- 1State Key Laboratory for Biology of Plant Diseases and Insect Pests, Institute of Plant Protection, Chinese Academy of Agricultural Sciences, Beijing, China
- 2Institute of Food Science and Technology, Hebei Agricultural University, Baoding, China
Two-component regulatory systems (TCSs) play pivotal roles in bacteria sensing many different stimuli from environment. Here, we investigated the role of the LytSR TCS in spore formation in Bacillus thuringiensis (Bt) subsp. kurstaki HD73. lacZ gene fusions revealed that the transcription of the downstream genes, lrgAB, encoding two putative membrane-associated proteins, is regulated by LytSR. The sporulation efficiency of a lytSR mutant was significantly lower than that of wild-type HD73. A confocal microscopic analysis demonstrated that LytSR modulates the process of forespore engulfment. Moreover, the transcription of the lytSR operon is regulated by the mother-cell transcription factor SigE, whereas the transcription of the sporulation gene spoIIP was reduced in the lytSR mutant, as demonstrated with a β-galactosidase activity assay. These results suggest that LytSR modulates forespore engulfment by affecting the transcription of the spoIIP gene in Bt.
Introduction
As a type of specialized differentiated cell, spores are used by Bacillus to survive starvation and harsh conditions. Bacillus subtilis is the best-studied spore-forming bacterium. Its endospore is formed by an unusual mechanism involving asymmetric cell division, followed by the engulfment of the cells and the spore morphogenesis (Errington, 2003). The formation of the asymmetric septum is a key event in spore development. Two sigma factors, σF and σE, are instrumental in setting the cell-specific programs of gene expression in motion. Some σE- and σF-dependent genes are also involved in the prespore engulfment process (Errington, 2003). σE is initially produced as an inactive pro-σE precursor and is specifically activated only in the mother cell. The σE regulon includes genes necessary for engulfment (Tan and Ramamurthi, 2014). During engulfment, peptidoglycan degradation machinery composed of SpoIID, SpoIIM, and SpoIIP is initially required for septal-wall thinning and subsequently for the movement of the engulfing membranes (Ohara et al., 2015). The completion of engulfment is a key event governing the later stages of spore development. In the prespore, a third sporulation-specific sigma factor, σG, becomes active at this time, and this sigma factor controls the final stages of development inside the spore. The final mother–cell-specific sigma factor, σK, is regulated at multiple levels and is involved in the formation of the spore coat and in spore maturation (Errington, 2003; Hilbert and Piggot, 2004).
The two-component regulatory system (TCS), which typically consists of a membrane-spanning histidine kinase (HK) sensor and a cytoplasmic response regulator (RR), also plays a critical role in bacterial adaptation, survival, and virulence by sensing changes in the external environment and modulating gene expression in response to a variety of stimuli (Skerker et al., 2005). Studies have found that the transition of B. subtilis from vegetative growth to sporulation is governed by the master transcription factor Spo0A, which is regulated by a complex phosphorelay involving five autophosphorylating histidine kinases (KinA–E), which respond to different types of environmental stress. Spo0A is not a simple TCS containing a kinase and a regulator. Phosphorylated Spo0A is an essential positive regulator of the initiation of sporulation (Burbulys et al., 1991; Jiang et al., 2000; Fujita and Losick, 2003). However, it is not known whether other TCSs are involved in the subsequent spore formation stage, which consists of asymmetric cell division and engulfment in Bacillus.
The Bacillus cereus group of closely related Gram-positive, spore-forming bacteria includes B. cereus, a common cause of human food poisoning, B. thuringiensis (Bt), an insect pathogen, and B. anthracis, the etiological agent of anthrax in mammals (Schnepf et al., 1998; Stenfors Arnesen et al., 2008). The general functions of some TCSs in B. cereus strains have been studied. For example, PP2C-type phosphatase RsbY receives its input from the multi sensor hybrid kinase RsbK, and RsbKY has been shown to regulate the activity of the alternative sigma factor B (van Schaik et al., 2005; de Been et al., 2010). SpsRK is active in response to glucose-6 phosphate and regulates the activity of the spsABC operon, which is involved in sugar phosphate transport (Song et al., 2012). In B. anthracis, LytSR regulates murein hydrolase activity, whereas the lrgAB genes, which are regulated by LytSR, affect stationary-phase survival and sporulation efficiency (Chandramohan et al., 2009). The parental strain has a sporulation efficiency of 88%, whereas the sporulation efficiency of the lrgAB mutant is only 5%, suggesting that the lrgAB gene products have a dramatic impact on sporulation in B. anthracis (Chandramohan et al., 2009). However, how LytSR affects sporulation remains unclear.
The functions of the LytSR TCS in Bt were investigated in this study using Bt subsp. kurstaki HD73. Our results show that the downstream genes, lrgAB, are regulated by LytSR, which is under the control of the mother cell transcription sigma factor SigE. LytSR modulates the subsequent forespore engulfment process and regulates the expression of the sporulation gene spoIIP.
Materials and Methods
Bacterial Strains, Media, and DNA Manipulation
The bacterial strains and plasmids used in this study are listed in Table 1. Bt strain HD73 (accession no. CP004069) was used in this study (Liu et al., 2013). The Bt strains were transformed by electroporation, as previously described (Lereclus et al., 1989). Escherichia coli and the Bt strains were cultured in Luria-Bertani (LB) medium or Schaeffer's sporulation medium (SSM, 8 g of nutrition broth, 0.12% MgSO4 [m/v], 0.1% KCl [m/v], 0.01 M NaOH, 0.1 M MnCl2, 0.01 M Ca(NO3)2, and 0.01 M FeSO4 in 1 L of H2O; Schaeffer et al., 1965) with shaking (220 rpm) at 37 and 30°C, respectively. The antibiotic concentrations used for bacterial selection were 100 μg/ml kanamycin and 10 μg/ml erythromycin for Bt and 100 μg/ml ampicillin for E. coli. DNA manipulation as previously described (Peng et al., 2014). Oligonucleotide primers were listed in Table 2.
