- 1Department of Integrative Biology, Oklahoma State University, Stillwater, OK, USA
- 2Department of Microbiology and Molecular Genetics, Oklahoma State University, Stillwater, OK, USA
- 3Department of Veterinary Pathobiology, Oklahoma State University, Stillwater, OK, USA
Chlamydia trachomatis is the leading cause of bacterial sexually transmitted infections (STIs) and preventable blindness. Untreated, asymptomatic infection as well as frequent re-infection are common and may drive pelvic inflammatory disease, ectopic pregnancy, and infertility. In vivo models of chlamydial infection continue to be instrumental in progress toward a vaccine and further elucidating the pathogenesis of this intracellular bacterium, however significant gaps in our understanding remain. Chlamydial host cell exit occurs via two mechanisms, lysis and extrusion, although the latter has yet to be reported in vivo and its biological role is unclear. The objective of this study was to investigate whether chlamydial extrusions are shed in vivo following infection with multiple strains of Chlamydia. We utilized an established C3H/HeJ murine cervicovaginal infection model with C. trachomatis serovars D and L2 and the Chlamydia muridarum strain MoPn to monitor the (i) time course of infection and mode of host cell exit, (ii) mucosal and systemic immune response to infection, and (iii) gross and histopathology following clearance of active infection. The key finding herein is the first identification of chlamydial extrusions shed from host cells in an in vivo model. Extrusions, a recently appreciated mode of host cell exit and potential means of dissemination, had been previously observed solely in vitro. The results of this study demonstrate that chlamydial extrusions exist in vivo and thus warrant further investigation to determine their role in chlamydial pathogenesis.
Introduction
Chlamydia trachomatis is a pathogen of global significance, causing ocular trachoma, and urogenital infections. The species is comprised of at least 15 different serovars that are categorized into two separate biovars, (i) trachoma, which causes ocular disease (serovars A to C) and genital mucosa disease (serovars D to K), and (ii) lymphogranuloma venereum (LGV; serovars L1 to L3; Schachter, 1999). In the United States urogenital C. trachomatis infections are the most common bacterial sexually transmitted infections (STIs) reported by the CDC (Centers for Disease Prevention, 2011). Despite treatment, ongoing complications may arise after infection including pelvic inflammatory disease (PID), endometriosis, tubal scarring, ectopic pregnancy, and potentially cervical cancer (Smith et al., 2001; Brunham and Rey-Ladino, 2005).
Chlamydiae are a group of obligate intracellular bacteria that possess a unique biphasic life cycle consisting of two alternating forms: infectious, non-replicative, extracellular elementary bodies (EBs) and non-infectious, metabolically active, replicative reticulate bodies (RBs; Moulder, 1991). Upon binding of the EB to the host cell, the EB is endocytosed and contained within a membrane bound vacuole, termed an inclusion, and avoids lysosomal fusion (Fields and Hackstadt, 2002). From within the protection of the inclusion, EBs quickly differentiate into metabolically active RBs, which replicate via polarized budding process (Abdelrahman et al., 2016). Upon the host cell swelling in response to the growing numbers of Chlamydia, the Chlamydia exit the cell by one of two mechanisms: lysis or extrusion (Todd and Caldwell, 1985; Scidmore et al., 1996; Hybiske and Stephens, 2007, 2008; Lutter et al., 2013) for subsequent rounds of infection. Extrusion studies to date have been performed in vitro utilizing many cell types and chlamydial strains (Todd and Caldwell, 1985; Hybiske and Stephens, 2007, 2008; Chin et al., 2012; Lutter et al., 2013; Zuck et al., 2016a,b); however, the role of extrusions in chlamydial pathogenesis in vivo has not yet been described.
To investigate the mechanisms and outcomes of C. trachomatis infections, a female mouse urogenital infection model has been extensively studied with Chlamydia muridarum strains (Barron et al., 1981; Swenson et al., 1983; Ramsey et al., 2009) and human C. trachomatis urogenital isolates (Tuffrey et al., 1986). Several attributes of this model closely resemble acute genital tract infections in human females. Typically mice resolve chlamydial infections within 4 weeks and develop subsequent immunity (Morrison and Caldwell, 2002), which can differ between C. trachomatis strains and C. muridarum strains (Lyons et al., 2005; Morrison et al., 2011; De Clercq et al., 2013).
To date, much knowledge has been gained regarding infectivity, pathogenicity, fertility, and potential vaccine candidates for C. muridarum (MoPn) and C. trachomatis infections in the murine model (Perry and Hughes, 1999; Pal et al., 2001; Morrison and Caldwell, 2002; Kari et al., 2011; Morrison et al., 2011; Schautteet et al., 2011; Yu et al., 2012; De Clercq et al., 2013). In the literature there is a wide range of techniques, protocols, end point assays, chlamydial strains used for infection, and different mouse strains potentially confounding the interpretation of data across different studies. Thorough comparative studies across chlamydial strains have recently been published to address these issues, such as the one published by Morrison et al. (2011). However, there are no reports to date that have investigated the presence of chlamydial extrusions shed in vivo. Therefore, the objective of this study was to monitor the course of murine chlamydial infection while investigating the mode by which organisms are shed in vivo following infection across multiple strains of Chlamydia. Here, we present a comparison of the urogenital serovar D-LC, the LGV serovar L2 of C. trachomatis, and MoPn (Nigg) of C. muridarum infection in C3H/HeJ mice, a currently accepted mouse strain for optimal modeling of a robust cervicovaginal infection (Bernstein-Hanley et al., 2006). Overall, the findings are consistent with the expected time course of chlamydial infections, immune response generated, and resultant pathology while providing novel data on the mode of host-cell chlamydial shedding from the murine cervicovaginal tract. We document the first account of chlamydial extrusions in an in vivo model. This novel finding warrants further research in chlamydial pathogenesis.
Methods
Chlamydial Strains and Cell Culture
C. trachomatis serovars D-Late Clearance (D-LC; Sturdevant et al., 2010), L2 (LGV 434), and C. muridarum mouse pneumonitis (MoPn, Nigg) were propagated in HeLa 229 cells and purified by Renografin density gradient centrifugation as previously described (Caldwell et al., 1981). HeLa cells were grown in RPMI 1640 + 10% fetal bovine serum (FBS) at 37°C with 5% CO2 and 1 ug/mL cycloheximide added as needed.
