- 1National Engineering Research Center for Non-food Biorefinery, Guangxi Academy of Sciences, Nanning, China
- 2College of Life Science and Technology, Guangxi University, Nanning, China
- 3Department of Microbiology, University of Nigeria, Nsukka, Nigeria
- 4State Key Laboratory of Non-food Biomass and Enzyme Technology, Guangxi Academy of Sciences, Nanning, China
Aspergillus fumigatus is the most reported causative pathogen associated with the increasing global incidences of aspergilloses, with the health of immunocompromised individuals mostly at risk. Monitoring the pathogenicity of A. fumigatus strains to identify virulence factors and evaluating the efficacy of potent active agents against this fungus in animal models are indispensable in current research effort. Caenorhabditis elegans has been successfully utilized as an infection model for bacterial and dimorphic fungal pathogens because of the advantages of being time-efficient, and less costly. However, application of this model to the filamentous fungus A. fumigatus is less investigated. In this study, we developed and optimized a stable and reliable C. elegans model for A. fumigatus infection, and demonstrated the infection process with a fluorescent strain. Virulence results of several mutant strains in our nematode model demonstrated high consistency with the already reported pathogenicity pattern in other models. Furthermore, this C. elegans-A. fumigatus infection model was optimized for evaluating the efficacy of current antifungal drugs. Interestingly, the azole drugs in nematode model prevented conidial germination to a higher extent than amphotericin B. Overall, our established C. elegans infection model for A. fumigatus has potential applications in pathogenicity evaluation, antifungal agents screening, drug efficacy evaluation as well as host-pathogen interaction studies.
Introduction
Aspergillus fumigatus is a saprophytic environmental fungus with ubiquitous airborne spores. It is also an opportunistic fungal pathogen responsible for mycoses including invasive aspergillosis (IA) mostly in immunocompromised patients (Van De Veerdonk et al., 2017; Fang and Latge, 2018). IA is a very severe systemic infection with an estimated global incidence of over 200,000 per annum (Geissel et al., 2018) and a mortality rate close to 100% in most groups of patients who did not receive treatment (Darling and Milder, 2018). Unfortunately, cases of aspergillosis have also been reported in immunocompetent patients (Stevens and Melikian, 2011). The emergence of drug resistant strains to the wide range of currently available drugs has posed a serious challenge as cases are rising globally (Prigitano et al., 2017, 2019; Abdolrasouli et al., 2018). Reports of A. fumigatus resistance to azole drugs (Hagiwara et al., 2017; Sharma et al., 2019), polyene drugs such as amphotericin B (Ashu et al., 2018), and echinocandin drugs (Beer et al., 2018) are increasing in both clinical (Prigitano et al., 2017; Abdolrasouli et al., 2018; Hagiwara et al., 2018) and environmental isolates (Vaezi et al., 2018; Prigitano et al., 2019). With the limited repertoire of antifungal drug classes, there is therefore the need to urgently discover new therapeutic option for this infection.
Notable strategy will involve studying key virulence factors in suitable model as a tool for understanding pathogenesis as well as screening and testing the efficacy of antifungal agents. Caenorhabditis elegans possesses great advantages such as simple life cycle, ease of cultivation and manipulation, relatively low space as well as no ethical requirement, making it an excellent model for a broad range of pathological diseases and infection. Indeed, C. elegans has been widely utilized as a host model for pathogenic bacterial infections, such as C. elegans-Pseudomonas aeruginosa (Darby et al., 1999; Tan et al., 1999; Zaborin et al., 2009), C. elegans-Salmonella spp. (Labrousse et al., 2000), C. elegans-Yersinia pestis (Styer et al., 2005), C. elegans-Staphylococcus aureus (Thompson and Brown, 2017), C. elegans-Streptococcus pyogenes (Jansen et al., 2002), and C. elegans-Acinetobacter baumanii (Vallejo et al., 2015). It has also been adopted for screening of active antimicrobial compounds and discovery of effective agents against several human bacterial (Kong et al., 2014; Tharmalingam et al., 2018) and fungal pathogens (Tampakakis et al., 2008; Okoli et al., 2009). Furthermore, C. elegans model has been applied for evaluating the pathogenicity of a number of clinically relevant fungal pathogens, including Candida albicans, Candida krusei, Candida parapsilosis, Candida glabrata (Breger et al., 2007), Histoplasma capsulatum (Johnson et al., 2009), Cryptococcus neoformans (Mylonakis et al., 2002), and Penicillium marneffei (Huang et al., 2014).
Currently Galleria mellonella (Gomez-Lopez et al., 2014), Bombyx mori (Nakamura et al., 2017), Drosophila melanogaster (Lionakis and Kontoyiannis, 2012), mice (Paulussen et al., 2015), and guinea pigs (Wiederhold et al., 2015) have been utilized as in vivo models for A. fumigatus infection. On the contrary, little attention has been given to the application of C. elegans model in filamentous fungal infections, except for the only report in A. fumigatus (Okoli and Bignell, 2015). Part of the limitations to its use is that the co-culture technique of infecting C. elegans is only suitable for bacterial or dimorphic fungal pathogens but not appropriate for filamentous fungi, as rapid conidia germination and intensive hyphal filamentation would interfere with the survival assessment of worms in killing assay. Despite the fact that Okoli and Bignell have demonstrated the possibility of adopting the C. elegans model for A. fumigatus infection, further optimization is required to increase the efficiency in worms-spores separation and shorten operation time. This will enable suitable applications of the model in high-throughput screening for novel antifungal compounds, evaluating pathogenicity as well as testing the efficacy of current antifungal agents. In this study, we established a stable and reliable C. elegans-A. fumigatus infection model, confirmed by evaluating the pathogenicity of several A. fumigatus mutants strains. The model was also optimized for evaluating the efficacy of currently used antifungal agents.
Results
Establishment and Optimization of C. elegans Model for A. fumigatus Infection
C. elegans has been successfully utilized as an infection model for several clinically relevant fungal pathogens, such as C. albicans, C. glabrata, C. neoformans, and H. capsulatum (Mylonakis et al., 2002; Breger et al., 2007; Johnson et al., 2009), but with limited application in filamentous fungal pathogens like A. fumigatus. Considering the fast germination feature of A. fumigatus in Brain Heart Infusion medium (BHI), we adopted the pre-infection technique on solid plates as described by Okoli and Bignell (35). An extensive washing technique using hand-made device of filter membrane-attached-on-tube was applied to remove conidia that were not ingested by the worms at pre-infection assay. The pre-infection time, conidia concentration for adequate infection and suitable worm numbers for both pre-infection and post-infection were optimized. The whole procedure for establishing the infection model is summarized in Figure 1. Initially two C. elegans strains were used as the hosts: the single fem-3(q96) mutant which is incapable of producing progeny at 25°C, and the glp-4(bn2); sek-1(km4) double mutant that is also unable to produce progeny at 25°C as well as being immunocompromised. In order to monitor the conidial ingestion and infection progression stages, we chose A. fumigatus Af293-dsRed strain with continuously red fluorescent signal (Jhingran et al., 2012), and the widely used parental strain KU80Δ for nematode model establishment (Table 1). Both Af293-dsRed and KU80Δ were able to kill and significantly reduce survival rate of C. elegans fem-3(q96) strain compared to heat-killed KU80Δ and Escherichia coli OP50, the preferred food of nematode (P < 0.0001) (Figure 2A, Supplementary Table 1). Interestingly, significant statistical difference was also observed between the survival rates of KU80Δ and Af293-dsRed (P < 0.0001) infections, indicating that Af293-dsRed is less virulent than KU80Δ. Similar susceptibility patterns and statistical values were obtained in the double mutant glp-4(bn2); sek-1(km4) host (Figure 2B, Supplementary Table 2). However, the worm survival rates with Af293-dsRed and KU80Δ in glp-4(bn2); sek-1(km4) mutant were much lower than that in fem-3(q96), demonstrating that the immunocompromised mutant worm is indeed more susceptible to fungal infections. To further substantiate the ability of our C. elegans-A. fumigatus infection model to be employed in evaluating pathogenicity of A. fumigatus strains, clinical strain Af293, which is also the parental strain of Af239-dsRed, was used for infection. The result showed that this strain displayed the same virulence as the Af293-dsRed strain, but was significantly less virulent than the KU80Δ strain (P < 0.0001) (Figure 2B, Supplementary Table 2). After 72 h of killing assay, the survival rates of glp-4(bn2); sek-1(km4) with Af293-dsRed, KU80Δ and Af293 were reduced to 6, 8, and 5%, respectively (Supplementary Table 2). The glp-4(bn2);sek-1(km4) strain was therefore used as the host strain for all the subsequent experiments.