Construction of lytSR and lrgAB Mutants
DNA fragments corresponding to the downstream and upstream regions of the lytSR genes (HD73_5856 and HD73_5855) were amplified by PCR using chromosomal DNA from Bt HD73 as the template and the lytSR-1F/lytSR-1R and lytSR-2F/lytSR-2R primer pairs, respectively. The corresponding DNA fragments were fused with overlapping PCR using primers lytSR-1F and lytSR-2R, and the PCR product was digested with BamHI and EcoRI. The fragments were purified and ligated with the temperature-sensitive suicide plasmid pMAD (Arnaud et al., 2004) digested with the same enzymes, to yield the recombinant plasmid pMAD-ΔlytSR, which was used to transformed into host strains with electroporation. The confirmed transformants were incubated at 39–41°C. Colonies lacking erythromycin resistance were selected and one mutant strain, HDΔlytSR, was verified with PCR.
The upstream (562-bp) and downstream (561-bp) fragments of lrgAB (HD73_5854 and HD73_5853) were PCR amplified with the primer pairs lrgAB-1F/lrgAB-1R and lrgAB-2F/lrgAB-2R, respectively, and using Bt HD73 genomic DNA as the template. The kanamycin (Kan)-resistance gene (1,473 bp) was amplified using primers Kan-R and Kan-F. The deletion-insertion mutant cassette was amplified with overlapping PCR using the upstream and downstream fragments and the Kan-resistance gene as the templates, with primers lrgAB-1F and lrgAB-2R. The lrgAB deletion-insertion mutant cassette was inserted into the BamHI and EcoRI restriction sites of the pMAD plasmid to generate the recombinant plasmid pMAD-ΔlrgAB, which was then used to transform Bt HD73 cells with electroporation. Transformants were grown at 30°C in LB plate containing erythromycin and kanamycin, and then transferred to liquid LB containing kanamycin at 39°C. The cells were then plated on LB agar plates. Colonies with kanamycin resistance but lacking erythromycin resistance were selected, and one mutant strain, HDΔlrgAB, was verified with PCR.
Genetic Complementation of the lrgAB and lytSR Deletion Mutants
The oligonucleotide primer pairs lrgABhf-F/lrgABhf-R and lytSRhf-F/lytSRhf-R were used to amplify the lrgAB gene with its own promoter PlrgAB, and the lytSR gene with its promoter PlytSR. The resultant fragments were digested with PstI/BamHI and SalI/EcoRI, respectively, and then integrated into the shuttle vector pHT315 (Arantes and Lereclus, 1991) to generate pHTlrgAB and pHTlytSR, respectively. The genetically complemented mutant strains ΔlrgAB(lrgAB) and ΔlytSR(lytSR) were generated by introducing pHTlrgAB and pHTlytSR into HDΔlrgAB and -HDΔlytSR, respectively.
Construction of spoIID, spoIIM, and spoIIP Mutants
spoIID (HD73_5692), spoIIM (HD73_4392), and spoIIP (HD73_2232) mutants were constructed similar to lrgAB as described above, but using the primer pairs spoIID-1F/spoIID-1R, spoIID-2F/spoIID-2R, kanD-F/kanD-R, spoIIM-1F/spoIIM-1R, spoIIM-2F/spoIIM-2R, kanM-F/kanM-R, spoIIP-1F/spoIIP-1R, spoIIP-2F/spoIIP-2R, and kanP-F/kanP-R, respectively. The recombinant plasmids pMAD-ΔspoIID, pMAD-ΔspoIIM, and pMAD-ΔspoIIP were electroporated into Bt HD73 cells. Colonies with kanamycin resistance but lacking erythromycin resistance were selected, and mutant strains, HDΔspoIID, HDΔspoIIM, and HDΔspoIIP, were verified with PCR.
Growth Curve Assays
Overnight cultures of each strain grown in LB medium were used as starters for growth curve analyses. The exponential growth phase cells were washed in phosphate-buffered saline and then inoculated into SSM or M9 medium supplemented with tryptophan (50 μg/ml) and pyruvate (6 g/l) to an optical density at 600 nm (OD600) of 0.1. The cultures were incubated at 30°C with shaking at 220 rpm, and growth was monitored by measuring the absorbance at 600 nm at different timepoints. Values represent the means of at least three independent replicates. Error bars represent standard deviations.
Determination of Sporulation Efficiency
The HD73, HDΔlrgAB, ΔlrgAB(lrgAB), HDΔlytSR, and ΔlytSR(lytSR) strains were grown in SSM to T28 (T0 is the end of the exponential phase, and Tn is n hours after T0) at 30°C with vigorous shaking. The number of viable cells was counted as the total colony-forming units (CFU) on the LB plates. The number of spores was determined as the number of heat-resistant (65°C for 30 min) CFU on the LB plates. Sporulation efficiency was defined as the ratio of the number of spores to the number of viable cells, multiplied by 100. Values represent the means of at least three independent replicates. The data were analyzed with SPSS (version 19.0) using a t-test. Error bars represent standard deviations. P-values are indicated in the figure legend.
Laser Scanning Confocal Microscopy
The vital membrane dye FM4-64 (Molecular Probes, Inc., Eugene, OR, USA) was dissolved in dimethyl sulfoxide to a final concentration of 100 μM. The cells were stained with FM4-64 (100 μM) for 1 min on ice (Yang J. et al., 2013). To assess engulfment, 0.5 ml of cells cultured to T12 were pelleted and resuspended in 0.1 ml of H2O. An aliquot (2 μl) of this cell suspension was placed on a slide and stained with FM4-64 (100 μM) and MitoTracker Green FM (MTG, 100 nM; from Molecular Probes) for 1 min, and then scanned (476–490 nm excitation and 510–667 nm emission) with a confocal laser scanning microscope (Leica TCS SL; Leica Microsystems, Wetzlar, Germany). Each strain was scanned independently at least three times and each scan was then viewed in at least five fields. The rate of incomplete engulfment was defined as the ratio of the number of incompletely engulfed cells (stained with FM4-64 in the mother cell) to the total number of cells. The values given are the means of at least three independent replicates.
Construction of Promoter Fusions with lacZ
To assess the transcriptional activity of PlrgAB and PlytSR promoters, putative promoter fragments (633 and 845 bp, respectively) were cloned from Bt HD73 genomic DNA using the primer pairs PlrgAB-F/PlrgAB-R and PlytSR-F/PlytSR-R, respectively. The PstI/BamHI fragments of PlrgAB and PlytSR were separately integrated into vector pHT304-18Z, which is the Bt-E. coli shuttle harboring a promoterless lacZ gene (Agaisse and Lereclus, 1994) to generate plasmids pHT304PlrgAB and pHT304PlytSR, respectively. The former was introduced into Bt strain HD73 and the HDΔlytSR mutant, whereas the latter was introduced into Bt strain HD73 and the HDΔsigE mutant (Du et al., 2012). Resultant HD(PlrgAB), ΔlytSR(PlrgAB), HD(PlytSR), and ΔsigE(PlytSR) strains were selected with erythromycin and verified with PCR.