Cervicovaginal Infection of Mice
Six week old female inbred C3H/HeJ mice (innate immune-deficient) were purchased from Jackson Laboratories (Bar Harbor, Maine). All animal work was performed according to The Animal Care and Use Guidelines and approved by the Institution of Animal Care and Use Committee at Oklahoma State University. At 10 and 7 days prior to infection, all mice were subcutaneously injected with 2.5 mg of medroxyprogesterone acetate (Upjohn, Kalamazoo, MI) to synchronize estrus. Mice were divided into four groups (n = 15 per group): sham, C. trachomatis serovar D-LC, C. trachomatis serovar L2, and C. muridarum (MoPn). Mice were infected intravaginally with 1 × 106 EBs in 5 μL sucrose-phosphate-glutamate (SPG). Control mice were sham infected via intravaginal administration of 5 μL of SPG alone. Of the 15 mice within each group, 10 mice were dedicated to enumeration of IFU while the remaining five mice per group were strictly dedicated to the recovery of extrusions shed from the cervicovaginal tract. An additional experiment was conducted using mice (n = 10) treated with medroxyprogesterone acetate and infected with L2, as described above, to further confirm the presence of extrusions shed in vivo.
Quantification of Recoverable IFUs from the Cervicovaginal Tract
At days 3, 7, 14, 21, 28, 35, and 59 days post infection, cervicovaginal tracts were swabbed via 8 rotations to the right and 8 rotations to the left (Puritan Diagnostics, HydraFlock 6″ 15 cm swabs; Guilford, ME) and each swab was added to tubes containing 600 μL SPG and two glass beads on ice, as adopted from previously described studies (Shaw et al., 2001; Cheng et al., 2008; Chen et al., 2014). Swab samples were vortexed vigorously to liberate EBs from the swab followed by serial dilution and inoculation of confluent HeLa cell monolayers in 24 well plates (CellTreat Scientific, MA). Plates were centrifuged at 700 × g for 1 h to promote EB entry. Inoculated HeLa monolayers were incubated in RPMI 1640 + 10% FBS for 24–34 h at 37°C with 5% CO2. Plates were fixed with methanol and stained with anti-MOMP (MOMP antibody recognizes both C. trachomatis and C. muridarum, courtesy of Dr. Harlan Caldwell) followed by anti-mouse DyLight 488 (Jackson Immunoresearch, Westgrove PA). Twenty fields of view were counted using a Leica MI6000B fluorescent microscope. Total recoverable IFUs/mouse were calculated for each sample.
Microscopy of Extrusions Shed from the Cervicovaginal Tract
Collection and Preparation of Extrusions for Live Cell Imaging
At days 10, 17, and 24 post-infection the cervicovaginal tracts were swabbed as described above then placed into 100 μL of RPMI 1640 + 10% FBS on ice. Swab samples were gently swirled to release extrusions; samples were not vortexed due to the fragile nature of extrusions. Samples were centrifuged at 75 × g and supernatant removed. The pellet was resuspended in 100 μL RMPI+10% FBS and stained with Hoechst NucBlue® Live ReadyProbes® (1:200) Reagent and FM 4-64 (Thermo Fisher Scientific; 5 μg/mL). Wet mounts were prepared and visualized under oil using an Olympus IX81 Spinning Disc Confocal microscope; differential interference contrast (DIC) and live cell fluorescent images of extrusions were captured.
Collection and Preparation of Extrusions for Fixed Cell Imaging
To confirm the presence of chlamydial EBs housed inside extrusions an additional murine study was performed using L2-infected mice, as described above. At 17 and 20 days post-infection cervicovaginal tracts were swabbed for extrusions (n = 5), as described above, or lavage (n = 5) performed by gently washing the vaginal tract with 60 μL RPMI 1640 + 10% FBS twice to produce a 120 μL lavage sample per mouse for analysis. Pooled swab and pooled lavage samples were centrifuged at 75 × g to enrich extrusions. The pellets were resupended in 200 μL RPMI 1640 + 10% FBS and stained with aldehyde fixable analog, FM 4-64FX (Thermo Fisher Scientific; 5 μg/mL), for 10 min. Samples were then centrifuged at 75 × g and fixed with 4% paraformaldehyde. Chlamydial EBs were labeled with anti-L2 and anti-rabbit DyLight 488 secondary antibodies (Jackson Immuno Research); nuclei were stained with DAPI (4′,6′-diamidino-2-phenylindole; Thermo Fisher Scientific; 0.5 μg/ml). Next, samples were mixed with ProLong Diamond Antifade Mountant (Thermo Fisher Scientific), mounted onto glass slides and viewed using a Leica DMI6000B microscope.
Enzyme-Linked Immunosorbent Assay (ELISA)
To measure the murine mucosal and systemic immune response to infection, vaginal washes and sera were collected 31 and 35 days post infection, respectively, for ELISA. Vaginal washes were obtained by gently washing the vaginal vault with 60 μL 0.5% BSA-PBS twice and stored at −20°C until the ELISA was performed. Briefly, 96-well polystyrene plates (Immulon 2HB; Thermo, Milford, MA) were coated with 1 μg of formalin fixed EBs (serovar D-LC, serovar L2, and MoPn) per well in 100 μL TBS (50 mM Tris buffer pH 7.5, 0.15 M NaCl) overnight at 4°C. Following adsorption of fixed EBs, wells were washed to remove unbound EBs then blocked with 200 μL 2% BSA in 0.012 M Tris pH 7.4, 0.14 M NaCl, 3.0 mM KCl, 0.05% Tween 20 for 90 min at 37°C. Plates were washed and serial dilutions of vaginal washes and sera were added to appropriately matched wells (e.g., vaginal washes from serovar D-LC infected mice added to plates with serovar D-LC fixed EBs). Plates were incubated for 90 min at 37°C. Chlamydial-specific antibodies were detected in vaginal wash and sera samples using alkaline phosphatase-conjugated anti-mouse IgA and IgG isotype-specific antibodies (Southern Biotech Associates, Birmingham, AL), respectively. P-nitrophenyl phosphate (PNPP) was used as a substrate and resulting absorbance was read at 405 nm (BioTek Synergy, Winooski, VT). Vaginal washes and sera from sham mice were used as negative controls. Antibody titers were considered positive at the highest sample dilution that displayed an absorbance that was ≥3X the absorbance of the concentrated sham samples.