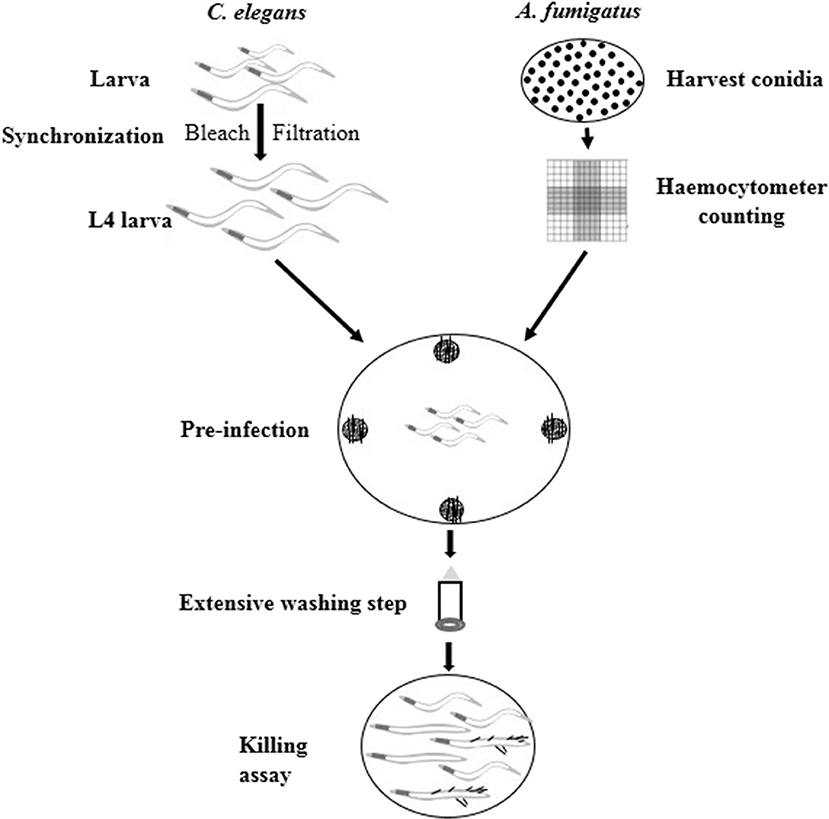
Figure 1. Schematic procedure of C. elegans-based A. fumigatus infection model. Synchronized L4 stage worms were put into NGM plates containing A. fumigatus spores at the four edges for 16 h pre-infection. Then extensive washing step was applied to remove conidia that were not ingested by worms. Finally the washed worms were added to BHI medium for killing assay.
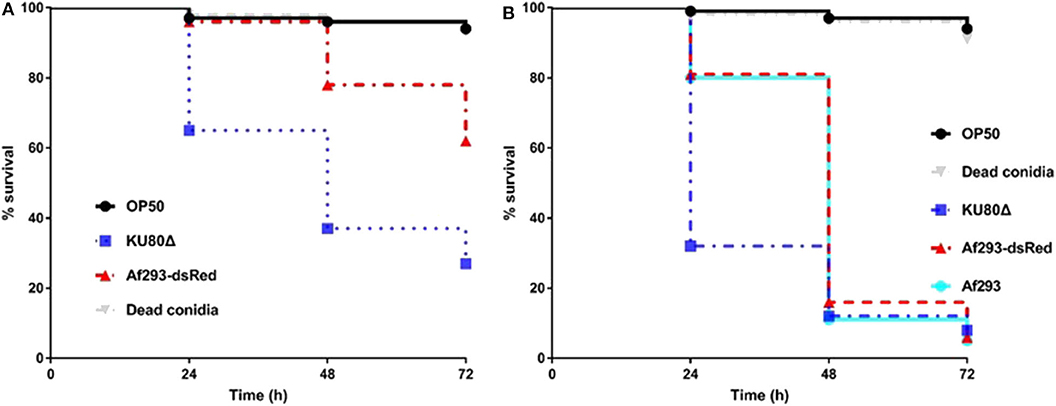
Figure 2. Survival rate comparison in C. elegans hosts. (A), Kaplan-Meier survival plots of fem-3(q96) fed by E.coli OP50 or conidia of indicated A. fumigatus strains. (B), Kaplan-Meier survival plots of glp-4(bn2); sek-1(km4) fed by E.coli OP50 or conidia of indicated A. fumigatus strains. Compared to OP50 and dead conidia the Af293-dsRed and KU80Δ strains exhibited significant pathogenicity [P < 0.0001, Log-rank (Mantel-Cox) test] in both fem-3(q96) and glp-4(bn2); sek-1(km4) worms. Three biological repeats (each with triplicates) were conducted for each strain.
Infection and Progression Stages of A. fumigatus in C. elegans
To understand the infection and progression stages of A. fumigatus in C. elegans, Af293-dsRed strain was used to infect glp-4(bn2); sek-1(km4) worms and spores progression was monitored using fluorescence microscope. As shown in Figure 3, the ingestion of conidia was clearly evident throughout the nematode intestine at the beginning of killing assay (Figure 3, Supplementary Figure 1). The conidia started to germinate with hyphae protruding from head, body and tail of C. elegans by 24 h (Figure 3, Supplementary Figure 2). When the killing assay reached 48 h, the hyphae became longer filaments protruding all over the worms' cuticle (Figure 3, Supplementary Figure 3). Furthermore, by 72 h, long hyphae diffused all through the body of the worms, making them appear as “ghosts” bodies (Figure 3). Survival of C. elegans was judged mainly by movement and sinusoidal shape as dead worms could not move and appeared straightened or slightly bent in shape, and mostly accompanied with fungal filaments.
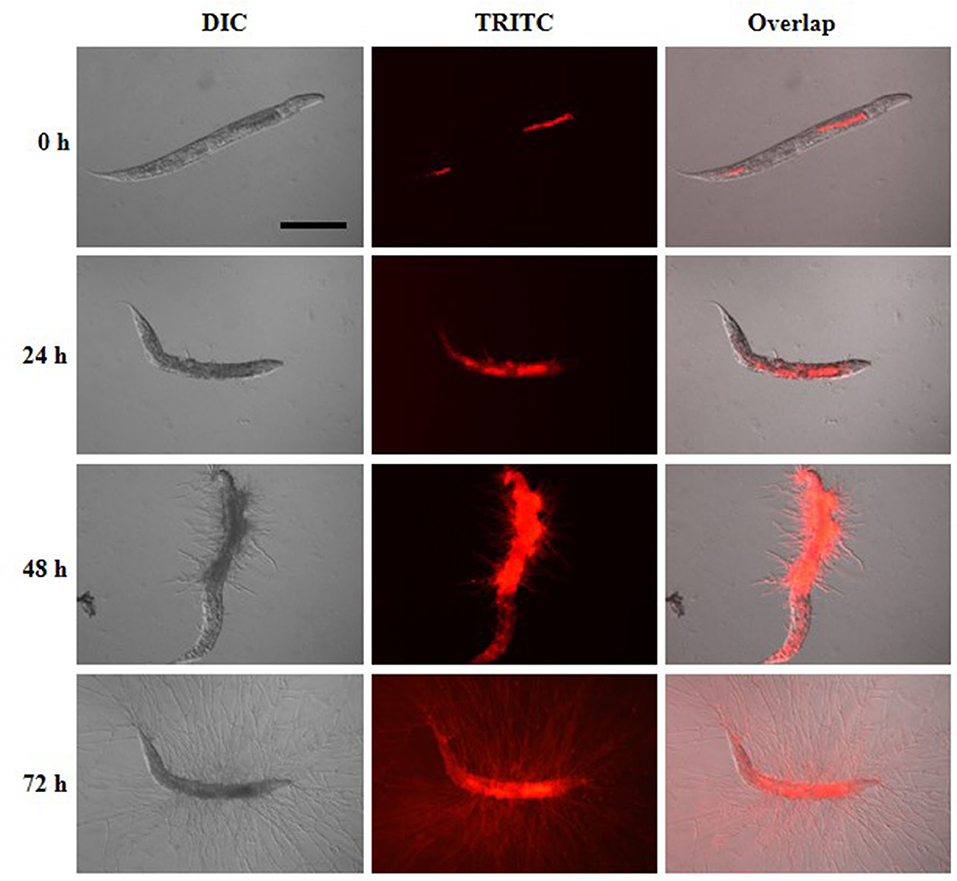
Figure 3. Infection and progression stages of Af293-dsRed in infection to glp-4(bn2); sek-1(km4) worms. Images were taken under DIC and TRITC channels at killing time of 0, 24, 48, and 72 h. Scale bar is 200 μm.
