- 1Faculty of Medicine, University of Khartoum, Khartoum, Sudan
- 2College of Medicine, Princess Nourah Bint Abdulrahman University, Riyadh, Saudi Arabia
- 3Institut du Cerveau, INSERM, CNRS, Sorbonne Université, Paris, France
- 4Division of Pediatric Neurology, Department of Pediatrics, College of Medicine, King Saud University, Riyadh, Saudi Arabia
- 5Department of Biochemistry, Faculty of Medicine, National University, Khartoum, Sudan
- 6Ecole Pratique des Hautes Etudes, EPHE, PSL Research University, Paris, France
- 7Department of Molecular Biology, Institute of Endemic Diseases, University of Khartoum, Khartoum, Sudan
- 8Department of Neurology and Epileptology, Hertie Institute for Clinical Brain Research, Tuebingen, Germany
- 9APHP, Pitié-Salpêtrière Hospital, Department of genetics, Paris, France
Background: Arginases catalyze the last step in the urea cycle. Hyperargininemia, a rare autosomal-recessive disorder of the urea cycle, presents after the first year of age with regression of milestones and evolves gradually into progressive spastic quadriplegia and cognitive dysfunction. Genetic studies reported various mutations in the ARG1 gene that resulted in hyperargininemia due to a complete or partial loss of arginase activity.
Case Presentation: Five patients from an extended highly consanguineous Sudanese family presented with regression of the acquired milestones, spastic quadriplegia, and mental retardation. The disease onset ranged from 1 to 3 years of age. Two patients had epileptic seizures and one patient had stereotypic clapping. Genetic testing using whole-exome sequencing, done for the patients and a healthy parent, confirmed the presence of a homozygous novel missense variant in the ARG1 gene [GRCh37 (NM_001244438.1): exon 4: g.131902487T>A, c.458T>A, p.(Val153Glu)]. The variant was predicted pathogenic by five algorithms and affected a highly conserved amino acid located in the protein domain ureohydrolase, arginase subgroup. Sanger sequencing of 13 sampled family members revealed complete co-segregation between the variant and the disease distribution in the family in line with an autosomal-recessive mode of inheritance. Biochemical analysis confirmed hyperargininemia in five patients.
Conclusion: This study reports the first Sudanese family with ARG1 mutation. The reported variant is a loss-of-function missense mutation. Its pathogenicity is strongly supported by the clinical phenotype, the computational functional impact prediction, the complete co-segregation with the disease, and the biochemical assessment.
Background
Arginases (EC 3.5.3.1) are proteins involved in the last reaction of the urea cycle in which they catalyze the hydrolysis reaction of L-arginine to form urea and ornithine. There are two isoforms of the arginase enzyme (arginases 1 and 2) encoded by the ARG1 [*608313] and ARG2 [*107830] genes, respectively. Arginase one predominates in the liver, constituting 98% of the hepatic arginase activity. Its deficiency results in hyperargininemia/arginase deficiency [#207800], which is an autosomal-recessive (AR) inborn error of metabolism. Arginase deficiency is the least common of all urea cycle disorders. The incidence of hyperargininemia has been estimated to vary between 1:350,000 and 1:1,000,000 (1). There is scarcely any data available about the incidence/prevalence in Sudan and other Sub-Saharan African countries.
Most commonly, the condition manifests clinically between 1 and 3 years of age, although adult-onset arginase deficiency has been occasionally reported (2). Classically, patients suffer from developmental delay and regression of the acquired milestones. The condition may then evolve gradually into spastic quadriparesis and intellectual disability if left untreated (3). The condition can be almost non-progressive, which can sometimes lead to confusion with cerebral palsy (4). Biochemically, there is hyperargininemia with periodic hyperammonemia, although, in contrast to other urea cycle disorders, neonatal hyperammonemia and its clinical features are relatively uncommon in arginase deficiency. Affected children are frequently asymptomatic at birth and in their early childhood.
