- Department of Biomedical Engineering, University of Memphis, Memphis, TN, United States
The implantation of a biomaterial quickly initiates a tissue repair program initially characterized by a neutrophil influx. During the acute inflammatory response, neutrophils release neutrophil extracellular traps (NETs) and secrete soluble signals to modulate the tissue environment. In this work, we evaluated chloroquine diphosphate, an antimalarial with immunomodulatory and antithrombotic effects, as an electrospun biomaterial additive to regulate neutrophil-mediated inflammation. Electrospinning of polydioxanone was optimized for rapid chloroquine elution within 1 h, and acute neutrophil-biomaterial interactions were evaluated in vitro with fresh human peripheral blood neutrophils at 3 and 6 h before quantifying the release of NETs and secretion of inflammatory and regenerative factors. Our results indicate that chloroquine suppresses NET release in a biomaterial surface area–dependent manner at the early time point, whereas it modulates signal secretion at both early and late time points. More specifically, chloroquine elution down-regulates interleukin 8 (IL-8) and matrix metalloproteinase nine secretion while up-regulating hepatocyte growth factor, vascular endothelial growth factor A, and IL-22 secretion, suggesting a potential shift toward a resolving neutrophil phenotype. Our novel repurposing of chloroquine as a biomaterial additive may therefore have synergistic, immunomodulatory effects that are advantageous for biomaterial-guided in situ tissue regeneration applications.
Introduction
Biomaterial-guided in situ tissue regeneration utilizes a tissue engineering approach to guide the regeneration of diseased, damaged, or missing tissues (Li et al., 2015). However, compared to exogenous delivery in traditional tissue engineering, in situ tissue regeneration relies on the body’s endogenous cells and signals to drive the repair and regeneration processes. Electrospun biomaterials have great potential for guiding in situ tissue regeneration because their extracellular matrix (ECM)–mimicking fibers can be fabricated from a variety of biocompatible polymers and modified to suit tissue-specific applications (Lannutti et al., 2007; Kelleher and Vacanti, 2010). Additionally, fabrication of electrospun biomaterials is relatively simple, cost-effective, and easy to scale up for mass production. As such, electrospun templates are an excellent platform for developing biomaterials that guide in situ tissue regeneration with the potential for a far-reaching clinical impact.
Independent of location, the implantation of a biomaterial quickly initiates a tissue repair program to restore homeostasis that is initially characterized by an influx of neutrophils (Anderson et al., 2008; Chen and Nuñez, 2010). During the acute inflammatory response, neutrophils condition the microenvironment through multiple mechanisms before recruiting additional immune cells in the tissue repair program. Two of their most significant effector functions include the extrusion of neutrophil extracellular traps (NETs) and the secretion of soluble signals (Brinkmann et al., 2004; Lacy, 2006; Selders et al., 2017). Together, these mechanisms shape the microenvironment for resolution and healing or further perturbations of homeostasis.
NETs are composed of DNA that is complexed with antibacterial neutrophil-derived proteins, including histones, neutrophil elastase, and myeloperoxidase (MPO) (Brinkmann et al., 2004). They are released in response to bacterial signals for the purpose of killing pathogens and are also released in response to inflammatory mediators, such as interleukin 8 (IL-8) and tissue necrosis factor α (TNF-α), activated endothelial cells, and platelets (Gupta et al., 2005, 2010; Clark et al., 2007; Neeli et al., 2008). While they are indispensable for preventing pathogen dissemination, the dysregulated release of NETs is associated with aberrant effects in sterile inflammation due to the localization of noxious cargo that can damage host cells (Gupta et al., 2010; Kaplan and Radic, 2012; Chrysanthopoulou et al., 2014; Gregory et al., 2015). Of particular interest to tissue engineers is the ability of NETs to initiate thrombosis and fibrosis, both of which can be detrimental to functional tissue regeneration (Maugeri et al., 2006; Fuchs et al., 2010; Martinod et al., 2013; Chrysanthopoulou et al., 2014; Gregory et al., 2015; Riehl et al., 2016). In fact, our group has previously shown that NETs are released in response to the surface area–dependent, topographical cues of electrospun polydioxanone (PDO) biomaterials, functioning as a preconditioning event in the tissue repair program (Fetz et al., 2017).
Similar to the release of NETs, neutrophil degranulation and the secretion of soluble signals are meant to neutralize pathogens and initiate the tissue repair program, but they can become dysregulated, leading to tissue damage (Lacy, 2006; Selders et al., 2017). The combination of neutrophil recruitment and tissue damage is typically attributed to the release of proteinases that break down the ECM, such as matrix metalloproteinase 9 (MMP-9), and the release of proinflammatory chemotactic factors, such as IL-8 (Harada et al., 1994; Nagaoka and Hirota, 2000; Martinez et al., 2004; Rayment et al., 2008; Liu et al., 2009). With continual neutrophil recruitment and degranulation, the acute response can develop into a nonresolving, chronic response through a perpetual cycle of recruitment and activation (Martinez et al., 2004; Rayment et al., 2008). Despite these potential deleterious outcomes, neutrophil degranulation and the secretion of signaling molecules have also been shown to be tissue-restorative and proangiogenic (Ardi et al., 2007; Deryugina et al., 2014; Phillipson and Kubes, 2019). Neutrophils secrete vascular endothelial growth factor A (VEGF-A) and hepatocyte growth factor (HGF), both of which support and guide angiogenesis (Taichman et al., 1997; McCourt et al., 2001). Moreover, MMP-9 is also proangiogenic and has been shown to initiate and guide endothelial cell sprouting (Ardi et al., 2007; Christoffersson et al., 2012). Taken together, these data suggest that regulating neutrophil NET release and their secretion of signaling molecules at the onset of the tissue repair program is pertinent for regulating acute neutrophil-driven inflammation (Jhunjhunwala et al., 2015; Herath et al., 2017; Lin et al., 2017; Fetz et al., 2020).
In this work, we evaluated chloroquine diphosphate as an electrospun biomaterial additive to regulate in vitro NET release and the secretion of proinflammatory and prohealing mediators from human neutrophils. Chloroquine is a Food and Drug Administration–approved, antimalarial drug that has more recently been investigated as an immunomodulatory and antithrombotic drug for treating rheumatoid arthritis, systemic lupus erythematosus, and cancer (Wozniacka et al., 2006; Manic et al., 2014; Schrezenmeier and Dörner, 2020; Roldan et al., 2020). Furthermore, chloroquine has been shown to inhibit NET formation, indicating its potential benefit as an additive for regulating biomaterial-induced NET release and inflammation (Smith et al., 2014; Boone et al., 2015, 2018; Murthy et al., 2019). Here, we show that electrospun PDO biomaterials loaded with chloroquine modulate template-induced NET release and the inflammatory response from human neutrophils. We found that chloroquine suppresses NET release in a surface area–dependent manner at early time points while modulating proinflammatory and healing signals at both early and late time points. Ultimately, our findings demonstrate a novel repurposing of chloroquine as a template additive for in situ tissue engineering that modulates the in vitro acute inflammatory response to biomaterials.
Materials and Methods
Biomaterial Fabrication
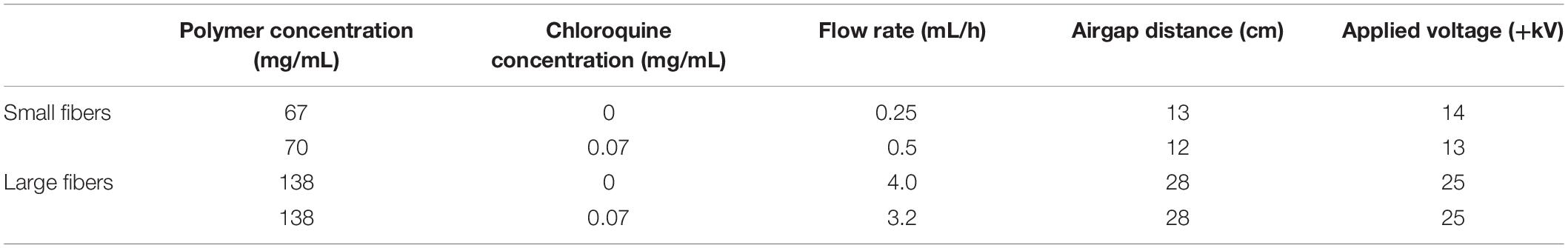
PDO (cat. no. 6100, Bezwada Biomedical, Hillsborough, NJ, United States) was dissolved overnight in 1,1,1,3,3,3-hexafluoro-2-propanol (cat. no. 003409-1KG, Oakwood Chemical, Estill, SC, United States) at varying concentrations (Table 1) to generate biomaterials composed of small and large fibers, previously shown to regulate NET release through their surface area–dependent, topographical cues (Fetz et al., 2017, 2018). Chloroquine diphosphate (cat. no. 0219391910, MP Biomedicals, Solon, OH, United States) was added to the solutions at a concentration of 0.07 mg/mL and dissolved for 1.5 h with gentle agitation before electrospinning. Then, the solutions were loaded into a syringe with a 22.5-gauge blunt needle for the 67 mg/mL PDO solution and an 18-gauge blunt needle for all other solutions and electrospun with optimized parameters (Table 1) to produce small and large fibers as desired (Fetz et al., 2018). Fibers were collected on a 20 × 75 × 5-mm grounded, stainless-steel rectangular mandrel that was rotating 1,250 revolutions/min and translating 6.5 cm/s over 13 cm. Additional higher concentrations of chloroquine were incorporated into the electrospun biomaterials during optimization (Supplementary Methods). For all experiments, 8-mm-diameter discs of the electrospun biomaterials were cut using a biopsy punch (cat. no. P825, Acuderm Inc., Ft. Lauderdale, FL, United States) and stored in a desiccator until use. Prior to cell culture, the biomaterials were irradiated with ultraviolet light at a wavelength of 365 nm using an 8-W lamp (cat. no. EN280L, Spectroline, Westbury, NY, United States) at a working distance of 9.5 cm. The samples were disinfected for 10 min on each side in a sterile, laminar flow hood and kept disinfected until cell culture.
