- Department of Microbiology, Immunology and Infectious Diseases, University of Calgary, Calgary, AB, Canada
Members of the Burkholderia cepacia complex (Bcc) are important in medical, biotechnological, and agricultural disciplines. These bacteria naturally occur in soil and water environments and have adapted to survive in association with plants and animals including humans. All Bcc species are opportunistic pathogens including Burkholderia cenocepacia that causes infections in cystic fibrosis and chronic granulomatous disease patients. The adaptation of B. cenocepacia to the host environment was assessed in a rat chronic respiratory infection model and compared to that of high cell-density in vitro grown cultures using transcriptomics. The distribution of genes differentially expressed on chromosomes 1, 2, and 3 was relatively proportional to the size of each genomic element, whereas the proportion of plasmid-encoded genes differentially expressed was much higher relative to its size and most genes were induced in vivo. The majority of genes encoding known virulence factors, components of types II and III secretion systems and chromosome 2-encoded type IV secretion system were similarly expressed between in vitro and in vivo environments. Lower expression in vivo was detected for genes encoding N-acyl-homoserine lactone synthase CepI, orphan LuxR homolog CepR2, zinc metalloproteases ZmpA and ZmpB, LysR-type transcriptional regulator ShvR, nematocidal protein AidA, and genes associated with flagellar motility, Flp type pilus formation, and type VI secretion. Plasmid-encoded type IV secretion genes were markedly induced in vivo. Additional genes induced in vivo included genes predicted to be involved in osmotic stress adaptation or intracellular survival, metal ion, and nutrient transport, as well as those encoding outer membrane proteins. Genes identified in this study are potentially important for virulence during host–pathogen interactions and may be associated with survival and adaptation to the host environment during chronic lung infections.
Introduction
Members of the Burkholderia cepacia complex (Bcc) are commonly found in soil and aquatic environments (LiPuma, 2010; Loutet and Valvano, 2010). Seventeen Bcc species have been identified, all of which have the potential to be opportunistic pathogens, although Burkholderia cenocepacia is the most clinically significant. B. cenocepacia causes lung infections resulting in significantly decreased survival rates in cystic fibrosis and chronic granulomatous disease patients (Mahenthiralingam et al., 2005). The organism is intrinsically multidrug resistant and can persist in the lungs of CF patients for many years (Mahenthiralingam et al., 2008). In some patients, infection with B. cenocepacia can progress to what is termed “cepacia syndrome.” Cepacia syndrome is associated with a rapid deterioration in lung function associated with necrotizing pneumonia, bacteremia and sepsis that can result in death (Isles et al., 1984).
Many virulence factors have been identified in B. cenocepacia including extracellular enzymes, toxins, secretions systems, iron acquisition systems, cell–cell communication (quorum sensing, QS) systems, regulatory proteins as well as genes contributing to motility, biofilm formation, adhesion, cell invasion, intracellular survival, and bacterial protection from host factors (for review see Loutet and Valvano, 2010). Several infection models have been employed to identify and characterize the contribution of numerous genes to pathogenesis (Uehlinger et al., 2009). B. cenocepacia exhibits virulence against Caenorhabditis elegans (Kothe et al., 2003), Galleria mellonella (Seed and Dennis, 2008), Acanthamoeba (Marolda et al., 1999), Dictyostelium discoideum (Aubert et al., 2008), Danio rerio (Vergunst et al., 2010), Drosophila melanogaster (Castonguay-Vanier et al., 2010), and alfalfa seedlings (Bernier et al., 2003). Chronic respiratory infection models have been developed in mice and rats to investigate pathogenesis of Bcc species. The rat chronic respiratory infection model described by Cash et al. (1979) involves transtracheal delivery of agar-embedded bacteria directly into the lung allowing for bacterial persistence and pathology to be measured. This chronic infection model has been used to identify Bcc species and bacterial strains that persisted or caused lung pathology from less virulent strains such as mutants in ornibactin biosynthesis, uptake and utilization, zinc metalloproteases, and genes encoding other enzymes, transcriptional regulators, and lipopolysaccharide (Sokol et al., 1999, 2000; Bernier et al., 2003, 2008; Corbett et al., 2003; Baldwin et al., 2004; Bernier and Sokol, 2005; Kooi et al., 2006; Loutet et al., 2006; Flannagan et al., 2007). These studies have revealed the importance of individual genes or systems to virulence but have not assessed bacterial gene expression during infection.
Transcriptional profiling using custom B. cenocepacia microarrays and RNA sequencing technology have enabled in vitro gene expression studies to be performed at a genome level. Transcriptional profiling has been used to examine gene expression in different environmental conditions such as those mimicking CF sputum or soil, or in response to antimicrobials (Drevinek et al., 2008; Yoder-Himes et al., 2009, 2010; Peeters et al., 2010; Bazzini et al., 2011; Coenye et al., 2011; Sass et al., 2011). In addition to further characterizing genes previously known to be important in virulence, these studies have also identified many genes with potential importance in virulence. Our current understanding of B. cenocepacia physiology, pathogenesis, and survival is incomplete since the B. cenocepacia genome, which is over 8 Mb, contains genes encoding many uncharacterized proteins. Identifying such proteins and determining their functional significance will improve our abilities to target such proteins for therapeutic purposes. To date, no studies have profiled B. cenocepacia gene expression at the whole genome level directly from infected cells/tissues or during infection of a susceptible host. To further understand B. cenocepacia adaptation to the host environment, we have used microarrays to examine the B. cenocepacia gene expression signature in the rat chronic respiratory infection model and compared this to high cell-density laboratory-grown cultures.
Materials and Methods
Bacterial Strains and Growth Conditions for in vitro Samples
Burkholderia cenocepacia K56-2 is a CF isolate that belongs to the ET12 lineage (RAPD type 2) and is clonally related to the sequenced strain J2315 (Mahenthiralingam et al., 2000; Baldwin et al., 2004; Holden et al., 2009). To generate in vitro samples, K56-2 cultures were grown at 37°C, in 10 ml Miller’s Luria broth (LB; Invitrogen, Burlington, ON, Canada) with shaking in 125 ml Erlenmeyer flasks to stationary phase (16 h) as previously described (O’Grady et al., 2009). Bacterial growth was assessed by determining the optical density (OD) at 600 nm.
Animal Studies
Animal infections were performed using the rat agar bead respiratory infection model (Cash et al., 1979). Adult male Sprague-Dawley rats (150–180 g; Charles River, QC, Canada) were inoculated transtracheally with approximately 107 CFU of K56-2. At 3 days postinfection, infected lungs were aseptically removed, stored at 4°C overnight in 15 ml of RNA later (Ambion, Streetsville, ON, Canada), and subsequently maintained at −70°C to prevent RNA degradation. Animal experiments were conducted according to the guidelines of the Canadian Council of Animal Care for the care and use of experimental animals under protocol M08089 approved by the University of Calgary Animal Care Committee.
RNA Manipulations
Total RNA from in vitro samples was prepared as previously described (O’Grady et al., 2009) using a RiboPure bacterial RNA isolation kit according to manufacturer’s instructions (Ambion). For in vivo samples, total RNA from infected lungs was isolated using Tri Reagent (Invitrogen) as recommended by the manufacturer. Total RNA samples were enriched for bacterial RNA using a MicrobEnrich kit (Ambion) and purified using a MegaClear kit (Ambion). Enriched and purified bacterial RNA was depleted of 16S and 23S rRNAs using a MicrobExpress kit (Ambion) to isolate mRNA according to manufacturer’s instructions to provide enhanced sensitivity for microarray experiments. DNase treatment was performed on all RNA samples using DNA-Free (Ambion), and samples were confirmed by PCR using Taq polymerase (Invitrogen) to be free of DNA prior to cDNA synthesis.
Microarray Analysis
In vitro-derived total RNA and in vivo-derived mRNA samples were indirectly labeled with the CyScribe Post-Labelling Kit (GE Healthcare) and cDNA synthesis performed as described by Sass et al. (2011) with the following modifications. Three independent RNA samples were used for in vitro samples and two mRNA samples (each consisting of an mRNA pool isolated from two infected rats to reduce variability between animals) were used for in vivo samples. Approximately 10 μg total RNA was labeled for each in vitro sample and 8 μg mRNA was labeled for each in vivo sample. The reference pool for microarray experiments consisted of B. cenocepacia J2315 genomic DNA isolated and labeled as described (Sass et al., 2011). The B. cenocepacia J2315 custom microarray, with each probe printed four times using the Agilent Sure Print 4 × 44 microarray platform, was used (Drevinek et al., 2008; Sass et al., 2011). Approximately 700–1000 ng labeled cDNA from the in vitro and in vivo samples and 55 ng labeled control genomic DNA was used per microarray. Hybridization, washing, and scanning were performed as described using the Two Color Microarray Based Gene Expression Analysis Protocol (Agilent) and the data analyzed using GeneSpring GX version 7.3.1. All labeling, hybridization, and scanning were performed by the Mahenthiralingam Laboratory, Cardiff University, Wales. Initial data were preprocessed by employing the enhanced Agilent FE import method. Probes specific to J2315 were filtered on a 1.5-fold change in expression between conditions to identify clusters of differentially regulated genes related to specific functions or potentially organized in operons. To eliminate potential differences in RNA between samples, data were normalized to control samples and mean log2 ratios (in vivo/in vitro) calculated from replicates were used and reported as expression ratios. Mean log2 ratios were also filtered on twofold changes in expression between in vivo and in vitro conditions to identify more stringently differentially regulated genes. The in vitro- or in vivo-derived K56-2 cDNA produced a signal that was detected by at least 94% of the probes on the microarray. Operon prediction and gene annotation or predicted protein function were retrieved from the B. cenocepacia J2315 genome at http://www.burkholderia.com (Winsor et al., 2008) or http://www.microbesonline.org (Dehal et al., 2009). The entire microarray data set has been deposited in the Array Express database http://www.ebi.ac.uk/arrayexpress under accession number E-MEXP-3367.