Pathogenicity Pattern of A. fumigatus Mutant Strains in C. elegans Model
Following establishing the C. elegans-A. fumigatus infection model, we hypothesized that the model would be able to assess the pathogenicity of A. fumigatus mutant strains whose virulence have been tested in other infection models. Six mutant strains were collected including triple agsΔ mutant, ΔpksP mutant, ΔmrsA mutant, ΔleuB mutant, and ΔtptA mutant strains in which attenuated pathogenicity has been previously reported in mice or G. mellonella infection models (Pihet et al., 2009; Beauvais et al., 2013; Long et al., 2016, 2018; Huang et al., 2019), and Δafmid1 mutant with an augmented virulence reported in mice model (Jiang et al., 2014) (Table 1). The triple agsΔ mutant, obtained by sequential deletions of the three α-1,3 glucan synthase genes (AGS1, AGS2, and AGS3), is unable to synthesize α-(1, 3)-glucan, a major cell wall polysaccharide component (Henry et al., 2012; Beauvais et al., 2013). Compared to the parental KU80Δ strain, the triple agsΔ mutant displayed significant attenuated virulence (P < 0.0001) with the survival rate of 56 ± 6.9 by 72 h in killing assay (Figure 4A, Supplementary Table 3). ΔpksP is the knockout strain of polyketide synthase gene pksp, which is involved in cell wall melanin synthesis (Pihet et al., 2009; Bayry et al., 2014). There was statistically significant difference between the virulence of ΔpksP mutant and the KU80Δ (P < 0.05), with a higher virulence observed with KU80Δ indicating that this mutant is less virulent than KU80Δ. MrsA and TptA are mitochondrial transporters involved in iron adaptation and homeostasis whereas LeuB is a Zn2Cys6-type transcription factor for leucine biosynthesis and iron acquisition (Long et al., 2016, 2018; Huang et al., 2019). Reduced pathogenicity was recorded in our nematode model for ΔmrsA, ΔleuB, and ΔtptA strains, with high significant difference (P < 0.0001) when compared to KU80Δ (Figure 4A, Supplementary Table 3). However, Δafmid1 strain, which is devoid of the plasma ion channel for replenishing extracellular calcium (Jiang et al., 2014), showed no significant virulence difference with KU80Δ (P > 0.05) (Figure 4A, Supplementary Table 3). This observation is in contrast to its enhanced pathogenicity above KU80Δ reported in murine model (Jiang et al., 2014). Among the mutant strains, the triple agsΔ is the least virulent, followed by ΔtptA and ΔleuB strains, and then the less attenuated ΔpksP and ΔmrsA strains. Interestingly, the hyphal filamentation rates at 24 h of killing assay followed the same pattern as the survival rates. The Δafmid1 mutant and the parental strain KU80Δ had the highest filamentation rate whereas the triple agsΔ mutant had the least (Figure 4B, Supplementary Table 3). It was observed that the more virulent the strain is, the higher the hyphal filamentation rate, suggesting that the death of the infected worms was mainly due to the germination and hyphal growth of conidia inside the worms which eventually led to puncture of worms when the filaments started to protrude. We noted that the filamentation of these mutant strains in C. elegans model did not correlate with in vitro growth in BHI medium, of which the triple agsΔ mutant exhibited the fastest germination, followed by KU80Δ, Δpksp, ΔleuB, and Δafmid1 mutant strains, while the ΔmrsA and ΔtptA mutant strains were the slowest (Supplementary Figure 4).
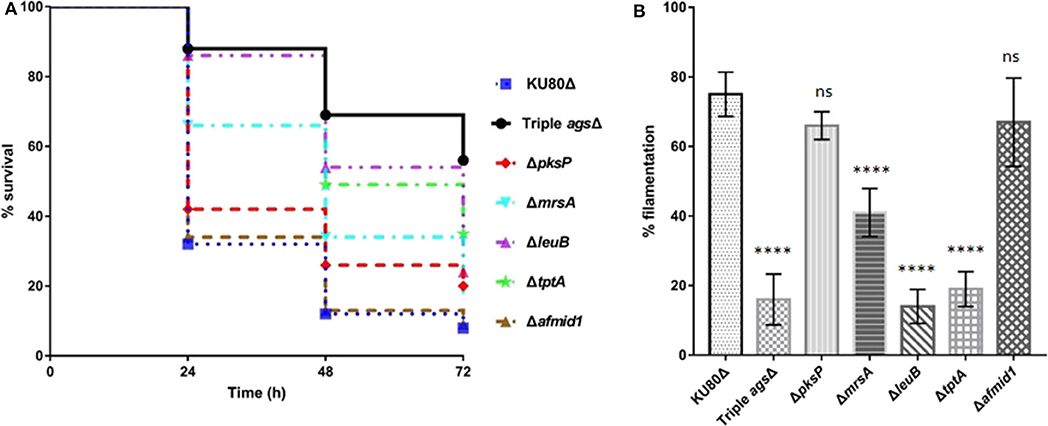
Figure 4. Survival curve and hyphal filamentation rate of A. fumigatus mutant strains in C. elegans model. (A), Kaplan-Meier survival plots of glp-4(bn2); sek-1(km4) on solid BHI plates after 16 h pre-infection with conidia of indicate A. fumigatus mutant strains. (B), Hyphal filamentation rates resulting from infection by indicated A. fumigatus mutant strains at 24 h in killing assay. Three biological repeats (each with triplicates) were conducted for each strain. One-way ANOVA was used for statistical analysis of filamentation rate, where p > 0.05 showing not significant, ns; p ≤ 0.0001 showing highly significant, ****, compared to the KU80Δ strain.
Evaluation of Antifungal Drug Efficacy in C. elegans-A. fumigatus Infection Model
Having established and confirmed the virulence pattern of our C. elegans-A. fumigatus infection model, the next challenge was to determine if clinically used antifungal drugs could rescue the aspergillosis infection and death in the nematode model. To achieve a uniformity exposure of the worms to the antifungals as well as ensure sufficient bioavailability of the drug, we optimized the pre-infection time for the liquid killing assay. Pre-infections of 8, 12, and 16 h on solid medium were tested and 8 h was chosen as the best duration for the worms to ingest sufficient conidia while also allowing adequate antifungal treatment test. Amphotericin B (AmB), itraconazole (ItrZ), and voriconazole (VoZ) were used in our C. elegans antifungal assay since they are the three commonly used drugs for aspergillosis treatment in clinical settings. All the three drugs showed excellent efficacy and increased the survival of infected worms when compared to the DMSO control treatments. AmB exhibited statistically significant rescue at concentration of 1 μg/ml or higher when compared to DMSO control (P < 0.0001) (Figure 5A, Supplementary Table 4). Similar significant rescue was obtained for ItrZ and VoZ at ≥2 μg/ml and ≥0.5 μg/ml, respectively, compared to DMSO control (P < 0.0001) (Figures 5B,C, Supplementary Tables 5, 6). The rescue effects of these antifungal drugs illustrated that our C. elegans-A. fumigatus infection model is suitable for evaluating antifungal drug efficacy.