Histopathology
Mouse uteri were excised and immersion fixed in 10% buffered neutral formalin. Following fixation, they were processed entire and embedded en bloc into paraffin followed by sections cut at 4 μm and staining with hematoxylin and eosin (H&E). Sections were examined and scored via light microscopy by an American College of Veterinary Pathologists (ACVP) certified veterinary pathologist with the following numerical designations: 0, normal; 1, minimal change; 2, mild change; 3, moderate change; 4, severe change. Scored parameters (mean ± SD) included an overall impression, periglandular mucinous change, hydrosalpinx, uterine luminal, and stratum compactum inflammation. Additionally, endometrial luminal epithelial height was subjected to morphometric analyses focused on epithelial height as determined by calibrated measurements via Olympus CellSens software coupled to an Olympus DP70 microscope camera. Each reproductive tract was evaluated regionally and morphometrically at six locations (proximal, mid-, and distal locations of the two uterine horns) determined to be uniform and free of tissue bends and curves; epithelial height (mean μm ± SD) was measured from the basement membrane to the most apical cytoplasmic membrane.
Statistics
One way ANOVA (Prism 3.0) was used to analyze the recoverable IFU and immunoglobulin data sets of sham vs. each of the three infected groups (D-LC, L2, MoPn) followed by Tukey post hoc analysis (significance at p < 0.05). One way ANOVA of morphometric measurements of epithelial height and post hoc Student's two-way t-test with equal variances were performed between sham infected (n = 3) and each (n = 3) of the infected groups (D-LC, L2, MoPn) with significance assigned most conservatively at p < 0.016 (0.05/three tests).
Results
Recoverable IFUs during C. trachomatis and C. muridarum Infection
Medroxyprogesterone acetate treated female 6 week old C3H/HeJ mice were cervicovaginally infected with 1 × 106 IFUs of either C. trachomatis D-LC, C. trachomatis L2, or C. muridarum. Infectious burden and duration was monitored via swabbing the vaginal vault (n = 10) and culturing in HeLa cells at weekly intervals through day 35 post infection (and again on day 59 to confirm that clearance of infection was complete). Quantification of recoverable IFUs are shown in Figure 1. Infection by all three strains was self-limiting with peaks of infection occurring on day 14 after which infectious burden decreased steadily. C. trachomatis L2 resulted in infection with the greatest recoverable IFUs (p < 0.001 L2 vs. D-LC; p < 0.001 L2 vs. MoPn) while C. muridarum and C. trachomatis serovar D-LC presented with similar infection burdens throughout the course of infection. At day 59 post infection mice displayed no evidence of infection (data not shown); mice were euthanized by cervical dislocation and reproductive tissues harvested for gross and histological evaluation.
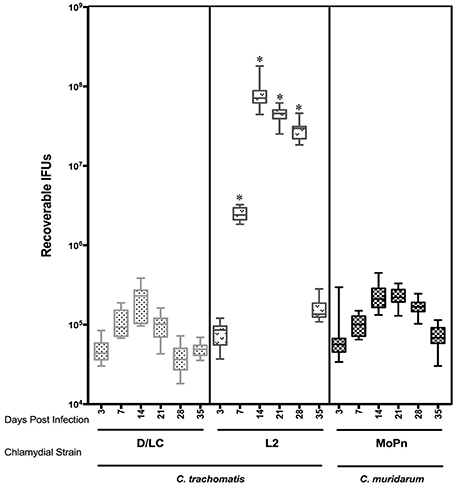
Figure 1. Comparison of recoverable infectious forming units (IFUs) obtained from C3H/HeJ mice between C. trachomatis (serovars D-LC, L2) and C. muridarum (MoPn) infection. Mice were intravaginally infected with 1X 106 EBs of corresponding Chlamydial strain. Recoverable IFUs were obtained by swabbing vaginal tracts and enumerating on HeLa cell monolayers. IFU data are expressed (mean ± SE) for each Chlamydial strain from day 3 to35 post infection.*p < 0.001 (L2 vs. D-LC, L2 vs. MoPn), One way ANOVA.
Extrusions Recovered from the Cervicovaginal Tract
Vaginal swabs (n = 5) obtained 24 days post infection were examined by DIC and fluorescent microscopy. Stains for live cells (Hoescht) and plasma membranes (FM 4-64) were used to visualize cells. Round Chlamydia-filled plasma membrane enclosed vesicles were identified as well as vaginal epithelial cells and cellular debris. Representative images are found in Figure 2A. These Chlamydia-filled membrane surrounded structures devoid of nuclei were consistent in size and shape with previously described extrusions (Hybiske and Stephens, 2007; Lutter et al., 2013; Zuck et al., 2016a). During live microscopy the Chlamydia were vibrating within the extrusion changing extrusion shape from circular to amorphous suggesting the extrusion membranes are highly pliable. The occurrence of Chlamydia extrusions were observed across all strains (D-LC, L2, and MoPn). The overall morphological appearance and size of extrusions were consistent between chlamydial strains. Sham infected mice were devoid of any extrusions. Additional studies were conducted to verify the presence of chlamydial EBs housed inside extrusions via fixed cell imaging. Vaginal swab and lavage samples collected and fixed at 17 and 20 days post L2-infection were analyzed by fluorescent microscopy of the host plasma membrane (FM 4-64FX), chlamydial EBs (anti-L2), and host nuclei (DAPI). As illustrated in Figure 2B, there is positive staining (red) for host plasma membrane surrounding chlamydial EBs (green) within a vesicular structure devoid of host nuclei. Collectively, our microscopy data provide definitive proof of chlamydial extrusions shed from the murine cervicovaginal tract.
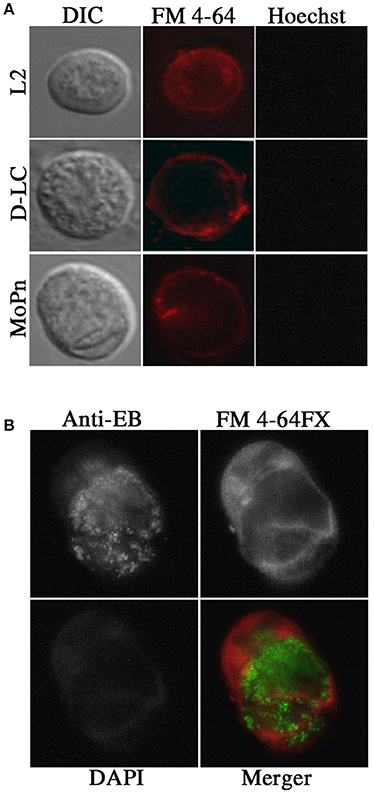
Figure 2. Presence of extrusions in vivo after infection with Chlamydia by live and fixed cell microscopy. (A) Live cell microscopy identified potential extrusions shed from mice infected with each chlamydial strain tested. Olympus Laser Scanning Confocal microscopy was used to obtain DIC, nuclear (Hoechst; blue) and plasma membrane (FM 4–64; red) images on 60X with oil. (B) Fixed cell imaging was used to confirm extrusions shed from the cervicovaginal tracts infected with C. trachomatis L2. Indirect immunoflourescent images of fixed extrusions were obtained using a Leica DMI6000B at 40X magnification. Staining of plasma membrane (FM 4–64 FX; red), L2 EBs (anti-L2; Green), nuclei (DAPI; blue), and a merged image are shown.
