- Department of Clinical and Biological Sciences, University of Torino, Orbassano, Italy
Dictyostelium cells are forest soil amoebae, which feed on bacteria and proliferate as solitary cells until bacteria are consumed. Starvation triggers a change in life style, forcing cells to gather into aggregates to form multicellular organisms capable of cell differentiation and morphogenesis. As a soil amoeba and a phagocyte that grazes on bacteria as the obligate source of food, Dictyostelium could be a natural host of pathogenic bacteria. Indeed, many pathogens that occasionally infect humans are hosted for most of their time in protozoa or free-living amoebae, where evolution of their virulence traits occurs. Due to these features and its amenability to genetic manipulation, Dictyostelium has become a valuable model organism for studying strategies of both the host to resist infection and the pathogen to escape the defense mechanisms. Similarly to higher eukaryotes, iron homeostasis is crucial for Dictyostelium resistance to invasive bacteria. Iron is essential for Dictyostelium, as both iron deficiency or overload inhibit cell growth. The Dictyostelium genome shares with mammals many genes regulating iron homeostasis. Iron transporters of the Nramp (Slc11A) family are represented with two genes, encoding Nramp1 and Nramp2. Like the mammalian ortholog, Nramp1 is recruited to phagosomes and macropinosomes, whereas Nramp2 is a membrane protein of the contractile vacuole network, which regulates osmolarity. Nramp1 and Nramp2 localization in distinct compartments suggests that both proteins synergistically regulate iron homeostasis. Rather than by absorption via membrane transporters, iron is likely gained by degradation of ingested bacteria and efflux via Nramp1 from phagosomes to the cytosol. Nramp gene disruption increases Dictyostelium sensitivity to infection, enhancing intracellular growth of Legionella or Mycobacteria. Generation of mutants in other “iron genes” will help identify genes essential for iron homeostasis and resistance to pathogens.
Introduction
Dictyostelium discoideum is a member of the Amoebozoa (Schilde and Schaap, 2013). The cells live as unicellular amoebae in deciduous forest soil, feeding on bacteria that are taken up by phagocytosis and dividing by binary fission. Exhaustion of the food supply triggers a shift from growth to development, resulting in cells gathering by chemotaxis into aggregates of several thousands of cells. The tight aggregates transform into elongated sausage-like multicellular organisms, called slugs, in which cells differentiate into pre-stalk and pre-spore subtypes. After extensive migration, the slug eventually culminates into a fruiting body, consisting of a slender stalk of vacuolated cells bearing on top a ball of fully differentiated spores (Kessin, 2001).
Due to their life cycle, easy handling and genetic tractability, D. discoideum (in the followings Dictyostelium) has long been a preferred model organism for studying basic processes, such as motility and chemotaxis, cell-cell communication and adhesion, cell differentiation, pattern formation and morphogenesis (Bozzaro, 2013). Their ability to phagocytose has been exploited, in the last decade, to investigate dynamics and regulatory pathways of phagocytosis as well as interactions with an increasing number of clinically relevant bacterial pathogens, including Legionella pneumophila, Mycobacterium avium or marinum, and Pseudomonas aeruginosa (Bozzaro et al., 2008, 2013; Cosson and Soldati, 2008; Clarke, 2010; Bozzaro and Eichinger, 2011; Steinert, 2011). In the light of these recent developments, in this review we will discuss the opportunities offered by Dictyostelium in investigating the role of divalent metal homeostasis, with a special emphasis on iron, for cell growth, and defense against pathogenic bacteria.
Dictyostelium: Professional Phagocyte and Pathogen Host
Wild type Dictyostelium strains are strictly dependent on bacteria for growth, though a few selected laboratory strains are able to grow in liquid axenic media by fluid-phase endocytosis, mostly macropinocytosis (Kessin, 2001; Maniak, 2001). Thousands of prokaryotic species are present in the forest soil; how many of them can serve as food for Dictyostelium is unknown, but the cells appear to be rather omnivorous. Soil bacteria which have been isolated in association with wild type strains include close relatives of Burkholderia xenovorans, Stenotrophomonas maltophilia, Enterobacter sakazakii, Pseudomonas fluorescens, and Flavobacterium johnsoniae (Brock et al., 2011). Under laboratory conditions, the cells are able to graze on a very large variety of Gram-negative and Gram-positive bacteria, including different species of Enterobacter, Serratia, Salmonella, Yersinia, Proteus, Aeromonas, Alcaligenes, Acinetobacter, Staphylococcus, Listeria, and Bacillus (Depraitere and Darmon, 1978). They are also capable of modulating their response to different types of bacteria by activating specific sets of gene transcripts (Farbrother et al., 2006; Carilla-Latorre et al., 2008; Sillo et al., 2008, 2011). In a recent paper, (Nasser et al., 2013) have studied global transcriptional response of wild type and selected mutant cells to a series of Gram-negative and Gram-positive bacteria, and they could show that cells respond differently to these two large families of bacteria. By analysing the transcriptional response to live, in contrast to dead bacteria, they were also able to identify selective gene pathways needed for defense, rather than growth, on either Gram-negative or Gram-positive bacteria, such as the activation of different sets of lysozymes or a set of glycoproteins apparently required for growth on Gram-positive bacteria.
Phagocytosis, both on agar plates or under shaking in simple salt solution, is very efficient. Only a handful of bacteria strains are not phagocytosed, but very few systematic studies have been published in this regard. Legionella pneumophila is taken up by macropinocytosis, thus its uptake by natural wild type strains is negligible (Peracino et al., 2010; Balest et al., 2011). Bacillus anthracis was reported not to be phagocytosed (Depraitere and Darmon, 1978), but a recent report described its uptake, though no details were offered on the efficiency of phagocytosis (Nasser et al., 2013). Human pathogenic Escherichia coli strains were found to be graze-resistant when co-cultured with Dictyostelium cells at high, but not low, density. Whether this was due to the high bacterial density inhibiting phagocytosis or killing the cells was not assessed, except for one strain (Adiba et al., 2010). A correlation between phagocytosis efficiency and bacterial density or previous growth conditions has been also described for Salmonella typhimurium (Sillo et al., 2011), Aeromonas spp. (Froquet et al., 2007) and for Klebsiella pneumoniae (March et al., 2013). Cells are also able to discriminate between edible and less appetizing bacteria. When co-cultured with S. typhimurium and E. coli, cells depleted E. coli from the medium leaving quite unaltered the Salmonella (Sillo et al., 2011). Co-culturing with live Gram-negative bacteria was also shown to prime the cells to grow on dead Gram-positive bacteria that otherwise were not utilized as food source (Nasser et al., 2013).
It is nowadays accepted that free-living amoebae and protozoa, by predating bacteria, can become a driving force for the evolution of pathogenic traits, paradigmatic cases being Acanthamoeba castellanii parasitic interaction with Legionella pneumophila (Barker and Brown, 1994; Levin, 1996; Steinert et al., 2000); or the variation of the rfb virulence locus in Salmonella enterica mediated by intestinal protozoan predation (Wildschutte et al., 2004). The ability of many pathogens to grow in macrophages and cause human diseases appears thus to be a consequence of their adaptation and survival in the normally hostile amoeboid niche (Steinert et al., 2000; Greub and Raoult, 2004; Casadevall, 2008).
Being a bacterial predator, it would be rather surprising that Dictyostelium would be an exception on this regard, though Dictyostelium parasites naturally occurring in the wild have not been described, likely due to lack of systematic studies. At least for the fungus Cryptococcus neoformans it was, however, shown that passage through Dictyostelium cells enhanced its virulence in mice (Steenbergen et al., 2003). Invasive pathogenic microbes which have been shown to be able to grow in Dictyostelium cells include, in addition to Cryptococcus, Legionella (Haegele et al., 2000; Solomon and Isberg, 2000), Mycobacteria (Solomon et al., 2003) and Burkholderia species (Hasselbring et al., 2011). S. typhimurium co-cultured under optimal nutrient conditions has been shown to enter wild type cells and kill the cells, but its intracellular replication has been documented in autophagic Dictyostelium mutants only (Jia et al., 2009; Sillo et al., 2011). Similarly, K. pneumoniae is pathogenic for some Dictyostelium mutants, not for the parental wild type strain (Benghezal et al., 2006). Capsulated Neisseria meningitidis (Colucci et al., 2008) as well as pathogenic Escherichia coli strains (Adiba et al., 2010) have been shown to resist degradation, but their intracellular growth has not been documented.
