- 1Graduate School of Environment and Information Sciences, Yokohama National University, Yokohama, Kanagawa, Japan
- 2Faculty of Environment and Information Sciences, Yokohama National University, Yokohama, Kanagawa, Japan
- 3Department of Earth Sciences, Graduate School of Science, Chiba University, Chiba, Japan
Introduction: Microbial communities are important components of glacier and snowpack ecosystems that influence biogeochemical cycles and snow/ice melt. Recent environmental DNA surveys have revealed that chytrids dominate the fungal communities in polar and alpine snowpacks. These could be parasitic chytrids that infect snow algae as observed microscopically. However, the diversity and phylogenetic position of parasitic chytrids has not been identified due to difficulties in establishing their culture and subsequent DNA sequencing. In this study, we aimed to identify the phylogenetic positions of chytrids infecting the snow algae, Chloromonas spp., bloomed on snowpacks in Japan.
Methods: By linking a microscopically picked single fungal sporangium on a snow algal cell to a subsequent sequence of ribosomal marker genes, we identified three novel lineages with distinct morphologies.
Results: All the three lineages belonged to Mesochytriales, located within “Snow Clade 1”, a novel clade consisting of uncultured chytrids from snow-covered environments worldwide. Additionally, putative resting spores of chytrids attached to snow algal cells were observed.
Discussion: This suggests that chytrids may survive as resting stage in soil after snowmelt. Our study highlights the potential importance of parasitic chytrids that infect snow algal communities.
1. Introduction
Seasonal snowpacks are inhabited by diverse organisms of various taxonomic groups, including photosynthetic primary producers (snow algae and cyanobacteria) (Takeuchi, 2001; Hoham and Remias, 2020) and heterotrophic microorganisms (bacteria, fungi, and micro invertebrates) (Yakimovich et al., 2020; Fiołka et al., 2021; Kobayashi et al., 2023). During snowmelt, snow algae grow on snow surfaces, which covers the surface with green or red color (Hoham and Duval, 2001). Colored snow is typically dominated by algae belonging to the phylum Chlorophyta (Procházková et al., 2019; Hoham and Remias, 2020; Nakashima et al., 2021; Ono et al., 2021). Snow algae also accelerate snowmelt, as they can reduce the albedo of snow surface, resulting in increased absorption of solar radiation by snow (Lutz et al., 2016; Ganey et al., 2017; Hotaling et al., 2021). Several taxonomic studies have been performed on snow algae to elucidate their ecology (Matsuzaki et al., 2015, 2018, 2019; Segawa et al., 2018; Procházková et al., 2019). Some bacteria and fungi have symbiotic or parasitic relationships with snow algae, and they may affect snow algal population (Terashima et al., 2017). Recently, some studies have attempted to characterize the snow algae–bacteria relationships using 16S rRNA gene amplicon sequencing and co-cultivation with snow algae (Terashima et al., 2017; Krug et al., 2020). However, few studies have focused on the snow algae–fungus relationship, and the taxonomic knowledge of this relationship is particularly poor.
Recent environmental DNA analyses have revealed that diverse fungi inhabit alpine snowpacks worldwide and chytrids often dominate fungal communities (Freeman et al., 2009; Schmidt et al., 2012; Naff et al., 2013; Brown et al., 2015; Yakimovich et al., 2020). Novel clades composed mainly of environmental DNA sequences from snow-covered regions were identified in Chytridiomycota (Naff et al., 2013). However, as these chytrid sequences were directly obtained from soil and snow samples and not from cultures, their morphology, life cycles, and ecology remain unknown.
Microscopic observations have shown that chytrids infect algae in alpine snowpacks and glaciers. Kol (1942) reported that a chytrid, morphologically identified as Rhizophidium sphaerocarpum, infects the glacier alga Ancylonema nordenskioldii on a glacier in Alaska. Additionally, chytrids infecting the snow alga Sanguina nivaloides (formerly known as Chlamydomonas nivalis) have been observed in polar and alpine snowpacks (Kobayashi and Okubo, 1954; Kol, 1968; Yakimovich et al., 2020; Fiołka et al., 2021; Kobayashi et al., 2023). As observed in lakes, these chytrids may suppress algal populations and affect trophic dynamics in snow ecosystems (Kagami et al., 2007; Frenken et al., 2017). However, as they have not yet been cultured, their diversity and phylogenetic positions have not been determined.
This study aimed to determine the phylogenetic position of parasitic chytrids that infect snow algae. We used single-spore PCR to sequence uncultured chytrids in the bloom of snow alga Chloromonas spp. in alpine snowpacks in Japan. This method enabled us to directly link microscopic observations to DNA sequencing without culturing (Ishida et al., 2015; Kagami et al., 2021; Van den Wyngaert et al., 2022). This study provides the first phylogenetic evidence of parasitic chytrids infecting snow algae in alpine regions.
2. Materials and methods
2.1. Study site
The field study was performed on Mt. Gassan, Yamagata prefecture in Japan at two sites, Site A (38° 29′ N, 140° 00′ E 770 m above sea level (a. s. l.); Figure 1A) and Site B (38° 31′ N, 140° 00′ E 1,150 m a. s. l.; Figure 1B). A large amount of snow accumulates in Mount Gassan every winter (Kariya, 2005). At Sites A and B, green snow appeared from late April to mid-June (Figures 2A,C). The vegetation at the study site is dominated by mountain broad-leaved deciduous trees, including Fagus crenata shown in Figure 1A, up to an elevation of 1,500 m a. s. l.
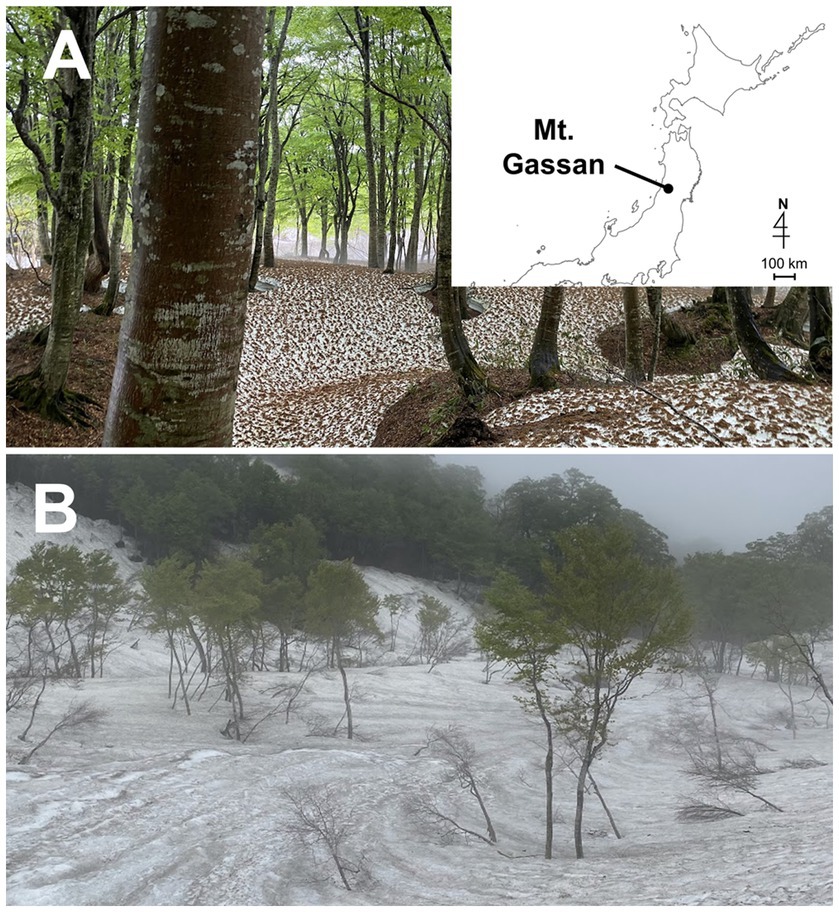

Figure 1. Photographs of the landscape of the sampling sites on Mt. Gassan in Japan. (A): Site A (770 m a. s. l.). The map shows the location of Mt. Gassan. (B): Site B (1150 m a. s. l.).