Mucosal and Systemic Immune Response to C. trachomatis and C. muridarum
To compare the mucosal immune response across multiple strains, vaginal washes were obtained from mice 31 days post infection with either C. trachomatis serovar D-LC (n = 10), C. trachomatis serovar L2 (n = 10), or C. muridarum MoPn (n = 10). Vaginal IgA antibody titers against strain-specific EBs were measured by ELISA (Figure 3). All mice infected with MoPn produced an anti-Chlamydia mucosal immune response with IgA titers ranging from 64 to 128, while those infected with serovar D-LC displayed, on average, lower titers with substantial variability (p < 0.05 MoPn vs. D-LC). Mice infected with serovar L2 consistently produced negligible IgA titers (Figure 3A) that were significantly lower than titers produced by D-LC and MoPn-infected mice (p < 0.05 L2 vs. D-LC; p < 0.001 L2 vs. MoPn). Sera were collected from sham and infected mice (n = 5 per group) 35 days post infection. Mice infected with MoPn exhibited similar titers for both anti-Chlamydia IgG2a and IgG1 antibodies with little variation between individual mice. In contrast, mice infected with serovar D-LC and L2 generated highly variable titers of IgG2a and IgG1 without statistically significant differences between groups (Figure 3B).
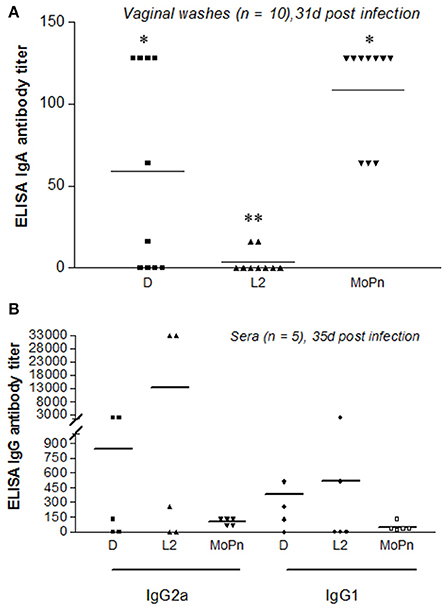
Figure 3. Mucosal and systemic antibody responses following infection with C. trachomatis (serovars D, L2) and C. muridarum (MoPn). (A). Vaginal washes were collected 31 days post infection (n = 10) and assayed for the presence of anti-chlamydial secretory IgA. (B) Sera were collected 35 days post infection (n = 5) and assayed for the presence of anti-chlamydial IgG2a and IgG1. Antibody titers are expressed for individual mice as the highest dilution that produced ≥3-fold the absorbance reading of the control (sham-infected) vaginal wash and sera samples. Bars represent mean antibody titer per group.**p < 0.001 (L2 vs. MoPn), *p < 0.05 (D vs. L2; D vs. MoPn), One way ANOVA.
Histopathological and Morphometric Assessment of Reproductive Tracts Following C. trachomatis and C. muridarum Infection
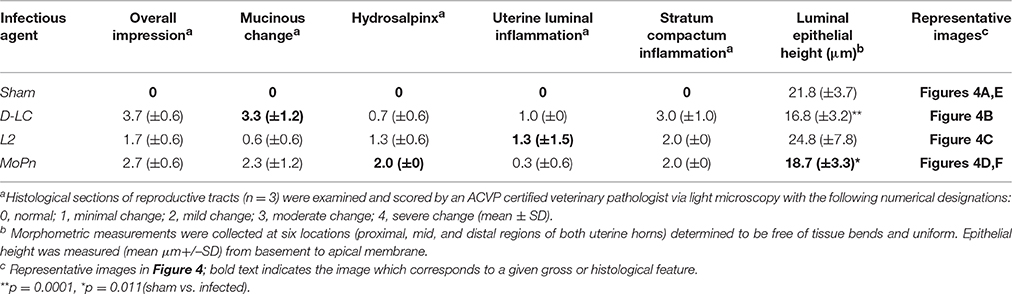
Reproductive tracts (n = 3 per group) were collected 59 days post infection (sham, D-LC, L2, and MoPn) whereupon images of gross morphology were captured followed by formalin fixation for downstream histological evaluation and morphometric measurements. Gross morphology consistently revealed the greatest degree of hydrosalpinx in MoPn-infected mice (Figure 4F). Sections of uterine tissue were examined and the scores assigned by an AVCP certified veterinary pathologist via light microscopy are shown in Table 1. The overall impression scores of uterine tissues were minimal to moderate pathology for all infected mice. Moderate pathology included periglandular mucinous change (Figure 4B), stratum compactum inflammation, and decreased (p = 0.0001) luminal epithelial height (Table 1) in D-LC infected mice relative to sham. Mice infected with MoPn displayed oviduct dilation (Figure 4D) and decreased (p = 0.011) luminal epithelial height (Table 1). The overall impression of tissues and degree of mucinous change in L2 infected mice were minimal; the luminal epithelial height did not differ from the sham infected (Table 1).
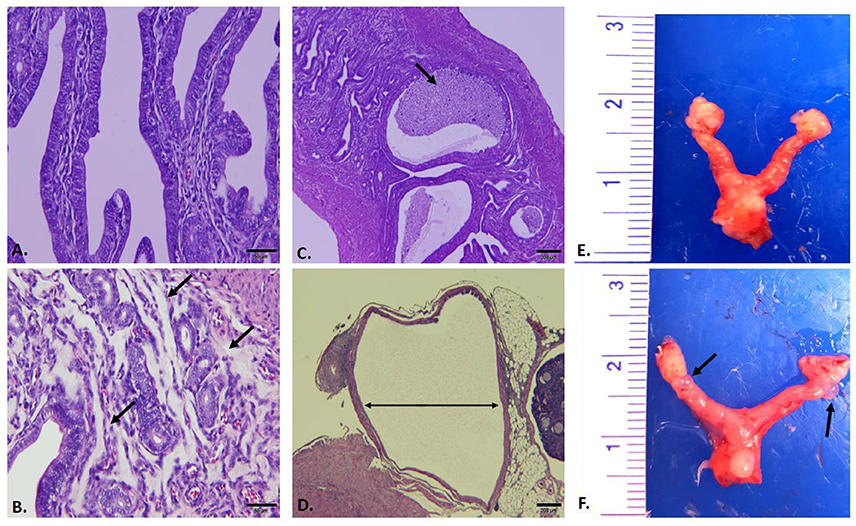
Figure 4. Gross and Histopathological Assessment of Reproductive Tracts Post Infection. Mice (n = 3 per group) were euthanized 59 days post infection and entire reproductive tracts were removed for gross assessment followed by formalin fixation for histological evaluation. Images of H&E stained sections of uterine tissue were captured using an Olympus DP70. (A) Sham infected uterine tissue (20X). (B) C. trachomatis serovar D-LC infected; arrows to areas of mucinous inflammation (20X). (C) C. trachomatis serovar L2 infected; arrows to areas of luminal inflammation (4X). (D) C. muridarum MoPn infected; arrow indicating oviduct dilation (4X). (E) Sham infected whole reproductive tract. (F) C. muridarum MoPn infected whole reproductive tract; arrows to areas displaying hydrosalpinx. See Table 1 for pathological scoring and morphometric measurements of reproductive tracts.
