- 1Key Laboratory of Growth Regulation and Translational Research of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou, China
- 2Westlake Laboratory of Life Sciences and Biomedicine, Hangzhou, China
- 3Institute of Biology, Westlake Institute for Advanced Study, Hangzhou, China
- 4Key Laboratory of Structural Biology of Zhejiang Province, School of Life Sciences, Westlake University, Hangzhou, China
Both Hippo signaling pathways and cell polarity regulation are critical for cell proliferation and the maintenance of tissue homeostasis, despite the well-established connections between cell polarity disruption and Hippo inactivation, the molecular mechanism by which aberrant cell polarity induces Hippo-mediated overgrowth remains underexplored. Here we use Drosophila wing discs as a model and identify the Wnd-Nmo axis as an important molecular link that bridges loss-of-cell polarity-triggered Hippo inactivation and overgrowth. We show that Wallenda (Wnd), a MAPKKK (mitogen-activated protein kinase kinase kinase) family member, is a novel regulator of Hippo pathways in Drosophila and that overexpression of Wnd promotes growth via Nemo (Nmo)- mediated Hippo pathway inactivation. We further demonstrate that both Wnd and Nmo are required for loss-of-cell polarity-induced overgrowth and Hippo inactivation. In summary, our findings provide a novel insight on how cell polarity loss contributes to overgrowth and uncover the Wnd-Nmo axis as an essential additional branch that regulates Hippo pathways in Drosophila.
Introduction
Proper control of cell proliferation is fundamental for correct organ development, disruption of which would cause tumorigenesis. Drosophila imaginal discs, the precursor of the adult organ during the larval stage, are a powerful model to study tissue growth, and numerous signaling pathways essential for cell proliferation have been uncovered using this system (Gokhale and Pfleger, 2019; Gou et al., 2020), including the Hippo pathway, a key regulator of cell proliferation and organ size (Pan et al., 2018; Zheng and Pan, 2019). The core components of the Hippo signaling pathway consist of serine/threonine kinases Hippo (Hpo) and Warts (Wts) and the transcriptional coactivator Yorkie (Yki) (Justice et al., 1995; Xu et al., 1995; Harvey et al., 2003; Pantalacci et al., 2003; Udan et al., 2003; Wu et al., 2003; Huang et al., 2005). Upon phosphorylation by Hpo, Wts is activated and subsequently phosphorylates Yki to restrict its nuclear entrance, thereby preventing the transcription of pro-proliferative target genes, including expanded (ex), bantam (ban), and Cyclin E (cycE) (Pan, 2010; Snigdha et al., 2019; Zheng and Pan, 2019). Recent advances on this evolutionary conserved pathway have revealed comprehensive roles of Hippo signaling in regulating a wide range of biological functions, ranging from cell adhesion and mechanical tension to regeneration and immune surveillance (Fallahi et al., 2016; Chen, 2019; Ma et al., 2019; Zheng and Pan, 2019).
Cell polarity maintenance is essential for tissue homeostasis, and dysregulation of apical-basal polarity has been linked to various developmental disorders and cancer (Wodarz and Nathke, 2007). Studies in Drosophila have revealed that several apical-basal cell polarity modules regulate tissue growth via the Hippo pathway, including lethal-2-giant larvae (Lgl)-atypical protein kinase C (aPKC), Crumbs (Crb), and Scribbled (Scrib)/Disc large (Dlg) (Richardson and Portela, 2017). We recently identified the E3 ubiquitin ligase POSH (Plenty of SH3s) as a key regulator of Hippo pathway, which integrates the signal from the cell polarity protein Crb to negatively regulate Ex (Expanded)-mediated Hippo activation in Drosophila (Ma et al., 2018). We also identified Wallenda (Wnd), a member of mitogen activated protein kinase kinase kinase (MAPKKK) family, as an essential downstream component that links loss-of-polarity-induced cell invasion phenotype to JNK pathway activation (Ma et al., 2016). Although recent studies on MAP3K13, the human homolog of Wnd, have revealed its tumor promoting roles in head and neck squamous cell carcinoma (HNSCC) and hepatocellular carcinoma (HCC) patients (Edwards et al., 2017; Zhang et al., 2020), the mechanism by which Wnd/MAP3K13 regulates tissue growth remains underexplored.
In this study, we uncovered Wnd as a novel regulator of Hippo signaling. Our results showed that overexpression of Wnd induces cell proliferation in the Drosophila wing imaginal disc via inactivating the Hippo signaling. Knockdown of wnd impedes Rho1 expression or loss-of-polarity induced Hippo inactivation. We further identified Nemo (Nmo) as an essential downstream mediator of Wnd in regulating Hippo signaling. Our data demonstrate that Wnd-Nmo represents a new axis that bridges cell-polarity-loss-induced growth and Hippo pathway inactivation in Drosophila.
Results and Discussion
Wnd/MAP3K13 Negatively Regulates Hippo Signaling
Our previous studies have revealed that both Rho1 and Wnd are essential for tumorigenesis by inducing JNK signaling mediated cell death and cell invasion in wing imaginal discs (Ma et al., 2015b, 2016). Interestingly, we noticed that Rho1-induced actin accumulation could only be impeded by depletion of Wnd but remained unaffected by complete inhibition of JNK signaling, suggesting that Wnd has additional roles other than the JNK pathway activation. To further explore the role of Wnd in regulating tumorigenesis, we examined the potential role of MAP3K13 (human homolog of Wnd) through data mining using several open-access online servers. We found that copy number variants (CNV) of MAP3K13 are dysregulated in various human cancers (Figure 1A), especially in lung squamous cell carcinoma (LUSC) and ovarian serous cystadenocarcinoma (OV). Accordingly, transcripts of MAP3K13 are significantly up regulated in LUSC and OV (Figure 1C) and the high expression level of MAP3K13 is correlated with low survival probability in endometrial cancer patients (Figure 1B). Together, these results indicate that MAP3K13 is a potential oncogene.