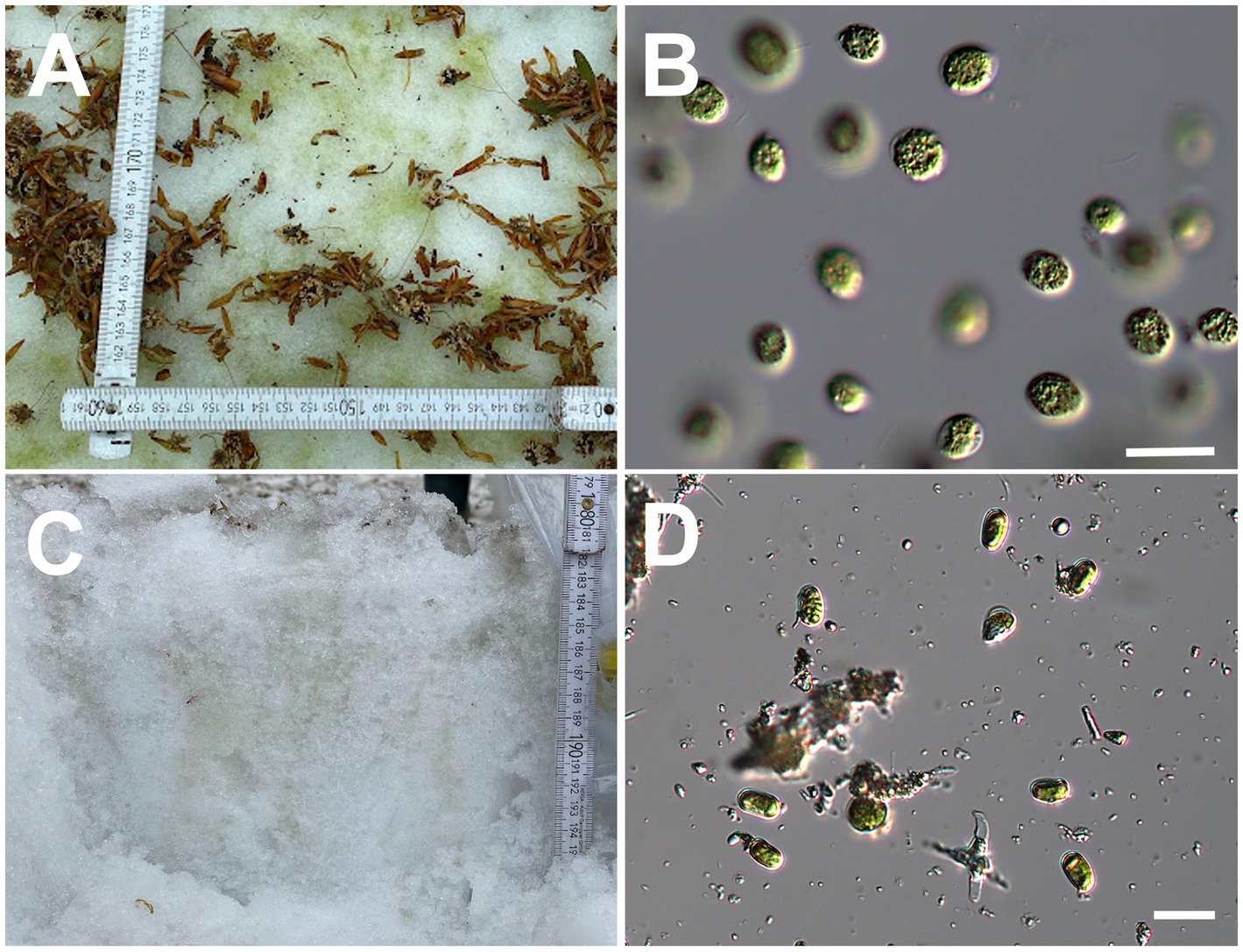

Figure 2. (A): Green snow at Site A and (B): micrograph of the green snow at Site A. (C): Vertical section of green snow at Site B and (D): micrograph of the green snow at Site B. All scale bars: 20 μm.
2.2. Sample collection
Samples were collected from green snow that appeared in May 2021 and May 2022 at the two sites (Figures 2A,C). The 2021 samples were collected in 15 mL plastic tubes, stored in a box filled with snow, and transported to a laboratory. Some tubes were stored at −80°C for DNA analysis in a deep freezer (MDF-C8V1-PJ, Panasonic, Japan). The remaining tubes were incubated at 3°C under dim light 20 μmol photons m−2 s−1 and 12:12 h light:dark cycle in an incubator (SLC-25A, Mitsubishi Electric Engineering, Japan). The incubated samples were observed under a light microscope (IX71, Olympus, Japan) every few days to identify parasitic chytrids infecting Chloromonas spp.
2.3. Microscopic observation
Many snow algal cells of Chloromonas spp. were present in the green snow at both the sites (Figures 2B,D). After calcofluor white fluorescent (CFW) –wheat germ agglutinin (WGA) double staining (Klawonn et al., 2023), the samples were observed under a microscope to identify parasitic chytrids infecting Chloromonas spp. Green snow samples were aliquoted into 1 mL Eppendorf tubes, and 5 μg mL−1 of CFW (Fluorescent Brightener 28, Sigma Aldrich, United States) and 5 μg mL−1 of WGA (Wheat Germ Agglutinin, Alexa Fluor™ 488 conjugate, Thermo Fisher Scientific, United States) were added. CFW and WGA can be used simultaneously because they have different binding target structures and absorb at different wavelengths, making the detection of chitin in fungal cell walls reliable (Klawonn et al., 2023). Photographs were taken using a digital camera (Advancam 305 color, Carl Zeiss, Germany) at differential interference contrast, blue fluorescence excitation (CFW), and green fluorescence excitation (WGA) on a fluorescence microscope (AXIO Imager. M2, Carl Zeiss, Germany). Based on the micrographs, the chytrids were morphologically typed (Figure 3). Chytrids with thick cell walls that accumulated single or multiple intracellular lipid globules were classified as the resting spore (Seto et al., 2017). To calculate the prevalence of chytrid infection in algal cells, we counted the number of algal cells with and without chytrids in six samples collected at Site A in May 2021. More than 200 algal cells were counted in each sample. The prevalence of infection was calculated as the number of infected algal cells divided by the total number of algal cells counted.