The ARG1 gene is located on chromosome six mapping to the cytogenetic location 6q23.2. The gene harbors eight exons. Hyperargininemia is associated with pathogenic variants in both homozygous and compound heterozygous states, with no obvious genotype/phenotype correlation. The first case in whom the genetic basis of argininemia was characterized was a Japanese patient with compound heterozygous mutations (5). A study published in 2018 by Diez-Fernandez et al. carried out sequence analysis compiling 66 published and novel variants of the ARG1 gene and found that the most encountered category was point mutations (missense, non-sense, and splicing), although other structural alterations were also found. Missense variants were the most common mutation type distributed all over the coding region of the gene (6–8). Genetic studies of hyperargininemia are extremely rare in Sub-Saharan Africa and are non-existent in Sudan. In this study, we are reporting the first case from Sudan with ARG1 mutation. A novel homozygous missense mutation in the ARG1 gene was identified in five spastic quadriplegic patients from an extended highly consanguineous Sudanese family.
Case Presentation
Five affected patients from three branches of family F15 are reported in this study. The patients had a history of regression of milestones following a normal early development till the age at onset, which ranged from 1 to 3 years. The clinical picture was dominated by spastic quadriplegia complicated by cognitive impairment and sphincter disturbances in the majority of patients. Visual and hearing assessments were normal, except in one patient who had conductive deafness in the right ear (patient 138). Despite the remarkable homogeneity of the clinical presentation in all branches, two patients had epileptic seizures (patients 138 and 142) and one patient (patient 138) had stereotypic clapping (Table 1). Extrapyramidal and cerebellar symptoms and signs were not detected in all five patients. No