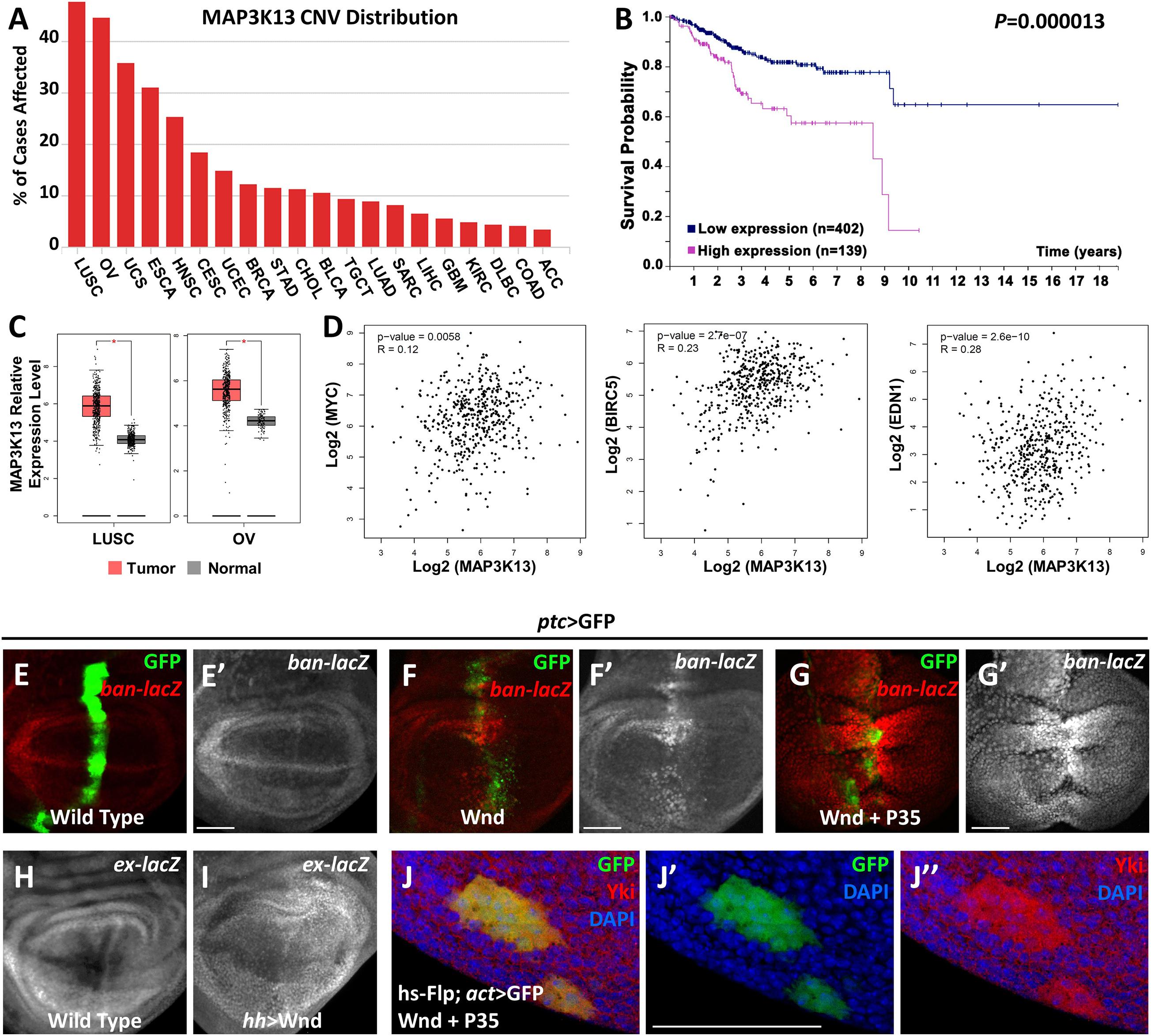
Figure 1. MAP3K13/Wnd activates expression of Hippo pathway target genes. (A) The distribution of copy number variation (CNV) of MAP3K13 across different cancer types. (B) Analysis of the prognostic significance of high and low expression level of MAP3K13 in endometrial cancer. (C) Box plots showing the relative expression level of MAP3K13 in lung squamous cell carcinoma (LUSC) and ovarian serous cystadenocarcinoma (OV). (D) Expression correlation between MAP3K13 and MYC, BIRC5, or EDN1, three target genes downstream of the transcriptional co-activators YAP/TAZ, in LUSC. (E–J″) Wing imaginal discs from third instar larva are shown. Compared with wild type controls (E,E′), expression of Wnd for 2–3 days (F,F′) or co-expression of wnd and P35 (G,G′) driven by ptc-Gal4 (GFP positive area) elevates ban expression level. Expression of Wnd in the posterior region of wing disc elevates ex expression level (I), compared with wild type controls (H). (J–J″) Yki nuclear localization is increased in heat shock-induced clones expressing wnd. Scale bars, 50 μm.
Next, we tried to further address the function of Wnd in tissue growth control in vivo using Drosophila. Given that ectopic wnd expression can induce the up-regulation of wingless (wg) and cycE (Ma et al., 2016), two known downstream target genes of the Hippo pathway, we tested the possibility that wnd could regulate the Hippo pathway in Drosophila. We examined the transcriptional change of two additional Hippo signaling reporters, ban and ex (Cho et al., 2006; Thompson and Cohen, 2006), and found that upon Wnd overexpression under ptc promoter along the anterior-posterior boundary of wing imaginal discs, an obvious up-regulation of ban-lacZ was seen (Figures 1F,F′), compared with the wild-type control (Figures 1E,E′). Given that ectopic expression of Wnd also leads to apoptosis as reported previously (Ma et al., 2015b), we inhibited apoptosis simultaneously by co-expression of P35 with wnd and observed a stronger up-regulation of ban-lacZ (Figures 1G,G′). Similarly, overexpression of wnd by hh-Gal4 in the posterior region of wing imaginal discs significantly up-regulated ex-lacZ level (Figures 1H,I). More importantly, we also observed strong nuclear Yki localization in heat-shock-induced wnd expressing clones compared with endogenous control outside the GFP positive clones (Figures 1J–J″). Furthermore, our data mining results suggested that the up-regulation of MAP3K13 in LUSC is positively correlated with increased expression of MYC, BIRC5, and EDN1 (Figure 1D), three known target genes downstream of YAP/TAZ (Choi et al., 2018), indicating a potential conserved role of MAP3K13 in regulating Hippo pathway in human cancers. Interestingly, we also observed an increased level of cytoplasmic Yki, suggesting that there is an overall increase of the cellular Yki amount, which we could not explain at this point, and for which further investigation is required to dissect the underlying mechanism. Therefore, we conclude that Wnd enhances Yki nuclear localization and up-regulates Hippo pathway target genes.
Wnd Acts Downstream of Cell Polarity Loss-Rho1 Axis
Our previous study revealed a physical interaction between Wnd and the Rho GTPase Rho1 in regulating cell invasion (Ma et al., 2016), and we identified Rho1 as an important regulator of Hippo signaling-mediated growth (Ma et al., 2015a). These clues raised the possibility that Wnd and Rho1 are somehow linked in the regulation of the Hippo pathway. Therefore, we investigated whether Wnd is also required for Rho1-mediated Hippo inactivation in Drosophila. Overexpression of Rho1 with P35 by ptc-Gal4 in wing imaginal discs induces obvious cell proliferation and enhanced Yki nuclear localization (Figures 2B,B′), both of which are significantly impeded by knockdown of wnd (Figures 2C,C′). Consistently with this, we found that compared with the wild type control (Figures 2D,D′), knockdown of wnd also reduces Rho1 overexpression-induced ban-lacZ up-regulation (Figures 2E–F′ and Supplementary Figure 2), suggesting that Wnd is required for Rho1-induced Hippo signaling inactivation.
