- 1Department of Paediatrics, Chris Hani Baragwanath Academic Hospital and School of Clinical Medicine, University of the Witwatersrand, Johannesburg, South Africa
- 2Institute of Infection and Immunity, St. Georges, University of London, London, United Kingdom
- 3Medical Research Council (MRC) Clinical Trials Unit at University College London (UCL), London, United Kingdom
Introduction: Multidrug-resistant, Gram-negative infections, particularly due to carbapenem resistant organisms (CRO), have increased globally. Few studies have reported on the burden of CRO in neonates from low-middle income countries (LMIC). This study aimed to determine the incidence and mortality rates of culture-confirmed Gram-negative infections, with a special focus on CRO in a neonatal unit from a LMIC.
Materials and Methods: Positive bacterial cultures from sterile sites of infants admitted in the neonatal unit from the 1st January 2018 to 31st December 2019, were reviewed retrospectively. Type of organism, susceptibility and outcomes were recorded. Data on Gram-negative isolates, including the CRO, were extracted. Rates and outcomes were analysed.
Results: There were 2219 neonates with organisms isolated from sterile sites (blood and cerebrospinal fluid), accounting for 30% of all admissions, giving a neonatal sepsis incidence of 17.9/1000 patient-days. There was a total of 1746 positive isolates (excluding coagulase negative Staphyloccocus). Of these, 1706 (98%) were isolated from blood, and 40 (2%) from cerebrospinal fluid. Overall, 1188 (68%) were Gram-negative, 371 (21%) Gram-positive and 187 (10.7%) fungal isolates. The common Gram-negatives were Acinetobacter baumannii (526/1188;44%) and Klebsiella pneumoniae (469/1188;40%). Carbapenem resistance was observed in 359 (68%) of the Acinetobacter baumannii (CRAB) and in 103 (18%) of the Enterobacterales (CRE) isolates, with 98% of CRE being Klebsiella pneumoniae (CR-Klebs). Twenty-four (41%) of Pseudomonas species were carbapenem resistant. Overall, carbapenem resistance was seen in 42% of all Gram-negative organisms. The rate of CRAB and CRE were 2.9 and 0.8/1000 patient-days respectively. The overall, all-cause in-hospital mortality rate in infants with Gram-negative isolates was 22%, with higher mortality rate in those infected with CRO compared to non-CRO (34% vs 13%; OR 3.44; 95% CI 2.58–4.60; p < 0.001). The mortality rate in infants with CRE was higher than those with CRAB (48% vs 33%; OR 1.85; 95% CI 1.18–2.89; p = 0.007).
Conclusion: We observed a high incidence of positive cultures from sterile sites. The common organisms isolated were Gram-negatives, and among these carbapenem resistance was high and was associated with high mortality. Mortality was higher in infants with CRE compared to those with CRAB.
Introduction
Globally, approximately 5.8 million children under five years of age die every year (1), with close to half (45%) of these deaths occurring during the neonatal period (first 28 days of life). Low-middle income countries (LMIC) carry the burden of more than 90% of these deaths, with the highest neonatal mortality rates concentrated in Sub-Saharan Africa and South Asia. Infections are the third most common cause of deaths in neonates (2). A large proportion of neonates acquire infections during their hospital stay (3). The reported incidence of hospital associated infections (HAI) in neonatal units worldwide ranges from 6% to 38% of admitted patients, with the burden of disease being higher in LMIC (4, 5).
Organisms most commonly responsible for HAI in LMIC are Gram-negative pathogens, namely Klebsiella species, Escherichia coli, Pseudomonas species and Acinetobacter baumannii. There has been a global increase in antimicrobial resistance among a number of these organisms with increased use of carbapenems (6, 7). High usage of carbapenems to treat Gram-negative MDRO, has resulted in the emergence of carbapenem resistant organisms (CRO), posing a global health challenge, especially in LMIC, where therapeutic options are limited.
One of the important mechanisms underlying carbapenem resistance is the production of a specific group of beta lactamases, known as the carbapenemases, which have the ability to hydrolyze carbapenems. Beta lactamases are grouped into four classes based on their amino acid sequence (class A, B, C and D). Carbapenem resistance may emerge through expression of beta lactamases according to their molecular structure. The three key groups of carbapenemases, of clinical and epidemiological significance include the Klebsiella pneumoniae carbapenemase (KPC), New Delhi metallo-beta lactamases (NDM) and oxacillinases (OXA). KPC, a class A carbapenemase, is able to hydrolyze a broad spectrum of beta-lactams, including penicillins, cephalosporins, carbapenems and aztreonam. NDM belongs to class B carbapenamases and hydrolyze all beta-lactams, except for the monobactams such as aztreonam. Class D carbapenemases consist of the OXA beta lactamases, of which the most commonly found in Enterobacteriaceae are the OXA-48-like enzymes, which weakly hydrolyze carbapenems and typically spare the expanded spectrum cephalosporins (8, 9).
The commonly reported CRO are the carbapenem resistant Enterobacterales (CRE), in particular Klebsiella species, and the non-lactose fermenters namely carbapenem resistant A. baumannii (CRAB) and Pseudomonas species (10, 11). Although the literature reports many case series of CRO from the adult population, there have been only limited case reports highlighting the problem of CRO, especially CRE and CRAB in neonates (11–15).
Mortality rates secondary to HAI caused by Gram-negative MDRO, are reported to be as high as 40% (16–18). Moreover, infections with CRO have been reported to have higher mortality rates than those infected with carbapenem susceptible organisms (CSO) (19, 20). There is insufficient literature on the incidence and outcomes of multi-drug resistant Gram-negative infections in neonates from LMIC. This study aimed to determine the incidence and mortality rates of culture-confirmed Gram-negative infections, with a special focus on CRO in a single large tertiary neonatal unit in a LMIC.
Materials And Methods
Study Design and Study Population
This was a retrospective case series conducted over two years in the neonatal unit at Chris Hani Baragwanath Academic Hospital (CHBAH), in Johannesburg, South Africa. Infants admitted to the neonatal unit from 1st January 2018 to the 31st December 2019 who had organisms isolated from blood and/or cerebrospinal fluid (CSF) were eligible for enrolment. All isolates from these infants were included in the analysis, except for coagulase-negative Staphylococci (CoNS), Bacillus species and Corynebacterium species, as these were considered in our setting to be contaminants (21). Only infants infected with proven significant Gram-negative infections were included in the final analysis. Isolates could not be stratified according to postnatal age of infection (early onset versus late onset), as information on this was not available for analysis.
Study Setting
Chris Hani Baragwanath Academic Hospital is a public, tertiary hospital situated in Johannesburg, South Africa. It is the referral hospital for community health centers in Soweto and surrounding areas, and a referral center for surgical services for neonates from local district and regional hospitals in southern Gauteng and North West province. The hospital conducts approximately 20,000 of the 33,000 annual births in Soweto. The neonatal unit at CHBAH has 185 neonatal beds consisting of 18 intensive care beds (level 3 nursery), 48 high care beds (level 2 nursery), 100 standard care beds (level 2 and 1 nursery) and 19 kangaroo mother care (KMC) beds (level 1 nursery).