Discussion
The pathological consequences and reproductive sequelae of C. trachomatis are well-known; however, the mechanisms related to pathogenesis resulting in Chlamydia-induced damage are not fully understood. Within this study, three different chlamydial strains (C. trachomatis D-LC, L2, and C. muridarum MoPn) were compared for infectivity, mode of host cell exit, immune response, and sequelae in the cervicovaginal infection model using C3H/HeJ mice. Mice infected with C. trachomatis L2 exhibited the highest infectious burden compared to C. trachomatis D-LC and C. muridarum. This level of increased infectious burden was not surprising as C. trachomatis L2 has consistently exhibited a higher level of recoverable Chlamydia compared to other C. trachomatis serovars (Morrison et al., 2011). However, there is greater variability when comparing cervicovaginal infections by other C. trachomatis strains to C. muridarum. Our data demonstrated an infectious burden for C. trachomatis serovar D and C. muridarum MoPn that is consistent with a previous report (Ito and Lyons, 1999) but not others (Morrison et al., 2011). Morrison et al. (2011) reported less infectious burden with C. trachomatis serovar D than C. muridarum during genital challenge. These differences, however, may be attributed to their use of different strains of C. muridarum (Weiss) and C. trachomatis serovar D (UW-3/CX) compared to our study. The Weiss strain has been shown by others (Ramsey et al., 2009) to be less virulent than the Nigg strain, therefore offering a possible explanation for the lower infectious burden in our MoPn-infected mice.
The significant finding in this study was the identification of chlamydial extrusions in vivo. Swabbing the murine vaginal vault recovered extrusions from infections by all three chlamydial strains (C. trachomatis D-LC, L2, and C. muridarum MoPn), as demonstrated by live cell microscopy. However, live cell staining is limited since antibodies to Chlamydia cannot penetrate live membranes. Therefore, the shedding of chlamydial EBs enclosed within extrusions was validated via fixed cell staining of vaginal lavages collected from L2-infected mice. Consistent with SEM and confocal microscopy data in previous in vitro reports of extrusions (Hybiske and Stephens, 2007, 2008; Lutter et al., 2013; Zuck et al., 2016a,b), the extrusions we collected were filled with Chlamydia, enclosed in the host plasma membrane, and devoid of host nuclei. In vivo host cell exit of all three tested chlamydial strains via the extrusion pathway was hypothesized based on in vitro data demonstrating that all chlamydial species tested to date produce extrusions (Hybiske and Stephens, 2007, 2008; Lutter et al., 2013; Zuck et al., 2016a,b), however documentation was technically challenging in vivo. Extrusions are produced readily and in high numbers by monolayers in cell culture (Lutter et al., 2013; Zuck et al., 2016b); on the other hand, visualization of extrusions produced and shed from the murine cervicovaginal tract was less abundant. The disparity in quantity of extrusions shed in vivo vs. in vitro was expected due to both biological and technical reasons. Extrusions may be produced in high numbers in vivo, but they are likely short lived in the harsh environment of the genital tract (Amjadi et al., 2014; Deruaz and Luster, 2015); pH and mucinous changes, secretory antibodies, and/or inflammatory mediators may decrease the quantity of recoverable extrusions. Likewise, extrusions are physically fragile; mechanical abrasion during swabbing and/or vaginal lavage protocols jeopardize the integrity of the extrusions, as well as downstream procedures for fixation of cells in suspension. Further refinement of protocols to recover extrusions will be an important next step in this area of research. Despite biological and technical challenges of harvesting significant numbers of extrusions in vivo, the fact that they are recovered from the murine vaginal mucosa reveals a previously unappreciated mechanism that may participate in chlamydial pathogenesis in vivo. Recent in vitro studies demonstrate significant advantages for Chlamydia that are packaged into extrusions, including enhanced extracellular survival (Zuck et al., 2016b) and the ability to avoid the immune system by surviving inside macrophages (Zuck et al., 2016a). Confirming these advantages in an animal model would greatly advance our understanding of chlamydial strategies for survival.
Prevention of C. trachomatis STI is needed since infection is often unrecognized, re-infection is common and, whether treated or not, may lead to permanent complications, such as pelvic inflammatory disease, ectopic pregnancy, and tubal infertility. Several decades of effort toward vaccine development has generated extensive knowledge of the immune response to C. trachomatis in mice, yet replicating it to effectively convey protection has proved to be difficult. The overriding challenges stem from the complexity of targeting the bi-phasic life cycle of this obligate intracellular pathogen and stimulating immunity at the site of infection, the genital mucosa. There are several players of the innate and adaptive immune system that participate in clearance of infection, however it is well established that γ-IFN secretion by CD4+ Th1 cells is, (i) the primary requisite to elicit protection against C. muridarum (Perry et al., 1997; Li et al., 2008; Gondek et al., 2009; Darville and Hiltke, 2010), and (ii) a major, but not categorical, player in adaptive immunity against C. trachomatis (Morrison et al., 2011).
