- Pacific Northwest National Laboratory, Energy and Environment Directorate, Richland, WA, United States
A waste product from plutonium production at Hanford, the radioisotope iodine-129 (129I), is an environmental concern due to its long half-life, mobility, and hazardous potential to humans through bioaccumulation in the thyroid gland. Consequently, understanding the biological mechanisms and contributors to iodine speciation is important in order to increase our knowledge of iodine mobility and the overall risk to human health and the environment, and to evaluate remediation strategies for contaminated areas, as current remediation methods are insufficient and unsustainable. Although iodide (I−) is thermodynamically favored in the geological support material based on existing pH and Eh ranges at the Hanford Site, the dominant species of iodine found in groundwater is iodate (). While microbial activity has been shown to catalyze the oxidation of I−, this process has not been demonstrated by naturally occurring microbes found in the subsurface at the Hanford Site. Four microbial isolates enriched from Hanford groundwater were shown to oxidize I− to molecular iodine (I2) when grown on sugars and organic acids. Glucose proved to be the best substrate for growth, oxidative enzyme production, and I− oxidation. Multi-copper oxidases, such as laccase, have been shown to oxidize I−, and were produced during growth on glucose, xylose, and lactate. These results indicate that bacteria capable of iodide oxidation are present in the Hanford groundwater. Further research will be required to determine whether I2 produced is spontaneously oxidized to .
Introduction
The Hanford Site in Richland, Washington, contains two separate iodine-129 (129I) contaminant plumes (Figure 1), a result of plutonium production during World War II and throughout the Cold War. The two plumes contain average concentrations of 3.5 pCi/L and span an area of over two square miles (the federal drinking water standard for 129I is 1 pCi/L). While 127I is a necessary micronutrient for thyroid hormone production in humans, the radioisotope, 129I, is thought to be potentially toxic through bioaccumulation in the food chain and in the thyroid gland of humans, leading to cancer.
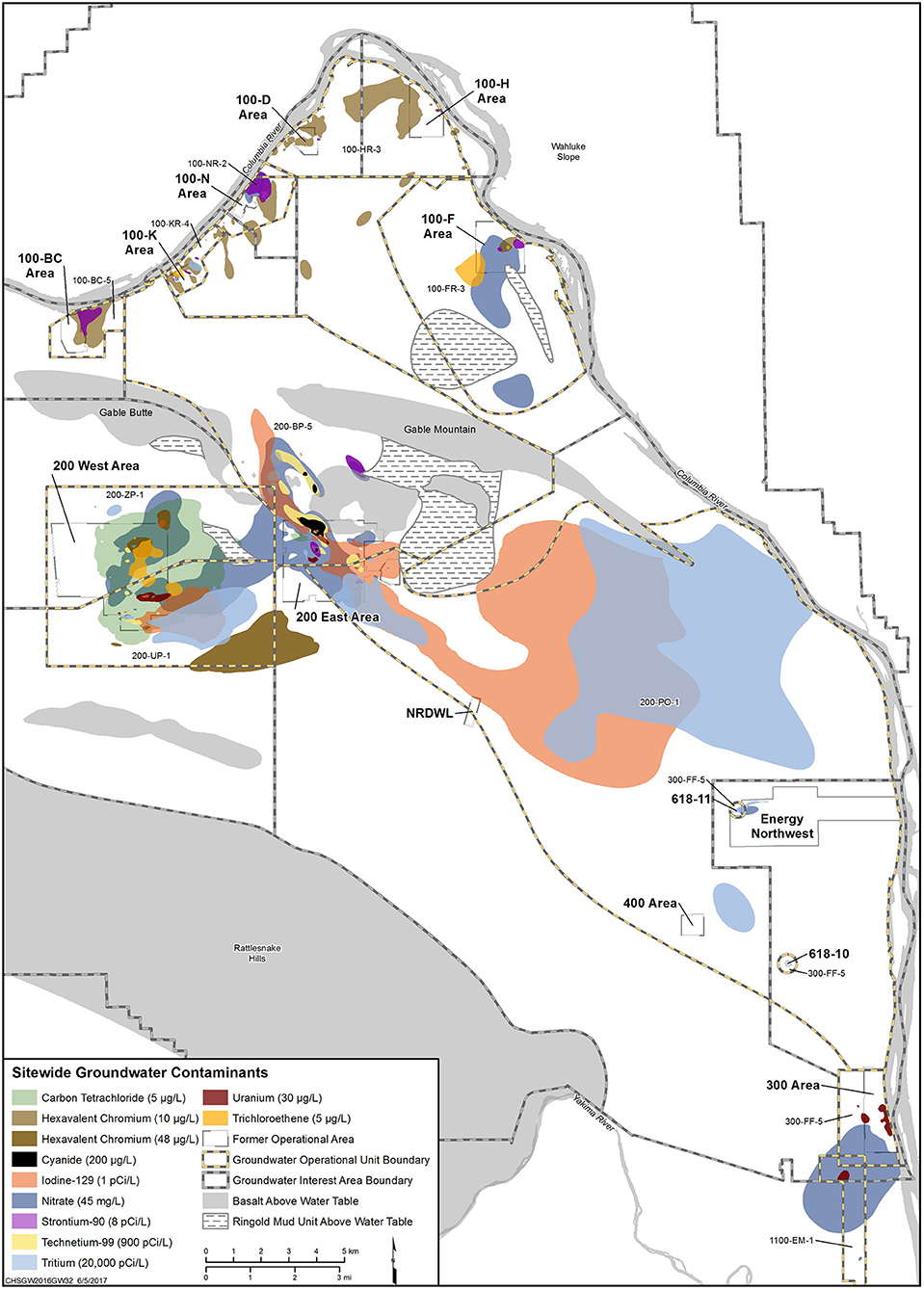
Figure 1. Contaminant plumes emanating from the 200 West and East areas of the Hanford Site. The 129I plume is shown in burnt orange for both sites.