sensory nervous system abnormality was detected, although assessment could not be done properly. Brain images (MRI) were normal in all patients. The disease had a progressive nature, as expected, with variable severity. The most severe course was observed in one patient who died in the first half of the second decade.
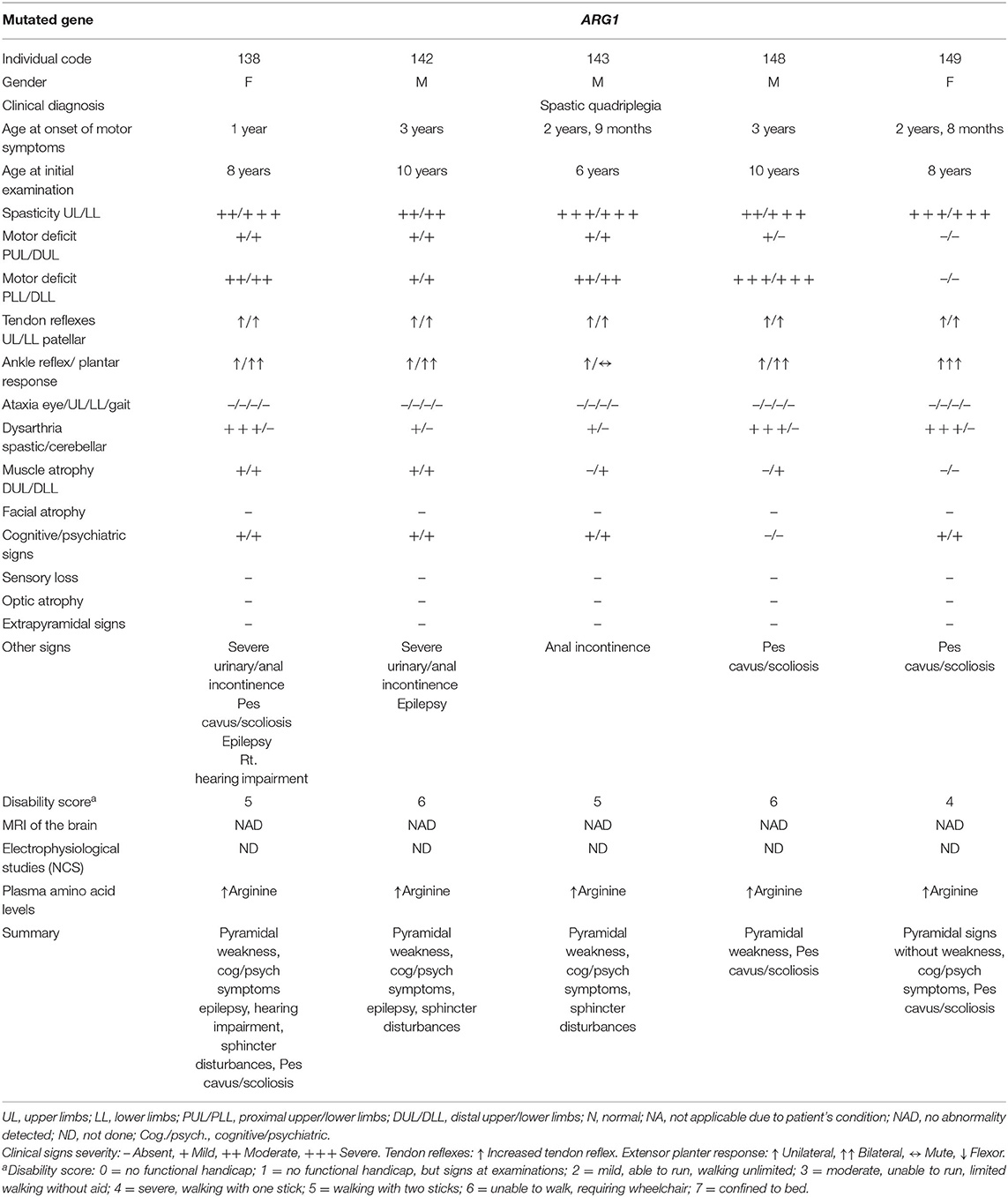
Table 1. Clinical data of five patients from family F15 with hyperargininemia due to a mutation in the ARG1 gene.
Materials and Methods
DNA Extraction
DNA was extracted from saliva collected from five patients and nine healthy parents and relatives using Oragene.Discover® DNA collection kits (DNA Genotek Inc.®, Ottawa, ON, Canada) according to the manufacturer's prepIT.L2P manual protocol.
Whole-Exome Sequencing
Whole-exome sequencing (WES) was conducted for all five patients and one healthy parent. WES was performed on 150-bp paired-end reads using the Hiseq property of the NextSeq-500 sequencer (Illumina®, San Diego, CA, USA). Genomics Workbench (CLC Bio®, Aarhus, Denmark) was used for quality control and variant calling. Both probability-based and quality-based algorithms were used to identify single nucleotide variants (SNVs) and insertions/deletions (Indels). For gene/variant prioritization, a minimum depth of 30 × and 1% as the minor allele frequency cutoff were selected. Nonsense, frameshift, splice site, and predicted pathogenic missense variants were filtered according to the autosomal-recessive mode of inheritance predicted from the pedigree analysis.
Sanger Sequencing
Sanger sequencing using the BIGDYE chemistry on an ABI3730 sequencer (Applied Biosystems®) was carried out on 13 affected and healthy family members to validate the WES results and confirm co-segregation between the pathogenic allele and the disease within the family. Chromas Lite® software (Technelysium®, South Brisbane, QLD, Australia) and Seqscape® (Applied Biosystems®) were utilized for sequence visualization and analysis.