Since Dictyostelium can be easily grown in association with bacteria on agar plates, a plaque assay, and in some cases growth assay under shaking, have been used by several labs for screening microbial virulence genes, using either wild type or in some cases mutant cells. Virulence traits have thus been identified in E. coli (Adiba et al., 2010), Pseudomonas aeruginosa (Cosson et al., 2002; Pukatzki et al., 2002; Alibaud et al., 2008), Vibrio cholerae (Pukatzki et al., 2006; Miyata et al., 2011; Zheng et al., 2011), Stenotrophomonas aeruginosa and S. malthophylia (Alonso et al., 2004; Adamek et al., 2011), K. pneumoniae (Benghezal et al., 2006; Pan et al., 2011; March et al., 2013), Burkholderia coenocepacia and B. pseudomallei (Aubert et al., 2008; Hasselbring et al., 2011).
In contrast to these bacteria, Legionella pneumophila, Mycobacterium avium, or M. marinum grow rapidly intracellularly, and their interaction with Dictyostelium cells has been extensively studied [for recent reviews see (Bozzaro and Eichinger, 2011; Hilbi et al., 2011; Steinert, 2011)]. In vivo imaging of the dynamics of infection, favored by the large array of fluorescent probes against cytoskeletal and organelle proteins, has shown that the process is highly conserved between Dictyostelium and macrophages. Both Legionella and Mycobacteria manipulate the endocytic pathway to hinder fusion of the pathogen-containing phagosome with acidic and lysosomal vesicles, favoring association with other compartments, such as the endoplasmic reticulum (Fajardo et al., 2004; Lu and Clarke, 2005; Ragaz et al., 2008; Peracino et al., 2010) and mitochondria (Francione et al., 2009), generating a replication vacuole. Whereas massive intracellular growth of Legionella leads to cell lysis, M. marinum has been shown to escape intact cells by a novel non-lytic exocytic mechanism that may be active also in macrophages (Hagedorn et al., 2009). Genomic-wide transcriptional changes during infection and proteomic analysis of the Legionella-containing vacuole (LCV) have also helped in characterizing the dynamics of infection (Farbrother et al., 2006; Li et al., 2009; Shevchuk et al., 2009; Urwyler et al., 2009).
Infection assays have been exploited particularly with Legionella to identify and/or characterize virulence genes. Thus, it has been shown that Legionella mutants defective in sigmaS or its effector LqsR, which regulate expression of several genes involved in virulence, motility and transmission, are defective for growth in Dictyostelium and macrophages (Tiaden et al., 2007; Hovel-Miner et al., 2009). The Icm/Dot TFSS (Type Four Secretion System) substrate SdhA appears to hinder cell apoptosis, thus the mutant is defective for growth in macrophages and to a lower extent in Dictyostelium (Laguna et al., 2006). Mutants in the Icm/Dot effectors SidJ or SidjA (Liu and Luo, 2007), SidC or SidA (Ragaz et al., 2008) display impaired recruitment of endoplasmic reticulum to the LCV, with differential effects on intracellular growth, which was depressed in macrophages and Dictyostelium or Dictyostelium only for SidJ or SidjA, respectively, but unaltered in the SidC or SidA mutants. The Icm/Dot TFSS effectors LepA and LepB have been shown to regulate non-lytic exocytosis from the host (Chen et al., 2004), whereas Leg proteins regulate LCV traffic, by disrupting the endocytic vesicle traffic of the host (de Felipe et al., 2008). The envelope-associated protein EnhC was found to be required for bacterial growth in macrophages, but not in Dictyostelium (Liu et al., 2008). Among the ICM/Dot IV injected effectors, AnkyrinB (AnkB) was shown to be anchored via farnesylation to the LCV and be required for docking polyubiquinated host proteins, favoring intracellular growth, both in macrophages and in Dictyostelium (Al-Quadan and Kwaik, 2011).
From the site of the host, the large number of available Dictyostelium mutants has been exploited with Legionella or Mycobacteria to identify host resistance factors, whose genetic disruption leads to enhanced pathogen proliferation. They include cytoskeletal and signal transduction proteins, transcription factors, autophagy and mitochondrial proteins [for recent reviews see: (Bozzaro and Eichinger, 2011; Steinert, 2011)]. Among the several host cell factors so far identified, the iron transporters of the Nramp family have been investigated in detail.
Dictyostelium Nramp Iron Transporters in Bacterial Infection
Since the discovery of an allelic form of the gene encoding natural resistance associated membrane protein (Nramp)1, which conferred susceptibility to various intracellular microbes (Vidal et al., 1995), the number of studies on this metal transporter family has increased dramatically (Cellier, 2012). The Nramp family is widely distributed, from bacteria to humans (Courville et al., 2006). In mammals, Nramp1 (Slc11A1) expression is restricted to macrophages, but a second Nramp protein, (Nramp2, Slc11A2, or DMT1), is localized in the plasma membrane of several tissues, and mutations have been linked to severe microcytic anemia and serum and hepatic overload (Courville et al., 2006; Shawki et al., 2012). Eukaryotic Nramp proteins are Fe2+ and Mn2+, possibly Co2+, transporters (Forbes and Gros, 2001; Nevo and Nelson, 2006), whereas the bacterial homologs (MntH subfamily) transports mainly Mn2+ (Papp-Wallace and Maguire, 2006; Cellier, 2012).
The Dictyostelium genome harbors two genes encoding members of this family, which have been named Nramp1 and Nramp2. Nramp1 is the ortholog of mammalian Nramp1, and like the macrophage counterpart, it is localized exclusively in phagosomes or macropinosomes. The protein is recruited from trans-Golgi to phago- or macropinosomes shortly after their closure, and is then retrieved from the vesicles during their post-lysosomal maturation (Peracino et al., 2006). The nramp1 gene is expressed during growth, up-regulated upon incubation with bacteria and down-regulated upon starvation. In contrast, the Nramp2 protein is phylogenetically closer to α-proteobacteria MntH and to Nramp proteins from yeast, fungi and protists, and is exclusively localized in the membrane of the contractile vacuole (Peracino et al., 2013). The contractile vacuole is a bladder and tubular network, which in Dictyostelium, like other free-living amoebae and protozoa exposed to sudden environmental changes, regulates osmolarity. Under hypotonic conditions, water is pumped into the contractile vacuole, giving rise to large vacuoles that fuse with the plasma membrane, expelling their content. Under hypertonic conditions, bladder and tubules flatten, releasing water in the cytosol (Gerisch et al., 2002; Heuser, 2006).
The contractile vacuole membrane is studded with the V-H+ ATPase, which pumps protons inside the lumen (Heuser et al., 1993; Clarke et al., 2002). The V-H+ ATPase is also rapidly recruited to phagosomes or macropinosomes, shortly after their engulfment (Clarke et al., 2002; Peracino et al., 2006). Thus, both Nramp1 and Nramp2, though in two different compartments, co-localize with the vacuolar ATPase that can provide the electrogenic potential regulating their transport activity. Though still debated, it is likely that the proton gradient generated by the activity of the V-ATPase favors Nramp-dependent iron transport via a symport rather than antiport mechanism (Forbes and Gros, 2001; Courville et al., 2006; Nevo and Nelson, 2006). Experiments with purified phagosomes from Dictyostelium wild type and Nramp1-null cells supported this hypothesis, showing also that the protein was essential for iron transport (Peracino et al., 2006). It is likely that Nramp2 in the contractile vacuole acts similarly to Nramp1, but this has not been proven yet.