Neonates born at CHBAH requiring admission are admitted to the neonatal unit from the delivery room. Neonates born at other facilities, who have never left that facility, are referred to the neonatal unit at CHBAH for tertiary care. These admitted babies remain in the neonatal unit until they are discharged home, transferred back to the referring hospital or until the time of death. Babies from the community are not admitted to the neonatal unit. Patient days are calculated by taking a census of patients in the neonatal unit on a daily basis.
The unit protocol is that blood cultures are taken if an infant has any clinical signs or symptoms suggestive of infection on admission, or at any time during their hospital stay. A lumbar puncture is done either at the time of the work-up for sepsis in neonates with suspected HAI or when there is a confirmed blood stream infection (BSI). For suspected early onset sepsis, neonates are started empirically on ampicillin and gentamicin on admission to the neonatal unit. For suspected HAI, patients are started empirically on a combination of piperacillin-tazobactam and amikacin or meropenem if deemed to be severely ill by the attending physician. If the patient is already on meropenem for 24-48 hours and shows clinical deterioration with signs of haemodynamic instability, severe metabolic acidosis or increased ventilatory support requirement, then colistin is added empirically.
The local microbiology laboratory uses an automated continuous monitoring blood culture system (BacT/Alert system, BioMerieux, Marcy l’Etoile, France). If bacterial growth is detected, a Gram stain is performed and the sample sub-cultured onto appropriate media and incubated overnight. Further identification and antimicrobial susceptibilities with minimum inhibitory concentrations (MIC) are performed using either the Kirby-Baeur method or the automated system, Microscan, Siemens, USA and interpreted according to the Clinical Laboratory Standards Institute standards (22). Organisms with in-vitro resistance to more than one antimicrobial agent are referred to as multidrug resistant organisms (MDRO). Microbiology results are communicated telephonically between the microbiologists and the neonatologists daily detailing results with patient’s names, hospital number and ward. When a culture result is received, the patient is identified, and treatment is started with the appropriate antimicrobial.
Data Collection
All positive microbiological cultures identified from blood and/or CSF, including identification of isolates, and antibiotic susceptibilities, were retrieved from National Health Laboratory Services (NHLS) microbiology database. Information on primary outcome was retrieved from the patients’ files. All data retrieved were captured into an MS-Excel spreadsheet.
Study Definitions
As only limited patient records were available retrospectively for analysis of the clinical and laboratory characteristics, isolates that were CoNS, Corynebacterium species, and Bacillus species were excluded from the analysis, as they were considered to be contaminants (21). All other pathogens were considered significant if grown from a blood or CSF culture. An isolate was defined as a CRO if the organism displayed complete or intermediate resistance to all classes of antimicrobials, including one or more of the carbapenem antibiotics, meropenem, ertapenem and imipenem, and in this group colistin susceptibility was determined. Isolates that were susceptible to the carbapenem antibiotics and any other class of antimicrobials were defined as CSO. Primary outcome was defined as all-cause mortality before hospital discharge.
Statistical Analysis
Descriptive statistics were used to determine infection rates due to CRO and the all-cause mortality rates in infants with CRO. Incidence rate was calculated per 1000 patient-days and mortality rate per 100 patients with infections. Logistic regression analysis was performed to compare mortality rates between CRO and CSO on all Gram-negative infections combined and by individual Gram-negative species separately. Differences between the two groups were considered to be statistically significant when the p-value was less than 0.05 or the 95% confidence intervals of odds ratios were not including the number 1. Statistical analysis was performed using STATA version 15.
Ethics approval to conduct this study was obtained from the Human Research Ethics Committee, of the University of the Witwatersrand, Johannesburg.
Results
Incidence
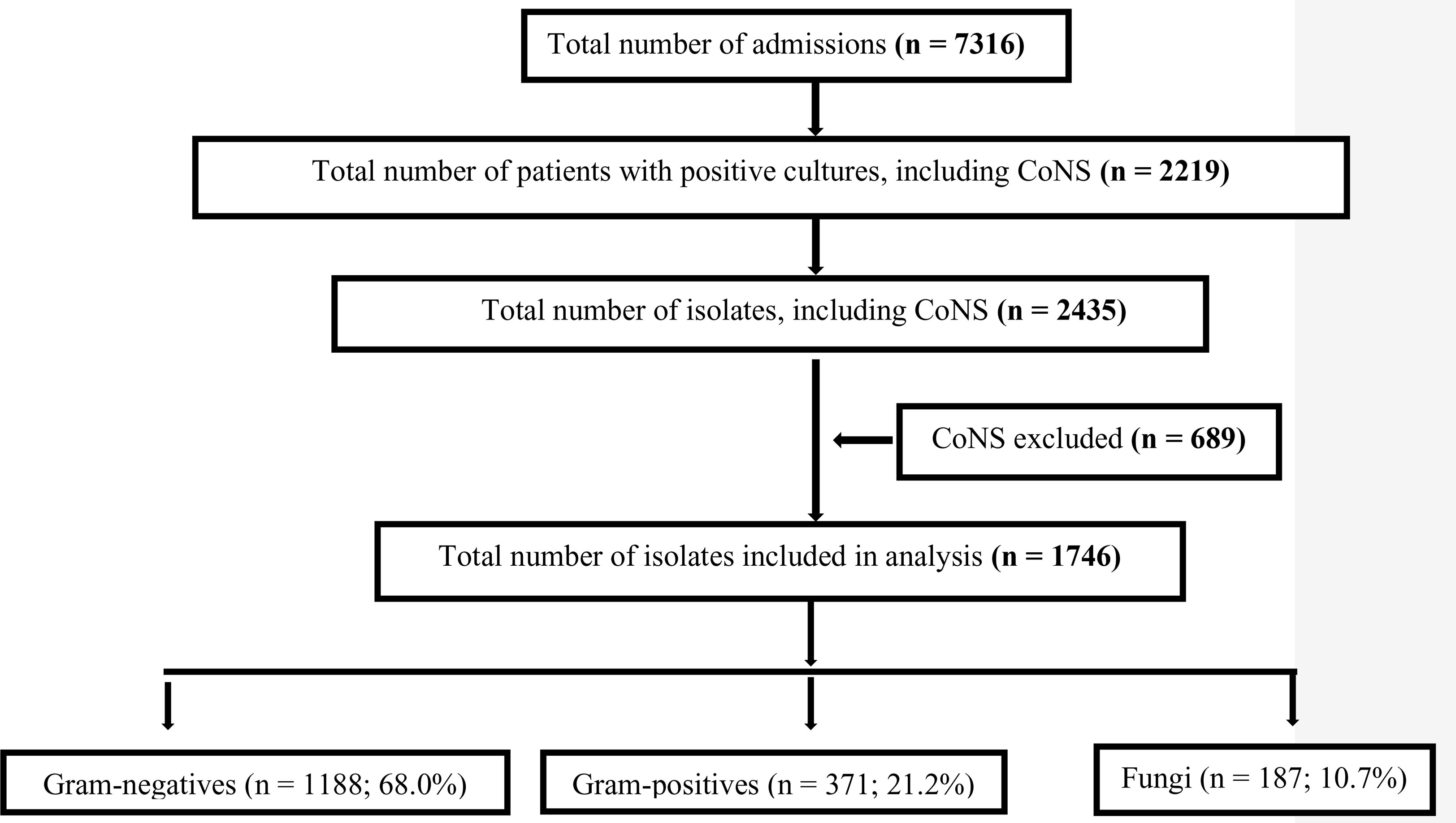
There was a total of 7316 admissions to the neonatal unit over the two-year study period including a total of 123812 patient-days (Figure 1). In this retrospective study, no data was available on the numbers of babies presenting with the clinical signs of sepsis requiring empiric treatment. There were 2219 neonates that were screened for sepsis where a pathogen, including CoNS, was isolated from a blood or CSF culture (30% of all admissions). The overall incidence of culture positive neonatal sepsis was 17.9 cases per 1000 patient-days. There were a total of 2435 isolates identified from 2219 patients. After excluding CoNS, the incidence rates for significant culture positive isolates were 19.7 per 1000 patient days and Gram-negatives accounted for 68% of these isolates, with an incidence of 9.6 per 1000 patient days.
