- 1GQE-Le Moulon, INRA, Université Paris-Sud, CNRS, AgroParisTech, Université Paris-Saclay, Gif-sur-Yvette, France
- 2Micalis Institute, INRA, AgroParisTech, CIRM-Levures, Université Paris-Saclay, Jouy-en-Josas, France
- 3SPO, INRA, Montpellier SupAgro, Univ Montpellier, Montpellier, France
Microbial communities are essential for the maintenance and functioning of ecosystems, including fermented food ecosystems. The analysis of food microbial communities is mainly focused on lactic acid bacteria (LAB), while yeast diversity is less understood. Here, we describe the fungal diversity of a typical food fermented product, sourdough bread. The species diversity of 14 sourdoughs collected from bakeries located all over France was analyzed. Bakeries were chosen to represent diverse bakery practices and included bakers and farmer-bakers. Both non-culture-based (pyrosequencing of Internal Transcribed Spacer 1 amplicons) and culture-based methods were used. While both identification methods were in agreement regarding the dominant yeast species of each sourdough, the ITS1 metabarcoding analysis identified an increased number of fungal species in sourdough communities. Two third of the identified sequences obtained from sourdoughs were Saccharomycetales, mostly in the Kazachstania genus. No Saccharomycetales species was shared by all the sourdoughs, whereas five other fungal species, mainly known plant pathogens, were found in all sourdoughs. Interestingly, Saccharomyces cerevisiae, known as “baker’s yeast,” was identified as the dominant species in only one sourdough. By contrast, five Kazachstania species were identified as the dominant sourdough species, including one recently described Kazachstania species, Kazachstania saulgeensis and an undescribed Kazachstania sp. Sourdoughs from farmer-bakers harbored Kazachstania bulderi, Kazachstania unispora and two newly described Kazachstania species, while sourdough from bakers mostly carried Kazachstania humilis as the dominant species. Such yeast diversity has not been found in sourdoughs before, highlighting the need to maintain different traditional food practices to conserve microbial diversity.
Introduction
Microbes colonize all types of environments and are necessary for the function of all ecosystems, including human domesticated environments. They are commonly used for the fermentation of a wide range of primary products as diverse as milk, cereals, legumes, plant roots, vegetables, fish, and meat (for a review see Tamang et al., 2016). They play key roles in food texture and flavor (see for bread, Martinez-Anaya et al., 1990; Damiani et al., 1996; Hansen and Schieberle, 2005; Csoma et al., 2010; Birch et al., 2013) as well as in bio-preservation and nutritional compounds (Caplice and Fitzgerald, 1999; Bourdichon et al., 2012; Gobbetti et al., 2014, 2018; Gänzle and Ripari, 2016). The microbial community involved in the fermentation process include lactic acid bacteria (LAB) as well as fungi, particularly yeasts (reviewed in De Vuyst et al., 2014: De Vuyst et al., 2016; Carbonetto et al., 2018).
Microbial communities of fermented food products have long been studied using culture-based microbiological methods, in which a restricted number of cultivable clones are isolated after growth on different media, allowing the detection of only a fraction of the microbial population. In the last decade, metabarcoding approach, based on the sequencing of bacteria or eukaryotic markers from environmental DNA, became highly popular (reviewed in van Hijum et al., 2013; Kergourlay et al., 2015; De Filippis et al., 2017). This approach first focused on bacterial species and was more recently used to study fungi (reviewed in De Filippis et al., 2017). As a result, the number of publications describing fungal diversity using metabarcoding analysis remains lower than those describing bacteria (De Filippis et al., 2017). Fungal metabarcoding analysis of fermented products mostly focused on dairy products and alcoholic beverages (De Filippis et al., 2017). The analysis of fungal communities of plant-fermented food is less numerous but includes the cacao bean (Illeghems et al., 2012), the green olive (Arroyo-López et al., 2016), douchi (Yang et al., 2016), pu-erh tea (Zhang et al., 2016), kombucha (Reva et al., 2015; Coton et al., 2017), paocai (Liang et al., 2018), and sourdough (Minervini et al., 2015). The internal transcribed spacers (ITS1 and ITS2), including two non-coding parts of the ribosomal DNA unit situated between the small subunit (SSU) and the large subunit (LSU) rRNA genes, are the most frequently used fungal barcodes (Schoch et al., 2012; Vu et al., 2016, 2019). These barcodes have been recognized as a primary barcode in fungi and appears to better discriminate yeast species compared to the D1/D2 domains of the LSU region (Martin and Rygiewicz, 2005; Schoch et al., 2012; Vu et al., 2016). Amplification of either ITS1 or ITS2 was shown to be informative in describing fungal community composition. However, the analysis of fungal diversity is more difficult compared to the bacterial diversity for a variety of reason. First, the D1/D2 domains might be too much conserved to discriminate species. The ITS sequences are better to discriminate species but they harbor size polymorphisms, which render sequence alignment and comparison more tricky. Second, some ITS sequences are so small that they can be lost in the process of amplification and library making. Third, the ITS databases are either curated but not exhaustive (such as UNITE) or present a lot of redundancy (such as NCBI).
Among plant-fermented food, bread is of importance both from a cultural and a nutritive point of view. Leavened bread can be made using either yeast starter or using sourdough. Three types of sourdoughs are defined. Type I sourdough is a mixture of flour, water, bacteria and yeasts. It is a naturally fermented sourdough and is the traditional sourdough. Type II and type III sourdoughs are industrial types of sourdough. Usually, type I sourdough contains one dominant LAB species and one dominant yeast species, as well as a few other minor microbial species (Lhomme et al., 2015a, 2016; Minervini et al., 2015; Zhang et al., 2011 2015; Dertli et al., 2016; Liu et al., 2016, 2018; Michel et al., 2016; De Vuyst et al., 2017). According to the literature, the most frequent LAB dominant species of sourdough are Lactobacillus sanfranciscensis, L. brevis or L. plantarum, and the most frequent yeast species are Saccharomyces cerevisiae, Kazachstania humilis (Jacques et al., 2016), Pichia kudriavzevii, Torulaspora delbruecki, and Wickerhamomyces anomalus (reviewed in De Vuyst et al., 2017; Carbonetto et al., 2018). Bacterial species diversity appears to be shaped by several drivers, including the bakery environment and technical processes (Scheirlinck et al., 2007; Minervini et al., 2014, 2015, 2018; reviewed in Gobbetti et al., 2016; Van Kerrebroeck et al., 2017). While the factors driving bacterial diversity have been studied, those shaping the distribution of yeasts remain poorly explored.
In Europe, the microbial species diversity of wheat sourdoughs has been well described in Italy (Corsetti et al., 2001; Foschino et al., 2004; Minervini et al., 2011; Gobbetti and Gänzle, 2012; Lattanzi et al., 2013; Minervini et al., 2014, 2015; Pontonio et al., 2016) and Belgium (Scheirlinck et al., 2007, 2009; Vrancken et al., 2010). Although bread is a traditional food product in France, few studies have analyzed sourdoughs’ microbial diversity (Lhomme et al., 2015a,b, 2016; Michel et al., 2016). In France, bread is often made from wheat either by farmer-bakers or by bakers. The dedicated term “farmer-baker” describe someone involved in both farming and bread production (Demeulenaere and Bonneuil, 2010).
Bread making practices are diverse, especially among bakeries that make bread with organic flour. Previous culture and non-culture-based analysis of bacterial species diversity in French sourdoughs revealed a low level of species diversity, even among sourdoughs made with organic flour and with different bread-making practices (Lhomme et al., 2015b, 2016; Michel et al., 2016). Lactobacillus sanfranciscensis was found as the main dominant species. The objective of this study was to describe the pattern of fungal species diversity in a collection of type I sourdoughs made with organic flour coming from bakeries with different bread-making practices and from different French regions using culture- and non-culture-based methods.
Materials and Methods
Field Survey and Sourdough Collection
A total of 14 bakers located in different French regions participated to this study. The study was carried out jointly by two French researcher teams. One worked on bacterial species, the other one on yeasts. The data on bacterial species diversity and biochemical properties of sourdoughs and breads have been previously published (Lhomme et al., 2015b). We analyzed the yeast species diversity as well as the bread-making practices.
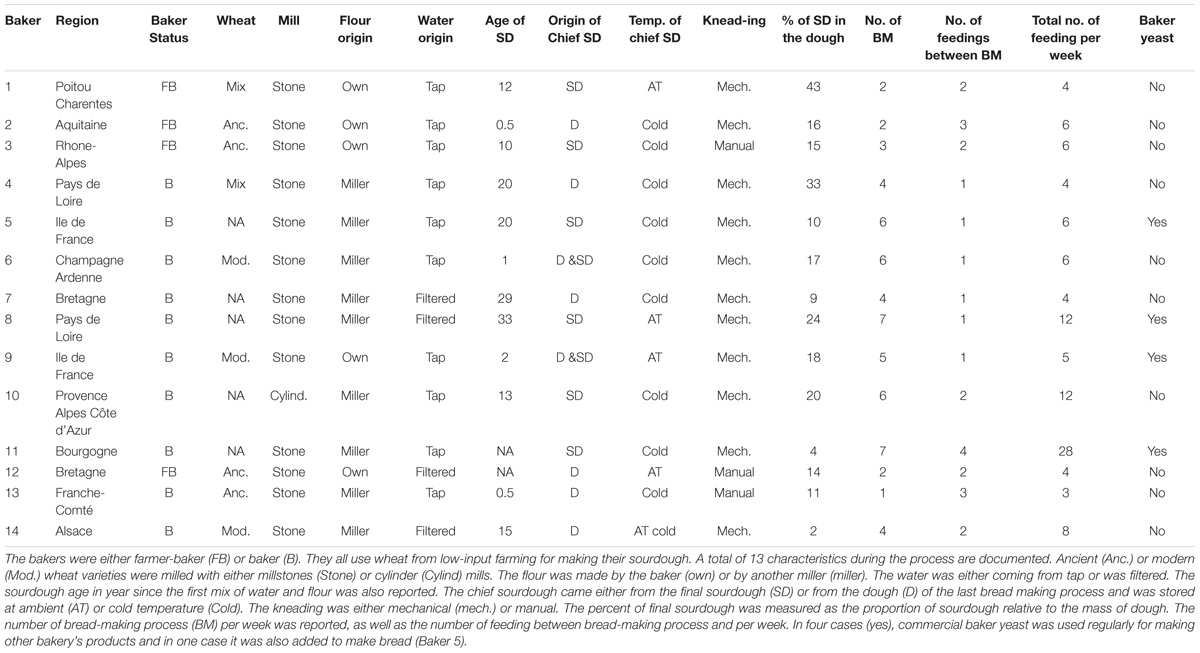
All bakers produce bread in a low-input agro-food ecosystem. They explained their bread-making practices during diverse interviews. The output of these interviews is presented in the results section and provides details on the specific practices of the bakers we studied. Broadly, each bread making process starts from the chief sourdough, which comes either from a piece of dough or a piece of final sourdough from the previous bread-making process. Water and flour are added to the chief sourdough to constitute the final sourdough. This is called feeding or back-slopping. The flour is obtained by milling ancient and/or modern wheat varieties using cylinders or stone mills. Once the final sourdough is reactivated, kneading can start by mixing the final sourdough with flour, water, and salt. In some cases, the baker also adds commercial baker’s yeast. After kneading, the first fermentation lasts for a variable amount of time, depending on bakers. The dough is then divided and shaped. A second fermentation occurs before cooking. Bread is cooked in wood-fired or electric ovens. From one bread making process to the next, the chief sourdough is stored either at room temperature or in the refrigerator. During this process, thirteen well-documented characteristics were retained from the interview to reflect the diversity of bakery practices (Table 1).
