J. L., and Pelicic, V. (2015). Exceptionally widespread nanomachines composed of type IV pilins: the prokaryotic Swiss Army knives. FEMS Microbiol. Rev. 39, 1–21. doi: 10.1093/femsre/fuu001
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Chen, I., Christie, P. J., and Dubnau, D. (2005). The ins and outs of DNA transfer in bacteria. Science 310, 1456–1460. doi: 10.1126/science.1114021
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Chimileski, S., Franklin, M. J., and Papke, R. T. (2014). Biofilms formed by the archaeon Haloferax volcanii exhibit cellular differentitaion and social motility, and facilitate horizontal gene transfer. BMC Biol. 12:65. doi: 10.1186/s12915-014-0065-5
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Esquivel, R. N., and Pohlschröder, M. (2014). A conserved type IV pilin signal peptide H-domain is critical for the post-translational regulation of flagella-dependent motility. Mol. Microbiol. 93, 494–504. doi: 10.1111/mmi.12673
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Esquivel, R. N., and Pohlschröder, M. (2015). Archaeal type IV pili and their involvement in biofilm formation. Front. Microbiol. 6:190. doi: 10.3389/fmicb.2015.00190
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Esquivel, R. N., Xu, R., and Pohlschröder, M. (2013). Novel archaeal adhesion pilins with a conserved N terminus. J. Bacteriol. 195, 3808–3818. doi: 10.1128/JB.00572-13
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Fröls, S., Ajon, M., Wagner, M., Teichmann, D., Zolghadr, B., Folea, M., et al. (2008). UV-inducible cellular aggregation of the hyperthermophilic archaeon Sulfolobus solfataricus is mediated by pili formation. Mol. Microbiol. 70, 938–952. doi: 10.1111/j.1365-2958.2008.06459.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Fröls, S., Dyall-Smith, M., and Pfeifer, F. (2012). Biofilm formation by haloarchaea. Environ. Microbiol. 14, 3159–3174. doi: 10.1111/j.1462-2920.2012.02895.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Gerl, L., and Sumper, M. (1988). Halobacterial flagellins are encoded by a multigene family. J. Biol. Chem. 263, 13246–13251.
Giltner, C. L., Nguyen, Y., and Burrows, L. L. (2012). Type IV pilin proteins: versatile molecular modules. Microbiol. Mol. Biol. Rev. 76, 740–772. doi: 10.1128/MMBR.00035-12
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Guan, Z., Naparstek, S., Calo, D., and Eichler, J. (2012). Protein glycosylation as an adaptive response in Archaea: growth at different salt concentrations leads to alterations in Haloferax volcanii S-layer glycoprotein N-glycosylation. Environ. Microbiol. 14, 743–753. doi: 10.1111/j.1462-2920.2011.02625.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Henche, A. L., Ghosh, A., Yu, X., Jeske, T., Egelman, E., and Albers, S. V. (2012). Structure and function of the adhesive type IV pilus of Sulfolobus acidocaldarius. Environ. Microbiol. 14, 3188–3202. doi: 10.1111/j.1462-2920.2012.02898.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Hodgkin, J., and Kaiser, D. (1979). Genetics of gliding motility in Myxococcus xanthus (Myxobacterales): two gene systems control movement. Mol. Gen. Gentics 171, 177–191. doi: 10.1007/BF00270004
Jarrell, K. F. (2012). Control of archaellation in Sulfolobus acidocaldarius: unravelling of the regulation of surface structure biosynthesis in Archaea begins. Mol. Microbiol. 86, 1–5. doi: 10.1111/j.1365-2958.2012.08191.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jarrell, K. F., and Albers, S. V. (2012). The archaellum: an old motility structure with a new name. Trends Microbiol. 20, 307–312. doi: 10.1016/j.tim.2012.04.007
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jarrell, K. F., Stark, M., Nair, D. B., and Chong, J. P. (2011). Flagella and pili are both necessary for efficient attachment of Methanococcus maripaludis to surfaces. FEMS Microbiol. Lett. 319, 44–50. doi: 10.1111/j.1574-6968.2011.02264.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Jarrell, K. F., Ding, Y., Nair, D. B., and Siu, S. (2013). Surface appendages of Archaea: structure, function, genetics and assembly. Life 3, 86–117. doi: 10.3390/life3010086
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lassak, K., Ghosh, A., and Albers, S. V. (2012a). Diversity, assembly and regulation of archaeal type IV pili-like and non-type-IV pili-like surface structures. Res. Microbiol. 163, 630–644. doi: 10.1016/j.resmic.2012.10.024