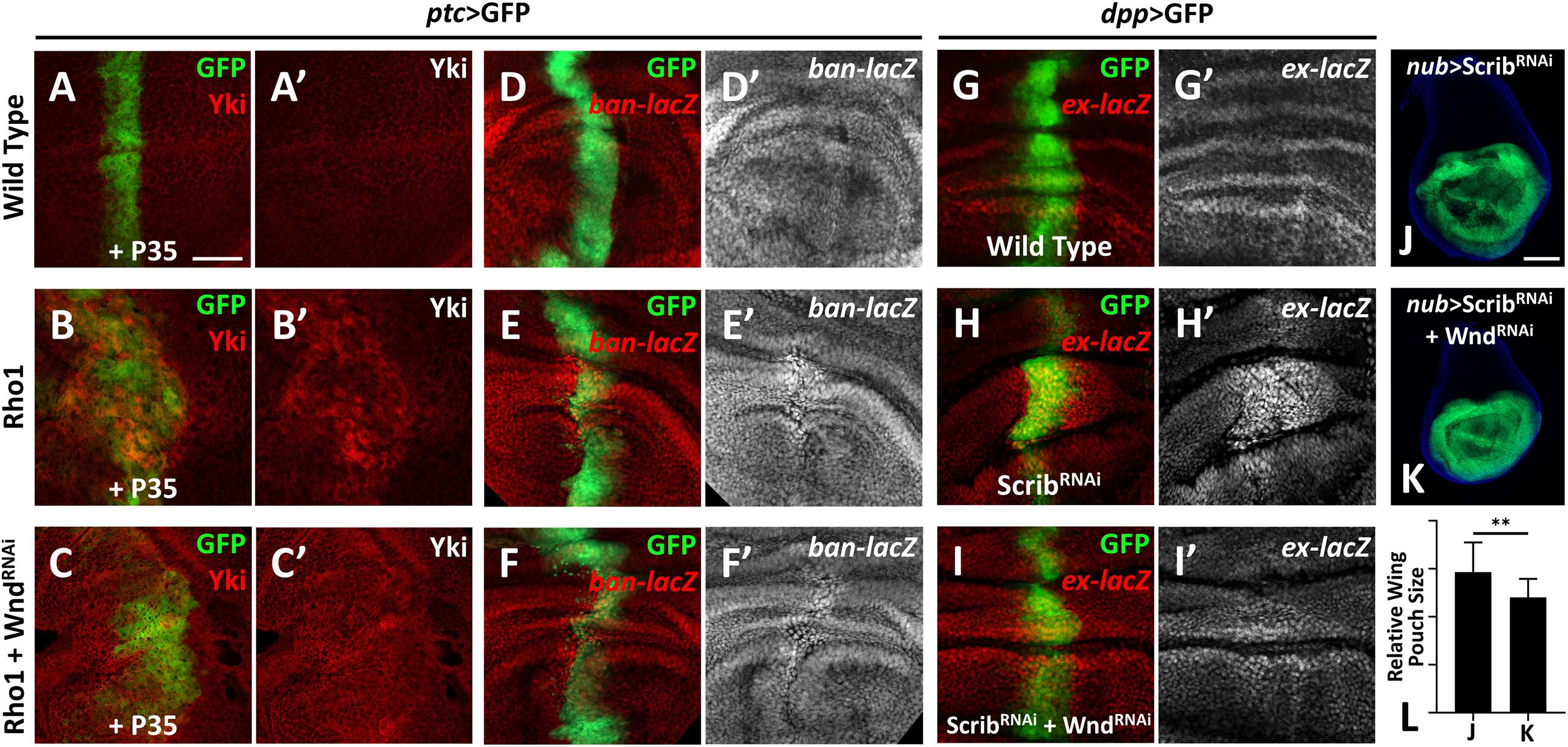
Figure 2. Wnd is essential for Rho1 activation and cell polarity loss-induced Hippo inactivation. (A–K) Wing imaginal discs from third instar larva with genes or RNA is expressed in GFP positive area are shown. (A–C′) Compared with wild type controls (A,A′), expression of Rho1 together with P35 enhances Yki nuclear localization (B,B′), which is rescued by knocking down wnd (C,C′). (D–F′) Compared with wild type controls (D,D′), expression of Rho1 induced ban upregulation (E,E′) is suppressed by knocking down wnd (F,F′). (G–I′) Compared with wild type controls (G,G′), scribble (scrib) knockdown upregulates ex transcription (H,H′) and this is impeded by wnd knockdown (I,I′). (J) Knocking down scrib in the wing pouch region causes tissue overgrowth (J), which is suppressed by wnd knockdown (K). Statistical analysis of (J,K) is shown in (L). Scale bars for (A–I′), 50 μm; for (J,K), 100 μm. **P < 0.01. (A Student t-test was used to calculate statistical significance, mean + SD, n = 15).
Loss of scrib has been reported to upregulate Yki target genes (Verghese et al., 2012). To test whether Wnd is also required for scrib-loss induced Hippo inactivation, we monitored ex-lacZ level in wing imaginal discs. Knockdown of scrib by dpp-Gal4 induces evident upregulation of ex-lacZ (Figures 2H,H′), in comparison with the wild type control (Figures 2G,G′), while knockdown simultaneously of wnd reduces the ex-lacZ level to a certain degree (Figures 2I,I′). Additionally, knockdown of scrib by nub-Gal4 leads to significant overgrowth of the wing pouch region (Figure 2J), which is also inhibited by wnd knockdown (Figures 2K,L). In summary, these results indicate that Wnd acts downstream of cell polarity-loss-induced Hippo inactivation.