In each bakery, the final sourdough of one bread-making run was sampled twice. One sample was taken from the surface of the sourdough and the other from the center. All samples were kept at 4°C for 3–7 or sent by post-mail at ambient temperature and then kept cold before the culture-based analysis was initiated. Yeasts were first isolated and identified by the culture-based technique, while part of the sourdough was kept at -20°C for the non-culture-based analysis.
Culture-Based Method of Yeast Species Identification
Yeast enumeration, isolation and DNA extraction were performed according to Lhomme et al. (2016).
Two pieces of sourdough (one was taken from the surface of the sourdough and the other from the center) were diluted independently and sowed on different petri dishes. From 39 to 40 isolates per sourdough were then cloned, stored and further analyzed, except for sourdough 14, from which only 12 isolates were characterized. Each cloned isolate represents a strain. The strains were isolated by randomly picking colonies on different petri dishes.
With 40 strains analyzed, the probability of detection of one species representing 10% of the sourdough species is over 98% but falls at 65% when only 10 strains are analyzed.
For DNA extraction, strains were grown at 30°C in 1 ml of YE (1% YE, 2% glucose) shaken at 200 rpm. After digestion of the pellet for 1 h at 37°C using 10 units of zymolyase (Euromedex, Souffelweyersheim, France) in 0.5 mL of sorbitol 1 M, Na2EDTA 0.1 M, pH 7.5 buffer, a standard method of DNA extraction was used (Lhomme et al., 2016). All the 519 yeast isolates were identified on the basis of their pattern of AluI digestion of the amplified rDNA Non-Transcribed Spacer 2 (NTS2) (Nguyen and Gaillardin, 1997). Species identification was confirmed by sequencing the D1/D2 rDNA region (Kurtzman and Robnett, 2003) for 60 strains representatives of the different AluI profiles and by sequencing the actin coding gene, ACT1 for 46 strains (Daniel and Meyer, 2003). All the oligonucleotide PCR primers used were obtained from Invitrogen (Invitrogen, Cergy Pontoise, France) and PCR products were Sanger sequenced on both strands by Eurofins (Supplementary Table S1).
DNA sequence alignments were carried out using the pairwise alignment method for the forward and reverse complement sequences of each strain in order to obtain a consensus sequence. Identification queries were fulfilled by a BLAST search of the National Center for Biotechnology Information database (NCBI, Bethesda, United States) and the YeastIP databases1. A similarity of more than 99% to gene sequences of type strains was used as the criterion for species identification.
Non-culture Based Method of Fungal Identification
Synthetic Sourdough
As control for pyrosequencing analysis, we created a synthetic sourdough. One representative strain of nine species (Candida carpophila, Hyphopicchia pseudoburtonii, Kazachstania humilis, Kazachstania bulderi, Kazachstania saulgeensis, Kazachstania unispora, Rhodotorula mucilaginosa, Saccharomyces cerevisiae, Torulaspora delbrueckii) previously isolated from French organic sourdoughs (Jacques et al., 2016; Lhomme et al., 2016) was included in the synthetic sourdough (Supplementary Table S2). After 12 h in 5 mL of YE media at 30°C, cell counts for each strain were determined using a Coulter counter (Beckman Coulter, Brea, CA United States). All strains were mixed at the same density (107 UFC/g of sourdough) together with organic wheat flour (0.76 g, organic T110 wheat flour from Biocer) and distilled water (1 g). This synthetic sourdough, was then stored at -20°C for 10 days to mimic the storage process of bakeries sourdough samples. No yeast growth phase in the sourdough was included before storage to avoid adding a factor leading to inequality between yeast density.
Total DNA Extraction
The DNA extraction was carried out on a sample, taken independently from the ones used for the culture-dependent analysis. For each sourdough, 200 mg of sourdough was used to extract DNA with the PowerSoil DNA Isolation Kit (MO BIO laboratories, Inc., Carlsbad, CA, United States)).
To improve the yield, the following changes were made to the manufacturer’s extraction protocol: after adding C1 solution to an aliquot of sourdough in the bead tubes, the tubes were incubated at 65°C for 10 min and then horizontal mixed at 30 Hz for 5 min. The tubes were then put on ice before continuing with the kit protocol. The final elution step with C6 solution was carried out twice with the addition of 40 μL (total 80 μL).
Amplicon Preparation for Pyrosequencing
Choice of primer pairs
Pyrosequencing was carried out on the ITS1 rDNA region. Among 29 ITS1 primers published in the literature (Supplementary Table S3), we selected the ITS primers that could specifically amplify fungal DNA but not plant DNA. To do so, we first built an ITS sequences database by blasting the S288C Saccharomyces cerevisiae ITS1-5.8S-ITS2 sequence against the NCBI non-redundant nucleotide database, including any sequences with both an e-value < 10-4 and a > 80% identity. We also added Poaceae ITS sequences. Overall, there were 1995 sequences in this database. Second, we blasted 29 primers from the literature (Supplementary Table S3) on this database (parameters: e-value < 10-2; > 80% identity). From this analysis, six primers theoretically allowed amplification of fungal but not Poaceae ITS regions: forward primers NSI1 (Gardes and Bruns, 1993) and NSA3 (Martin and Rygiewicz, 2005), and reverse primers ITS4 and NLB4 (Gardes and Bruns, 1993), 58A2R and NLC2 (Martin and Rygiewicz, 2005). Two of these hypothetical fungal-specific primer pairs (NSA3-NLC2 and NSA3-58A2R) were tested on DNA extracted from 17 natural sourdough yeast strains representing the ten species previously identified in natural French organic sourdoughs (Jacques et al., 2016; Lhomme et al., 2016) and from three plant species: wheat, maize and nigella (Supplementary Table S2). The primers pair, NSA3 (5′–AAACTCTGTCGTGCTGGGGATA–3′) and 58A2R (5′–CTGCGTTCTTCATCGAT–3′) (Martin and Rygiewicz, 2005) gave fungal specific amplification and was selected for ITS pyrosequencing.
Amplicon preparation
Amplicons for pyrosequencing were prepared in two steps: first, ITS1 amplification and second, ITS1 amplification with barcode addition. ITS1 region was amplified from 100 to 200 ng of total sourdough DNA incubated in a 50 μL of the following mix reaction: 0.2 mM of each dNTP, 1 mM of MgCl2, 15 μM of each primer, 2.6 unit of Taq Polymerase (Roche Expand High Fidelity, Roche, Mannheim, Germany), 1X Roche Expand High Fidelity buffer. Two consecutive amplification regimes were performed for the first ITS1 amplification step. After an initial denaturation at 95°C for 2 min, 10 cycles of amplification were conducted as follows: denaturation at 95°C for 15 s, annealing at 60°C for 30 s, and extension at 72 °C for 45 s. The final extension was at 72°C for 45 s. A second 10 cycles of amplification was performed: denaturation at 95°C for 15 s, annealing at 60°C for 30 s, and extension at 72°C for 45 s with addition of 5 s extension time at each subsequent cycle. The final extension was at 72°C for 7 min.
Addition of specific barcodes was performed in 50 μL reactions containing 2 μL of the previous PCR product, 0.2 mM of each dNTP, 0.5 mM of MgCl2, 0.3 μM of each specific primer containing the specific index and the Eurofins MIDs (Supplementary Table S4), 2.6U de Taq Expand High Fidelity of Roche and 1X Buffer Expand High Fidelity. After an initial denaturation at 95°C for 2 min, 5 cycles of amplification were conducted as follows: denaturation at 95°C for 15 s, annealing at 60°C for 30 s, and extension at 72°C for 45 s. The final extension was at 72°C for 7min.
Each sample was column purified with the MinElute PCR Purification Kit (Qiagen, 28006), followed by concentration with a speed vacuum (Savant SPD IIIV Speed Vac Concentrator, Thermo Electron Corporation) for 20 min to obtain 60 ng/μL as measured by NanoDrop (NanoDrop 2000 UV-Vis Spectrophotometer, Thermo Fisher Scientific). Finally, amplicons were sent to Eurofins2 for the titanium FLX pyrosequencing (Roche technology, compared with other new-generation sequencing systems in Liu et al., 2012).
Metabarcoding Data Analysis
The metabarcoding data were processed with the QIIME (Quantitative Insights Into Microbial Ecology) v 1.8.0 (Caporaso et al., 2010) pipeline.
Database
UNITE fungal ITS database version 7 (17,421 sequences; downloaded at http://unite.ut.eerelease_01.08.2015, Kõljalg et al., 2013) was completed with 13 ITS1 Sanger sequences obtained from five natural sourdough strains and eight type strain sequences (Supplementary Table S5) corresponding to 11 previously described bread sourdough species. The additional sequences were non-redundant with the downloaded UNITE database sequences. The UNITE database completed with the sourdough strains contains 432 Saccharomycetes reference sequences. Hereafter in this article, we term the downloaded UNITE database “UNITE only” and the UNITE database completed with the sourdough strain sequences “UNITE completed.”
Filtering, quality trimming and clustering into OTU
To conserve high quality sequences only, we trimmed and filtered the reads. For filtering, raw pyrosequencing fna and quality files were transformed to fastq files. Reads were truncated with the PRINSEQ tool at the 3′ position at the quality threshold of 34, and any read <127 bp was discarded. The following steps were performed using QIIME: chimera detection using usearch61 against the uchime_reference dataset3; operational taxonomic unit (OTU) selection using uclust (Edgar, 2010) with the open reference OTU-picking workflow and a threshold of 99% pairwise identity; and taxonomy assignment using the RDP Classifier 2.2 (Wang et al., 2007) and using the RDP Classifier 2.2 (Wang et al., 2007) and BLAST against the “UNITE completed” database.
Assignment was performed on the most representative sequence of each OTU according to the “UNITE completed” database (17,434 sequences) and cut in the SSU region at the sequence CGTAAGGT (Supplementary Table S5) so as not to bias assignment in favor of the SSU region, which is well conserved between species.
Data Analysis
The datasets are available under the URL http://doi.org/10.5281/zenodo.1472295, under a creative commons attribution.
The results regarding Sourdough 14 gave peculiar data (see section Identification of Yeast Species in Sourdough by the Culture-Based Method) and was thus excluded from most of the analysis described below. Indeed, few microbial isolates could be retrieved from the sourdough. In addition, metabarcoding data revealed only two reads assigned to saccharomycete species over 8444, while this class is supposed to contain the dominant sourdough yeast flora.
Analysis of Bread-Making Practices
To describe the diversity of bakery practices, we performed a multiple correspondence analysis (MCA) on 13 bread-making process characteristics described for each baker (Table 1). Bakers were either farmer-bakers (producing their own wheat grains) or bakers (only making bread). This “status” variable was not used to build the MCA axis and was only used as a Supplementary variable. To avoid classes with only one observation, we recoded variables as follows: conservation at “cold & room temperature” were included in the “cold” group; the modalities “dough + SD” and “dough” of the sourdough origin variable were grouped together in the “dough” modality; the total number of feedings per week was recoded as “frequent” if it exceeded 6 or “rare” otherwise; the number of bread makings per week was recoded as “frequent” (>3) or “rare”; the fraction of sourdough in the dough was recoded as “low” (<20%) or “high”; the sourdough age was recoded as “recent” (2 years or under), “average” (between 2 and 20 years), or “old” (>20 years); and the number of feedings between bread makings was recoded as “1,” “2,” and “3+4”. For each variable, missing values were treated as a modality and coded “UN”. Then, using the first two axes of the MCA, we clustered the bakeries into groups using a k-means algorithm and used this “practice type” variable in the following sections.
Analysis of Microbial Species Diversity
The microbial species richness was characterized over all sourdough and fungal orders. The other alpha diversity as well as beta diversity indices were carried out only on Saccharomycetales species, as they may be involved in fermentation and they were the only species found using the culture-based method.