Results
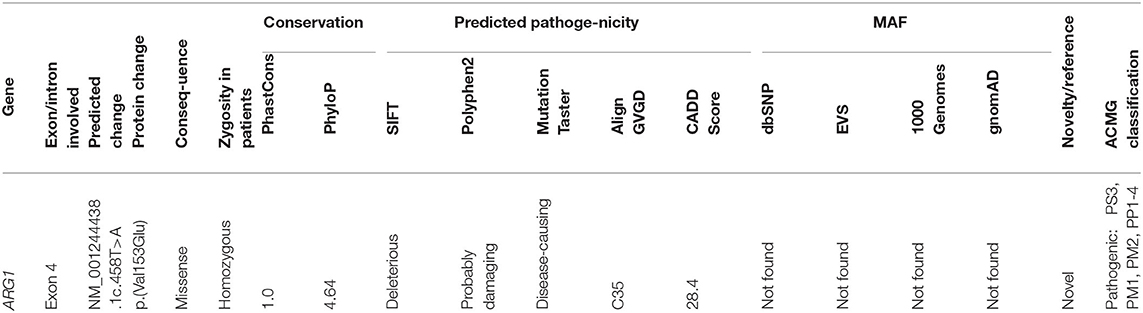
After filtering out the common and non-coding variants, WES data analysis detected only one homozygous variant that was shared by all five patients, which was heterozygous in the healthy parent. This was a novel missense transversion variant [GRCh37 (NM_001244438.1): exon 4: g.131902487T>A, c.458T>A, p.(Val153Glu)] in the ARG1 gene. The mutation was predicted pathogenic by five algorithms (SIFT = deleterious, Polyphen2 = probable damaging, mutation taster = disease causing, align GVGD = C35, and CADD score = 28.4) (Table 2). The variant was absent from gnomAD or dbSNP. Sanger sequencing revealed complete co-segregation between the pathogenic allele and the disease distribution, being a homozygous mutant in all the five patients and a homozygous reference or heterozygous carriers in nine related healthy individuals, including all parents who were available for sampling. The variant resulted in the replacement of the highly conserved (12 species) hydrophobic amino acid valine, located in the protein domain ureohydrolase, arginase subgroup, by the acidic amino acid glutamate (Figure 1). The moderately radical physiochemical difference [Grantham distance of 121 (0–215)] exists between the two amino acids. No other homozygous or compound heterozygous variants were found in genes known to be involved in other neurological disorders.
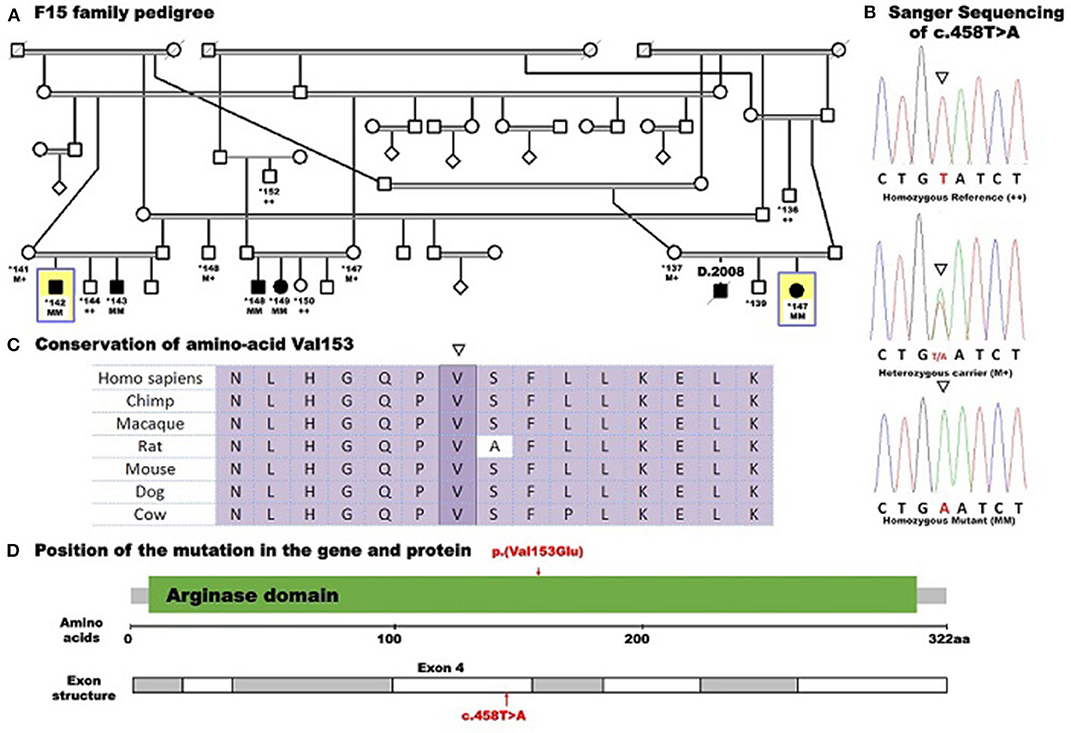
Figure 1. Pedigree of family F15 caused by a missense mutation in ARG1 segregating with the disease distribution in the whole family presenting with spastic tetraplegia and mental retardation. Whole-exome sequencing was performed for five affected individuals (138, 142, 143, 148, and 149) and the mother (137) of patient 138. Example Sanger sequencing of the mutation in a patient (homozygous mutant), a heterozygous carrier, and a control homozygous reference allele with conserved amino acid sequence is shown. Pedigree symbols: asterisk indicates sampled individual; yellow square indicates the index patient. Genotype symbols: ++ Homozygous reference genotype; M+ Heterozygous genotype; MM Homozygous mutant genotype. Others are standard medical pedigree symbols.
