- 1Key Laboratory of East China Sea Fishery Resources Exploitation, Ministry of Agriculture, East China Sea Fisheries Research Institute, Chinese Academy of Fishery Sciences, Shanghai, China
- 2Guangdong Provincial Key Laboratory of Marine Biotechnology, Shantou University, Shantou, China
Scavenger receptors are cell surface membrane-bound receptors that typically bind multiple ligands and promote the removal of endogenous proteins and pathogens. In this study, we characterized a novel scavenger receptor-like protein, namely, SpBark. SpBark was upregulated in hemocytes after challenges with bacteria, suggesting that it might be involved in antibacterial defense. SpBark is a type I transmembrane protein with four extracellular domains, including three scavenger receptor cysteine-rich domains (SRCRDs) and a C-type lectin domain (CTLD). Western blot assay showed that SpBark CTLD possessed a much stronger binding activity to tested microbes than the three SRCRDs. It also exhibited apparent binding activities to lipopolysaccharide (LPS) and acetylated low-density lipoprotein (ac-LDL), whereas the other SRCRDs showed much lower or no binding activities to these components. Agglutination activities were observed in the presence of Ca2+ by incubating microorganisms with SpBark CTLD instead of SRCRDs. These results suggested that SpBark CTLD was the major binding site for ac-LDL and LPS. Coating Vibrio parahemolyticus with SpBark CTLD promoted bacterial clearance in vivo. This finding indicated that SpBark might participate in the immune defenses against Gram-negative bacteria through a certain mechanism. The promotion of bacterial clearance by SpBark was further determined using SpBark-silenced crabs injected with V. parahemolyticus. SpBark knockdown by injection of SpBark dsRNA remarkably suppressed the clearance of bacteria in hemolymph. Meanwhile, it also severely restrained the phagocytosis of bacteria. This finding suggested that SpBark could modulate the phagocytosis of bacteria, and the promotion of bacterial clearance by SpBark was closely related to SpBark-mediated phagocytosis activity. The likely mechanism of bacterial clearance mediated by SpBark was as follows: SpBark acted as a pattern recognition receptor, which could sense and bind to LPS on the surface of invading bacteria with its CTLD in hemolymph. The binding to LPS made the bacteria adhere to the surface of hemocytes. This process would facilitate phagocytosis of the bacteria, resulting in their removal. This study provided new insights into the hemocyte phagocytosis mechanisms of invertebrates and the multiple biological functions of Bark proteins.
Introduction
Due to the lack of adaptive immunity, invertebrates rely on innate immunity to resist microbial invasion (1). Pattern recognition receptors (PRRs) participate in host immune defense by binding to specific ligands, resulting in the occurrence of innate immune responses (2, 3). These PRRs comprise structurally diverse domains that can recognize different pathogens by sensing specific pathogen-associated molecular patterns, including microbial polysaccharides, glycolipids, lipoproteins, nucleotides, and nucleic acids. To date, various PRRs, such as C-type lectins, Toll-like receptors, and scavenger receptors, have been identified in many invertebrates and vertebrate species (4–7).
Scavenger receptors are cell surface receptors that typically bind multiple ligands and promote the removal of non-self or altered-self targets (8). These proteins constitute a “superfamily” of membrane-bound receptors that were initially thought to bind and internalize modified low-density lipoprotein (mLDL). They were later found to play a role in binding to various ligands, including endogenous proteins and pathogens (9, 10). Scavenger receptors are currently proposed to be categorized into 12 classes on the basis of their structural diversities and biological functions (11). Each member in different classes exhibits a similar structure, including the intracellular portion, transmembrane domain, and extracellular region (10). The extracellular region always contains one or more functional domains, such as collagenous domain, scavenger receptor cysteine-rich domain (SRCRD), and C-type lectin domain (CTLD), thereby exhibiting a wide range of biological functions.