Diversity indices were only calculated on the metabarcoding data. Indeed, only 40 strains were analyzed by culture-based method which may bias estimates. For each sourdough, we computed the species richness as well as the Shannon and Simpson alpha diversity indices. The Shannon and Simpson indices value were also converted to effective number of species (Jost, 2006). Between-sample community dissimilarity was quantified by computing weighted UniFrac distances (Lozupone and Knight, 2005) using the Gunifrac R-package on the rarefied OTU abundance matrix. We used the OTU phylogenetic tree constructed with Candida boleticola species as the outgroup.
Based on the UniFrac distance matrix, the sourdoughs were clustered into groups using hierarchical clustering. The link between practices and community composition was studied by performing non-parametric analysis of variance on the Gunifrac distance matrix as well as exact Fisher tests between each practice variable and the group of sourdoughs clustered according to the UniFrac distance. To account for multiple testing, we adjusted the p-value with the false discovery rate (FDR) method (Benjamini and Hochberg, 1995).
Analysis of Sourdough Properties and Their Link With Bread-Making Practices
To study the relation between the microbial population densities and the biochemical properties of the sourdough, we performed a principal component analysis (PCA) on the average values of the yeast and LAB population densities (on a log10 scale), the pH, the lactic acid and acetic acid contents, the TTA (Lhomme et al., 2015b). Sourdough 14 was not used to build the PCA axes but used as a supplementary individual. To investigate the relationship between bread making practices, the community composition, and the population densities and biochemical properties of the sourdough, we used the bread-making practices groups built from the MCA and the community group as supplementary variables.
Results
Characterization of Bakery Practices
The MCA performed on the bakery practices is represented in Figure 1. The first axis (explaining 25% of the variance) opposed intensive practices (frequent bread-makings, frequent sourdough back-slopping, use of commercial yeast in the bakery, flour originating from a miller, mechanical kneading, unknown wheat cultivar) against extensive practices (manual kneading, use of own flour, rare bread-making processes and sourdough back-sloppings, use of ancient wheat populations) (Figure 1A). The second axis of the MCA was mostly explained by the characteristics of the sourdough (origin, age, percent in the dough) as well as the mill type (Figure 1B).
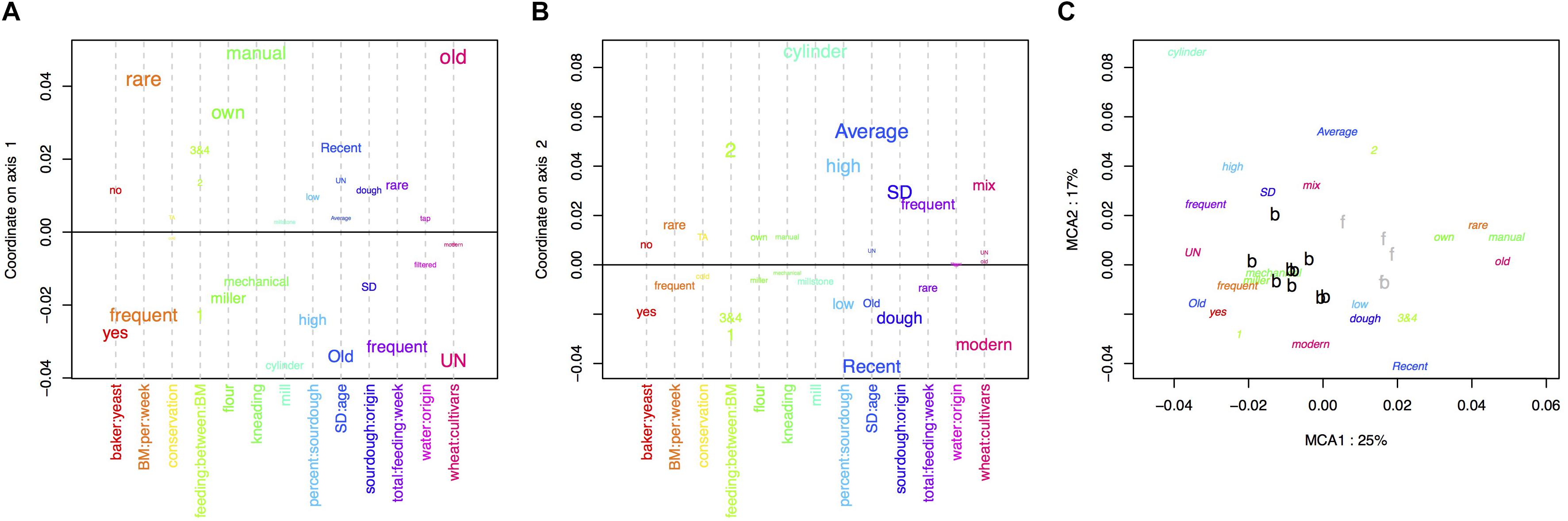
Figure 1. Multiple correspondence analysis (MCA) on the 13 characteristics of the bread-making process described for each baker (A) variables explaining the first axis, (B) variables explaining the second axis, (C) distribution of baker practices on the first two axis.
The clustering into two groups separated bakers having intensive practices from bakers having extensive practices (Figure 1C). All bakers but one were in the “intensive practices” group, and all farmer-bakers plus one baker were in the “extensive practices” group. Hereafter, we named these two groups “baker-like” and “farmer-baker-like” and encoded them “B” and “F,” respectively. We can note that this clustering was largely influenced by the number of bread making processes per week and match perfectly this variable. Increasing the number of groups to three increased the fraction of between-group variance from 50 to 73%. However, this would result in a group with only two bakers, which would make statistical analysis and interpretation difficult. For this reason, we decided to discard further interpretation of the second axis of the MCA and to keep the first axis clustering into two groups in the following analyses.
Identification of Yeast Species in Sourdough by the Culture-Based Method
AluI digestions of the NTS2 region were obtained on 519 yeast strains isolated from 13 natural organic sourdoughs. Eight yeast species were identified corresponding to seven AluI digestion patterns (Table 2). One of them was recently discovered: Kazachstania saulgeensis was found in Baker 1’s sourdough and was called by the name of its town of origin, Saulgé (Jacques et al., 2016; Sarilar et al., 2017). Another profile corresponds to a yet undescribed Kazachstania species, found in Baker 2’s sourdough. The ITS sequence of this undescribed species, represented by strain B2_TP1_4 (sequence Accession Number KX458458.1), had 100% identity with the sequence of strain IMBRR1 (sequence Accession Number KC118125.1), isolated from Bulgarian boza, a cereal-based fermented beverage. Based on the rRNA sequence comparison, the most closely related Kazachstania species to strain Kazachstania sp. B2_TP1_4 is Kazachstania exigua. They share 96% identity in the ITS sequence and 99% identity in the 26S D1/D2 LSU rDNA sequence (large subunit rRNA gene) between the K. exigua neotype strain CBS379NT and the strain B2_TP1_4. Based on the RPB2 coding gene sequence comparison, the most closely related Kazachstania species to strain Kazachstania sp. B2_TP1_4 (Sequence Accession Number LT158256.1) is Kazachstania australis CBS 194T (Sequence Accession Number LT158259.1), with 93% identity followed by K. exigua CBS 379NT (Sequence Accession Number LT158276.1) with 92% identity.
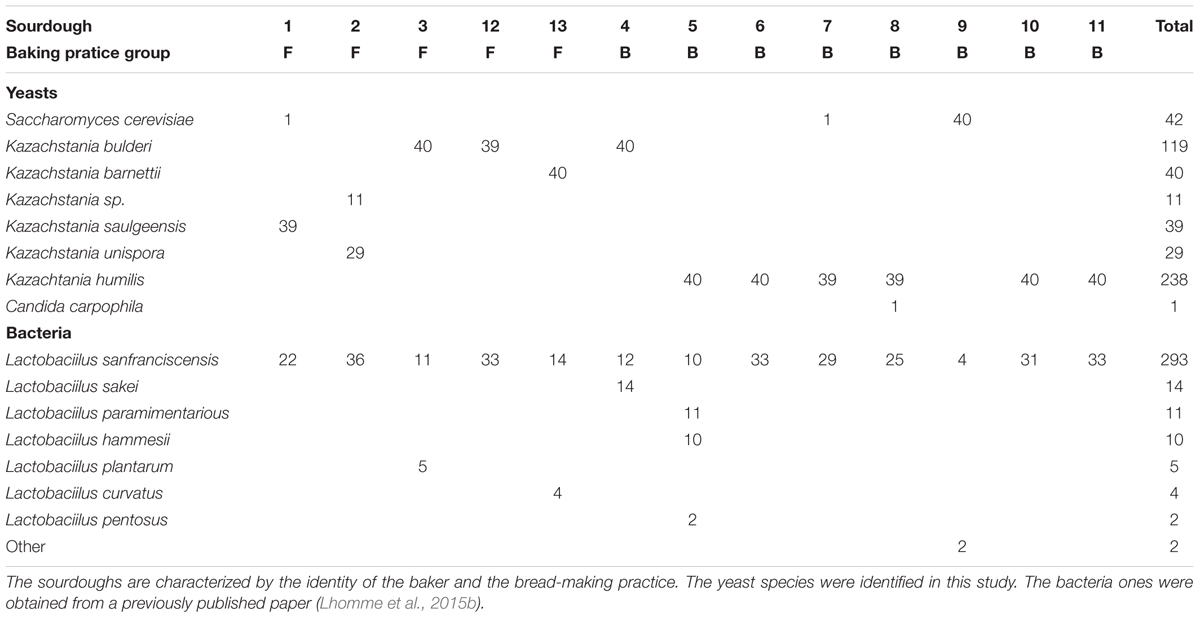
Table 2. Yeasts and bacterial species counts in 13 natural sourdoughs as studied by the culture-based method.
Among the 519 strains (Table 2), 45.9% belonged to K. humilis and 22.9% belonged to K. bulderi. The remaining were S. cerevisiae (8.1%), K. barnettii (7.7%), K. saulgeensis (7.5%), K. unispora (5.6%), Kazachstania sp. (2.1%), and Candida carpophila (0.2%). No yeast species was shared by all 14 sourdoughs. K. humilis was the most widely distributed species. It was detected as a unique species in four sourdoughs and as the dominant species together with other yeast species in two sourdoughs. Kazachstania bulderi was found as a unique species in three sourdoughs. S. cerevisiae was identified as a unique species in one sourdough and as a minor species (1 isolate) in sourdoughs 1 and 7. The new species K. saulgeensis was dominant in sourdough 1, while the Kazachstania sp. strains described above (represented by B2_TP1_4) were found together with Kazachstania unispora strains in sourdough 2. It is important to note that, overall, only 42 (from sourdoughs 1, 7, and 9) out of 519 isolates belonged to the S. cerevisiae species that is usually used in commercial starters as a leavening agent in bakery products.
Sourdough Fungal Composition Described by Non-culture-Based Analysis
Synthetic Sourdough
A synthetic sourdough (sourdough 15), made of flour and nine yeast species previously found to be present in French sourdoughs, was analyzed as a control for our method of pyrosequencing. The eight Saccharomycetes species (Candida carpophila, Hyphopicchia pseudoburtonii, Kazachstania humilis, Kazachstania bulderi, Kazachstania saulgeensis, Kazachstania unispora, Saccharomyces cerevisiae, Torulaspora delbrueckii) as well as the single Basidiomycota species (Rhodotorula mucilaginosa) mixed in equal density in the synthetic sourdough were all detected but in different proportions, from 6 reads for C. carpophila to 296 reads for K. humilis. In addition, the synthetic sourdough contained other yeast species, which probably came from the flour. Overall, it contained 40.31% Dothideomycetes class, mainly represented by the Capnodiales order (31.8% of the total synthetic sourdough reads) and 14.59% Sordariomycetes class, including the Hypocreales and Xylariales orders (7.51 and 7.01% of the total synthetic sourdough reads, respectively). The Basidiomycota phylum represented 18.37% of the reads. Surprisingly, the Saccharomycetales order represented only 9.85% of the reads (972 reads), suggesting that the Saccharomycetales were not dominant in the flour and not well extracted from the yeasts added to the synthetic sourdough.