Both Nramp1 and Nramp2 are dispensable for phagocytosis or growth on non-pathogenic bacteria, but their disruption enhances intracellular growth of Legionella pneumophila and, at least for Nramp1, also Mycobacterium avium (Peracino et al., 2006, 2013). Constitutive Nramp1 overexpression depresses Legionella, not, however, Mycobacterium, growth. Interestingly, endogenous Nramp1 gene expression is down-regulated during Legionella, but not Mycobacterium infection (Peracino et al., 2006). These differences may be related to differences between both pathogens in the establishment of their replication vacuole. In Mycobacteria infection, the replication vacuole transiently recruits the vacuolar ATPase during the first 90 min post-infection, retrieves it between 2 and 12 h, avoiding recruitment of lysosomal enzymes while acquiring markers of a post-lysosomal compartment (Hagedorn and Soldati, 2007). Legionella instead forms a replicative vacuole that avoids fusion with acidic vesicles, associates with mitochondria and recruits proteins of the endoplasmic reticulum and other trafficking routes, similarly to what occurs in macrophages (Fajardo et al., 2004; Lu and Clarke, 2005; Francione et al., 2009). Fusion with vesicles decorated with Nramp1 is not inhibited, but recruitment of the V-H+ ATPase, and thus vacuole acidification, is delayed of several hours, though eventually occurs after 12–24 h post-infection (Peracino et al., 2010). Down-regulating Nramp1 expression, could thus help maintaining in the long run a replication-friendly vacuole, if acidification would occur.
Iron is an essential element for virtually all cells, and pathogens such as Legionella, Mycobacteria, or Salmonella are known to assimilate significant amounts of iron for their metabolism and virulence (Pope et al., 1996; Robey and Cianciotto, 2002; Rodriguez, 2006; Pandey and Rodriguez, 2012; Soldati and Neyrolles, 2012). A systematic analysis of human pathogenic Escherichia coli strains showed that E. coli genes involved in iron metabolism, such as irp, fyuA, and IroN, favored resistance to predation by Dictyostelium cells (Adiba et al., 2010). Thus, depleting iron from the phagosome via Nramp1 can be an effective host defense strategy to starve the pathogen for iron. Conversely, hindering co-recruitment of the V-H+ ATPase by the pathogen, as shown for Legionella, not only avoids acidification of the vacuole, thus neutralizing Nramp1-dependent iron efflux, but could even favor iron influx in the vacuole via Nramp1, thus turning Nramp1 to Legionella advantage (Peracino et al., 2010) (Figure 1). Experiments with isolated phagosomes showed indeed that inactivating the vacuolar ATPase resulted in passive iron flux (Peracino et al., 2006).
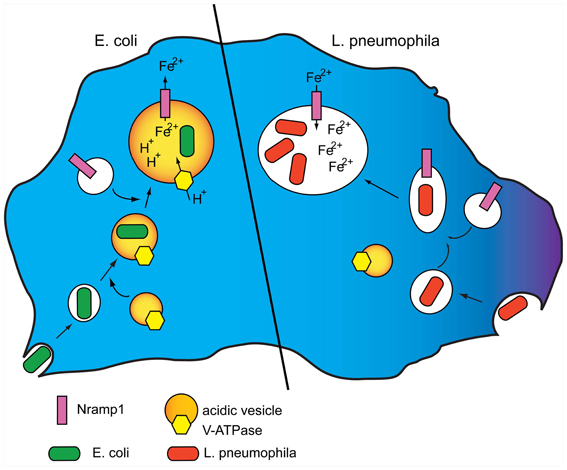
Figure 1. Model for Nramp1 activity and its manipulation by Legionella. (Left) Nramp1 and the V-H+ ATPase are recruited to phagosomes shortly after their uptake. The activity of the vacuolar ATPase generates a proton gradient in the maturing phagosome that provides the electrogenic force necessary for Nramp1 to transport iron, and possibly other divalent metals, to the cytosol, thus depleting the bacteria from an essential nutrient element. (Right) L. pneumophila is taken up in Dictyostelium cells by macropinocytosis. Following uptake, the pathogen inhibits fusion of its vacuole with acidic vesicles bearing the V-H+ ATPase. Nramp1 recruitment is not affected, but lack of the electrogenic force neutralizes Nramp1-dependent iron efflux, and can even lead to passive influx of cytosolic labile iron to the advantage of the pathogen. For original data see: Peracino et al. (2010).
Intracellular growth of L. pneumophila is enhanced in Nramp1-null mutants and inhibited in cells overexpressing Nramp1, but the latter effect was found to be reversed by phosphatidylinositide-3 kinase (PI3K) inhibitors or by genetic ablation of PI3K and the phosphatase PTEN (Peracino et al., 2010). The PI(4,5)P2 or PI(3,4,5)P3 phosphatase Dd5P4 (OCRL1) also enhanced Legionella growth (Weber et al., 2009), and PI3K inhibitors stimulated Legionella infection also in macrophages (Peracino et al., 2010). Membrane phosphatidylinositides are important regulators of actin assembly/disassembly in the plasma membrane and in phago- or macropinosomes as well as phago- and macropinosome fusion with vesicles of the endo-lysosomal pathway. It was shown that altering phosphoinositide metabolism in Legionella infection resulted in even stronger inhibition of the Legionella-containing macropinosome with acidic vesicles, a process apparently stimulated by PI3P formation (Clarke et al., 2010; Peracino et al., 2010). This has led to the hypothesis that Legionella hinders fusion of the Legionella-containing macropinosome with acidic vesicles, but not vesicles bearing Nramp1, by inhibiting PI(3)P formation either by secreting a 3-P phosphatase or by anchoring a 3-P phosphatase of the host to the replication vacuole.
The Genetic Basis of Iron Homeostasis in Dictyostelium
Dictyostelium cell growth is sensitive to iron depletion as well as to iron loading above 0.2 mM. When axenic wild type cells are incubated in minimal medium without iron, growth decreases dramatically and after 4–5 generations stops completely. Nramp1 or nramp2 knockout mutants fail to duplicate already after 2 generations. High iron concentrations reduce the growth rate in the wild type, to a lower extent in the nramp single knockout mutants, but only minimally in the double KO mutant. These results suggest that inactivating nramp1 and nramp2 leads to a lower intracellular level of bioavailable cellular iron, independently of the total amount that may enter the cell (Peracino et al., 2013). Dictyostelium development occurs under starving conditions in simple salt solutions, or even in water; addition of divalent transition metals is not required. During development, the only source of energy and organelle recycling is the autophagic breakdown of cellular components, which is responsible for the observed decrease in cell size (Kessin, 2001). Autophagy could also be the major source of iron during development, unless excess labile iron is accumulated during growth in the contractile vacuole, and the latter acts as iron reserve. It is worth mentioning that Nramp2 gene expression is stimulated by starvation, with maximal mRNA accumulation reached during aggregation and slug formation (Peracino et al., 2013).
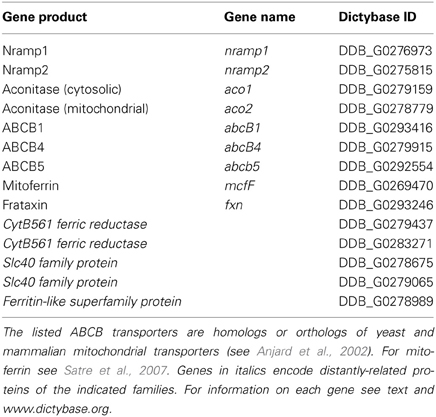
In addition to the Nramp iron transporters, the Dictyostelium genome encodes many proteins involved in cellular iron homeostasis (Table 1). Homologs of the mitochondrial iron transporter mitoferrin (Satre et al., 2007), Fe-S and heme ABCB transporters (Anjard et al., 2002), and the iron sensor frataxin as well as a cytosolic and a mitochondrial aconitases are present. The cytosolic aconitase (Aco1) is highly similar to mammalian Iron Regulatory Protein (IRP), raising the possibility that it may bind iron regulatory elements (IREs), thus regulating iron-dependent genes (Anderson et al., 2012). Two distantly-related ferroportin (Slc40A1 or IREG1)-like proteins exist, but no homologs for trasferrin or transferrin receptors, the systemic iron traffic regulator hepcidin or the hepcidin inhibitor hemojuvelin are found, in agreement with the notion that iron is mainly derived from bacterial digestion. Ferritin or mitoferritin homologs are also not evident, though a putative ferritin-like protein, but with very low homology to other ferritin-like proteins, is encoded in the genome.