Quantitative RT-PCR
RNA for quantitative RT-PCR (qRT-PCR) was derived independently of that used for microarray analysis. Briefly, total RNA was isolated from three independent in vitro cultures prepared as described above. In a separate animal experiment to that used to prepare the microarray samples, enriched and purified total RNA was isolated as described above from three infected rats yielding three independent in vivo samples. Oligonucleotide primers (Table 1) were designed with Primer3 (Rozen and Skaletsky, 2000) and were synthesized by the University of Calgary Core DNA Services (Calgary, AB, Canada). BCAL0421 (gyrB) encoding DNA gyrase subunit B, previously used as a housekeeping gene in the Bcc multilocus sequence typing scheme (Baldwin et al., 2005) was used as a control as described previously (Peeters et al., 2010). Expression of gyrB was not significantly altered according to microarray analysis (data not shown). RT-PCR was performed using an iScript Select cDNA synthesis kit (Bio-Rad, Mississauga, ON, Canada). Quantification and melting curve analyses were performed with SsoFast Evagreen supermix with low ROX on an iCycler (Bio-Rad) according to manufacturer’s instructions. For each of the three in vitro and in vivo cDNA samples, qRT-PCRs were performed in triplicate, normalized to the control gene, gyrB. Data were calculated as previously described (Schmittgen and Livak, 2008) and represented as fold change of the in vivo samples relative to the in vitro samples.
Results
Genes on all Genomic Elements are Induced in Response to the Host Environment
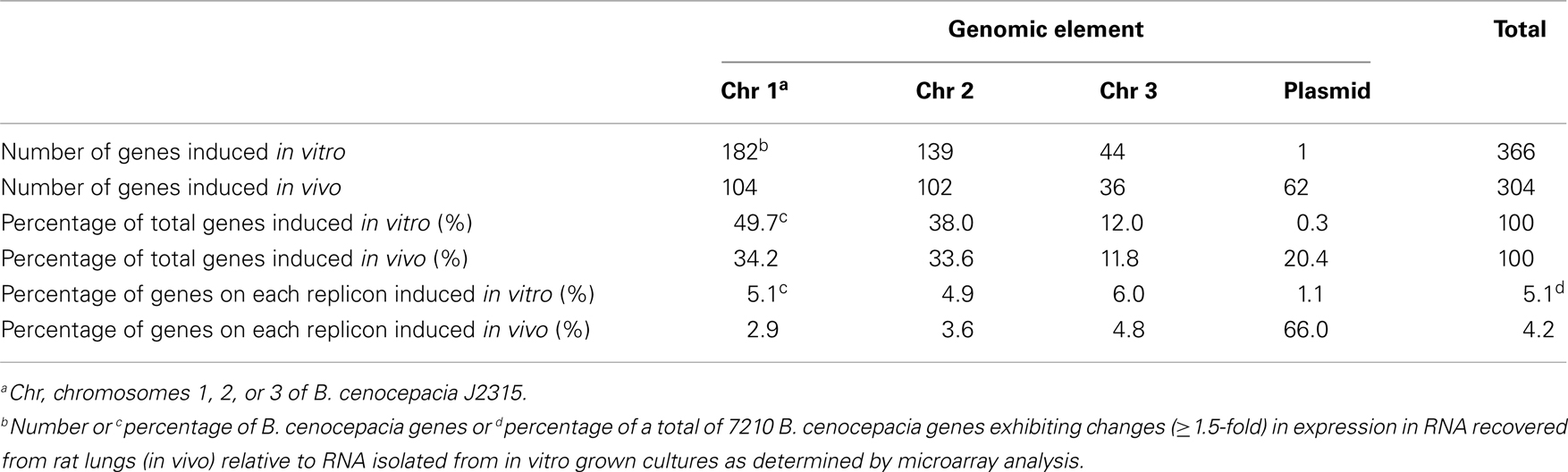
Global gene expression profiles were generated using microarrays from B. cenocepacia cultures recovered from rat lungs 3 days postinfection using a chronic respiratory infection model and compared to those of B. cenocepacia cultures grown to high cell-density in vitro. Using a fold change cut off of ≥1.5, we identified 366 genes that were induced in vitro and 304 genes that were induced in vivo (Table 2). The B. cenocepacia J2315 genome is comprised of four genetic elements: chromosome 1, 3.87 Mb; chromosome 2, 3.22 Mb; chromosome 3, 0.88 Mb; and a plasmid, 0.09 Mb (Holden et al., 2009). Differential expression was observed for genes present on the three chromosomes as well as the plasmid. The number of genes induced in vitro or induced in vivo on each genomic element and the percentage of the total number of genes induced in vitro or in vivo located on each genomic element is shown in Table 2. For in vitro induced genes, the distribution of changes across the genome was relatively proportional to the size of each genomic element, i.e., a decreasing percentage of genes showed altered expression from chromosomes 1 through 3 and to the plasmid. Interestingly, more than 20% of genes induced in vivo were plasmid genes indicating this group of genes was highly overrepresented (Table 2). Consistent with this observation, for chromosomes 1 through 3, the percentage of genes on each replicon induced in vivo was similar and ranged from 2.9 to 4.8%, in marked contrast to the plasmid where 66% of plasmid-encoded genes were induced in vivo (Table 2).
A Majority of Characterized Virulence Genes are Similarly Expressed between in vitro and in vivo Environments
At least 28 genes have been characterized in B. cenocepacia that are known to be important for virulence and belong to functional groups including stress resistance, extracellular enzymes or secreted toxins, QS, transcriptional regulation, as well as genes involved in heme uptake, iron acquisition, and the synthesis of structural components such as lipopolysaccharide, porins, and lectins (Loutet and Valvano, 2010). Analysis of these virulence genes showed that expression of the majority of these genes was similar between in vitro and in vivo conditions (Figure 1). The expression of cepI, encoding an N-acyl-homoserine lactone (AHL) synthase, was somewhat lower in vivo and this observation was consistent with lower expression of CepIR-regulated genes including those encoding extracellular zinc metalloproteases ZmpA and ZmpB, the orphan LuxR homolog CepR2 and the LysR-type transcriptional regulator ShvR (Figure 1). Two other genes known to be influenced by CepIR such as the major catalase/peroxidase encoded by katB and an acyl-CoA dehydrogenase encoded by BCAS0208 were similarly expressed in the in vitro and in vivo environments (Figure 1). The BCAS0208 mutant caused less lung pathology than wild type in the rat chronic respiratory infection model (Subramoni et al., 2011).
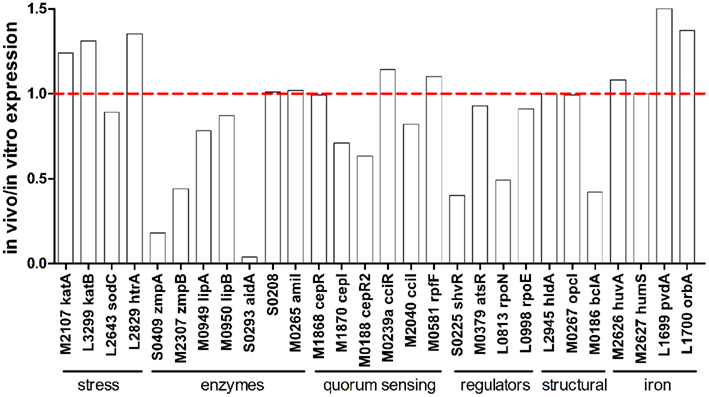
Figure 1. In vivo expression of characterized virulence genes. Expression ratio of RNA recovered from rat lungs (in vivo) relative to RNA isolated from in vitro grown cultures as determined by microarray analysis. The “BCA” designation has been removed from names of genes encoded on chromosomes 1, 2, and 3 for image clarity.
Limited iron availability in mammals is circumvented by infectious pathogens by the production of iron binding and transport complexes such as heme binding proteins and siderophores. Although genes involved in heme transport (huvA and hmuS) were not differentially expressed between in vivo and in vitro environments (Figure 1), huvA mutants exhibited survival defects in the rat chronic respiratory infection model (Hunt et al., 2004). Genes involved in ornibactin biosynthesis and transport were also expressed at similar levels in both environments, although ornibactin mediated iron uptake is required for persistence in the rat chronic respiratory infection model (Visser et al., 2004). Among the characterized virulence genes, the lowest in vivo expression ratio (0.04) was observed for BCAS0293 (aidA; Figure 1). The aidA gene encodes a protein that significantly contributes to virulence against C. elegans (Huber et al., 2004), but an aidA mutation had no effect on virulence in the rat chronic respiratory infection model (Uehlinger et al., 2009).
Secretion Systems are Selectively Regulated between in vitro and in vivo Environments
Burkholderia cenocepacia has one type II, type III, and type VI protein secretion systems (T2SS, T3SS, and T6SS, respectively) that contribute to pathogenesis, and two type IV secretion systems (T4SS), one of which has been shown to be important in virulence. Expression of genes encoding components of each of these systems varied between in vitro and in vivo environments.
The T2SS is composed at least 12 ORFs on three gsp operons and is involved in secretion of extracellular zinc metalloproteases ZmpA, ZmpB, and other extracellular proteins that have enzymatic activity such as phospholipase C, hemolysin, lipase, and polygalacturonase (Fehlner-Gardiner et al., 2002; Kothe et al., 2003; Gingues et al., 2005; Somvanshi et al., 2010). Expression of the three gsp operons encoding the T2SS was similar between in vitro and in vivo conditions (Figure 2A). Apart from the lower expression of zmpA and zmpB in vivo (Figure 1), expression of other genes encoding enzymes secreted by the T2SS described above was not different between in vitro and in vivo conditions (data not shown). The B. cenocepacia T3SS genes are organized in two operons on chromosome 2 thought to be responsible for secretion of effector proteins that have yet to be identified (Tomich et al., 2003; Glendinning et al., 2004). Mutation of bcscN, encoding an ATP-binding protein, reduced bacterial survival, and lung inflammation in a mouse agar bead infection model (Tomich et al., 2003). In our study, the mean expression ratio of genes in the bcscQ and bcscV operons was 1.03 and 0.99, respectively, in the in vivo compared to in vitro conditions (Figure 2B) indicating that there was no difference in expression.