Biomaterial Characterization
The biomaterials were imaged with a scanning electron microscope, and scanning electron micrographs (SEMs) were analyzed in FibraQuant 1.3 software (nanoTemplate Technologies, LLC) to quantify fiber diameter as previously described (Fetz et al., 2018). Briefly, 150 semiautomated random measurements per SEM were taken to determine the average and corresponding standard deviation for fiber diameter.
Chloroquine Elution From Biomaterials
The elution of chloroquine from the biomaterials was quantified over the first 24 h using a microplate reader to measure absorbance as described (Lima et al., 2018). The biomaterials (n = 4) were placed in a 96-well cell culture plate, and 150 μL of 1 × Hanks buffered salt solution (HBSS, calcium, magnesium, and phenol red free, cat. no. 14175095, Thermo Fisher Scientific, Waltham, MA, United States) was added to each well. After incubating at 37°C for 30 min, 1, 3, 6, and 24 h, the supernatant was removed and refreshed with 150 μL of HBSS. The absorbance of the collected supernatant was read on a SpectraMax i3x Multi-Mode Microplate Reader at 330 nm, and the chloroquine concentration was interpolated from a standard dilution ranging from 333 to 0 μg/mL in HBSS (Supplementary Figure 1). In addition to concentration, the average percent release and standard deviation were calculated for each biomaterial.
Isolation and Culture of Primary Human Neutrophils
Heparinized whole blood from healthy donors was obtained by venipuncture from Tennessee Blood Services. As purchased or donated samples are not traceable back to the donor, it does not qualify as human subjects research as determined by the University of Memphis Institutional Review Board on November 22, 2016. Neutrophils were then isolated as previously described using Isolymph® density separation (Neeli and Radic, 2013; Fetz et al., 2017, 2018). After isolation, neutrophils were resuspended in HBSS with 10 mM HEPES and 0.2% autologous serum at a concentration of 1 million neutrophils/mL. The disinfected biomaterials (n = 3) were placed in a 96-well plate, and 40 μL of the cell culture media was added to each well to hydrate the biomaterials. Negative and positive tissue culture plastic (TCP) wells (n = 3) received 30 μL of the cell culture media prior to cell seeding. Subsequently, 100 μL of cell culture media containing 100,000 neutrophils was added to each well followed by 10 μL of heparin (cat. no. H3393, Sigma–Aldrich, St. Louis, MO, United States) at a final concentration of 10 U/mL heparin. Heparin was added to dissociate NET-associated MPO as previously described (Parker et al., 2012; Minden-Birkenmaier et al., 2020). The negative vehicle and positive controls added to TCP wells were 0.15% dimethylsulfoxide in 10 μL of HBSS and 100 nM phorbol 12-myristate 13-acetate (cat. no. P8139, Sigma–Aldrich, St. Louis, MO, United States) in 10 μL of HBSS, respectively. The neutrophils were cultured at 37°C and 5% CO2 for 3 and 6 h. Following incubation, the samples were placed on ice for 10 min to inhibit neutrophil stimulation prior to processing. Three experiments were performed with unique donors (male, between 18 and 40 years of age), and the results were pooled for analysis.
Quantification of NETs and Secreted Signals
Supernatants were collected and assayed using a ProcartaPlex multiplex immunomagnetic assay (cat. no. PPX, Thermo Fisher Scientific, Waltham, MA, United States) on a MAGPIX® microplate reader (Luminex Corporation, Austin, TX, United States). The assayed analytes included angiopoietin, fibroblast growth factor 2, granulocyte colony-stimulating factor (CSF), HGF, IL-1β, IL-1 receptor antagonist, IL-6, IL-8, IL-10, IL-22, monocyte chemoattractant protein 1, MMP-9, MPO, TNF-α, and VEGF-A. To quantify percent NET release, the concentration of MPO was normalized to the concentration of MPO in the positive control at 6 h (Parker et al., 2012; Minden-Birkenmaier et al., 2020).
Fluorescent Microscopy
Samples were fixed with 10% buffered formalin (cat. no. SF1004, Thermo Fisher Scientific, Waltham, MA, United States) and stained with 5 μM SYTOX orange (cat. no. S34861, Thermo Fisher Scientific, Waltham, MA, United States) and NucBlueTM Fixed Cell ReadyProbesTM Reagent (DAPI, cat. no. R37606, Thermo Fisher Scientific, Waltham, MA, United States) as described (Minden-Birkenmaier et al., 2020). Briefly, samples were sequentially incubated with each stain for 5 min at room temperature. Three washes with 1 × phosphate-buffered saline for 5 min each were performed between each step. Cells and NETs were visualized on an Olympus BX43 fluorescent microscope.
Statistical Analysis
Statistical significance between fiber diameters was tested with a Kruskal–Wallis and Dunn multiple-comparisons test. All other statistical significance was tested with an analysis of variance and Holm–Sidak multiple-comparisons test. Statistical analyses were performed in Prism version 8.4.3 (GraphPad Software, San Diego, CA, United States) at a significance level of 0.05. Data are reported as mean ± standard deviation.
Results
Electrospun PDO Rapidly Elutes Chloroquine
PDO was electrospun to create biomaterials with small and large fibers (Figure 1A). Based on our previous work, the small- and large-fiber biomaterials in this study represent materials that trigger two distinct neutrophil NET responses and two distinct potentials for tissue regeneration (Fetz et al., 2017, 2018). In order to make comparisons independent of fiber size, the polymer concentration was adjusted for the small- and large-fiber biomaterials so that the addition of chloroquine did not alter the resulting fibers (Figure 1B). Any differences in the neutrophil inflammatory response can therefore be attributed to chloroquine elution. Both the small- and large-fiber biomaterials rapidly eluted chloroquine with near 100% elution within the first hour and no detectable increase after 3 h (Figure 1C), which is ideal and desired for targeting the acute neutrophil response during inflammation (Lammermann et al., 2013). The burst elution, driven by segregation of the charged drug to the outer surface of the electrospun fibers (Sun et al., 2008), equated to a concentration of 11.8 ± 1.33 μM and 12.2 ± 1.63 μM chloroquine for the small- and large-fiber biomaterials, respectively (Figure 1D). This elution was optimized by changing chloroquine incorporation during biomaterial fabrication to achieve an eluted concentration near those previously reported in the literature (Smith et al., 2014; Germic et al., 2017). Additional biomaterials were also fabricated to elute higher concentrations of chloroquine (Supplementary Figure 2).
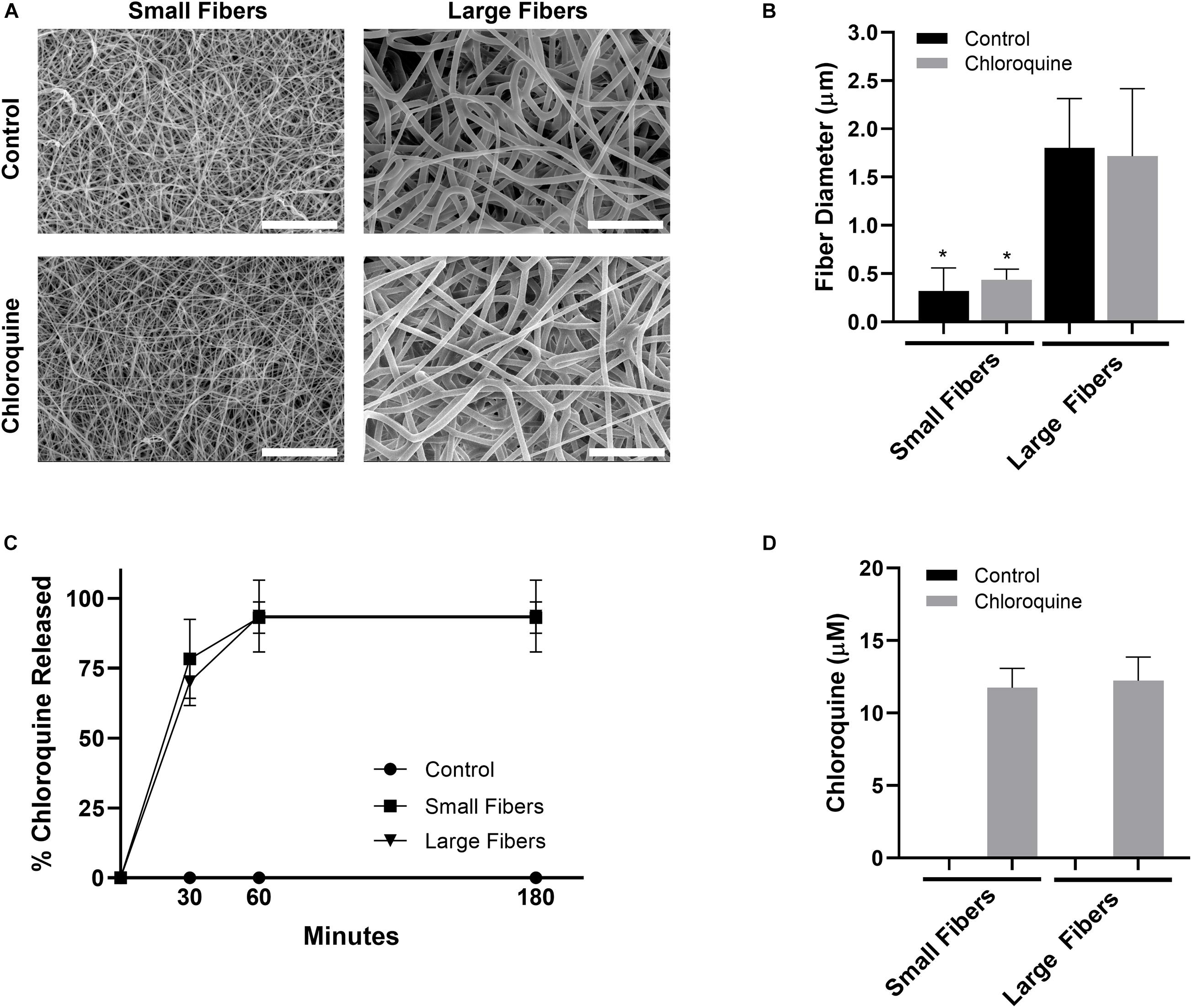
Figure 1. Chloroquine incorporation into electrospun biomaterials results in uniform fibers that rapidly elute the additive. (A) Representative SEMs of the control and chloroquine-loaded biomaterials. Micrographs were acquired at 1,000× magnification and scale bars are 30 μm. (B) Fiber diameters of the electrospun biomaterials. Measurements (n = 150) were taken in FibraQuant 1.3 software. (C) Percent chloroquine released from the biomaterials and (D) eluted chloroquine concentration at 3 h. There was no increase in concentration after 3 h. See Supplementary Figure 1 for the standard curve used to interpolate concentration (n = 4) from absorbance. Graphs show mean ± standard deviation. *p < 0.0001 was determined using a Kruskal–Wallis and Dunn multiple-comparisons test. Raw data are available in Supplementary Spreadsheet 1.