Types of Organisms Isolated From Sterile Sites
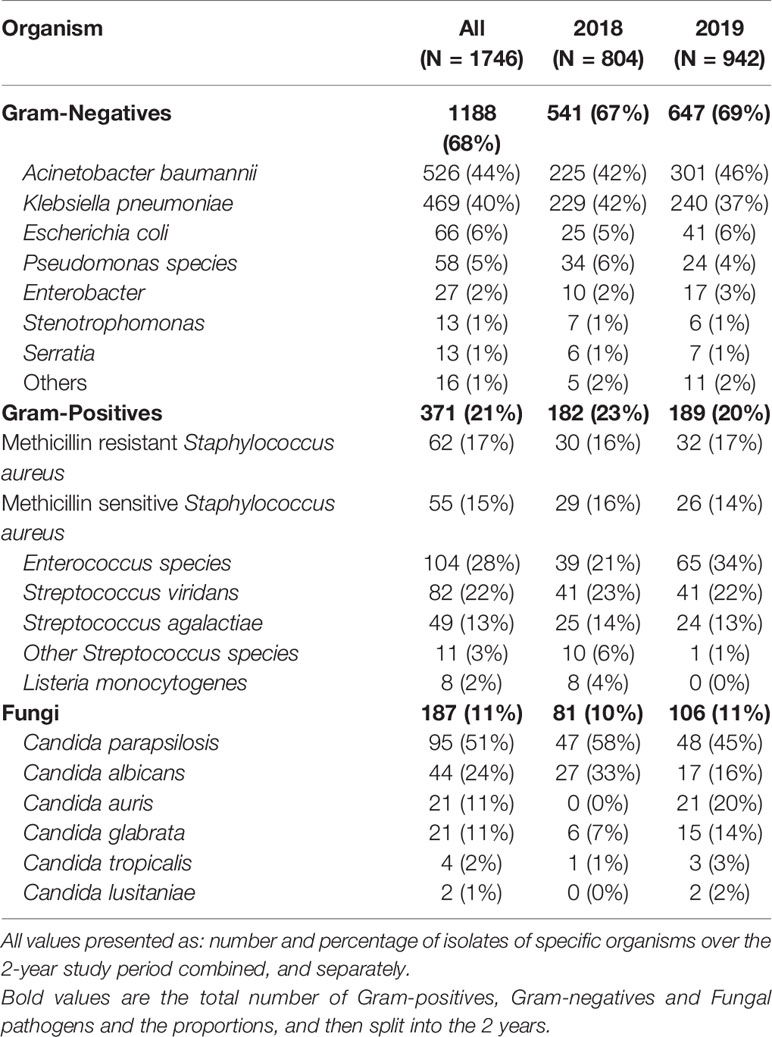
The pathogens isolated from sterile sites are presented in Table 1. One thousand seven hundred and fourty-six pathogens were isolated, of which 1706 pathogens (98%) were from blood and 40 pathogens (2%) were from CSF. Overall, A. baumannii (44%) and K. pneumoniae (40%) were the most common organisms isolated from sterile sites of neonates, accounting for 57% of all positive cultures due to pathogenic organisms, and 84% of all Gram-negatives. Among the Gram-positive organisms, the most common organisms were Streptococcus species (38%), Staphylococcus aureus (32%) and Enterococcus species (28%). The most common fungal pathogen isolated was Candida parapsilosis (51%).
Carbapenem Resistance Among Gram-Negative Isolates
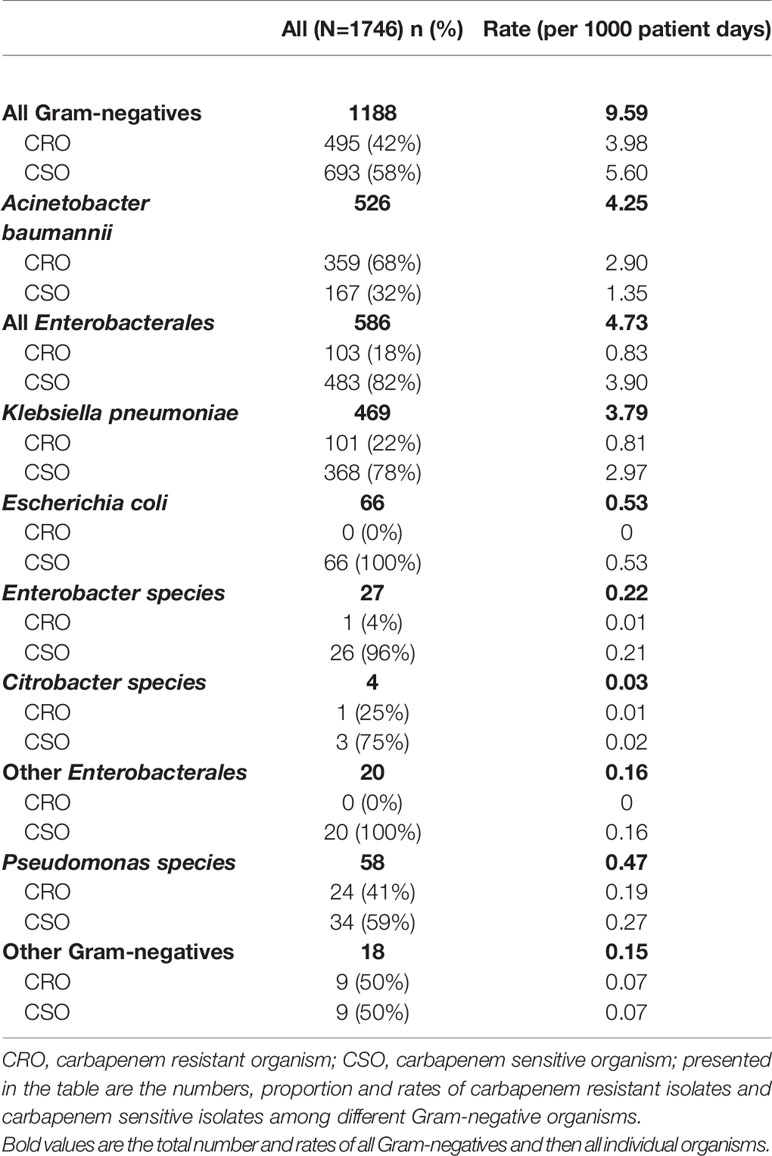
Carbapenem resistance was detected in 495 (42%) of all Gram-negative isolates, with an overall CRO rate of 4.0 per 1000 patient-days (Table 2). Among the A. baumannii isolates, 359 (68%) were carbapenem resistant, giving a carbapenem resistant A. baumannii (CRAB) rate of 2.9 per 1000 patient days. Among the K. pneumoniae isolates, 101 (22%) were carbapenem resistant. Carbapenem resistance was seen in one Citrobacter freundii isolate and one Enterobacter cloacae isolate. There was no carbapenem resistance seen among any other Enterobacterales. Overall, the rate of CRE was 0.8 per 1000 patient days. Among the Pseudomonas species, 24 (41%) were carbapenem resistant. A large majority (89%) of the positive cultures yielding CRO were from patients admitted in the level 2 and level 3 nurseries (Figure 2), compared to 80% of CSO infections.