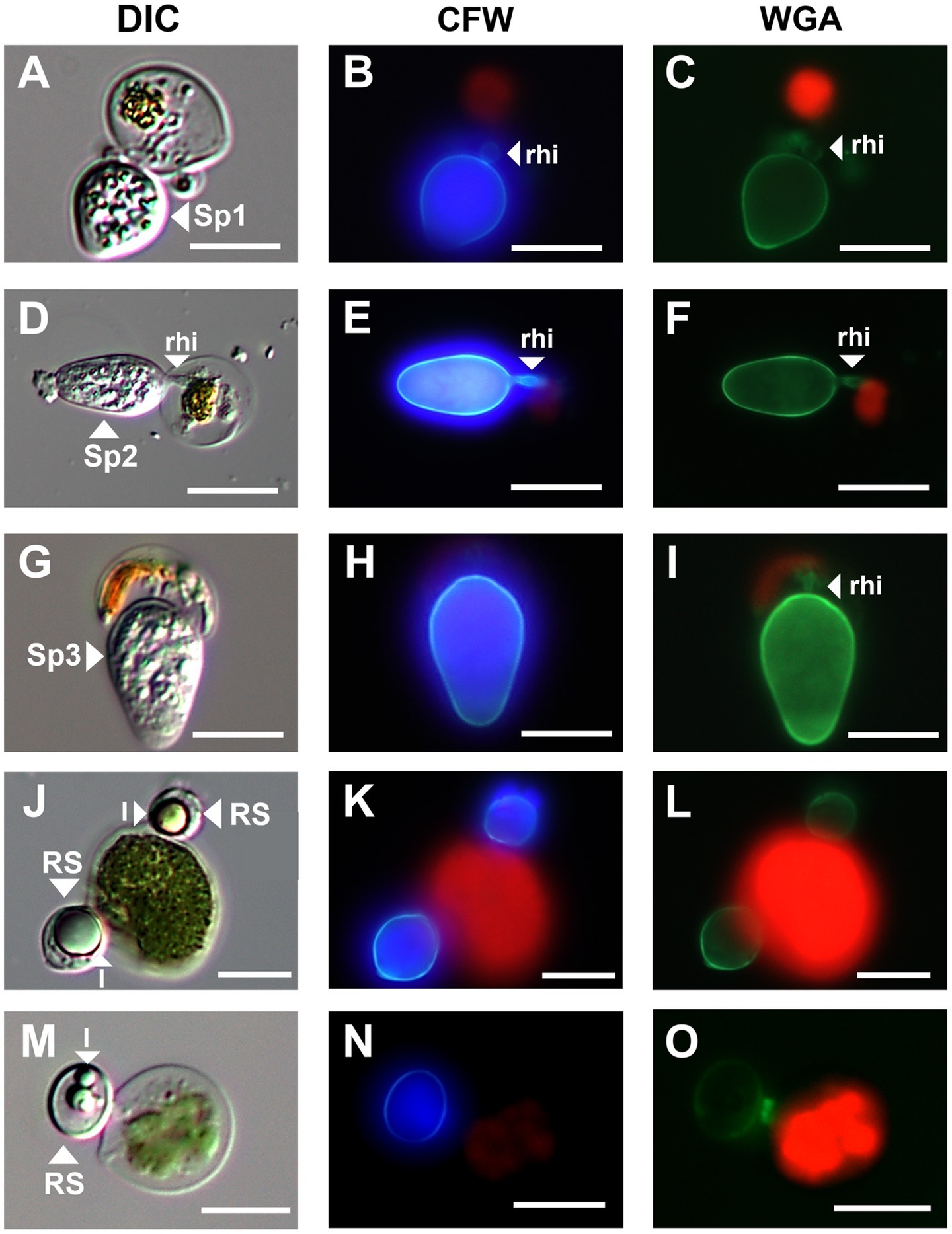
Figure 3. Micrographs of three sporangium type and resting spore of parasitic chytrids infecting Chloromonas spp. (A–C); Sp1, (D–F); Sp2, (G–I); Sp3, (J–O); resting spores. All scale bars are 10 μm. DIC, differential interference contrast; CFW, under blue fluorescent excitation after CFW staining; WGA, under green fluorescent excitation after WGA staining; rhi, rhizoids; l, lipid globules; RS, resting spore.
2.4. Single-spore PCR
2.4.1. Microscopic observation and isolation of parasitic chytrids
The cryopreserved and incubated samples were used for single-cell isolation. The cryopreserved samples were thawed at room temperature (approximately 20°C) before analysis. To stain the fungal cell wall, 5 μL of CFW was added to 1 mL of the samples. Chytrids infecting snow algae Chloromonas spp. were observed under an inverted fluorescence microscope (IX71, Olympus, Japan) and photographed using a digital camera (DP21, Olympus, Japan). The chytrid and host cells were isolated using a capillary pipette at room temperature. The isolated cells were washed three times with autoclaved deionized water to remove contamination of other fungi, and the cells were individually transferred into 200 μL PCR tubes with a small amount of deionized water.
2.4.2. DNA extraction
DNA extraction was performed using the HotSHOT method (Truett et al., 2000). PCR tubes containing isolated cells were filled with 5 μL of alkaline lysis buffer (20 mM EDTA and 0.2 mM NaOH; pH 12) and heat-treated in a thermal cycler (MiniAmp Plus, Thermo Fisher Scientific, United States) at 96°C for 10 min and 5°C for 5 min. Subsequently, 5 μL of neutralization buffer (40 mM Tris hydrochloride, pH 5) was added and stored at −80°C.
2.4.3. PCR amplification and sequencing
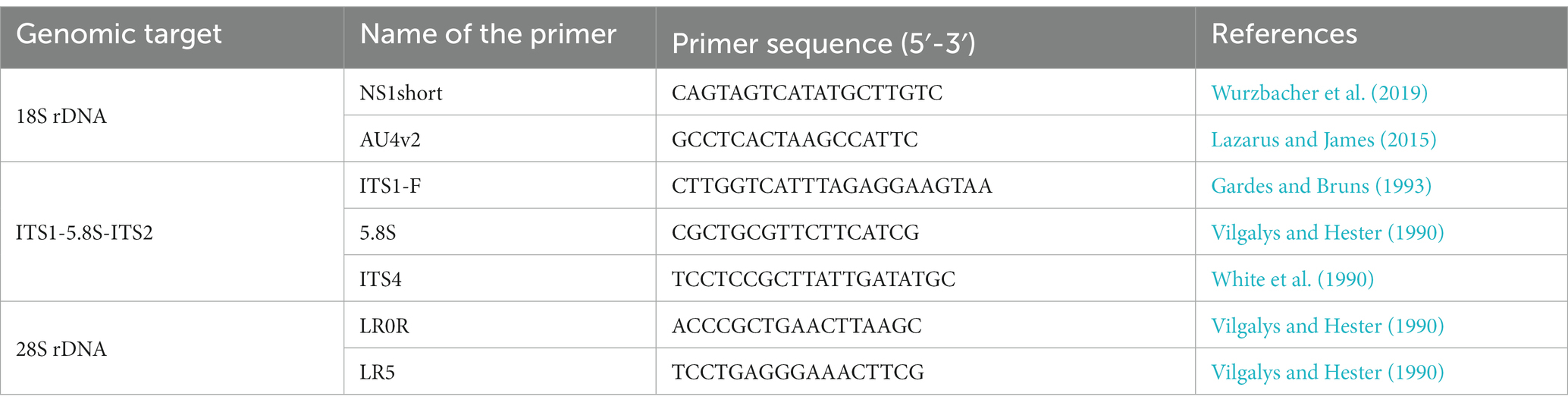
The extracted DNA was subjected to PCR amplification of the ribosomal RNA gene (rDNA) region in a thermal cycler using several fungus-specific primer sets (Table 1) and the DNA polymerase KOD FX Neo (TOYOBO, Japan). PCR was performed in 10 μL volume composed of 1.3 μL of sterile ultrapure water, 5 μL of 2× PCR Buffer for KOD FX Neo, 2 μL of dNTPs (2 mM), 0.2 μL of each primer (10 μM), 0.2 μL of KOD FX Neo, 0.1 μL of bovine serum albumin (50 mg mL−1, Wako, Japan), and 1 μL of extracted DNA. The thermal cycle for PCR was (1) 95°C for 5 min, (2) 12 cycles of denaturation at 95°C for 30 s, annealing at 54–48°C for 30 s (decrease of 0.5°C per cycle), extension at 68°C for 4 min, and (3) 23 cycles of 95°C for 30 s, 48°C for 30 s, and 68°C for 4 min. The PCR products were purified using ExoSAP-IT (Affymetrix, United States). Sequencing was performed using the commercial service of Eurofins Genomics in Japan with the sequencing primers listed in Table 2. Differences in the PCR success rate in isolated chytrids at sporangial stage and at resting spore stage were assessed by Fisher’s exact probability test. The statistical analysis was performed using R software (version 4.2.1; R Core Team, 2022).1
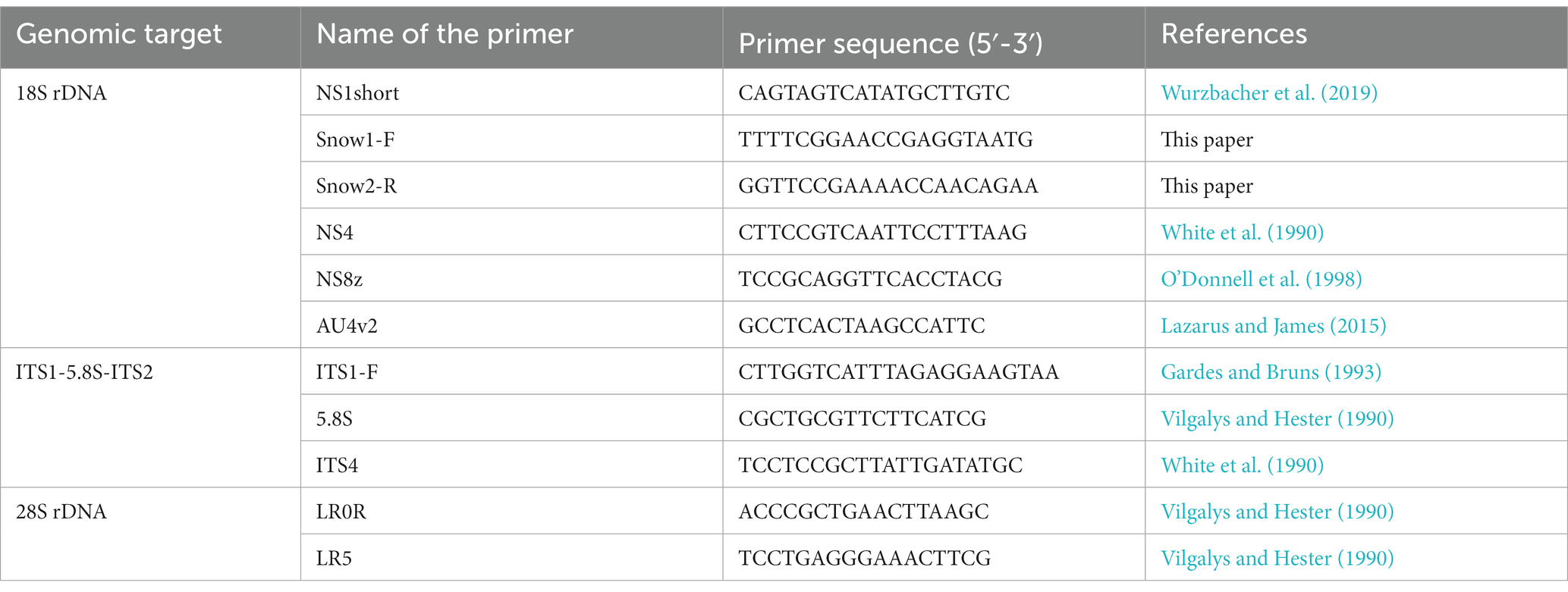
Table 2. List of primers used for sanger sequencing, which Snow1-F and Snow2-R are primers newly applied in this study.