The constructions of PspoIID, PspoIIM, and PspoIIP (650, 437, and 668 bp, respectively) with lacZ fusions are similar to PlrgAB as described above, but using the primer pairs PspoIID-F/PspoIID-R, PspoIIM-F/PspoIIM-R, and PspoIIP-F/PspoIIP-R, respectively. The recombinant plasmids pHT304PspoIID, pHT304PspoIIM, and pHT304PspoIIP were introduced into Bt strain HD73 and the HDΔlytSR mutant. The resultant strains ΔlytSR(PspoIID), ΔlytSR(PspoIIM), and ΔlytSR(PspoIIP) were selected with erythromycin and verified with PCR.
β-Galactosidase Activity Assay
Bt strains carrying lacZ transcriptional fusions were cultured in liquid SSM and 2-ml samples were collected at 1-h intervals. The cells were pelleted and resuspended in 0.5 ml of Z buffer (Peng et al., 2014) at 4°C, then lysed with a Mini-Beadbeater cell disrupter (BioSpec, Bartlesville, OK, USA) and centrifuged at 10,000 × g for 7 min at 4°C. β-Galactosidase activity was determined as previously described (Perchat et al., 2011). The reported values are the means of at least three independent assays. The data were analyzed with SPSS (version 19.0) using a t-test. Error bars represent standard deviations.
Results
PlrgAB Promoter Transcription is Regulated by LytSR
lrgAB is located downstream from the lytSR genes in Bt HD73 (Figure 1A). The Bt lytS (HD73_5856, sensor histidine kinase), lytR (HD73_5855, response regulator), lrgA (HD73_5854, holin-like protein), and lrgB (HD73_5853, holin-like protein) genes encode proteins that share 50, 44, 44, and 54% amino acid sequence identity, respectively, with homologs in Staphylococcus aureus (Patel and Golemi-Kotra, 2015), and 66, 65, 62, and 78% amino acid sequence identity with homologs in B. subtilis (van den Esker et al., 2017). Alignments of these proteins from Bt, S. aureus, and B. subtilis are shown in Supplementary Figure 1. The two-component system LytSR/LytST contained the conserved His_kinase domain and the response regulator receiver domain in Bt, S. aureus, and B. subtilis (Supplementary Figure 1).
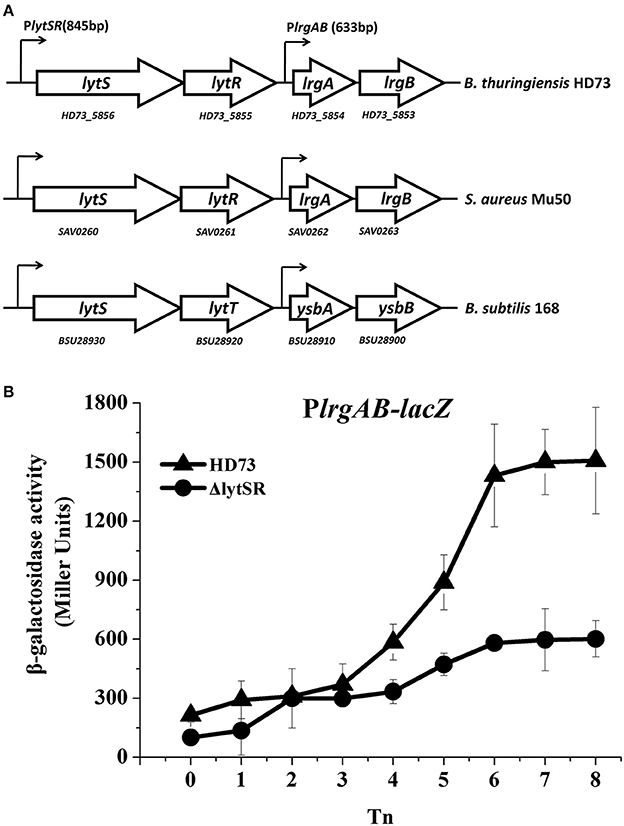
Figure 1. PlrgAB transcription in wild-type Bt HD73 and the lytSR mutant. (A) Gene organization at the lytSR–lrgAB locus in Bt HD73, S. aureus and B. subtilis. White arrows represent open reading frames (ORFs); small arrows denote the lengths of promoters upstream from the lytS and lrgA genes in Bt. (B) β-galactosidase activity from the lrgAB promoter (PlrgAB) in HD73 (▴) and lytSR mutant (•) grown in SSM. T0 is the end of the exponential phase; Tn is n hours after T0. Values represent the means of at least three independent replicates; error bars represent standard deviations.
To investigate the transcription from and regulation of the PlrgAB promoter in Bt, Bt strain HD73, and the lytSR mutant HDΔlytSR were transformed with a PlrgAB-lacZ fusion construct. The results of the β-galactosidase assay showed that the transcriptional activity of PlrgAB increased from T4 to T8 in the HD73 strain in SSM, whereas it did not increase dramatically in the HDΔlytSR mutant (Figure 1B), suggesting that the transcription of the lrgAB genes is positively regulated by LytSR during the late sporulation process.
LytSR Modulates Bt Forespore Engulfment
Previous studies have shown that LytSR/LytST is involved in pyruvate utilization (Zhu et al., 2010; van den Esker et al., 2017). We also compared the growth of the lytSR mutant with that of wild-type strain HD73 in the presence of pyruvate. Results showed that ΔlytSR was unable to grow in M9 medium supplemented with pyruvate, whereas the wild-type reached an OD600 of 0.9 after 20 h of incubation (Figure 2A), suggesting that LytSR is involved in pyruvate utilization in Bt. However, no differences in the growth curves of ΔlytSR and the wild-type were observed in SSM (Figure 2B). Thus, in order to eliminate the effects of growth medium, we selected SSM for further analyses of the sporulation efficiency.