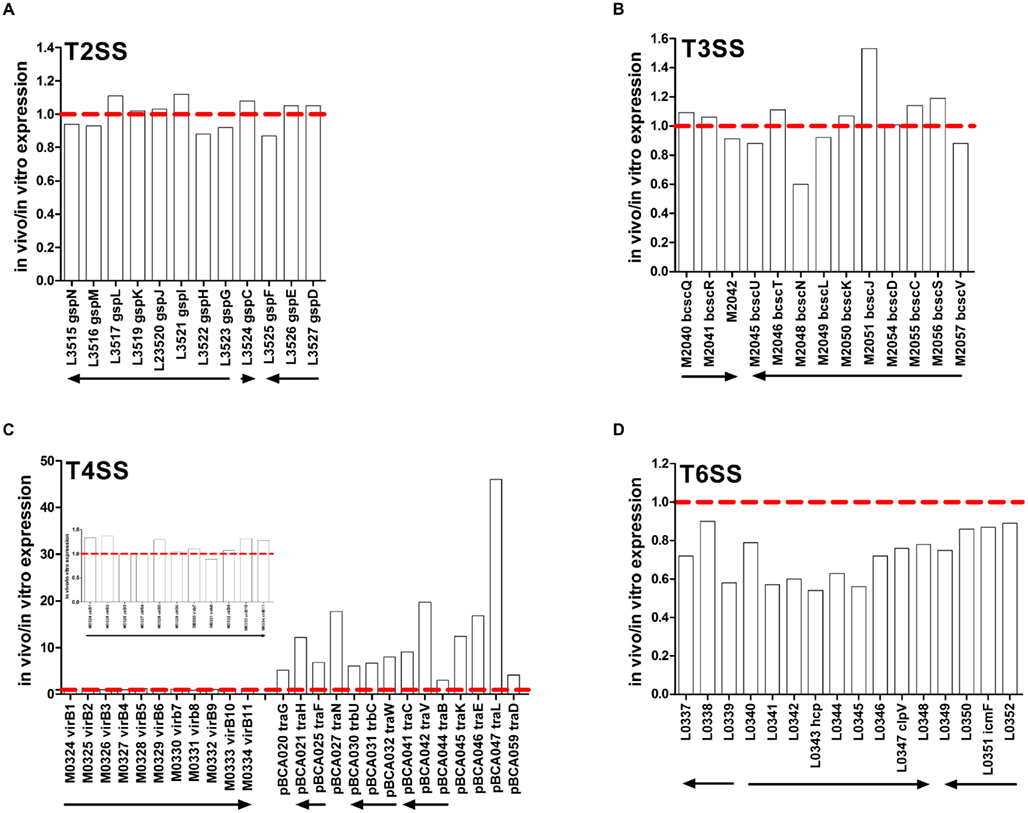
Figure 2. In vivo expression of genes encoding secretion systems. Expression ratio of RNA recovered from rat lungs (in vivo) relative to RNA isolated from in vitro grown cultures as determined by microarray analysis. (A) T2SS, (B) T3SS, (C) T4SS, (D) T6SS. Inset in (C) is chromosome 2-encoded T4SS genes with expanded y-axis. The “BCA” designation has been removed from names of genes encoded on chromosomes 1, 2, and 3 for image clarity. Putative operons are indicated by arrows.
Two gene clusters located on chromosome 2 and the plasmid have been identified to encode components of T4SS. Interestingly, the plasmid-encoded T4SS was induced in vivo. The bc-VirB/D4 T4SS on chromosome 2 shares homology with the Agrobacterium tumefaciens T4SS and is involved in plasmid mobilization (Engledow et al., 2004). The second T4SS gene cluster exists on a 92.7-kb plasmid that is found in relatively few B. cenocepacia strains including J2315 and K56-2 (Engledow et al., 2004) but not AU1054 or MCO-3 (Winsor et al., 2008). This plasmid-encoded T4SS contributes to the plant tissue watersoaking (ptw) phenotype and disease symptoms in onion tissue (Engledow et al., 2004) and increased survival of B. cenocepacia in macrophages and airway epithelial cells (Sajjan et al., 2008). Expression of genes on the chromosome 2-encoded T4SS were similar in the in vitro and in vivo conditions (Figure 2C). In contrast, several genes that are part of the plasmid-encoded T4SS were markedly induced in vivo at levels ranging from 3- to 46.1-fold (Figure 2C). Higher in vivo expression of pBCA025 encoding the putative conjugative transfer protein TraF and pBCA045 encoding the putative exported protein TraK was confirmed using qRT-PCR (Table 3). These data indicated differential regulation of chromosome 2- and plasmid-encoded T4SS between in vitro and in vivo conditions.
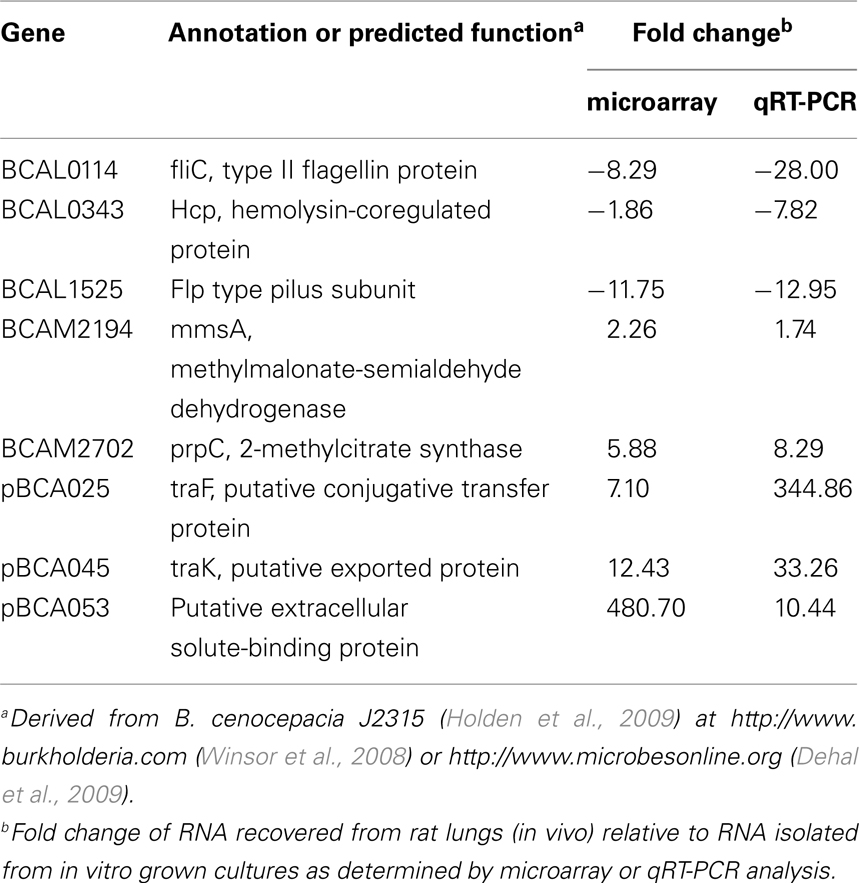
Table 3. Microarray and qRT-PCR analysis of selected genes showing differential expression from in vivo compared to in vitro grown cultures.
The B. cenocepacia T6SS comprises 16 genes organized in three adjacent operons on chromosome 1. The T6SS contributes to survival of B. cenocepacia in the rat chronic respiratory infection model (Hunt et al., 2004) and influences infection of macrophages (Aubert et al., 2008). Expression of BCAL0339 and BCAL0346 was lower in B. cenocepacia growing in medium supplemented with CF sputum compared to control cultures (Drevinek et al., 2008). In our study, expression of six T6SS genes was lower in vivo compared to in vitro conditions. The BCAL0340–0348 operon exhibited the lowest expression in vivo (0.66) compared to the other two T6SS operons (Figure 2D). The BCAL0340 operon includes genes encoding the ClpV-like chaperone (BCAL0347) and the hemolysin-coregulated protein (Hcp) (BCAL0343; Aubert et al., 2008). The ClpV-like chaperone is required for secretion of Hcp in Pseudomonas aeruginosa (Mougous et al., 2006). The hcp gene showed the lowest in vivo expression (0.54) of any T6SS gene and the low hcp expression in vivo was confirmed using qRT-PCR (Figure 2D; Table 3).
Motility and Flp Type Pilus-Encoding Genes are Induced in vitro
Bacterial motility, attachment, and invasion via flagellar- and pilus-encoding genes are known to be important in virulence (Tomich et al., 2002; Urban et al., 2004). Expression of 24 flagellar-associated genes from eight different operons distributed across chromosome 1 was lower in vivo, with the lowest in vivo/in vitro expression ratio (0.23) observed for fliC, encoding type II flagellin (Figure 3A). Lower expresssion of fliC in vivo compared to in vitro conditions was independently confirmed using qRT-PCR (Table 3).