In the environment, iodine predominantly exists in five different forms: iodide (I−), iodine (I2), hypoiodous acid (HIO), iodate (), and organo-iodine species that are often associated with organic compounds such as humic acid. Various reactions of iodine speciation have been summarized by Kaplan et al. (2013). At the 200 Area of the Hanford Site, I− is expected to be the dominant iodine species based on the current ranges of pH and Eh in the complex subsurface geologic environments found on the Central Plateau (Hou et al., 2009). However, through the groundwater sampling of monitoring wells within the 200 West Area and further speciation of these samples, was found to be the dominant species (70.6%), while organo-iodine (25.8%) and I− (3.6%) were found in far lower quantities (Zhang et al., 2013). Microbially-catalyzed reactions impacting iodine cycling in subsurface environments are shown in Figure 2. Bacteria can catalyze the initial step of I− oxidation to I2, which would then spontaneously be oxidized to HOI− and finally .
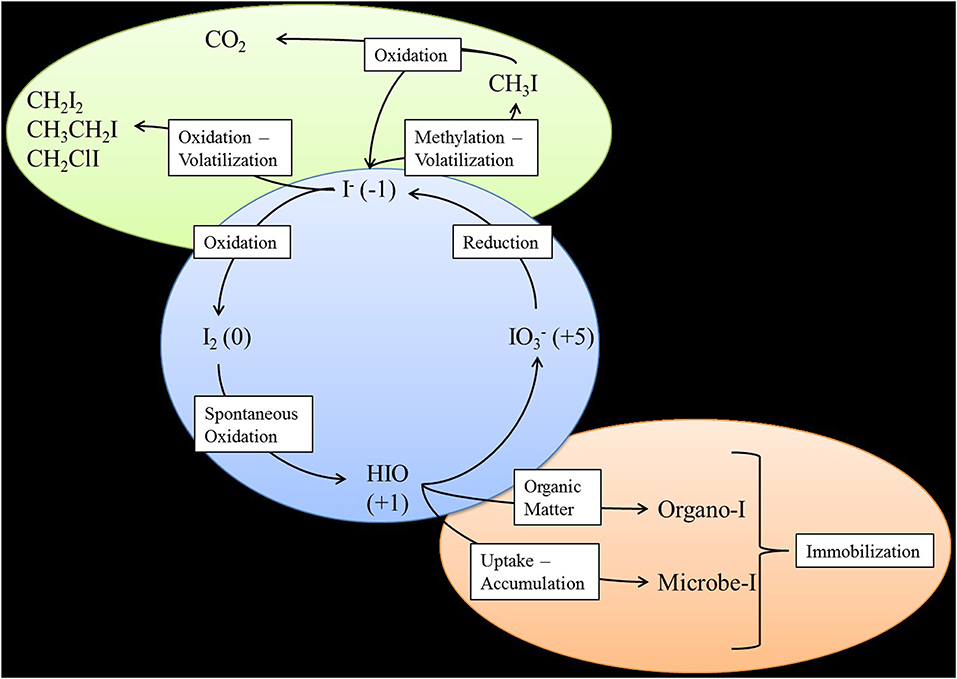
Figure 2. Microbial iodine cycling in the environment. Green ellipse represents the atmosphere. Blue circle represents aqueous environments. Brown ellipse represents the soil or sediment environment.
For decades, bacterial oxidation of I− has been recognized as an important part of the iodine cycle in marine environments (Tsunogai and Sase, 1969). In the early 1970s, scientists observed oxidation of I− to iodine in an aquarium, causing mass fish mortality; they discovered Pseudomonas iodooxidans had an extracellular peroxidase enzyme that, in the presence of polysaccharides, was shown to catalyze the I− oxidation reaction (Gozlan and Margalith, 1973, 1974).
Two primary, and well-studied, environments where iodide-oxidizing bacteria have been found include natural gas brines and marine environments (Amachi et al., 2005b; Žic et al., 2008; Arakawa et al., 2012). Iodine-oxidizing bacteria have also been isolated from corroding pipes at an iodine production facility in China (Wakai et al., 2014). Roseovarius tolerans is an iodide-oxidizing bacterium that was isolated from natural gas brines, where populations of iodide-oxidizing bacteria were up to 105 colony forming units per milliliter of water (Amachi et al., 2005b). The oxidation of I− to molecular iodine was catalyzed by an extracellular enzyme that required oxygen. Two bacteria closely related to R. tolerans have also been isolated from marine systems, and were able to produce free iodine from I− (Fuse et al., 2003). Three Roseovarius species capable of I− oxidation to iodine have also been isolated from both water samples and biofilms collected from a natural gas plant. The marine bacterium Roseobacter species AzwK-3Gb, which has been characterized as a manganese-oxidizing bacterium, has also been shown to oxidize I− to iodine (Li et al., 2014). In the latter study, I− oxidation was catalyzed by production of superoxides in the microbial growth medium.
In addition to brines and marine environments, a number of bacteria were isolated from sediments from a 129I-contaminated aquifer at the Savannah River Site, and were also shown to oxidize I− to iodine. Hydrogen peroxide-dependent I− oxidation was enhanced by the addition of organic acids, indicating that peroxy carboxylic acids were a strong oxidant for I−. As pH was decreased from 9 to 4, H2O2-dependent I− oxidation increased exponentially (Li et al., 2012b).
Production of the enzyme laccase, or other laccase-like multicopper oxidase (LMCO) enzymes by fungi and bacteria, has also been shown to catalyze the oxidation of I− (Kulys et al., 2005; Suzuki et al., 2012; Ihssen et al., 2014). Iodide oxidation by Roseovarius species discussed above is thought to be catalyzed by an LMCO enzyme (Amachi et al., 2005b; Arakawa et al., 2012). An α-Proteobacteria designated strain Q-1 that was isolated from iodide-rich natural gas brine water was shown to oxidize I− to iodine using an oxidase-like enzyme (Suzuki et al., 2012). Molecular and enzymatic characterization of the enzyme showed similarity to multicopper oxidase enzymes. Laccases and other LMCO enzymes have also been shown to catalyze conversion of inorganic iodine to volatile and non-volatile organo-iodine species (Figure 2).
In the current study, bacteria that were enriched from sediment traps that were incubated in groundwater with varying historic 129I concentrations were screened for the ability to oxidize I−. Bacterial isolates demonstrating the highest levels of I− oxidation were assayed for growth on four different carbon sources, production of LMCO via the 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) (ABTS) assay, and I− oxidation as indicated by the formation of molecular iodine.