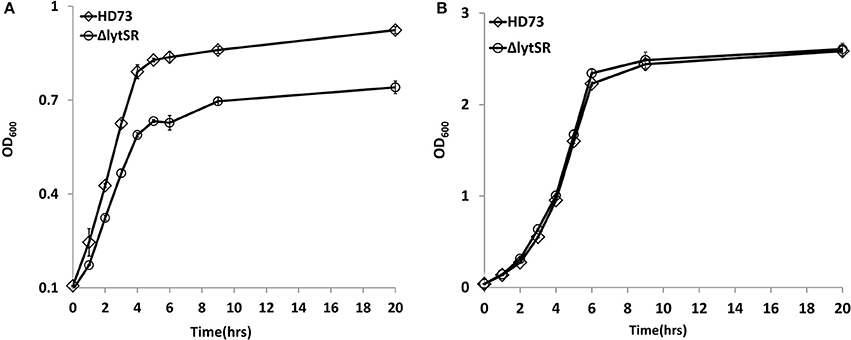
Figure 2. Growth curves assay. Wild-type HD73 (♢) and lytSR mutant cells (◦) were grown in M9 supplemented with pyruvate (A), and SSM (B). Values represent the means of at least three independent replicates; Error bars represent standard deviations.
Because the lrgAB genes have a dramatic impact on sporulation in B. anthracis (Chandramohan et al., 2009), we predicted that the lytSR or lrgAB mutation would affect the ability of the Bt cells to undergo sporulation. Therefore, the abilities of the lrgAB and lytSR mutants to sporulate were assessed. The wild-type strain HD73 had a sporulation efficiency of 85 ± 4% after growth to T28 in SSM (Figure 3). The sporulation efficiency was not significantly different between HD73 and either HDΔlrgAB (72 ± 11%) or ΔlrgAB(lrgAB) (76 ± 9%), whereas it was significantly reduced in both HDΔlytSR (47 ± 3%, P ≤ 0.001) and, the genetically complemented strain ΔlytSR(lytSR) (54 ±4%, P ≤ 0.01). Based on the P-values (P ≤ 0.05, Figure 3) between ΔlytSR and ΔlytSR(lytSR), ΔlytSR(lytSR) showed a partly restored sporulation function. These results indicate that lytSR affects spore formation and the regulation of the genes involved in sporulation.
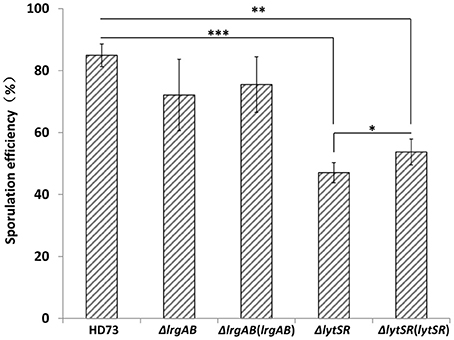
Figure 3. Analysis of sporulation efficiency. Sporulation efficiencies of wild-type HD73, ΔlytSR, ΔlrgAB, ΔlrgAB(lrgAB), and ΔlytSR(lytSR) were compared. Sporulation efficiency was defined as the ratio of the number of spores to the number of viable cells, multiplied by 100. Values represent the means of at least three independent replicates. The data were analyzed with SPSS (version 19.0) using a t-test. Error bars represent standard deviations. *P ≤ 0.05; **P ≤ 0.01; ***P ≤ 0.001.
To determine the effect of LytSR on sporulation in Bt HD73, the cell membranes of Bt HD73 and its mutants were stained with the vital dye FM4-64, which labels the plasma membranes of living cells, and the process of spore formation was visualized with confocal microscopy. In cells grown to T3 in SSM, the polar septum was curved in the wild-type and mutant cells, whereas some cells of HDΔlytSR had an incomplete septum at the distal pole (Figure 4). At T12, the process of engulfment was completed in the forespores of the wild-type (Figure 5, arrow 1) and HDΔlrgAB cells. In these cases, the spores were not labeled with FM4-64, but were stained with MTG, and only the outer membranous outline of the living cells could be observed. In the mutant HDΔlytSR, a proportion of the cells had completed the process of engulfment, but 52 ± 3% cells were arrested in forespore engulfment (Figure 5, arrow 2), and a bipolar septum phenotype was observed.
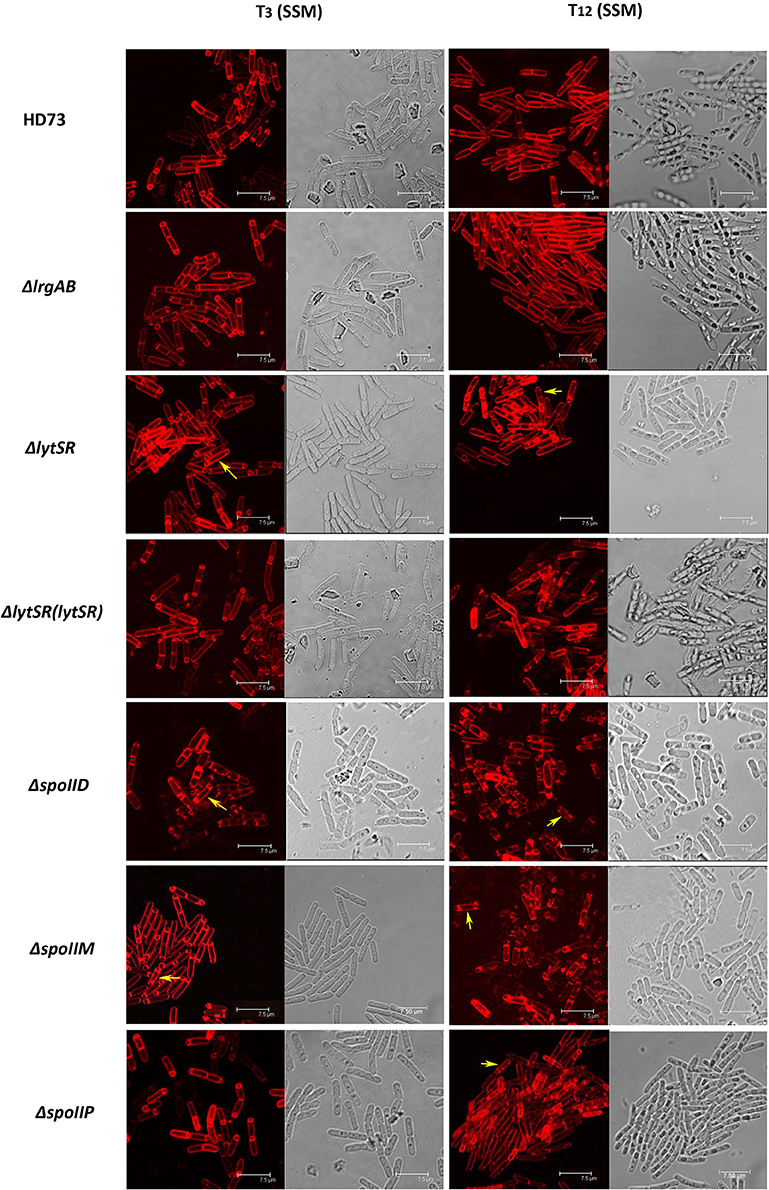
Figure 4. Sporulation process in Bt HD73 in SSM. Laser scanning confocal micrographs of Bt wild-type HD73 cells and ΔlrgAB, ΔlytSR, ΔlytSR(lytSR), ΔspoIID, ΔspoIIM, and ΔspoIIP cells grown in SSM to T3 and T12 (30°C). Cell membrane is visible as red fluorescence. Yellow arrow indicates a bipolar septum. Bar, 7.5 μm.