Natural Sourdoughs
A total of 689 OTUs corresponding to 202,225 reads were selected for analysis because they were represented by more than five reads after chimera detection. Each sourdough sample contained between 104 (Sourdough 10) and 351 (Sourdough 7) OTUs (Supplementary Table S6).
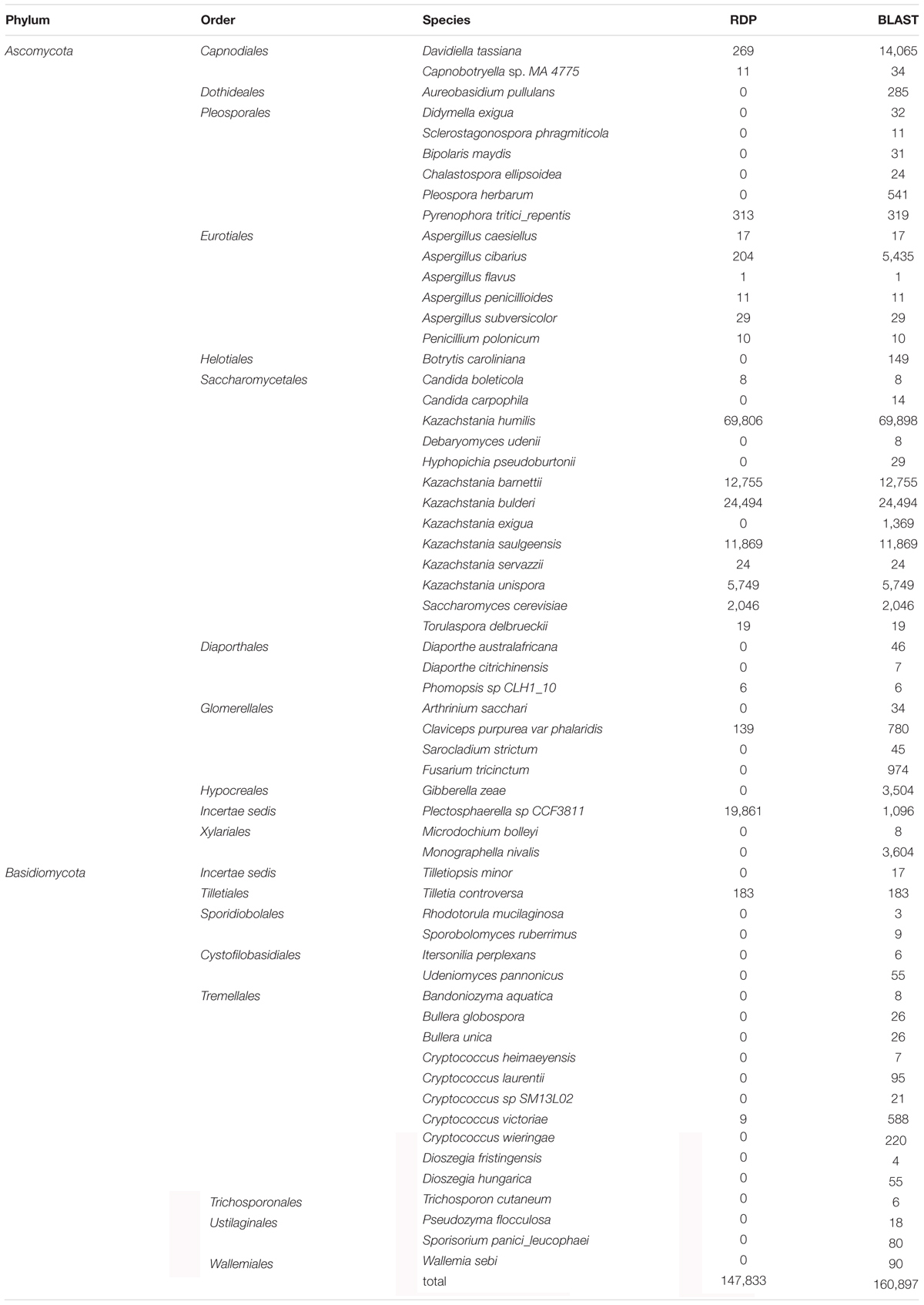
The assignment was done using either RDP or BLAST after completing the UNITE database with ITS sequences obtained from sourdough strains (Table 3). Database completion improved the rate of OTU assignment at the species level (data not shown). Most reads were assigned to Ascomycota (99.0% with RDP, 94.2% with BLAST) while Basidiomycota (0.2% with RDP, 1% with BLAST) and Anthophyta (0.1% with RDP, none with BLAST) were much less represented. A total of 4.7% (with BLAST) and 0.7% (with RDP) of the reads remained unidentified at the phylum level. Overall, at the species level, BLAST was able to assign more OTUs then RDP (Table 3). It allowed the detection of 32 species that RDP did not. For this reason, only BLAST assignments are presented below.
Over all sourdoughs, the Saccharomycetes class represented 66.7% (128,282) of the reads (Supplementary Table S6). Dothideomycetes was the second most represented class, with 18.1% of the reads. The other classes were represented by less than 5% of the reads. With the exception of Sourdough 14, the Saccharomycetales order was the most common in each natural sourdough, representing 26% (1,890 reads out of 7212 reads in Sourdough 9) to 99% (13,569 reads out of 13723 reads in Sourdough 10) of the reads (Figure 2). Other orders were also found : the Capnodiales order represented from 3% (43 reads, Sourdough 10) to 48% (3,481 reads, Sourdough 9) of reads in each sourdough; Pleosporales from 0.01% (35 reads, Sourdough 10) to 0.7% (130 reads in Sourdough 7); Eurotiales from 0% (0 reads, Sourdough 1 and 13) to less than 0.6% in the other sourdoughs (less than 100 reads per sourdough); Hypocreales from 0.1% (14 reads, Sourdough 10) to 8% (889 reads, Sourdough 2); and Xylariales from less than 0.1% (2 reads, sourdough 10) to 11% (1,929 reads, Sourdough 7) (Figure 2).
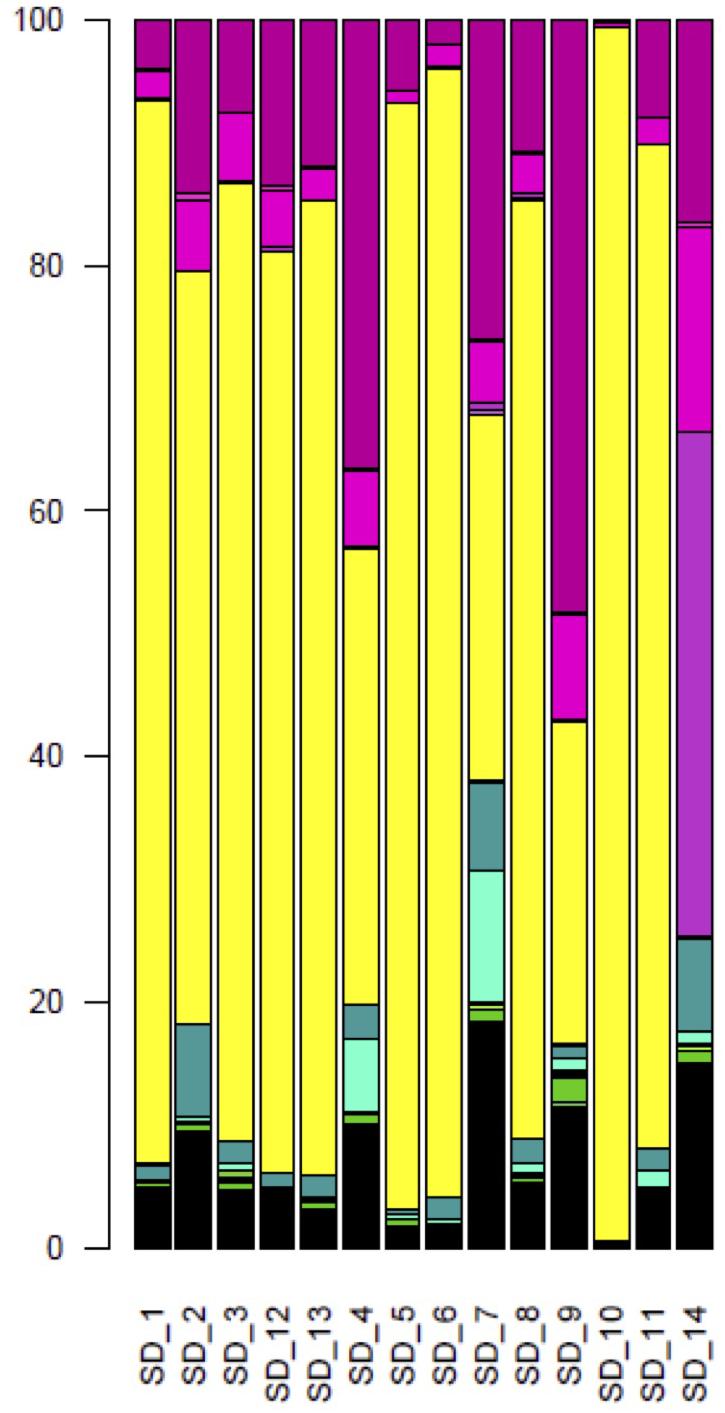
Figure 2. Composition of fungal orders, after BLAST assignments, for each natural sourdough. In yellow, Saccharomycetales; in medium violet red, Capnodiales; in hot magenta, Pleosporales; in fuchsia, Eurotiales; in Persian green, Hypocreales; in bright turquoise, Xylariales; in free speech green, Tremellales; in razzle dazzle rose, Dothideales; in apple, Tilletiales; in green yellow, Filobasidiales; in orchid, Helotiales; in La Palma, Sporidiobolales; in lime, Wallemiales; in fern, Ustilaginales; in turquoise blue, Diaporthales; in Kelly green, Cystofilobasidiales; in Pacific blue, Glomerellales; in myrtle, Trichosporonales; in bright green, Exobasidiales.
At the species level, a total of 160,897 reads (79.6% of the total number of reads) were assigned to 60 species (Supplementary Table S7). Overall, only 20.3% of the reads were assigned to non-Saccharomycetales species. Five non-Saccharomycetales species, known plant pathogens, were identified in all sourdoughs: Davidiella tassiana (Capnodiales order), Claviceps purpurea (Hypocreales), Fusarium poae (Hypocreales), Monographella nivalis (Xylariales) and Cryptococcus victoriae (Filobasidiales). However, these were identified at less than 1% of each sample in Sourdoughs 1 to 13. By contrast, most of the reads (79.7%) were assigned to the Saccharomycetales species, with 13 species detected. Over these 13 species, eight were also detected by culture-based method, including seven dominant species (K. humilis, K. bulderi, K. barnettii, Kazachstania sp., K. saulgeensis, K. unispora, S. cerevisiae) and one rare species (Candida carpophila). The K. humilis species was the most frequent species and was found in 12 out of 14 sourdoughs. In five sourdoughs, this species was dominant and was represented by more than 50% of the sourdough reads. In Sourdough 7, this species was represented by 30% of the reads, but this sourdough also had the highest number of unassigned reads. Finally, in six sourdoughs, this species was represented by less than 1% of the reads. The species K. bulderi was also recurrently found at high frequency in three sourdoughs (from 38 to 77.8% of the sourdough reads) and was detected at low frequency (below 2%) in five sourdoughs. Several other Kazachstania species were detected as dominant species in a single sourdough. This was the case for K. barnettii (79.1% of the reads in sourdough 13), K. saulgeensis (86.3% of the reads in sourdough 1), and K. unispora (48.8% of the reads in sourdough 2). The species K. servazzii, previously found in French sourdough as dominant species (Lhomme et al., 2015a), was detected twice but only at a frequency below 0.2%. Interestingly, the species S. cerevisiae was detected as the dominant species only once (25.6% of the reads in sourdough 9) although it was found at a frequency below 1% in nine other sourdoughs.