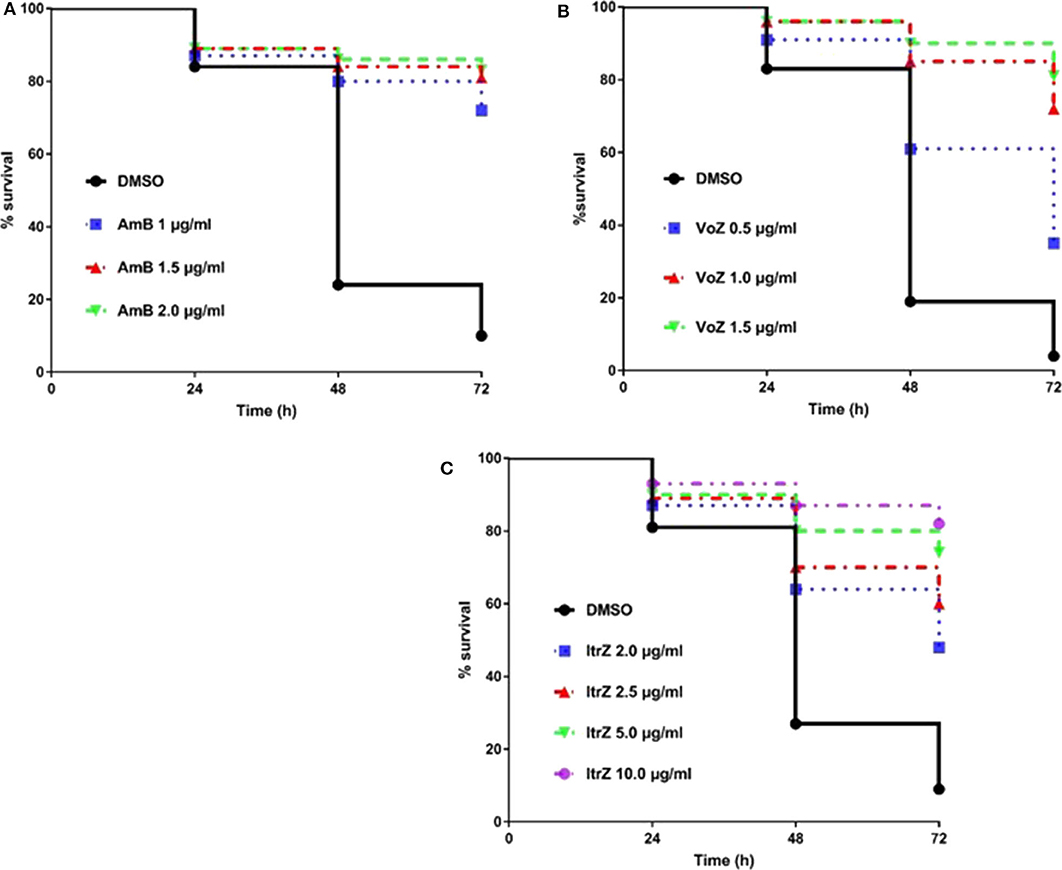
Figure 5. Kaplan-Meier survival plots of C. elegans infected by KU80Δ in the presence of antifungal drugs. The glp-4(bn2); sek-1(km4) worms were pre-infected with KU80Δ for 8 h then transferred into liquid killing media containing different concentrations of AmB (A), VoZ (B), and ItrZ (C). Three biological repeats (each with triplicates) were conducted for each concentration.
Killing Mode of Antifungal Drugs in C. elegans-A. fumigatus Infection Model
To analyze the killing mode of the antifungal drugs in C. elegans-A. fumigatus infection model, Af293-dsRed strain was used for infection of glp-4(bn2);sek-1(km4) worms. As shown in Figure 6, worms treated with DMSO had hyphal filaments protruding from their cuticles even at 24 h in killing assay. The fungal load from worms treated with the three drugs was significantly reduced as shown from the worm numbers with fluorescent signals (Figure 6). Moreover, dead worms from the AmB treated group possessed advanced hyphal growth whereas dead worms from ItrZ or VoZ treated group had strong accumulated fluorescent signals inside the body but without protruding hyphal filaments (Figure 6). It is interesting to note that ItrZ and VoZ exhibited superior antifungal activity against A. fumigatus by preventing the hyphal growth in C. elegans model when compared to AmB treatment.
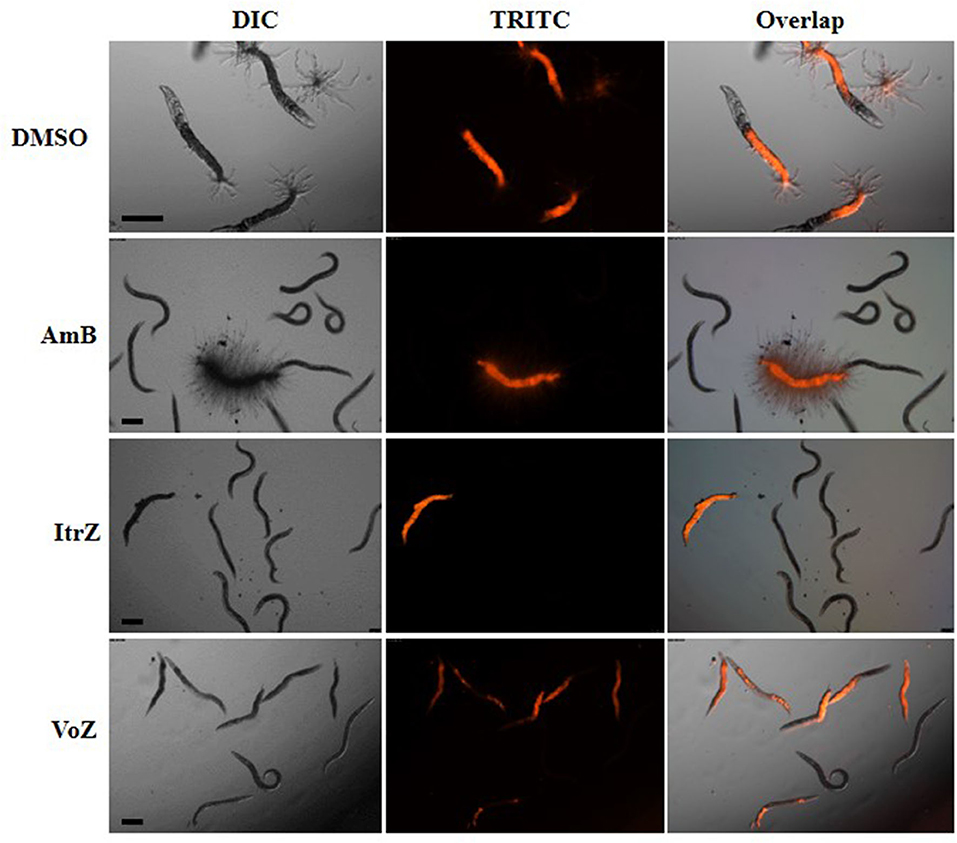
Figure 6. Effect of antifungal treatment on Af293-dsRed infection to glp-4(bn2); sek-1(km4) worms. Images were taken under DIC and TRITC channels from DMSO treatment by 24 h, 1.5 μg/ml AmB treatment by 48 h, 2 μg/ml ItrZ treatment by 72 h and 0.5 μg/ml VoZ treatment by 72 h. Scale bar is 200 μm.
Discussion
C. elegans as a model organism has been applied for evaluating pathogenicity, studying host-pathogen interactions, testing the efficacy of potential anti-infective compounds, and screening new antimicrobial agents against pathogenic bacteria and fungi. Okoli and Bignell had initially reported that C. elegans-based infection model could be applied to the filamentous fungus A. fumigatus (Okoli and Bignell, 2015). However, sufficient details on the infection process has not been provided to validate the model.
In this study through a series of optimization trials we developed a stable and efficient C. elegans-A. fumigatus infectious model. At the beginning of setting up the infection of the worms with A. fumigatus spores, fluorescence imaging showed that each of the single and double mutant worms all ingested varying amount of conidia. In addition to this challenge, it was also laborious and difficult to count the exact spore numbers from 100 or 1,000 of worms, which would involve crushing the worms and separating spores from worm lysate for counting. However, we noticed that the conidia concentration, worm numbers and the pre-infection time affected the amount of ingested fungal spores and survival of worms. Therefore, we standardized the concentration of spores and worm numbers, fixed the pre-infection time as 16 h for pathogenicity evaluation and applied conidia in four different points on the pre-infection plates to enhance the chances of conidia ingestion by the worms. As shown in Supplementary Tables 1–6, the survival rate of worms in killing assay displayed acceptable standard deviations from three biological repeats of each with triplicates, confirming the repeatability of the infection assay using our optimized protocol even if each worm may not ingest the same amount of spores. A device of filter membrane-attached-on-tube was developed to allow fast extensive worms-spores separation washing, thereby enabling better removal of conidia that were not ingested by the worms. The extensive washing after pre-infection also assisted easier observation of the killing assay without much obstruction from in vitro aggressive hyphal growth (Figure 1). Fluorescent Af293-dsRed strain provided further support to our findings that infection could emanate from any part of the worms including head, neck, abdomen, and tail regions through hyphal filaments protrusion (Figure 3), not only through the tail region as initially reported (Okoli and Bignell, 2015).