SRCRDs are present in a variety of host defense-related proteins, including several classes of scavenger receptors (11–14). They are ~100–110 amino acids long with 6–8 conserved cysteines predicted to form 3–4 disulfide bonds. SRCRD-containing proteins can bind a variety of ligands, including mLDL, structure proteins of certain viruses, bacteria, lipopolysaccharides (LPS), lipoteichoic acid (LTA), apoptotic cells, and certain acute phase reactants (15–19). Furthermore, some SRCRD-containing scavenger receptors have been validated as versatile proteins mediating diverse immune responses (20–22).
Most CTLD-containing proteins participate in immunity on the basis of non-self-recognition and binding abilities, which are accomplished by the characteristic CTLDs (7, 23). The structure of CTLDs is maintained by relying on four conserved cysteine residues, which form two disulfide bridges (24). Certain CTLD-containing scavenger receptors play a role in immune defense. LOX-1 and dectin-1 are two scavenger receptors of class E. They are type II transmembrane proteins with CTLDs displaying a scavenger receptor activity (25, 26). Dectin-1 receptor recognizes various bacterial, fungal, and plant carbohydrates. LOX-1 has been implicated in recognizing other ligands, including apoptotic cells and bacteria. The mouse SRCL (belonging to class SR-A4) is a bacteria-binding receptor containing a CTLD that plays a role in host defense (27).
Currently, an increasing number of scavenger receptors and scavenger receptor-like proteins with diverse structures have been characterized in invertebrates. Some of them were demonstrated to participate in immune defense. Drosophila Croquemort and DmSR-CI are the class B and C members of the scavenger receptor family, respectively. Both of them are involved in the phagocytosis of bacteria (15, 28). Another insect scavenger receptor TmSR-C plays a pivotal role in phagocytosing fungi and bacteria (29). Cf SR, a mollusk SRCRD-containing scavenger receptor, can bind to several kinds of microbial polysaccharides serving as a versatile PRR involved in immune recognition (16). In crustaceans, several scavenger receptors were identified as PRRs participating in antibacterial responses by enhancing phagocytosis of bacteria (30–32). Different from these identified scavenger receptors involved in immunity, a putative transmembrane scavenger receptor-like protein named Bark beetle (Bark) was demonstrated to participate in epithelial cell adhesion and mounting of a core complex of septate junctions (33). To date, few reports demonstrate that Bark and Bark-like proteins play a role in immune defense even though their extracellular regions contain defense-related domains, such as SRCRD and CTLD.
In this study, we identified a Bark-like protein (SpBark) in an invertebrate animal (mud crab), Scylla paramamosain. This protein shares a similar structure with Drosophila bark protein possessing three SRCR domains and a CTLD in the extracellular region. To determine whether SpBark could function as a PRR involved in antibacterial immunity, the binding activities of its four domains to mLDL and microbial polysaccharides were tested, and SpBark-mediated bacterial clearance activity in vivo was also investigated. This study demonstrates the role of SpBark in antibacterial defense and provides new insights into the diverse biological functions of Bark-like proteins.
Materials and Methods
Reagents and Chemicals
Taq DNA polymerases, RNAiso Plus, and First Strand cDNA Synthesis Kit were obtained from TaKaRa (Dalian, China). LPS from Escherichia coli 0111:B4 and peptidoglycan (PGN) and LTA from Staphylococcus aureus were obtained from Sigma (St. Louis, MO, USA). Trehalose from Saccharomyces cerevisiae was purchased from Amresco (Solon, OH, USA).