2.5. Molecular phylogenetic analysis
The sequences obtained were assembled using Chromas Pro ver. 2.1.10 (Technelysium, Australia). The assembled sequences of 18S, ITS, and 28S rDNA were deposited under the accession numbers LC761280–LC761291. BLAST searches performed using the rDNA sequences obtained indicated that all isolates belonged to the order Mesochytriales of Chytridiomycota. Therefore, we performed phylogenetic analysis of 18S rDNA using the new sequences obtained in this study because this marker region is the richest in reference sequences for Mesochytriales. For molecular phylogenetic analysis, a dataset of 18S rDNA sequences from Mesochytriales, Gromochytriales, and Polyphagales was used (Table 3). Lobulomycetales were included as outgroups. Sequences were aligned using MAFFT (Katoh et al., 2019) with default parameters. Ambiguously aligned sequences were trimmed using trimAl (Capella-Gutiérrez et al., 2009). Molecular phylogenetic analysis using the maximum likelihood method was performed using the IQ-TREE online server (Trifinopoulos et al., 2016).
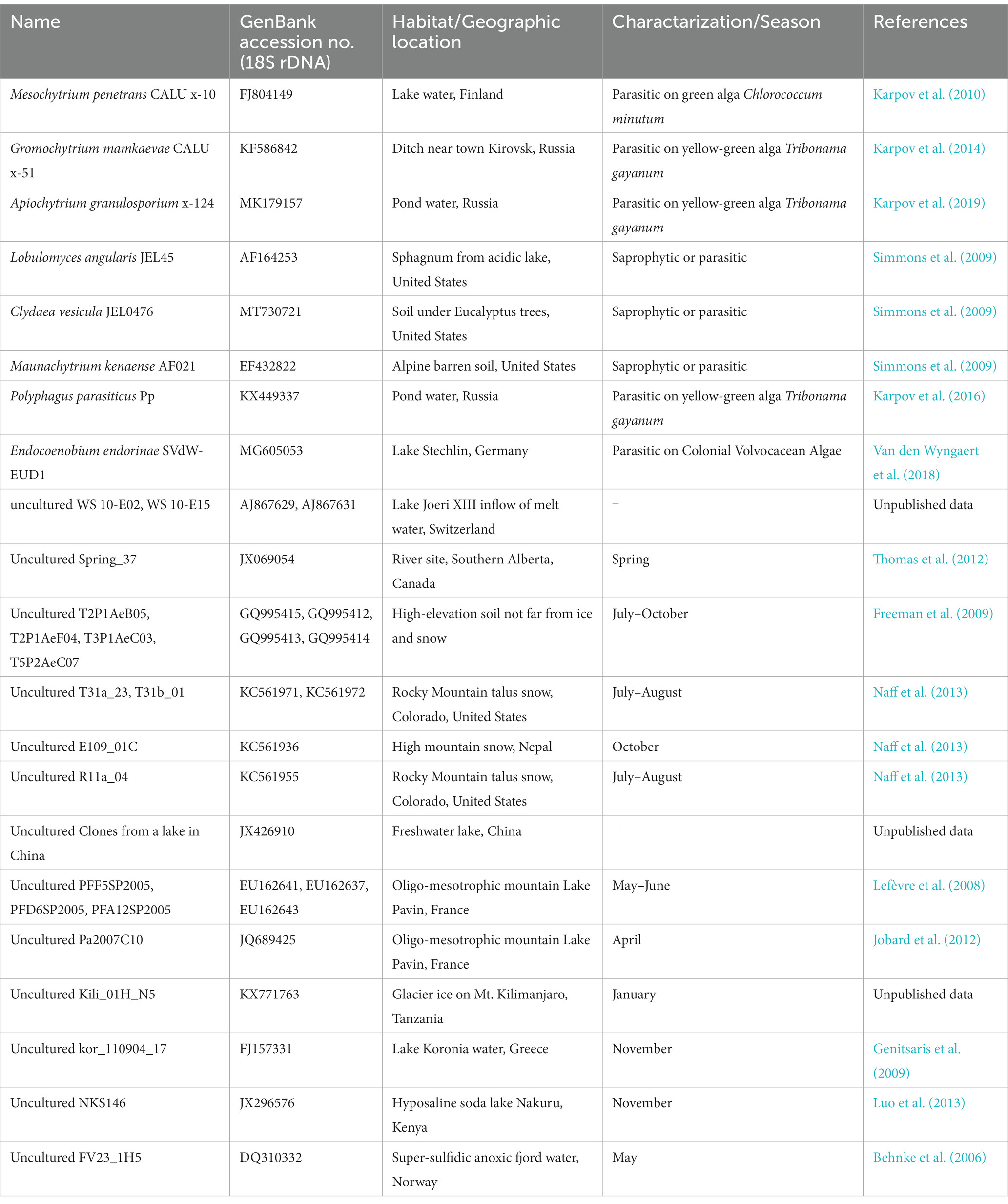
Table 3. List of known species and environmental DNA sequences belonging to Mesochytriales, Gromochytriales, Polyphagales, and Lobulomycetales used in the molecular phylogenetic tree (Figure 4).
3. Results
3.1. Observation of parasitic chytrids infecting snow algae
The green snow samples collected in this study revealed the presence of chytrids at the sporangial stage and putative resting spores attached to the surface of vegetative cells of Chloromonas spp. (Figure 3). Chytrids at the sporangial stage were further categorized into three types based on their shape (Figure 3). Sporangium type 1 (Sp1) was egg shaped with a rounded swollen rhizoid (Figures 3A–C). Sporangium type 2 (Sp2) was ellipsoidal and had rod-shaped rhizoids (Figures 3D–F). Sporangium type 3 (Sp3) was immersed in host cells and had a rod-shaped rhizoid structure (Figures 3G–I). Putative resting spores of chytrids were clearly visible with thick cell walls and single or multiple large lipid globules (Figures 3J–O). Based on the observations of six green snow samples, the average prevalence of chytrid infection in the total vegetative algal cells observed was calculated to be 5.34%.