Previous studies have demonstrated that hyphal filamentation is an important virulence factor in fungal pathogenicity in mammal (Mitchell et al., 2007; Ariyachet et al., 2013; Ghosh et al., 2015) and nematode models (Breger et al., 2007; Pukkila-Worley et al., 2009; Huang et al., 2014). Similarly, in our C. elegans-A. fumigatus infection model, the in vivo germination and growth of conidia into filaments led to the quick death of worms (as the epidermal cuticles were disrupted) as well as limited worm's movement due to worm-hyphae adhesion to the medium surface. In both the single and double mutant worms, strain KU80Δ exhibited higher virulence than Af293 and Af293-dsRed strain, corresponding to higher hyphal filamentation rate at 24 h in killing assay (Figure 2, Supplementary Tables 1, 2). To further corroborate the role of hyphal filamentation in the virulence of A. fumigatus in C. elegans, our results from heat-killed conidia gave no hyphal filamentation as expected and displayed no difference from E.coli OP50 with regard to the survival rate of worms (Figure 2, Supplementary Tables 1, 2).
Assessing the pathogenicity of A. fumigatus strains is one of the main objective of setting up the nematode model. Among the selected six A. fumigatus mutant strains, triple agsΔ mutant, ΔpksP, ΔmrsA, and ΔtptA mutants all displayed significant attenuated virulence compared to the parental KU80Δ strain (Figure 4), consistent with their virulence patterns already shown in immunocompromised murine model of aspergillosis (Beauvais et al., 2013; Bayry et al., 2014; Long et al., 2016; Huang et al., 2019). Similar attenuated virulence was obtained for ΔleuB mutant in our C. elegans model which is in agreement with previously described pathogenicity pattern in G. mellonella model (Long et al., 2018). An exception was the Δafmidl mutant which was hypervirulent in murine model (Jiang et al., 2014) but exhibited no significant difference in our C. elegans model when compared to the KU80Δ strain. Currently we could not explain why Δafmidl virulence was not enhanced in C. elegans model but we strongly suspect that it could be due to the vast differences in in vivo environment between the nematode and mice, including the immune system and concentration of calcium ion, which requires further investigation. Again the correlation between virulence and hyphal filamentation was confirmed in infections caused by the six mutant strains (Figure 4B, Supplementary Table 3). However, it is noteworthy that some A. fumigatus strains, such as the ΔleuB and ΔtptA mutant strains, did not produce much filamentation at 24 h but eventually became relatively virulent by 72 h in killing assay, implying that hyphal filamentation is not the sole factor responsible for virulence, which further buttresses the need for molecular characterization. To the best of our knowledge, this is the first time that the pathogenicity of A. fumigatus strains in C. elegans infection model is being compared with that from other animal models. Our stable and reliable nematode infection model would be suitable for evaluating the virulence of more A. fumigatus mutant strains, such as mutants from the ongoing COFUN project, which plans to generate knockout mutants of all the coding genes in A. fumigatus (https://www.phe-culturecollections.org.uk/products/fungi/cofun-aspergillus-fumigatus-gene-wide-knock-out-collection.aspx).
The major drawbacks of the nematode model system include the inability to culture at 37°C and the absence of an adaptive immune response. Even the innate immune responses against pathogenic organisms in murine and C. elegans are different. Unlike in mammalian systems in which the transcription factor NF-κB, the Toll-like receptor (TLR) adaptor protein MYD88 and other components of the TLR signaling pathway are key to the innate immune system, C. elegans lacks these components. The C. elegans immune system however, mounts several conserved signaling pathways, including the p38 mitogen activated protein kinase (MAPK) cassette, the transforming growth factor (TGFβ), the insulin-like DAF-2-DAF-16, as well as the bZIP transcription factor ZIP-2 to defend against pathogens (Pukkila-Worley and Ausubel, 2012). Nevertheless, it is still unclear what the exact pathogenic triggers are and how the C. elegans detected and responded to pathogens. In murine model, the conidial cell wall of the triple agsΔ mutant was covered by a glycoprotein matrix, and PAMPs such as chitin and β-glucan were exposed, thus stimulating host immune response, which led to reduced virulence (Beauvais et al., 2013). Attenuated virulence of the Δpksp mutant in murine aspergillosis model is reported to be due to β-glucan exposure and the activated autophagy pathway of the LC3-associated phagocytosis (LAP) (Akoumianaki et al., 2016). The immune responses caused by the other four mutant strains are not clear yet in murine or G. mellonella models. Nevertheless, it will be desirable to uncover the immune defense of C. elegans in more detail by combining histological dissection of intestine epithelium cell structure and global transcription analysis of worms infected by KU80Δ and those mutant strains, particularly by the triple agsΔ mutant and the Δpksp mutant.
C. elegans model has been utilized in the evaluation of the efficacy of antifungal agents against infections caused by dimorphic fungi, such as C. albicans, C. krusei, C. parapsilosis, and P. marneffei (Breger et al., 2007; Huang et al., 2014) but not with the filamentous fungus, A. fumigatus. We have also demonstrated the possibility of adopting the C. elegans model for in vivo evaluation of antifungal agents on nematode aspergillosis. Shorter pre-infection time and liquid killing medium were applied in the assay to assess drug effectiveness. The minimum treatment doses of AmB, ItrZ, and VoZ in C. elegans were 1, 2, and 0.5 μg/ml, respectively, close to the in vitro MIC values of the tested antifungal drugs (Subcommittee on Antifungal Susceptibility Testing of the ESCMID European Committee for Antimicrobial Susceptibility Testing, 2008). The evaluation assay can easily be scaled up from 24-well-plates to 96-well or 384-well-plates for high-throughput screening of other bioactive compounds against A. fumigatus. Interestingly the killing mode of antifungal drugs in C. elegans model was different from that in the in vitro culture. The macrolide polyene AmB is a well-known broad-spectrum fungicidal agent (Hamill, 2013; Ashu et al., 2018) while azole drugs, i.e., VoZ and ItrZ, are mainly fungistatic against A. fumigatus (Meletiadis et al., 2007). However, visible hyphal filamentation from a number of infected worms graduated to “ghost stage” under AmB treatment whereas only diffused internal hyphal growth and early germinated conidia were observed from worms treated with VoZ and ItrZ (Figure 6). This is suggestive that VoZ and ItrZ may be an attractive alternative to AmB in treatment of invasive pulmonary aspergillosis and reducing occurrence of disseminated aspergillosis in in vivo applications.
In summary, our established C. elegans-A. fumigatus infection model could serve as a preliminary screening for pathogenic phenotypes of A. fumigates strains. The application of this model to evaluate the efficacy of antifungal agents on nematode aspergillosis could be the groundwork to develop future antifungal agents against A. fumigatus.
Materials and Methods
Strains, Media, and Culture Conditions
The A. fumigatus strains used in this study are summarized in Table 1. The strains were grown on A. fumigatus complete medium (Subcommittee on Antifungal Susceptibility Testing of the ESCMID European Committee for Antimicrobial Susceptibility Testing, 2008) slants composed of 1 g/l yeast extract, 2 g/l peptone, 10 g/l glucose, 1.5 g/l casein hydrolysate acid, 1 ml/l trace element solution, 20 ml/l 50X salt solution. After incubation at 37°C for 24–72 h (depending on mutant strains) conidia were harvested with 0.2% (v/v) Tween 20 aqueous solution, centrifuged and then Tween 20 aqueous solution was decanted out. The conidia were then re-suspended in M9 buffer and standardized to 1.25 × 108 spores/ml in Eppendorf tube.