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Lassak, K., Neiner, T., Ghosh, A., Klingl, A., Wirth, R., and Albers, S. V. (2012b). Molecular analysis of the crenarchaeal flagellum. Mol. Microbiol. 83, 110–124. doi: 10.1111/j.1365-2958.2011.07916.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Losensky, G., Vidakovic, L., Klingl, A., Pfeifer, F., and Fröls, S. (2014). Novel pili-like surface structures of Halobacterium salinarum strain R1 are crucial for surface adhesion. Front. Microbiol. 5:755. doi: 10.3389/fmicb.2014.00755
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Moissl, C., Rachel, R., Briegel, A., Engelhardt, H., and Huber, R. (2005). The unique structure of archaeal “hami,” highly complex cell appendages with nano-grappling hooks. Mol. Microbiol. 56, 361–370. doi: 10.1111/j.1365-2958.2005.04294.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Nair, D. B., Chung, D. K., Schneider, J., Uchida, K., Aizawa, S., and Jarrell, K. F. (2013). Identification of an additional minor pilin essential for piliation in the archaeon Methanococcus maripaludis. PLoS ONE 8:e83961. doi: 10.1371/journal.pone.0083961
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Naor, A., Lapierre, P., Mevarech, M., Papke, R. T., and Gophna, U. (2012). Low species barriers in halophilic archaea and the formation of recombinant hybrids. Curr. Biol. 22, 1444–1448. doi: 10.1016/j.cub.2012.05.056
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Nather, D. J., Rachel, R., Wanner, G., and Wirth, R. (2006). Flagella of Pyrococcus furiosus: multifunctional organelles, made for swimming, adhesion to various surfaces, and cell-cell contacts. J. Bacteriol. 188, 6915–6923. doi: 10.1128/JB.00527-06
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Nather-Schindler, D. J., Schopf, S., Bellack, A., Rachel, R., and Wirth, R. (2014). Pyrococcus furiosus flagella: biochemical and transcriptional analyses identify the newly detected flaB0 gene to encode the major flagellin. Front. Microbiol. 5:695. doi: 10.3389/fmicb.2014.00695
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Nickell, S., Hergel, R., Baumeisiter, W., and Rachel, R. (2003). Pyrodictium cannulae enter the periplasmic space but do not enter the cytoplasm, as revealed by cryo-electron tomography. J. Struct. Biol. 141, 34–42. doi: 10.1016/S1047-8477(02)00581-6
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Orell, A., Peeters, E., Vassen, V., Jachlewski, S., Schalles, S., Siebers, B., et al. (2013). Lrs14 transcriptional regulators influence biofilm formation and cell motility of Crenarchaea. ISME J. 7, 1886–1898. doi: 10.1038/ismej.2013.68
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Patenge, N., Berendes, A., Engelhardt, H., Schuster, S. C., and Oesterhelt, D. (2001). The fla gene cluster is involved in the biogenesis of flagella in Halobacterium salinarum. Mol. Microbiol. 41, 653–663. doi: 10.1046/j.1365-2958.2001.02542.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Peabody, C. R., Chung, Y. J., Yen, M. R., Vidal-Ingigliardi, D., Pugsley, A. P., and Saier, M. H. (2003). Type II protein secretion and its relationship to bacterial type IV pili and archaeal flagella. Microbiology 149, 3051–3072. doi: 10.1099/mic.0.26364-0
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Perras, A. K., Wanner, G., Klingl, A., Mora, M., Auerbach, A. K., Heinz, V., et al. (2014). Grappling archaea: ultrastructural analyses of an uncultivated, cold-loving archaeon, and its biofilm. Front. Microbiol. 5:397. doi: 10.3389/fmicb.2014.00397
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Pohlschröder, M., Ghosh, A., Tripepi, M., and Albers, S. V. (2011). Archaeal type IV pilus-like structures–evolutionarily conserved prokaryotic surface organelles. Curr. Opin. Microbiol. 14, 357–363. doi: 10.1016/j.mib.2011.03.002
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Reimann, J., Lassak, K., Khadouma, S., Ettema, T. J., Yang, N., Driessen, A. J., et al. (2012). Regulation of archaella expression by the FHA and von Willebrand domain-containing proteins ArnA and ArnB in Sulfolobus acidocaldarius. Mol. Microbiol. 86, 24–36. doi: 10.1111/j.1365-2958.2012.08186.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Rosenshine, I., Tchelet, R., and Mevarech, M. (1989). The mechanism of DNA transfer in the mating system of an archaebacterium. Science 245, 1387–1389.