Nmo Genetically Acts Downstream of Wnd
Next, we further dissected the potential mechanism by which wnd overexpression inactivates Hippo signaling. As a MAPKKK, logically, Wnd may activate certain protein kinase(s) that are known to genetically interact with Hippo signaling. However, the classical known kinases of the Hippo pathway are all negative regulators of Yki, suggesting that Wnd might act through a non-canonical kinase target to regulate the Hippo pathway. The only exception known so far is a MAPK family protein named Nmo, a serine/threonine protein kinase essential for cell death, planar cell polarity, neuronal function, and circadian clock regulation (Mirkovic et al., 2002; Merino et al., 2009; Yu et al., 2011; Collu et al., 2018). It is recently reported that Nemo-Like Kinase (NLK, human homolog of Nmo) phosphorylates YAP and subsequently positively regulate its transcriptional activity (Moon et al., 2017). It is worth noting that both Wnd and Nmo are essential for the overgrowth phenotype of neuromuscular junction (NMJ) caused by a mutation in a ubiquitin E3 ligase named Highwire (Collins et al., 2006; Wu et al., 2007; Merino et al., 2009), indicating a potential genetic interaction between wnd and nmo. Therefore, we examined whether Nmo is genetically involved in Wnd-mediated Yki activation.
Given that wnd overexpression induces cell death by activating JNK pathway, we co-expressed P35 to inhibit Wnd-induced cell death. Overexpression of P35 alone by ptc-Gal4 leads to no change on Wg level comparing to the wild type (Figures 3A–B′), iwhile co-expression of wnd and P35 and observed a significant up-regulation of wg expression along the ptc stripe (GFP positive) (Figures 3D,D′), which was dramatically reduced by simultaneously knocking down nmo (Figures 3E,E′). Strikingly, we found that nmo knockdown not only reduced the nuclear Yki accumulation induced by co-expression of wnd and P35, but also caused sharp decrease of Yki protein level (Figures 3I–J″), whereas knockdown of nmo alone did not cause obvious phenotypes (Figures 3C,C′,H–H″, and Supplementary Figure 4). Since the increased nuclear Yki localization causally leads to cell proliferation, we also examined cell proliferation by PH3 staining. We found that though there is no significant difference of proliferation rate in P35 overexpression and nmo knockdown wing discs compared with the wild type control (Figures 3P–R′,U), the increased proliferation caused by wnd and P35 co-expression was significantly suppressed by nmo knockdown (Figures 3S–U). Consistent with genetic interactions between wnd and nmo in Drosophila, we found that upregulation of MAP3K13 in LUSC and OV are positively correlated with increased expression of NLK (Supplementary Figure 1). Yki level reduction would induce apoptosis by downregulating the expression of diap1 (Huang et al., 2005). Compared with controls (Figures 3K,K″), overexpression of P35 (Figures 3L,L″) or knockdown of nmo (Figures 3M,M″) has no significant changes on apoptosis. However, we also observed non-autonomous apoptosis when Wnd and P35 are co-overexpressed (Figures 3N,N′), which is commonly seen when JNK signaling is hyperactivated (Uhlirova et al., 2005; Perez-Garijo et al., 2013; La Marca and Richardson, 2020). Surprisingly, we found that Nmo inhibition caused a shift toward a strong autonomous apoptosis increase in the wnd and P35 co-expression wing discs (Figures 3O,O′). Given that loss of Yki activity facilitates cell death (Huang et al., 2005; Liu et al., 2016), we speculate that in the Wnd and P35 co-expressing wing disc, nmo knockdown leads to the downregulation of Yki level and therefore increases apoptosis. Taken together, these data indicate that Nmo is an essential downstream effector of Wnd in regulating Hippo signaling.
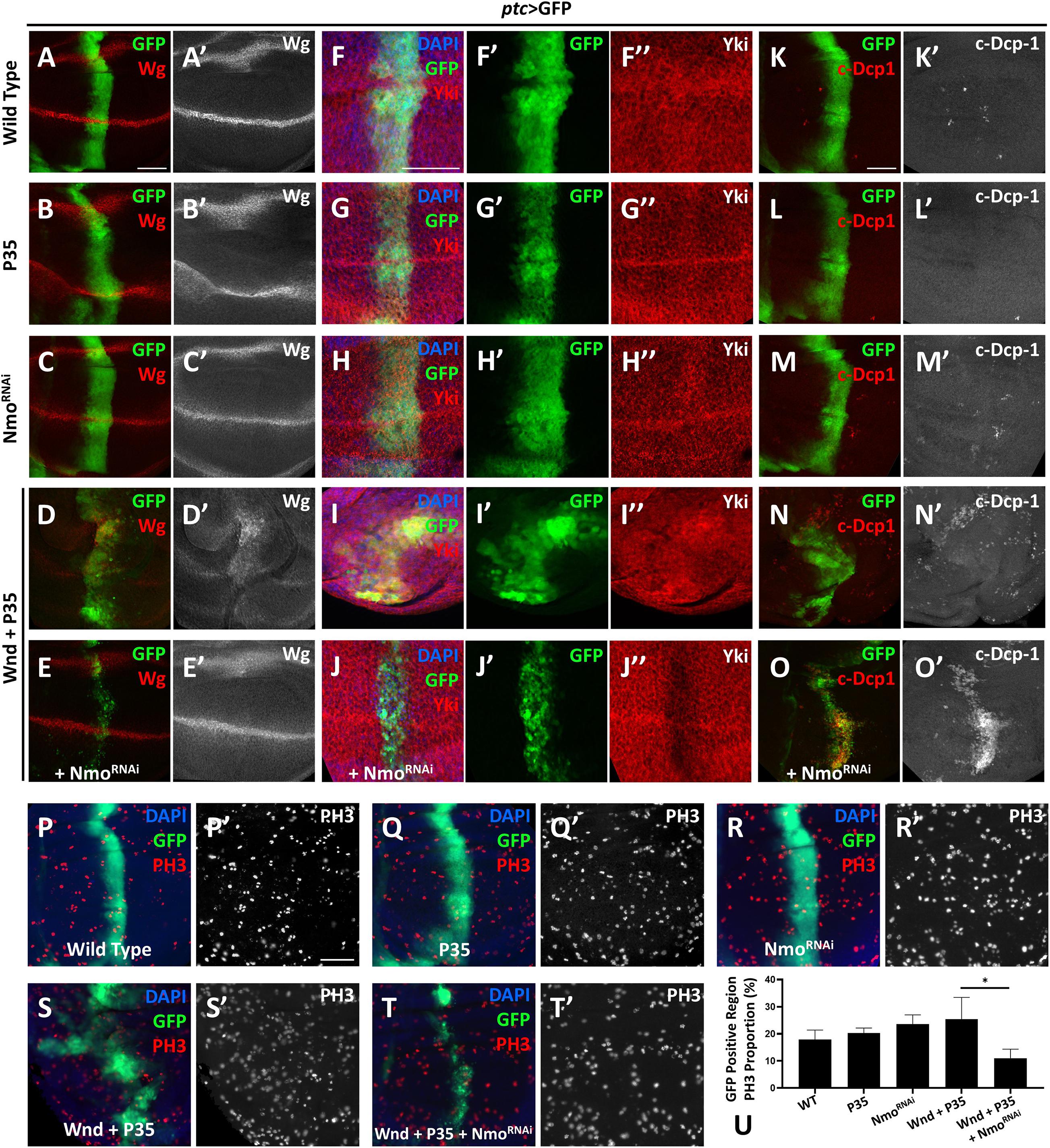
Figure 3. Nmo acts genetically downstream of wnd. Wing imaginal discs from third instar larva with genes or RNAi expressed in GFP positive area driven by ptc-Gal4 are shown. (A–E′) Compared with wild type (A,A′) or P35 expression alone control (B,B′), co-expression of wnd and P35 elevates Wingless (Wg) level (D,D′), which is rescued by knocking down Nmo (E,E′), while nmo knockdown alone shows little effect (C,C′). (F–J″) Compared with wild type (F–F″) or P35 expression alone control (G–G″), co-expression of wnd and P35 enhances Yki nuclear localization, which is completely rescued by nmo knockdown (J–J″), while Nmo knockdown alone has little effect (H–H″). (K–O) Compared with wild-type controls (K,K′) or P35 expression alone controls (L,L′), co-expression of Wnd and P35 induce non-autonomous apoptosis (shown by c-Dcp-1 staining) throughout the wing disc (N,N′). Although nmo knockdown alone does not affect apoptosis rate (M,M′), upon Nmo knockdown under Wnd and P35 co-expression background, fewer apoptosis signals are found outside GFP positive region while severe apoptosis is detected in an autonomous manner (O,O′). (P–T′) PH3 staining images are shown to demonstrate cell proliferation. Compared with wild type (P,P′) or P35 expression alone control (Q,Q′), co-expression of wnd and P35 elevates cell proliferation rate (S,S′), which is suppressed by knocking down nmo (T,T′), while nmo knockdown alone does not show significant difference (R,R′). Scale bars, 50 μm. (U) Statistic analysis of the relative PH3 + number in panel P–T. ∗P < 0.05 (unpaired t-test with Welch’s correction was used to calculate statistical significance mean + SD, n ≥ 4).