Materials and Methods
Screening of Microbial Isolates for Iodide Oxidation
Bacteria enriched from materials incubated in 129I plumes at the Hanford Site were screened for the ability to oxidize I− to the more oxidized iodine compounds, such as I2. Isolates were cultured aerobically in modified M9 minimal medium (2.27 mM ammonium sulfate, 0.02 M sodium chloride, 0.2 mM magnesium sulfate heptahydrate, 0.15 g/L yeast extract, 0.012 M phosphate buffer, 3 mL/L Hutner's mineral solution, 0.2% glucose or xylose, pH 7.5) at 30°C and shaken at 130 rpm. Pseudomonas putida strain mt-2 was used as a positive bacterial control, since laccase production has been demonstrated in this bacterium (Ahmad et al., 2010). Iodide oxidation was monitored using the formation of I2 through spectrophotometric analysis of triiodide (see below).
Batch Iodide Oxidation Experiments
Four isolates that demonstrated I− oxidation during screening were further tested to determine the effect of carbon source on I− oxidation, as well as production of multicopper oxidase enzymes previously associated with I− oxidation. Carbon sources tested were 0.2% (w/v) glucose, 0.2% (w/v) xylose, 10 mM acetate, and 10 mM lactate. Cultures were grown overnight (~10–16 h) and used to inoculate batch experiments. Experiments were performed in 125 mL Erlenmeyer flasks with a working volume of 50 mL, with 0.2 mM potassium iodide (KI) as the source of I−. Negative controls contained modified M9 medium without bacteria. Upon inoculation, the flasks were incubated at 30°C and shaken (130 rpm), and samples were taken every 24 h for a total of 96 h.
Sample Analysis
Cell density of the cultures was determined by measuring the optical density using a spectrophotometer set at a wavelength of 600 nm. For other analyses, samples (5 mL) were taken and the culture supernatant was clarified by centrifugation at 6,000 rpm for 10 min, followed by filtration through a 0.2-μm filter membrane. Samples were stored at 4°C until analysis.
Iodide Oxidation Assay
An assay developed to monitor triiodide formation () was used on the sampled culture supernatant from each batch experiment as an indicator of I− oxidation. This method measures iodine (I2) formed from the oxidation of I− by bacteria by adding I− in excess to form triiodide () (Amachi et al., 2005b). Briefly, the assay was prepared by adding the following to a 96-well plate: 50 μl Nanopure H2O (18 mΩ), 20 μl of 1 M phosphate buffer (pH 7.5), 10 μl 0.005 mM hydrogen peroxide (H2O2), 100 μl of culture supernatant, and 20 μl of 100 mM potassium iodide (KI). Positive controls were prepared for the assay using 1 mL of modified M9 medium with 0.2 mM KI and 3.75 mg of a fungal laccase enzyme from Trametes versicolor. The absorbance was read at a wavelength of 353 nm (the maximum absorption of ) after 20 min of incubation on a shaker table at 30°C (Li et al., 2012b). The concentration of iodine (I2) in samples was then estimated using the Beer-Lambert Law using a molecular coefficient for of 25 mM−1cm−1. Addition of assay components resulted in a 2X dilution, which was taken into account during calculations to determine the concentration of I2 in the samples.
Enzyme Assays
To evaluate the presence of LMCO enzymes in correlation to I− oxidation, the 2,2′-azino-bis(3-ethylbenzothiazoline-6-sulphonic acid) (ABTS) assay was performed on the clarified supernatant. ABTS oxidation was monitored using 100 μl of an ABTS solution (0.2 M sodium acetate, 3.6 mM ABTS) mixed with 100 μl of filtered culture supernatant in a 96-well plate and incubated in the dark for 5 min at room temperature. Samples were then read in the spectrophotometer at 420 nm (the maximum absorption of oxidized ABTS). Following the assay, absorbance was converted to concentration of oxidized ABTS using a molecular coefficient of 36 mM−1 cm−1.
Sequencing of 16S rRNA and Phylogenetic Analysis
Bacteria screened for I− oxidation potential were identified by amplifying the 16S rRNA gene using colony PCR and the bacterial consensus primers 8F and 1492R. Colony PCR was performed with an initial denaturation at 95°C for 5 min (to lyse cells and extract DNA), followed by 30 cycles of denaturation at 94°C for 1 min, annealing at 55°C for 1 min, extension at 72°C for 2 min, followed by a final 10 min extension at 72°C. The PCR mixture, generated using Taq PCR Core Kit (www.qiagen.com), contained a small amount of bacterial colony material (template), 1 μL of each primer (25 mM), 5 μL of 10 × PCR buffer, 1 μL of BSA (15 mg/ml), 1 μL of dNTPs (10 mM each), and 0.25 μL Taq DNA polymerase (5 U/μL), in a final reaction volume of 50 μL. PCR products were purified with Qiagen's QIAquick PCR Purification Kit according to the manufacturer's instructions and sequenced by a 3130XL DNA sequencer using primers 8F, 341F, 907R, and 1492R, and assembled accordingly using BioEdit. The obtained 16S rRNA sequence was subjected to BLAST search to determine 16S rRNA similarities with other sequences deposited into GenBank.
Results
Iodide Oxidation Screening
Bacteria screened for I− oxidation (Table 1) are representatives of the classes of bacteria found from sequencing of the 16S rRNA gene. Conditions used for enrichment from the sediments incubated in the groundwater yielded cultivable representatives of these different groups. Bacteria were enriched from sediment traps incubated in a 129I plume beneath the Hanford Site. Isolates from these enrichments were classified via sequencing of the 16S rRNA gene, and affiliation of each strain is shown in Table 1. During an initial screen for I− oxidation capacity, 11 of the 14 isolates tested were able to oxidize I− to I2, as indicated by the formation of triiodide when excess KI was added to the cleared culture supernatant (Figures 3A,B). These results show that a number of both gram-positive and gram-negative bacteria are able to oxidize I−. Two isolates (DVZ18 and DVZ21) were only able to oxidize I− when grown on glucose (Figure 3A). Likewise, two of the nine isolates able to oxidize I−, showed I2 formation when grown on xylose only (DVZ23 and DVZ52) (Figure 3B). The highest levels of I− oxidation were attributed to isolate DVZ2, which is most closely related to Bacillus thuringiensis. Other isolates demonstrating good I− oxidation were DVZ19 (Microbacterium invictum), DVZ21 (Arthrobacter ilicis), and DVZ24 (Pseudomonas mosselii). DVZ24 showed the best I− oxidation when grown on either substrate. Ps. putida strain mt-2 also demonstrated I− oxidation, but levels were much lower than the Hanford isolates. These four bacterial isolates were selected for further analysis in an effort to identify the potential for I− oxidation when supplied with different carbon sources.