Biochemical Assessment
Biochemical assessment of the amino acid levels in the serum of all the five patients showed an increased level of arginine. A second confirmatory test was performed for the proband only and showed an increased serum arginine level, confirming the first test.
Counseling and Management
The family received genetic counseling, which included discussions with the senior family members (parents and grandparents) about consanguinity to provide the family a better understanding of the nature of their genetic disorders, increase their awareness about the role played by consanguinity in inheritance, and to eliminate any misperceptions. Good response to counseling regarding consanguinity was observed, particularly in the female members of the family, and a new generation is non-consanguineously married.
The patients were put on physiotherapy, with epilepsy treatment for the patients with seizures. However, most of the time, the family was faced with limited financial capabilities, which hindered their compliance with the follow-up and the physiotherapy.
A specialist was consulted for dietary modulation to start the patients on a restricted protein diet and arginine-free amino acid supplements. Two obstacles faced the use of special formulas with dietary restrictions: the availability of the formulas and the finance. The required special formulas were not available in Sudan, and attempts to obtain them from a neighboring country were not successful. In addition, the family was unable to afford either the cost of the formulas regularly from other countries or the necessary tests used for the monitoring of patients when started on treatment.
Discussion and Conclusion
Arginase 1 deficiency can lead to a devastating neurological and cognitive dysfunction in children, though it is a potentially treatable condition (9). The age at onset was typical for all patients, and the clinical features of the five reported Sudanese patients, which included progressive spastic quadriplegia, regression of milestones, and epilepsy, are similar to what was reported about the presentation of a loss-of-function mutation in the arginase 1 enzyme (7, 10, 11). However, the five Sudanese patients maintained normal head circumference and normal physical growth despite the variable degrees of mental impairment witnessed in all of them. These features are different from what was reported from many Japanese and Chinese patients in whom microcephaly and poor physical growth were the major complicating features (1, 5, 12). The conservation of the nucleotide affected by the mutation reported in this study, (c.458T>A), as evidenced by two elevated conservation scores (PhastCons and PhyloP), and the prediction of pathogenicity by five algorithms (SIFT, Polyphen2, mutation taster and align GVGD, and CADD score) supported the significance of the variant which was classified as pathogenic according to the American College of Medical Genetics and Genomics (ACMG) classification (Table 2). The mutation (c.458T>A) is located in exon 4, and its pathogenic effect is in line with studies which have revealed missense mutations in certain conserved regions to be pathologic. This was further supported by a sequence analysis study which demonstrated clustering of the pathogenic missense variants in exons 1, 4, and 7. The involvement of these conserved regions in the catalytic activities of the enzyme makes mutations in these sequences likely to be pathogenic. This does not apply to truncating mutations that occur randomly along the gene, leading to non-sense-mediated protein decay and consequent hyperargininemia by complete loss of function (8, 13).
The high conservation of the mutated hydrophobic amino acid valine over mammalian species, its replacement by the acidic glutamate with radical Grantham distance existing between the amino acids, and the location of the mutated amino acid in the ureohydrolase domain, arginase subgroup further favor the pathogenicity of the change.
In the absence of other evidence of involvement of any other variants, the matching of the clinical phenotype and the biochemical profile of the patients to the published data, and the complete co-segregation of this variant with the disease, we considered the ARG1 mutation as causative in this family.
In the Middle East region, there are some reported cases of hyperargininemia from Saudi Arabia, Palestine (14), and Bahrain (15). However, studies from Sub-Saharan Africa are extremely rare. This is the first study from Sudan to describe a novel missense mutation in the ARG1 gene causing hyperargininemia, spastic quadriplegia, and cognitive dysfunction. Little is known about the incidence and the prevalence of the disease in Sudan, raising the need for more studies.