Pattern of Fungal Species Diversity Among Sourdoughs
The within-sourdough yeast species diversity (alpha diversity) (Table 4) and the between-sourdough yeast species diversity (beta-diversity) was analyzed using metabarcoding data only since it allows a deeper analysis of the fungal community. Only the reads identified as Saccharomycetales were retained (Supplementary Table S7), as these species may be involved in fermentation. Sourdough 14 was discarded from this analysis, as only 2 reads were identified as Saccharomycetales in this sourdough.
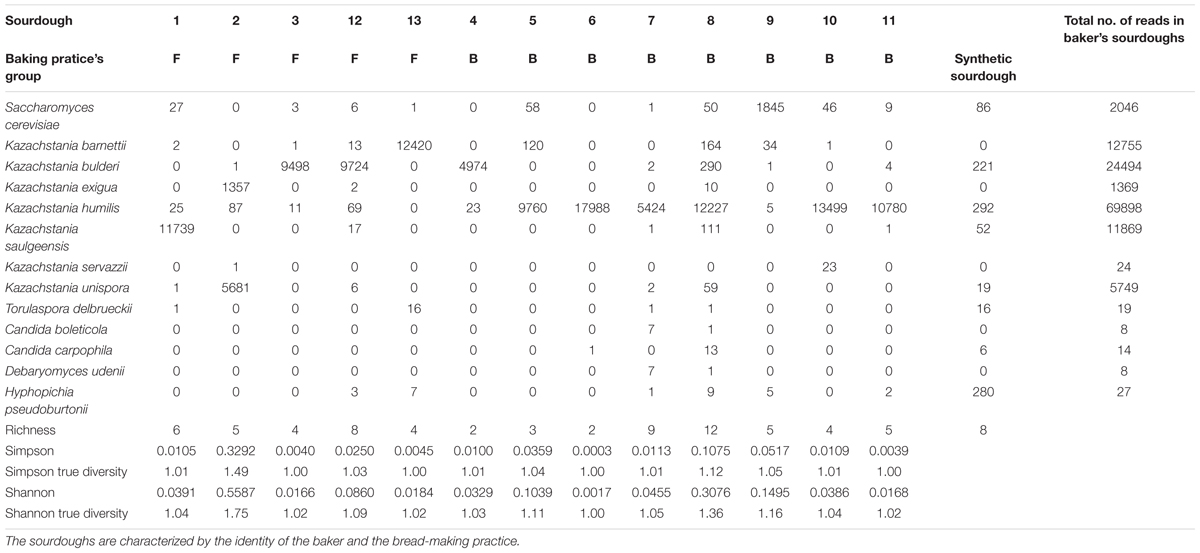
Table 4. Number of reads assigned to each species of the Saccharomycetale order and alpha diversity in 13 natural sourdoughs as studied by pyrosequencing.
We first analyzed the alpha diversity (Tables 2, 4). The number of Saccharomycetales species ranged from 2 to 12 within sourdoughs. All sourdoughs but one (Sourdough 2) had a single dominant species (Tables 2, 4). The Simpson and Shannon indices were low (below 0.06 and 0.15, respectively) and the effective number of species was close to one for all sourdoughs except two (Table 4). These two sourdoughs had a higher diversity for different reasons. One (Sourdough 8) had a dominant species with a frequency of 0.94 and 11 other rare species (Table 4). The second (Sourdough 2) had two species with a frequency of 0.19 and 0.8 as well as three additional rare species (Table 4). The effective number of species estimated from the Simpson and Shannon indices were of 1,12 and 1,36 respectively for sourdough 8 and 1,49 and 1,75 for sourdough 2. There was no evidence of difference in alpha diversity between sourdoughs of the two groups of bakery practices (Table 4).
Given that all sourdoughs but one were dominated by only one species, the weighted UniFrac distances between sourdoughs were strongly related to the phylogenetic distances between dominating species (Figure 3). The 13 sourdough communities were clustered into five groups: the five sourdoughs dominated by K. humilis, the three sourdoughs dominated by K. bulderi, the two sourdoughs dominated by K. barnettii or K. saulgeensis and the two groups represented by a sourdough dominated by either K. unispora or S. cerevisiae. These groups did not cluster sourdoughs according to their geographical origin.
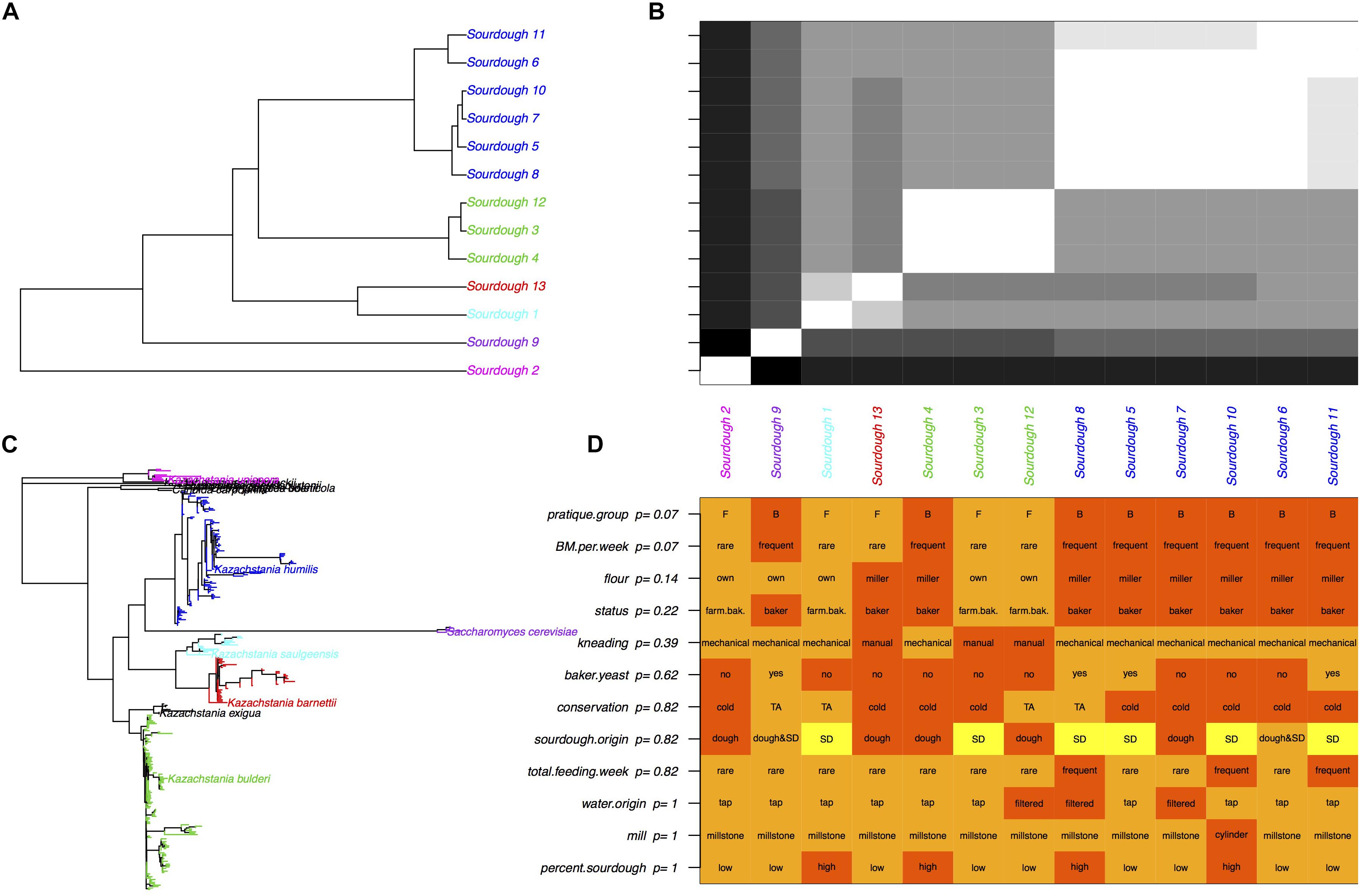
Figure 3. Yeast species diversity among sourdoughs and its link to bakery practices. (A) Tree of sourdough yeast communities based on the UniFrac distances. Colors correspond to the “yeasts’ community” group variable derived from a hierarchical clustering, (B) UniFrac distances between sourdoughs. White corresponds to close communities and black to the most distant communities. (C) Phylogenetic tree based on the OTU sequences. Colors correspond to the dominate species of each sourdough yeast community presented in (A,B). (D) Baker practices and adjusted p-values obtained with an exact Fisher test between the practice and the “community” group. FDR correction was used to account for multiple testing.
Overall, we did not found a significant correlation between the UniFrac distances and the distances between bakers’ practices estimated by their difference in axis 1 coordinates on the MCA (Mantel test p = 0.19). Due to the small sample size and the strong dependence between practices variables, we could not select one or several specific practices variables that would explain the yeasts diversity of sourdough communities (Fisher exact tests, Figure 3). Instead, we examined the link between the sourdough community groups (five groups described above and Figure 3) and the practice group (bakers-like versus farmer-bakers like practices). The UniFrac distance differed between practice groups (p = 0.007) and these groups were not independent from the sourdough community groups (Fisher exact test, p = 0.01). A majority of sourdoughs collected from sourdough made with bakers-like practice (6 out of 8) were dominated by K. humilis, whereas none of the sourdoughs sampled from farmer-bakers practices’ bakeries were (Fisher exact text, p = 0.07). However, all sourdoughs but one (Sourdough 13) had a few reads identified as K. humilis. A part from the strong presence of K. humilis in sourdoughs from bakers but not from farmer-bakers, we were not able to distinguish species patterns related to specific practices.
Finally, we carried out a PCA to visualize the link between sourdough acidity (pH, TTA, lactic acid, acetic acid), yeast and bacteria density, bakery practice group and the five UniFrac groups of sourdoughs (Figure 4). The physico-chemical characteristics of these sourdoughs were previously reported in Lhomme et al. (2015b). The first two components explained 58% of the variation. The first axis was mostly explained by the variation in microbial density, pH and TTA, while the second axis reflected variation in lactic and acetic acid (Figure 4A). There was no clustering according to the bakery practice group (Figure 4B) or the yeast species composition of the sourdough (Figure 4C).