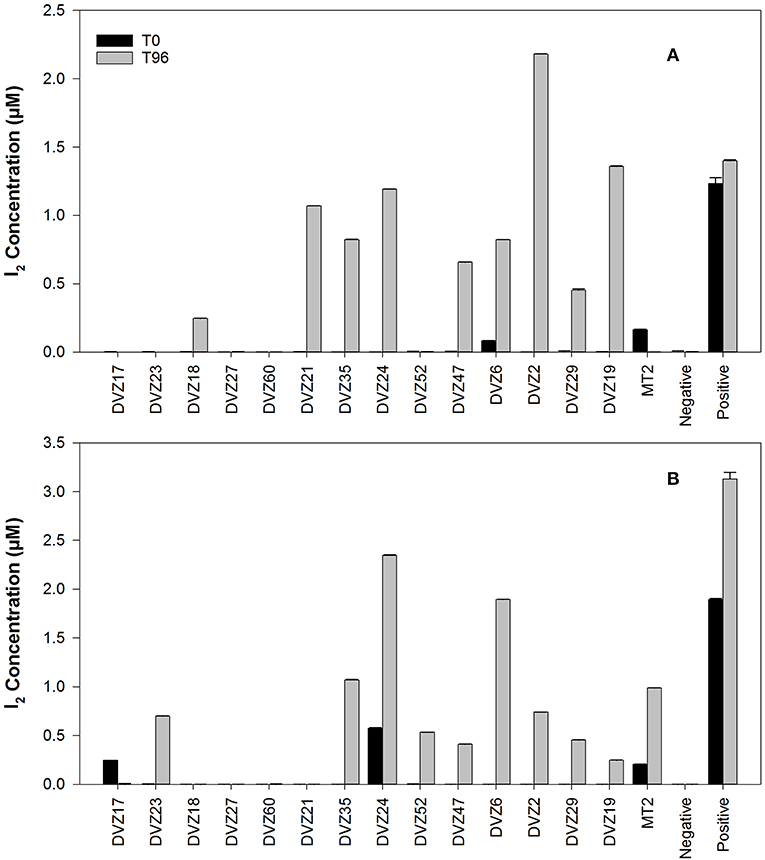
Figure 3. I2 production from batch experiments to determine the ability of various bacteria isolated from Hanford sediment to oxidize I−. Carbon sources: (A) glucose, and (B) xylose.
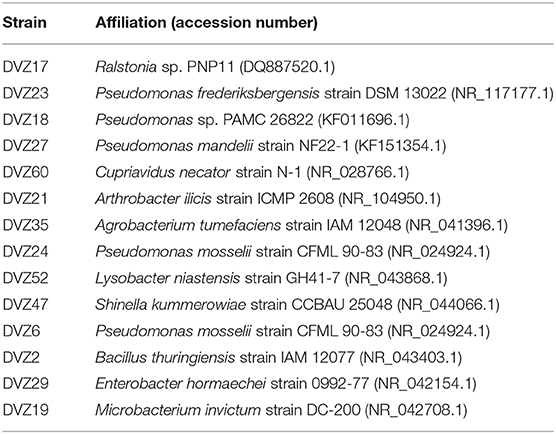
Table 1. Identity of Hanford bacterial isolates that were tested for the ability to oxidize I− to more oxidized iodine species.
Growth of Isolates on Different Carbon Sources
All four Hanford isolates grew at different rates and to different maximum optical densities during the experiment (Figures 4A–D). Optical density was the highest when the isolates were grown on glucose, while growth was similar when the isolates were supplied with xylose or the organic acids, lactate, and acetate. Isolates DVZ2 and DVZ24 showed the shortest lag phases when grown on glucose and transitioned into logarithmic growth within the first 5 h of inoculation when grown on glucose (Figure 4D). Isolate DVZ21 grew to the highest cell density when grown on glucose, while DVZ2 was lowest. Isolate DVZ19 showed the longest lag phase and transitioned into logarithmic growth at around 20 h. All isolates grew to much lower final cell densities when grown on xylose (Figure 4C). As with glucose, DVZ21 and DVZ24 grew the fastest and to the highest cell densities on xylose. Isolate DVZ19 again showed the longest lag phase of the isolates tested. Similar trends in growth were noted when all of the isolates were grown on lactate and acetate (Figures 4A,B). DVZ21 and DVZ24 grew the fastest and produced the highest cell densities on these organic acids, and overall cell densities were less than half compared to when the isolates were grown on glucose.