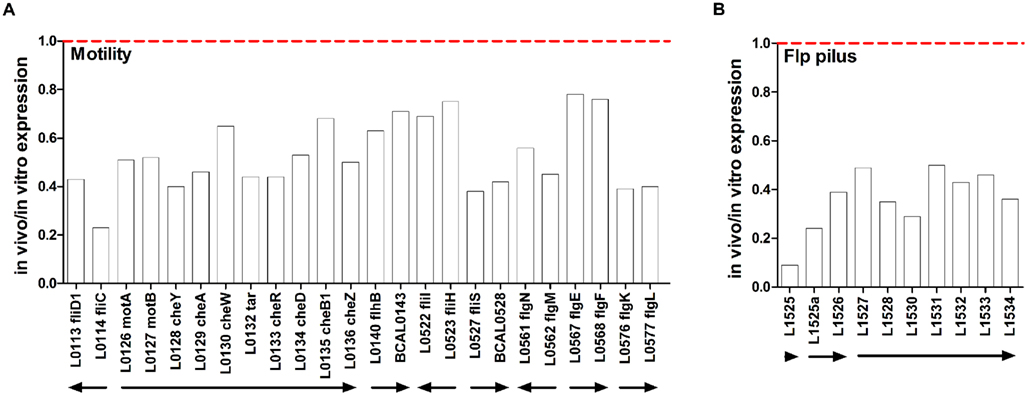
Figure 3. In vivo expression of motility and pilus genes. Expression ratio of RNA recovered from rat lungs (in vivo) relative to RNA isolated from in vitro grown cultures as determined by microarray analysis. (A) Flagellar-associated genes, (B) Flp type pilus genes. The “BCA” designation has been removed from names of genes encoded on chromosomes 1, 2, and 3 for image clarity. Putative operons are indicated by arrows.
The genomic locus from BCAL1520–1537 encodes components of a subclass of type IVb prepilins, called a Flp type pilus, that is similar to the flp–tad–rcp locus that is involved in adherence and biofilm formation in Actinobacillus actinomycetemcomitans (Kachlany et al., 2001; Inoue et al., 2003) and aggregation and biofilm formation in P. aeruginosa (de Bentzmann et al., 2006). Ten genes encoding components of the chromosome 1-encoded Flp type pilus had lower in vivo expression. The lowest expression was observed for BCAL1525 encoding a Flp type pilus subunit and this trend was confirmed using qRT-PCR (Figure 3B; Table 3).
Identification of Genes Potentially Important in the Host Environment
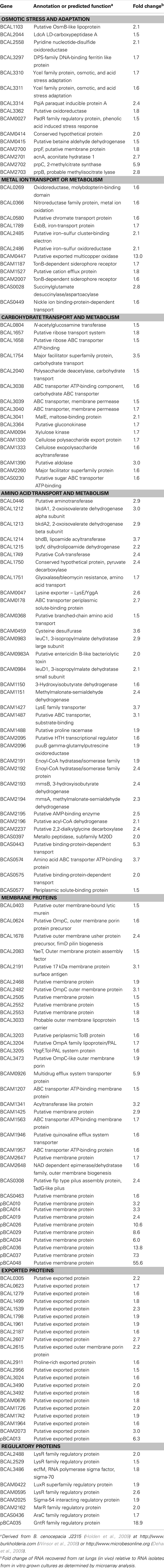
Approximately 300 genes were identified with at least a 1.5-fold change increase in expression in vivo compared to in vitro grown cultures (Table A1 in Appendix). Selected genes and their fold change differences are shown in Table 4. Many of these genes have not been previously characterized in B. cenocepacia. The most common putative functions of these in vivo induced genes were related to adaptation to stress or a host environment, metabolism, or nutrient acquisition (Table 4).
Novel Genes Induced in vivo
A four gene operon (BCAM2703–2700) containing genes involved in the methylcitrate cycle, required for propionyl-CoA metabolism, and fatty-acid utilization, were markedly induced in vivo (Table 4). Induced in vivo expression of BCAM2702 (prpC) encoding 2-methylcitrate synthase was confirmed using qRT-PCR (Table 3). Genes involved in the methylcitrate and glyoxylate cycles are required for virulence in Mycobacterium tuberculosis, which relies more on fatty acids than carbohydrates during infection (Munoz-Elias and McKinney, 2005). Genes involved in the methylcitrate cycle are upregulated in M. tuberculosis isolated from murine macrophages (Schnappinger et al., 2003) and are important for growth in macrophages but not for intracellular survival (Munoz-Elias et al., 2006). It is unknown whether the methylcitrate cycle plays a role in B. cenocepacia intracellular survival in macrophages. An uncharacterized seven gene operon (BCAM2196–BCAM2191) containing genes putatively involved in lipid metabolism was also induced in vivo (Table 4), suggesting that fatty-acid metabolism or utilization may be important in B. cenocepacia lung infections. Using qRT-PCR we confirmed expression of BCAM2194 (mmsA) encoding methylmalonate-semialdehyde dehydrogenase was induced in vivo (Table 3). A four gene operon (BCAL1212–1215) induced in vivo encodes genes for a 2-oxo acid dehydrogenase complex (Table 4). The dihydrolipoamide dehydrogenase gene component of a similar complex was shown to be important for persistence and virulence in Streptococcus pneumoniae infection models likely due to having a role in capsule synthesis rather than metabolism of 2-oxo acids (Smith et al., 2002).
BCAM0415 encodes a betaine aldehyde dehydrogenase (BADH; Table 4). In P. aeruginosa, BADH has been shown to be induced by choline and choline precursors (Velasco-Garcia et al., 2006a) which are abundant in infected lung tissues (Wright and Clements, 1987). In addition to playing a role in assimilating carbon and nitrogen from choline, BADH produces glycine betaine which can protect bacteria from high osmolarity stress and oxidative stress in infected tissues. BADH has been proposed as a therapeutic target for P. aeruginosa since inactivation of this enzyme leads to intracellular accumulation of betaine aldehyde, which is toxic, and the inability to grow in medium with choline (Velasco-Garcia et al., 2006b; Zaldivar-Machorro et al., 2011). Homologs of other genes induced by osmotic stress in bacteria were also identified as being induced in vivo (Table 4). BCAL1103, encodes an OsmB-like protein. OsmB is induced by osmotic stress and stationary phase growth conditions in E. coli (Jung et al., 1990; Boulanger et al., 2005). BCAL3310 and BCAL3311 are predicted to be co-transcribed YceI family proteins, homologs of which have been shown to be induced in response to osmotic stress in E. coli (Weber et al., 2006) and acid stress in Helicobacter pylori (Sisinni et al., 2010). BCAL2558, a putative pyridine nucleotide-disulfide oxidoreductase with some similarity to TrxB (thioredoxin reductase) homologs, was induced twofold in vivo. TrxB genes are involved in cellular redox processes and defense against oxidative stress and are important in intracellular survival in some pathogens (Bjur et al., 2006; Potter et al., 2009). BCAL3314 encodes a homolog of PqiA-like proteins, which are induced by paraquat and other superoxide generators in E. coli (Koh and Roe, 1995). BCAL3297 encodes a DPS-family DNA-binding ferritin. Homologs of these proteins are involved in resistance as well as iron sequestration (Calhoun and Kwon, 2011).
Although many of the in vivo induced outer membrane protein encoding genes are uncharacterized, a few have homology to proteins with predicted functions. BCAL3203, L3204, and L3205 form part of the Tol-PAL system membrane complex that is required for membrane integrity and has been implicated in the pathogenesis of several Gram-negative bacteria (Bowe et al., 1998; Godlewska et al., 2009; Paterson et al., 2009). TolB (BCAL3203) is a periplasmic protein involved in biopolymer transport. BCAL3205 is a YbgF homolog which is the last gene of the Tol-PAL complex and interacts with TolA (Krachler et al., 2010). BCAL3204 has been annotated as OmpA/PAL. PAL has been shown to contribute to virulence in several Gram-negative bacteria and in E. coli has been shown to be released into the bloodstream contributing to septic shock (Hellman et al., 2002; Liang et al., 2005). A 17 kDa OmpA-like protein has recently been shown to be an immunodominant antigen following intranasal immunization with a B. cenocepacia outer membrane protein preparation in mice (Makidon et al., 2010). Although the immunoreactive protein reported to be an OmpA-like protein was not conclusively identified, the partial amino acid sequence determined from a peptide of this molecular mass isolated from SDS-polyacrylamide gels, has 95.8% identity to BCAL3204. There are at least six other OmpA-like proteins in B. cenocepacia with varying degrees of sequence identify; however, PAL has been shown to highly immunogenic in other bacteria (Godlewska et al., 2009). Therefore it is possible that the immunodominant antigen identified by Makidon et al. (2010) is PAL. BCAL2191, which was increased threefold in vivo (Table 4) is predicted to be an outer membrane lipoprotein with similarity to 17 kDa surface antigens in other species and therefore it is also possible that this protein contributed to the observed reaction with antiserum on Western blots in the study by Makidon et al. (2010). Several other proteins involved in biogenesis of membrane and other cell surface components were also identified (Table 4) including BCAL2083, a YaeT homolog, which in E. coli is an essential gene required for outer membrane assembly (Werner and Misra, 2005). BCAL2482 is a putative outer membrane porin (OmpC) and is in the same predicted operon as BCAL2486 and BCAL2485, which are iron–sulfur oxidoreductase and iron–sulfur electron transport proteins, respectively. All three genes are induced at least twofold in vivo (Table 4).
Although ornibactin biosynthesis and uptake genes were expressed at similar levels in the in vitro and in vivo conditions used in this study, a number of other genes potentially involved in metal ion transport and metabolism were identified as being induced in vivo (Table 4). These included exbB, genes coding for iron–sulfur proteins and receptors for unknown siderophores. One of the most highly induced genes in vivo was BCAM0447 which encodes a putative multicopper oxidase (MCO). MCO genes are found in a number of genomes but have only recently been characterized. The MCO protein of P. aeruginosa has been shown to be involved in the oxidation of ferrous to ferric iron and may be important in iron acquisition (Huston et al., 2002). MCO homologs are also involved in copper resistance and dissemination in mice in S. typhimurium (Achard et al., 2010) and copper tolerance in Campylobacter jejuni (Hall et al., 2008).
Genes encoding proteins of unknown function induced in vivo are shown in Table 4 and Table A1 in Appendix. Many of the expressed genes encode outer membrane proteins (11) and exported proteins (24) that could contribute to cell surface alterations or virulence. Genes encoding six hypothetical proteins were unique to B. cenocepacia (Table A1 in Appendix), whereas, 23 genes encoding hypothetical proteins were conserved in one or more members of the Bcc, of which 11 were also conserved in Burkholderia pseudomallei (Table A1 in Appendix). It is possible that these proteins are involved in adaptation, survival, or virulence in lung infections although further studies are required to determine their potential importance.