Nmo Is Essential for Impaired Cell Polarity or Rho1-Induced Hippo Inactivation
The data described above show that Nmo is required for Wnd-induced Yki activation, and Wnd is required for Rho1 activation and cell polarity-loss-induced Hippo inactivation. Next, we asked whether Nmo is also required for scrib loss and Rho1 activation-induced tissue growth and Hippo inactivation. Expression of Rho1 in the posterior region of wing discs under the control of hh-Gal4 autonomously up-regulates ex transcription and induces tissue overgrowth (Figures 4A–B″), which are significantly reduced by co-expression of nmo and RNAi (Figure 4C″ and Supplementary Figure 3). Consistently, we found that reducing Nmo activity significantly impeded scrib. RNAi induced ex-lacZ upregulation (Figures 4D′,E′). Together, these results indicate that Nmo is an essential regulator of impaired cell polarity and Rho1-induced Hippo inactivation in Drosophila wing disc.
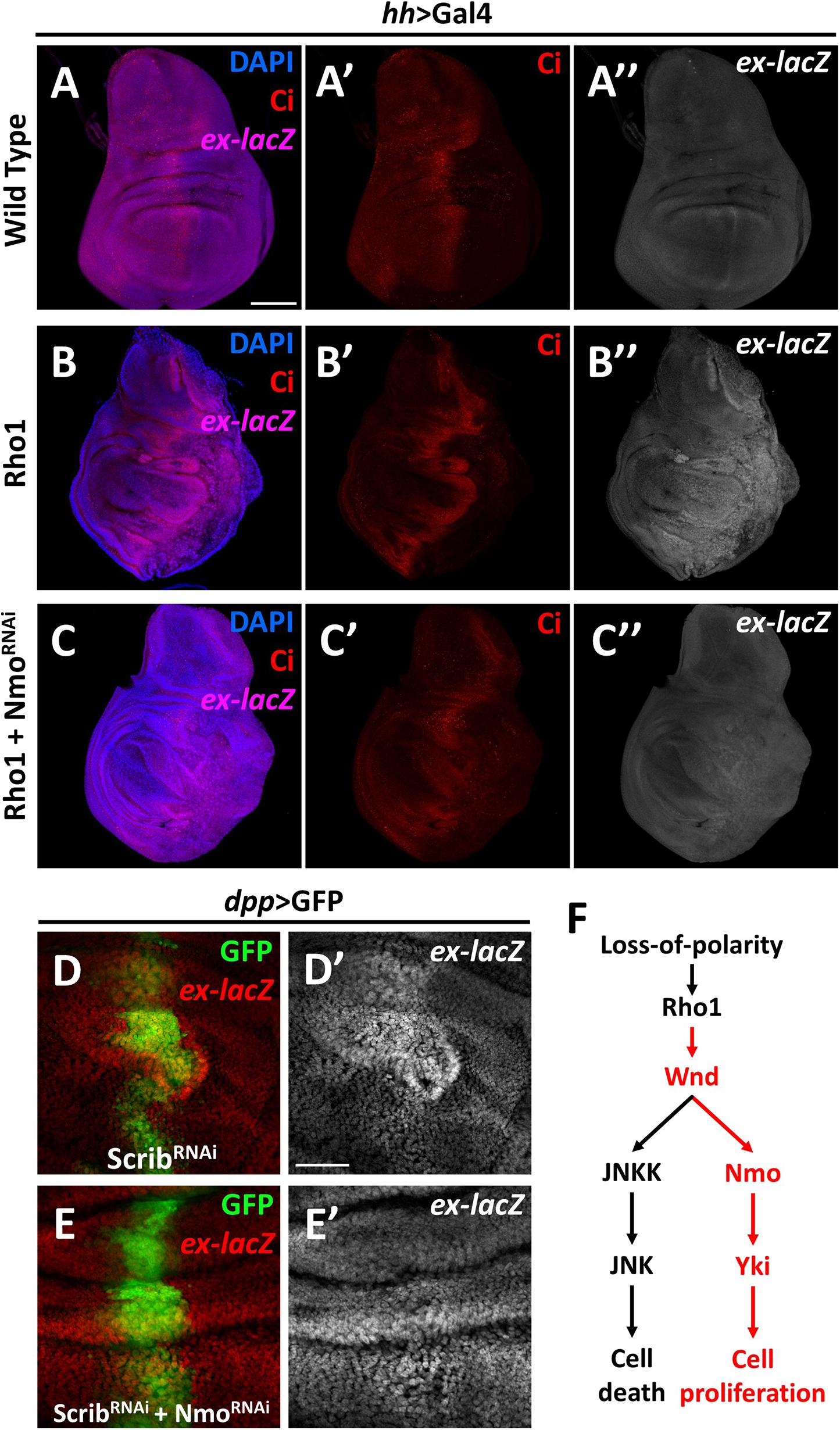
Figure 4. Nmo is essential for loss-of-cell polarity induced growth and Yki activation Wing imaginal discs from third instar larva are shown. (A–C″) Gene or RNAi is expressed driven by hh-Gal4 in the posterior region (opposite to ci-stained zone). Compared with wild type control (A–A″), expression of Rho1 elevates ex-lacZ expression level (B–B″), which is suppressed by nmo knockdown (C–C″). (D–E′) RNAi lines are driven by dpp-Gal4 in the GFP positive region. scrib knockdown caused ex transcription upregulation (D,D′) is rescued by nmo knockdown (E,E′). (F) A schematic model showing the regulation of Hippo signaling by the Wnd-Nmo axis. Scale bars, 50 μm.