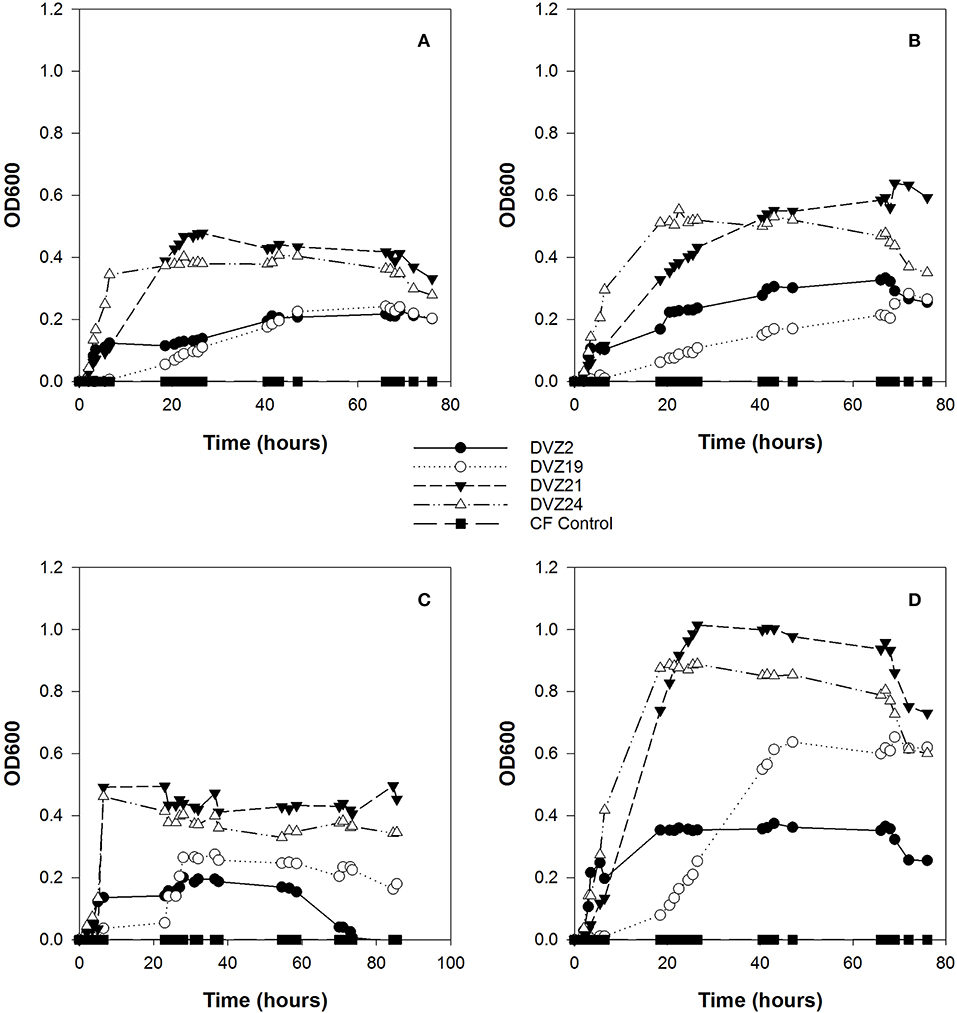
Figure 4. Optical density of Hanford isolates grown on different carbon sources: (A) acetate, (B) lactate, (C) xylose, and (D) glucose.
Iodide Oxidation Assays
Spectrophotometric assays used to monitor the formation of iodine (I2) in batch experiments measured through the formation of triiodide () displayed significantly different results for each of the carbon sources tested (Figures 5A–D). When tested on glucose, all four isolates showed I2 production during the experiment (Figure 5D). DVZ19 was the only isolate that did not show I2 production at every sampling point. A similar trend developed when the isolates were grown on xylose, oxidation was noted in three of the isolates, but not DVZ19 (Figure 5C). Interestingly, I2 formation was not detected until the final two sampling points at 72 and 96 h. When the isolates were grown with lactate as the carbon source, only isolates DVZ2 and DVZ24 were able to oxidize I−, and I2 production was minimal (Figure 5B). Isolates grown on acetate did not show evidence of I− oxidation, since no I2 was measured during the analyses (Figure 5A). While different levels of I2 formation were generated, positive controls using the fungal laccase, indicated that the assay was working as expected.
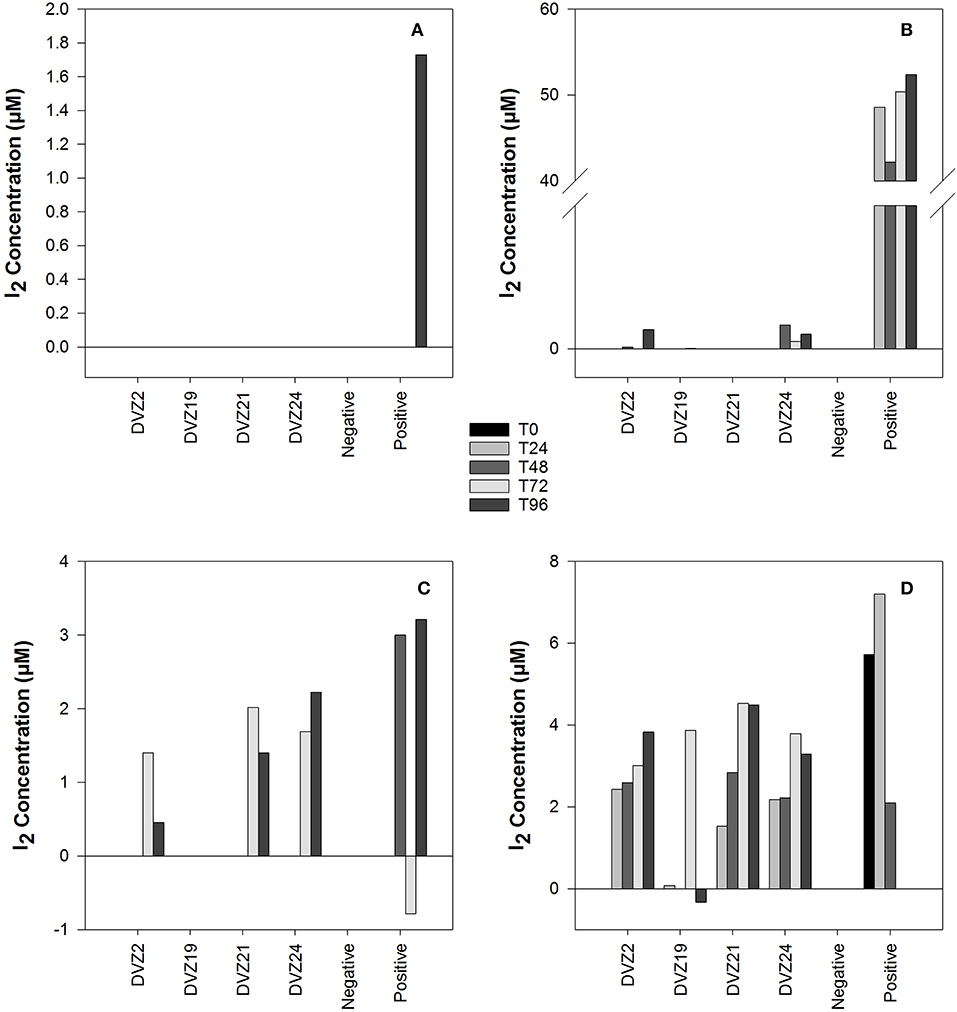
Figure 5. I2 production by Hanford isolates grown on different carbon sources in the presence of I−: (A) acetate, (B) lactate, (C) xylose, and (D) glucose.