Bacterial digestion in phago-lysosomes will result in ferric ions that need to be reduced for transport via Nramp1 in the cytosol. In macrophages, where the major source of iron is represented by aged erythrocytes and bacteria, this is accomplished by ferric reductases of the STEAP family (Wang and Pantopoulos, 2011), which have no orthologs in Dictyostelium genome. Two putative ferric reductases of the domon-cytB561 family are, however, encoded in the genome, one of which is highly expressed during growth (Table 1). Whether any of them is localized in the phagosome, and may be responsible for ferric ion reduction, is under study. It is clear, in any case, that cells manage to reduce ferric ions, as ferric chloride added to minimal medium stimulates cell growth (Peracino et al., 2013). As summarized in the model in Figure 2, it can be hypothesized that following ferric ion reduction, iron is exported from the phagosomes via Nramp1 in a proton-gradient dependent co-transport. Most cytosolic iron will be transported to the mitochondria via mitoferrin, to be incorporated into Fe-S clusters and heme groups, a process likely regulated by frataxin, as in mammalian cells (Wang and Pantopoulos, 2011). Excess labile iron in the cytosol could accumulate in the contractile vacuole, either to be released following CV discharge upon fusion with the membrane or to be recycled to the cytosol via Nramp2, particularly under starving conditions, when the only source of iron is intracellular. In this model, the contractile vacuole is proposed to act as homeostatic vesicle-bound iron store, taking over ferritin function. In principle, also a plasma membrane iron exporter, such as ferroportin, would be dispensable in this model. If the model is correct, new questions arise: how is iron transport in the contractile vacuole regulated? Is Nramp2 the only iron transporter and does it mediate both import and export? How does the electrogenic potential generated by the V-H+ ATPase in the CV membrane regulate Nramp2 activity, taking in consideration that protons are efficiently buffered in the contractile vacuole lumen (Clarke and Heuser, 1997), contrary to what occurs for Nramp1 in the phago-lysosomes?
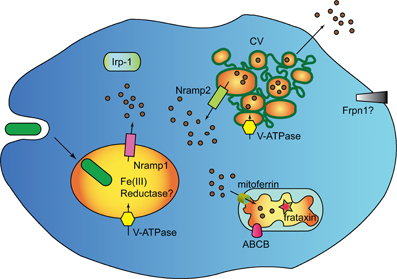
Figure 2. Genes regulating iron homeostasis in Dictyostelium. The major, if not unique, source if iron for Dictyostelium cells are engulfed bacteria which are degraded in phago-lysosomes. The Nramp1 transporter is recruited to phagosomes shortly after uptake and is retrieved during post-lysosomal maturation. Nramp2 is localized in the membrane of the contractile vacuole (CV). It is assumed that Nramp2 acts like Nramp1, due to co-localization of the V-H+ ATPase in both phagosome and CV. The CV lumen is, however, not acidic, due to rapid buffering of the pumped protons, thus Nramp2 activity and directionality of transport could be modulated by other unknown factors. In addition to the Nramp transporters, the Dictyostelium genomes harbors homologs of the mitochondrial iron transporter mitoferrin, the iron sensor frataxin, and mitochondrial Fe-S and heme exporters of the ABCB family. In addition to mitochondrial ferrochelatase and aconitase (not shown), the genome encodes a homolog of mammalian cytosolic aconitase IRP-1, which may act as iron-regulatory-protein by binding to iron-regulatory elements (IRE) in target proteins. Distant homologs of the membrane iron exporter ferroportin are also encoded in the genome, as well as two putative ferric reductases. It is not known whether any of the ferric reductases is located in phagosomes. Dictyostelium does not possess homologs of transferrin or transferrin receptors nor mitoferritin or ferritin, but a very distantly related ferritin-like protein. It is proposed that the CV acts as membrane-bound labile iron store, secreting excess iron by fusing with the plasma membrane or transporting iron back in the cytosol via Nramp2, when needed. In this model, the CV takes over ferritin, and possibly also ferroportin, function.
Not Only Iron
In addition to iron, major divalent transitions metals that have been involved in host-pathogen interactions are manganese, copper, and zinc (Kehl-Fie and Skaar, 2010; Botella et al., 2012; Soldati and Neyrolles, 2012). Whereas iron and manganese are depleted from mature phagosomes, copper, and possibly zinc have been suggested to be pumped in to intoxicate bacteria; increased concentrations of both metals have been reported in mycobacterial infection after stimulation of infected macrophages with cytokines (Wagner et al., 2006).
Very little is known about these metals in Dictyostelium. In contrast to iron, cell growth is unaffected by both manganese depletion or millimolar addition to minimal medium, and manganese does not rescue cell growth in iron-depleted medium (Peracino, unpublished results). Manganese can, however, stimulate cell aggregation and cell differentiation, as it directly stimulates the activity of adenylyl cyclase (Loomis et al., 1978; Hagmann, 1985) and of a bi-functional glycosyltransferase that regulates O2-dependent development (Trinchera and Bozzaro, 1996; West et al., 2010). Dictyostelium development is also sensitive to heavy metals present in the soil. Hg in particular inhibits development at concentrations of 50 mg per kg dry soil, compared to about 6-, 16-, and 32-fold higher concentrations for Fe, Zn, and Cu, respectively (Ponte et al., 2000; Balbo and Bozzaro, 2009; Rodriguez-Ruiz et al., 2013).
The Dictyostelium genome encodes several putative copper and zinc transporters. A Menkes type Cu2+ ATPase and a putative p80 copper transporter have been shown to be localized in the plasma membrane and in phagosomes. The unusual resistance to Cu has been linked to the high secretory efficiency of the Cu2+ ATPase (Burlando et al., 2002), which in the phagosome could pump Cu2+ ions in the lumen, favoring a potentially toxic effect of this metal on bacteria. A chemical assay for detection of metallothioneins, which could also account for copper resistance, failed to detect any activity, suggesting that Dictyostelium cells do not express these proteins (Burlando et al., 2002). The Dictyostelium genome harbors, however, one gene (DDB_G0288281), which has been annotated as putative metallothionein. Metallothioneins are small cysteine-rich proteins capable of binding heavy metals through the thiol group of their cysteine residues. Unfortunately, their primary structure is extremely variable and their secondary and tertiary structures are highly heterogeneous, making it difficult to recognize homologs among phyla, sometimes even among species (Coyle et al., 2002). Thus, whether the DDB_G0288281gene product is truly a metallothionein is open.
The p80 copper transporter is present in the plasma membrane and in endocytic vesicles. The protein is retrieved from the phagosome just after closure, but is then recruited again to mature phagosomes, probably by fusion with endocytic vesicles (Ravanel et al., 2001). In M. marinum infection, p80 has been shown to be recruited to the pathogen-replication vacuole at the onset of intravacuolar growth (Hagedorn and Soldati, 2007). Based on homology with other channel transporters, p80 is expected to transport copper outside of the vacuole, but no functional studies have been done, nor it is known whether disruption or overexpression of the gene affects host-pathogen interactions. The Dictyostelium zinc transporter family includes 12 members, most of which are expressed in late development and are assumed to be involved in cell differentiation (Sunaga et al., 2008). No data are available on potential localization of zinc transporters in phagosomes and their role during growth or host-pathogen interactions.