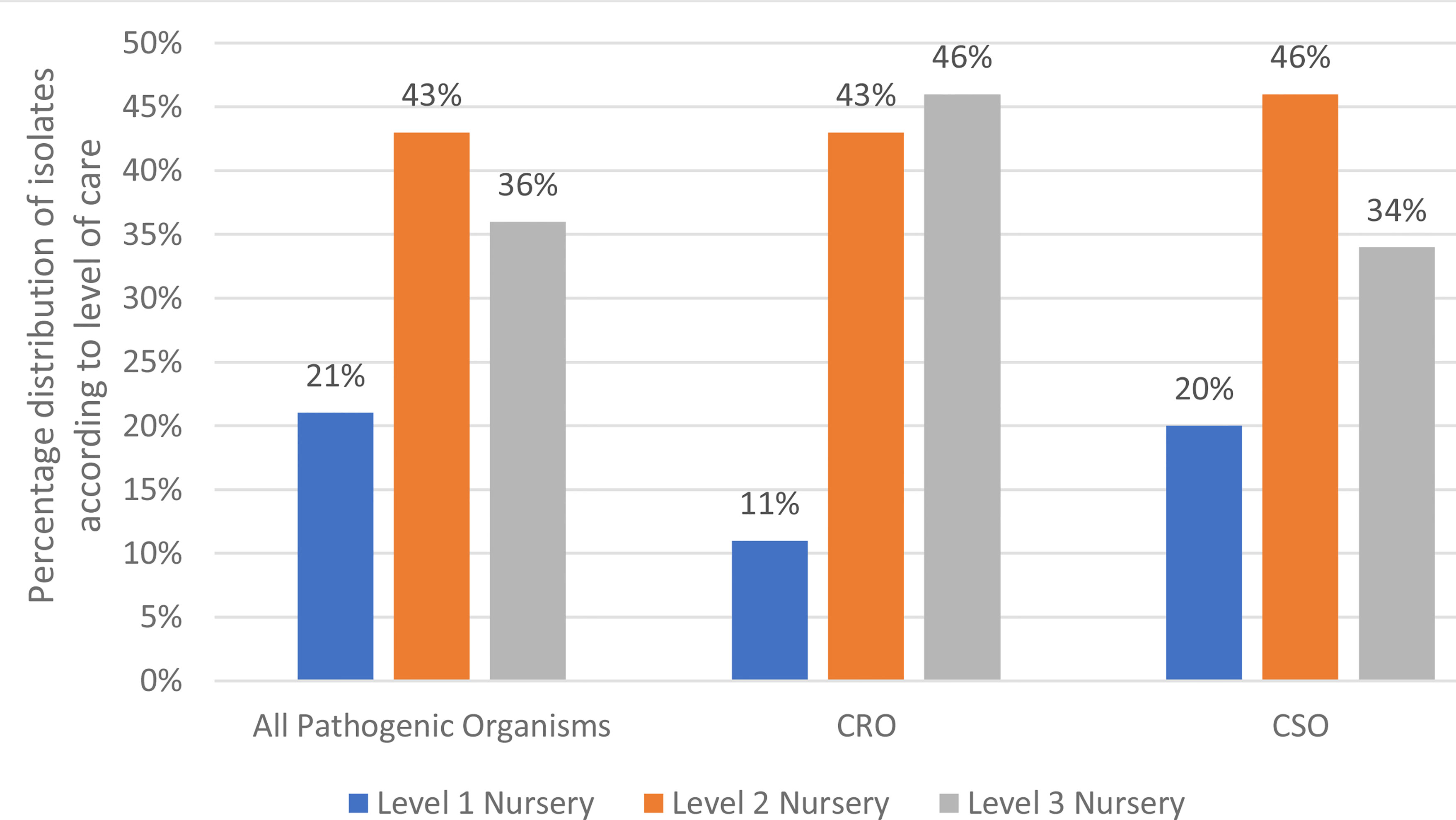
Figure 2 Level of care at time of positive culture. CRO- Carbapenem Resistant Organism; CSO, Carbapenem Sensitive Organism; Level 1 Nursery, Standard Care Unit; Level 2 Nursery, High Care Unit; Level 3 Nursery, Neonatal Intensive Care Unit.