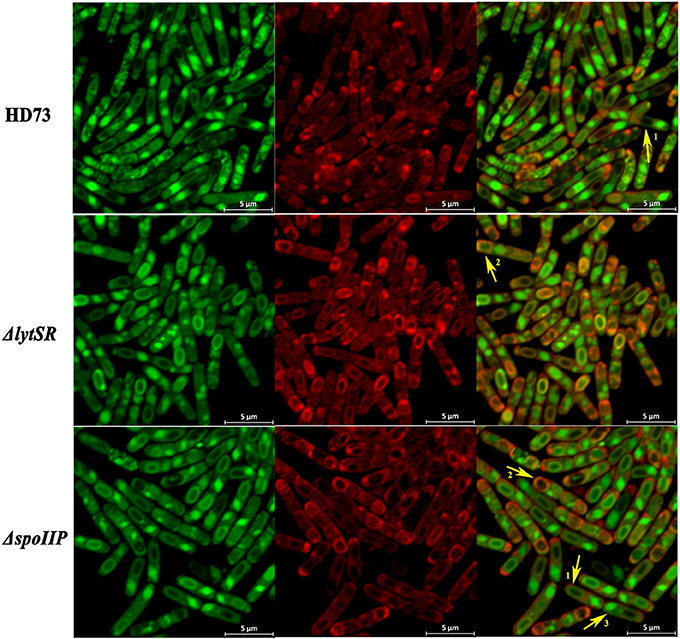
Figure 5. Schematic representation of the membrane fusion assay. Laser scanning confocal micrographs of Bt wild-type HD73, ΔlytSR, and ΔspoIIP cells grown in SSM to T12 (30°C). Red lines represent membranes stained with FM4-64 and MitoTracker Green FM (MTG), and green lines indicate membranes stained with MTG only. Arrow 1 points to cells that have completed the process of engulfment; only the mother–cell membranes are stained with FM4-64, but MTG stained both the forespore and mother–cell membranes. Arrow 2 points to cells that have undergone incomplete engulfment, and the membrane fusion is stained with FM4-64 and MTG. Arrow 3 points to the crystal protein stained with MTG only.
lytSR Transcription is Controlled by SigE
HDΔlytSR cells were unable to initiate engulfment or form bipolar septa. The mother–cell-specific sigma factor SigE plays a critical role in the formation of an asymmetric septum and in forespore engulfment (Errington, 2003). Therefore, we predicted that SigE would also affect the transcription of lytSR. The results of the β-galactosidase assay indicate that the transcriptional activity of PlytSR increased rapidly from T4 to T10 in wild-type HD73, whereas it increased much more slowly in the HDΔsigE mutant grown in SSM (Figure 6), suggesting that the transcription of lytSR is controlled by the mother-cell-specific sigma factor SigE.
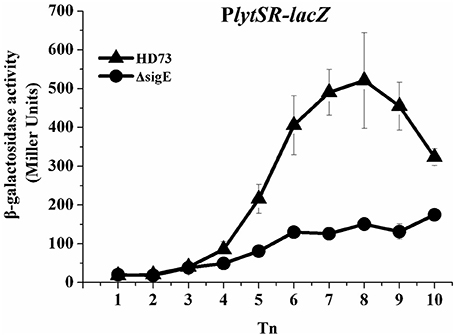
Figure 6. Transcription of PlytSR promoter in Bt. Wild-type HD73 (▴) and sigE mutant cells (•) were grown in SSM. T0 is the end of the exponential phase, and Tn is n hours after T0. Values represent the means of at least three independent replicates; error bars represent standard deviations.
LytSR Affects spoIIP Expression
In B. subtilis, the sporulation genes spoIID, spoIIM, and spoIIP are controlled by SigE (Eichenberger et al., 2001) and may also be involved in suppressing septum formation at the distal pole of the sporangium (Chastanet and Losick, 2007). To determine whether LytSR affects the process of spore engulfment by regulating the expression of spoIID, spoIIM, and spoIIP, the promoters of these genes were fused to lacZ and the β-galactosidase activity was assessed in wild-type HD73 cells, lytSR and sigE mutants. The results showed that the transcriptional activities of spoIID, spoIIM, and spoIIP were sharply reduced or abolished in the sigE mutant grown in SSM (Figure 7), suggesting that the transcription of spoIID, spoIIM, and spoIIP is directly controlled by SigE in Bt. The transcription of PspoIID and PspoIIM transcription did not differ between the wild-type and mutants grown in SSM (Figures 7A,B). However, PspoIIP activity was dramatically reduced in the lytSR mutant grown in SSM (Figure 7C). These results suggest that the transcription of spoIIP is affected by LytSR.
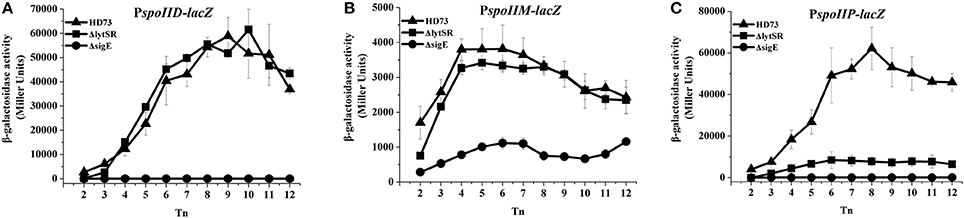
Figure 7. Transcription of PspoIID, PspoIIM, and PspoIIP promoters in Bt. Transcription of PspoIID (A), PspoIIM (B), and PspoIIP (C) in wild-type HD73 (▴), lytSR mutant (■), and sigE mutant (•) cells grown in SSM. T0 is the end of the exponential phase, and Tn is n hours after T0. Values represent the means of at least three independent replicates; error bars represent standard deviations.