Chloroquine Elution Inhibits NET Release on Small Fibers
Neutrophils were isolated from the whole blood of healthy donors and seeded on electrospun biomaterials with or without chloroquine to trigger biomaterial-induced NET release. As anticipated, neutrophils had an increased propensity to form NETs on the small fibers compared to the large fibers at 3 h (Figure 2A). More importantly, the elution of chloroquine from the small-fiber biomaterials significantly reduced NET release to the level of the large fibers at 3 h while having no effect upon elution from the large fibers (Figure 2B). By 6 h, the difference between small and large fibers was less pronounced, and increased NET release was observed on both chloroquine-eluting biomaterials, suggesting a temporal, therapeutic window for inhibiting acute NET release (Figure 2B).
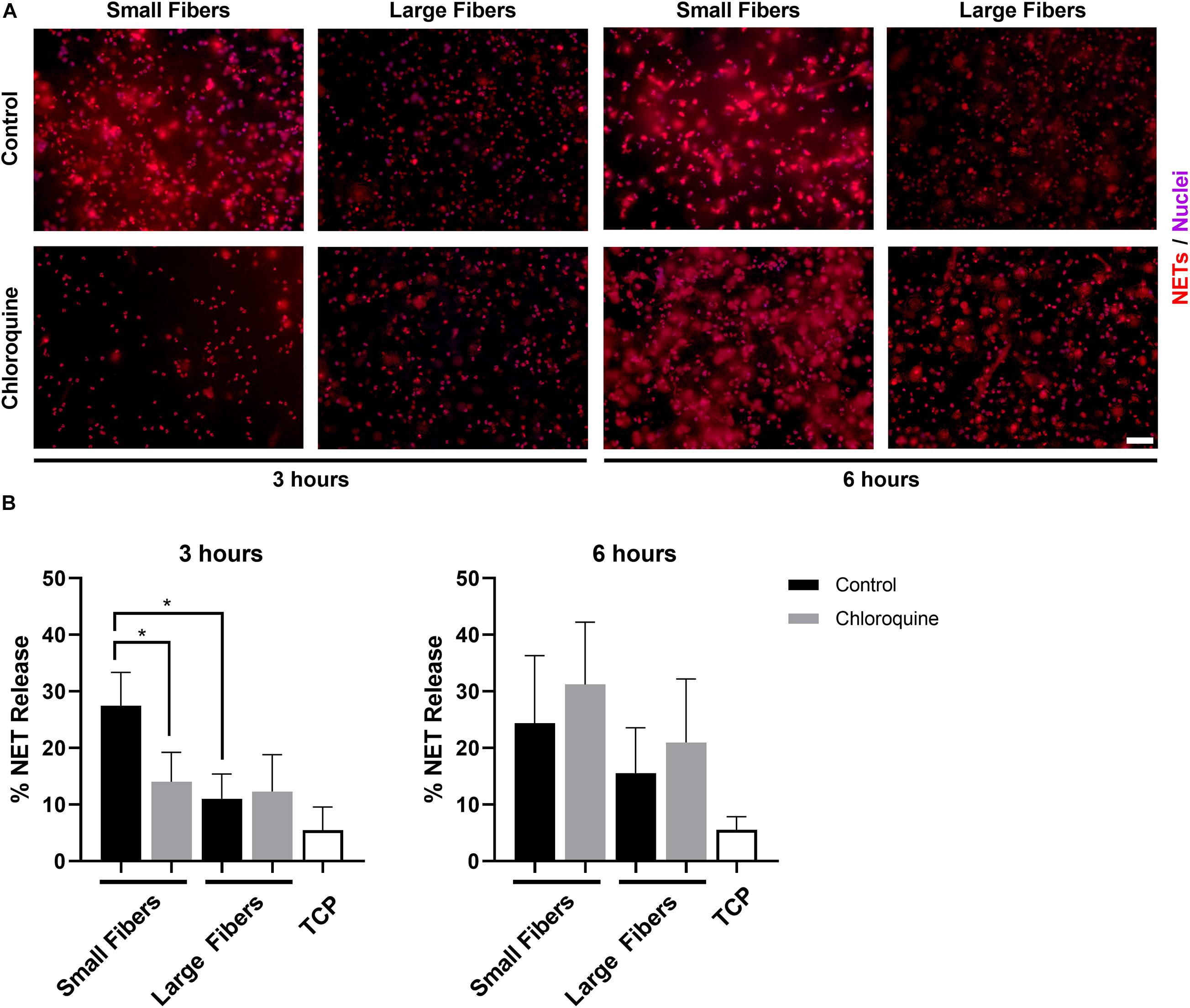
Figure 2. Chloroquine elution inhibits NET release on small fibers but has no effect on large fibers. (A) Fluorescent micrographs of neutrophils on the electrospun biomaterials at 3 and 6 h after seeding. Staining of NETs (red) and nuclei (purple) reveals that chloroquine elution from the small fibers attenuates NET formation at the early time point, but not at the late time point. Conversely, chloroquine elution from the large fibers does not modulate NET release. Scale bar is 50 μm. (B) Percent NET release at 3 h (left) and 6 h (right) as quantified by the ELISA for NET-disassociated MPO. The quantification of percent NET release (n = 3) indicates that chloroquine elution from the small fibers reduces NET release to the level of the large fibers at 3 h with no effect at 6 h. The data represent the mean ± standard deviation of three independent experiments with unique donors. *p < 0.0001 was determined using an analysis of variance (ANOVA) and Holm–Sidak multiple-comparisons test. Raw data are available in Supplementary Spreadsheet 1.
Chloroquine Elution Decreases Inflammatory Signal Secretion
Given the documented anti-inflammatory effects, we also evaluated if chloroquine elution would modulate the inflammatory response through the secretion of soluble signals using a multiplexed immunomagnetic assay. Of the assayed inflammatory analytes, only IL-8 and MMP-9 were detected in the supernatant (Figure 3). IL-8 is the archetypal neutrophil chemoattractant secreted by damaged cells as well as neutrophils during an acute inflammatory response. At both 3 and 6 h, IL-8 secretion was significantly greater on the small-fiber biomaterials compared to the large fibers (Figure 3A). As the small fibers appear to up-regulate NET release in a proinflammatory response, it is not surprising that IL-8 secretion mimicked the trends in NET release. However, despite observing a temporal inhibition of acute NET release at 3 h only, the elution of chloroquine from both small and large fibers continued to significantly suppress IL-8 secretion at 6 h. These data suggest there is independent regulation of NET release and IL-8 synthesis and secretion in the context of biomaterial-induced activation, which may be important for reducing aberrant neutrophil recruitment during the tissue repair program (Fujishima et al., 1993; de Oliveira et al., 2013; Degroote et al., 2020). Similar to IL-8, MMP-9 secretion was significantly greater on the small-fiber biomaterials compared to the large fibers with chloroquine elution suppressing secretion at 3 h (Figure 3B). However, unlike IL-8, MMP-9 secretion was near equivalent on all biomaterials by 6 h, suggesting a temporal modulation of MMP-9. As a promiscuous endopeptidase, elevated MMP-9 is correlated with tissue degradation and chronic inflammation, so its acute suppression by chloroquine may prevent triggering a continuum of matrix destruction during the initial inflammatory response (Nagaoka and Hirota, 2000; Rayment et al., 2008; Liu et al., 2009).