The Sudanese population has one of the highest overall consanguinity rates in the world with the highest first cousin/double first cousin marriages in the Arabic world (16, 17). Increasing evidence of augmenting AR inherited disorders including hyperargininemia is accumulating despite the scarcity of the statistics and the insufficiency of data about most of the genetic disorders in Sudan (18–22). This necessitates further studies to be conducted and campaigns to be launched in order to increase public awareness about the impact of consanguinity and its contribution to the shaping of the genetics of the individuals, families, and populations.
The lack of neonatal metabolic screening programs in Sudan leads to delayed diagnosis with the development of several neurological complications that include loss of sphincter control, inability to walk, and intellectual disability, which could have been avoided by early diagnosis and proper management (23, 24).
However, this study also highlighted deeper issues in the provision of the required care for patients with metabolic disorders in a limited-resource country like Sudan. Consequently, there is a necessity to conduct a larger-scale research project exploring some treatable metabolic disorders in Sudan in order to collect the required data that will be the basis for a national newborn screening program for these disorders. It draws attention to the importance of establishing specialized centers for metabolic disorders, which would increase the opportunity for early diagnosis and provide all the requirements for patients' care at an affordable cost. This will efficiently help decrease the disability as a consequence of potentially treatable causes in Sudanese children, especially with the advent of more advanced options of therapy like rescue mRNA therapy in addition to the existing treatment options (25, 26).
Data Availability Statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found at: https://www.ncbi.nlm.nih.gov/, VCV000916528.1.
Ethics Statement
The studies involving human participants were reviewed and approved by The Ethical Committee of Medical Campus, University of Khartoum, Sudan according to the recommendations of the Helsinki Declaration. Written informed consent to participate in this study was provided by the participants' legal guardian/next of kin. Written informed consent was obtained from the individual(s) and/or minor(s)' legal guardian/next of kin for participation in the study and for the publication of any potentially identifiable images or data included in this article.
Author Contributions
LE, GS, AEA, AB, ME, and MI formulated and designed the study and contributed to the interpretation of data. AB, GS, AEA, ME, and MI granted funds and participated in manuscript editing, review, and critique. LE, IM, AH, ME, MS, AY, RS, MK, and ASIA contributed to the clinical, radiologic, and biochemical data collection and interpretation. LE, GS, MK, and AY contributed to the bioinformatic analysis. All authors read the manuscript, assented to be submitted, and accepted and agreed to be responsible for all aspects of this work.
Funding
This study was funded by the European Union through the H2020 program (SOLVE-RD to GS) and the French ASL-HSP Association (to GS). LE has been granted the scholarship from Campus France, University of Khartoum and the Sudanese Ministry of Higher Education. MS was supported by Researchers Supporting Project Number RSP-2020/38, King Saud University, Riyadh, Saudi Arabia.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank the French Embassy in Sudan, Campus France, University of Khartoum, and the Sudan Ministry of Higher Education for the scholarship (to LE), the European Union through the H2020 program (SOLVE-RD to GS), and the French ASL-HSP Association (to GS).
References
1. Wong D, Cederbaum S, Crombez EA. Arginase deficiency. In: Pagon RA, Adam MP, Ardinger HH, Bird TD, Dolan CR, Fong C-T, et al. editors. GeneReviews(®) [Internet]. Seattle, WA: University of Washington, Seattle (1993) Available online at: http://www.ncbi.nlm.nih.gov/books/NBK1159/ (accessed Jan 22, 2015).
2. Cowley DM, Bowling FG, McGill JJ, Dongen J van, Morris D. Adult-onset arginase deficiency. J Inherit Metab Dis. (1998) 21:677–8. doi: 10.1023/A:1005492819527
3. Scaglia F, Lee B. Clinical, biochemical, and molecular spectrum of hyperargininemia due to arginase I deficiency. Am J Med Genet C Semin Med Genet. (2006) 142C:113–20. doi: 10.1002/ajmg.c.30091
4. Scheuerle AE, Mcvie R, Beaudet AL, Shapira SK. Arginase deficiency presenting as cerebral palsy. Pediatrics. (1993) 91:995–6.