All-Cause Mortality Rates
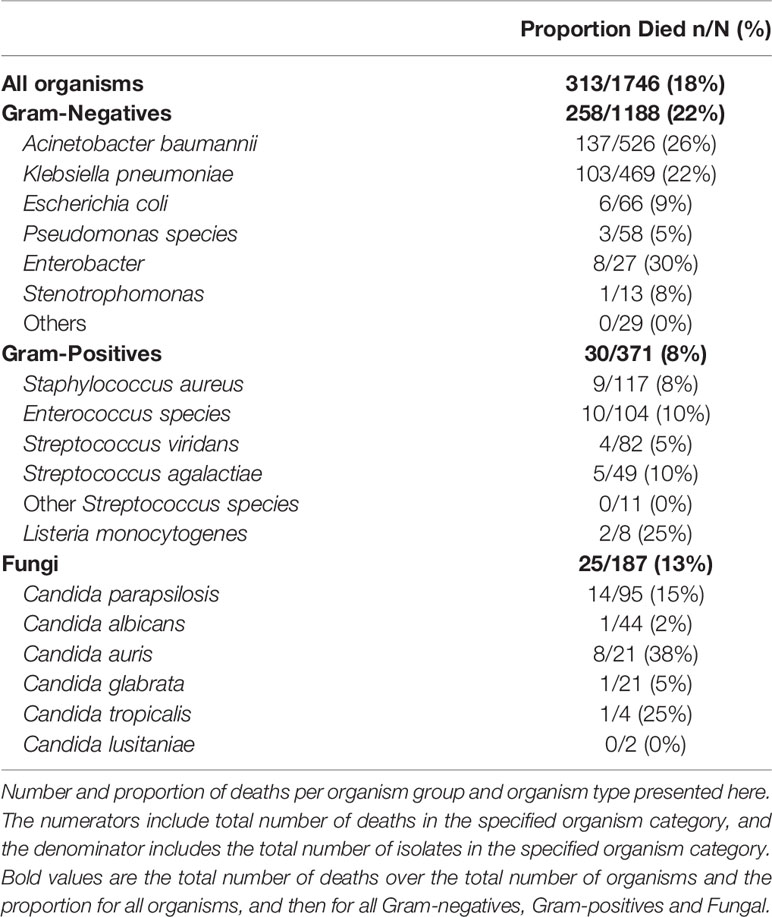
The overall all-cause in hospital mortality rate in infants with a significant organism detected was 18%; 22% for Gram-negative organisms, 8% for Gram-positive organisms and 13% for Fungi (Table 3). Among the Gram-negative organisms, mortality rates in infants infected with CRO was 34% compared to 13% in those infected with CSO (OR 3.44; 95% CI 2.58 – 4.60; p < 0.001; Table 4). Mortality rates of infants infected with carbapenem resistant K. pneumoniae was 48%, compared to 15% for infants with carbapenem sensitive K. pneumoniae infections (OR 5.15; 95% CI 3.16 - 8.37; P < 0.001). Similarly, mortality rates in infants infected with carbapenem resistant A. baumannii was 33% compared to 11% in infants with carbapenem sensitive A. baumannii (OR 3.81; 95% CI 2.25 - 6.45; p < 0.001). Infants with carbapenem resistant K. pneumoniae had a higher mortality rate compared to those with carbapenem resistant A. baumannii (48% vs 33%; OR 1.85; 95% CI 1.18 – 2.89; p = 0.007).
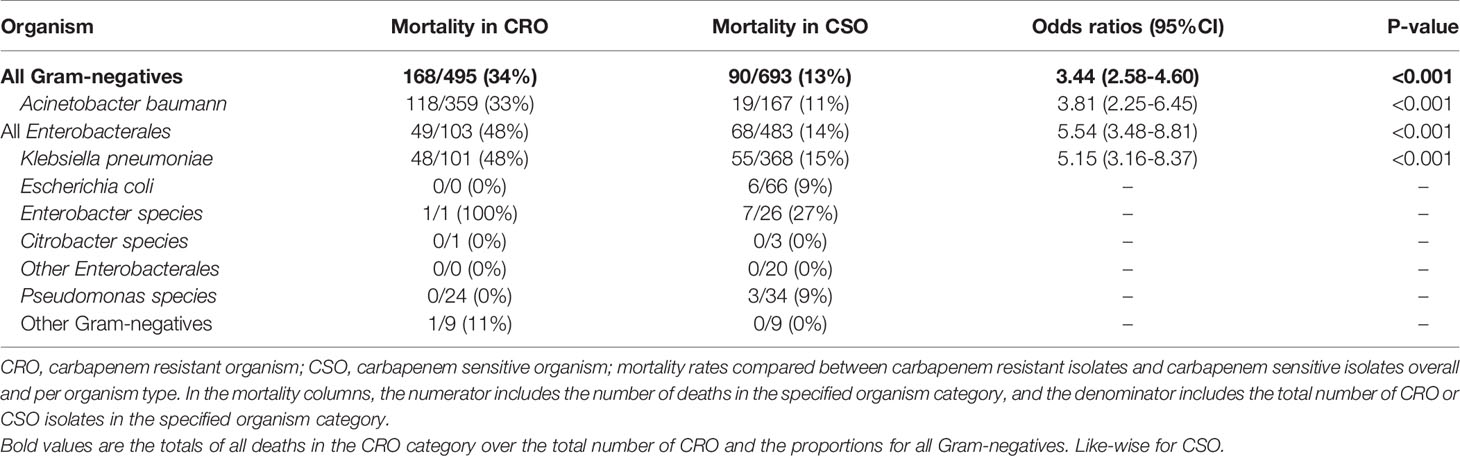
Table 4 All-cause mortality rates of different Gram-negative organisms according to their carbapenem susceptibility.
Discussion
Main Findings
This study aimed to quantify the incidence and mortality rates of culture-confirmed Gram-negative infections, especially CRO infections, in a large neonatal unit from a LMIC. We found there was a very high incidence of culture confirmed sepsis episodes, which were identified in around 30% of all admissions. Gram-negative organisms accounted for the great majority of the isolates, and carbapenem resistance was high. Overall, there was a high all-cause mortality rate in patients with sterile site culture positive infections. Gram-negative infections were associated with the highest mortality rates. Patients with CRO had a significantly higher mortality rate compared to patients with CSO. Carbapenem resistant K. pneumoniae was associated with higher mortality compared to patients with carbapenem resistant A. baumannii.
Comparison to Previous Literature
The incidence of culture confirmed sepsis reported in this study at 17.9 per 1000 patient days is six times higher than that previously reported from HIC at 3 per 1000 patient days and 50 per 1000 admissions (23–25). This incidence is also higher than that observed in similar settings from two facilities in South Africa which reported incidences of 3.9/1000 patient days (5) and 14/1000 patient days (26), and also much higher than studies from other LMIC reported in the literature (27, 28). However, some of these studies only reported hospital acquired blood stream infections beyond three days of hospital stay. This study looked at all isolates from blood and CSF from time of admission to the neonatal unit.
Gram-negative organisms were found to be the predominant pathogens in this study, contributing to 68% of all isolates excluding contaminants. This is similar to findings from the Delhi Neonatal Infection Study (DeNIS), which reported that 75% of isolates were Gram-negative organisms (16). Gram-negative organisms have been shown to be the more predominant pathogens in LMIC compared to high income countries, where Gram-positive organisms predominate (5, 16, 28–30). Among the Gram-negative organisms, A. baumannii and K. pneumoniae were the most common pathogens, accounting for almost 85% of Gram-negative infections. Both A. baumannii and K. pneumonia have emerged as leading pathogens in many neonatal units and have been responsible for many hospital outbreaks (16, 31–34).