In summary, our data here revealed a novel role of Wnd in promoting cell proliferation through inhibiting the Hippo pathway activity. We further demonstrated that Nmo, a MAPK family kinase, is required for the regulation of the Hippo pathway downstream of Wnd (Figure 4F). As a MAPKKK, Wnd is orthologous to both MAP3K13 and MAP3K12 in humans. Studies of these two proteins mainly focus on neuronal development, which is also the main topic of Wnd-related studies, while several studies on MAP3K13 also revealed its relation to tumor development (Han et al., 2016; Edwards et al., 2017; Chen et al., 2018; Wlaschin et al., 2018; Zhang et al., 2020). In this study, we genetically linked Wnd to the Hippo pathway for the first time and provided in vivo evidence that Wnd promotes cell proliferation in some contexts. Nmo (NLK) is involved in diverse developmental and cellular processes and was recently identified as a positive regulator of Yki/YAP activity (Moon et al., 2017; Daams and Massoumi, 2020). Despite its sequence characteristics as a MAPK, no MAPKK has been reported to phosphorylate NLK, and in contrast it can auto-phosphorylate itself. It is possible that at least in Drosophila, a Wnd-Nmo axis may exist. Interestingly, in accordance with our findings, a recent study identified both Wnd and Nmo in the same genetic screen, aiming to reveal the underlying mechanisms of Alk (Anaplastic lymphoma kinase) oncogenic signaling (Wolfstetter et al., 2020). It is also noteworthy that we found wnd and P35 overexpression-induced growth phenotype can be significantly suppressed by co-expression of wts (Supplementary Figure 5), indicating that an additional regulating mechanism may exist downstream of Wnd. Therefore, we cannot exclude the possibility that Wnd-Nmo could act on Yki via the inactivation of Wts or other kinase(s), and further investigation is required to dissect the underlying mechanism of Wnd-mediated Hippo pathway regulation. Taking advantage of the Drosophila model, our findings here suggest the exciting prospect that similar mechanisms may exist in human cancer progression, and further investigation is required to elucidate the potential link between MAP3K13-NLK axis and polarity loss induced YAP/TAZ activation in tumorigenesis.
Materials and Methods
Drosophila Strains
Drosophila stocks were reared on standard media at 25°C unless otherwise indicated. For Figures 1F–I, tub-Gal80ts was used, flies were first raised at 18°C to restrict Gal4 activity for 5–6 days, then shifted to 29°C for 2–3 days to inactivate Gal80ts. The following strains were used for this study: ptc-Gal4, UAS-GFP, UAS-p35, ban-lacZ, ex-lacZ, UAS-Rho1 (#7334), and UAS-wnd.RNAi (#27525) were obtained from the Bloomington Stock Center. UAS-scrib.RNAi (v27424) and UAS-nmo.RNAi (v3002) were collected from the Vienna Drosophila Resource Center. UAS-Wts was a gift from Shian Wu (Nankai University, Tianjin, China), UAS-Wnd (Collins et al., 2006) was a gift from Aaron DiAntonio (Washington University, St. Louis, MO).
Clonal Analysis
Flp-out ectopic expression clones in Figure 1J were generated by crossing UAS-Wnd; UAS-p35 with y w hs-FLP; act > y+ > Gal4,UAS-GFP. Clones were induced at the second instar: heat shock for 10 min at 37°C 48–72 h after egg laying (AEL), dissection were performed 36 h after clone induction.
Immunohistochemistry
Third instar larvae were dissected to collect their wing imaginal discs. Tissues were fixed with 4% formaldehyde and then washed with PBST (PBS + Triton, 1,000:3) for 5 min for 3 times at room temperature (RT). Block in PBST with Goat Serum (Solarbio SL038, 1:10) at RT for 30 min, then treat with 1st antibody at 4°C overnight. The following antibodies were used: rabbit anti-Yki (gist from Duojia Pan, 1:500); mouse anti-β-Gal (Promega, 1:500); mouse anti-wingless (DSHB 4D4, 1:100); rabbit anti-cleaved Dcp-1 (CST 9578, 1:100); Rat anti-Ci (DSHB 2A1 1:50); rabbit anti-Phospho-Histone H3 (Ser10) (CST 9701, 1:200). Next, wash the tissue with PBST for 10 min for 3 times at RT, then treat with the second antibody at RT for 2 h. Wash with PBST for 5 min for 3 times. For experiments involving nuclear localization of Yki and for Figure 4, additional DAPI staining was conducted by treating the tissue with DAPI (Beyotime C1,002) in PBST (1:1,000) at RT for 10 min. Finally, move the tissue onto a slide glass and mount with mounting media with DAPI (VECTASHIELD H-1,800) and then covered by cover glass.
Imaging and Analysis
Wing discs were imaged with the Zeiss Axio Observer microscope. Image J (Fiji) was used to count PH3 the number of positive cells. All statistical analyses were performed using GraphPad Prism 8. The experiments are repeated for at least 4 times except for Figures 3F–G″, which is repeated for 3 times. Selected pictures are representative ones. Adobe Photoshop (22.1) was used to process and adjust images. Data were statistically analyzed by the Student t-test, the unpaired t-test with Welch’s correction, or the Kruskal-Wallis test, showed in bar graph as mean + SD.
Database Analyses
CNV analysis in Figure 2A was processed on the Genomic Data Commons Data Portal1, a robust data-driven platform for access to cancer data for analysis, based on The Cancer Genome Atlas (TCGA) database. Only gain-of-number results are shown. Prognostic analysis in Figure 1B was conducted through The Human Protein Atlas2, a program aiming to map all the human proteins. This platform automatically found three cancer types that show significant difference in prognosis under low or high MAP3K13 expression level. The selected endometrial cancer result is a representative one. Expression analysis in Figure 1C and correlation analysis in Figure 1D was carried out by using the GEPIA3, an interacting web server to analyze RNA sequencing data provided by TCGA and Genotype-Tissue Expression (GTEx). We choose LUSC and OV for expression analysis and LUSC for correlation analysis because these two cancer types are the top two hits in the CNV analysis.