Plasmid-Associated Genes
Interestingly, the most highly induced genes in vivo were located on the plasmid where the vast majority of the genes were expressed at much higher levels in vivo than in vitro (Figure 4). Of the plasmid genes annotated in the J2315 sequence (Winsor et al., 2008), 62 genes had higher expression in the lung infection model. Only one gene, pBCA055, had higher expression levels in vitro, and the following genes had similar expression: pBCA003–007, 061, 063, 064, 066–075, 078–081, 083–086, 091–094.
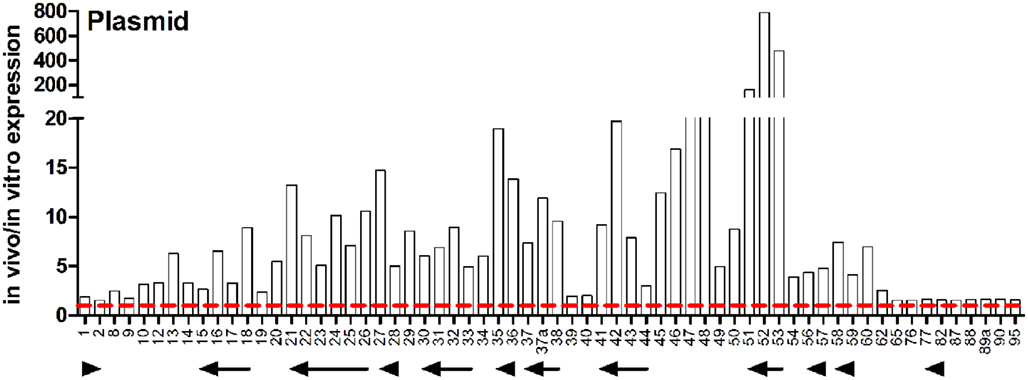
Figure 4. In vivo expression of plasmid-encoded genes. Expression ratio of RNA recovered from rat lungs (in vivo) relative to RNA isolated from in vitro grown cultures as determined by microarray analysis. The “pBCA” designation has been removed from names of plasmid-encoded genes for image clarity. Putative operons are indicated by arrows.
Many of the highly induced genes are part of the plasmid-encoded T4SS, which has been shown to play a role in both plant pathogenesis and survival in eukaryotic cells (Engledow et al., 2004; Sajjan et al., 2008). Expression ratios of genes known or predicted to be a part of the T4SS are shown in Figure 2C and described above. The presence of the plasmid-encoded T4SS in the B. cenocepacia ET12 lineage strains J2315 and K56-2 but not AU1054 or MCO-3 that entirely lack a plasmid is an interesting characteristic. Gene expression of pBCA054 encoding a LuxR family regulatory protein was higher in vivo. Interestingly, the most closely related pBCA054 orthologs are found in B. pseudomallei and Burkholderia mallei, rather than in other members of the Bcc. pBCA001–002 are parAB-like homologs that are putatively involved in chromosome partitioning. pBCA017 is similar to the zeta toxin family of toxin–antitoxin complexes which are involved in programmed cell death to prevent proliferation of plasmid free cells (Gerdes et al., 2005). In addition to plasmid maintenance, toxin–antitoxin pairs can also be involved in responding to nutrient stress. Zeta toxins have recently been shown to target peptidoglycan synthesis triggering autolysis (Mutschler et al., 2011). Zeta toxins are typically paired with epsilon antitoxins; however, there does not appear to be an epsilon homolog adjacent to pBCA017. In some cases, a chromosomal antitoxin can neutralize the plasmid toxin, but in this case toxin expression would not favor plasmid maintenance (Van Melderen and Saavedra De Bast, 2009). Alternatively the toxin can be integrated into other regulatory networks or serve to reduce the overall population to increase nutrient availability for the survivors. Three genes forming an operon (pBCA053–051) exhibited the highest induction of any group of genes in vivo (Figure 4). pBCA053 encodes a extracellular solute-binding protein involved in dicarboxylate transporter carbohydrate metabolism and we confirmed higher in vivo expression of this gene using qRT-PCR (Table 3). The second and third genes in the operon encode an exported protein and a protein with homology to LamB/YcsF family proteins, respectively. In addition to the hypothetical proteins noted above, four putative exported proteins, nine putative membrane proteins, 12 conserved hypothetical proteins and 10 hypothetical proteins encoded on the plasmid were induced in vivo (Table A1 in Appendix). Few genes on this plasmid have been studied in detail opening the possibility for identifying proteins with potentially novel functions.
Discussion
In this study, we have identified the gene expression signature of B. cenocepacia during lung infections. To the best of our knowledge, this is the first study to apply transcriptomics for any member of the Bcc to study gene expression during infection of a susceptible host. Differential gene expression was observed for characterized virulence genes as well as potential novel virulence genes between in vitro and in vivo environments.
Altered in vivo gene expression was observed for genes encoding enzymes, regulators, structural appendages as well as those contributing to ornibactin biosynthesis, and quorum sensing systems. Lower in vivo expression was observed for AHL-dependent QS controlled genes that are directly (e.g., aidA) and indirectly (e.g., shvR) regulated at the transcriptional level by CepR (Weingart et al., 2005; O’Grady et al., 2011). These observations suggest that more favorable conditions exist for CepIR-dependent regulation of selected genes in high cell-density (∼109) laboratory-grown cultures compared to the lower cell-density (∼105) in the lung infections, although it is possible that higher expression of QS regulated genes occurs in selected locations in the lungs where bacteria are present in high cell-density biofilms. Since cepI and CepR-regulated genes including zmpA, zmpB, and shvR have been shown to be important for virulence in the rat chronic respiratory infection model (Corbett et al., 2003; Sokol et al., 2003; Kooi et al., 2006; Bernier et al., 2008), it is clear that these genes are expressed at sufficient levels to play a role in infection. The majority of characterized virulence genes were similarly expressed in the in vivo and in vivo conditions. This suggests that expression of these genes is just as important in high cell-density cultures and during lung infections. The contribution of these individual genes has been characterized in one or more infection models highlighting their importance in B. cenocepacia pathogenesis. Similar expression of characterized virulence genes during growth in vivo in hamsters and in vitro has previously been observed for B. pseudomallei (Tuanyok et al., 2006).
Increased expression of some genes belonging to the T3SS was observed in the closely related pathogens B. mallei and B. pseudomallei during infection of mice and hamsters, respectively (Kim et al., 2005; Tuanyok et al., 2006). In the present study, expression of T2SS and T3SS genes was similar between in vitro and in vivo environments. Genes in these secretion systems appear to be expressed at moderate levels in both in vitro and in vivo environments. We previously showed expression of the T2SS genes gspC and gspG was influenced by growth medium composition (O’Grady et al., 2011). A previous study was not able to identify growth conditions that altered expression of T3SS genes suggesting these genes are constitutively expressed (Engledow et al., 2004). The in vivo growth conditions provided a stimulus for expression of genes in the plasmid-encoded T4SS but did not affect expression of the T4SS genes on chromosome 2. A mutation in the chromosome 2-encoded T4SS was shown not to contribute to bacterial persistence or histopathology in the rat chronic respiratory infection model (Bernier and Sokol, 2005). To date, no studies have observed such a dramatic increase in expression of plasmid-encoded T4SS genes suggesting that specific environmental signal(s) in the lung environment enabled increased expression of these genes to be detected. It was shown that the plasmid-encoded T4SS contributed to onion tissue maceration through secretion of one or more effectors (Engledow et al., 2004). Whether this plasmid-encoded T4SS or its effectors have a role in mammalian cell/tissue damage has yet to be determined. We observed some T6SS genes had lower in vivo expression, in particular those genes on the BCAL0340 operon that includes a gene encoding the secreted effector Hcp. Previous work identified a transposon insertion in each of the three operons of the T6SS locus affected survival of B. cenocepacia in the rat chronic respiratory infection model (Hunt et al., 2004).
Using a mouse agar bead infection model, a flagellin mutant failed to cause mortality compared to wild type (Urban et al., 2004). It was also shown that motility mutants were less able to invade epithelial cells (Tomich et al., 2002). Recent work showed expression of flagellar- and chemotaxis-associated genes and motility was reduced in B. cenocepacia strains of the ET12 lineage that were isolated from CF patients (Sass et al., 2011). However, a previous study showed transcription of flagellar-associated genes was increased in B. cenocepacia J2315 cultured in medium supplemented with CF sputum (Drevinek et al., 2008). Conflicting data regarding expression of flagellar-associated genes in these two studies likely reflect the experimental conditions employed where increased expression of flagellar-associated genes was detected in rapidly growing cultures (Drevinek et al., 2008). The phenotypic characteristics of the B. cenocepacia non-motile CF isolates are similar to P. aeruginosa clinical isolates which often acquire loss-of-function mutations associated with motility during chronic lung infection (Mahenthiralingam et al., 1994). It has also been shown that P. aeruginosa exhibited decreased transcription of flagellar-associated genes when cultured in CF sputum (Wolfgang et al., 2004). In our study, we detected lower in vivo expression of genes involved in motility and Flp type pilus formation. This result was likely due to differences in culture conditions between in vitro and in vivo environments. The agar bead infection model bypasses the colonization step during infection (Cash et al., 1979). Our data suggest expression of these genes is not required in an established infection taking place in the lower respiratory tract. Therefore, decreased expression of these genes was expected since expression of these genes is an energy-expensive process and is more likely associated with rapidly growing cultures than cultures recovered from chronic lung infection.