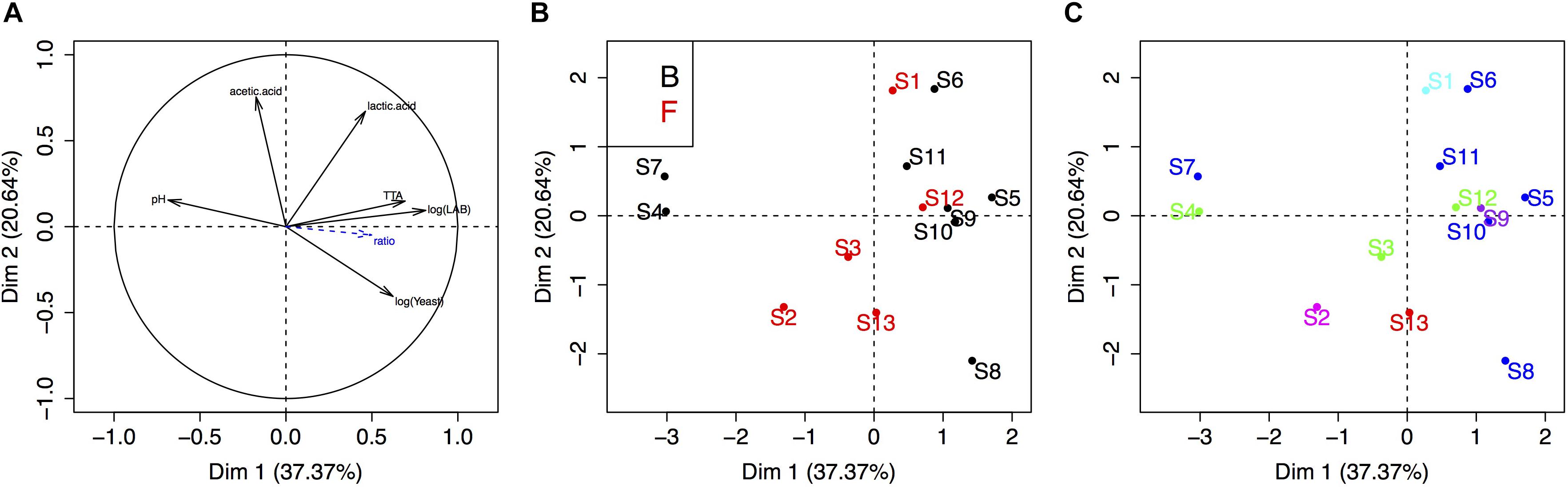
Figure 4. Principal component analysis of the sourdough’s biochemical and microbial properties. (A) Correlation circle of sourdough properties (acidity, population densities, etc.). (B,C) First two axis of the PCA analysis showing the distribution of sourdoughs according to the “practice” group (B, Bakers and F, farmer-baker) or the “yeast community” group (dark blue, sourdough dominated by K. humilis; green, by K. bulderi; red, by K. barnettii; purple, by S. cerevisiae; light blue, by K. saulgeensis; pink, by K. unispora and a yet undescribed Kazachstania species, see Figure 3).
Pattern of Yeast and Bacterial Species Diversity
The bacterial species diversity of these 14 sourdoughs have been analyzed previously (Lhomme et al., 2015b). The published data on the culture-based analysis of bacteria are presented Table 2. All sourdoughs contained Lactobacillus sanfranciscensis. This species was dominant (Freq > 0.5) in all sourdoughs but two when analyzed by culture-based method and dominant in all sourdoughs when analyzed by 16S metabarcoding (Lhomme et al., 2015b). The lack of diversity for the dominant LAB species limited any analysis of yeast-bacteria co-occurrence in this study. It also limits the testing of any link between metabolic properties and LAB species content. Four sourdoughs had more than one bacterial species, as detected by culture-based method. These sourdoughs (B3, B4, B5, B9) did not appear to have specific metabolic and microbial density properties. Indeed, they were not distributed on specific positions on the PCA compared to the ones where only L. sanfranciscensis was isolated (Figure 4).
Discussion
In this work, we analyzed the yeast diversity in a collection of 14 sourdoughs collected from bakers with different bread-making practices. We developed a metabarcoding method for analyzing fungal communities in bread sourdough. From our knowledge only two studies have used so far metabarcoding to characterize fungi in sourdoughs made of cereals (Minervini et al., 2015; Raimondi et al., 2017, see review in Weckx et al., 2018). They used regions of the LSU rDNA as barcode. We used ITS1 that has been shown to better discriminate closely-related yeast species (Vu et al., 2016). We tested our method on a synthetic sourdough composed of flour and known sourdough yeast species in equal proportion. After data analysis of this synthetic sourdough, all the added yeast species were recovered showing that ITS1 barcode allows the discrimination of these yeast species. However, the species were detected in different proportions, revealing the limit of quantitative analysis using the pyrosequencing approach. The added species were found in low frequencies (9.4% of the total sourdough’s reads). The low representation of Saccharomycetales in the synthetic sourdough could be related to a DNA extraction, PCR or amplicon sequencing problem. The high proportion of non-Saccharomycetales reads also revealed the natural flour fungal composition. When analyzing the fungal distribution in natural sourdoughs, the Saccharomycetales were dominant compared to the non–Saccharomycetales species. In natural sourdough, where fermentation has occurred, the density of non-fermentative fungi, such as wheat pathogens, may be reduced by the production of metabolic inhibitors such as alcohol, acids and carbon dioxide during the fermentation process (Caplice and Fitzgerald, 1999).
By adding sourdough strain sequences to the public database, we improved the rate of OTU assignment at the species level. However, the “UNITE completed” database was not sufficient to assign all OTU reads at the species level. A total of 79.6% of the total number of reads were assigned to 60 species. The other reads were not assigned at species level. This might be related to a lack of power of ITS1 to discriminate species. It might also be related to a lack of reference sequences in databases. Different databases exist to assign fungal rDNA regions: INSD (international nucleotide sequence database), GenBank (Benson et al., 2006), ENA, DDBJ (Nakamura et al., 2013), SILVA (Pruesse et al., 2007), Ribosomal Database Project II (Cole et al., 2014), FUSARIUM-ID (Geiser et al., 2004), YEASTIP (Weiss et al., 2013) and UNITE databases. We chose the UNITE database because it is a commonly used and curated database for fungal barcoding. In addition, it contained non-translated parts of the rDNA region (ITS1 -5.8S- ITS2, Kõljalg et al., 2013). The presence of SSU parts in the reference database may alter assignments in favor of sequences containing SSU parts. However, the UNITE database still needs to be improved. For instance, taxonomical definitions need to be modified for “Candida” or “Incertae sedis”. More ITS1 sequences still need to be completed to contain sequences corresponding to all of the 100,000 described fungal species (Ainsworth, 2008). It would be particularly easy to include the 1100 known species of Saccharomycetales (432 in our completed version).
Culture-based and non-culture-based approaches gave similar results for the dominant species except for one species, the not-yet described Kazachstania species also discovered in boza from Bulgaria (Gouliamova et al., unpublished), and identified as K. exigua by the metabarcoding method. We did not add its ITS1 sequence to complete the UNITE database since the species was not described yet. The ITS1 sequence of this species and K. exigua are close, explaining the assignment to K. exigua. Strains of this species were previously identified as K. exigua by a previous publication that used a region of the LSU as barcode (Gouliamova et al., 2012). The K. exigua species has been described in many different sourdoughs (De Vuyst et al., 2016; Carbonetto et al., 2018). Based on the partial sequence of the large rDNA subunit (D1D2) and based on the ITS1 sequence, it clustered with this new species, as well as with the recently described species K. saulgeensis (Jacques et al., 2016; Carbonetto et al., 2018). Additional studies are needed to shed light on this closely related complex of yeast species.
As found previously, a single yeast species dominated the fungal community in each sourdough. As expected in fermented products (Lv et al., 2013; Illeghems et al., 2012; Tamang et al., 2016), Saccharomycetales species dominated in all the natural sourdoughs with the exception of Sourdoughs 9 and 14, which had a lower number of reads. Interestingly, the most commonly known bakery yeast species, S. cerevisiae, was not the major species, and no other species of the Saccharomyces sensus stricto clade was found. Only four of the participating bakers used commercial baker’s yeast. This might explain the overall low occurrence of S. cerevisiae. For three of them, S. cerevisiae was not isolated from their sourdough. These bakers used as chief sourdough, a piece of final sourdough of the previous bread-making process. This piece is collected before kneading, the step during which commercial yeast is usualy added. Therefore, our results suggest that commercial yeast does not invade the sourdough when used for other purposes in the bakeries. This finding should be confirmed on a higher number of samples or with dedicated experiments.
Most of the sourdoughs’ yeast species belonged to the Kazachstania clade. Overall, the Kazachstania species represented 91% of the 519 strains and 65.59% of the reads from natural sourdoughs. Seven Kazachstania species were detected (K. humilis, K. bulderi, K. barnettii, K. unispora, K. saulgeensis, K. servazzii and an as-yet undescribed Kazachstania sp), including a recently isolated species, K. saulgeensis (Jacques et al., 2016). The Kazachstania humilis and K. exigua species have been detected in sourdoughs in many different studies (De Vuyst et al., 2016; Carbonetto et al., 2018). The other ones are less commonly found.
Using the beta diversity index, we classified sourdoughs into five groups. The first group was characterized by the presence of K. humilis as the dominant species. The second group contained sourdoughs with K. bulderi as the dominant species. The other groups included the remaining species. The sourdoughs made by bakers harbored mostly K. humilis but also, in one case, S. cerevisiae or K. bulderi. The sourdoughs made by farmer-baker practices harbored a higher Kazachstania species diversity (K. bulderi, K. unispora, K. saulgeensis, K. barnettii, and the as-yet undescribed Kazachstania sp). These results highlight the importance of conserving diverse bakery practices to maintain yeast species diversity. However, additional sourdoughs should be analyzed to confirm this finding and understand if they are specific practices which drive the diversity and structure of these sourdough microbial communities.
No geographical structuration of the distribution of the dominant yeast species was found in our French sample. This could be due to a lack of statistical power given the small size of our sample. Numerous studies have been carried out on wheat sourdoughs’ microbial diversity in Europe, especially in Belgium and Italy. It was found that S. cerevisiae was the most frequent dominant yeast species, followed by Wickerhamomyces anomalus and K. humilis in Belgium and Italy, respectively (for a review, see De Vuyst et al., 2016; Carbonetto et al., 2018). In our French samples, we found that K. humilis was the most widespread dominant yeast species, followed by K. bulderi. We did not detect W. anomalus. Overall, the French sourdoughs studied here harbored higher species richness than Italian and Belgium sourdoughs. A previous study on 16 French sourdoughs collected in non-organic bakeries showed that all but one sourdough contained S. cerevisiae as the dominant species (Lhomme et al., 2015a). Our study reveals a much higher yeast species diversity. This result contrasted with the one on bacteria. Indeed, the L. sanfranciscensis species was the dominant bacterial species in 41 out of 45 French sourdoughs (Lhomme et al., 2015a,b, 2016; Michel et al., 2016). The origin of the yeast species diversity is still unknown. The drivers for the establishment and composition of LAB sourdough diversity have been reviewed (Gobbetti et al., 2016; Gänzle and Ripari, 2016). Numerous factors drive the LAB species diversity, such as the house microbiota, the flour species, and other ingredients. Drift has been shown to drive change in sourdough bacterial species in laboratory experiments (Lhomme et al., 2014; Rizzello et al., 2015). Several studies revealed that dispersion from the bakery environment or from the flour may also explained part of the composition of the sourdough bacterial community (Minervini et al., 2015; Minervini et al., 2018). Moreover, differences in sourdough environment, such as pH, cereal composition of the flour or fermentation time, may drive sourdough microbial community divergence (see for a review Gänzle and Ripari, 2016). The factors that drive yeast species diversity have been less explored. Additional studies, including a wider collection of sourdoughs or experimentations on different bakery practices, need to be carried out to better understand the origin and conservation of yeast species and genetic diversity.