Data Availability Statement
Publicly available datasets were analyzed in this study. This data can be found here: The Cancer Genome Atlas (TCGA) (https://tcga-data.nci.nih.gov/tcga/; dbGaP accession number: phs000178.v1.p1).
Author Contributions
XM conceived the study. XW, HL, and WX performed the experiments. XW, HL, and XM analyzed the data and wrote the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by start-up funds from Westlake University to XM, Westlake Laboratory of Life Sciences and Biomedicine (Grant 10128A092001), and Team for Growth Control and Size Innovative Research (Grant 201804016).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Drs. Aaron DiAntonio, Duojia Pan, the Bloomington Stock Center, the Vienna Drosophila RNAi Center, the Core Facility of Drosophila Resource and Technology at the Shanghai Institute of Biochemistry, and Cell Biology of the Chinese Academy of Sciences for providing fly stocks and reagents.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fcell.2021.658288/full#supplementary-material
Footnotes
- ^ https://portal.gdc.cancer.gov
- ^ https://www.proteinatlas.org
- ^ http://gepia.cancer-pku.cn/index.html
References
Chen, L. (2019). Non-canonical hippo signaling regulates immune responses. Adv. Immunol. 144, 87–119. doi: 10.1016/bs.ai.2019.07.001
Chen, M., Geoffroy, C. G., Meves, J. M., Narang, A., Li, Y., Nguyen, M. T., et al. (2018). Leucine zipper-bearing kinase is a critical regulator of astrocyte reactivity in the adult mammalian CNS. Cell. Rep. 22, 3587–3597. doi: 10.1016/j.celrep.2018.02.102
Cho, E., Feng, Y., Rauskolb, C., Maitra, S., Fehon, R., and Irvine, K. D. (2006). Delineation of a fat tumor suppressor pathway. Nat. Genet. 38, 1142–1150. doi: 10.1038/ng1887
Choi, W., Kim, J., Park, J., Lee, D. H., Hwang, D., Kim, J. H., et al. (2018). YAP/TAZ initiates gastric tumorigenesis via upregulation of MYC. Cancer Res. 78, 3306–3320. doi: 10.1158/0008-5472.can-17-3487
Collins, C. A., Wairkar, Y. P., Johnson, S. L., and DiAntonio, A. (2006). Highwire restrains synaptic growth by attenuating a MAP kinase signal. Neuron 51, 57–69. doi: 10.1016/j.neuron.2006.05.026
Collu, G. M., Jenny, A., Gaengel, K., Mirkovic, I., Chin, M. L., Weber, U., et al. (2018). Prickle is phosphorylated by nemo and targeted for degradation to maintain prickle/spiny-legs isoform balance during planar cell polarity establishment. Plos Genet. 14:e1007391. doi: 10.1371/journal.pgen.1007391
Daams, R., and Massoumi, R. (2020). Nemo-like kinase in development and diseases: insights from mouse studies. Int. J. Mol. Sci. 21:9203. doi: 10.3390/ijms21239203
Edwards, Z. C., Trotter, E. W., Torres-Ayuso, P., Chapman, P., Wood, H. M., Nyswaner, K., et al. (2017). Survival of head and neck cancer cells relies upon LZK kinase-mediated stabilization of mutant p53. Cancer Res. 77, 4961–4972. doi: 10.1158/0008-5472.can-17-0267
Fallahi, E., O’Driscoll, N. A., and Matallanas, D. (2016). The MST/hippo pathway and cell death: a non-canonical affair. Genes 7:28. doi: 10.3390/genes7060028
Gokhale, R., and Pfleger, C. M. (2019). The power of drosophila genetics: the discovery of the hippo pathway. Methods Mol. Biol. 1893, 3–26. doi: 10.1007/978-1-4939-8910-2_1
Gou, J., Stotsky, J. A., and Othmer, H. G. (2020). Growth control in the drosophila wing disk. Wiley Interdiscip. Rev. Syst. Biol. Med. 12:e1478. doi: 10.1002/wsbm.1478
Han, H., Chen, Y., Cheng, L., Prochownik, E. V., and Li, Y. (2016). microRNA-206 impairs c-Myc-driven cancer in a synthetic lethal manner by directly inhibiting MAP3K13. Oncotarget 7, 16409–16419. doi: 10.18632/oncotarget.7653
Harvey, K. F., Pfleger, C. M., and Hariharan, I. K. (2003). The drosophila mst ortholog, hippo, restricts growth and cell proliferation and promotes apoptosis. Cell 114, 457–467. doi: 10.1016/s0092-8674(03)00557-9
Huang, J., Wu, S., Barrera, J., Matthews, K., and Pan, D. (2005). The Hippo signaling pathway coordinately regulates cell proliferation and apoptosis by inactivating Yorkie, the drosophila homolog of YAP. Cell 122, 421–434. doi: 10.1016/j.cell.2005.06.007
Justice, R. W., Zilian, O., Woods, D. F., Noll, M., and Bryant, P. J. (1995). The drosophila tumor suppressor gene warts encodes a homolog of human myotonic dystrophy kinase and is required for the control of cell shape and proliferation. Genes Dev. 9, 534–546. doi: 10.1101/gad.9.5.534
La Marca, J. E., and Richardson, H. E. (2020). Two-faced: roles of JNK signalling during tumourigenesis in the drosophila model. Front. Cell. Dev. Biol. 8:42. doi: 10.3389/fcell.2020.00042
Liu, S., Sun, J., Wang, D., Pflugfelder, G. O., and Shen, J. (2016). Fold formation at the compartment boundary of drosophila wing requires Yki signaling to suppress JNK dependent apoptosis. Sci. Rep. 6:38003.