Recently, attention has been driven to a potential involvement of Mg2+ in nutritional immunity, based on results showing that synthesis of the M. tuberculosis virulence factor isoTb, which is involved in phagosome maturation arrest, was up- or down-regulated by Mg2+ overloading or depletion, respectively (Mann et al., 2011; Soldati and Neyrolles, 2012). In Dictyostelium, a V P-ATPase was identified as the tagged gene in the kil2 mutant, characterized by reduced phagosomal protease activity and defective growth on Klebsiella pneumoniae (Lelong et al., 2011). Addition of Mg2+ rescued the mutant, suggesting that the V P-ATPase could be a magnesium pump responsible for optimal Mg2+ concentration in the phagosome. Intriguingly, the kil2 mutant was able to grow on other bacteria species and did not display increased susceptibility to M. marinum infection.
Conclusion
As obligate phagocytes, at least during the growth phase of their life cycle, Dictyostelium cells resemble macrophages for their ability to engulf bacteria and dead cells, to discriminate between self and non-self and to fight potential pathogens. They also share with macrophages several “iron genes” regulating cellular iron metabolism, though lacking genes involved in systemic iron homeostasis. The organism is thus a useful model for investigating iron homeostasis at cellular, rather than systemic, level and the role of iron (and other divalent metals) in host-pathogen interactions. The potential absence of ferritin, and possibly also ferroportin, homologs raises the question of how labile iron is neutralized and stored in the cell. The possibility that the contractile vacuole acts as iron (divalent metal ions?) store and/or sink, thus taking over the functions of ferritin and ferroportin, is very suggestive, opening novel potential roles for this compartment also in free-living amoebae and protozoa, but needs to be proven. Investigating the mechanism of action of Nramp2 can be of help in this context as well as developing more sensitive probes for detecting iron in intracellular compartments. The highly conserved function of Nramp1 in the phago-lysosomal membrane, from Dictyostelium to macrophages, corroborates the central role of iron control for the host and the pathogen in this compartment. The finding that Nramp2 KO mutants are sensitive to infection almost as much as Nramp1 KO mutants raises the question of potential cross-talk between the phago-lysosome (or the pathogen-replication vacuole) and the contractile vacuole in iron homeostasis and in resistance to pathogens. The ease in generating and analysing mutants in Dictyostelium will help in the near future in dissecting the role of mitochondrial and cytosolic iron genes in these intracellular interactions and in extending these studies to genes regulating metabolism of other divalent metals relevant for host-pathogen interactions.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
Work in the lab was supported by research grants of the University of Torino and by the Compagnia San Paolo (12-CSP-C03-065).
References
Adamek, M., Overhage, J., Bathe, S., Winter, J., Fischer, R., and Schwartz, T. (2011). Genotyping of environmental and clinical Stenotrophomonas maltophilia isolates and their pathogenic potential. PLoS ONE 6:e27615. doi: 10.1371/journal.pone.0027615
Adiba, S., Nizak, C., van Baalen, M., Denamur, E., and Depaulis, F. (2010). From grazing resistance to pathogenesis: the coincidental evolution of virulence factors. PLoS ONE 5:e11882. doi: 10.1371/journal.pone.0011882
Alibaud, L., Kohler, T., Coudray, A., Prigent-Combaret, C., Bergeret, E., Perrin, J., et al. (2008). Pseudomonas aeruginosa virulence genes identified in a Dictyostelium host model. Cell. Microbiol. 10, 729–740. doi: 10.1111/j.1462-5822.2007.01080.x
Alonso, A., Morales, G., Escalante, R., Campanario, E., Sastre, L., and Martinez, J. (2004). Overexpression of the multidrug efflux pump SmeDEF impairs Stenotrophomonas maltophilia phylogeny. J. Antimicrob. Chemother. 53, 432–434. doi: 10.1093/jac/dkh074
Al-Quadan, T., and Kwaik, Y. (2011). Molecular characterization of exploitation of the polyubiquitination and farnesylation machineries of Dictyostelium discoideum by the AnkB F-box effector of Legionella pneumophila. Front. Microbiol. 2:23. doi: 10.3389/fmicb.2011.00023
Anderson, C., Shen, M., Eisenstein, R., and Leibold, E. (2012). Mammalian iron metabolism and its control by iron regulatory proteins. Biochim. Biophys. Acta 1823, 1468–1483. doi: 10.1016/j.bbamcr.2012.05.010
Anjard, C., Consortium, T. D. S., and Loomis, W. F. (2002). Evolutionary analyses of ABC transporters of Dictyostelium discoideum. Eukaryot. Cell 1, 643–652. doi: 10.1128/EC.1.4.643-652.2002
Aubert, D. F., Flannagan, R. S., and Valvano, M. A. (2008). A novel sensor kinase-response regulator hybrid controls biofilm formation and type VI secretion system activity in Burkholderia cenocepacia. Infect. Immun. 76, 1979–1991. doi: 10.1128/IAI.01338-07
Balbo, A., and Bozzaro, S. (2009). A novel bioassay for evaluating soil bio-hazards using Dictyostelium as biosensor: validation and application to the Bio-Bio project. Fresenius Environ. Bull. 17, 1137–1143.
Balest, A., Peracino, B., and Bozzaro, S. (2011). Legionella pneumophila infection is enhanced in a RacH-null mutant of Dictyostelium. Commun. Integr. Biol. 4, 194–197. doi: 10.4161/cib.4.2.14381
Barker, J., and Brown, M. (1994). Trojan horses of the microbial world: protozoa and the survival of bacterial pathogens in the environment. Microbiology 140, 1253–1259. doi: 10.1099/00221287-140-6-1253
Benghezal, M., Fauvarque, M. O., Tournebize, R., Froquet, R., Marchetti, A., Bergeret, E., et al. (2006). Specific host genes required for the killing of Klebsiella bacteria by phagocytes. Cell. Microbiol. 8, 139–148. doi: 10.1111/j.1462-5822.2005.00607.x
Botella, H., Stadthagen, G., Lugo-Villarino, G., de Chastellier, C., and Neyrolles, O. (2012). Metallobiology of host-pathogen interactions: an intoxitaging new insight. Trends Microbiol. 20, 106–112. doi: 10.1016/j.tim.2012.01.005
Bozzaro, S. (2013). The model organism Dictyostelium discoideum. Methods Mol. Biol. 983, 17–37. doi: 10.1007/978-1-62703-302-2_2
Bozzaro, S., Bucci, C., and Steinert, M. (2008). Phagocytosis and host-pathogen interactions in Dictyostelium with a look at macrophages. Int. Rev. Cell Mol. Biol. 271, 253–300. doi: 10.1016/S1937-6448(08)01206-9
Bozzaro, S., and Eichinger, L. (2011). The professional phagocyte Dictyostelium discoideum as a model host for bacterial pathogens. Curr. Drug Targets 12, 942–954. doi: 10.2174/138945011795677782
Bozzaro, S., Peracino, B., and Eichinger, L. (2013). Dictyostelium host response to Legionella infection: strategies and assays. Methods Mol. Biol. 954, 417–438. doi: 10.1007/978-1-62703-161-5_26
Brock, D. A., Douglas, T. E., Queller, D. C., and Strassmann, J. E. (2011). Primitive agriculture in a social amoeba. Nature 469, 393–396. doi: 10.1038/nature09668
Burlando, B., Evangelisti, V., Dondero, F., Pons, G., Camakaris, J., and Viarengo, A. (2002). Occurrence of Cu-ATPase in Dictyostelium: possible role in resistance to copper. Biochem. Biophys. Res. Commun. 291, 476–483. doi: 10.1006/bbrc.2002.6463
Carilla-Latorre, S., Calvo-Garrido, J., Bloomfield, G., Skelton, J., Kay, R. R., Ivens, A., et al. (2008). Dictyostelium transcriptional responses to Pseudomonas aeruginosa: common and specific effects from PAO1 and PA14 strains. BMC Microbiol. 8:109. doi: 10.1186/1471-2180-8-109
Casadevall, A. (2008). Evolution of intracellular pathogens. Ann. Rev. Microbiol. 62, 19–33. doi: 10.1146/annurev.micro.61.080706.093305
Cellier, M. (2012). Nutritional immunity: homology modeling of Nramp metal import. Adv. Exp. Med. Biol. 946, 335–351. doi: 10.1007/978-1-4614-0106-3_19
Chen, J., Defelipe, K. S., Clarke, M., Lu, H., Anderson, O. R., Segal, G., et al. (2004). Legionella effectors that promote nonlytic release from protozoa. Science 303, 1358–1361. doi: 10.1126/science.1094226
Clarke, M. (2010). Recent insights into host-pathogen interactions from Dictyostelium. Cell. Microbiol. 12, 283–291. doi: 10.1111/j.1462-5822.2009.01413.x
Clarke, M., and Heuser, J. (1997). “Water and ion transport,” in Dictyostelium—A Model System for Cell and Developmental Biology, eds Y. Maeda, K. Inouye, and I. Takeuchi (Tokyo: Universal Academy Press), 75–91.