Shahapure, R., Driessen, R. P., Haurat, M. F., Albers, S. V., and Dame, R. T. (2014). The archaellum: a rotating type IV pilus. Mol. Microbiol. 91, 716–723. doi: 10.1111/mmi.12486
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Streif, S., Staudinger, W. F., Marwan, W., and Oesterhelt, D. (2008). Flagellar rotation in the archaeon Halobacterium salinarum depends on ATP. J. Mol. Biol. 384, 1–8. doi: 10.1016/j.jmb.2008.08.057
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Syutkin, A. S., Pyatibratov, M. G., Galzitskaya, O. V., Rodriguez-Valera, F., and Fedorov, O. V. (2014). Haloarcula marismortui archaellin genes as ecoparalogs. Extremophiles 18, 341–349. doi: 10.1007/s00792-013-0619-4
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Szabó, Z., Sani, M., Groeneveld, M., Zolghadr, B., Schelert, J., Albers, S. V., et al. (2007a). Flagellar motility and structure in the hyperthermoacidophilic archaeon Sulfolobus solfataricus. J. Bacteriol. 189, 4305–4309. doi: 10.1128/JB.00042-07
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Szabó, Z., Stahl, A. O., Albers, S. V., Kissinger, J. C., Driessen, A. J., and Pohlschröder, M. (2007b). Identification of diverse archaeal proteins with class III signal peptides cleaved by distinct archaeal prepilin peptidases. J. Bacteriol. 189, 772–778. doi: 10.1128/JB.01547-06
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Tripepi, M., Esquivel, R. N., Wirth, R., and Pohlschröder, M. (2013). Haloferax volcanii cells lacking the flagellin FlgA2 are hypermotile. Microbiology 159, 2249–2258. doi: 10.1099/mic.0.069617-0
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Tripepi, M., Imam, S., and Pohlschröder, M. (2010). Haloferax volcanii flagella are required for motility but are not involved in PibD-dependent surface adhesion. J. Bacteriol. 192, 3093–3102. doi: 10.1128/JB.00133-10
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Tripepi, M., You, J., Temel, S., Onder, O., Brisson, D., and Pohlschröder, M. (2012). N-glycosylation of Haloferax volcanii flagellins requires known Agl proteins and is essential for biosynthesis of stable flagella. J. Bacteriol. 194, 4876–4887. doi: 10.1128/JB.00731-12
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Vandyke, D. J., Wu, J., Ng, S. Y., Kanbe, M., Chaban, B., Aizawa, S., et al. (2008). Identification of a putative acetyltransferase gene, MMP0350, which affects proper assembly of both flagella and pili in the archaeon Methanococcus maripaludis. J. Bacteriol. 190, 5300–5307. doi: 10.1128/JB.00474-08
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Vassart, A., Van Wolferen, M., Orell, A., Hong, Y., Peeters, E., Albers, S. V., et al. (2013). Sa-Lrp from Sulfolobus acidocaldarius is a versatile, glutamine-responsive, and architectural transcriptional regulator. Microbiologyopen 2, 75–93. doi: 10.1002/mbo3.58
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Wang, Y. A., Yu, X., Ng, S. Y., Jarrell, K. F., and Egelman, E. H. (2008). The structure of an archaeal pilus. J. Mol. Biol. 381, 456–466. doi: 10.1016/j.jmb.2008.06.017
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zolghadr, B., Klingl, A., Rachel, R., Driessen, A. J., and Albers, S. V. (2011). The bindosome is a structural component of the Sulfolobus solfataricus cell envelope. Extremophiles 15, 235–244. doi: 10.1007/s00792-010-0353-0
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zolghadr, B., Weber, S., Szabó, Z., Driessen, A. J., and Albers, S. V. (2007). Identification of a system required for the functional surface localization of sugar binding proteins with class III signal peptides in Sulfolobus solfataricus. Mol. Microbiol. 64, 795–806. doi: 10.1111/j.1365-2958.2007.05697.x
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Zusman, D. R., Scott, A. E., Yang, Z., and Kirby, J. R. (2007). Chemosensory pathways, motility and development in Myxococcus xanthus. Nat. Rev. Microbiol. 5, 862–872. doi: 10.1038/nrmicro1770
PubMed Abstract | Full Text | CrossRef Full Text | Google Scholar
Keywords: haloarchaea, type IV pili, archaellum, surface adhesion, archaeal biofilm formation, microbial cell-to-cell interactions
Citation: Chimileski S and Papke RT (2015) Getting a hold on archaeal type IV pili: an expanding repertoire of cellular appendages implicates complex regulation and diverse functions. Front. Microbiol. 6:362. doi: 10.3389/fmicb.2015.00362
Received: 08 January 2015; Accepted: 10 April 2015;
Published: 05 May 2015.
Edited by:
Biswarup Mukhopadhyay, Virginia Tech, USAReviewed by:
Sonja-Verena Albers, University of Freiburg, GermanyJerry Eichler, Ben-Gurion University of the Negev, Israel
Mechthild Pohlschröder, University of Pennsylvania, USA
Copyright © 2015 Chimileski and Papke. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Scott Chimileski,c2NvdHQuY2hpbWlsZXNraUB1Y29ubi5lZHU=
 Scott Chimileski
Scott Chimileski R. Thane Papke
R. Thane Papke