We identified numerous genes that were induced during lung infections. Many of these genes encode proteins with functions related to metabolism, physiology, or adaptation to a stressful environment. While homologs of some of these proteins have been studied in other pathogens, these proteins have not been specifically studied in B. cenocepacia. Several B. cenocepacia ET12 lineage strains contain at least a 45-kb fragment of the plasmid found in K56-2 and J2315 (Engledow et al., 2004) while strains AU1054 and MCO-3 lack a plasmid (Winsor et al., 2008). While plasmid-minus derivatives of B. cenocepacia J2315 or K56-2 have not been reported, it would be interesting to determine what influence absence of the plasmid may have on infection considering the vast majority of plasmid-encoded genes were induced in vivo. Further confirmatory experiments are required to substantiate trends for additional genes that exhibited altered expression in the in vivo environmental conditions. Revealing the changes in gene expression that occur in bacterial cells during infection is a first step in understanding the response of bacterial cells to the host environment. Increased expression of genes during infection suggests these genes promote bacterial survival and adaptation in the lungs and potentially influence virulence. The identification of potential novel virulence genes among these in vivo induced genes provides an opportunity to characterize these genes in more detail in future studies. Determining what growth conditions alter the expression of these genes and how they are regulated in B. cenocepacia will shed light on their expression pattern. Increased expression of genes during lung infection could be due to a change in environmental cues that enable transcriptional activation by a positive regulator(s) or derepression by a negative regulator(s). For potentially novel virulence genes, it will be important to construct mutations and examine their influence on virulence-related phenotypes and pathogenesis in one or more infection models. This study provides an insight into B. cenocepacia gene expression in vivo and may provide opportunities to devise strategies to reduce or control B. cenocepacia lung infections.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
These studies were supported by research grants from Cystic Fibrosis Canada, Cystic Fibrosis Foundation Therapeutics (CFFT) (grant SOKOL06V0), and Canadian Institutes of Health Research (grant MOP-42510) to PAS. EPO was the recipient of a Cystic Fibrosis Canada fellowship. We thank S. A. McKeon and D. F. Viteri for performing the animal experiments and D. T. Nguyen for excellent technical assistance. Microarray processing and initial data assessment was provided by the Mahenthiralingam Laboratory, Cardiff University, with support from CFFT.
References
Achard, M. E., Tree, J. J., Holden, J. A., Simpfendorfer, K. R., Wijburg, O. L., Strugnell, R. A., Schembri, M. A., Sweet, M. J., Jennings, M. P., and McEwan, A. G. (2010). The multi-copper-ion oxidase CueO of Salmonella enterica serovar Typhimurium is required for systemic virulence. Infect. Immun. 78, 2312–2319.
Aubert, D. F., Flannagan, R. S., and Valvano, M. A. (2008). A novel sensor kinase-response regulator hybrid controls biofilm formation and type VI secretion system activity in Burkholderia cenocepacia. Infect. Immun. 76, 1979–1991.
Baldwin, A., Mahenthiralingam, E., Thickett, K. M., Honeybourne, D., Maiden, M. C., Govan, J. R., Speert, D. P., LiPuma, J. J., Vandamme, P., and Dowson, C. G. (2005). Multilocus sequence typing scheme that provides both species and strain differentiation for the Burkholderia cepacia complex. J. Clin. Microbiol. 43, 4665–4673.
Baldwin, A., Sokol, P. A., Parkhill, J., and Mahenthiralingam, E. (2004). The Burkholderia cepacia epidemic strain marker is part of a novel genomic island encoding both virulence and metabolism-associated genes in Burkholderia cenocepacia. Infect. Immun. 72, 1537–1547.
Bazzini, S., Udine, C., Sass, A., Pasca, M. R., Longo, F., Emiliani, G., Fondi, M., Perrin, E., Decorosi, F., Viti, C., Giovannetti, L., Leoni, L., Fani, R., Riccardi, G., Mahenthiralingam, E., and Buroni, S. (2011). Deciphering the role of RND efflux transporters in Burkholderia cenocepacia. PLoS ONE 6, e18902. doi: 10.1371/journal.pone.0018902
Bernier, S. P., Nguyen, D. T., and Sokol, P. A. (2008). A LysR-type transcriptional regulator in Burkholderia cenocepacia influences colony morphology and virulence. Infect. Immun. 76, 38–47.
Bernier, S. P., Silo-Suh, L., Woods, D. E., Ohman, D. E., and Sokol, P. A. (2003). Comparative analysis of plant and animal models for characterization of Burkholderia cepacia virulence. Infect. Immun. 71, 5306–5313.
Bernier, S. P., and Sokol, P. A. (2005). Use of suppression-subtractive hybridization to identify genes in the Burkholderia cepacia complex that are unique to Burkholderia cenocepacia. J. Bacteriol. 187, 5278–5291.
Bjur, E., Eriksson-Ygberg, S., Aslund, F., and Rhen, M. (2006). Thioredoxin 1 promotes intracellular replication and virulence of Salmonella enterica serovar Typhimurium. Infect. Immun. 74, 5140–5151.
Boulanger, A., Francez-Charlot, A., Conter, A., Castanie-Cornet, M. P., Cam, K., and Gutierrez, C. (2005). Multistress regulation in Escherichia coli: expression of osmB involves two independent promoters responding either to sigmaS or to the RcsCDB His-Asp phosphorelay. J. Bacteriol. 187, 3282–3286.
Bowe, F., Lipps, C. J., Tsolis, R. M., Groisman, E., Heffron, F., and Kusters, J. G. (1998). At least four percent of the Salmonella typhimurium genome is required for fatal infection of mice. Infect. Immun. 66, 3372–3377.
Calhoun, L. N., and Kwon, Y. M. (2011). Structure, function and regulation of the DNA-binding protein Dps and its role in acid and oxidative stress resistance in Escherichia coli: a review. J. Appl. Microbiol. 110, 375–386.
Cash, H. A., Woods, D. E., McCullough, B., Johanson, W. G. Jr., and Bass, J. A. (1979). A rat model of chronic respiratory infection with Pseudomonas aeruginosa. Am. Rev. Respir. Dis. 119, 453–459.
Castonguay-Vanier, J., Vial, L., Tremblay, J., and Deziel, E. (2010). Drosophila melanogaster as a model host for the Burkholderia cepacia complex. PLoS ONE 5, e11467. doi: 10.1371/journal.pone.0011467
Coenye, T., Van Acker, H., Peeters, E., Sass, A., Buroni, S., Riccardi, G., and Mahenthiralingam, E. (2011). Molecular mechanisms of chlorhexidine tolerance in Burkholderia cenocepacia biofilms. Antimicrob. Agents Chemother. 55, 1912–1919.
Corbett, C. R., Burtnick, M. N., Kooi, C., Woods, D. E., and Sokol, P. A. (2003). An extracellular zinc metalloprotease gene of Burkholderia cepacia. Microbiology 149, 2263–2271.
de Bentzmann, S., Aurouze, M., Ball, G., and Filloux, A. (2006). FppA, a novel Pseudomonas aeruginosa prepilin peptidase involved in assembly of type IVb pili. J. Bacteriol. 188, 4851–4860.
Dehal, P. S., Joachimiak, M. P., Price, M. N., Bates, J. T., Baumohl, J. K., Chivian, D., Friedland, G. D., Huang, K. H., Keller, K., Novichkov, P. S., Dubchak, I. L., Alm, E. J., and Arkin, A. P. (2009). MicrobesOnline: an integrated portal for comparative and functional genomics. Nucleic Acids Res. 38, D396–D400.
Drevinek, P., Holden, M. T., Ge, Z., Jones, A. M., Ketchell, I., Gill, R. T., and Mahenthiralingam, E. (2008). Gene expression changes linked to antimicrobial resistance, oxidative stress, iron depletion and retained motility are observed when Burkholderia cenocepacia grows in cystic fibrosis sputum. BMC Infect. Dis. 8, 121. doi: 10.1186/1471-2334-8-121
Engledow, A. S., Medrano, E. G., Mahenthiralingam, E., LiPuma, J. J., and Gonzalez, C. F. (2004). Involvement of a plasmid-encoded type IV secretion system in the plant tissue watersoaking phenotype of Burkholderia cenocepacia. J. Bacteriol. 186, 6015–6024.
Fehlner-Gardiner, C. C., Hopkins, T. M., and Valvano, M. A. (2002). Identification of a general secretory pathway in a human isolate of Burkholderia vietnamiensis (formerly B. cepacia complex genomovar V) that is required for the secretion of hemolysin and phospholipase C activities. Microb. Pathog. 32, 249–254.
Flannagan, R. S., Aubert, D., Kooi, C., Sokol, P. A., and Valvano, M. A. (2007). Burkholderia cenocepacia requires a periplasmic HtrA protease for growth under thermal and osmotic stress and for survival in vivo. Infect. Immun. 75, 1679–1689.
Gerdes, K., Christensen, S. K., and Lobner-Olesen, A. (2005). Prokaryotic toxin-antitoxin stress response loci. Nat. Rev. Microbiol. 3, 371–382.
Gingues, S., Kooi, C., Visser, M. B., Subsin, B., and Sokol, P. A. (2005). Distribution and expression of the ZmpA metalloprotease in the Burkholderia cepacia complex. J. Bacteriol. 187, 8247–8255.
Glendinning, K. J., Parsons, Y. N., Duangsonk, K., Hales, B. A., Humphreys, D., Hart, C. A., and Winstanley, C. (2004). Sequence divergence in type III secretion gene clusters of the Burkholderia cepacia complex. FEMS Microbiol. Lett. 235, 229–235.
Godlewska, R., Wisniewska, K., Pietras, Z., and Jagusztyn-Krynicka, E. K. (2009). Peptidoglycan-associated lipoprotein (Pal) of Gram-negative bacteria: function, structure, role in pathogenesis and potential application in immunoprophylaxis. FEMS Microbiol. Lett. 298, 1–11.
Hall, S. J., Hitchcock, A., Butler, C. S., and Kelly, D. J. (2008). A multicopper oxidase (Cj1516) and a CopA homologue (Cj1161) are major components of the copper homeostasis system of Campylobacter jejuni. J. Bacteriol. 190, 8075–8085.