3.2. Single-spore PCR
In total, 79 chytrid cells were isolated from the snow algal hosts, of which 57 cells could be categorized into the three aforementioned sporangial stages: Sp1 (n = 11), Sp2 (n = 37), and Sp3 (n = 4). The other five sporangial cells could not be assigned to any sporangial type because they were immature small sporangia. We also isolated 22 cells from the putative resting spore stage of parasitic chytrids (Supplementary Table S1).
Single-spore PCR was performed using the 72 cells, and at least one of the three rDNA regions was successfully amplified in 33 cells (Supplementary Table S1). Chytrids at the resting spore stage had a significantly lower PCR success rate than those at the sporangial stage (value of p = 2.135 x 10–5, Supplementary Table S1).
Sanger sequencing was used to determine the partial or assembled sequences of rDNA regions of the 32 chytrid cells. Eleven cells were sequenced for their 18S rDNA regions. Four cells were Sp1, five were Sp2, and two were Sp3 (Supplementary Table S1). The sequences differed between the types but were identical within types.
3.3. Molecular phylogenetic analysis
Our molecular phylogenetic analysis showed that the 11 isolated cells of parasitic chytrids infecting snow algae clustered into three novel lineages (Sp1, Sp2, and Sp3) in Mesochytriales (Figure 4). They were included in Snow Clade 1 consisting of sequences from alpine snow-related environments located worldwide that are deeply divergent from the clade containing the known species Mesochytrium penetrans. Sp1 is a sister group of clades composed of environmental sequences obtained from lakes in Switzerland fed by abundant snow meltwater, river water from southwestern Alberta, Canada, and talus snows from the Rocky Mountains in Colorado, United States. Sp2 is closely related to the environmental sequences from glaciers on the summit of Mt. Kilimanjaro, Tanzania. Sp3 formed an independent clade that was divergent from the clade containing Sp1 and Sp2.
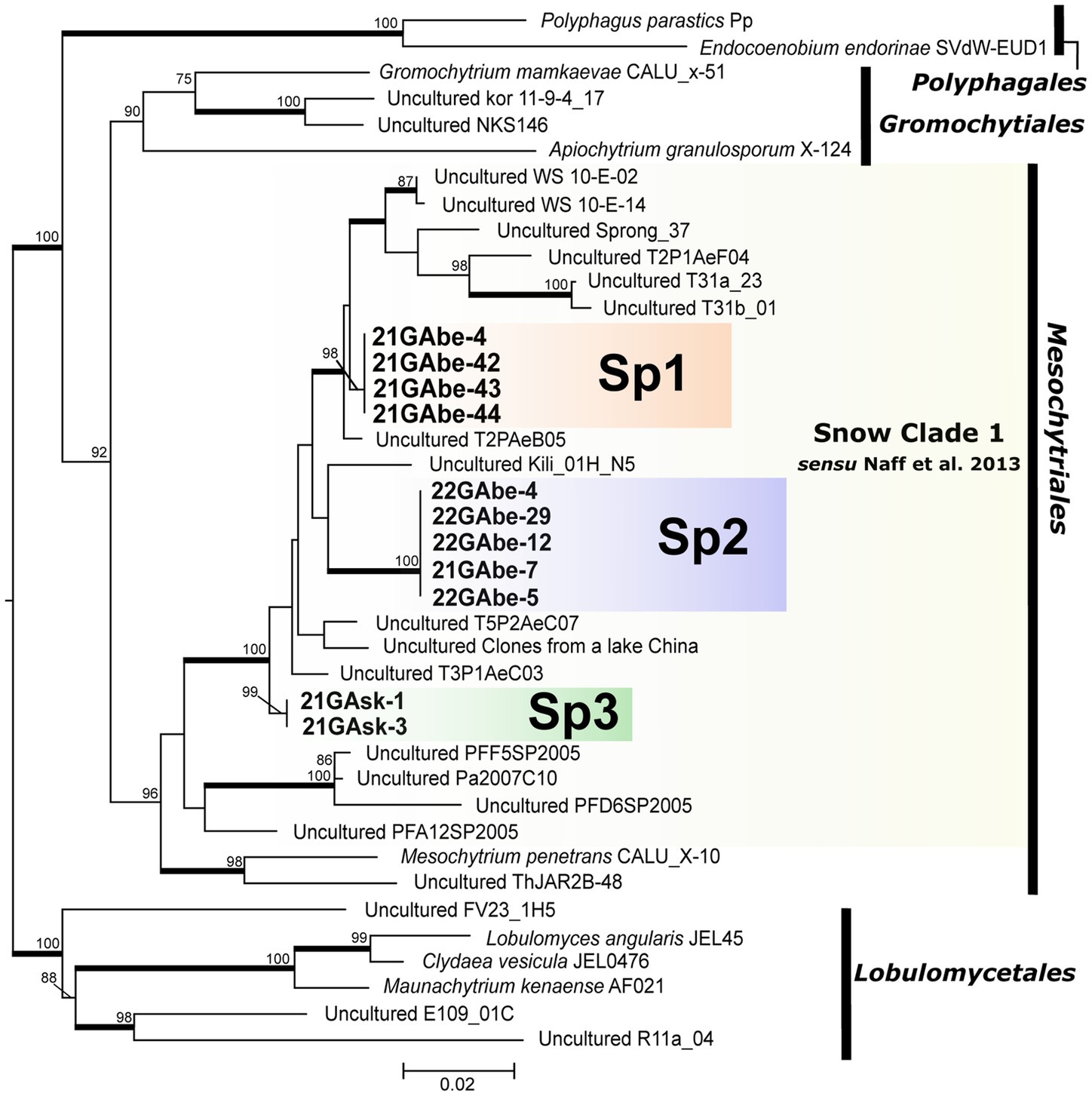
Figure 4. Maximum likelihood molecular phylogenetic tree showing the phylogenetic position of chytrid species belonging to Polyphagales, Gromochytriales, Mesochytriales, and Lobulomycetales using 18S rDNA. Lobulomycetales is the outgroup. Bold letters indicate samples detected in this study. Yellow shading indicates Snow Clade 1 proposed by Naff et al. (2013). Numbers above branches indicate values computed with UFBoot and only UFBoot ≥70% is indicated. Nodes supported by the SH-aLRT ≥95% are highlighted with a bold line. Genbank accession numbers for other sequences used in the phylogenetic analysis, as well as information on the source of isolation (culture strain) and detection point (environmental DNA sequence) are listed in Table 3.
4. Discussion
This study successfully identified the phylogenetic position of three novel parasitic chytrids that infect snow algae in Japan. Mesochytriales, to which the three novel lineages belong, includes only one described species, Mesochytrium penetrans, which infected the green alga Chlorococcum minutum, isolated from a pond in Russia (Gromov et al., 2000; Karpov et al., 2010, 2014). Mesochytriales is one of the most understudied orders in the phylum Chytridiomycota and comprises many environmental DNA sequences (Karpov et al., 2014). Particularly, a group of environmental DNA sequences detected only in snow-related environments has been reported as Snow Clade 1 (Naff et al., 2013). It has been hypothesized that chytrids belonging to Snow Clade 1 spend at least a part of their life in snow and use either abundant snow algae or pollen as the nutritional resource (Naff et al., 2013). In this study, 11 chytrid cells infecting the snow algae Chloromonas spp. were found within Snow Clade 1 as independent lineages. This and earlier sequencing data suggest that chytrids infecting snow algae are phylogenetically diverse and present in various regions of the world. Furthermore, it can be assumed that the lineage composed of snow-derived environmental DNA sequences within Snow Clade 1 are parasitic chytrids infecting snow algae.