Although clearance will indeed occur in the absence of a humoral response (Su et al., 1997) the capacity of systemic and mucosal neutralizing antibodies to reduce bacterial load is relevant in accelerating protective immunity (Morrison and Morrison, 2005; Armitage et al., 2014; Olsen et al., 2015). Bacterial entry and transmission occur at the genital mucosa which, unlike other mucosal sites, lacks lymphoid architecture. Migration of chlamydia-specific IL-17 secreting CD4+ T cells (Th17 cells) to the genital mucosa has recently been recognized as a critical step for the enhancement of mucosal IgA secretion (Cunningham et al., 2008; Hirota et al., 2013; Armitage et al., 2014). We show that mice intravaginally infected with serovar L2 produced negligible IgA in the vaginal tract while exhibiting the highest infectious burden, relative to serovar D-LC and MoPn. This inverse correlation between vaginal IgA and recoverable L2 IFU is consistent with a recent study in minipigs infected with serovar L2 wherein significantly lower vaginal IgA was observed together with a high infectious burden. Furthermore, their subunit antigen immunization with a Th1/Th17 adjuvant approach induced significant vaginal IgA that enhanced clearance of C. trachomatis (Lorenzen et al., 2015). Our data show mice infected with MoPn consistently produced high levels of IgA in the vaginal tract while shedding significantly lower IFU at day 7 through 28 post-infection, relative to L2-infected mice. Serovar D-LC-infected mice also shed fewer organisms from the vaginal tract compared to L2-infected mice, but exhibited substantial variation in the levels of vaginal IgA. This side-by-side analysis captures the significant differences in the mucosal response to chlamydial strains warranting further investigation into the mechanism whereby serovar L2-infected mice differ in their mucosal IgA response, which may simply reflect a delayed response as shown by others (Morrison et al., 2011) or involve a reduction in homing to and/or function of Th17 cells at the genital mucosa.
Our comparison of the systemic immune response consistently revealed similar IgG2a and IgG1 titers between individual mice infected with MoPn. In contrast, mice infected with D-LC and L2 exhibited highly variable titers ranging from non-responders to exceptionally high titers (>11,000 in L2 infected) at 35 days post infection. The mean titer values of IgG2a vs. IgG1 in D-LC and L2 infected mice suggest a favoring of a Th1 (IgG2a) response, but the data were not statistically significant. The contiguous antibody titers (both mucosal and systemic) produced by all MoPn-infected mice may be the result of infection within its natural host as opposed to infection of mice with human serovars.
According to Table 1 all chlamydial strains resulted in some degree of uterine pathology indicating minimal to mild overall changes in L2-infected mice while D-LC and MoPn-infected mice exhibited mild to moderate pathology. The most significant observation was a decrease in luminal epithelial height in both D-LC and MoPn infected mice suggesting atrophy upon excessive inflammatory damage. In contrast, mice infected with L2 displayed similar luminal epithelial height as the sham infected. Considering the high infectious burden in L2-infected mice (relative to D-LC and MoPn) it is interesting that the evaluation of gross and histological morphology indicated minimal overall changes in L2-infected mice while D-LC and MoPn exhibited mild to moderate pathology. L2-infected mice also appeared normal upon assessment for mucinous changes. In contrast, serovar D-LC produced the largest mucinous change, but the least degree of hydrosalpinx. This is overall consistent with other studies comparing C. trachomatis and C. muridarum although the reasons for the differences in local inflammation are not fully understood; the complete vs. partial set of chlamydial cytotoxin genes offer a possible explanation (Belland et al., 2001). MoPn has been shown to contain genes that encode proteins with homology to large cytotoxins and exerts cytopathic effects in HeLa cells, however serovar L2 lacks a substantial portion of these cytotoxin genes and does not produce cytopathic effects in vitro. Serovar D displays a lesser degree of genetic deletion and retains the capacity to drive intermediate cytopathic effects in vitro (Belland et al., 2001). This genetic variation in chlamydial cytotoxins may be a factor in the histopathological differences observed across strains tested herein. Additionally, histopathological differences may arise due to high IgA titers in the vaginal tracts of D-LC and MoPn-infected mice that drive inflammatory damage while L2-infected mice, which produced negligible IgA, experienced less inflammatory damage. This could be another example of the “double-edged sword” concept of cell mediated immune responses that are necessary to clear infection, but cause excessive tissue damage in the process. It has been reported that Th17 cells, which enhance mucosal IgA, stimulate excessive neutrophil-mediated inflammatory damage (Darville and Hiltke, 2010). Therefore, it would be interesting to repeat our study to investigate the cytokine and chemokine profile produced by CD4+ Th1 and Th17 cell populations. This would advance our understanding of strain-specific mechanisms that result in different reproductive sequelae as well as contribute to vaccine efforts toward promoting a Th1 immune response with minimal collateral tissue damage at the reproductive tract.
Future Directions
Our data are the first to identify chlamydial extrusions shed from a murine cervicovaginal infection model. Though, the biological role of chlamydial extrusions during infection is not known, the overall morphology of extrusions appears conserved between in vitro and in vivo infection. An important next step includes the refinement of in vivo techniques to collect extrusions in quantities sufficient for functional assays between chlamydial strains and serovars. Moreover, establishing animal infection models that retain fluorescently labeled Chlamydia strains (e.g., L2-GFP) will provide means of quantification and a more comprehensive analysis of the extrusions produced during in vivo infections.
Author Contributions
EL: project conceptualization, data acquisition, data analysis, data interpretation, revision of manuscript, principle investigator; JS: project conceptualization, data acquisition, data analysis, data interpretation, revision of manuscript, co-principal investigator, AB: data acquisition, data analysis TS: data acquisition, data analysis, revision of manuscript; NA: data acquisition, data analysis
Funding
This study was supported by a grant from the US National Institutes of Health (1R15AI119906-01) to EL and JS.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors would like to thank Dr. Harlan Caldwell for helpful advice and for generously providing the C. trachomatis D-LC strain and anti-MOMP (L2-I-5) antibody.
References
Abdelrahman, Y., Ouellette, S. P., Belland, R. J., and Cox, J. V. (2016). Polarized cell division of Chlamydia trachomatis. PLoS Pathog. 12:e1005822. doi: 10.1371/journal.ppat.1005822
Amjadi, F., Salehi, E., Mehdizadeh, M., and Aflatoonian, R. (2014). Role of the innate immunity in female reproductive tract. Adv. Biomed. Res. 3:1. doi: 10.4103/2277-9175.124626
Armitage, C. W., O'meara, C. P., Harvie, C. G. M, Timms, P., Wijburg, O. L., and Beagley, K. W. (2014). Evaluation of intra- and extra-epithelial secretory IgA in chlamydial infections. Immunology 143, 520–530. doi: 10.1111/imm.12317
Barron, A. L., White, H. J., Rank, R. G., Soloff, B. L., and Moses, E. B. (1981). A new animal model for the study of Chlamydia trachomatis genital infections: infection of mice with the agent of mouse pneumonitis. J. Infect. Dis. 143, 63–66.