LytSR Mainly Modulates Bt Forespore Engulfment by Regulating spoIIP Expression
To determine whether LytSR modulates Bt forespore engulfment by regulating spoIIP expression, we observed the phenotypes of the spoIID, spoIIM, and spoIIP mutants in SSM. In cells grown to T3 or T12 in SSM, the polar septum was curved or had completed the process of engulfment in the wild-type, whereas some HDΔspoIIM, and HDΔspoIID cells displayed an incomplete septum at the distal pole (Figure 4). At T12, the phenotype of HDΔspoIIP was similar to that of HDΔlytSR, and the only difference was that more HDΔspoIIP cells (68 ± 5%) than HDΔlytSR cells (48 ± 3%) had completed the process of engulfment (Figure 5). In contrast, almost all the HDΔspoIID and HDΔspoIIM cells arrested in forespore engulfment, and bipolar septa were also observed (Figure 4), so this phenotype is similar to that of the spoIID and spoIIM mutants of B. subtilis (Pogliano et al., 1999). The β-galactosidase activity assay also revealed that PspoIIP was dramatically reduced in the lytSR mutant grown in SSM (Figure 7). All these results indicate that LytSR modulates Bt forespore engulfment, mainly by affecting spoIIP expression.
Discussion
The sporulation efficiency assay and a confocal microscopic analysis showed that spore formation was unaffected in the Bt lrgAB mutant. This differs from the dramatic impact of this mutation on sporulation efficiency observed in B. anthracis (Chandramohan et al., 2009), although orthologues of the lrgAB locus of Bt HD73 are conserved in the genomes of the B. cereus group (Supplementary Figure 2). These genes share high sequence similarity and a similar organization with those of the lrgAB locus. However, in the Bt lytSR mutant, sporulation efficiency was markedly reduced and spore engulfment was lower than wild-type, and a bipolar septum was observed in some cells grown in SSM. These results indicate that LytSR does not modulate the process of spore formation by regulating of lrgAB, but probably by controlling the expression of other genes.
LytSR and LrgAB are widely conserved in both the B. cereus group and amongst other bacterial species (Supplementary Figure 2). In S. aureus, the LytSR are involved in the regulation of bacterial programmed cell death, biofilm formation, and adaptation to cationic antimicrobial peptides (Brunskill and Bayles, 1996; Rice et al., 2005; Sharma-Kuinkel et al., 2009; Yang S. J. et al., 2013; Lehman et al., 2015), while in Staphylococcus epidermidis, they play a role in regulating extracellular murein hydrolase activity, bacterial cell death, and pyruvate utilization (Zhu et al., 2010). In B. subtilis, the lytSR and lrgA homologs lytST and ysbA are not involved in programmed cell death, but are essential for pyruvate transport or utilization (van den Esker et al., 2017). We also found that mutation of lytSR has an effect on pyruvate utilization in M9 medium in Bt (Figure 2A). However, no differences in the growth curves of ΔlytSR and the wild-type were observed in SSM (Figure 2B). We further demonstrated that the LytSR are involved in the process of spore engulfment in Bt in SSM. These results indicate that LytSR does not modulate the process of spore formation by affecting the pyruvate utilization.
A high proportion (61%) of B. subtilis sigE mutant cells had complete septa near both the poles and failed to undergo engulfment. SigE direct controls the transcription of the sporulation genes spoIID, spoIIM, and spoIIP (Eichenberger et al., 2001). Single mutants of these genes prevent engulfment as they are defective in the dissolution of the peptidoglycan layer between the two membranes of the polar septum. Instead, the septal membrane bulges through the incompletely degraded cell wall layer. In double mutants, the bulge is less prominent, and only in the absence of all three proteins does septum formation occur at both poles at a frequency similar to that observed in the sigE mutant (Eichenberger et al., 2001; Meyer et al., 2010; Tan and Ramamurthi, 2014). The transcription of spoIID, spoIIM, and spoIIP is controlled by SigE in Bt (Figure 7) as in B. subtilis, and the transcriptional activity of spoIIP was sharply reduced in the lytSR mutant compared with that in the wild-type strain grown in SSM. However, the transcriptional activities of spoIIM and spoIID in lytSR mutant did not differ from those in the wild-type strain. This observation suggests that the effect of lytSR on spoIIP expression does not result from the direct activity of LytR on the transcription of spoIIP. The effect of the lytSR mutation on spoIIP expression might be attributable to the low availability of active SigE in the mother–cell compartment of the mutant strain. The transcription of spoIIP requires SigE. However, the amount of SigE required for the full expression of spoIIM, spoIID, and spoIIP might differ, as has been demonstrated for the genes of the Spo0A regulon, which are distributed in two classes: those that are regulated at a low dose of Spo0A-P and those that require a high dose to be activated or repressed (Fujita et al., 2005). In a similar way, spoIIP transcription might require larger amounts of SigE than the transcription of spoIIM and spoIID. Therefore, the SigE defect in the lytSR mutant would have a more dramatic effect on spoIIP expression than on spoIIM or spoIID expression.
The transcriptional analysis of lytSR in the sigE mutant and of spoIID, spoIIM, and spoIIP in the lytSR mutant revealed that the lytSR operon is controlled by SigE and that the efficacy of spoIIP transcription depends, directly or indirectly, on lytSR. We have demonstrated that LytSR affects spore formation by preventing the correct engulfment of the forespore. However, we did not determine whether this effect is responsible for the defect in spoIIP expression or, reciprocally, if it is caused by weak spoIIP expression. In B. subtilis, SpoIIP is targeted to the septal membrane by SpoIIM, where it interacts with SpoIID, which also localizes to the membrane via its interaction with SpoIIP. SpoIIP and SpoIID have complementary enzymatic activities, which are similar to those of LytB and LytC (CwlB), respectively, the major vegetative autolysins involved in peptidoglycan degradation (Shida et al., 2001; Chastanet and Losick, 2007; Gutierrez et al., 2010). Therefore, we infer that LytSR modulates spore engulfment by directly or indirectly inducing the transcription of the sporulation gene spoIIP. However, many other genes are also involved in engulfment and must be examined in future studies because they may be more directly responsible for the sporulation phenotype.