Our three species were morphologically distinguished from the previously described species, Chytridium (Chy.) neochlamydococci, parasitic chytrid of the snow alga Chlamydomonas (Chla.) nivalis (=Sanguina nivaloides) (Kobayashi and Okubo, 1954). Chytridium neochlamydococci formed epibiotic and citriform sporangia of 8–10 μm in length, and rhizoids with 2–3 branches from the base, which was originally found from the snow at Ozegahara Moor in Japan (Kobayashi and Okubo, 1954). Contrary to Chy. neochlamydococci, our Sp3 uniquely had sporangia partially embedded in the host cell. The Sp1 and Sp2 differed in other aspects; (i) the Sp1 had egg-shaped sporangia, (ii) the Sp2 had ellipsoidal and larger (10–15 μm in length), (iii) both our species had swollen or rod-shaped rhizoids without branches. The other described chytrid, Chy. f. cryophile (Kol, 1968), a parasite of Chla. nivalis, was not comparable due to the lack of morphological descriptions. Since the host of the chytrid was different from the hosts of our three lineages Chloromonas spp., they were probably different species.
Although parasitic chytrids generally reproduce under aquatic conditions, there are sequences in Snow Clade 1 that have been abundantly detected in soils after the disappearance of snow cover (Freeman et al., 2009). These sequences may have been derived from chytrids in their resting spore stage. In this study, a putative resting-spore stage of chytrids was identified in a snowpack. They were positive for CFW–WGA double staining and had typical structures of resting spores, thick cell walls, and large lipid globules accumulated within the cells, allowing them to survive in soils exposed to high temperatures and desiccation after snowmelt. Likewise, snow algae commonly encyst to form persistent resting spores and adhere to the soil upon melting of the snowpack (Hoham and Duval, 2001; Řezanka et al., 2008; Remias et al., 2010). The resting spores are believed to germinate at the snow-soil interface during the next snowmelt season, swim up to the snow surface, and reproduce (Hoham and Duval, 2001; Hoham and Remias, 2020). This strategy allows them to survive even at unsuitable reproduction times.
Based on these results, we hypothesized that parasitic chytrids infecting snow algae begin to grow after germination of the resting spores of chytrids during the snowmelt season, reproduce for a limited period, and then, rest in soil. Parasitic chytrids that infect algae in lakes are believed to form resting spores at the end of algal blooms, allowing them to survive in the absence of their hosts (Van Donk and Ringelberg, 1983). Germination of the resting spores of parasitic chytrids is also observed in several species within the genus Synchytrium, which infect cucurbitaceous plants. These resting spores germinate when they are exposed to high temperatures and moisture for more than a certain period of time (Raghavendra Rao and Pavgi, 1979). Further studies on parasitic chytrids infecting snow algae are needed to understand their life histories and influence on algal populations.
No phylogenetic information on the resting spores of chytrids was found in this study because single-spore PCR and sequencing were unsuccessful for all 22 cells. Resting spores of chytrids form a thick cell wall, and the dissolution of cell wall using the HotSHOT method may be insufficient. Additionally, as there is only one nucleus in resting spores, the amount of DNA is significantly lower than that in chytrids at the sporangial stage, which contain multiple nuclei. To unify the sporangial and resting spore stages in the same lineage, it is necessary to establish a two-member culture system with the host, and induce this in experiments. This will allow us to determine the consistent life histories of parasitic chytrids that infect snow algae. Furthermore, genome sequencing of cultured strains may provide genetic insights into the evolutionary history of chytrid adaptation to snow environments.
To the best of our knowledge, this study provides the first phylogenetic evidence of parasitic chytrids infecting snow algae in alpine regions. Identification using culture-independent single-spore PCR provided insights into the detailed diversity of these chytrids. However, we could not assess the impact of chytrids on the population dynamics of snow algae and snowpack ecosystems. In this study, the prevalence of chytrid infection in the vegetative cells of Chloromonas spp. in green snow samples was 5.34%. This low prevalence does not necessarily indicate a low impact of chytrids on algal dynamics. In lakes, chytrids significantly suppress algal populations, but prevalence of infection was not always high. There is often a time lag between the spread of chytrids infecting algae in the lake and peak algal bloom (Gsell et al., 2013). Spatial variation in the prevalence of infection could be significant, as reported for a glacier in which more glacial algal cells were infected by chytrids in water-filled holes on the glacier (cryoconite holes) than those on the ice surface (Kobayashi et al., 2023). In alpine snowpacks, the increased water content of snow may also spread chytrids that infect snow algae. Since the beginning of the 21st century, alpine snow cover has been declining globally due to climate change (Hock et al., 2019). Changes in snowfall level due to global warming have been reported in the high mountainous areas of Japan (Kawase et al., 2020). Therefore, in these areas, chytrids infecting snow algae may have increased prevalence and inhibit the albedo-decreasing effects of snow algae.
Our findings highlight the importance of parasitic chytrids that infect snow algae in various regions of the world. Further studies warranted to understand the impact of chytrid infection on snow algal dynamics and predict how their dynamics will be affected by climate change.
Data availability statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/Supplementary material.
Author contributions
HN, KS, and MK designed the study. HN, NT, and MK collected the samples. HN performed microscopic observation and single-spore PCR. HN and KS performed molecular phylogenetic analysis. HN wrote the manuscript with major input from KS, NT, and MK. All authors discussed the results and contributed to the final manuscript.
Funding
This study was supported by a JSPS KAKENHI Grant-in-Aid for Scientific Research (22H03731, 20K21840, and 19H01143).
Acknowledgments
The authors thank Masato Ono, who helped with sampling, and Jun Uetake, Takahiro Segawa, and our colleagues at Yokohama National University and Chiba University for their useful discussions. The authors would like to thank Editage (www.editage.com) for English language editing.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Supplementary material
The Supplementary material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2023.1201230/full#supplementary-material
Footnotes
References
Behnke, A., Bunge, J., Barger, K., Breiner, H.-W., Alla, V., and Stoeck, T. (2006). Microeukaryote community patterns along an O2/H2S gradient in a supersulfidic anoxic fjord (Framvaren, Norway). Appl. Environ. Microbiol. 72, 3626–3636. doi: 10.1128/AEM.72.5.3626-3636.2006
Brown, S. P., Olson, B. J. S. C., and Jumpponen, A. (2015). Fungi and algae co-occur in snow: an issue of shared habitat or algal facilitation of heterotrophs? Arct. Antarct. Alp. Res. 47, 729–749. doi: 10.1657/AAAR0014-071
Capella-Gutiérrez, S., Silla-Martínez, J. M., and Gabaldón, T. (2009). trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25, 1972–1973. doi: 10.1093/bioinformatics/btp348
Fiołka, M. J., Takeuchi, N., Sofińska-Chmiel, W., Wójcik-Mieszawska, S., Irvine-Fynn, T., and Edwards, A. (2021). Morphological and spectroscopic analysis of snow and glacier algae and their parasitic fungi on different glaciers of Svalbard. Sci. Rep. 11:21785. doi: 10.1038/s41598-021-01211-8
Freeman, K. R., Martin, A. P., Karki, D., Lynch, R. C., Mitter, M. S., Meyer, A. F., et al. (2009). Evidence that chytrids dominate fungal communities in high-elevation soils. Proc. Natl. Acad. Sci. U. S. A. 106, 18315–18320. doi: 10.1073/pnas.0907303106
Frenken, T., Alacid, E., Berger, S. A., Bourne, E. C., Gerphagnon, M., Grossart, H.-P., et al. (2017). Integrating chytrid fungal parasites into plankton ecology: research gaps and needs. Environ. Microbiol. 19, 3802–3822. doi: 10.1111/1462-2920.13827
Ganey, G. Q., Loso, M. G., Burgess, A. B., and Dial, R. J. (2017). The role of microbes in snowmelt and radiative forcing on an Alaskan icefield. Nat. Geosci. 10, 754–759. doi: 10.1038/ngeo3027
Gardes, M., and Bruns, T. D. (1993). ITS primers with enhanced specificity for basidiomycetes – application to the identification of mycorrhizae and rusts. Mol. Ecol. 2, 113–118. doi: 10.1111/j.1365-294X.1993.tb00005.x
Genitsaris, S., Kormas, K. A., and Moustaka-Gouni, M. (2009). Microscopic eukaryotes living in a dying Lake (Lake Koronia, Greece). FEMS Microbiol. Ecol. 69, 75–83. doi: 10.1111/j.1574-6941.2009.00686.x
Gromov, B. V., Mamkaeva, K. A., and Pljusch, A. V. (2000). Mesochytrium penetrans gen. et sp. nov. (Chytridiales) – a parasite of the green alga Chlorococcum minutum (Chlorococcales), with an unusual behaviour of the sporangia. Nova_Hedwigia 71, 151–160. doi: 10.1127/nova/71/2000/151
Gsell, A. S., De Senerpont Domis, L. N., Naus-Wiezer, S. M. H., Helmsing, N. R., Van Donk, E., and Ibelings, B. W. (2013). Spatiotemporal variation in the distribution of chytrid parasites in diatom host populations. Freshw. Biol. 58, 523–537. doi: 10.1111/j.1365-2427.2012.02786.x
Hock, R., Rasul, G., Adler, C., Cáceres, B., Gruber, S., Hirabayashi, Y., et al. (2019). “High Mountain Areas,” in IPCC Special Report on the Ocean and Cryosphere in a Changing Climate eds H. -O. Pörtner, D. C. Roberts, V. Masson-Delmotte, P. Zhai, M. Tignor, E. Poloczanska, et al. (Cambridge: Cambridge University Press), 131–202. doi: 10.1017/9781009157964.004
Hoham, R. W., and Duval, B. (2001). “Microbial ecology of snow and freshwater ice with emphasis on snow algae” in Snow ecology: an interdisciplinary examination of snow-covered ecosystems. eds. H. G. Jones, J. W. Pomeroy, D. A. Walker, and R. W. Hoham (Cambridge: Cambridge University Press), 168–288.