Belland, R. J., Scidmore, M. A., Crane, D. D., Hogan, D. M., Whitmire, W., McClarty, G., et al. (2001). Chlamydia trachomatis cytotoxicity associated with complete and partial cytotoxin genes. Proc. Natl. Acad. Sci. U.S.A. 98, 13984–13989. doi: 10.1073/pnas.241377698
Bernstein-Hanley, I., Balsara, Z. R., Ulmer, W., Coers, J., Starnbach, M. N., and Dietrich, W. F. (2006). Genetic analysis of susceptibility to Chlamydia trachomatis in mouse. Genes Immun. 7, 122–129. doi: 10.1038/sj.gene.6364285
Brunham, R. C., and Rey-Ladino, J. (2005). Immunology of Chlamydia infection: implications for a Chlamydia trachomatis vaccine. Nat. Rev. Immunol. 5, 149–161. doi: 10.1038/nri1551
Caldwell, H. D., Kromhout, J., and Schachter, J. (1981). Purification and partial characterization of the major outer membrane protein of Chlamydia trachomatis. Infect. Immun. 31, 1161–1176.
Centers for Disease Prevention (2011). CDC Grand Rounds: Chlamydia prevention: challenges and strategies for reducing disease burden and sequelae. MMWR Morb. Mortal. Wkly. Rep. 60, 370–373.
Chen, W., Mao, K., Hua-Huy, T., Bei, Y., Liu, Z., and Dinh-Xuan, A. T. (2014). Fasudil inhibits prostate cancer-induced angiogenesis in vitro. Oncol. Rep. 32, 2795–2802. doi: 10.3892/or.2014.3491
Cheng, W., Shivshankar, P., Li, Z., Chen, L., Yeh, I. T., and Zhong, G. (2008). Caspase-1 contributes to Chlamydia trachomatis-induced upper urogenital tract inflammatory pathologies without affecting the course of infection. Infect. Immun. 76, 515–522. doi: 10.1128/IAI.01064-07
Chin, E., Kirker, K., Zuck, M., James, G., and Hybiske, K. (2012). Actin recruitment to the Chlamydia inclusion is spatiotemporally regulated by a mechanism that requires host and bacterial factors. PLoS ONE 7:e46949. doi: 10.1371/journal.pone.0046949
Cunningham, K. A., Carey, A. J., Finnie, J. M., Bao, S., Coon, C., Jones, R., et al. (2008). Poly-immunoglobulin receptor-mediated transport of IgA into the male genital tract is important for clearance of Chlamydia muridarum infection. Am J. Reprod. Immunol. 60, 405–414. doi: 10.1111/j.1600-0897.2008.00637.x
Darville, T., and Hiltke, T. J. (2010). Pathogenesis of genital tract disease due to Chlamydia trachomatis. J. Infect. Dis. 201(Suppl. 2), S114–S125. doi: 10.1086/652397
De Clercq, E., Kalmar, I., and Vanrompay, D. (2013). Animal models for studying female genital tract infection with Chlamydia trachomatis. Infect. Immun. 81, 3060–3067. doi: 10.1128/IAI.00357-13
Deruaz, M., and Luster, A. D. (2015). Chemokine-mediated immune responses in the female genital tract mucosa. Immunol. Cell Biol. 93, 347–354. doi: 10.1038/icb.2015.20
Fields, K. A., and Hackstadt, T. (2002). The chlamydial inclusion: escape from the endocytic pathway. Annu. Rev. Cell Dev. Biol. 18, 221–245. doi: 10.1146/annurev.cellbio.18.012502.105845
Gondek, D. C., Roan, N. R., and Starnbach, M. N. (2009). T cell responses in the absence of IFN-gamma exacerbate uterine infection with Chlamydia trachomatis. J. Immunol. 183, 1313–1319. doi: 10.4049/jimmunol.0900295
Hirota, K., Turner, J. E., Villa, M., Duarte, J. H., Demengeot, J., Steinmetz, O. M., et al. (2013). Plasticity of Th17 cells in Peyer's patches is responsible for the induction of T cell-dependent IgA responses. Nat. Immunol. 14, 372–379. doi: 10.1038/ni.2552
Hybiske, K., and Stephens, R. S. (2007). Mechanisms of host cell exit by the intracellular bacterium Chlamydia. Proc. Natl. Acad. Sci. U.S.A. 104, 11430–11435. doi: 10.1073/pnas.0703218104
Hybiske, K., and Stephens, R. S. (2008). Exit strategies of intracellular pathogens. Nat. Rev. Microbiol. 6, 99–110. doi: 10.1038/nrmicro1821
Ito, J. I., and Lyons, J. M. (1999). Role of gamma interferon in controlling murine chlamydial genital tract infection. Infect. Immun. 67, 5518–5521.
Kari, L., Whitmire, W. M., Olivares-Zavaleta, N., Goheen, M. M., Taylor, L. D., Carlson, J. H., et al. (2011). A live-attenuated chlamydial vaccine protects against trachoma in nonhuman primates. J. Exp. Med. 208, 2217–2223. doi: 10.1084/jem.20111266
Li, W., Murthy, A. K., Guentzel, M. N., Seshu, J., Forsthuber, T. G., Zhong, G., et al. (2008). Antigen-specific CD4+ T cells produce sufficient IFN-gamma to mediate robust protective immunity against genital Chlamydia muridarum infection. J. Immunol. 180, 3375–3382. doi: 10.4049/jimmunol.180.5.3375
Lorenzen, E., Follmann, F., Bøje, S., Erneholm, K., Olsen, A. W., Agerholm, J. S., et al. (2015). Intramuscular priming and intranasal boosting induce strong genital immunity through secretory IgA in minipigs infected with Chlamydia trachomatis. Front. Immunol. 6:628. doi: 10.3389/fimmu.2015.00628
Lutter, E. I., Barger, A. C., Nair, V., and Hackstadt, T. (2013). Chlamydia trachomatis inclusion membrane protein CT228 recruits elements of the myosin phosphatase pathway to regulate release mechanisms. Cell Rep. 3, 1921–1931. doi: 10.1016/j.celrep.2013.04.027
Lyons, J. M., Ito, J. I., Peña, A. S., and Morré, S. A. (2005). Differences in growth characteristics and elementary body associated cytotoxicity between Chlamydia trachomatis oculogenital serovars D and H and Chlamydia muridarum. J. Clin. Pathol. 58, 397–401. doi: 10.1136/jcp.2004.021543
Morrison, R. P., and Caldwell, H. D. (2002). Immunity to murine chlamydial genital infection. Infect. Immun. 70, 2741–2751. doi: 10.1128/IAI.70.6.2741-2751.2002
Morrison, S. G., Farris, C. M., Sturdevant, G. L., Whitmire, W. M., and Morrison, R. P. (2011). Murine Chlamydia trachomatis genital infection is unaltered by depletion of CD4+ T cells and diminished adaptive immunity. J. Infect. Dis. 203, 1120–1128. doi: 10.1093/infdis/jiq176
Morrison, S. G., and Morrison, R. P. (2005). A predominant role for antibody in acquired immunity to chlamydial genital tract reinfection. J. Immunol. 175, 7536–7542. doi: 10.4049/jimmunol.175.11.7536
Moulder, J. W. (1991). Interaction of chlamydiae and host cells in vitro. Microbiol. Rev. 55, 143–190.