Few previous studies have reported on the emergence of CRO in the neonatal units (28, 35). The Burden of Antibiotic Resistance in Neonates from Developing Societies (BARNARDS) study reported on the antimicrobial resistance profiles of Gram-negative bacteria from 7 LMIC, and found that 54% of isolated bacteria were resistant to at least one antibiotic within four to six classes of antibiotics. Widespread carriage of carbapenem resistance genes and virulence factors was also observed (36). Another recent web-based survey, involving 39 neonatal units from 12 countries, looked at period prevalence of blood culture isolates and their resistance patterns. Carbapenem resistance rates ranged from 0% - 80%, with CRO rates in South Africa reported at 39% (14), similar to rate of 42% observed in this study. The high rates of carbapenem resistant A. baumannii (68%) and K. pneumoniae (22%) are similar to those reported in the DeNIS study where 78% of A. baumannii and 35% of Klebsiella species were found to be carbapenem resistant. However, the rate of carbapenem resistant K. pneumoniae from this study is much higher than that recently reported from another public tertiary hospital in South Africa (37). Carbapenem resistance among Pseudomonas isolates in our study was slightly higher, at 41%, compared to 31% reported in the DeNIS study (16). High rates of antimicrobial resistance among Gram-negative organisms have also been reported in other studies (28, 30)
Most pathogenic organisms, including the CRO, were isolated from neonates receiving higher levels of neonatal care. Patients receiving level 2 and level 3 neonatal care are usually a sicker subgroup of infants, requiring various invasive and non-invasive interventions, including frequent handling, blood sampling, intubation, central vascular catheterization, surgical procedures etc., predisposing them to sepsis. This finding is supported by previous studies which have shown that intensive care as well as high illness severity scores are risk factors for acquiring infections with MDRO (38, 39)
All-cause mortality rate in Gram-negative infections, reported in this study was 21%. Mortality was significantly higher in infections with CRO (34%) compared to CSO (13%). Various studies have reported higher mortality due to MDRO sepsis. A study from Jordan reported a significantly higher mortality rate in MDRO sepsis compared to non-MDRO sepsis (60% vs. 13%) (40). The DeNIS study also reported that mortality in culture-positive sepsis was 13.4% when caused by MDRO compared to 9.8% when caused by non-MDRO (16). In this study mortality in patients with carbapenem resistant Klebsiella was 48%, higher than the mortality rate of 33%, reported in another South African neonatal unit (37). The direct causality of multidrug resistance, including carbapenem resistance, on mortality cannot be ascertained, as it is possible that the sicker neonates, who are at higher risk of dying, are also at higher risk of acquiring infections with MDRO. Studies looking at risk factors for acquiring multidrug resistant infections have shown that previous need for mechanical ventilation, intensive care as well as high illness severity scores are independent predictors for acquiring infections with MDRO, suggesting that patients with MDRO and CRO are a sicker subgroup to start with (38, 39). Another possible explanation for the increased mortality seen in patients with CRO is the likely delay in starting treatment with the appropriate antimicrobial, as none of these organisms would be susceptible to the empiric treatment regimens used. A recent study has shown that inappropriate empiric antibiotic therapy is associated with increased mortality in young children and neonates (41). However, empiric prescribing of broad-spectrum antibiotics to adequately treat possible infections with MDRO and CRO, can also drive antibiotic resistance, and therefore, a balance has to be struck between appropriate prescribing of empiric antibiotics without driving further antibiotic resistance. Since individual patient data was not available for analysis in our study, we were unable to determine the possible association between empiric treatment discordance and mortality. In our study, higher mortality was seen in carbapenem resistant K. pneumoniae, compared to carbapenem resistant A. baumannii (48% vs 33%). Higher mortality in K. pneumoniae infections compared to A. baumannii infections has been reported in other studies (27).
The limitations of this study are mostly due to the nature of the retrospective study design. This study only reports on positive bacterial isolates from sterile sites. Although multiple isolates of the same pathogen from the same patient were not duplicated, this study did not take into consideration the clinical case definitions of sepsis. Information on molecular typing of strains and mechanisms of resistance was not available for analysis. Blood sampling techniques could not be determined. CoNS were excluded as potential contaminants due to lack of clinical data and repeat blood culture data. Individual patient-level data was not available which meant that case fatality rates could not be determined nor patient-level factors investigated. Consequently, we cannot determine with confidence the likely causality of the carbapenem resistance on mortality, and the possibility remains that some of the neonates with carbapenem resistant infections may have been less clinically well before infection than those with carbapenem susceptible infections. The main strength of this study is that it is one of very few studies providing data on the high burden of Gram-negative BSI and carbapenem resistance from a neonatal unit in a LMIC.
In conclusion, in this study from a large population of neonates from a public, tertiary neonatal unit in a LMIC, there was a high rate of positive cultures from sterile sites observed over a two- year period, with a predominance of Gram-negative organisms. This study confirms the high burden of multi-drug resistant Gram-negative sepsis in LMIC and their associated high mortality rate, especially in infants infected with carbapenem resistant organisms. Improving measures at infection control and prevention, strict adherence to antimicrobial stewardship practices, and ongoing surveillance and reporting should be emphasized. Trials to guide future recommendations on empiric treatment regimens for the treatment of MDR neonatal sepsis should be prioritized.
Data Availability Statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation.
Ethics Statement
The studies involving human participants were reviewed and approved by University of the Witwatersrand’s Human Research Ethics Committee. Written informed consent from the participants’ legal guardian/next of kin was not required to participate in this study in accordance with the national legislation and the institutional requirements.
Author Contributions
The study was conceptualized by RT and SV and the manuscript write up was supervised and edited by SV, MS, and MC. Data collection was performed by CO-E, NM and RT. The majority of the manuscript was written by RT. All authors contributed to the article and approved the submitted version.
Funding
This research was privately funded by RT. MC is supported by core support from the Medical Research Council UK to the MRC Clinical Trials Unit [MC_UU_12023/22 and MC_UU_00004/09].
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher’s Note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
Acknowledgments
We would like to thank Dr Firdose Nakwa for providing us with the statistics on neonatal admissions and number of patient days over the study period.
Abbreviations
BSI, Blood Stream Infection; CHBAH, Chris Hani Baragwanath Academic Hospital; CoNS, Coagulase Negative Staphylococcus; CRAB, Carbapenem Resistant Acinetobacter Baumannii; CRE, Carbapenem-Resistant Enterobacteriaceae; CR-Klebs, Carbapenem-Resistant Klebsiella; CRO, Carbapenem Resistant Organisms; CSF, Cerebrospinal Fluid; CSO, Carbapenem Sensitive Organisms; HAI, Healthcare Associated Infections; HIC, High Income Country; KMC, Kangaroo Mother Care; KPC, Klebsiella Pneumoniae Carbapenemase; LMIC, Low-Middle Income Country; MDRO, Multi Drug Resistant Organisms; MIC, Minimum Inhibitory Concentration; NDM, New Delhi Metallo-β-Lactamase; NHLS, National Health Laboratory Services; NICU, Neonatal Intensive Care Unit; OXA, Oxacillinases; WHO, World Health Organization.
References
1. GBD 2015 Child Mortality Collaborators. Global, Regional, National, and Selected Subnational Levels of Stillbirths, Neonatal, Infant, and Under-5 Mortality, 1980-2015: A Systematic Analysis for the Global Burden of Disease Study 2015. Lancet Lond Engl (2016) 388(10053):1725–74. doi: 10.1016/S0140-6736(16)31575-6
2. United Nations. The Millenium Development Goals Report (2015). Available at: http://mdgs.un.org/unsd/mdg/Resources/Static/Products/Progress2015/English2015.pdf.