Author Contributions
CU carried out the experiments, analyzed all the data, and drafted the manuscript. PM analyzed the pyrosequencing data. SC analyzed the new species. JL and DS designed the project, analyzed all the data and revised the manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We acknowledge financial support from ANR Bakery–13-ALID-0005 (French National Research Agency), région Ile de France (DIM ASTREA). This work is part of the Ph.D. thesis of CU (Urien, 2015). We thank the 14 bakers for their collaboration on the project and for sharing their bread-making knowledge and pleasure in good bread with us. Thanks are also addressed to Aurélie Bourgais, Candice Aulard and Stéphane Guezenec for their technical assistance and to Laurence Bérard and Mathieu Thomas for their help to study baker’s practices.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2019.00201/full#supplementary-material
Footnotes
- ^ http://genome.jouy.inra.fr/yeastip/
- ^ http://www.eurofins.com/en.aspx
- ^ https://unite.ut.ee/sh_files/UNITE_public_01.08.2015.fasta.zip
References
Arroyo-López, F. N., Medina, E., Ruiz-Bellido, M. Á., Romero-Gil, V., Montes-Borrego, M., and Landa, B. B. (2016). Enhancement of the knowledge on fungal communities in directly brined Aloreña de Málaga Green olive fermentations by metabarcoding analysis. PLoS One 11:e0163135. doi: 10.1371/journal.pone.0163135
Benjamini, Y., and Hochberg, Y. (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300. doi: 10.1111/j.2517-6161.1995.tb02031.x
Benson, D. A., Karsch-Mizrachi, I., Lipman, D. J., Ostell, J., and Wheeler, D. L. (2006). GenBank. Nucleic Acids Res. 34, D16–D20. doi: 10.1093/nar/gkj157
Birch, A. N., Petersen, M. A., Arneborg, N., and Hansen, Å. S. (2013). Influence of commercial baker’s yeasts on bread aroma profiles. Food Res. Int. 52, 160–166. doi: 10.1016/j.foodres.2013.03.011
Bourdichon, F., Casaregola, S., Farrokh, C., Frisvad, J. C., Gerds, M. L., Hammes, W. P., et al. (2012). Food fermentations: microorganisms with technological beneficial use. Int. J. Food Microbiol. 154, 87–97. doi: 10.1016/j.ijfoodmicro.2011.12.030
Caplice, E., and Fitzgerald, G. F. (1999). Food fermentations: role of microorganisms in food production and preservation. Int. J. Food Microbiol. 50, 131–149. doi: 10.1016/S0168-1605(99)00082-3
Caporaso, J. G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E. K., et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336. doi: 10.1038/nmeth.f.303
Carbonetto, B., Ramsayer, J., Nidelet, T., Legrand, J., and Sicard, D. (2018). Bakery yeasts, a new model for studies in ecology and evolution. Yeast 35, 591–603. doi: 10.1002/yea.3350
Cole, J. R., Wang, Q., Fish, J. A., Chai, B., McGarrell, D. M., Sun, Y., et al. (2014). Ribosomal database project: data and tools for high throughput rRNA analysis. Nucleic Acids Res. 42, D633–D642. doi: 10.1093/nar/gkt1244
Corsetti, A., Lavermicocca, P., Morea, M., Baruzzi, F., Tosti, N., and Gobbetti, M. (2001). Phenotypic and molecular identification and clustering of lactic acid bacteria and yeasts from wheat (species Triticum durum and Triticum aestivum) sourdoughs of Southern Italy. Int. J. Food Microbiol. 64, 95–104. doi: 10.1016/S0168-1605(00)00447-5
Coton, M., Pawtowski, A., Taminiau, B., Burgaud, G., Deniel, F., Coulloumme-Labarthe, L., et al. (2017). Unraveling microbial ecology of industrial-scale kombucha fermentations by metabarcoding and culture-based methods. FEMS Microbiol. Ecol. 93, 1–16. doi: 10.1093/femsec/fix048
Csoma, H., Zakany, N., Capece, A., Romano, P., and Sipiczki, M. (2010). Biological diversity of Saccharomyces yeasts of spontaneously fermenting wines in four wine regions: comparative genotypic and phenotypic analysis. Int. J. Food Microbiol. 140, 239–248. doi: 10.1016/j.ijfoodmicro.2010.03.024
Damiani, P., Gobbetti, M., Cossignani, L., Corsetti, A., Simonetti, M. S., and Rossi, J. (1996). The sourdough microflora. characterization of hetero- and homofermentative lactic acid bacteria, yeasts and their interactions on the basis of the volatile compounds produced. LWT Food Sci. Technol. 29, 63–70. doi: 10.1006/fstl.1996.0009
Daniel, H.-M., and Meyer, W. (2003). Evaluation of ribosomal RNA and actin gene sequences for the identification of ascomycetous yeasts. Int. J. Food Microbiol. 86, 61–78. doi: 10.1016/S0168-1605(03)00248-4
De Filippis, F. D., Parente, E., and Ercolini, D. (2017). Metagenomics insights into food fermentations. Microbial. Biotechnol. 10, 91–102. doi: 10.1111/1751-7915.12421
De Vuyst, L., Harth, H., Van Kerrebroeck, S., and Leroy, F. (2016). Yeast diversity of sourdoughs and associated metabolic properties and functionalities. Int. J. Food Microbiol. 239, 26–34. doi: 10.1016/j.ijfoodmicro.2016.07.018
De Vuyst, L., Van Kerrebroeck, S., Harth, H., Huys, G., Daniel, H.-M., and Weckx, S. (2014). Microbial ecology of sourdough fermentations: diverse or uniform? Food Microbiol. 37, 11–29. doi: 10.1016/j.fm.2013.06.002
De Vuyst, L., Van Kerrebroeck, S., and Leroy, F. (2017). “Chapter two - microbial ecology and process technology of sourdough fermentation,” in Advances in Applied Microbiology, eds S. Sariaslani and G. M. Gadd (Cambridge, MA: Academic Press), 49–160. doi: 10.1016/bs.aambs.2017.02.003
Demeulenaere, E., and Bonneuil, C. (2010). Les Mondes Agricoles en Politique. De la fin des Paysans au Retour de la Question Agricole. Available at: https://hal.archives-ouvertes.fr/hal-00528010 [accessed August 6, 2018].
Dertli, E., Mercan, E., Arıcı, M., Yılmaz, M. T., and Sagdiç, O. (2016). Characterisation of lactic acid bacteria from Turkish sourdough and determination of their exopolysaccharide (EPS) production characteristics. LWT Food Sci. Technol. 71, 116–124. doi: 10.1016/j.lwt.2016.03.030
Edgar, R. C. (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26, 2460–2461. doi: 10.1093/bioinformatics/btq461
Foschino, R., Gallina, S., Andrighetto, C., Rossetti, L., and Galli, A. (2004). Comparison of cultural methods for the identification and molecular investigation of yeasts from sourdoughs for Italian sweet baked products. FEMS Yeast Res. 4, 609–618. doi: 10.1016/j.femsyr.2003.12.006
Gänzle, M., and Ripari, V. (2016). Composition and function of sourdough microbiota: from ecological theory to bread quality. Int. J. Food Microbiol. 239, 19–25. doi: 10.1016/j.ijfoodmicro.2016.05.004
Gardes, M., and Bruns, T. D. (1993). ITS primers with enhanced specificity for basidiomycetes–application to the identification of mycorrhizae and rusts. Mol. Ecol. 2, 113–118. doi: 10.1111/j.1365-294X.1993.tb00005.x
Geiser, D. M., Jiménez-Gasco, M., del, M., Kang, S., Makalowska, I., Veeraraghavan, N., et al. (2004). FUSARIUM-ID v. 1.0: a DNA sequence database for identifying emphasis type = ”Italic” fusarium emphasis. Eur. J. Plant Pathol. 110, 473–479. doi: 10.1023/B:EJPP.0000032386.75915.a0
Gobbetti, M., De Angelis, M., Di Cagno, R., Calasso, M., Archetti, G., and Rizzello, C. G. (2018). Novel insights on the functional/nutritional features of the sourdough fermentation. Int. J. Food Microbiol. (in press). doi: 10.1016/j.ijfoodmicro.2018.05.018
Gobbetti, M., Minervini, F., Pontonio, E., Di Cagno, R., and De Angelis, M. (2016). Drivers for the establishment and composition of the sourdough lactic acid bacteria biota. Int. J. Food Microbiol. 239, 3–18. doi: 10.1016/j.ijfoodmicro.2016.05.022
Gobbetti, M., Rizzello, C. G., Di Cagno, R., and De Angelis, M. (2014). How the sourdough may affect the functional features of leavened baked goods. Food Microbiol. 37, 30–40. doi: 10.1016/j.fm.2013.04.012
Gouliamova, D. E., Dimitrov, R. A., and Stoilova-Disheva, M. M. (2012). DNA barcoding of yeasts from selected bulgarian food products. Biotechnol. Biotechnol. Equip. 26, 32–34. doi: 10.5504/50YRTIMB.2011.0006
Hansen, A., and Schieberle, P. (2005). Generation of aroma compounds during sourdough fermentation: applied and fundamental aspects. Trends Food Sci. Technol. 16, 85–94. doi: 10.1016/j.tifs.2004.03.007
Illeghems, K., De Vuyst, L., Papalexandratou, Z., and Weckx, S. (2012). Phylogenetic analysis of a spontaneous cocoa bean fermentation metagenome reveals new insights into its bacterial and fungal community diversity. PLoS One 7:e38040. doi: 10.1371/journal.pone.0038040
Jacques, N., Sarilar, V., Urien, C., Lopes, M. R., Morais, C. G., Uetanabaro, A. P. T., et al. (2016). Three novel ascomycetous yeast species of the kazachstania clade, kazachstania saulgeensis sp. nov., kazachstaniaserrabonitensis sp. nov. and kazachstania australis sp. nov. reassignment of candida humilis to kazachstania humilis f.a. comb. nov. and candida pseudohumilis to kazachstania pseudohumilis f.a. comb. nov. Int. J. Syst. Evol. Microbiol. 66, 5192–5200. doi: 10.1099/ijsem.0.001495
Kergourlay, G., Taminiau, B., Daube, G., and Champomier Vergès, M.-C. (2015). Metagenomic insights into the dynamics of microbial communities in food. Int. J. Food Microbiol. 213, 31–39. doi: 10.1016/j.ijfoodmicro.2015.09.010
Kõljalg, U., Nilsson, R. H., Abarenkov, K., Tedersoo, L., Taylor, A. F. S., Bahram, M., et al. (2013). Towards a unified paradigm for sequence-based identification of fungi. Mol. Ecol. 22, 5271–5277. doi: 10.1111/mec.12481
Kurtzman, C. P., and Robnett, C. J. (2003). Phylogenetic relationships among yeasts of the “Saccharomyces complex” determined from multigene sequence analyses. FEMS Yeast Res. 3, 417–432. doi: 10.1016/S1567-1356(03)00012-6
Lattanzi, A., Minervini, F., Di Cagno, R., Diviccaro, A., Antonielli, L., Cardinali, G., et al. (2013). The lactic acid bacteria and yeast microbiota of eighteen sourdoughs used for the manufacture of traditional Italian sweet leavened baked goods. Int. J. Food Microbiol. 163, 71–79. doi: 10.1016/j.ijfoodmicro.2013.02.010
Lhomme, E., Mezaize, S., Ducasse, M. B., Chiron, H., Champomier-Vergès, M. C., Chaillou, S., et al. (2014). A polyphasic approach to study the dynamics of microbial population of an organic wheat sourdough during its conversion to gluten-free sourdough. Int. Microbiol. 17, 1–9. doi: 10.2436/20.1501.01.202
Lhomme, E., Lattanzi, A., Dousset, X., Minervini, F., De Angelis, M., Lacaze, G., et al. (2015a). Lactic acid bacterium and yeast microbiotas of sixteen French traditional sourdoughs. Int. J. Food Microbiol. 215, 161–170. doi: 10.1016/j.ijfoodmicro.2015.09.015
Lhomme, E., Orain, S., Courcoux, P., Onno, B., and Dousset, X. (2015b). The predominance of Lactobacillus sanfranciscensis in French organic sourdoughs and its impact on related bread characteristics. Int. J. Food Microbiol. 213, 40–48. doi: 10.1016/j.ijfoodmicro.2015.05.010
Lhomme, E., Urien, C., Legrand, J., Dousset, X., Onno, B., and Sicard, D. (2016). Sourdough microbial community dynamics: an analysis during French organic bread-making processes. Food Microbiol. 53, 41–50. doi: 10.1016/j.fm.2014.11.014
Liang, H., Chen, H., Zhang, W., Yu, C., Ji, C., and Lin, X. (2018). Investigation on microbial diversity of industrial zhacai paocai during fermentation using high-throughput sequencing and their functional characterization. LWT 91, 460–466. doi: 10.1016/j.lwt.2018.01.088
Liu, L., Li, Y., Li, S., Hu, N., He, Y., Pong, R., et al. (2012). Comparison of next-generation sequencing systems. BioMed. Res. Int. 2012:251364. doi: 10.1155/2012/251364
Liu, T., Li, Y., Chen, J., Sadiq, F. A., Zhang, G., Li, Y., et al. (2016). Prevalence and diversity of lactic acid bacteria in Chinese traditional sourdough revealed by culture dependent and pyrosequencing approaches. LWT Food Sci. Technol. 68, 91–97. doi: 10.1016/j.lwt.2015.12.025
Liu, T., Li, Y., Sadiq, F. A., Yang, H., Gu, J., Yuan, L., et al. (2018). Predominant yeasts in Chinese traditional sourdough and their influence on aroma formation in Chinese steamed bread. Food Chem. 242, 404–411. doi: 10.1016/j.foodchem.2017.09.081
Lv, X.-C., Huang, X.-L., Zhang, W., Rao, P.-F., and Ni, L. (2013). Yeast diversity of traditional alcohol fermentation starters for Hong Qu glutinous rice wine brewing, revealed by culture-dependent and culture-independent methods. Food Control 34, 183–190. doi: 10.1016/j.foodcont.2013.04.020
Lozupone, C., and Knight, R. (2005). UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235. doi: 10.1128/AEM.71.12.8228-8235.2005
Martin, K. J., and Rygiewicz, P. T. (2005). Fungal-specific PCR primers developed for analysis of the ITS region of environmental DNA extracts. BMC Microbiol. 5:28. doi: 10.1186/1471-2180-5-28
Martinez-Anaya, M. A., Pitarch, B., Bayarri, P., and de Barber, C. (1990). Microflora of the sourdoughs of wheat flour bread. X. Interactions between yeasts and lactic acid bacteria in wheat doughs and their effects on bread quality. Cereal Chem. 67, 85–91.