The C. elegans strains fem-3(q96) and glp-4(bn2); sek-1(km4) were used in this study. The fem-3(q96) and glp-4(bn2) mutations are temperature sensitive and thus prevents them from producing progeny at 25°C while the sek-1(km4) mutation makes the worms immunocompromised. Worms were grown at 15°C in 90 mm Nematode Growth Medium (Fedorova et al., 2008) plates seeded with E. coli OP50.
Setting up C. elegans-A. fumigatus Infection Model
NGM medium plus antibiotics (NGM+) was used for pre-infection assay according to methods modified from Okoli and Bignell (2015). Ampicillin, streptomycin, and kanamycin were added to NGM at 100, 100, and 45 μg/ml, respectively. Unstarved worms for at least three generations were filtered through 11 μm pore-sized membrane filter (Merck Millipore Ltd.) using M9, to collect L1 worms for synchronization. Filtrated L1 worms were spread on NGM plates seeded with OP50, and then incubated at 25°C for 48 h for synchronization.
Synchronized L4 worms were washed off from the NGM plates with M9 and transferred to 50 ml tube to sediment for 10 min. The M9-OP50 mix was gently removed using micropipette. Three more sedimentation washes were performed and then standardized worms (about 200–500) were dispensed into triplicate 90 mm NGM+ plates and gently spread to dry. Worms were allowed to move around on the plates for 30 min before 25 μl of standardized conidia (1.25 × 108 spores/ml) were added at the four cardinal points of each NGM+ plates making a total of 100 μl conidia per plate. The plates were then incubated at 25°C for 16 h pre-infection period to allow worms ingest conidia. Control was setup with OP50 instead of conidia.
A 30% Brain Heart Infusion (BHI) plus 200 μg/ml ampicillin, 200 μg/ml streptomycin, and 90 μg/ml kanamycin in M9 solid medium (referred to as BHI+S) was used for solid killing assay. Pre-infected worms were washed off with M9 into 50 ml tube. Then we developed a hand-made “filter membrane-attached-on-tube” device with a 35 μm pore-sized membrane (Sango Biotech) designed into 15 ml Eppendorf tubes to wash out conidia that were not ingested by worms. The filter separated conidia that were not ingested from C. elegans to drastically reduce the amount of conidia that would otherwise interfere with the experiment when germinated on BHI+S plates. About 80–200 washed worms were dispensed into 90 mm BHI+S plates in triplicates. Living and dead worms with or without hyphal filament were recorded at 24 h in killing assay using dissecting microscope (Motic SMZ-168 series) whereas only dead and living worm numbers were recorded at 48 and 72 h of killing assay.
Microscopic Imaging
We used two approaches to prepare our worms for fluorescent microscopy. The first approach majorly applied to worms at 0 h in killing assay (immediately after 16 h of pre-infection). We made 2% agarose pad on slides and used worm picker to randomly collect worms from BHI+S plates. Worms were transferred to the agarose gel in M9 plus levamisole, then observed and recorded under both fluorescence and DIC channels of Leica upright microscope (Leica Microsystems, STP 8000). The second approach involved cutting out and transferring the BHI+S agar with worms to slides between 24 and 72 h in killing assay. This was very necessary as protrusion of hyphal filaments made worms attached to the medium.
The Procedure for Evaluating Antifungal Drugs Efficacy
The 8 h Pre-infection time and liquid killing assay were chosen for antifungal drugs evaluation and BHI+S without agar (referred to as BHI+L) was used as the killing medium. Amphotericin B, itraconazole, and voriconazole (MedChemExpress) were prepared in DMSO and diluted to 20 μg/ml with BHI+L (referred to as BHI+L+). Antifungal drug concentrations ranging from 0.5 to 2.0 μg/ml for amphotericin B, 1.0–2.0 μg/ml for voriconazole, and 1.0–10 μg/ml for itraconazole were utilized in 24-well-plates. BHI+L+ were dispensed into triplicates of 24-well-plates to a volume of 320 μl. Then 80 μl of pre-infected worms (~30–60) were added into each well to make the final volume of 400 μl. DMSO was used as control. Dead and living worm numbers were recorded using Leica inverted microscope (Leica DMC 5400). Antifungal phenotypic images were captured by the inverted microscope under both fluorescent and DIC channels.
Data Analysis
All infection experiments were performed in triplicates and all the experimental numerical data were expressed as the means ± S.D. The Kaplan-Meier survival curves were plotted by GraphPad Prism 7.0. Statistical P-values for survival rates were calculated by Log-rank (Mantel-Cox) test. One-way ANOVA was used for statistical analysis of filamentation rates. Images analysis were performed using ImageJ software.
Data Availability Statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation, to any qualified researcher.
Author Contributions
WF and BW conceptualized the study. WF helped with data curation. CA and QQ did the formal analysis. CA, QQ, AM, and JO carried out the investigation. QQ and WF contributed to software. CJ, WF, and BW supervised the study. CA wrote the original draft. AO, WF, and BW reviewed and edited the manuscript. All authors contributed to the article and approved the submitted version.
Funding
This work was funded by National Natural Science Foundation of China (31960032), Guangxi Natural Science Foundation (2018GXNSFAA138012) and Research Start-up Funding of Guangxi Academy of Sciences (2017YJJ025) to WF, Bagui Scholar Program Fund (2016A24) of Guangxi Zhuang Autonomous Region to CJ, National Natural Science Foundation of China (31960129) and Research Start-up Funding of Guangxi Academy of Sciences (2017YJJ026) to BW.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Prof. Jean-Paul Latgé from Institute of Pasteur for providing triple agsΔ and ΔpksP mutant, Prof. David S. Askew from University of Cincinnati and Prof. Tobias M. Hohl from Fred Hutchinson Cancer Research Center for providing the Af293-dsRed mutant, Prof. Ling Lu from Nanjing Normal Univeristy for providing the ΔmrsA, ΔtptA, ΔleuB, and Δafmid1 mutants, Caenorhabditis Genetics Center funded by the NIH Office of Research Infrastructure Programs (P40 OD010440) for providing C.elegans strains.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fcimb.2020.00320/full#supplementary-material
References
Abdolrasouli, A., Scourfield, A., Rhodes, J., Shah, A., Elborn, J. S., Fisher, M. C., et al. (2018). High prevalence of triazole resistance in clinical Aspergillus fumigatus isolates in a specialist cardiothoracic centre. Int. J. Antimicrob. Agents 52, 637–642. doi: 10.1016/j.ijantimicag.2018.08.004
Akoumianaki, T., Kyrmizi, I., Valsecchi, I., Gresnigt, M. S., Samonis, G., Drakos, E., et al. (2016). Aspergillus cell wall melanin blocks LC3-associated phagocytosis to promote pathogenicity. Cell Host Microbe 19, 79–90. doi: 10.1016/j.chom.2015.12.002
Ariyachet, C., Solis, N. V., Liu, Y., Prasadarao, N. V., Filler, S. G., and Mcbride, A. E. (2013). SR-like RNA-binding protein Slr1 affects Candida albicans filamentation and virulence. Infect. Immun. 81, 1267–1276. doi: 10.1128/IAI.00864-12