5. Haraguchi Y, Aparicio JM, Takiguchi M, Akaboshi I, Yoshino M, Mori M, et al. Molecular basis of argininemia. Identification of two discrete frame-shift deletions in the liver-type arginase gene. J Clin Invest. (1990) 86:347–50. doi: 10.1172/JCI114707
6. Diez-Fernandez C, Rüfenacht V, Gemperle C, Fingerhut R, Häberle J. Mutations and common variants in the human arginase 1 (ARG1) gene: impact on patients, diagnostics, and protein structure considerations [Internet]. Hum Mutat. (2018) 39:1029–50. doi: 10.1002/humu.23545
7. Sin YY, Baron G, Schulze A, Funk CD. Arginase-1 deficiency. J Mol Med. (2015) 93:1287–96. doi: 10.1007/s00109-015-1354-3
8. Vockley JG, Goodman BK, Tabor DE, Kern RM, Jenkinson CP, Grody WW, et al. Loss of function mutations in conserved regions of the human arginase I gene. Biochem Mol Med. (1996) 59:44–51. doi: 10.1006/bmme.1996.0063
9. Bachmann C. Outcome and survival of 88 patients with urea cycle disorders: a retrospective evaluation. Eur J Pediatr. (2003) 162:410–6. doi: 10.1007/s00431-003-1188-9
10. Schlune A, vom Dahl S, Häussinger D, Ensenauer R, Mayatepek E. Hyperargininemia due to arginase I deficiency: the original patients and their natural history, and a review of the literature. Amino Acids. (2015) 47:1751–62. doi: 10.1007/s00726-015-2032-z
11. Wu T, Li X, Ding Y, Liu Y, Song J, Wang Q, et al. [Seven patients of argininemia with spastic tetraplegia as the first and major symptom and prenatal diagnosis of two fetuses with high risk]. Zhonghua Er Ke Za Zhi. (2015) 53:425–30.
12. Wu T-F, Liu Y-P, Li X-Y, Wang Q, Ding Y, Ma Y-Y, et al. Five novel mutations in ARG1 gene in chinese patients of argininemia. Pediatr Neurol. (2013) 49:119–23. doi: 10.1016/j.pediatrneurol.2013.04.026
13. Cohen YH, Bargal R, Zeigler M, Markus-Eidlitz T, Zuri V, Zeharia A. Hyperargininemia: a family with a novel mutation in an unexpected site. JIMD Rep. (2012) 5:83–8. doi: 10.1007/8904_2011_101
14. Hertecant JL, Al-Gazali LI, Karuvantevida NS, Ali BR. A novel mutation in ARG1 gene is responsible for arginase deficiency in an Asian family. Saudi Med J. (2009) 30:1601–3.
15. Bakhiet M, AlAwadi AMI, AlHammadi MM, Ali MF, Butti N. A case report of neurological complications owing to lately diagnosed hyperargininemia emphasizing the role of national neonatal screening policies in the kingdom of Bahrain. Medicine. (2018) 97:e10780. doi: 10.1097/MD.0000000000010780
16. Tadmouri GO, Nair P, Obeid T, Al Ali MT, Al Khaja N, Hamamy HA. Consanguinity and reproductive health among Arabs. Reprod Health. (2009) 6:17. doi: 10.1186/1742-4755-6-17
17. Bittles AH, Black ML. Consanguinity, human evolution, and complex diseases. Proc Natl Acad Sci. (2010) 107(suppl. 1):1779–86. doi: 10.1073/pnas.0906079106
18. Elsayed LEO, Mohammed IN, Hamed AAA, Elseed MA, Johnson A, Mairey M, et al. Hereditary spastic paraplegias: identification of a novel SPG57 variant affecting TFG oligomerization and description of HSP subtypes in Sudan. Eur J Hum Genet EJHG. (2016) 25:100–10. doi: 10.1038/ejhg.2016.108