Clarke, M., Kohler, J., Arana, Q., Liu, T. Y., Heuser, J., and Gerisch, G. (2002). Dynamics of the vacuolar H+-ATPase in the contractile vacuole complex and the endosomal pathway of Dictyostelium cells. J. Cell Sci. 115, 2893–2905.
Clarke, M., Maddera, L., Engel, U., and Gerisch, G. (2010). Retrieval of the vacuolar H-ATPase from phagosomes revealed by live cell imaging. PLoS ONE 5:e8585. doi: 10.1371/journal.pone.0008585
Colucci, A. M., Peracino, B., Tala, A., Bozzaro, S., Alifano, P., and Bucci, C. (2008). Dictyostelium discoideum as a model host for meningococcal pathogenesis. Med. Sci. Monit. 14, BR134–BR140.
Cosson, P., and Soldati, T. (2008). Eat, kill or die: when amoeba meets bacteria. Curr. Opin. Microbiol. 11, 271–276. doi: 10.1016/j.mib.2008.05.005
Cosson, P., Zulianello, L., Join-Lambert, O., Faurisson, F., Gebbie, L., Benghezal, M., et al. (2002). Pseudomonas aeruginosa virulence analyzed in a Dictyostelium discoideum host system. J. Bacteriol. 184, 3027–3033. doi: 10.1128/JB.184.11.3027-3033.2002
Courville, P., Chaloupka, R., and Cellier, M. (2006). Recent progress in structure-function analyses of Nramp proton-dependent metal-ion transporters. Biochem. Cell Biol. 84, 960–978. doi: 10.1139/o06-193
Coyle, P., Philcox, J., Carey, L., and Rofe, A. (2002). Metallothionein: the multipurpose protein. Cell. Mol. Life Sci. 2002, 627–647. doi: 10.1007/s00018-002-8454-2
de Felipe, K. S., Glover, R. T., Charpentier, X., Anderson, O. R., Reyes, M., Pericone, C. D., et al. (2008). Legionella eukaryotic-like type IV substrates interfere with organelle trafficking. PLoS Pathog. 4:e1000117. doi: 10.1371/journal.ppat.1000117.
Depraitere, C., and Darmon, M. (1978). Croissance de l'amibe sociale Dictyostelium discoideum sur differentes especes bacteriennes. Ann. Microbiol. (Institute Pasteur) 129B, 451–461.
Fajardo, M., Schleicher, M., Noegel, A., Bozzaro, S., Killinger, S., Heuner, K., et al. (2004). Calnexin, calreticulin and cytoskeleton-associated proteins modulate uptake and growth of Legionella pneumophila in Dictyostelium discoideum. Microbiology 150, 2825–2835. doi: 10.1099/mic.0.27111-0
Farbrother, P., Wagner, C., Na, J. B., Tunggal, B., Morio, T., Urushihara, H., et al. (2006). Dictyostelium transcriptional host cell response upon infection with Legionella. Cell. Microbiol. 8, 438–456. doi: 10.1111/j.1462-5822.2005.00633.x
Forbes, J., and Gros, P. (2001). Divalent-metal transport by NRAMP proteins at the interface of host-pathogen interactions. Trends Microbiol. 9, 397–403. doi: 10.1016/S0966-842X(01)02098-4
Francione, L., Smith, P. K., Accari, S. L., Taylor, P. E., Bokko, P. B., Bozzaro, S., et al. (2009). Legionella pneumophila multiplication is enhanced by chronic AMPK signalling in mitochondrially diseased Dictyostelium cells. Dis. Model Mech. 2, 479–489. doi: 10.1242/dmm.003319
Froquet, R., Cherix, N., Burr, S. E., Frey, J., Vilches, S., Tomas, J. M., et al. (2007). Alternative host model to evaluate Aeromonas virulence. Appl. Environ. Microbiol. 73, 5657–5659. doi: 10.1128/AEM.00908-07
Gerisch, G., Heuser, J., and Clarke, M. (2002). Tubular-vesicular transformation in the contractile vacuole system of Dictyostelium. Cell Biol. Int. 26, 845–852. doi: 10.1006/cbir.2002.0938
Greub, G., and Raoult, D. (2004). Microorganisms resistant to free-living amoebae. Clin. Microbiol. Rev. 17, 413–433. doi: 10.1128/CMR.17.2.413-433.2004
Haegele, S., Kohler, R., Merkert, H., Schleicher, M., Hacker, J., and Steinert, M. (2000). Dictyostelium discoideum: a new host model system for intracellular pathogens of the genus Legionella. Cell. Microbiol. 2, 165–171. doi: 10.1046/j.1462-5822.2000.00044.x
Hagedorn, M., Rohde, K. H., Russell, D. G., and Soldati, T. (2009). Infection by tubercular mycobacteria is spread by nonlytic ejection from their amoeba hosts. Science 323, 1729–1733. doi: 10.1126/science.1169381
Hagedorn, M., and Soldati, T. (2007). Flotillin and RacH modulate the intracellular immunity of Dictyostelium to Mycobacterium marinum infection. Cell. Microbiol. 9, 2716–2733. doi: 10.1111/j.1462-5822.2007.00993.x
Hagmann, J. (1985). Adenylate cyclase of Dictyostelium discoideum: solubilization and Mn2+-dependency. Cell Biol. Int. Reports 9, 491–494. doi: 10.1016/0309-1651(85)90157-2
Hasselbring, B. M., Patel, M. K., and Schell, M. A. (2011). Dictyostelium discoideum as a model system for identification of Burkholderia pseudomallei virulence factors. Infect. Immun. 79, 2079–2088. doi: 10.1128/IAI.01233-10
Heuser, J. (2006). Editorial: evidence for recycling of contractile vacuole membrane during osmoregulation in Dictyostelium amoebae—a tribute to Gunther Gerisch. Eur. J. Cell Biol. 85, 859–871. doi: 10.1016/j.ejcb.2006.05.011
Heuser, J., Zhu, Q. L., and Clarke, M. (1993). Proton pumps populate the contractile vacuoles of Dictyostelium amoebae. J. Cell Biol. 121, 1311–1327. doi: 10.1083/jcb.121.6.1311
Hilbi, H., Weber, S., and Finsel, I. (2011). Anchors for effectors: subversion of phosphoinositide lipids by Legionella. Front. Microbiol. 2:91. doi: 10.3389/fmicb.2011.00091
Hovel-Miner, G., Pampou, S., Faucher, S., Clarke, M., Morozova, I., Morozov, P., et al. (2009). SigmaS controls multiple pathways associated with intracellular multiplication of Legionella pneumophila. J. Bacteriol. 191, 2461–2473. doi: 10.1128/JB.01578-08