3. Giannoni E, Agyeman PKA, Stocker M, Posfay-Barbe KM, Heininger U, Spycher BD, et al. Neonatal Sepsis of Early Onset, and Hospital-Acquired and Community-Acquired Late Onset: A Prospective Population-Based Cohort Study. J Pediatr (2018) 201:106–14.e4. doi: 10.1016/j.jpeds.2018.05.048
4. Dong Y, Speer CP. Late-Onset Neonatal Sepsis: Recent Developments. Arch Dis Child Fetal Neonatal Ed (2015) 100(3):F257–63. doi: 10.1136/archdischild-2014-306213
5. Dramowski A, Madide A, Bekker A. Neonatal Nosocomial Bloodstream Infections at a Referral Hospital in a Middle-Income Country: Burden, Pathogens, Antimicrobial Resistance and Mortality. Paediatr Int Child Health (2015) 35(3):265–72. doi: 10.1179/2046905515Y.0000000029
6. Hsia Y, Lee BR, Versporten A, Yang Y, Bielicki J, Jackson C, et al. Use of the WHO Access, Watch, and Reserve Classification to Define Patterns of Hospital Antibiotic Use (AWaRe): An Analysis of Paediatric Survey Data From 56 Countries. Lancet Glob Health (2019) 7(7):e861–71. doi: 10.1016/S2214-109X(19)30071-3
7. Prusakov P, Goff DA, Wozniak PS, Cassim A, Scipion CEA, Urzúa S, et al. A Global Point Prevalence Survey of Antimicrobial Use in Neonatal Intensive Care Units: The No-More-Antibiotics and Resistance (NO-MAS-R) Study. EClinicalMed (2021) 32:100727. doi: 10.1016/j.eclinm.2021.100727
8. Eichenberger EM, Thaden JT. Epidemiology and Mechanisms of Resistance of Extensively Drug Resistant Gram-Negative Bacteria. Antibiot Basel Switz (2019) 8(2):37. doi: 10.3390/antibiotics8020037
9. Iovleva A, Doi Y. Carbapenem-Resistant Enterobacteriaceae. Clin Lab Med (2017) 37(2):303–15. doi: 10.1016/j.cll.2017.01.005
10. Folgori L, Bielicki J, Heath PT, Sharland M. Antimicrobial-Resistant Gram-Negative Infections in Neonates: Burden of Disease and Challenges in Treatment. Curr Opin Infect Dis (2017) 30(3):281–8. doi: 10.1097/QCO.0000000000000371
11. Datta S, Roy S, Chatterjee S, Saha A, Sen B, Pal T, et al. A Five-Year Experience of Carbapenem Resistance in Enterobacteriaceae Causing Neonatal Septicaemia: Predominance of NDM-1. PloS One (2014) 9(11):e112101. doi: 10.1371/journal.pone.0112101
12. Zhu J, Sun L, Ding B, Yang Y, Xu X, Liu W, et al. Outbreak of NDM-1-Producing Klebsiella Pneumoniae ST76 and ST37 Isolates in Neonates. Eur J Clin Microbiol Infect Dis Off Publ Eur Soc Clin Microbiol (2016) 35(4):611–8. doi: 10.1007/s10096-016-2578-z
13. Kumar A, Randhawa VS, Nirupam N, Rai Y, Saili A. Risk Factors for Carbapenem-Resistant Acinetobacter Baumannii Blood Stream Infections in a Neonatal Intensive Care Unit, Delhi, India. J Infect Dev Ctries (2014) 8(08):1049–54. doi: 10.3855/jidc.4248
14. Li G, Bielicki JA, Ahmed ASMNU, Islam MS, Berezin EN, Gallacci CB, et al. Towards Understanding Global Patterns of Antimicrobial Use and Resistance in Neonatal Sepsis: Insights From the NeoAMR Network. Arch Dis Child (2020) 105(1):26–31. doi: 10.1136/archdischild-2019-316816
15. Bielicki JA, Sharland M, Heath PT, Walker AS, Agarwal R, Turner P, et al. Evaluation of the Coverage of 3 Antibiotic Regimens for Neonatal Sepsis in the Hospital Setting Across Asian Countries. JAMA Netw Open (2020) 3(2):e1921124. doi: 10.1001/jamanetworkopen.2019.21124
16. Investigators of the Delhi Neonatal Infection Study (DeNIS) collaboration. Characterisation and Antimicrobial Resistance of Sepsis Pathogens in Neonates Born in Tertiary Care Centres in Delhi, India: A Cohort Study. Lancet Glob Health (2016) 4(10):e752–760. doi: 10.1016/S2214-109X(16)30148-6
17. Ozsurekci Y, Aykac K, Cengiz AB, Basaranoglu ST, Sancak B, Karahan S, et al. Bloodstream Infections in Children Caused by Carbapenem-Resistant Versus Carbapenem-Susceptible Gram-Negative Microorganisms: Risk Factors and Outcome. Diagn Microbiol Infect Dis (2017) 87(4):359–64. doi: 10.1016/j.diagmicrobio.2016.12.013
18. Peters L, Olson L, Khu DTK, Linnros S, Le NK, Hanberger H, et al. Multiple Antibiotic Resistance as a Risk Factor for Mortality and Prolonged Hospital Stay: A Cohort Study Among Neonatal Intensive Care Patients With Hospital-Acquired Infections Caused by Gram-Negative Bacteria in Vietnam. PloS One (2019) 14(5):e0215666. doi: 10.1371/journal.pone.0215666
19. Stewardson AJ, Marimuthu K, Sengupta S, Allignol A, El-Bouseary M, Carvalho MJ, et al. Effect of Carbapenem Resistance on Outcomes of Bloodstream Infection Caused by Enterobacteriaceae in Low-Income and Middle-Income Countries (PANORAMA): A Multinational Prospective Cohort Study. Lancet Infect Dis (2019) 19(6):601–10. doi: 10.1016/S1473-3099(18)30792-8
20. Nour I, Eldegla HE, Nasef N, Shouman B, Abdel-Hady H, Shabaan AE. Risk Factors and Clinical Outcomes for Carbapenem-Resistant Gram-Negative Late-Onset Sepsis in a Neonatal Intensive Care Unit. J Hosp Infect (2017) 97(1):52–8. doi: 10.1016/j.jhin.2017.05.025
21. Vergnano S, Buttery J, Cailes B, Chandrasekaran R, Chiappini E, Clark E, et al. Neonatal Infections: Case Definition and Guidelines for Data Collection, Analysis, and Presentation of Immunisation Safety Data. Vaccine (2016) 34(49):6038–46. doi: 10.1016/j.vaccine.2016.03.046
22. Institute, CLSI. Performance Standards for Antimicrobial Susceptibility Testing; Seventeenth to Twenty-First Informational Supplements, in CLSI Documents M100-S17-21. Wayne, Pennsylvania, USA: CLSI (2017).