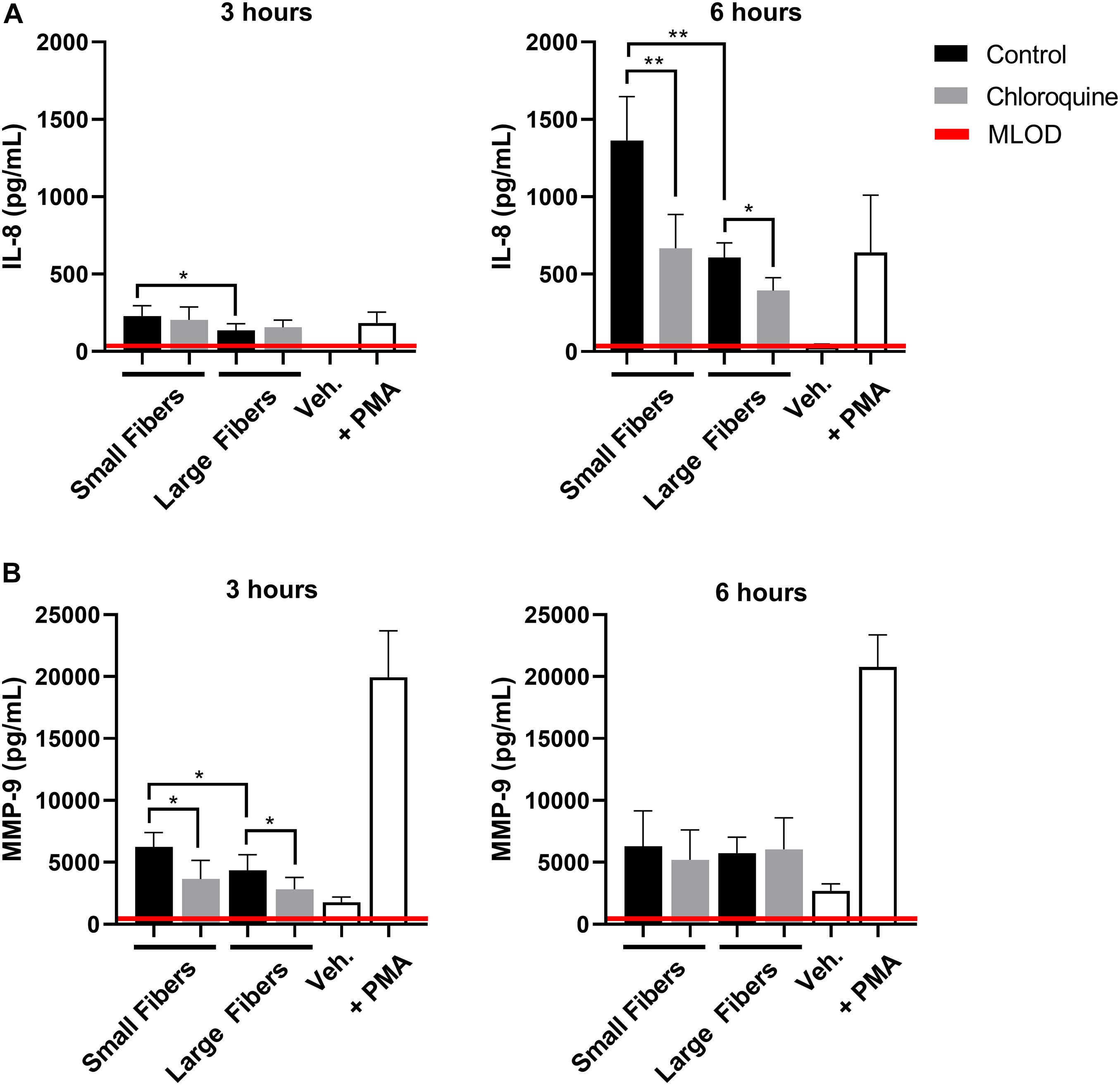
Figure 3. Chloroquine-eluting biomaterials suppress inflammatory IL-8 and MMP-9 secretion from neutrophils. (A) IL-8 and (B) MMP-9 secretion at 3 h (left) and 6 h (right) after neutrophil seeding. Chloroquine elution down-regulated IL-8 secretion at both time points, whereas it only attenuates the acute secretion of MMP-9. The data (n = 3) represent the mean ± standard deviation of three independent experiments with unique donors. *p < 0.05 and **p < 0.0001 were determined using an analysis of variance (ANOVA) and Holm–Sidak multiple-comparisons test. Raw data are available in Supplementary Spreadsheet 1.
Chloroquine Elution Increases Regenerative Signal Secretion
Although well characterized for its anti-inflammatory effects, chloroquine is not well studied for its potential regenerative effects. Therefore, we also evaluated if chloroquine elution would regulate the secretion of regenerative signals from biomaterial-interacting neutrophils. While several regenerative and anti-inflammatory analytes were assayed, only HGF, VEGF-A, and IL-22 were detected (Figure 4). The secretion of HGF (Figure 4A) and VEGF-A (Figure 4B) followed very similar trends at 3 h, with both having significantly greater secretion on large fibers compared to small fibers, which is the inverse of NET release. Additionally, the elution of chloroquine from the small fibers increased the secretion of HGF and VEGF-A to the level of the large fibers. By 6 h, the trends remained the same with an overall increase in the magnitude of secretion. As they are classic angiogenic signals (Taichman et al., 1997; McCourt et al., 2001; Wislez et al., 2003; Fridlender et al., 2009; He et al., 2016), these data indicate that chloroquine elution may help establish a more regenerative microenvironment around a biomaterial. Likewise, IL-22 secretion followed the same trends as HGF and VEGF-A, but its secretion was not detectable until 6 h, suggesting an absence of readily available stores (Dyring-Andersen et al., 2017). Nonetheless, as a proliferative and proangiogenic signal (Dyring-Andersen et al., 2017; Protopsaltis et al., 2019), the increased IL-22 secretion observed in response to chloroquine elution from the small fibers further suggests that chloroquine modulates the neutrophil phenotype toward healing and regeneration.
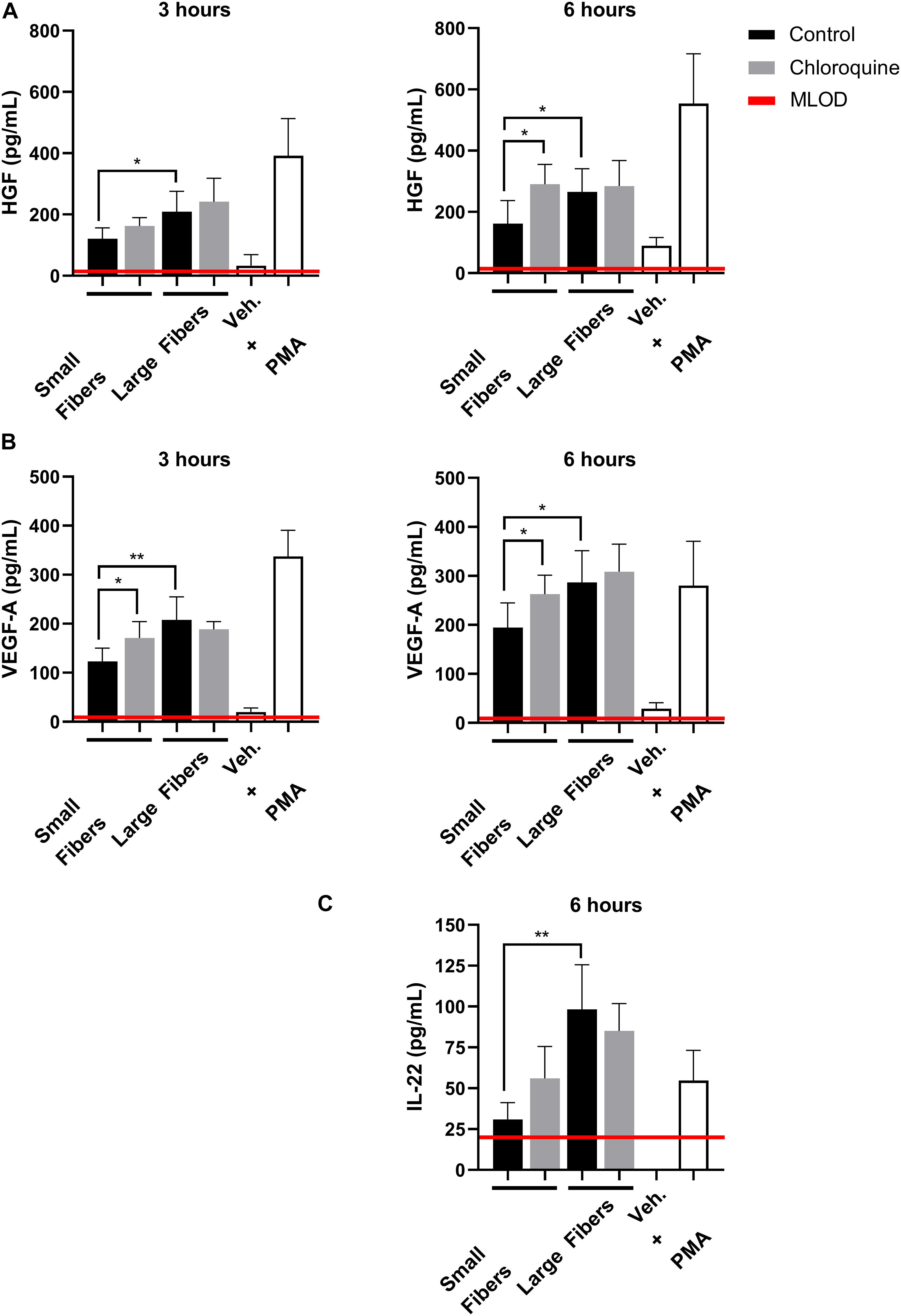
Figure 4. Chloroquine-eluting biomaterials increase regenerative HGF, VEGF-A, and IL-22 secretion from neutrophils. (A) HGF and (B) VEGF-A were up-regulated with chloroquine elution at 3 h (left) and 6 h (right) after seeding, whereas (C) IL-22 was only detectable and up-regulated at 6 h after seeding. The data (n = 3) represent the mean ± standard deviation of three independent experiments with unique donors. *p < 0.05 and **p < 0.0001 were determined using an analysis of variance (ANOVA) and Holm–Sidak multiple-comparisons test. Raw data are available in Supplementary Spreadsheet 1.
Discussion
Electrospun biomaterials are excellent candidates for guided in situ tissue regeneration applications because they have a biomimetic structure that can also function as a drug delivery system to modulate acute inflammation (Kelleher and Vacanti, 2010; Minden-Birkenmaier et al., 2017). In recent years, neutrophils have gained attention as an important part of the acute inflammatory response to a biomaterial and the initiation of the tissue repair program (Fetz et al., 2017, 2020; Selders et al., 2017; Minden-Birkenmaier et al., 2020). Neutrophil recruitment, the release of NETs, and the secretion of soluble mediators can have diametrically opposed effects, leading to the resolution of inflammation and healing or chronic inflammation and fibrosis (Selders et al., 2017). Indeed, seminal work in neutrophil biology has highlighted the phenotypic plasticity of neutrophils and their ability to regulate the tissue environment (Buckley et al., 2006; Fridlender et al., 2009; Horckmans et al., 2016). Given the emphasis on endogenous cells for in situ tissue regeneration, regulation of the neutrophil response during acute inflammation is of utmost importance for cell integration and regeneration.