Michel, E., Monfort, C., Deffrasnes, M., Guezenec, S., Lhomme, E., Barret, M., et al. (2016). Characterization of relative abundance of lactic acid bacteria species in French organic sourdough by cultural, qPCR and MiSeq high-throughput sequencing methods. Int. J. Food Microbiol. 239, 35–43. doi: 10.1016/j.ijfoodmicro.2016.07.034
Minervini, F., Cagno, R. D., Lattanzi, A., Angelis, M. D., Antonielli, L., Cardinali, G., et al. (2011). The lactic acid bacteria and yeast microbiota of nineteen sourdoughs used for the manufacture of traditional/typical Italian breads: interactions between ingredients and microbial species diversity. Appl. Environ. Microbiol. 78, 1251–1264. doi: 10.1128/AEM.07721-11
Minervini, F., De Angelis, M., Di Cagno, R., and Gobbetti, M. (2014). Ecological parameters influencing microbial diversity and stability of traditional sourdough. Int. J. Food Microbiol. 171, 136–146. doi: 10.1016/j.ijfoodmicro.2013.11.021
Minervini, F., Lattanzi, A., De Angelis, M., Celano, G., and Gobbetti, M. (2015). House microbiotas as sources of lactic acid bacteria and yeasts in traditional italian sourdoughs. Food Microbiol. 52, 66–76. doi: 10.1016/j.fm.2015.06.009
Minervini, F., Lattanzi, A., Dinardo, F. R., De Angelis, M., and Gobbetti, M. (2018). Wheat endophytic lactobacilli drive the microbial and biochemical features of sourdoughs. Food Microbiol. 70, 162–171. doi: 10.1016/j.fm.2017.09.006
Nakamura, Y., Cochrane, G., and Karsch-Mizrachi, I. (2013). The international nucleotide sequence database collaboration. Nucleic Acids Res. 41, D21–D24. doi: 10.1093/nar/gks1084
Nguyen, H.-V., and Gaillardin, C. (1997). Two subgroups within the saccharomyces bayanus species evidenced by PCR amplification and restriction polymorphism of the non-transcribed spacer 2 in the ribosomal DNA unit. Syst. Appl. Microbiol. 20, 286–294. doi: 10.1016/S0723-2020(97)80075-6
Pontonio, E., Rizzello, C. G., Di Cagno, R., Dousset, X., Clément, H., Filannino, P., et al. (2016). How organic farming of wheat may affect the sourdough and the nutritional and technological features of leavened baked goods. Int. J. Food Microbiol. 239, 44–53. doi: 10.1016/j.ijfoodmicro.2016.07.013
Pruesse, E., Quast, C., Knittel, K., Fuchs, B. M., Ludwig, W., Peplies, J., et al. (2007). SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res. 35, 7188–7196. doi: 10.1093/nar/gkm864
Raimondi, S., Amaretti, A., Rossi, M., Fall, P. A., Tabanelli, G., Gardini, F., et al. (2017). Evolution of microbial community and chemical properties of a sourdough during the production of colomba, an italian sweet leavened baked product. LWT 86, 31–39. doi: 10.1016/j.lwt.2017.07.042
Reva, O. N., Zaets, I. E., Ovcharenko, L. P., Kukharenko, O. E., Shpylova, S. P., Podolich, O. V., et al. (2015). Metabarcoding of the kombucha microbial community grown in different microenvironments. AMB Express 5:124. doi: 10.1186/s13568-015-0124-5
Rizzello, C. G., Cavoski, I., Turk, J., Ercolini, D., Nionelli, L., Pontonio, E., et al. (2015). Organic cultivation of triticum turgidum subsp. Durum is reflected in the flour-sourdough fermentation-bread axis. Appl. Environ. Microbiol. 81, 3192–3204. doi: 10.1128/AEM.04161-14
Sarilar, V., Sterck, L., Matsumoto, S., Jacques, N., Neuvéglise, C., Tinsley, C. R., et al. (2017). Genome sequence of the type strain CLIB 1764T ( = CBS 14374T) of the yeast species kazachstania saulgeensis isolated from French organic sourdough. Genomics Data 13, 41–43. doi: 10.1016/j.gdata.2017.07.003
Scheirlinck, I., Meulen, R. V., der Vuyst, L. D., Vandamme, P., and Huys, G. (2009). Molecular source tracking of predominant lactic acid bacteria in traditional Belgian sourdoughs and their production environments. J. Appl. Microbiol. 106, 1081–1092. doi: 10.1111/j.1365-2672.2008.04094.x
Scheirlinck, I., Van der Meulen, R., Van Schoor, A., Vancanneyt, M., De Vuyst, L., Vandamme, P., et al. (2007). Influence of geographical origin and flour type on diversity of lactic acid bacteria in traditional belgian sourdoughs. Appl. Environ. Microbiol. 73, 6262–6269. doi: 10.1128/AEM.00894-07
Schoch, C. L., Seifert, K. A., Huhndorf, S., Robert, V., Spouge, J. L., Levesque, C. A., et al. (2012). Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc. Natl. Acad. Sci. U.S.A. 109, 6241–6246. doi: 10.1073/pnas.1117018109
Tamang, J. P., Watanabe, K., and Holzapfel, W. H. (2016). Review: diversity of microorganisms in global fermented foods and beverages. Front. Microbiol. 7:377. doi: 10.3389/fmicb.2016.00377
Urien, C. (2015). Diversité des espèces de levures dans des levains Naturels Français Produits à partir de farine issue de l’Agriculture Biologique : Une étude Pilote Pour Analyser les Pratiques Boulangères et les patterns des Communautés Microbiennes. Ph.D. Thesis, [Gif-sur-Yvette]: Université Paris Sud-Paris, French.
van Hijum, S. A., Vaughan, E. E., and Vogel, R. F. (2013). Application of state-of-art sequencing technologies to indigenous food fermentations. Curr. Opin. Biotechnol. 24, 178–186. doi: 10.1016/j.copbio.2012.08.004
Van Kerrebroeck, S., Maes, D., and De Vuyst, L. (2017). Sourdoughs as a function of their species diversity and process conditions, a meta-analysis. Trends Food Sci. Technol. 68, 152–159. doi: 10.1016/j.tifs.2017.08.016
Vrancken, G., De Vuyst, L., Van der Meulen, R., Huys, G., Vandamme, P., and Daniel, H.-M. (2010). Yeast species composition differs between artisan bakery and spontaneous laboratory sourdoughs. FEMS Yeast Res. 10, 471–481. doi: 10.1111/j.1567-1364.2010.00621.x
Vu, D., Groenewald, M., de Vries, M., Gehrmann, T., Stielow, B., Eberhardt, U., et al. (2019). Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Stud. Mycol. 92, 135–154. doi: 10.1016/j.simyco.2018.05.001
Vu, D., Groenewald, M., Szöke, S., Cardinali, G., Eberhardt, U., Stielow, B., et al. (2016). DNA barcoding analysis of more than 9 000 yeast isolates contributes to quantitative thresholds for yeast species and genera delimitation. Stud. Mycol. 85, 91–105. doi: 10.1016/j.simyco.2016.11.007
Wang, Q., Garrity, G. M., Tiedje, J. M., and Cole, J. R. (2007). Naïve bayesian classifier for rapid assignment of rrna sequences into the new bacterial taxonomy. Appl. Environ. Microbiol. 73, 5261–5267. doi: 10.1128/AEM.00062-07
Weckx, S., Van Kerrebroeck, S., and De Vuyst, L. (2018). Omics approaches to understand sourdough fermentation processes. Int. J. Food Microbiol. doi: 10.1016/j.ijfoodmicro.2018.05.029 [Epub ahead of print].
Weiss, S., Samson, F., Navarro, D., and Casaregola, S. (2013). YeastIP: a database for identification and phylogeny of Saccharomycotina yeasts. FEMS Yeast Res. 13, 117–125. doi: 10.1111/1567-1364.12017
Yang, L., Yang, H., Tu, Z., and Wang, X. (2016). High-throughput sequencing of microbial community diversity and dynamics during douchi fermentation. PLoS One 11:e0168166. doi: 10.1371/journal.pone.0168166
Zhang, G., Sadiq, F. A., Zhu, L., Liu, T., Yang, H., Wang, X., et al. (2015). Investigation of microbial communities of chinese sourdoughs using culture-dependent and DGGE approaches. J. Food Sci. 80, M2535–M2542. doi: 10.1111/1750-3841.13093
Zhang, J., Liu, W., Sun, Z., Bao, Q., Wang, F., Yu, J., et al. (2011). Diversity of lactic acid bacteria and yeasts in traditional sourdoughs collected from western region in inner mongolia of china. Food Control 22, 767–774. doi: 10.1016/j.foodcont.2010.11.012
Keywords: microbial communities, metabarcoding, food microbial ecosystem, yeasts, Kazachstania, fermented food, bakers, farmer-bakers
Citation: Urien C, Legrand J, Montalent P, Casaregola S and Sicard D (2019) Fungal Species Diversity in French Bread Sourdoughs Made of Organic Wheat Flour. Front. Microbiol. 10:201. doi: 10.3389/fmicb.2019.00201
Received: 07 August 2018; Accepted: 24 January 2019;
Published: 18 February 2019.
Edited by:
Michael Gänzle, University of Alberta, CanadaReviewed by:
Feng-Yan Bai, Institute of Microbiology (CAS), ChinaSimon Van Kerrebroeck, Vrije University Brussel, Belgium
Copyright © 2019 Urien, Legrand, Montalent, Casaregola and Sicard. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Delphine Sicard, ZGVscGhpbmUuc2ljYXJkQGlucmEuZnI=
 Charlotte Urien
Charlotte Urien Judith Legrand
Judith Legrand Pierre Montalent1
Pierre Montalent1 Serge Casaregola
Serge Casaregola Delphine Sicard
Delphine Sicard