ABTS Oxidation
Activity of LMCO, such as laccase, are commonly assayed using the ABTS oxidation assay. This assay was applied to the clarified supernatant from I− oxidation experiments using the four Hanford isolates. ABTS oxidation was the highest when the isolates were grown on glucose (Figure 6D). ABTS was the highest for isolates DVZ21 and DVZ24, maxing out at 3 and 4 M, respectively. Isolate DVZ19 showed low levels of ABTS oxidation over the duration of the test. Isolate DVZ2 did not show any ABTS oxidation during the experiment. ABTS oxidation when the isolates were grown on xylose was substantially lower, never attaining values over 1 M (Figure 6C). When grown on lactate, only DVZ24 was able to maintain prolonged ABTS oxidation; however, levels were still below 1 M (Figure 6B). All isolates appeared to oxidize ABTS at time zero, but not after. As would be expected from the analysis looking at I2 production, with the exception of DVZ19, none of the isolates oxidized ABTS when grown on acetate (Figure 6A).
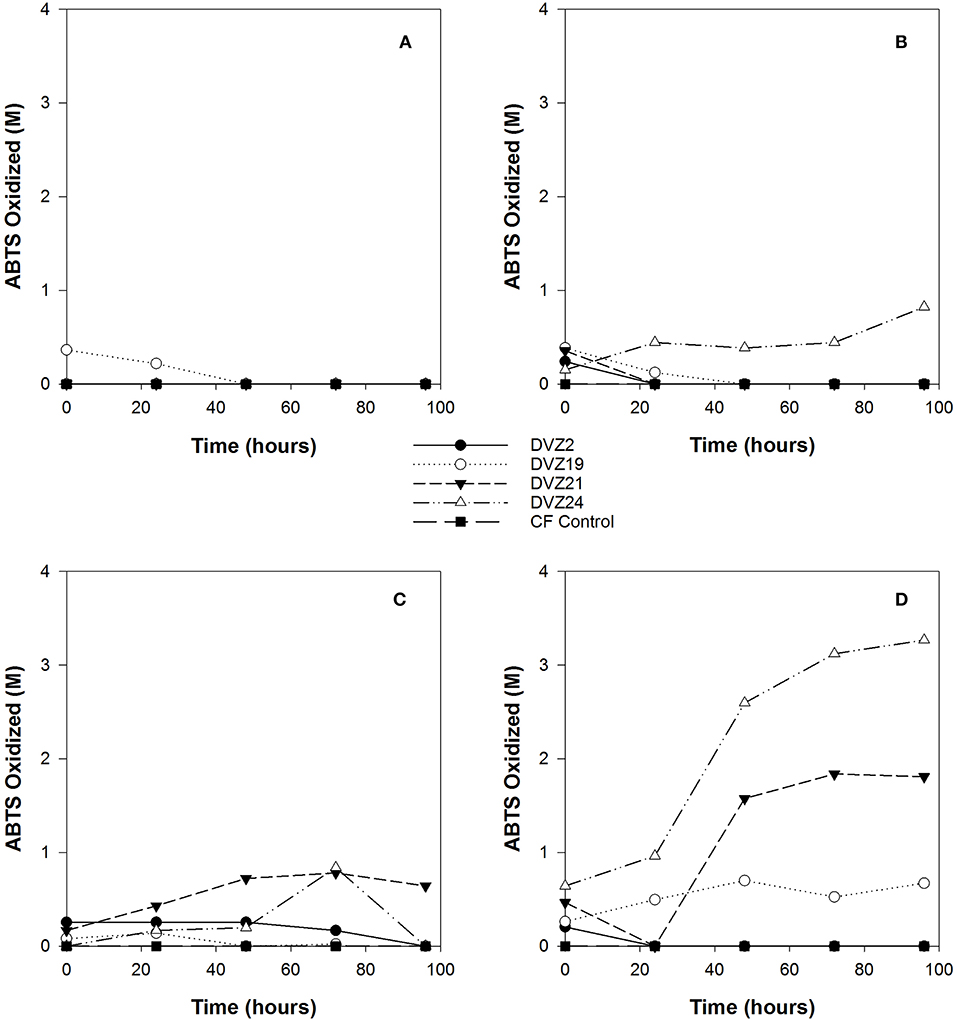
Figure 6. ABTS oxidation by bacteria grown on different carbon sources in the presence of I−. (A) acetate, (B) lactate, (C) xylose, and (D) glucose.
Discussion
Previous work has shown that microbial metabolic activity influences the speciation of iodine through multiple redox reactions (Amachi et al., 2005a,b, 2007; Amachi, 2008; Allard et al., 2009). Eleven of 14 bacterial isolates enriched from sediments incubated in 129I-impacted groundwater showed the ability to oxidize I− to I2, demonstrated by the formation of triiodide when either xylose or glucose was supplied for growth. Four isolates that demonstrated high levels of I− oxidation were selected for additional analyses. In this study, oxidation of I− was observed at different levels for the different isolates and was shown to be dependent on the carbon source used for growth. Glucose was shown to produce higher optical densities and metabolic activity (monitored in growth curves, and both the ABTS and I− oxidation assay, respectively). Higher LMCO (e.g., laccase) activity, which can be qualitatively estimated using ABTS oxidation, was demonstrated for three of the isolates when grown in 0.2% glucose, while lower ABTS oxidation was seen when 0.2% xylose, 10 mM lactate, and 10 mM acetate were used as the carbon source. Laccase-like multicopper oxidase presence and involvement in I− oxidation was evaluated by comparing oxidized ABTS to iodine concentration (μM) in culture supernatant over a 96-h period, which showed strong correlation of LCMO presence to oxidized I−. Isolates that produced the highest concentrations of iodine also oxidized the greatest concentrations of ABTS, suggesting high levels of LMCO present and active in the culture supernatant. The highest increase in oxidation of I- occurred in the first 24–48 h of growth (varying depending on bacterial isolate) and increased slowly until the end of the 96 h incubation, which correlates with the estimated LMCO presence in samples quantified using the ABTS assay. In the first 24–48 h, laccase enzyme is produced exponentially but levels off or decreases after 48 h. Additionally, oxidation of I− does not appear to increase from time 72–96 h, indicating that metabolic activity has slowed with decreased production of enzyme. One exception was for DVZ2, which showed no ABTS oxidation but produced I2. Sequencing of the 16S rRNA gene indicated that DVZ2 was most closely related to Bacillus thuringiensis strain IAM 12077. While the presence of a cell surface localized LMCO was not determined for DVZ2, spore coat proteins, such as CotA in Bacillus subtilis have demonstrated laccase-like activity (Hullo et al., 2001). Since bacteria and spores were removed from the growth medium prior to testing any LMCO activity associated with the cell would have been removed. In contrast to the current study, Li et al. (2012a) showed that the I− oxidation activity for a number of Bacilli was found in the culture supernatant rather than in the crude cell extract.