In this work, we developed electrospun PDO biomaterials that elute chloroquine to regulate biomaterial-induced neutrophil activation. PDO is an appealing polymer for electrospun biomaterials because of its biocompatibility, degradation rate, and mechanical properties (Boland et al., 2005; McClure et al., 2009). Additionally, chloroquine is an inexpensive drug historically used to treat malaria that has more recently garnered attention for its immunomodulatory and antithrombotic effects (Wozniacka et al., 2006; Moore et al., 2011; Manic et al., 2014; Schrezenmeier and Dörner, 2020). As a weak base that concentrates in acidic vesicles, it is classified as an inhibitor of lysosomes, lysosomal degradation, and endosomal Toll-like receptor signaling, as well as an inhibitor of autophagy (Sundelin and Terman, 2002; Frustaci et al., 2012; Mauthe et al., 2018). Recently, several groups have shown that chloroquine can be used to inhibit NET release (Smith et al., 2014; Boone et al., 2015, 2018; Germic et al., 2017; Murthy et al., 2019). As it is implicated in tissue fibrosis and thrombosis, NET formation is an appealing pharmacological target, especially in the context of biomaterial-induced NET release (Demers et al., 2012; Chrysanthopoulou et al., 2014; Gregory et al., 2015; Döring et al., 2017; Meng et al., 2017). Our group has shown that neutrophils have an increased propensity to form NETs on the surface of small-fiber electrospun biomaterials, leading to fibrotic encapsulation, whereas large-fiber biomaterials down-regulate NET release and guide tissue integration (Fetz et al., 2017). Jhunjhunwala et al. observed similar outcomes with implanted microcapsules that up-regulated NET release (Jhunjhunwala et al., 2015), thus indicating the need to engineer biomaterials that attenuate NET formation during the acute inflammatory response.
In these experiments, we incorporated chloroquine into the electrospun biomaterials to modulate NET release. At a dose 99.9% lower than the daily oral dose for malaria prophylaxis (Solitro and MacKeigan, 2016; Della Porta et al., 2020), we found that chloroquine eluted from small-fiber biomaterials down-regulated NET release to the level of the large-fiber biomaterials, whereas chloroquine elution from large fibers had no effect on NET release at the early time point. These findings are quite interesting, given that both materials eluted the same concentration of drug with near-identical release profiles. We observed a similar effect upon incorporating Cl-amidine into the electrospun biomaterials, which inhibits the enzyme peptidyl arginine deiminase 4 that is involved in NET formation (Leshner et al., 2012; Fetz et al., 2018). In both cases, these data suggest several mechanisms may be governing the release of NETs on electrospun biomaterials. Additionally, by the later time point, increased NET release was observed on both chloroquine-eluting biomaterials, suggesting a temporal, therapeutic window for inhibiting acute NET release. As the half-life of chloroquine is estimated to be 13–55 days, the observed increase in NET formation at 6 h again suggests that other regulatory mechanisms are involved in biomaterial-induced NET release (Moore et al., 2011; Della Porta et al., 2020). Last, biomaterials that eluted higher concentrations of chloroquine (Supplementary Figure 2) were developed to determine the therapeutic range of NET inhibition by chloroquine. Our data (Supplementary Figure 3) indicate that increasing the chloroquine concentration did not inhibit biomaterial-induced NET release and may have resulted in cytotoxic effects as previously reported (Pliyev and Menshikov, 2012).
Although we and others have shown that chloroquine can inhibit NET release, one group has found that a concentration of chloroquine similar to ours did not inhibit NET formation (Smith et al., 2014; Boone et al., 2015, 2018; Germic et al., 2017; Murthy et al., 2019). When chloroquine was shown to be effective at blocking NET release, neutrophils were stimulated with platelet activating factor, lipopolysaccharide, or our electrospun biomaterials (Smith et al., 2014; Boone et al., 2015). When chloroquine was shown to be ineffective, neutrophils were primed with granulocyte–macrophage CSF and stimulated with C5a, which initiates distinctly different, vital NET release, or the release of mitochondrial NETs (Yousefi et al., 2009; Germic et al., 2017). These data indicate that the therapeutic efficacy of chloroquine for inhibiting NET formation is stimuli dependent and that there may be some overlap in the signaling pathways for biomaterial-induced NET release and other reported triggers of NET release (Farley et al., 2012; Alemán et al., 2016; Khan et al., 2017). Further work is needed to determine the specific signaling pathway involved in biomaterial-induced NET release, but our current data suggest involvement of surface-adsorbed IgG (Fetz et al., 2019).
In addition to inhibiting NETs, chloroquine has been reported to have immunomodulatory effects by altering the secretion of proinflammatory mediators. IL-1β, IL-8, MMP-9, and transforming growth factor β have all been shown to decrease with chloroquine treatment to attenuate inflammation (Delgado et al., 2006; Zhang et al., 2010; Sharma et al., 2017; Fujita et al., 2019). Likewise, in this work, we found that IL-8 and MMP-9 secretions were suppressed on the chloroquine-eluting biomaterials. IL-8 secretion is significantly up-regulated in neutrophils several hours after stimulation before returning to baseline levels, after which monocytes become the major source of IL-8 in vivo (Fujishima et al., 1993). Although neutrophils are necessary for tissue healing (Stirling et al., 2009; Horckmans et al., 2016; Lin et al., 2017), these data suggest that suppression of IL-8 by chloroquine at both early and later time points may down-regulate acute inflammation by reducing pernicious neutrophil chemotaxis (Végran et al., 2011). Similar to IL-8, MMP-9 is robustly secreted from neutrophils during acute inflammation and functions to degrade the ECM for enhanced cell motility (Lu et al., 2011; Deryugina et al., 2014). Consequently, MMP-9 can drive both tissue destruction through excessive ECM degradation and the rapid induction of angiogenesis (Ladwig et al., 2002; Ardi et al., 2007, 2009; Rayment et al., 2008; Liu et al., 2009; Hsu et al., 2015). While angiogenesis is paramount for tissue regeneration, elevated levels of MMP-9 could perpetuate inflammation and cyclic matrix destruction, so its suppression by chloroquine may approximate levels more conducive to regeneration, although this remains to be determined.
In conjunction with proinflammatory signals, we also evaluated the potential impact of chloroquine elution on regenerative signals. To our knowledge, no one has yet to explore this aspect of chloroquine’s anti-inflammatory effects. We found that both HGF and VEGF-A secretions were increased with chloroquine elution from the biomaterials. HGF is a proangiogenic growth factor that promotes regeneration and homeostasis while inhibiting chronic inflammation and fibrosis in various tissues (McCourt et al., 2001; Wislez et al., 2003; Matsumoto et al., 2014; He et al., 2016). Similarly, VEGF-A is the canonical angiogenic signal secreted by neutrophils that has also been shown to recruit a proangiogenic subset of neutrophils (Taichman et al., 1997; Fridlender et al., 2009; Christoffersson et al., 2012, 2017). IL-22 closely followed the trends for HGF and VEGF-A, but was only detectable at the later time point, likely because of the time needed to up-regulate synthesis and secretion (Chen et al., 2016). IL-22 has been shown to be both protective and inflammatory, depending on the disease and model (Zindl et al., 2013; Dyring-Andersen et al., 2017; Zenewicz, 2018). However, it was more recently found to support angiogenesis in the tumor microenvironment by inducing endothelial cell proliferation, survival, and chemotaxis (Protopsaltis et al., 2019). Together, the increased secretion of HGF, VEGF-A, and IL-22 on chloroquine-eluting biomaterials suggests a previously unrecognized aspect of chloroquine’s immunomodulatory effects.
Taken together, our novel incorporation of chloroquine into electrospun biomaterials illustrates the potential therapeutic benefit of this drug for biomaterial-guided, in situ tissue regeneration applications. Although we are the first to repurpose it as an electrospun template additive for tissue engineering applications, chloroquine has been incorporated into a coating for urine catheters to reduce sterile inflammation by reducing neutrophil necrosis and IL-8 secretion (Puyo et al., 2018). Likewise, our data indicate that local delivery through electrospun PDO biomaterials may down-regulate acute neutrophil-driven inflammation while simultaneously up-regulating their regenerative phenotype. The mechanisms underlying chloroquine’s regulation of NET formation and signal secretion as well as its impact on macrophage phenotype and its in vivo efficacy are the subject of further investigations. Additional future work includes evaluation of these biomaterials with platelets and platelet–neutrophil interactions to begin elucidating if chloroquine’s antithrombotic properties are correlated with its inhibition of NET formation (Ouseph et al., 2015; Boone et al., 2018). In conclusion, we have shown that our chloroquine-eluting biomaterials regulate acute neutrophil-driven inflammatory responses in vitro by down-regulating NET release and inflammatory signals while up-regulating regenerative signals. These responses may have synergistic effects that are advantageous for biomaterial-guided in situ tissue regeneration.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding author/s.
Ethics Statement
The studies involving human participants were reviewed by the University of Memphis Institutional Review Board. Written informed consent for participation was not required for this study in accordance with the national legislation and the institutional requirements.
Author Contributions
AF and GB: conceptualization and funding acquisition. AF and SW: methodology and data curation. AF: writing—original draft preparation. AF, SW, and GB: writing—review and editing. GB: supervision and project administration. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Institute of Biomedical Imaging and Bioengineering of the National Institutes of Health under Award No. R15EB022345. Additionally, this work was supported by the National Science Foundation Graduate Research Fellowship Program under Grant no. 1451514.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fbioe.2021.652055/full#supplementary-material
References
Alemán, O. R., Mora, N., Cortes-Vieyra, R., Uribe-Querol, E., and Rosales, C. (2016). Transforming growth factor-β-activated kinase 1 is required for human FcγRIIIb-induced neutrophil extracellular trap formation. Front. Immunol. 7:277. doi: 10.3389/fimmu.2016.00277
Anderson, J. M., Rodriguez, A., and Chang, D. T. (2008). Foreign body reaction to biomaterials. Proc. Semin. Immunol 20, 86–100.