Hoham, R. W., and Remias, D. (2020). Snow and glacial algae: a review1. J. Phycol. 56, 264–282. doi: 10.1111/jpy.12952
Hotaling, S., Lutz, S., Dial, R. J., Anesio, A. M., Benning, L. G., Fountain, A. G., et al. (2021). Biological albedo reduction on ice sheets, glaciers, and snowfields. Earth Sci. Rev. 220:103728. doi: 10.1016/j.earscirev.2021.103728
Ishida, S., Nozaki, D., Grossart, H.-P., and Kagami, M. (2015). Novel basal, fungal lineages from freshwater phytoplankton and lake samples. Environ. Microbiol. Rep. 7, 435–441. doi: 10.1111/1758-2229.12268
Jobard, M., Rasconi, S., Solinhac, L., Cauchie, H.-M., and Sime-Ngando, T. (2012). Molecular and morphological diversity of fungi and the associated functions in three European nearby lakes: true Fungi (Eumycota) in pelagic freshwaters. Environ. Microbiol. 14, 2480–2494. doi: 10.1111/j.1462-2920.2012.02771.x
Kagami, M., de Bruin, A., Ibelings, B. W., and Van Donk, E. (2007). Parasitic chytrids: their effects on phytoplankton communities and food-web dynamics. Hydrobiologia 578, 113–129. doi: 10.1007/s10750-006-0438-z
Kagami, M., Seto, K., Nozaki, D., Nakamura, T., Wakana, H., and Wurzbacher, C. (2021). Single dominant diatom can host diverse parasitic fungi with different degree of host specificity. Limnol. Oceanogr. 66, 667–677. doi: 10.1002/lno.11631
Kariya, Y. (2005). Holocene landscape evolution of a nivation hollow on Gassan volcano, northern Japan. Catena 62, 57–76. doi: 10.1016/j.catena.2005.02.004
Karpov, S. A., Kobseva, A. A., Mamkaeva, M. A., Mamkaeva, K. A., Mikhailov, K. V., Mirzaeva, G. S., et al. (2014). Gromochytrium mamkaevae gen. & sp. nov. and two new orders: Gromochytriales and Mesochytriales (Chytridiomycetes). Persoonia 32, 115–126. doi: 10.3767/003158514X680234
Karpov, S. A., Letcher, P. M., Mamkaeva, M. A., and Mamkaeva, K. A. (2010). Phylogenetic position of the genus Mesochytrium (Chytridiomycota) based on zoospore ultrastructure and sequences from the 18S and 28S rRNA gene. Nova_Hedwigia 90, 81–94. doi: 10.1127/0029-5035/2010/0090-0081
Karpov, S. A., Lуpez-Garcia, P., Mamkaeva, M. A., Vishnyakov, A. E., and Moreira, D. (2016). Chytridiomycete Polyphagus parasiticus: molecular phylogeny supports the erection of a new chytridiomycete order. Mikologiya 50, 362–366.
Karpov, S. A., Moreira, D., Mamkaeva, M. A., Popova, O. V., Aleoshin, V. V., and López-García, P. (2019). New member of Gromochytriales (Chytridiomycetes)— Apiochytrium granulosporum nov. gen. et sp. J. Eukaryotic Microbiol. 66, 582–591. doi: 10.1111/jeu.12702
Katoh, K., Rozewicki, J., and Yamada, K. D. (2019). MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief. Bioinform. 20, 1160–1166. doi: 10.1093/bib/bbx108
Kawase, H., Yamazaki, T., Sugimoto, S., Sasai, T., Ito, R., Hamada, T., et al. (2020). Changes in extremely heavy and light snow-cover winters due to global warming over high mountainous areas in Central Japan. Prog Earth Planet Sci 7:10. doi: 10.1186/s40645-020-0322-x
Klawonn, I., Dunker, S., Kagami, M., Grossart, H.-P., and Van den Wyngaert, S. (2023). Intercomparison of two fluorescent dyes to visualize parasitic fungi (Chytridiomycota) on phytoplankton. Microb. Ecol. 85, 9–23. doi: 10.1007/s00248-021-01893-7
Kobayashi, Y., and Okubo, M. (1954). “Studies on the aquatic fungi of the Ozegahara Moor,” in Scientific research of the Ozegahara Moor, ed. Scientific Researchers of the Ozegahara Moor, Japan Society for the Promotion of Science, 561–584.