Jia, K., Thomas, C., Akbar, M., Sun, Q., Adams-Huet, B., Gilpin, C., et al. (2009). Autophagy genes protect against Salmonella typhimurium infection and mediate insulin signaling-regulated pathogen resistance. Proc. Natl. Acad. Sci. U.S.A. 106, 14564–14569. doi: 10.1073/pnas.0813319106
Kehl-Fie, T., and Skaar, E. (2010). Nutritional immunity beyond iron: a rolefor manganese and zinc. Curr. Opin. Chem. Biol. 14, 218–224. doi: 10.1016/j.cbpa.2009.11.008
Kessin, R. H. (2001). Dictyostelium—Evolution, Cell Biology, and the Development of Multicellularity. Cambridge: Cambridge University Press. doi: 10.1017/CBO9780511525315
Laguna, R. K., Creasey, E. A., Li, Z., Valtz, N., and Isberg, R. R. (2006). A Legionella pneumophila-translocated substrate that is required for growth within macrophages and protection from host cell death. Proc. Natl. Acad. Sci. U.S.A. 103, 18745–18750. doi: 10.1073/pnas.0609012103
Lelong, E., Marchetti, A., Gueho, A., Lima, W. C., Sattler, N., Molmeret, M., et al. (2011). Role of magnesium and a phagosomal P-type ATPase in intracellular bacterial killing. Cell. Microbiol. 13, 246–258. doi: 10.1111/j.1462-5822.2010.01532.x
Levin, B. (1996). The evolution and maintenance of virulence in microparasites. Emerg. Infect. Dis. 2, 93–102. doi: 10.3201/eid0202.960203
Li, Z., Dugan, A. S., Bloomfield, G., Skelton, J., Ivens, A., Losick, V., et al. (2009). The amoebal MAP kinase response to Legionella pneumophila is regulated by DupA. Cell Host Microbe 6, 253–267. doi: 10.1016/j.chom.2009.08.005
Liu, M., Conover, G. M., and Isberg, R. R. (2008). Legionella pneumophila EnhC is required for efficient replication in tumour necrosis factor alpha-stimulated macrophages. Cell. Microbiol. 10, 1906–1923. doi: 10.1111/j.1462-5822.2008.01180.x
Liu, Y., and Luo, Z. (2007). The Legionella pneumophila effector SidJ is required for efficient recruitment of endoplasmic reticulum proteins to the bacterial phagosome. Infect. Immun. 75, 592–603. doi: 10.1128/IAI.01278-06
Loomis, W. F., Klein, C., and Brachet, P. (1978). The effect of divalent cations on aggregation of Dictyostelium discoideum. Differentiation 12, 83–89. doi: 10.1111/j.1432-0436.1979.tb00993.x
Lu, H., and Clarke, M. (2005). Dynamic properties of Legionella-containing phagosomes in Dictyostelium amoebae. Cell. Microbiol. 7, 995–1007. doi: 10.1111/j.1462-5822.2005.00528.x
Maniak, M. (2001). Fluid-phase uptake and transit in axenic Dictyostelium cells. Biochim. Biophys. Acta 1525, 197–204. doi: 10.1016/S0304-4165(01)00105-2
Mann, F., Vanderven, B., and Peters, R. (2011). Magnesium depletion triggers production of an immune modulating diterpenoid in Mycobacterium tuberculosis. Mol. Microbiol. 79, 1594–1601. doi: 10.1111/j.1365-2958.2011.07545.x
March, C., Cano, V., Moranta, D., Llobet, E., Pérez-Gutiérrez, C., Tomàs, J., et al. (2013). Role of bacterial surface structures on the interaction of Klebsiella pneumoniae with phagocytes. PLoS ONE 8:e56847. doi: 10.1371/journal.pone.0056847
Miyata, S. T., Kitaoka, M., Brooks, T. M., McAuley, S. B., and Pukatzki, S. (2011). Vibrio cholerae requires the type VI secretion system virulence factor VasX to kill Dictyostelium discoideum. Infect. Immun. 79, 2941–2949. doi: 10.1128/IAI.01266-10
Nasser, W., Santhanam, B., Miranda, E., Parikh, A., Juneja, K., Rot, G., et al. (2013). Bacterial discrimination by dictyostelid amoebae reveals the complexity of ancient interspecies interactions. Curr. Biol. 23, 862–872. doi: 10.1016/j.cub.2013.04.034
Nevo, Y., and Nelson, N. (2006). The NRAMP family of metal-ion transporters. Biochim. Biophys. Acta 1763, 609–620. doi: 10.1016/j.bbamcr.2006.05.007
Pan, Y. J., Lin, T. L., Hsu, C. R., and Wang, J. T. (2011). Use of a Dictyostelium model for isolation of genetic loci associated with phagocytosis and virulence in Klebsiella pneumoniae. Infect. Immun. 79, 997–1006. doi: 10.1128/IAI.00906-10
Pandey, R., and Rodriguez, G. (2012). A ferritin mutant of Mycobacterium tuberculosis is highly susceptible to killing by antibiotics and is unable to establish a chronic infection in mice. Infect. Immun. 80, 3650–3659. doi: 10.1128/IAI.00229-12
Papp-Wallace, K., and Maguire, M. (2006). Manganese transport and the role of manganese in virulence. Annu. Rev. Microbiol. 60, 187–209. doi: 10.1146/annurev.micro.60.080805.142149
Peracino, B., Balest, A., and Bozzaro, S. (2010). Phosphoinositides differentially regulate bacterial uptake and Nramp1-induced resistance to Legionella infection in Dictyostelium. J. Cell. Sci. 123, 4039–4051. doi: 10.1242/jcs.072124
Peracino, B., Buracco, S., and Bozzaro, S. (2013). The Nramp (Slc11A) proteins regulate development, resistance to pathogenic bacteria and iron homeostasis in Dictyostelium discoideum. J. Cell. Sci. 126, 301–311. doi: 10.1242/jcs.116210
Peracino, B., Wagner, C., Balest, A., Balbo, A., Pergolizzi, B., Noegel, A. A., et al. (2006). Function and mechanism of action of Dictyostelium Nramp1 (Slc11a1) in bacterial infection. Traffic 7, 22–38. doi: 10.1111/j.1600-0854.2005.00356.x
Ponte, E., Rivero, F., Fechheimer, M., Noegel, A., and Bozzaro, S. (2000). Severe developmental defects in Dictyostelium null mutants for actin-binding proteins. Mech. Dev. 91, 153–161. doi: 10.1016/S0925-4773(99)00292-0
Pope, C., O'Connell, W., and Cianciotto, N. (1996). Legionella pneumophila mutants that are defective for iron acquisition and assimilation and intracellular infection. Infect. Immunol. 64, 629–636.