Hellman, J., Roberts, J. D. Jr., Tehan, M. M., Allaire, J. E., and Warren, H. S. (2002). Bacterial peptidoglycan-associated lipoprotein is released into the bloodstream in Gram-negative sepsis and causes inflammation and death in mice. J. Biol. Chem. 277, 14274–14280.
Holden, M. T., Seth-Smith, H. M., Crossman, L. C., Sebaihia, M., Bentley, S. D., Cerdeno-Tarraga, A. M., Thomson, N. R., Bason, N., Quail, M. A., Sharp, S., Cherevach, I., Churcher, C., Goodhead, I., Hauser, H., Holroyd, N., Mungall, K., Scott, P., Walker, D., White, B., Rose, H., Iversen, P., Mil-Homens, D., Rocha, E. P., Fialho, A. M., Baldwin, A., Dowson, C., Barrell, B. G., Govan, J. R., Vandamme, P., Hart, C. A., Mahenthiralingam, E., and Parkhill, J. (2009). The genome of Burkholderia cenocepacia J2315, an epidemic pathogen of cystic fibrosis patients. J. Bacteriol. 191, 261–277.
Huber, B., Feldmann, F., Kothe, M., Vandamme, P., Wopperer, J., Riedel, K., and Eberl, L. (2004). Identification of a novel virulence factor in Burkholderia cenocepacia H111 required for efficient slow killing of Caenorhabditis elegans. Infect. Immun. 72, 7220–7230.
Hunt, T. A., Kooi, C., Sokol, P. A., and Valvano, M. A. (2004). Identification of Burkholderia cenocepacia genes required for bacterial survival in vivo. Infect. Immun. 72, 4010–4022.
Huston, W. M., Jennings, M. P., and McEwan, A. G. (2002). The multicopper oxidase of Pseudomonas aeruginosa is a ferroxidase with a central role in iron acquisition. Mol. Microbiol. 45, 1741–1750.
Inoue, T., Shingaki, R., Sogawa, N., Sogawa, C. A., Asaumi, J., Kokeguchi, S., and Fukui, K. (2003). Biofilm formation by a fimbriae-deficient mutant of Actinobacillus actinomycetemcomitans. Microbiol. Immunol. 47, 877–881.
Isles, A., Maclusky, I., Corey, M., Gold, R., Prober, C., Fleming, P., and Levison, H. (1984). Pseudomonas cepacia infection in cystic fibrosis: an emerging problem. J. Pediatr. 104, 206–210.
Jung, J. U., Gutierrez, C., Martin, F., Ardourel, M., and Villarejo, M. (1990). Transcription of osmB, a gene encoding an Escherichia coli lipoprotein, is regulated by dual signals. Osmotic stress and stationary phase. J. Biol. Chem. 265, 10574–10581.
Kachlany, S. C., Planet, P. J., Desalle, R., Fine, D. H., Figurski, D. H., and Kaplan, J. B. (2001). flp-1, the first representative of a new pilin gene subfamily, is required for non-specific adherence of Actinobacillus actinomycetemcomitans. Mol. Microbiol. 40, 542–554.
Kim, H. S., Schell, M. A., Yu, Y., Ulrich, R. L., Sarria, S. H., Nierman, W. C., and Deshazer, D. (2005). Bacterial genome adaptation to niches: divergence of the potential virulence genes in three Burkholderia species of different survival strategies. BMC Genomics 6, 174. doi: 10.1186/1471-2164-6-174
Koh, Y. S., and Roe, J. H. (1995). Isolation of a novel paraquat-inducible (pqi) gene regulated by the soxRS locus in Escherichia coli. J. Bacteriol. 177, 2673–2678.
Kooi, C., Subsin, B., Chen, R., Pohorelic, B., and Sokol, P. A. (2006). Burkholderia cenocepacia ZmpB is a broad-specificity zinc metalloprotease involved in virulence. Infect. Immun. 74, 4083–4093.
Kothe, M., Antl, M., Huber, B., Stoecker, K., Ebrecht, D., Steinmetz, I., and Eberl, L. (2003). Killing of Caenorhabditis elegans by Burkholderia cepacia is controlled by the cep quorum-sensing system. Cell. Microbiol. 5, 343–351.
Krachler, A. M., Sharma, A., Cauldwell, A., Papadakos, G., and Kleanthous, C. (2010). TolA modulates the oligomeric status of YbgF in the bacterial periplasm. J. Mol. Biol. 403, 270–285.
Liang, M. D., Bagchi, A., Warren, H. S., Tehan, M. M., Trigilio, J. A., Beasley-Topliffe, L. K., Tesini, B. L., Lazzaroni, J. C., Fenton, M. J., and Hellman, J. (2005). Bacterial peptidoglycan-associated lipoprotein: a naturally occurring toll-like receptor 2 agonist that is shed into serum and has synergy with lipopolysaccharide. J. Infect. Dis. 191, 939–948.
LiPuma, J. J. (2010). The changing microbial epidemiology in cystic fibrosis. Clin. Microbiol. Rev. 23, 299–323.
Loutet, S. A., Flannagan, R. S., Kooi, C., Sokol, P. A., and Valvano, M. A. (2006). A complete lipopolysaccharide inner core oligosaccharide is required for resistance of Burkholderia cenocepacia to antimicrobial peptides and bacterial survival in vivo. J. Bacteriol. 188, 2073–2080.
Loutet, S. A., and Valvano, M. A. (2010). A decade of Burkholderia cenocepacia virulence determinant research. Infect. Immun. 78, 4088–4100.
Mahenthiralingam, E., Baldwin, A., and Dowson, C. G. (2008). Burkholderia cepacia complex bacteria: opportunistic pathogens with important natural biology. J. Appl. Microbiol. 104, 1539–1551.
Mahenthiralingam, E., Campbell, M. E., and Speert, D. P. (1994). Nonmotility and phagocytic resistance of Pseudomonas aeruginosa isolates from chronically colonized patients with cystic fibrosis. Infect. Immun. 62, 596–605.
Mahenthiralingam, E., Coenye, T., Chung, J. W., Speert, D. P., Govan, J. R., Taylor, P., and Vandamme, P. (2000). Diagnostically and experimentally useful panel of strains from the Burkholderia cepacia complex. J. Clin. Microbiol. 38, 910–913.
Mahenthiralingam, E., Urban, T. A., and Goldberg, J. B. (2005). The multifarious, multireplicon Burkholderia cepacia complex. Nat. Rev. Microbiol. 3, 144–156.
Makidon, P. E., Knowlton, J., Groom, J. V. II, Blanco, L. P., LiPuma, J. J., Bielinska, A. U., and Baker, J. R. Jr. (2010). Induction of immune response to the 17 kDa OMPA Burkholderia cenocepacia polypeptide and protection against pulmonary infection in mice after nasal vaccination with an OMP nanoemulsion-based vaccine. Med. Microbiol. Immunol. 199, 81–92.
Marolda, C. L., Hauroder, B., John, M. A., Michel, R., and Valvano, M. A. (1999). Intracellular survival and saprophytic growth of isolates from the Burkholderia cepacia complex in free-living amoebae. Microbiology 145(Pt 7), 1509–1517.
Mougous, J. D., Cuff, M. E., Raunser, S., Shen, A., Zhou, M., Gifford, C. A., Goodman, A. L., Joachimiak, G., Ordonez, C. L., Lory, S., Walz, T., Joachimiak, A., and Mekalanos, J. J. (2006). A virulence locus of Pseudomonas aeruginosa encodes a protein secretion apparatus. Science 312, 1526–1530.
Munoz-Elias, E. J., and McKinney, J. D. (2005). Mycobacterium tuberculosis isocitrate lyases 1 and 2 are jointly required for in vivo growth and virulence. Nat. Med. 11, 638–644.
Munoz-Elias, E. J., Upton, A. M., Cherian, J., and McKinney, J. D. (2006). Role of the methylcitrate cycle in Mycobacterium tuberculosis metabolism, intracellular growth, and virulence. Mol. Microbiol. 60, 1109–1122.
Mutschler, H., Gebhardt, M., Shoeman, R. L., and Meinhart, A. (2011). A novel mechanism of programmed cell death in bacteria by toxin-antitoxin systems corrupts peptidoglycan synthesis. PLoS Biol. 9, e1001033. doi: 10.1371/journal.pbio.1001033
O’Grady, E. P., Nguyen, D. T., Weisskopf, L., Eberl, L., and Sokol, P. A. (2011). The Burkholderia cenocepacia LysR-type transcriptional regulator ShvR influences expression of quorum-sensing, protease, type II secretion, and afc genes. J. Bacteriol. 193, 163–176.
O’Grady, E. P., Viteri, D. F., Malott, R. J., and Sokol, P. A. (2009). Reciprocal regulation by the CepIR and CciIR quorum sensing systems in Burkholderia cenocepacia. BMC Genomics 10, 441. doi: 10.1186/1471-2164-10-441
Paterson, G. K., Northen, H., Cone, D. B., Willers, C., Peters, S. E., and Maskell, D. J. (2009). Deletion of tolA in Salmonella typhimurium generates an attenuated strain with vaccine potential. Microbiology 155, 220–228.
Peeters, E., Sass, A., Mahenthiralingam, E., Nelis, H., and Coenye, T. (2010). Transcriptional response of Burkholderia cenocepacia J2315 sessile cells to treatments with high doses of hydrogen peroxide and sodium hypochlorite. BMC Genomics 11, 90. doi: 10.1186/1471-2164-11-90
Potter, A. J., Kidd, S. P., Edwards, J. L., Falsetta, M. L., Apicella, M. A., Jennings, M. P., and McEwan, A. G. (2009). Thioredoxin reductase is essential for protection of Neisseria gonorrhoeae against killing by nitric oxide and for bacterial growth during interaction with cervical epithelial cells. J. Infect. Dis. 199, 227–235.