Kobayashi, K., Takeuchi, N., and Kagami, M. (2023). High prevalence of parasitic chytrids infection of glacier algae in cryoconite holes in Alaska. Sci. Rep. 13:3973. doi: 10.1038/s41598-023-30721-w
Kol, E. (1968). “Kryobiologie. Biologie und Limnologie des Schneesund Eises” in Die Binnengewa¨sser, Band XXIV. ed. I. Kryovegetation (Stuttgart: Schweizerbart’sche Verlagsbuchhandlung)
Krug, L., Erlacher, A., Markut, K., Berg, G., and Cernava, T. (2020). The microbiome of alpine snow algae shows a specific inter-kingdom connectivity and algae-bacteria interactions with supportive capacities. ISME J. 14, 2197–2210. doi: 10.1038/s41396-020-0677-4
Lazarus, K. L., and James, T. Y. (2015). Surveying the biodiversity of the Cryptomycota using a targeted PCR approach. Fungal Ecol. 14, 62–70. doi: 10.1016/j.funeco.2014.11.004
Lefèvre, E., Roussel, B., Amblard, C., and Sime-Ngando, T. (2008). The molecular diversity of freshwater picoeukaryotes reveals high occurrence of putative parasitoids in the plankton. PLoS One 3:e2324. doi: 10.1371/journal.pone.0002324
Luo, W., Kotut, K., and Krienitz, L. (2013). Hidden diversity of eukaryotic plankton in the soda Lake Nakuru, Kenya, during a phase of low salinity revealed by a SSU rRNA gene clone library. Hydrobiologia 702, 95–103. doi: 10.1007/s10750-012-1310-y
Lutz, S., Anesio, A. M., Raiswell, R., Edwards, A., Newton, R. J., Gill, F., et al. (2016). The biogeography of red snow microbiomes and their role in melting arctic glaciers. Nat. Commun. 7:11968. doi: 10.1038/ncomms11968
Matsuzaki, R., Kawai-Toyooka, H., Hara, Y., and Nozaki, H. (2015). Revisiting the taxonomic significance of aplanozygote morphologies of two cosmopolitan snow species of the genus Chloromonas (Volvocales, Chlorophyceae). Phycologia 54, 491–502. doi: 10.2216/15-33.1
Matsuzaki, R., Nozaki, H., and Kawachi, M. (2018). Taxonomic revision of Chloromonas nivalis (Volvocales, Chlorophyceae) strains, with the new description of two snow-inhabiting Chloromonas species. PLoS One 13:e0193603. doi: 10.1371/journal.pone.0193603
Matsuzaki, R., Nozaki, H., Takeuchi, N., Hara, Y., and Kawachi, M. (2019). Taxonomic re-examination of “Chloromonas nivalis (Volvocales, Chlorophyceae) zygotes” from Japan and description of C. muramotoi sp. nov. PLoS One 14:e0210986. doi: 10.1371/journal.pone.0210986
Naff, C. S., Darcy, J. L., and Schmidt, S. K. (2013). Phylogeny and biogeography of an uncultured clade of snow chytrids. Environ. Microbiol. 15, 2672–2680. doi: 10.1111/1462-2920.12116
Nakashima, T., Uetake, J., Segawa, T., Procházková, L., Tsushima, A., and Takeuchi, N. (2021). Spatial and temporal variations in pigment and species compositions of snow algae on Mt. Tateyama in Toyama prefecture, Japan. Front. Plant Sci. 12:689119. doi: 10.3389/fpls.2021.689119
O’Donnell, K., Cigelnik, E., and Benny, G. L. (1998). Phylogenetic relationships among the Harpellales and Kickxellales. Mycologia 90, 624–639. doi: 10.1080/00275514.1998.12026952
Ono, M., Takeuchi, N., and Zawierucha, K. (2021). Snow algae blooms are beneficial for microinvertebrates assemblages (Tardigrada and Rotifera) on seasonal snow patches in Japan. Sci. Rep. 11:5973. doi: 10.1038/s41598-021-85462-5
Procházková, L., Leya, T., Křížková, H., and Nedbalová, L. (2019). Sanguina nivaloides and Sanguina aurantia gen. et spp. nov. (Chlorophyta): the taxonomy, phylogeny, biogeography and ecology of two newly recognised algae causing red and orange snow. FEMS Microbiol. Ecol. 95:fiz064. doi: 10.1093/femsec/fiz064
Raghavendra Rao, N. N., and Pavgi, M. S. (1979). Germination of resting spores in Synchytrium species parasitic on cucurbitaceae. Mycopathologia 69, 3–10. doi: 10.1007/BF00428598
R Core Team (2022). R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria. Available at: https://www.R-project.org/
Remias, D., Karsten, U., Lütz, C., and Leya, T. (2010). Physiological and morphological processes in the alpine snow alga Chloromonas nivalis (Chlorophyceae) during cyst formation. Protoplasma 243, 73–86. doi: 10.1007/s00709-010-0123-y
Řezanka, T., Nedbalová, L., Sigler, K., and Cepák, V. (2008). Identification of astaxanthin diglucoside diesters from snow alga Chlamydomonas nivalis by liquid chromatography–atmospheric pressure chemical ionization mass spectrometry. Phytochemistry 69, 479–490. doi: 10.1016/j.phytochem.2007.06.025
Schmidt, S. K., Naff, C. S., and Lynch, R. C. (2012). Fungal communities at the edge: ecological lessons from high alpine fungi. Fungal Ecol. 5, 443–452. doi: 10.1016/j.funeco.2011.10.005
Segawa, T., Matsuzaki, R., Takeuchi, N., Akiyoshi, A., Navarro, F., Sugiyama, S., et al. (2018). Bipolar dispersal of red-snow algae. Nat. Commun. 9:3094. doi: 10.1038/s41467-018-05521-w
Seto, K., Kagami, M., and Degawa, Y. (2017). Phylogenetic position of parasitic chytrids on diatoms: characterization of a novel clade in chytridiomycota. J. Eukaryot. Microbiol. 64, 383–393. doi: 10.1111/jeu.12373
Simmons, D. R., James, T. Y., Meyer, A. F., and Longcore, J. E. (2009). Lobulomycetales, a new order in the Chytridiomycota. Mycol. Res. 113, 450–460. doi: 10.1016/j.mycres.2008.11.019
Takeuchi, N. (2001). The altitudinal distribution of snow algae on an Alaska glacier (Gulkana Glacier in the Alaska Range). Hydrol. Process. 15, 3447–3459. doi: 10.1002/hyp.1040
Terashima, M., Umezawa, K., Mori, S., Kojima, H., and Fukui, M. (2017). Microbial community analysis of colored snow from an alpine snowfield in Northern Japan reveals the prevalence of Betaproteobacteria with snow algae. Front. Microbiol. 8:1481. doi: 10.3389/fmicb.2017.01481
Thomas, M. C., Selinger, L. B., and Inglis, G. D. (2012). Seasonal diversity of planktonic protists in Southwestern Alberta Rivers over a 1-year period as revealed by terminal restriction fragment length polymorphism and 18S rRNA gene library analyses. Appl. Environ. Microbiol. 78, 5653–5660. doi: 10.1128/AEM.00237-12
Trifinopoulos, J., Nguyen, L. T., von Haeseler, A., and Minh, B. Q. (2016). W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res. 44, W232–W235. doi: 10.1093/nar/gkw256
Truett, G. E., Heeger, P., Mynatt, R. L., Truett, A. A., Walker, J. A., and Warman, M. L. (2000). Preparation of PCR-quality mouse genomic DNA with hot sodium hydroxide and tris (HotSHOT). BioTechniques 29, 52–54. doi: 10.2144/00291bm09
Van den Wyngaert, S., Ganzert, L., Seto, K., Rojas-Jimenez, K., Agha, R., Berger, S. A., et al. (2022). Seasonality of parasitic and saprotrophic zoosporic fungi: linking sequence data to ecological traits. ISME J. 16, 2242–2254. doi: 10.1038/s41396-022-01267-y
Van den Wyngaert, S., Rojas-Jimenez, K., Seto, K., Kagami, M., and Grossart, H.-P. (2018). Diversity and hidden host specificity of chytrids infecting colonial volvocacean algae. J. Eukaryot. Microbiol. 65, 870–881. doi: 10.1111/jeu.12632
Van Donk, E., and Ringelberg, J. (1983). The effect of fungal parasitism on the succession of diatoms in Lake Maarsseveen I (The Netherlands). Freshw. Biol. 13, 241–251. doi: 10.1111/j.1365-2427.1983.tb00674.x
Vilgalys, R., and Hester, M. (1990). Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 172, 4238–4246. doi: 10.1128/jb.172.8.4238-4246.1990
White, T. J., Bruns, T., Lee, S., and Taylor, J. (1990). “Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics” in PCR protocols: a guide to methods and applications. eds. M. A. Innis, D. H. Gelfand, J. J. Sninsky, and T. J. White (New York: Academic Press), 315–322.
Wurzbacher, C., Larsson, E., Bengtsson-Palme, J., van den Wyngaert, S., Svantesson, S., Kristiansson, E., et al. (2019). Introducing ribosomal tandem repeat barcoding for fungi. Mol. Ecol. Resour. 19, 118–127. doi: 10.1111/1755-0998.12944
Keywords: snow algae, fungi, chytrid, host–parasite interactions, snowpacks, polar and alpine ecosystems
Citation: Nakanishi H, Seto K, Takeuchi N and Kagami M (2023) Novel parasitic chytrids infecting snow algae in an alpine snow ecosystem in Japan. Front. Microbiol. 14:1201230. doi: 10.3389/fmicb.2023.1201230
Edited by:
Katsumi Matsuura, Tokyo Metropolitan University, JapanReviewed by:
Daniel Remias, University of Salzburg, AustriaTamotsu Hoshino, Hachinohe Institute of Technology, Japan
Copyright © 2023 Nakanishi, Seto, Takeuchi and Kagami. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hiroaki Nakanishi, bmFrYW5pc2hpLWhpcm9ha2ktcHlAeW51Lmpw; Maiko Kagami, a2FnYW1pLW1haWtvLWJkQHludS5hYy5qcA==
 Hiroaki Nakanishi
Hiroaki Nakanishi Kensuke Seto
Kensuke Seto Nozomu Takeuchi
Nozomu Takeuchi Maiko Kagami
Maiko Kagami