Ashu, E. E., Korfanty, G. A., Samarasinghe, H., Pum, N., You, M., Yamamura, D., et al. (2018). Widespread amphotericin B-resistant strains of Aspergillus fumigatus in Hamilton, Canada. Infect. Drug Resist. 11, 1549–1555. doi: 10.2147/IDR.S170952
Bayry, J., Beaussart, A., Dufrene, Y. F., Sharma, M., Bansal, K., Kniemeyer, O., et al. (2014). Surface structure characterization of Aspergillus fumigatus conidia mutated in the melanin synthesis pathway and their human cellular immune response. Infect. Immun. 82, 3141–3153. doi: 10.1128/IAI.01726-14
Beauvais, A., Bozza, S., Kniemeyer, O., Formosa, C., Balloy, V., Henry, C., et al. (2013). Deletion of the alpha-(1,3)-glucan synthase genes induces a restructuring of the conidial cell wall responsible for the avirulence of Aspergillus fumigatus. PLoS Pathog. 9:e1003716. doi: 10.1371/annotation/05c0ca66-4ed9-4c04-96c6-3addac835e04
Beer, K. D., Farnon, E. C., Jain, S., Jamerson, C., Lineberger, S., Miller, J., et al. (2018). Multidrug-resistant Aspergillus fumigatus carrying mutations linked to environmental fungicide exposure - three States, 2010-2017. MMWR Morb. Mortal. Wkly. Rep. 67, 1064–1067. doi: 10.15585/mmwr.mm6738a5
Breger, J., Fuchs, B. B., Aperis, G., Moy, T. I., Ausubel, F. M., and Mylonakis, E. (2007). Antifungal chemical compounds identified using a C. elegans pathogenicity assay. PLoS Pathog. 3:e18. doi: 10.1371/journal.ppat.0030018
Da Silva Ferreira, M. E., Kress, M. R., Savoldi, M., Goldman, M. H., Hartl, A., Heinekamp, T., et al. (2006). The akuB(KU80) mutant deficient for nonhomologous end joining is a powerful tool for analyzing pathogenicity in Aspergillus fumigatus. Eukaryotic Cell 5, 207–211. doi: 10.1128/EC.5.1.207-211.2006
Darby, C., Cosma, C. L., Thomas, J. H., and Manoil, C. (1999). Lethal paralysis of caenorhabditis elegans by Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. U.S.A. 96, 15202–15207. doi: 10.1073/pnas.96.26.15202
Darling, B. A., and Milder, E. A. (2018). Invasive aspergillosis. Pediatr. Rev. 39, 476–478. doi: 10.1542/pir.2017-0129
Fang, W., and Latge, J. P. (2018). Microbe profile: Aspergillus fumigatus: a saprotrophic and opportunistic fungal pathogen. Microbiology 164, 1009–1011. doi: 10.1099/mic.0.000651
Fedorova, N. D., Khaldi, N., Joardar, V. S., Maiti, R., Amedeo, P., Anderson, M. J., et al. (2008). Genomic islands in the pathogenic filamentous fungus Aspergillus fumigatus. PLoS Genet. 4:e1000046. doi: 10.1371/journal.pgen.1000046
Geissel, B., Loiko, V., Klugherz, I., Zhu, Z., Wagener, N., Kurzai, O., et al. (2018). Azole-induced cell wall carbohydrate patches kill Aspergillus fumigatus. Nat. Commun. 9:3098. doi: 10.1038/s41467-018-05497-7
Ghosh, A. K., Wangsanut, T., Fonzi, W. A., and Rolfes, R. J. (2015). The GRF10 homeobox gene regulates filamentous growth in the human fungal pathogen Candida albicans. FEMS Yeast Res. 15:fov093. doi: 10.1093/femsyr/fov093
Gomez-Lopez, A., Forastiero, A., Cendejas-Bueno, E., Gregson, L., Mellado, E., Howard, S. J., et al. (2014). An invertebrate model to evaluate virulence in Aspergillus fumigatus: the role of azole resistance. Med. Mycol. 52, 311–319. doi: 10.1093/mmy/myt022
Hagiwara, D., Arai, T., Takahashi, H., Kusuya, Y., Watanabe, A., and Kamei, K. (2018). Non-cyp51A azole-resistant Aspergillus fumigatus Isolates with mutation in HMG-CoA reductase. Emerging Infect. Dis. 24, 1889–1897. doi: 10.3201/eid2410.180730
Hagiwara, D., Miura, D., Shimizu, K., Paul, S., Ohba, A., Gonoi, T., et al. (2017). A Novel Zn2-Cys6 transcription factor AtrR plays a key role in an azole resistance mechanism of Aspergillus fumigatus by co-regulating cyp51A and cdr1B expressions. PLoS Pathog. 13:e1006096. doi: 10.1371/journal.ppat.1006096
Hamill, R. J. (2013). Amphotericin B formulations: a comparative review of efficacy and toxicity. Drugs 73, 919–934. doi: 10.1007/s40265-013-0069-4
Henry, C., Latge, J. P., and Beauvais, A. (2012). alpha1,3 glucans are dispensable in Aspergillus fumigatus. Eukaryotic Cell 11, 26–29. doi: 10.1128/EC.05270-11
Huang, J., Ma, Z., Zhong, G., Sheppard, D. C., Lu, L., and Zhang, S. (2019). The mitochondrial thiamine pyrophosphate transporter TptA promotes adaptation to low iron conditions and virulence in fungal pathogen Aspergillus fumigatus. Virulence 10, 234–247. doi: 10.1080/21505594.2019.1596505
Huang, X., Li, D., Xi, L., and Mylonakis, E. (2014). Caenorhabditis elegans: a simple nematode infection model for Penicillium marneffei. PLoS ONE 9:e108764. doi: 10.1371/journal.pone.0108764
Jansen, W. T., Bolm, M., Balling, R., Chhatwal, G. S., and Schnabel, R. (2002). Hydrogen peroxide-mediated killing of Caenorhabditis elegans by Streptococcus pyogenes. Infect. Immun. 70, 5202–5207. doi: 10.1128/IAI.70.9.5202-5207.2002
Jhingran, A, Mar, K. B., Kumasaka, D. K., Knoblaugh, S. E., Ngo, L. Y., Segal, B. H., et al. (2012). Tracing conidial fate and measuring host cell antifungal activity using a reporter of microbial viability in the lung. Cell Rep. 2, 1762–1773. doi: 10.1016/j.celrep.2012.10.026
Jiang, H., Shen, Y., Liu, W., and Lu, L. (2014). Deletion of the putative stretch-activated ion channel Mid1 is hypervirulent in Aspergillus fumigatus. Fungal Genet. Biol. 62, 62–70. doi: 10.1016/j.fgb.2013.11.003
Johnson, C. H., Ayyadevara, S., Mcewen, J. E., and Shmookler Reis, R. J. (2009). Histoplasma capsulatum and Caenorhabditis elegans: a simple nematode model for an innate immune response to fungal infection. Med. Mycol. 47, 808–813. doi: 10.3109/13693780802660532
Kong, C., Yehye, W. A., Abd Rahman, N., Tan, M. W., and Nathan, S. (2014). Discovery of potential anti-infectives against Staphylococcus aureus using a Caenorhabditis elegans infection model. BMC Complement. Altern. Med. 14:4. doi: 10.1186/1472-6882-14-4
Kyrmizi, I., Ferreira, H., Carvalho, A., Figueroa, J. A. L., Zarmpas, P., Cunha, C., et al. (2018). Calcium sequestration by fungal melanin inhibits calcium-calmodulin signalling to prevent LC3-associated phagocytosis. Nat Microbiol. 3, 791–803. doi: 10.1038/s41564-018-0167-x
Labrousse, A., Chauvet, S., Couillault, C., Kurz, C. L., and Ewbank, J. J. (2000). Caenorhabditis elegans is a model host for Salmonella typhimurium. Curr. Biol. 10, 1543–1545. doi: 10.1016/S0960-9822(00)00833-2
Lionakis, M. S., and Kontoyiannis, D. P. (2012). Drosophila melanogaster as a model organism for invasive aspergillosis. Methods Mol. Biol. 845, 455–468. doi: 10.1007/978-1-61779-539-8_32
Long, N., Orasch, T., Zhang, S., Gao, L., Xu, X., Hortschansky, P., et al. (2018). The Zn2Cys6-type transcription factor LeuB cross-links regulation of leucine biosynthesis and iron acquisition in Aspergillus fumigatus. PLoS Genet. 14:e1007762. doi: 10.1371/journal.pgen.1007762
Long, N., Xu, X., Qian, H., Zhang, S., and Lu, L. (2016). A putative mitochondrial iron transporter MrsA in Aspergillus fumigatus plays important roles in azole-, oxidative stress responses and virulence. Front. Microbiol. 7:716. doi: 10.3389/fmicb.2016.00716