Pukatzki, S., Kessin, R. H., and Mekalanos, J. J. (2002). The human pathogen Pseudomonas aeruginosa utilizes conserved virulence pathways to infect the social amoeba Dictyostelium discoideum. Proc. Natl. Acad. Sci. U.S.A. 99, 3159–3164. doi: 10.1073/pnas.052704399
Pukatzki, S., Ma, A. T., Sturtevant, D., Krastins, B., Sarracino, D., Nelson, W. C., et al. (2006). Identification of a conserved bacterial protein secretion system in Vibrio cholerae using the Dictyostelium host model system. Proc. Natl. Acad. Sci. U.S.A. 103, 1528–1533. doi: 10.1073/pnas.0510322103
Ragaz, C., Pietsch, H., Urwyler, S., Tiaden, A., Weber, S. S., and Hilbi, H. (2008). The Legionella pneumophila phosphatidylinositol-4 phosphate-binding type IV substrate SidC recruits endoplasmic reticulum vesicles to a replication-permissive vacuole. Cell. Microbiol. 10, 2416–2433. doi: 10.1111/j.1462-5822.2008.01219.x
Ravanel, K., de Chassey, B., Cornillon, S., Benghezal, M., Zulianello, L., Gebbie, L., et al. (2001). Membrane sorting in the endocytic and phagocytic pathway of Dictyostelium discoideum. Eur. Cell Biol. 80, 754–764. doi: 10.1078/0171-9335-00215
Robey, M., and Cianciotto, N. (2002). Legionella pneumophila feoAB promotes ferrous iron uptake and intracellular infection. Infect. Immunol. 70, 5659–5669. doi: 10.1128/IAI.70.10.5659-5669.2002
Rodriguez, G. (2006). Control of iron metabolism in Mycobacterium tuberculosis. Trends Microbiol. 14, 320–327. doi: 10.1016/j.tim.2006.05.006
Rodriguez-Ruiz, A., Marigomez, I., Boatti, L., and Viarengo, A. (2013). Dictyostelium discoideum developmental cycle (DDDC) assay: a tool for Hg toxicity assessment and soil health screening. Sci. Total Environ. 450–451, 39–50. doi: 10.1016/j.scitotenv.2013.01.060
Satre, M., Mattei, S., Aubry, L., Gaudet, P., Pelosi, L., Brandolin, G., et al. (2007). Mitochondrial carrier family: repertoire and peculiarities of the cellular slime mould Dictyostelium discoideum. Biochimie 89, 1058–1069. doi: 10.1016/j.biochi.2007.03.004
Schilde, C., and Schaap, P. (2013). The Amoebozoa. Methods Mol. Biol. 983, 1–15. doi: 10.1007/978-1-62703-302-2_1
Shawki, A., Knight, P., Maliken, B., Niespodzany, E., and Mackenzie, B. (2012). H(+)-coupled divalent metal-ion transporter-1: functional properties, physiological role and therapeutics. Curr. Top. Membr. 70, 169–214. doi: 10.1016/B978-0-12-394316-3.00005-3
Shevchuk, O., Batzilla, C., Hagele, S., Kusch, H., Engelmann, S., Hecker, M., et al. (2009). Proteomic analysis of Legionella-containing phagosomes isolated from Dictyostelium. Int. J. Med. Microbiol. 299, 489–508. doi: 10.1016/j.ijmm.2009.03.006
Sillo, A., Bloomfield, G., Balest, A., Balbo, A., Pergolizzi, B., Peracino, B., et al. (2008). Genome-wide transcriptional changes induced by phagocytosis or growth on bacteria in Dictyostelium. BMC Genomics 9:291. doi: 10.1186/1471-2164-9-291
Sillo, A., Matthias, J., Konertz, R., Bozzaro, S., and Eichinger, L. (2011). Salmonella typhimurium is pathogenic for Dictyostelium cells and subverts the starvation response. Cell. Microbiol. 13, 1793–1811. doi: 10.1111/j.1462-5822.2011.01662.x
Soldati, T., and Neyrolles, O. (2012). Mycobacteria and the intraphagosomal environment: take it with a pinch of salt(s)! Traffic 13, 1042–1052. doi: 10.1111/j.1600-0854.2012.01358.x
Solomon, J. M., and Isberg, R. R. (2000). Growth of Legionella pneumophila in Dictyostelium discoideum: a novel system for genetic analysis of host-pathogen interactions. Trends Microbiol. 8, 478–480. doi: 10.1016/S0966-842X(00)01852-7
Solomon, J. M., Leung, G. S., and Isberg, R. R. (2003). Intracellular replication of Mycobacterium marinum within Dictyostelium discoideum: efficient replication in the absence of host coronin. Infect. Immun. 71, 3578–3586. doi: 10.1128/IAI.71.6.3578-3586.2003
Steenbergen, J. N., Nosanchuk, J. D., Malliaris, S. D., and Casadevall, A. (2003). Cryptococcus neoformans virulence is enhanced after growth in the genetically malleable host Dictyostelium discoideum. Infect. Immun. 71, 4862–4872. doi: 10.1128/IAI.71.9.4862-4872.2003
Steinert, M. (2011). Pathogen-host interactions in Dictyostelium, Legionella, Mycobacterium and other pathogens. Semin. Cell Dev. Biol. 22, 70–76. doi: 10.1016/j.semcdb.2010.11.003
Steinert, N., Hentschel, U., and Hacker, J. (2000). Symbiosis and pathogenesis: evolution of the microbe-host interaction. Naturwissenschaften 87, 1–11. doi: 10.1007/s001140050001
Sunaga, N., Monna, M., Shimada, N., Tsukamoto, M., and Kawata, T. (2008). Expression of zinc transporter family genes in Dictyostelium. Int. J. Dev. Biol. 52, 377–381. doi: 10.1387/ijdb.072389ns
Tiaden, A., Spirig, T., Weber, S. S., Bruggemann, H., Bosshard, R., Buchrieser, C., et al. (2007). The Legionella pneumophila response regulator LqsR promotes host cell interactions as an element of the virulence regulatory network controlled by RpoS and LetA. Cell. Microbiol. 9, 2903–2920. doi: 10.1111/j.1462-5822.2007.01005.x
Trinchera, M., and Bozzaro, S. (1996). Dictyostelium cytosolic fucosyltransferase synthesizes H type 1 trisaccharide in vitro. FEBS Lett. 395, 68–72. doi: 10.1016/0014-5793(96)01003-4
Urwyler, S., Nyfeler, Y., Ragaz, C., Lee, H., Mueller, L. N., Aebersold, R., et al. (2009). Proteome analysis of Legionella vacuoles purified by magnetic immunoseparation reveals secretory and endosomal GTPases. Traffic 10, 76–87. doi: 10.1111/j.1600-0854.2008.00851.x
Vidal, S., Tremblay, M., Govoni, G., Gauthier, S., Sebastiani, G., Maio, D., et al. (1995). The Ity/Lsh/Bcg locus: natural resistance to infection with intracellular parasites is abrogated by disruption of the Nramp1 gene. J. Exp. Med. 182, 655–666. doi: 10.1084/jem.182.3.655
Wagner, D., Maser, J., Moric, I., Vogt, S., Kern, W., and Bermudez, L. (2006). Elemental analysis of the Mycobacterium avium phagosome in Balb/C mouse macrophages. Biochem. Biophys. Res. Commun. 344, 1346–1351. doi: 10.1016/j.bbrc.2006.04.048
Wang, J., and Pantopoulos, K. (2011). Regulation of cellular iron metabolism. Biochem. J. 434, 365–381. doi: 10.1042/BJ20101825
Weber, S. S., Ragaz, C., and Hilbi, H. (2009). The inositol polyphosphate 5-phosphatase OCRL1 restricts intracellular growth of Legionella, localizes to the replicative vacuole and binds to the bacterial effector LpnE. Cell. Microbiol. 11, 442–460. doi: 10.1111/j.1462-5822.2008.01266.x
West, C. M., Wang, Z. A., and van der Wel, H. (2010). A cytoplasmic prolyl hydroxylation and glycosylation pathway modifies Skp1 and regulates O2-dependent development in Dictyostelium. Biochim. Biophys. Acta 1800, 160–171. doi: 10.1016/j.bbagen.2009.11.006
Wildschutte, H., Wolfe, D., Tamewitz, A., and Lawrence, J. (2004). Protozoan predation, diversifying selection and evolution of antigenic diversity in Salmonella. Proc. Natl. Acad. Sci. U.S.A. 101, 10655–10649. doi: 10.1073/pnas.0404028101
Keywords: Dictyostelium, Legionella, Mycobacterium, Nramp1, Nramp2, iron homeostasis, iron genes, host-pathogen interactions
Citation: Bozzaro S, Buracco S and Peracino B (2013) Iron metabolism and resistance to infection by invasive bacteria in the social amoeba Dictyostelium discoideum. Front. Cell. Infect. Microbiol. 3:50. doi: 10.3389/fcimb.2013.00050
Received: 11 July 2013; Accepted: 22 August 2013;
Published online: 19 September 2013.
Edited by:
Mathieu F. Cellier, Institut National de la Recherche Scientifique, CanadaReviewed by:
Thierry Soldati, University of Geneva, SwitzerlandMichelle D. Snyder, Towson University, USA
Ricardo Escalante, Consejo Superior de Investigaciones Científicas, Spain
Copyright © 2013 Bozzaro, Buracco and Peracino. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Salvatore Bozzaro, Department of Clinical and Biological Sciences, University of Torino, AOU S. Luigi, Reg. Gonzole 10, 10043 Orbassano (TO), Italy e-mail:c2FsdmF0b3JlLmJvenphcm9AdW5pdG8uaXQ=