Ardi, V. C., Kupriyanova, T. A., Deryugina, E. I., and Quigley, J. P. (2007). Human neutrophils uniquely release TIMP-free MMP-9 to provide a potent catalytic stimulator of angiogenesis. Proc. Natl. Acad. Sci. U.S.A. 104, 20262–20267. doi: 10.1073/pnas.0706438104
Ardi, V. C., Van den Steen, P. E., Opdenakker, G., Schweighofer, B., Deryugina, E. I., and Quigley, J. P. (2009). Neutrophil MMP-9 proenzyme, unencumbered by TIMP-1, undergoes efficient activation in vivo and catalytically induces angiogenesis via a basic fibroblast growth factor (FGF-2)/FGFR-2 pathway. J. Biol. Chem. 284, 25854–25866. doi: 10.1074/jbc.M109.033472
Boland, E. D., Coleman, B. D., Barnes, C. P., Simpson, D. G., Wnek, G. E., and Bowlin, G. L. (2005). Electrospinning polydioxanone for biomedical applications. Acta Biomater. 1, 115–123. doi: 10.1016/j.actbio.2004.09.003
Boone, B. A., Murthy, P., Miller-Ocuin, J., Doerfler, W. R., Ellis, J. T., Liang, X., et al. (2018). Chloroquine reduces hypercoagulability in pancreatic cancer through inhibition of neutrophil extracellular traps. BMC Cancer. 18:678. doi: 10.1186/s12885-018-4584-2
Boone, B. A., Orlichenko, L., Schapiro, N. E., Loughran, P., Gianfrate, G. C., Ellis, J. T., et al. (2015). The receptor for advanced glycation end products (RAGE) enhances autophagy and neutrophil extracellular traps in pancreatic cancer. Cancer Gene. Ther. 22:326. doi: 10.1038/cgt.2015.21
Brinkmann, V., Reichard, U., Goosmann, C., Fauler, B., Uhlemann, Y., Weiss, D. S., et al. (2004). Neutrophil extracellular traps kill bacteria. Science 303, 1532–1535. doi: 10.1126/science.1092385
Buckley, C. D., Ross, E. A., McGettrick, H. M., Osborne, C. E., Haworth, O., Schmutz, C., et al. (2006). Identification of a phenotypically and functionally distinct population of long-lived neutrophils in a model of reverse endothelial migration. J. Leuko. Biol. 79, 303–311. doi: 10.1189/jlb.0905496
Chen, F., Cao, A., Yao, S., Evans-Marin, H. L., Liu, H., Wu, W., et al. (2016). mTOR mediates IL-23 induction of neutrophil IL-17 and IL-22 production. J. Immunol. 196, 4390–4399. doi: 10.4049/jimmunol.1501541
Chen, G. Y., and Nuñez, G. (2010). Sterile inflammation: sensing and reacting to damage. Nat. Rev. Immunol. 10, 826–837. doi: 10.1038/nri2873
Christoffersson, G., Lomei, J., O’Callaghan, P., Kreuger, J., Engblom, S., and Phillipson, M. (2017). Vascular sprouts induce local attraction of proangiogenic neutrophils. J. Leuko. Biol. 102, 741–751. doi: 10.1189/jlb.1MA0117-018R
Christoffersson, G., Vågesjö, E., Vandooren, J., Lidén, M., Massena, S., Reinert, R. B., et al. (2012). VEGF-A recruits a proangiogenic MMP-9-delivering neutrophil subset that induces angiogenesis in transplanted hypoxic tissue. Blood 120, 4653–4662. doi: 10.1182/blood-2012-04-421040
Chrysanthopoulou, A., Mitroulis, I., Apostolidou, E., Arelaki, S., Mikroulis, D., Konstantinidis, T., et al. (2014). Neutrophil extracellular traps promote differentiation and function of fibroblasts. J. Pathol. 233, 294–307. doi: 10.1002/path.4359
Clark, S. R., Ma, A. C., Tavener, S. A., McDonald, B., Goodarzi, Z., Kelly, M. M., et al. (2007). Platelet TLR4 activates neutrophil extracellular traps to ensnare bacteria in septic blood. Nat. Med. 13, 463–469. doi: 10.1038/nm1565
de Oliveira, S., Reyes-Aldasoro, C. C., Candel, S., Renshaw, S. A., Mulero, V., and Calado, Â (2013). Cxcl8 (IL-8) mediates neutrophil recruitment and behavior in the zebrafish inflammatory response. J. Immunol. 190, 4349–4359. doi: 10.4049/jimmunol.1203266
Degroote, R. L., Weigand, M., Hauck, S. M., and Deeg, C. A. (2020). IL8 and PMA trigger the regulation of different biological processes in granulocyte activation. Front. Immunol. 10:3064. doi: 10.3389/fimmu.2019.03064
Delgado, M. A., Poschet, J. F., and Deretic, V. (2006). Nonclassical pathway of Pseudomonas aeruginosa DNA-induced interleukin-8 secretion in cystic fibrosis airway epithelial cells. Infect. Immun. 74, 2975–2984. doi: 10.1128/IAI.74.5.2975-2984.2006
Della Porta, A., Bornstein, K., Coye, A., Montrief, T., Long, B., and Parris, M. A. (2020). Acute chloroquine and hydroxychloroquine toxicity: a review for emergency clinicians. Am. J. Emerg. Med. 38, 2209–2217. doi: 10.1016/j.ajem.2020.07.030
Demers, M., Krause, D. S., Schatzberg, D., Martinod, K., Voorhees, J. R., Fuchs, T. A., et al. (2012). Cancers predispose neutrophils to release extracellular DNA traps that contribute to cancer-associated thrombosis. Proc. Natl. Acad. Sci. U.S.A. 109, 13076–13081. doi: 10.1073/pnas.1200419109
Deryugina, E. I., Zajac, E., Juncker-Jensen, A., Kupriyanova, T. A., Welter, L., and Quigley, J. P. (2014). Tissue-infiltrating neutrophils constitute the major in vivo source of angiogenesis-inducing MMP-9 in the tumor microenvironment. Neoplasia 16, 771–788. doi: 10.1016/j.neo.2014.08.013
Döring, Y., Soehnlein, O., and Weber, C. (2017). Neutrophil extracellular traps in atherosclerosis and atherothrombosis. Circ. Res. 120, 736–743. doi: 10.1161/CIRCRESAHA.116.309692
Dyring-Andersen, B., Honoré, T. V., Madelung, A., Bzorek, M., Simonsen, S., Clemmensen, S. N., et al. (2017). IL-17A and IL-22 producing neutrophils in psoriatic skin. Br. J. Dermatol. 177:e321. doi: 10.1111/bjd.15533
Farley, K., Stolley, J. M., Zhao, P., Cooley, J., and Remold-O’Donnell, E. (2012). A serpinB1 regulatory mechanism is essential for restricting neutrophil extracellular trap generation. J. Immunol. 189, 4574–4581. doi: 10.4049/jimmunol.1201167
Fetz, A. E., Fantaziu, C. A., Smith, R. A., Radic, M. Z., and Bowlin, G. L. (2019). Surface area to volume ratio of electrospun polydioxanone templates regulates the adsorption of soluble proteins from human serum. Bioengineering 6:78. doi: 10.3390/bioengineering6030078
Fetz, A. E., Neeli, I., Buddington, K. K., Read, R. W., Smeltzer, M. P., Radic, M. Z., et al. (2018). Localized delivery of Cl-amidine from electrospun polydioxanone templates to regulate acute neutrophil NETosis: a preliminary evaluation of the PAD4 inhibitor for tissue engineering. Front. Pharmacol. 9:289. doi: 10.3389/fphar.2018.00289
Fetz, A. E., Neeli, I., Rodriguez, I. A., Radic, M. Z., and Bowlin, G. L. (2017). Electrospun template architecture and composition regulate neutrophil NETosis in vitro and in vivo. Tissue Engin. Part A 23, 1054–1063. doi: 10.1089/ten.TEA.2016.0452
Fetz, A. E., Radic, M. Z., and Bowlin, G. L. (2020). Neutrophils in biomaterial-guided tissue regeneration: Matrix reprogramming for angiogenesis. Tissue Engin. Part B doi: 10.1089/ten.TEB.2020.0028. [Epub ahead of print].
Fridlender, Z. G., Sun, J., Kim, S., Kapoor, V., Cheng, G., Ling, L., et al. (2009). Polarization of tumor-associated neutrophil phenotype by TGF-β:“N1” versus “N2”. TAN. Cancer Cell 16, 183–194. doi: 10.1016/j.ccr.2009.06.017
Frustaci, A., Morgante, E., Antuzzi, D., Russo, M. A., and Chimenti, C. (2012). Inhibition of cardiomyocyte lysosomal activity in hydroxychloroquine cardiomyopathy. Int. J. Cardiol. 157, 117–119. doi: 10.1016/j.ijcard.2012.03.112
Fuchs, T. A., Brill, A., Duerschmied, D., Schatzberg, D., Monestier, M., Myers, D. D., et al. (2010). Extracellular DNA traps promote thrombosis. Proc. Natl. Acad. Sci. U.S.A. 107, 15880–15885. doi: 10.1073/pnas.1005743107
Fujishima, S., Hoffman, A. R., Vu, T., Kim, K. J., Zheng, H., Daniel, D., et al. (1993). Regulation of neutrophil interleukin 8 gene expression and protein secretion by LPS, TNF-α, and IL-1β. J. Cell. Physiol. 154, 478–485. doi: 10.1002/jcp.1041540305
Fujita, Y., Matsuoka, N., Temmoku, J., Furuya, M. Y., Asano, T., Sato, S., et al. (2019). Hydroxychloroquine inhibits IL-1β production from amyloid-stimulated human neutrophils. Arthritis Res. Ther. 21:250. doi: 10.1186/s13075-019-2040-6
Germic, N., Stojkov, D., Oberson, K., Yousefi, S., and Simon, H. U. (2017). Neither eosinophils nor neutrophils require ATG 5-dependent autophagy for extracellular DNA trap formation. Immunology 152, 517–525. doi: 10.1111/imm.12790
Gregory, A. D., Kliment, C. R., Metz, H. E., Kim, K. H., Kargl, J., Agostini, B. A., et al. (2015). Neutrophil elastase promotes myofibroblast differentiation in lung fibrosis. J. Leuko. Biol. 98, 143–152. doi: 10.1189/jlb.3HI1014-493R
Gupta, A. K., Hasler, P., Holzgreve, W., Gebhardt, S., and Hahn, S. (2005). Induction of neutrophil extracellular DNA lattices by placental microparticles and IL-8 and their presence in preeclampsia. Hum. Immunol. 66, 1146–1154.