Meletiadis, J., Antachopoulos, C., Stergiopoulou, T., Pournaras, S., Roilides, E., and Walsh, T. J. (2007). Differential fungicidal activities of amphotericin B and voriconazole against Aspergillus species determined by microbroth methodology. Antimicrob. Agents Chemother. 51, 3329–3337. doi: 10.1128/AAC.00345-07
Mitchell, B. M., Wu, T. G., Jackson, B. E., and Wilhelmus, K. R. (2007). Candida albicans strain-dependent virulence and Rim13p-mediated filamentation in experimental keratomycosis. Invest. Ophthalmol. Vis. Sci. 48, 774–780. doi: 10.1167/iovs.06-0793
Mylonakis, E., Ausubel, F. M., Perfect, J. R., Heitman, J., and Calderwood, S. B. (2002). Nonlinear partial differential equations and applications: Killing of Caenorhabditis elegans by Cryptococcus neoformans as a model of yeast pathogenesis. Proc. Natl. Acad. Sci. U.S.A. 99, 15675–15680. doi: 10.1073/pnas.232568599
Nakamura, I., Kanasaki, R., Yoshikawa, K., Furukawa, S., Fujie, A., Hamamoto, H., et al. (2017). Discovery of a new antifungal agent ASP2397 using a silkworm model of Aspergillus fumigatus infection. J. Antibiot. 70, 41–44. doi: 10.1038/ja.2016.106
Okoli, I., and Bignell, E. (2015). Caenorhabditis elegans-Aspergillus fumigatus (nematode-mould) model for study of fungal pathogenesis. Br. Microbiol. Res. J. 7, 93–99. doi: 10.9734/BMRJ/2015/15838
Okoli, I., Coleman, J. J., Tampakakis, E., An, W. F., Holson, E., Wagner, F., et al. (2009). Identification of antifungal compounds active against Candida albicans using an improved high-throughput Caenorhabditis elegans assay. PLoS ONE 4:e7025. doi: 10.1371/journal.pone.0007025
Paulussen, C., Boulet, G., Bosschaerts, T., Cos, P., Fortin, A., and Maes, L. (2015). Efficacy of oleylphosphocholine (OlPC) in vitro and in a mouse model of invasive aspergillosis. Mycoses 58, 127–132. doi: 10.1111/myc.12286
Pihet, M., Vandeputte, P., Tronchin, G., Renier, G., Saulnier, P., Georgeault, S., et al. (2009). Melanin is an essential component for the integrity of the cell wall of Aspergillus fumigatus conidia. BMC Microbiol. 9:177. doi: 10.1186/1471-2180-9-177
Prigitano, A., Esposto, M. C., Biffi, A., De Lorenzis, G., Favuzzi, V., Koncan, R., et al. (2017). Triazole resistance in Aspergillus fumigatus isolates from patients with cystic fibrosis in Italy. J. Cyst. Fibros. 16, 64–69. doi: 10.1016/j.jcf.2016.06.006
Prigitano, A., Esposto, M. C., Romano, L., Auxilia, F., and Tortorano, A. M. (2019). Azole-resistant Aspergillus fumigatus in the Italian environment. J. Glob. Antimicrob. Resist. 16, 220–224. doi: 10.1016/j.jgar.2018.10.017
Pukkila-Worley, R., and Ausubel, F. M. (2012). Immune defense mechanisms in the Caenorhabditis elegans intestinal epithelium. Curr. Opin. Immunol. 24, 3–9. doi: 10.1016/j.coi.2011.10.004
Pukkila-Worley, R., Peleg, A. Y., Tampakakis, E., and Mylonakis, E. (2009). Candida albicans hyphal formation and virulence assessed using a Caenorhabditis elegans infection model. Eukaryotic Cell 8, 1750–1758. doi: 10.1128/EC.00163-09
Sharma, C., Nelson-Sathi, S., Singh, A., Radhakrishna Pillai, M., and Chowdhary, A. (2019). Genomic perspective of triazole resistance in clinical and environmental Aspergillus fumigatus isolates without cyp51A mutations. Fungal Genet. Biol. 132:103265. doi: 10.1016/j.fgb.2019.103265
Stevens, D. A., and Melikian, G. L. (2011). Aspergillosis in the ‘nonimmunocompromised’ host. Immunol. Invest. 40, 751–766. doi: 10.3109/08820139.2011.614307
Styer, K. L., Hopkins, G. W., Bartra, S. S., Plano, G. V., Frothingham, R., and Aballay, A. (2005). Yersinia pestis kills Caenorhabditis elegans by a biofilm-independent process that involves novel virulence factors. EMBO Rep. 6, 992–997. doi: 10.1038/sj.embor.7400516
Subcommittee on Antifungal Susceptibility Testing of the ESCMID European Committee for Antimicrobial Susceptibility Testing (2008). EUCAST technical note on the method for the determination of broth dilution minimum inhibitory concentrations of antifungal agents for conidia-forming moulds. Clin. Microbiol. Infect. 14, 982–984. doi: 10.1111/j.1469-0691.2008.02086.x
Tampakakis, E., Okoli, I., and Mylonakis, E. (2008). A C. elegans-based, whole animal, in vivo screen for the identification of antifungal compounds. Nat. Protoc. 3, 1925–1931. doi: 10.1038/nprot.2008.193
Tan, M. W., Mahajan-Miklos, S., and Ausubel, F. M. (1999). Killing of Caenorhabditis elegans by Pseudomonas aeruginosa used to model mammalian bacterial pathogenesis. Proc. Natl. Acad. Sci. U.S.A. 96, 715–720. doi: 10.1073/pnas.96.2.715
Tharmalingam, N., Rajmuthiah, R., Kim, W., Fuchs, B. B., Jeyamani, E., Kelso, M. J., et al. (2018). Antibacterial properties of four novel hit compounds from a methicillin-resistant staphylococcus aureus-caenorhabditis elegans high-throughput screen. Microb. Drug Resist. 24, 666–674. doi: 10.1089/mdr.2017.0250
Thompson, T. A., and Brown, P. D. (2017). Association between the agr locus and the presence of virulence genes and pathogenesis in Staphylococcus aureus using a Caenorhabditis elegans model. Int. J. Infect. Dis. 54, 72–76. doi: 10.1016/j.ijid.2016.11.411
Vaezi, A., Fakhim, H., Javidnia, J., Khodavaisy, S., Abtahian, Z., Vojoodi, M., et al. (2018). Pesticide behavior in paddy fields and development of azole-resistant Aspergillus fumigatus: should we be concerned? J. Mycol. Med. 28, 59–64. doi: 10.1016/j.mycmed.2017.12.007
Vallejo, J. A., Beceiro, A., Rumbo-Feal, S., Rodriguez-Palero, M. J., Russo, T. A., and Bou, G. (2015). Optimisation of the Caenorhabditis elegans model for studying the pathogenesis of opportunistic Acinetobacter baumannii. Int. J. Antimicrob. Agents. S0924-8579(15)00241-1. doi: 10.1016/j.ijantimicag.2015.05.021
Van De Veerdonk, F. L., Gresnigt, M. S., Romani, L., Netea, M. G., and Latgé, J.-P. (2017). Aspergillus fumigatus morphology and dynamic host interactions. Nat. Rev. Microbiol. 15, 661–674. doi: 10.1038/nrmicro.2017.90
Wiederhold, N. P., Najvar, L. K., Matsumoto, S., Bocanegra, R. A., Herrera, M. L., Wickes, B. L., et al. (2015). Efficacy of the investigational echinocandin ASP9726 in a guinea pig model of invasive pulmonary aspergillosis. Antimicrob. Agents Chemother. 59, 2875–2881. doi: 10.1128/AAC.04857-14
Keywords: Caenorhabditis elegans, Aspergillus fumigatus, infection model, hyphal filamentation, pathogenicity
Citation: Ahamefule CS, Qin Q, Odiba AS, Li S, Moneke AN, Ogbonna JC, Jin C, Wang B and Fang W (2020) Caenorhabditis elegans-Based Aspergillus fumigatus Infection Model for Evaluating Pathogenicity and Drug Efficacy. Front. Cell. Infect. Microbiol. 10:320. doi: 10.3389/fcimb.2020.00320
Received: 26 March 2020; Accepted: 26 May 2020;
Published: 26 June 2020.
Edited by:
Yong-Sun Bahn, Yonsei University, South KoreaReviewed by:
HEE-Soo Park, Kyungpook National University, South KoreaArianna Tavanti, University of Pisa, Italy
Copyright © 2020 Ahamefule, Qin, Odiba, Li, Moneke, Ogbonna, Jin, Wang and Fang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Bin Wang, YndhbmdAZ3hhcy5jbg==; Wenxia Fang, d2ZhbmdAZ3hhcy5jbg==
 Chukwuemeka Samson Ahamefule
Chukwuemeka Samson Ahamefule Qijian Qin1
Qijian Qin1 Anene N. Moneke
Anene N. Moneke Cheng Jin
Cheng Jin Bin Wang
Bin Wang Wenxia Fang
Wenxia Fang