Author Contributions
FS designed the research. QP and JW performed the experimental work. QP drafted the manuscript. JW, XC, and LQ constructed the mutants, analyzed the sporulation efficiency and perform the laser scanning confocal microscopy. FS, JZ, and HT critically revised the manuscript for intellectual content. All authors read and approved the final version of the manuscript.
Funding
This work was supported by grants from the National Natural Science Foundation of China (nos. 31530095) and the National Key Research and Development Program of China (2017YFD0200400).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors would like to thank Dr. Didier Lereclus from the Institut National de la Recherche Agronomique for his helpful suggestions.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fcimb.2017.00468/full#supplementary-material
References
Agaisse, H., and Lereclus, D. (1994). Structural and functional analysis of the promoter region involved in full expression of the cryIIIA toxin gene of Bacillus thuringiensis. Mol. Microbiol. 13, 97–107. doi: 10.1111/j.1365-2958.1994.tb00405.x
Arantes, O., and Lereclus, D. (1991). Construction of cloning vectors for Bacillus-thuringiensis. Gene 108, 115–120. doi: 10.1016/0378-1119(91)90495-W
Arnaud, M., Chastanet, A., and Débarbouillé, M. (2004). New vector for efficient allelic replacement in naturally nontransformable, low-GC-content, gram-positive bacteria. Appl. Environ. Microbiol. 70, 6887–6891. doi: 10.1128/AEM.70.11.6887-6891.2004
Stenfors Arnesen, L. P., Fagerlund, A., and Granum, P. E. (2008). From soil to gut: Bacillus cereus and its food poisoning toxins. FEMS Microbiol. Rev. 32, 579–606. doi: 10.1111/j.1574-6976.2008.00112.x
Brunskill, E. W., and Bayles, K. W. (1996). Identification and molecular characterization of a putative regulatory locus that affects autolysis in Staphylococcus aureus. J. Bacteriol. 178, 611–618. doi: 10.1128/jb.178.3.611-618.1996
Burbulys, D., Trach, K. A., and Hoch, J. A. (1991). Initiation of sporulation in B. subtilis is controlled by a multicomponent phosphorelay. Cell 64, 545–552. doi: 10.1016/0092-8674(91)90238-T
Chandramohan, L., Ahn, J.-S., Weaver, K. E., and Bayles, K. W. (2009). An overlap between the control of programmed cell death in Bacillus anthracis and sporulation. J. Bacteriol. 191, 4103–4110. doi: 10.1128/JB.00314-09
Chastanet, A., and Losick, R. (2007). Engulfment during sporulation in Bacillus subtilis is governed by a multi-protein complex containing tandemly acting autolysins. Mol. Microbiol. 64, 139–152. doi: 10.1111/j.1365-2958.2007.05652.x
de Been, M., Tempelaars, M. H., Van Schaik, W., Moezelaar, R., Siezen, R. J., and Abee, T. (2010). A novel hybrid kinase is essential for regulating the sigma B-mediated stress response of Bacillus cereus. Environ. Microbiol. 12, 730–744. doi: 10.1111/j.1462-2920.2009.02116.x
Du, L., Qiu, L., Peng, Q., Lereclus, D., Zhang, J., Song, F., et al. (2012). Identification of the promoter in the intergenic region between orf1 and cry8Ea1 controlled by sigma H factor. Appl. Environ. Microbiol. 78, 4164–4168. doi: 10.1128/AEM.00622-12
Eichenberger, P., Fawcett, P., and Losick, R. (2001). A three-protein inhibitor of polar septation during sporulation in Bacillus subtilis. Mol. Microbiol. 42, 1147–1162. doi: 10.1046/j.1365-2958.2001.02660.x
Errington, J. (2003). Regulation of endospore formation in Bacillus subtilis. Nat. Rev. Microbiol. 1, 117–126. doi: 10.1038/nrmicro750
Fujita, M., Gonzalez-Pastor, J. E., and Losick, R. (2005). High- and low-threshold genes in the Spo0A regulon of Bacillus subtilis. J. Bacteriol. 187, 1357–1368. doi: 10.1128/JB.187.4.1357-1368.2005
Fujita, M., and Losick, R. (2003). The master regulator for entry into sporulation in Bacillus subtilis becomes a cell-specific transcription factor after asymmetric division. Genes Dev. 17, 1166–1174. doi: 10.1101/gad.1078303
Gutierrez, J., Smith, R., and Pogliano, K. (2010). SpoIID-mediated peptidoglycan degradation is required throughout engulfment during Bacillus subtilis sporulation. J. Bacteriol. 192, 3174–3186. doi: 10.1128/JB.00127-10
Hilbert, D. W., and Piggot, P. J. (2004). Compartmentalization of gene expression during Bacillus subtilis spore formation. Microbiol. Mol. Biol. Rev. 68, 234–262. doi: 10.1128/MMBR.68.2.234-262.2004
Jiang, M., Shao, W. L., Perego, M., and Hoch, J. A. (2000). Multiple histidine kinases regulate entry into stationary phase and sporulation in Bacillus subtilis. Mol. Microbiol. 38, 535–542. doi: 10.1046/j.1365-2958.2000.02148.x
Lehman, M. K., Bose, J. L., Sharma-Kuinkel, B. K., Moormeier, D. E., Endres, J. L., Sadykov, M. R., et al. (2015). Identification of the amino acids essential for LytSR-mediated signal transduction in Staphylococcus aureus and their roles in biofilm-specific gene expression. Mol. Microbiol. 95, 723–737. doi: 10.1111/mmi.12902
Lereclus, D., Arantes, O., Chaufaux, J., and Lecadet, M. (1989). Transformation and expression of a cloned delta-endotoxin gene in Bacillus thuringiensis. FEMS Microbiol. Lett. 51, 211–217.