Ma, S., Meng, Z., Chen, R., and Guan, K. L. (2019). The hippo pathway: biology and pathophysiology. Annu. Rev. Biochem. 88, 577–604. doi: 10.1146/annurev-biochem-013118-111829
Ma, X., Chen, Y., Xu, W., Wu, N., Li, M., Cao, Y., et al. (2015a). Impaired hippo signaling promotes Rho1-JNK-dependent growth. Proc. Natl. Acad. Sci. U S A. 112, 1065–1070. doi: 10.1073/pnas.1415020112
Ma, X., Xu, W., Zhang, D., Yang, Y., Li, W., and Xue, L. (2015b). Wallenda regulates JNK-mediated cell death in drosophila. Cell. Death Dis. 6:e1737. doi: 10.1038/cddis.2015.111
Ma, X., Chen, Y., Zhang, S., Xu, W., Shao, Y., Yang, Y., et al. (2016). Rho1-Wnd signaling regulates loss-of-cell polarity-induced cell invasion in drosophila. Oncogene 35, 846–855. doi: 10.1038/onc.2015.137
Ma, X. J., Guo, X. W., Richardson, H. E., Xu, T., and Xue, L. (2018). POSH regulates hippo signaling through ubiquitin-mediated expanded degradation. P. Natl. Acad. Sci. USA. 115, 2150–2155. doi: 10.1073/pnas.1715165115
Merino, C., Penney, J., Gonzalez, M., Tsurudome, K., Moujahidine, M., O’Connor, M. B., et al. (2009). Nemo kinase interacts with mad to coordinate synaptic growth at the drosophila neuromuscular junction. J. Cell. Biol. 185, 713–725. doi: 10.1083/jcb.200809127
Mirkovic, I., Charish, K., Gorski, S. M., McKnight, K., and Verheyen, E. M. (2002). Drosophila nemo is an essential gene involved in the regulation of programmed cell death. Mech. Dev. 119, 9–20. doi: 10.1016/s0925-4773(02)00289-7
Moon, S., Kim, W., Kim, S., Kim, Y., Song, Y., Bilousov, O., et al. (2017). Phosphorylation by NLK inhibits YAP-14-3-3-interactions and induces its nuclear localization. EMBO Rep. 18, 61–71. doi: 10.15252/embr.201642683
Pan, Y., Alegot, H., Rauskolb, C., and Irvine, K. D. (2018). The dynamics of hippo signaling during drosophila wing development. Development 145:dev165712. doi: 10.1242/dev.165712
Pantalacci, S., Tapon, N., and Leopold, P. (2003). The salvador partner hippo promotes apoptosis and cell-cycle exit in drosophila. Nat. Cell. Biol. 5, 921–927. doi: 10.1038/ncb1051
Perez-Garijo, A., Fuchs, Y., and Steller, H. (2013). Apoptotic cells can induce non-autonomous apoptosis through the TNF pathway. Elife 2:e01004.
Richardson, H. E., and Portela, M. (2017). Tissue growth and tumorigenesis in drosophila: cell polarity and the Hippo pathway. Curr. Opin. Cell. Biol. 48, 1–9. doi: 10.1016/j.ceb.2017.03.006
Snigdha, K., Gangwani, K. S., Lapalikar, G. V., Singh, A., and Kango-Singh, M. (2019). Hippo signaling in cancer: lessons from drosophila models. Front. Cell. Dev. Biol. 7:85.
Thompson, B. J., and Cohen, S. M. (2006). The Hippo pathway regulates the bantam microRNA to control cell proliferation and apoptosis in drosophila. Cell 126, 767–774. doi: 10.1016/j.cell.2006.07.013
Udan, R. S., Kango-Singh, M., Nolo, R., Tao, C., and Halder, G. (2003). Hippo promotes proliferation arrest and apoptosis in the salvador/warts pathway. Nat. Cell. Biol. 5, 914–920. doi: 10.1038/ncb1050
Uhlirova, M., Jasper, H., and Bohmann, D. (2005). Non-cell-autonomous induction of tissue overgrowth by JNK/Ras cooperation in a drosophila tumor model. P. Natl. Acad. Sci. USA. 102, 13123–13128. doi: 10.1073/pnas.0504170102
Verghese, S., Waghmare, I., Kwon, H., Hanes, K., and Kango-Singh, M. (2012). Scribble acts in the drosophila fat-hippo pathway to regulate warts activity. PLoS One 7:e47173. doi: 10.1371/journal.pone.0047173
Wlaschin, J. J., Gluski, J. M., Nguyen, E., Silberberg, H., Thompson, J. H., Chesler, A. T., et al. (2018). Dual leucine zipper kinase is required for mechanical allodynia and microgliosis after nerve injury. Elife 7:e33910.
Wodarz, A., and Nathke, I. (2007). Cell polarity in development and cancer. Nat. Cell. Biol. 9, 1016–1024. doi: 10.1038/ncb433
Wolfstetter, G., Pfeifer, K., Backman, M., Masudi, T. A., Mendoza-Garcia, P., Chen, S., et al. (2020). Identification of the wallenda JNKKK as an Alk suppressor reveals increased competitiveness of Alk-expressing cells. Sci. Rep. 10:14954.
Wu, C., Daniels, R. W., and DiAntonio, A. (2007). DFsn collaborates with highwire to down-regulate the Wallenda/DLK kinase and restrain synaptic terminal growth. Neural Dev. 2:16. doi: 10.1186/1749-8104-2-16
Wu, S., Huang, J., Dong, J., and Pan, D. (2003). hippo encodes a Ste-20 family protein kinase that restricts cell proliferation and promotes apoptosis in conjunction with salvador and warts. Cell 114, 445–456. doi: 10.1016/s0092-8674(03)00549-x
Xu, T., Wang, W., Zhang, S., Stewart, R. A., and Yu, W. (1995). Identifying tumor suppressors in genetic mosaics: the drosophila lats gene encodes a putative protein kinase. Development 121, 1053–1063.
Yu, W., Houl, J. H., and Hardin, P. E. (2011). NEMO kinase contributes to core period determination by slowing the pace of the drosophila circadian oscillator. Curr. Biol. 21, 756–761. doi: 10.1016/j.cub.2011.02.037
Zhang, Q., Li, X., Cui, K., Liu, C., Wu, M., Prochownik, E. V., et al. (2020). The MAP3K13-TRIM25-FBXW7alpha axis affects c-Myc protein stability and tumor development. Cell. Death Differ. 27, 420–433. doi: 10.1038/s41418-019-0363-0
Keywords: Wnd, Hippo signaling, Nmo, growth, Drosophila
Citation: Wang X, Liang H, Xu W and Ma X (2021) Wallenda-Nmo Axis Regulates Growth via Hippo Signaling. Front. Cell Dev. Biol. 9:658288. doi: 10.3389/fcell.2021.658288
Received: 25 January 2021; Accepted: 04 March 2021;
Published: 16 April 2021.
Edited by:
Zengqiang Yuan, Peking Union Medical College, ChinaReviewed by:
Ana Carmena, Institute of Neurosciences of Alicante (IN), SpainJurgen Muller, University of Bradford, United Kingdom
Copyright © 2021 Wang, Liang, Xu and Ma. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xianjue Ma, bWF4aWFuanVlQHdlc3RsYWtlLmVkdS5jbg==
†These authors have contributed equally to this work and share first authorship
 Xianping Wang
Xianping Wang Hui Liang
Hui Liang Wenyan Xu2,3,4
Wenyan Xu2,3,4 Xianjue Ma
Xianjue Ma