Rozen, S., and Skaletsky, H. (2000). Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–386.
Sajjan, S. U., Carmody, L. A., Gonzalez, C. F., and LiPuma, J. J. (2008). A type IV secretion system contributes to intracellular survival and replication of Burkholderia cenocepacia. Infect. Immun. 76, 5447–5455.
Sass, A., Marchbank, A., Tullis, E., Lipuma, J. J., and Mahenthiralingam, E. (2011). Spontaneous and evolutionary changes in the antibiotic resistance of Burkholderia cenocepacia observed by global gene expression analysis. BMC Genomics 12, 373. doi: 10.1186/1471-2164-12-373
Schmittgen, T. D., and Livak, K. J. (2008). Analyzing real-time PCR data by the comparative C(T) method. Nat. Protoc. 3, 1101–1108.
Schnappinger, D., Ehrt, S., Voskuil, M. I., Liu, Y., Mangan, J. A., Monahan, I. M., Dolganov, G., Efron, B., Butcher, P. D., Nathan, C., and Schoolnik, G. K. (2003). Transcriptional adaptation of Mycobacterium tuberculosis within macrophages: insights into the phagosomal environment. J. Exp. Med. 198, 693–704.
Seed, K. D., and Dennis, J. J. (2008). Development of Galleria mellonella as an alternative infection model for the Burkholderia cepacia complex. Infect. Immun. 76, 1267–1275.
Sisinni, L., Cendron, L., Favaro, G., and Zanotti, G. (2010). Helicobacter pylori acidic stress response factor HP1286 is a YceI homolog with new binding specificity. FEBS J. 277, 1896–1905.
Smith, A. W., Roche, H., Trombe, M. C., Briles, D. E., and Hakansson, A. (2002). Characterization of the dihydrolipoamide dehydrogenase from Streptococcus pneumoniae and its role in pneumococcal infection. Mol. Microbiol. 44, 431–448.
Sokol, P. A., Darling, P., Lewenza, S., Corbett, C. R., and Kooi, C. D. (2000). Identification of a siderophore receptor required for ferric ornibactin uptake in Burkholderia cepacia. Infect. Immun. 68, 6554–6560.
Sokol, P. A., Darling, P., Woods, D. E., Mahenthiralingam, E., and Kooi, C. (1999). Role of ornibactin biosynthesis in the virulence of Burkholderia cepacia: characterization of pvdA, the gene encoding L-Ornithine N5 -oxygenase. Infect. Immun. 67, 4443–4455.
Sokol, P. A., Sajjan, U., Visser, M. B., Gingues, S., Forstner, J., and Kooi, C. (2003). The CepIR quorum-sensing system contributes to the virulence of Burkholderia cenocepacia respiratory infections. Microbiology 149, 3649–3658.
Somvanshi, V. S., Viswanathan, P., Jacobs, J. L., Mulks, M. H., Sundin, G. W., and Ciche, T. A. (2010). The type 2 secretion pseudopilin, gspJ, is required for multihost pathogenicity of Burkholderia cenocepacia AU1054. Infect. Immun. 78, 4110–4121.
Subramoni, S., Nguyen, D. T., and Sokol, P. A. (2011). Burkholderia cenocepacia ShvR-regulated genes that influence colony morphology, biofilm formation, and virulence. Infect. Immun. 79, 2984–2997.
Tomich, M., Griffith, A., Herfst, C. A., Burns, J. L., and Mohr, C. D. (2003). Attenuated virulence of a Burkholderia cepacia type III secretion mutant in a murine model of infection. Infect. Immun. 71, 1405–1415.
Tomich, M., Herfst, C. A., Golden, J. W., and Mohr, C. D. (2002). Role of flagella in host cell invasion by Burkholderia cepacia. Infect. Immun. 70, 1799–1806.
Tuanyok, A., Tom, M., Dunbar, J., and Woods, D. E. (2006). Genome-wide expression analysis of Burkholderia pseudomallei infection in a hamster model of acute melioidosis. Infect. Immun. 74, 5465–5476.
Uehlinger, S., Schwager, S., Bernier, S. P., Riedel, K., Nguyen, D. T., Sokol, P. A., and Eberl, L. (2009). Identification of specific and universal virulence factors in Burkholderia cenocepacia strains by using multiple infection hosts. Infect. Immun. 77, 4102–4110.
Urban, T. A., Griffith, A., Torok, A. M., Smolkin, M. E., Burns, J. L., and Goldberg, J. B. (2004). Contribution of Burkholderia cenocepacia flagella to infectivity and inflammation. Infect. Immun. 72, 5126–5134.
Van Melderen, L., and Saavedra De Bast, M. (2009). Bacterial toxin-antitoxin systems: more than selfish entities? PLoS Genet. 5, e1000437. doi: 10.1371/journal.pgen.1000437
Velasco-Garcia, R., Villalobos, M. A., Ramirez-Romero, M. A., Mujica-Jimenez, C., Iturriaga, G., and Munoz-Clares, R. A. (2006a). Betaine aldehyde dehydrogenase from Pseudomonas aeruginosa: cloning, over-expression in Escherichia coli, and regulation by choline and salt. Arch. Microbiol. 185, 14–22.
Velasco-Garcia, R., Zaldivar-Machorro, V. J., Mujica-Jimenez, C., Gonzalez-Segura, L., and Munoz-Clares, R. A. (2006b). Disulfiram irreversibly aggregates betaine aldehyde dehydrogenase – a potential target for antimicrobial agents against Pseudomonas aeruginosa. Biochem. Biophys. Res. Commun. 341, 408–415.
Vergunst, A. C., Meijer, A. H., Renshaw, S. A., and O’Callaghan, D. (2010). Burkholderia cenocepacia creates an intramacrophage replication niche in zebrafish embryos, followed by bacterial dissemination and establishment of systemic infection. Infect. Immun. 78, 1495–1508.
Visser, M. B., Majumdar, S., Hani, E., and Sokol, P. A. (2004). Importance of the ornibactin and pyochelin siderophore transport systems in Burkholderia cenocepacia lung infections. Infect. Immun. 72, 2850–2857.
Weber, A., Kogl, S. A., and Jung, K. (2006). Time-dependent proteome alterations under osmotic stress during aerobic and anaerobic growth in Escherichia coli. J. Bacteriol. 188, 7165–7175.
Weingart, C. L., White, C. E., Liu, S., Chai, Y., Cho, H., Tsai, C. S., Wei, Y., Delay, N. R., Gronquist, M. R., Eberhard, A., and Winans, S. C. (2005). Direct binding of the quorum sensing regulator CepR of Burkholderia cenocepacia to two target promoters in vitro. Mol. Microbiol. 57, 452–467.
Werner, J., and Misra, R. (2005). YaeT (Omp85) affects the assembly of lipid-dependent and lipid-independent outer membrane proteins of Escherichia coli. Mol. Microbiol. 57, 1450–1459.
Winsor, G. L., Khaira, B., Van Rossum, T., Lo, R., Whiteside, M. D., and Brinkman, F. S. (2008). The Burkholderia Genome Database: facilitating flexible queries and comparative analyses. Bioinformatics 24, 2803–2804.
Wolfgang, M. C., Jyot, J., Goodman, A. L., Ramphal, R., and Lory, S. (2004). Pseudomonas aeruginosa regulates flagellin expression as part of a global response to airway fluid from cystic fibrosis patients. Proc. Natl. Acad. Sci. U.S.A. 101, 6664–6668.
Wright, J. R., and Clements, J. A. (1987). Metabolism and turnover of lung surfactant. Am. Rev. Respir. Dis. 136, 426–444.
Yoder-Himes, D. R., Chain, P. S., Zhu, Y., Wurtzel, O., Rubin, E. M., Tiedje, J. M., and Sorek, R. (2009). Mapping the Burkholderia cenocepacia niche response via high-throughput sequencing. Proc. Natl. Acad. Sci. U.S.A. 106, 3976–3981.
Yoder-Himes, D. R., Konstantinidis, K. T., and Tiedje, J. M. (2010). Identification of potential therapeutic targets for Burkholderia cenocepacia by comparative transcriptomics. PLoS ONE 5, e8724. doi: 10.1371/journal.pone.0008724
Zaldivar-Machorro, V. J., Lopez-Ortiz, M., Demare, P., Regla, I., and Munoz-Clares, R. A. (2011). The disulfiram metabolites S-methyl-N,N-diethyldithiocarbamoyl sulfoxide and S-methyl-N,N-diethylthiocarbamoyl sulfone irreversibly inactivate betaine aldehyde dehydrogenase from Pseudomonas aeruginosa, both in vitro and in situ, and arrest bacterial growth. Biochimie 93, 286–295.
Appendix
Keywords: Burkholderia cenocepacia, Burkholderia cepacia complex, microarray, lung infection, rat chronic respiratory infection model, in vitro, in vivo
Citation: O’Grady EP and Sokol PA (2011) Burkholderia cenocepacia differential gene expression during host–pathogen interactions and adaptation to the host environment. Front. Cell. Inf. Microbio. 1:15. doi: 10.3389/fcimb.2011.00015
Received: 30 August 2011; Accepted: 20 November 2011;
Published online: 09 December 2011.
Edited by:
Joanna Goldberg, University of Virginia Health System, USAReviewed by:
Leonilde Morais Moreira, Instituto Superior Técnico, PortugalEshwar Mahenthiralingam, Cardiff University, UK
Copyright: © 2011 O’Grady and Sokol. This is an open-access article distributed under the terms of the Creative Commons Attribution Non Commercial License, which permits non-commercial use, distribution, and reproduction in other forums, provided the original authors and source are credited.
*Correspondence: Pamela A. Sokol, Department of Microbiology, Immunology and Infectious Diseases, Faculty of Medicine, University of Calgary, 3330 Hospital Drive NW, Calgary, AB, Canada T2N 4N1. e-mail:cHNva29sQHVjYWxnYXJ5LmNh