23. Scamardo MS, Dolce P, Esposito EP, Raimondi F, Triassi M, Zarrilli R. Trends, Risk Factors and Outcomes of Healthcare-Associated Infections in a Neonatal Intensive Care Unit in Italy During 2013–2017. Ital J Pediatr (2020) 46(1):34. doi: 10.1186/s13052-020-0799-3
24. Cailes B, Kortsalioudaki C, Buttery J, Pattnayak S, Greenough A, Matthes J, et al. Epidemiology of UK Neonatal Infections: The neonIN Infection Surveillance Network. Arch Dis Child - Fetal Neonatal Ed (2018) 103(6):F547–53. doi: 10.1136/archdischild-2017-313203
25. Verstraete EH, Mahieu L, De Coen K, Vogelaers D, Blot S. Impact of Healthcare-Associated Sepsis on Mortality in Critically Ill Infants. Eur J Pediatr (2016) 175(7):943–52. doi: 10.1007/s00431-016-2726-6
26. Ballot DE, Nana T, Sriruttan C, Cooper PA. Bacterial Bloodstream Infections in Neonates in a Developing Country. ISRN Pediatr (2012) 2012:1–6. doi: 10.5402/2012/508512
27. Jatsho J, Nishizawa Y, Pelzom D, Sharma R. Clinical and Bacteriological Profile of Neonatal Sepsis: A Prospective Hospital-Based Study. Int J Pediatr (2020) 2020:1–9. doi: 10.1155/2020/1835945
28. Johnson J, Robinson ML, Rajput UC, Valvi C, Kinikar A, Parikh TB, et al. High Burden of Bloodstream Infections Associated With Antimicrobial Resistance and Mortality in the Neonatal Intensive Care Unit in Pune, India. Clin Infect Dis (2020) 73(2):ciaa554. doi: 10.1093/cid/ciaa554
29. Ramasethu J. Prevention and Treatment of Neonatal Nosocomial Infections. Matern Health Neonatol Perinatol (2017) 3(1):5. doi: 10.1186/s40748-017-0043-3
30. Salama K, Gad A, El Tatawy S. Sepsis Profile and Outcome of Preterm Neonates Admitted to Neonatal Intensive Care Unit of Cairo University Hospital. Egypt Pediatr Assoc Gaz (2021) 69(1):8. doi: 10.1186/s43054-021-00055-1
31. Hosoglu S, Hascuhadar M, Yasar E, Uslu S, Aldudak B. Control of an Acinetobacter Baumannii Outbreak in a Neonatal ICU Without Suspension of Service: A Devastating Outbreak in Diyarbakir, Turkey. Infect (2012) 40(1):11–8. doi: 10.1007/s15010-011-0180-y
32. Eibach D, Belmar Campos C, Krumkamp R, Al-Emran HM, Dekker D, Boahen KG, et al. Extended Spectrum Beta-Lactamase Producing Enterobacteriaceae Causing Bloodstream Infections in Rural Ghana, 2007-2012. Int J Med Microbiol IJMM (2016) 306(4):249–54. doi: 10.1016/j.ijmm.2016.05.006
33. Tsai M-H, Chu S-M, Hsu J-F, Lien R, Huang H-R, Chiang M-C, et al. Risk Factors and Outcomes for Multidrug-Resistant Gram-Negative Bacteremia in the NICU. Pediatricsi (2014) 133(2):e322–9. doi: 10.1542/peds.2013-1248
34. Dramowski A, Aucamp M, Bekker A, Mehtar S. Infectious Disease Exposures and Outbreaks at a South African Neonatal Unit With Review of Neonatal Outbreak Epidemiology in Africa. Int J Infect Dis IJID Off Publ Int Soc Infect Dis (2017) 57:79–85. doi: 10.1016/j.ijid.2017.01.026
35. Chaurasia S, Sivanandan S, Agarwal R, Ellis S, Sharland M, Sankar MJ. Neonatal Sepsis in South Asia: Huge Burden and Spiralling Antimicrobial Resistance. BMJ (2019) 364:k5314. doi: 10.1136/bmj.k5314
36. BARNARDS Group, Sands K, Carvalho MJ, Portal E, Thomson K, Dyer C, et al. Characterization of Antimicrobial-Resistant Gram-Negative Bacteria That Cause Neonatal Sepsis in Seven Low- and Middle-Income Countries. Nat Microbiol (2021) 6(4):512–23. doi: 10.1038/s41564-021-00870-7
37. Ballot DE, Bandini R, Nana T, Bosman N, Thomas T, Davies VA, et al. A Review of -Multidrug-Resistant Enterobacteriaceae in a Neonatal Unit in Johannesburg, South Africa. BMC Pediatr (2019) 19(1):320. doi: 10.1186/s12887-019-1709-y
38. Mendel LC, Amity K, Katz DE, Lazarovitch T, Zaidenstein R, Marchaim D. The Epidemiology of Carbapenem Resistant Enterobacter Spp: A Case–Case–Control Matched Analysis. Infect Control Hosp Epidemiol (2021) 42(6):754–9. doi: 10.1017/ice.2020.1286
39. Palacios-Baena ZR, Giannella M, Manissero D, Rodríguez-Baño J, Viale P, Lopes S, et al. Risk Factors for Carbapenem-Resistant Gram-Negative Bacterial Infections: A Systematic Review. Clin Microbiol Infect (2021) 27(2):228–35. doi: 10.1016/j.cmi.2020.10.016
40. Yusef D, Shalakhti T, Awad S, Algharaibeh H, Khasawneh W. Clinical Characteristics and Epidemiology of Sepsis in the Neonatal Intensive Care Unit in the Era of Multi-Drug Resistant Organisms: A Retrospective Review. Pediatr Neonatol (2018) 59(1):35–41. doi: 10.1016/j.pedneo.2017.06.001
41. Cook A, Hsia Y, Russell N, Sharland M, Cheung K, Grimwood K, et al. Association of Empiric Antibiotic Regimen Discordance With 30-Day Mortality in Neonatal and Pediatric Bloodstream Infection—A Global Retrospective Cohort Study. Pediatr Infect Dis J (2021) 40(2):137–43. doi: 10.1097/INF.0000000000002910
Keywords: neonatal sepsis, multidrug resistance, low to middle income countries, carbapenem resistance, mortality
Citation: Thomas R, Ondongo-Ezhet C, Motsoaledi N, Sharland M, Clements M and Velaphi S (2022) Incidence and All-Cause Mortality Rates in Neonates Infected With Carbapenem Resistant Organisms. Front. Trop. Dis 3:832011. doi: 10.3389/fitd.2022.832011
Received: 09 December 2021; Accepted: 10 January 2022;
Published: 04 February 2022.
Edited by:
Daniel Gyamfi Amoako, National Institute for Communicable Diseases (NICD), South AfricaReviewed by:
Shravani Mitra, National Institute of Cholera and Enteric Diseases (ICMR), IndiaMohsina Huq, Qassim University, Saudi Arabia
Sumeyye Kazancioglu, Ankara City Hospital, Turkey
Copyright © 2022 Thomas, Ondongo-Ezhet, Motsoaledi, Sharland, Clements and Velaphi. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Reenu Thomas, cmVlbnUudGhvbWFzQHdpdHMuYWMuemE=
 Reenu Thomas
Reenu Thomas Claude Ondongo-Ezhet1
Claude Ondongo-Ezhet1 Nini Motsoaledi
Nini Motsoaledi Mike Sharland
Mike Sharland Michelle Clements
Michelle Clements Sithembiso Velaphi
Sithembiso Velaphi