Gupta, A. K., Joshi, M. B., Philippova, M., Erne, P., Hasler, P., Hahn, S., et al. (2010). Activated endothelial cells induce neutrophil extracellular traps and are susceptible to NETosis-mediated cell death. FEBS Lett. 584, 3193–3197. doi: 10.1016/j.febslet.2010.06.006
Harada, A., Sekido, N., Akahoshi, T., Wada, T., Mukaida, N., and Matsushima, K. (1994). Essential involvement of interleukin-8 (IL-8) in acute inflammation. J. Leuko. Biol. 56, 559–564.
He, M., Peng, A., Huang, X. Z., Shi, D. C., Wang, J. C., Zhao, Q., et al. (2016). Peritumoral stromal neutrophils are essential for c-Met-elicited metastasis in human hepatocellular carcinoma. Oncoimmunology 5:e1219828. doi: 10.1080/2162402X.2016.1219828
Herath, T. D., Larbi, A., Teoh, S. H., James Kirkpatrick, C., and Goh, B. T. (2017). Neutrophil-mediated enhancement of angiogenesis and osteogenesis in a novel triple cell co-culture model with endothelial cells and osteoblasts. J. Tissue Eng. Regen. Med. 12:e1221–e1236. doi: 10.1002/term.2521
Horckmans, M., Ring, L., Duchene, J., Santovito, D., Schloss, M. J., Drechsler, M., et al. (2016). Neutrophils orchestrate post-myocardial infarction healing by polarizing macrophages towards a reparative phenotype. Eur. Heart J. 38, 187–197. doi: 10.1093/eurheartj/ehw002
Hsu, A. T., Barrett, C. D., DeBusk, M. G., Ellson, C. D., Gautam, S., Talmor, D. S., et al. (2015). Kinetics and role of plasma matrix metalloproteinase-9 expression in acute lung injury and the acute respiratory distress syndrome. Shock 44, 128–136. doi: 10.1097/SHK.0000000000000386
Jhunjhunwala, S., Aresta-DaSilva, S., Tang, K., Alvarez, D., Webber, M. J., Tang, B. C., et al. (2015). Neutrophil responses to sterile implant materials. PLoS One. 10:e0137550. doi: 10.1371/journal.pone.0137550
Kaplan, M. J., and Radic, M. (2012). Neutrophil extracellular traps: Double-edged swords of innate immunity. J. Immunol. 189, 2689–2695. doi: 10.4049/jimmunol.1201719
Kelleher, C. M., and Vacanti, J. P. (2010). Engineering extracellular matrix through nanotechnology. J. R. Soc. Interface. 7, S717–S729. doi: 10.1098/rsif.2010.0345.focu
Khan, M. A., Farahvash, A., Douda, D. N., Licht, J.-C., Grasemann, H., Sweezey, N., et al. (2017). JNK activation turns on LPS-and Gram-negative bacteria-induced NADPH oxidase-dependent suicidal NETosis. Sci. Rep. 7:3409. doi: 10.1038/s41598-017-03257-z
Lacy, P. (2006). Mechanisms of degranulation in neutrophils. Allergy Asthma Clin. Immunol. 2:98. doi: 10.1186/1710-1492-2-3-98
Ladwig, G. P., Robson, M. C., Liu, R., Kuhn, M. A., Muir, D. F., and Schultz, G. S. (2002). Ratios of activated matrix metalloproteinase-9 to tissue inhibitor of matrix metalloproteinase-1 in wound fluids are inversely correlated with healing of pressure ulcers. Wound Repair Regen. 10, 26–37. doi: 10.1046/j.1524-475x.2002.10903.x
Lammermann, T., Afonso, P. V., Angermann, B. R., Wang, J. M., Kastenmuller, W., Parent, C. A., et al. (2013). Neutrophil swarms require LTB4 and integrins at sites of cell death in vivo. Nature 498, 371–375. doi: 10.1038/nature12175
Lannutti, J., Reneker, D., Ma, T., Tomasko, D., and Farson, D. (2007). Electrospinning for tissue engineering scaffolds. Mater. Sci. Eng. C 27, 504–509. doi: 10.1016/j.msec.2006.05.019
Leshner, M., Wang, S., Lewis, C., Zheng, H., Chen, X. A., Santy, L., et al. (2012). PAD4 mediated histone hypercitrullination induces heterochromatin decondensation and chromatin unfolding to form neutrophil extracellular trap-like structures. Front. Immunol. 3:307. doi: 10.3389/fimmu.2012.00307
Li, Q., Ma, L., and Gao, C. (2015). Biomaterials for in situ tissue regeneration: development and perspectives. J. Mater. Chem. 3, 8921–8938. doi: 10.1039/C5TB01863C
Lima, T., Feitosa, R., dos Santos-Silva, E., dos Santos-Silva, A., Siqueira, E., Machado, P., et al. (2018). Improving encapsulation of hydrophilic chloroquine diphosphate into biodegradable nanoparticles: a promising approach against herpes virus simplex-1 infection. Pharmaceutics 10:255. doi: 10.3390/pharmaceutics10040255
Lin, R. Z., Lee, C. N., Moreno-Luna, R., Neumeyer, J., Piekarski, B., Zhou, P., et al. (2017). Host non-inflammatory neutrophils mediate the engraftment of bioengineered vascular networks. Nat. Biotechnol. 1:0081. doi: 10.1038/s41551-017-0081
Liu, Y., Min, D., Bolton, T., Nubé, V., Twigg, S. M., Yue, D. K., et al. (2009). Increased matrix metalloproteinase-9 predicts poor wound healing in diabetic foot ulcers. Diabetes Care 32, 117–119. doi: 10.2337/dc08-0763
Lu, P., Takai, K., Weaver, V. M., and Werb, Z. (2011). Extracellular matrix degradation and remodeling in development and disease. Cold Spring Harb. Perspect. Biol. 3:a005058. doi: 10.1101/cshperspect.a005058
Manic, G., Obrist, F., Kroemer, G., Vitale, I., and Galluzzi, L. (2014). Chloroquine and hydroxychloroquine for cancer therapy. Mol. Cell Oncol. 1:e29911. doi: 10.4161/mco.29911
Martinez, F. O., Sironi, M., Vecchi, A., Colotta, F., Mantovani, A., and Locati, M. (2004). IL-8 induces a specific transcriptional profile in human neutrophils: synergism with LPS for IL-1 production. Eur. J. Immunol. 34, 2286–2292. doi: 10.1002/eji.200324481
Martinod, K., Demers, M., Fuchs, T. A., Wong, S. L., Brill, A., Gallant, M., et al. (2013). Neutrophil histone modification by peptidylarginine deiminase 4 is critical for deep vein thrombosis in mice. Proc. Natl. Acad. Sci. U.S.A. 110, 8674–8679. doi: 10.1073/pnas.1301059110
Matsumoto, K., Funakoshi, H., Takahashi, H., and Sakai, K. (2014). HGF–Met pathway in regeneration and drug discovery. Biomedicine 2, 275–300. doi: 10.3390/biomedicines2040275
Maugeri, N., Brambilla, M., Camera, M., Carbone, A., Tremoli, E., Donati, M., et al. (2006). Human polymorphonuclear leukocytes produce and express functional tissue factor upon stimulation 1. J. Thromb. Haemost. 4, 1323–1330. doi: 10.1111/j.1538-7836.2006.01968.x
Mauthe, M., Orhon, I., Rocchi, C., Zhou, X., Luhr, M., Hijlkema, K.-J., et al. (2018). Chloroquine inhibits autophagic flux by decreasing autophagosome-lysosome fusion. Autophagy 14, 1435–1455. doi: 10.1080/15548627.2018.1474314
McClure, M. J., Sell, S. A., Simpson, D. G., and Bowlin, G. L. (2009). Electrospun polydioxanone, elastin, and collagen vascular scaffolds: uniaxial cyclic distension. J. Eng. Fibers Fabr. 4, 18–25. doi: 10.1177/155892500900400204
McCourt, M., Wang, J., Sookhai, S., and Redmond, H. (2001). Activated human neutrophils release hepatocyte growth factor/scatter factor. Eur. J. Surg. Oncol. 27, 396–403. doi: 10.1053/ejso.2001.1133
Meng, H., Yalavarthi, S., Kanthi, Y., Mazza, L. F., Elfline, M. A., Luke, C. E., et al. (2017). In vivo role of neutrophil extracellular traps in antiphospholipid antibody–mediated venous thrombosis. Arthritis Rheum. 69, 655–667. doi: 10.1002/art.39938
Minden-Birkenmaier, B., Selders, G., Fetz, A., Gehrmann, C., and Bowlin, G. (2017). “Electrospun systems for drug delivery,” in Electrospun Materials for Tissue Engineering and Biomedical Applications, eds T. Uyar and E. Kny (Amsterdam: Elsevier), 117–145.