19. Elsayed LEO, Drouet V, Usenko T, Mohammed IN, Hamed AAA, Elseed MA, et al. A novel nonsense mutation in DNAJC6 expands the phenotype of autosomal-recessive juvenile-onset Parkinson's disease. Ann Neurol. (2016) 79:335–7. doi: 10.1002/ana.24591
20. Elsayed LEO, Mohammed IN, Hamed AAA, Elseed MA, Salih MAM, Yahia A, et al. Case report of a novel homozygous splice site mutation in PLA2G6 gene causing infantile neuroaxonal dystrophy in a Sudanese family. BMC Med Genet. (2018) 19:72. doi: 10.1186/s12881-018-0592-y
21. Yahia A, Elsayed L, Babai A, Salih MA, El-Sadig SM, Amin M, et al. Intra-familial phenotypic heterogeneity in a Sudanese family with DARS2-related leukoencephalopathy, brainstem and spinal cord involvement and lactate elevation: a case report. BMC Neurol. (2018) 18:175. doi: 10.1186/s12883-018-1180-7
22. Elsayed LEO, Ahmed AEM, Stevanin G. Neurogenetic disorders in Africa: hereditary spastic paraplegia: a case study. In: Rotimi CN, Ibrahim ME, editors. The Genetics of African Populations in Health and Disease [Internet]. Cambridge: Cambridge University Press (2019) p. 311–9. (Cambridge Studies in Biological and Evolutionary Anthropology). Available online at: https://www.cambridge.org/core/books/genetics-of-african-populations-in-health-and-disease/neurogenetic-disorders-in-africa-hereditary-spastic-paraplegia/8F6AD8D94D7F5CE40D4A5FDE69176263 doi: 10.1017/9781139680295.013
23. Häberle J, Boddaert N, Burlina A, Chakrapani A, Dixon M, Huemer M, et al. Suggested guidelines for the diagnosis and management of urea cycle disorders. Orphanet J Rare Dis. (2012) 7:32. doi: 10.1186/1750-1172-7-32
24. Uchino T, Endo F, Matsuda I. Neurodevelopmental outcome of long-term therapy of urea cycle disorders in Japan. J Inherit Metab Dis. (1998) 21(Suppl. 1):151–9. doi: 10.1023/A:1005374027693
25. Asrani KH, Cheng L, Cheng CJ, Subramanian RR. Arginase I mRNA therapy - a novel approach to rescue arginase 1 enzyme deficiency. RNA Biol. (2018) 15:914–22. doi: 10.1080/15476286.2018.1475178
Keywords: hyperargininemia, ARG1 gene, whole exome sequencing, Sudan, spastic quadriplegia
Citation: Elsayed LEO, Mohammed IN, Hamed AAA, Elseed MA, Salih MAM, Yahia A, Abubaker R, Koko M, Abd Allah ASI, Elbashir MI, Ibrahim ME, Brice A, Ahmed AE and Stevanin G (2020) Novel Homozygous Missense Mutation in the ARG1 Gene in a Large Sudanese Family. Front. Neurol. 11:569996. doi: 10.3389/fneur.2020.569996
Received: 05 June 2020; Accepted: 25 September 2020;
Published: 29 October 2020.
Edited by:
Andrea Legati, Carlo Besta Neurological Institute (IRCCS), ItalyReviewed by:
Mario Reynaldo Cornejo-Olivas, National Institute of Neurological Sciences (INCN), PeruShahid Mahmood Baig, National Institute for Biotechnology and Genetic Engineering, Pakistan
Copyright © 2020 Elsayed, Mohammed, Hamed, Elseed, Salih, Yahia, Abubaker, Koko, Abd Allah, Elbashir, Ibrahim, Brice, Ahmed and Stevanin. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Liena E. O. Elsayed, bGllbmEuZWxzYXllZEBnbWFpbC5jb20=
 Liena E. O. Elsayed
Liena E. O. Elsayed Inaam N. Mohammed1
Inaam N. Mohammed1 Mahmoud Koko
Mahmoud Koko Amal S. I. Abd Allah
Amal S. I. Abd Allah Muntaser E. Ibrahim
Muntaser E. Ibrahim Alexis Brice
Alexis Brice Giovanni Stevanin
Giovanni Stevanin