Liu, G., Song, L., Shu, C., Wang, P., Deng, C., Peng, Q., et al. (2013). Complete genome sequence of Bacillus thuringiensis subsp. kurstaki strain HD73. Genome Announc. 1:e00080–13. doi: 10.1128/genomeA.00080-13
Meyer, P., Gutierrez, J., Pogliano, K., and Dworkin, J. (2010). Cell wall synthesis is necessary for membrane dynamics during sporulation of Bacillus subtilis. Mol. Microbiol. 76, 956–970. doi: 10.1111/j.1365-2958.2010.07155.x
Ohara, K., Fukuda, T., Okada, H., Kitao, S., Ishida, Y., Kato, K., et al. (2015). Identification of significant amino acids in multiple transmembrane domains of human transient receptor potential ankyrin 1 (TRPA1) for activation by eudesmol, an oxygenized sesquiterpene in hop essential oil. J. Biol. Chem. 290, 3161–3171. doi: 10.1074/jbc.M114.600932
Patel, K., and Golemi-Kotra, D. (2015). Signaling mechanism by the Staphylococcus aureus two-component system LytSR: role of acetyl phosphate in bypassing the cell membrane electrical potential sensor LytS. F1000Res. 4:79. doi: 10.12688/f1000research.6213.1
Peng, Q., Yang, M., Wang, W., Han, L., Wang, G., Wang, P., et al. (2014). Activation of gab cluster transcription in Bacillus thuringiensis by gamma-aminobutyric acid or succinic semialdehyde is mediated by the Sigma 54-dependent transcriptional activator GabR. BMC Microbiol. 14:306. doi: 10.1186/s12866-014-0306-3
Perchat, S., Dubois, T., Zouhir, S., Gominet, M., Poncet, S., Lemy, C., et al. (2011). A cell-cell communication system regulates protease production during sporulation in bacteria of the Bacillus cereus group. Mol. Microbiol. 82, 619–633. doi: 10.1111/j.1365-2958.2011.07839.x
Pogliano, J., Osborne, N., Sharp, M. D., Abanes-De Mello, A., Perez, A., Sun, Y. L., et al. (1999). A vital stain for studying membrane dynamics in bacteria: a novel mechanism controlling septation during Bacillus subtilis sporulation. Mol. Microbiol. 31, 1149–1159. doi: 10.1046/j.1365-2958.1999.01255.x
Rice, K. C., Nelson, J. B., Patton, T. G., Yang, S. J., and Bayles, K. W. (2005). Acetic acid induces expression of the Staphylococcus aureus cidABC and lrgAB murein hydrolase regulator operons. J. Bacteriol. 187, 813–821. doi: 10.1128/JB.187.3.813-821.2005
Schaeffer, P., Millet, J., and Aubert, J.-P. (1965). Catabolic repression of bacterial sporulation. Proc. Natl. Acad. Sci. U.S.A. 54, 704–711. doi: 10.1073/pnas.54.3.704
Schnepf, E., Crickmore, N., Van Rie, J., Lereclus, D., Baum, J., Feitelson, J., et al. (1998). Bacillus thuringiensis and its pesticidal crystal proteins. Microbiol. Mol. Biol. Rev. 62, 775–806.
Sharma-Kuinkel, B. K., Mann, E. E., Ahn, J. S., Kuechenmeister, L. J., Dunman, P. M., and Bayles, K. W. (2009). The Staphylococcus aureus LytSR two-component regulatory system affects biofilm formation. J. Bacteriol. 191, 4767–4775. doi: 10.1128/JB.00348-09
Shida, T., Hattori, H., Ise, F., and Sekiguchi, J. (2001). Mutational analysis of catalytic sites of the cell wall lytic N-acetylmuramoyl-L-alanine amidases Cw1C and Cw1V. J. Biol. Chem. 276, 28140–28146. doi: 10.1074/jbc.M103903200
Skerker, J. M., Prasol, M. S., Perchuk, B. S., Biondi, E. G., and Laub, M. T. (2005). Two-component signal transduction pathways regulating growth and cell cycle progression in a bacterium: a system-level analysis. PLoS Biol. 3:e334. doi: 10.1371/journal.pbio.0030334
Song, F., Peng, Q., Brillard, J., Buisson, C., de Been, M., Abee, T., et al. (2012). A multicomponent sugar phosphate sensor system specifically induced in Bacillus cereus during infection of the insect gut. FASEB J. 26, 3336–3350. doi: 10.1096/fj.11-197681
Tan, I. S., and Ramamurthi, K. S. (2014). Spore formation in Bacillus subtilis. Environ. Microbiol. Rep. 6, 212–225. doi: 10.1111/1758-2229.12130
van den Esker, M. H., Kovacs, A. T., and Kuipers, O. P. (2017). YsbA and LytST are essential for pyruvate utilization in Bacillus subtilis. Environ. Microbiol. 19, 83–94. doi: 10.1111/1462-2920.13454
van Schaik, W., Tempelaars, M. H., Zwietering, M. H., de Vos, W. M., and Abee, T. (2005). Analysis of the role of RsbV, RsbW, and RsbY in regulating sigma(B) activity in Bacillus cereus. J. Bacteriol. 187, 5846–5851. doi: 10.1128/JB.187.16.5846-5851.2005
Yang, J., Peng, Q., Chen, Z., Deng, C., Shu, C., Zhang, J., et al. (2013). Transcriptional regulation and characteristics of a novel N-Acetylmuramoyl-l-alanine amidase gene involved in Bacillus thuringiensis mother cell lysis. J. Bacteriol. 195, 2887–2897. doi: 10.1128/JB.00112-13
Yang, S. J., Xiong, Y. Q., Yeaman, M. R., Bayles, K. W., Abdelhady, W., and Bayer, A. S. (2013). Role of the LytSR two-component regulatory system in adaptation to cationic antimicrobial peptides in Staphylococcus aureus. Antimicrob. Agents Chemother. 57, 3875–3882. doi: 10.1128/AAC.00412-13
Keywords: two-component system, LytSR, sporulation, spoIIP, Bacillus thuringiensis
Citation: Peng Q, Wu J, Chen X, Qiu L, Zhang J, Tian H and Song F (2017) Disruption of Two-component System LytSR Affects Forespore Engulfment in Bacillus thuringiensis. Front. Cell. Infect. Microbiol. 7:468. doi: 10.3389/fcimb.2017.00468
Received: 25 April 2017; Accepted: 23 October 2017;
Published: 03 November 2017.
Edited by:
Jan Potempa, University of Louisville, United StatesReviewed by:
Shonna Marie McBride, Emory University, United StatesRichard A. Daniel, Newcastle University, United Kingdom
Copyright © 2017 Peng, Wu, Chen, Qiu, Zhang, Tian and Song. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hongtao Tian, dGh0NjMxMDIyQDE2My5jb20=
Fuping Song, ZnBzb25nQGlwcGNhYXMuY24=
†These authors have contributed equally to this work.
 Qi Peng
Qi Peng Jianbo Wu
Jianbo Wu Xiaomin Chen
Xiaomin Chen Lili Qiu
Lili Qiu Jie Zhang
Jie Zhang Hongtao Tian
Hongtao Tian Fuping Song
Fuping Song