Olsen, A. W., Follmann, F., Erneholm, K., Rosenkrands, I., and Andersen, P. (2015). Protection against Chlamydia trachomatis infection and upper genital tract pathological changes by vaccine-promoted neutralizing antibodies directed to the VD4 of the major outer membrane protein. J. Infect. Dis. 212, 978–989. doi: 10.1093/infdis/jiv137
Pal, S., Peterson, E. M., and de la Maza, L. M. (2001). Susceptibility of mice to vaginal infection with Chlamydia trachomatis mouse pneumonitis is dependent on the age of the animal. Infect. Immun. 69, 5203–5206. doi: 10.1128/IAI.69.8.5203-5206.2001
Perry, L. L., Feilzer, K., and Caldwell, H. D. (1997). Immunity to Chlamydia trachomatis is mediated by T helper 1 cells through IFN-gamma-dependent and -independent pathways. J. Immunol. 158, 3344–3352.
Perry, L. L., and Hughes, S. (1999). Chlamydial colonization of multiple mucosae following infection by any mucosal route. Infect. Immun. 67, 3686–3689.
Ramsey, K. H., Sigar, I. M., Schripsema, J. H., Denman, C. J., Bowlin, A. K., Myers, G. A., et al. (2009). Strain and virulence diversity in the mouse pathogen Chlamydia muridarum. Infect. Immun. 77, 3284–3293. doi: 10.1128/IAI.00147-09
Schachter, J. (1999). “Infection and disease epidemiology,” in Chlamydia: Intracellular Biology, Pathogenesis, and Immunity, Chapter 6, ed R. S. Stephens (Washington, DC: ASM Press), 139–170.
Schautteet, K., De Clercq, E., and Vanrompay, D. (2011). Chlamydia trachomatis vaccine research through the years. Infect. Dis. Obstet. Gynecol. 2011:963513. doi: 10.1155/2011/963513
Scidmore, M. A., Fischer, E. R., and Hackstadt, T. (1996). Sphingolipids and glycoproteins are differentially trafficked to the Chlamydia trachomatis inclusion. J. Cell Biol. 134, 363–374.
Shaw, J. H., Grund, V. R., Durling, L., and Caldwell, H. D. (2001). Expression of genes encoding Th1 cell-activating cytokines and lymphoid homing chemokines by chlamydia-pulsed dendritic cells correlates with protective immunizing efficacy. Infect. Immun. 69, 4667–4672. doi: 10.1128/IAI.69.7.4667-4672.2001
Smith, J. S., Munoz, N., Franceschi, S., Eluf-Neto, J., Herrero, R., and Peeling, R. W. (2001). Chlamydia trachomatis and cervical squamous cell carcinoma. JAMA 285, 1704; author reply 1705–1706. doi: 10.1001/jama.285.13.1703
Sturdevant, G. L., Kari, L., Gardner, D. J., Olivares-Zavaleta, N., Randall, L. B., Whitmire, W. M., et al. (2010). Frameshift mutations in a single novel virulence factor alter the in vivo pathogenicity of Chlamydia trachomatis for the female murine genital tract. Infect. Immun. 78, 3660–3668. doi: 10.1128/IAI.00386-10
Su, H., Feilzer, K., Caldwell, H. D., and Morrison, R. P. (1997). Chlamydia trachomatis genital tract infection of antibody-deficient gene knockout mice. Infect. Immun. 65, 1993–1999.
Swenson, C. E., Donegan, E., and Schachter, J. (1983). Chlamydia trachomatis-induced salpingitis in mice. J. Infect. Dis. 148, 1101–1107.
Todd, W. J., and Caldwell, H. D. (1985). The interaction of Chlamydia trachomatis with host cells: ultrastructural studies of the mechanism of release of a biovar II strain from HeLa 229 cells. J. Infect. Dis. 151, 1037–1044.
Tuffrey, M., Falder, P., Gale, J., and Taylor-Robinson, D. (1986). Salpingitis in mice induced by human strains of Chlamydia trachomatis. Br. J. Exp. Pathol. 67, 605–616.
Yu, H., Karunakaran, K. P., Jiang, X., Shen, C., Andersen, P., and Brunham, R. C. (2012). Chlamydia muridarum T cell antigens and adjuvants that induce protective immunity in mice. Infect. Immun. 80, 1510–1518. doi: 10.1128/IAI.06338-11
Zuck, M., Ellis, T., Venida, A., and Hybiske, K. (2016a). Extrusions are phagocytosed and promote Chlamydia survival within macrophages. Cell Microbiol. doi: 10.1111/cmi.12683. [Epub ahead of print].
Keywords: chlamydia, extrusion, lymphogranuloma venereum, sexually transmitted infection, urogenital infection, MoPn, Serovar D
Citation: Shaw JH, Behar AR, Snider TA, Allen NA and Lutter EI (2017) Comparison of Murine Cervicovaginal Infection by Chlamydial Strains: Identification of Extrusions Shed In vivo. Front. Cell. Infect. Microbiol. 7:18. doi: 10.3389/fcimb.2017.00018
Received: 28 October 2016; Accepted: 16 January 2017;
Published: 03 February 2017.
Edited by:
Stacey Gilk, Indiana University School of Medicine, USAReviewed by:
Scot P. Ouellette, University of South Dakota, USAJoão Paulo Gomes, National Institutes of Health, Portugal
Copyright © 2017 Shaw, Behar, Snider, Allen and Lutter. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Erika I. Lutter, ZXJpa2EubHV0dGVyQG9rc3RhdGUuZWR1
†These authors have contributed equally to this work.
 Jennifer H. Shaw
Jennifer H. Shaw Amanda R. Behar
Amanda R. Behar Timothy A. Snider
Timothy A. Snider Noah A. Allen
Noah A. Allen Erika I. Lutter
Erika I. Lutter