Determining whether bacteria found in Hanford groundwater plumes were able to oxidize I− was the primary goal of the research. Research findings discussed for the current study show that when provided with adequate carbon, bacteria enriched from Hanford groundwater were able to oxidize I− to I2 which is the first step in the oxidation path which would produce in the aquifer (Figure 2). Additional analyses, to look at speciation of iodine in the samples would be required to prove whether the expected spontaneous oxidation to was occurring following I− oxidation by the Hanford isolates. Groundwater beneath the 200 Area at Hanford is considered oligotrophic and since exogenously added carbon was required for I− oxidation as shown in the current study, the dominance of in the Hanford groundwater would have required an oxidizable carbon source when the waste was disposed, or during transport through the vadose zone to the groundwater. Examples of simultaneous CCl4 disposal with the non-aqueous phase organic compounds such as dibutyl butyl phosphonate, tributyl phosphate, dibutyl phosphate, and machining lard oil in trenches at Hanford have been recorded (Nellis et al., 2009). Carbon tetrachloride contamination in Hanford groundwater is widespread and it may have acted as a solvent carrying these potential carbon sources through the vadose zone and likely into the groundwater. Once in the groundwater, indigenous microbes could have used the compounds for potential carbon compounds producing LCMO which would have potentially oxidized I− in the groundwater, or waste.
Conclusions
Plutonium production at the Hanford Site has led to contamination of groundwater with radionuclides, including 99Tc, U, and 129I. Radioiodine is a long-lived fission product that is mobile in groundwater and represents a hazard to human health and the environment. Speciation of iodine in Hanford groundwater is dominated by , but considering Eh and pH in the subsurface system, speciation should be dominated by I− not . Research was performed to determine whether microbial activity could drive the speciation of iodine currently found in groundwater beneath the Central Plateau of Hanford by oxidizing I− to I2, which is the initial step in the oxidizion pathway to . Demonstration of I2 formation showed that iodine-oxidizing microbes are present in the Hanford groundwater, and that I− oxidation can be stimulated when carbon is added. Findings from these studies represent the first evidence of microbial I− oxidation by bacteria isolated from the Hanford subsurface. Results indicate that if adequate carbon was present when the waste was dispersed to the environment, that bacteria present in the aquifer would have been able to grow, produce the enzymes necessary for I− oxidation, and convert the I− present in the waste to I2.
While the current study showed the potential for I− oxidation by Hanford bacterial isolates, future studies should include quantification of I−, , and any organo-iodine compounds formed. Experiments should also be performed looking at the effect of carbon sources found in the original waste on oxidation of I−. These experiments should be performed using saturated and unsaturated conditions, since some oxidation of the I− to I2 and then may have occurred as the waste migrated through the vadose zone.
Author Contributions
Experimental design and lab work was led by BL and MH. Field deployment of sediment traps was led by DS. Laboratory experiments were performed by EM and SB. BL, MH, and EM interpreted the data and wrote the paper.
Funding
This document was prepared by the Deep Vadose Zone Applied Field Research Initiative at Pacific Northwest National Laboratory. Funding for this work was provided by the U.S. Department of Energy Office of Environmental Management and Richland Operations Office. Pacific Northwest National Laboratory is operated by Battelle Memorial Institute for the U.S. Department of Energy under Contract DE-AC05-76RL01830.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
Ahmad, M., Taylor, C. R., Pink, D., Burton, K., Eastwood, D., Bending, G. D., et al. (2010). Development of novel assays for lignin degradation: comparative analysis of bacterial and fungal lignin degraders. Mol. Biosyst. 6, 815–821. doi: 10.1039/b908966g
Allard, S., Von Gunten, U., Sahli, E., Nicolau, R., and Gallard, H. (2009). Oxidation of iodide and iodine on birnessite (δ-MnO2) in the pH range 4–8. Water Res. 43, 3417–3426. doi: 10.1016/j.watres.2009.05.018
Amachi, S. (2008). Microbial contribution to global iodine cycling: volatilization, accumulation, reduction, oxidation, and sorption of iodine. Microb. Environ. 23, 269–276. doi: 10.1264/jsme2.ME08548
Amachi, S., Fujii, T., Shinoyama, H., and Muramatsu, Y. (2005a). Microbial influences on the mobility and transformation of radioactive iodine in the environment. J. Nucl. Radiochemical Sci. 6, 21–24. doi: 10.14494/jnrs2000.6.21
Amachi, S., Kawaguchi, N., Muramatsu, Y., Tsuchiya, S., Watanabe, Y., Shinoyama, H., et al. (2007). Dissimilatory iodate reduction by marine Pseudomonas sp. strain SCT. Appl. Environ. Microbiol. 73, 5725–5730. doi: 10.1128/AEM.00241-07