Minden-Birkenmaier, B. A., Smith, R. A., Radic, M. Z., van der Merwe, M., and Bowlin, G. L. (2020). Manuka honey reduces NETosis on an electrospun template within a therapeutic window. Polymers 12:1430. doi: 10.3390/polym12061430
Moore, B. R., Page-Sharp, M., Stoney, J. R., Ilett, K. F., Jago, J. D., and Batty, K. T. (2011). Pharmacokinetics, pharmacodynamics, and allometric scaling of chloroquine in a murine malaria model. Antimicrob. Agents Chemother. 55, 3899–3907. doi: 10.1128/AAC.00067-11
Murthy, P., Singhi, A. D., Ross, M. A., Loughran, P., Paragomi, P., Papachristou, G. I., et al. (2019). Enhanced neutrophil extracellular trap (NET) formation in acute pancreatitis contributes to disease severity and is reduced by chloroquine. Front. Immunol. 10:28. doi: 10.3389/fimmu.2019.00028
Nagaoka, I., and Hirota, S. (2000). Increased expression of matrix metalloproteinase-9 in neutrophils in glycogen-induced peritoneal inflammation of guinea pigs. Inflamm. Res. 49, 55–62. doi: 10.1007/s000110050559
Neeli, I., Khan, S. N., and Radic, M. (2008). Histone deimination as a response to inflammatory stimuli in neutrophils. J. Immunol. 180, 1895–1902. doi: 10.4049/jimmunol.180.3.1895
Neeli, I., and Radic, M. (2013). Opposition between PKC isoforms regulates histone deimination and neutrophil extracellular chromatin release. Front. Immunol. 4:38. doi: 10.3389/fimmu.2013.00038
Ouseph, M. M., Huang, Y., Banerjee, M., Joshi, S., MacDonald, L., Zhong, Y., et al. (2015). Autophagy is induced upon platelet activation and is essential for hemostasis and thrombosis. Blood 126, 1224–1233. doi: 10.1182/blood-2014-09-598722
Parker, H., Albrett, A. M., Kettle, A. J., and Winterbourn, C. C. (2012). Myeloperoxidase associated with neutrophil extracellular traps is active and mediates bacterial killing in the presence of hydrogen peroxide. J. Leuko. Biol. 91, 369–376. doi: 10.1189/jlb.0711387
Phillipson, M., and Kubes, P. (2019). The healing power of neutrophils. Trends Immunol. 40, 635–647. doi: 10.1016/j.it.2019.05.001
Pliyev, B. K., and Menshikov, M. (2012). Differential effects of the autophagy inhibitors 3-methyladenine and chloroquine on spontaneous and TNF-α-induced neutrophil apoptosis. Apoptosis 17, 1050–1065. doi: 10.1007/s10495-012-0738-x
Protopsaltis, N. J., Liang, W., Nudleman, E., and Ferrara, N. (2019). Interleukin-22 promotes tumor angiogenesis. Angiogenesis 22, 311–323. doi: 10.1007/s10456-018-9658-x
Puyo, C. A., Earhart, A., Staten, N., Huang, Y., Desai, A., Lai, H., et al. (2018). Mitochondrial DNA induces Foley catheter related bladder inflammation via Toll-like receptor 9 activation. Sci. Rep. 8:6377. doi: 10.1038/s41598-018-24818-w
Rayment, E. A., Upton, Z., and Shooter, G. K. (2008). Increased matrix metalloproteinase-9 (MMP-9) activity observed in chronic wound fluid is related to the clinical severity of the ulcer. Br. J. Dermatol. 158, 951–961. doi: 10.1111/j.1365-2133.2008.08462.x
Riehl, D. R., Roewe, J., Klebow, S., Esmon, N. L., Eming, S., Colucci, G., et al. (2016). Neutrophil extracellular traps drive bleomycin-induced lung fibrosis by regulating TGFβ1-dependent interactions of platelets and macrophages. FASEB J. 30:665.13. doi: 10.1096/fasebj.30.1_supplement.50.1
Roldan, E. Q., Biasiotto, G., Magro, P., and Zanella, I. (2020). The possible mechanisms of action of 4-aminoquinolines (chloroquine/hydroxychloroquine) against Sars-Cov-2 infection (COVID-19): a role for iron homeostasis? Pharmacol. Res. 158:104904. doi: 10.1016/j.phrs.2020.104904
Schrezenmeier, E., and Dörner, T. (2020). Mechanisms of action of hydroxychloroquine and chloroquine: implications for rheumatology. Nat. Rev. Rheumatol 16, 155–166. doi: 10.1038/s41584-020-0372-x
Selders, G. S., Fetz, A. E., Radic, M. Z., and Bowlin, G. L. (2017). An overview of the role of neutrophils in innate immunity, inflammation and host-biomaterial integration. Regen. Biomat. 4, 55–68. doi: 10.1093/rb/rbw041
Sharma, P., Yi, R., Nayak, A. P., Wang, N., Tang, F., Knight, M. J., et al. (2017). Bitter taste receptor agonists mitigate features of allergic asthma in mice. Sci. Rep. 7:46166. doi: 10.1038/srep46166
Smith, C. K., Vivekanandan-Giri, A., Tang, C., Knight, J. S., Mathew, A., Padilla, R. L., et al. (2014). Neutrophil extracellular trap–derived enzymes oxidize high-density lipoprotein: an additional proatherogenic mechanism in systemic lupus erythematosus. Arthritis Rheum. 66, 2532–2544. doi: 10.1002/art.38703
Solitro, A. R., and MacKeigan, J. P. (2016). Leaving the lysosome behind: novel developments in autophagy inhibition. Future Med. Chem. 8, 73–86. doi: 10.4155/fmc.15.166
Stirling, D. P., Liu, S., Kubes, P., and Yong, V. W. (2009). Depletion of Ly6G/Gr-1 leukocytes after spinal cord injury in mice alters wound healing and worsens neurological outcome. J. Neurosci. 29, 753–764. doi: 10.1523/JNEUROSCI.4918-08.2009
Sun, X. Y., Nobles, L. R., Borner, H. G., and Spontak, R. J. (2008). Field-driven surface segregation of biofunctional species on electrospun PMMA/PEO microfibers. Macromol. Rapid Comm. 29, 1455–1460. doi: 10.1002/marc.200800163
Sundelin, S. P., and Terman, A. (2002). Different effects of chloroquine and hydroxychloroquine on lysosomal function in cultured retinal pigment epithelial cells. APMIS 110, 481–489. doi: 10.1034/j.1600-0463.2002.100606.x
Taichman, N. S., Young, S., Cruchley, A. T., Taylor, P., and Paleolog, E. (1997). Human neutrophils secrete vascular endothelial growth factor. J. Leuko. Biol. 62, 397–400. doi: 10.1002/jlb.62.3.397
Végran, F., Boidot, R., Michiels, C., Sonveaux, P., and Feron, O. (2011). Lactate influx through the endothelial cell monocarboxylate transporter MCT1 supports an NF-κB/IL-8 pathway that drives tumor angiogenesis. Cancer Res. 71, 2550–2560. doi: 10.1158/0008-5472.CAN-10-2828
Wislez, M., Rabbe, N., Marchal, J., Milleron, B., Crestani, B., Mayaud, C., et al. (2003). Hepatocyte growth factor production by neutrophils infiltrating bronchioloalveolar subtype pulmonary adenocarcinoma: role in tumor progression and death. Cancer Res. 63, 1405–1412.
Wozniacka, A., Lesiak, A., Narbutt, J., McCauliffe, D., and Sysa-Jedrzejowska, A. (2006). Chloroquine treatment influences proinflammatory cytokine levels in systemic lupus erythematosus patients. LUPUS 15, 268–275. doi: 10.1191/0961203306lu2299oa
Yousefi, S., Mihalache, C., Kozlowski, E., Schmid, I., and Simon, H. U. (2009). Viable neutrophils release mitochondrial DNA to form neutrophil extracellular traps. Cell Death Differ. 16, 1438–1444. doi: 10.1038/cdd.2009.96
Zenewicz, L. A. (2018). IL-22: there is a gap in our knowledge. Immunohorizons 2, 198–207. doi: 10.4049/immunohorizons.1800006
Zhang, Q., Itagaki, K., and Hauser, C. J. (2010). Mitochondrial DNA is released by shock and activates neutrophils via p38 map kinase. Shock 34, 55–59. doi: 10.1097/SHK.0b013e3181cd8c08
Zindl, C. L., Lai, J. F., Lee, Y. K., Maynard, C. L., Harbour, S. N., Ouyang, W., et al. (2013). IL-22–producing neutrophils contribute to antimicrobial defense and restitution of colonic epithelial integrity during colitis. Proc. Natl. Acad. Sci. U.S.A. 110, 12768–12773. doi: 10.1073/pnas.1300318110
Keywords: tissue regeneration, electrospinning, tissue engineering, neutrophils, neutrophil extracellular traps, NETs, chloroquine
Citation: Fetz AE, Wallace SE and Bowlin GL (2021) Electrospun Polydioxanone Loaded With Chloroquine Modulates Template-Induced NET Release and Inflammatory Responses From Human Neutrophils. Front. Bioeng. Biotechnol. 9:652055. doi: 10.3389/fbioe.2021.652055
Received: 11 January 2021; Accepted: 19 March 2021;
Published: 27 April 2021.
Edited by:
Ryang Hwa Lee, Texas A&M University, United StatesReviewed by:
Yves Bayon, A Medtronic Company, FranceMasoud Mozafari, University of Toronto, Canada
Judite Barbosa, University of Porto, Portugal
Copyright © 2021 Fetz, Wallace and Bowlin. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Gary L. Bowlin, Z2xib3dsaW5AbWVtcGhpcy5lZHU=
 Allison E. Fetz
Allison E. Fetz Shannon E. Wallace
Shannon E. Wallace Gary L. Bowlin
Gary L. Bowlin