Amachi, S., Muramatsu, Y., Akiyama, Y., Miyazaki, K., Yoshiki, S., Hanada, S., et al. (2005b). Isolation of iodide-oxidizing bacteria from iodide-rich natural gas brines and seawaters. Microb. Ecol. 49, 547–557. doi: 10.1007/s00248-004-0056-0
Arakawa, Y., Akiyama, Y., Furukawa, H., Suda, W., and Amachi, S. (2012). Growth stimulation of iodide-oxidizing α-Proteobacteria in iodide-rich environments. Microb. Ecol. 63, 522–531. doi: 10.1007/s00248-011-9986-5
Fuse, H., Inoue, H., Murakami, K., Takimura, O., and Yamaoka, Y. (2003). Production of free and organic iodine by Roseovarius spp. FEMS Microbiol. Lett. 229, 189–194. doi: 10.1016/S0378-1097(03)00839-5
Gozlan, R., and Margalith, P. (1974). Iodide oxidation by Pseudomonas iodooxidans. J. Appl. Bacteriol. 37, 493–499. doi: 10.1111/j.1365-2672.1974.tb00474.x
Gozlan, R. S., and Margalith, P. (1973). Iodide oxidation by a marine bacterium. J. Appl. Bacteriol. 36, 407–417. doi: 10.1111/j.1365-2672.1973.tb04122.x
Hou, X., Hansen, V., Aldahan, A., Possnert, G., Lind, O. C., and Lujaniene, G. (2009). A review on speciation of iodine-129 in the environmental and biological samples. Anal. Chim. Acta 632, 181–196. doi: 10.1016/j.aca.2008.11.013
Hullo, M.-F., Moszer, I., Danchin, A., and Martin-Verstraete, I. (2001). CotA of Bacillus subtilis is a copper-dependent laccase. J. Bacteriol. 183, 5426–5430. doi: 10.1128/JB.183.18.5426-5430.2001
Ihssen, J., Schubert, M., Thöny-Meyer, L., and Richter, M. (2014). Laccase catalyzed synthesis of iodinated phenolic compounds with antifungal activity. PLoS ONE 9:e89924. doi: 10.1371/journal.pone.0089924
Kaplan, D. I., Denham, M. E., Zhang, S., Yeager, C., Xu, C., Schwehr, K. A., et al. (2013). Radioiodine biogeochemistry and prevalence in groundwater. Crit. Rev. Environ. Sci. Technol. 44, 2287–2335. doi: 10.1080/10643389.2013.828273
Kulys, J., Bratkovskaja, I., and Vidziunaite, R. (2005). Laccase-catalysed iodide oxidation in presence of methyl syringate. Biotechnol. Bioeng. 92, 124–128. doi: 10.1002/bit.20610
Li, H.-P., Brinkmeyer, R., Jones, W. L., Zhang, S., Xu, C., Ho, Y.-F., et al. (2012a). “Iodide oxidizing activity of bacteria from subsurface sediments of the Savannah River Site, SC, USA,” in Interdisciplinary Studies on Environmental Chemistry, eds M. Kawaguchi, K. Misaki, H. Sato, T. Yokokawa, T. Itai, T. M. Nguyen, J. Ono, and S. Tanabe (Tokyo: TERRAPUB), Vol. 6, 89–97.
Li, H.-P., Daniel, B., Creeley, D., Grandbois, R., Zhang, S., Xu, C., et al. (2014). Superoxide production by a manganese-oxidizing bacterium facilitates iodide oxidation. Appl. Environ. Microbiol. 80, 2693–2699. doi: 10.1128/AEM.00400-14
Li, H.-P., Yeager, C. M., Brinkmeyer, R., Zhang, S., Ho, Y.-F., Xu, C., et al. (2012b). Bacterial production of organic acids enhances H2O2-dependent iodide oxidation. Environ. Sci. Technol. 46, 4837–4844. doi: 10.1021/es203683v
Nellis, S. R., Yoon, H., Werth, C. J., Oostrom, M., and Valocchi, A. J. (2009). Surface and interfacial properties of nonaqueous-phase liquid mixtures released to the subsurface at the Hanford Site. Vadose Zone J. 8, 343–351. doi: 10.2136/vzj2008.0104
Suzuki, M., Eda, Y., Ohsawa, S., Kanesaki, Y., Yoshikawa, H., Tanaka, K., et al. (2012). Iodide oxidation by a novel multicopper oxidase from the alphaproteobacterium strain Q-1. Appl. Environ. Microbiol. 78, 3941–3949. doi: 10.1128/AEM.00084-12
Tsunogai, S., and Sase, T. (1969). Formation of iodide-iodine in the ocean. Deep Sea Res. Oceanogr. Abstr. 16, 489–496. doi: 10.1016/0011-7471(69)90037-0
Wakai, S., Ito, K., Iino, T., Tomoe, Y., Mori, K., and Harayama, S. (2014). Corrosion of iron by iodide-oxidizing bacteria isolated from brine in an iodine production facility. Microb. Ecol. 68 519–527. doi: 10.1007/s00248-014-0438-x
Zhang, S., Xu, C., Creeley, D., Ho, Y.-F., Li, H.-P., Grandbois, R., et al. (2013). Iodine-129 and iodine-127 speciation in groundwater at the Hanford Site, U.S.: iodate incorporation into calcite. Environ. Sci. Technol. 47, 9635–9642. doi: 10.1021/es401816e
Keywords: iodide oxidation, bacteria, Hanford, bioremediation, groundwater
Citation: Lee BD, Moser EL, Brooks SM, Saunders DL and Howard MH (2020) Microbial Contribution to Iodine Speciation in Hanford's Central Plateau Groundwater: Iodide Oxidation. Front. Environ. Sci. 7:145. doi: 10.3389/fenvs.2019.00145
Received: 20 August 2018; Accepted: 12 September 2019;
Published: 09 April 2020.
Edited by:
Prosun Bhattacharya, Royal Institute of Technology, SwedenReviewed by:
Prabhakar Sharma, Nalanda University, IndiaChen Xu, Texas A&M University, United States
Copyright © 2020 Lee, Moser, Brooks, Saunders and Howard. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Brady D. Lee, QnJhZHkuTGVlQHNybmwuZG9lLmdvdg==
†Present address: Brady D. Lee, Savannah River National Laboratory, Environmental, Materials, and Energy Sciences Directorate, Aiken, SC, United States
Erin L. Moser, Quinnipiac University School of Health Sciences, University of Michigan, Ann Arbor, MI, United States
 Brady D. Lee
Brady D. Lee Erin L. Moser†
Erin L. Moser†