Immune Challenges of Mud Crab and Tissue Collection
Mud crabs (~150 g each) purchased from an aquaculture farm in Chongming Country (Shanghai, China) were acclimated in 400 L tanks with aerated seawater for a week (mud crabs are not endangered or protected species, and no special permissions are required). Healthy crabs were randomly selected to investigate the tissue distribution and expression profiles of SpBark. For immune stimulation, Staphylococcus aureus and Vibrio parahemolyticus were cultured overnight in Luria–Bertani medium, collected by centrifugation, and re-suspended in PBS (10 mM Na2HPO4, 1.8 mM KH2PO4, 140 mM NaCl, and 2.7 mM KCl; pH 7.3). After washing twice with PBS, the resultant suspensions were used as bacterial inocula. Subsequently, 200 μL of each inoculum (2 × 107 CFU in PBS) was injected into the base of the right fifth leg of each crab. The corresponding control was challenged with the same volume of PBS. At each time point after injection (0, 2, 6, 12, 24, 48, and 72 h), hemolymph was drawn from the base of the legs by using a syringe preloaded with ice-cold anticoagulant buffer (0.45 M NaCl, 0.1 M glucose, 30 mM trisodium citrate, 26 mM citric acid, and 10 mM ethylenediaminetetraacetic acid; pH 4.6) (34). The hemocyte pellets were collected for RNA extraction after centrifugation at 800 × g for 10 min at 4°C. In addition, other tissues, including the heart, gills, hepatopancreas, stomach, intestine, connective tissue, and muscle, of healthy crabs were dissected; washed with sterile PBS; and collected for RNA extraction. For each tissue sample, at least three crabs were used to eliminate individual differences. Another two batches of previously isolated RNA samples were used to eliminate the differences among batches. All animal experimental procedures were conducted in accordance with the National Institutes of Health's Guide for the Care and Use of Laboratory Animals.
Total RNA Isolation and First Strand cDNA Synthesis
The total RNA from hemocytes and other collected tissues was isolated with RNAiso Plus, and DNase I (Promega, USA) was added into the extracted total RNA to remove the contaminated DNA. The first stand cDNA was synthesized using the First Strand cDNA Synthesis Kit (TaKaRa, Dalian) in accordance with the manufacturer's instructions.
SpBark cDNA Cloning
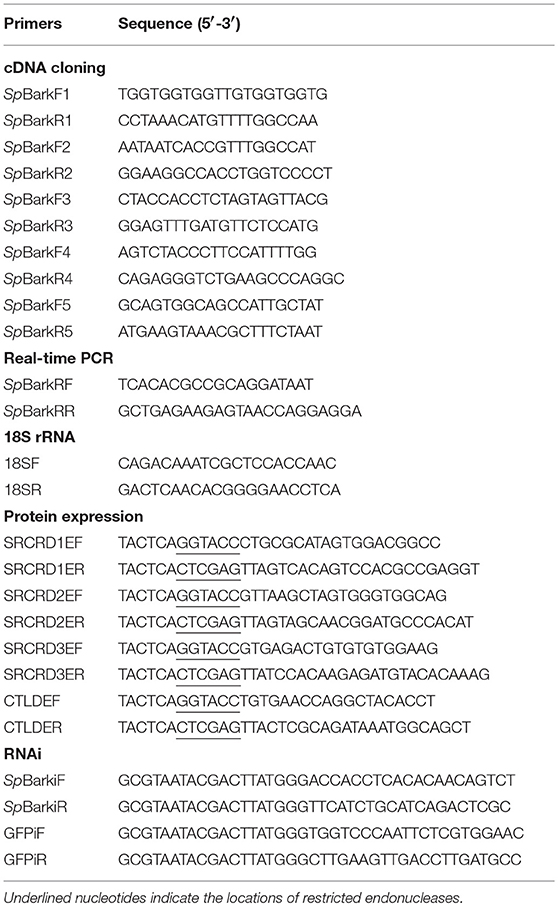
The original cDNA sequence of SpBark was obtained through high-throughput transcriptome sequencing with an RNA mixture isolated from the hemocytes, gill, and hepatopancreas, and it was further confirmed by PCR with five pairs of primers (Table 1). The PCR was performed under the following parameters: 95°C for 5 min; 35 cycles of 94°C for 30 s, 55°C for 30 s, and 72°C for 60 s; and finally 72°C for 10 min. The DNA products were separated by agarose gel electrophoresis, subcloned into a pMD-19T vector, and sequenced via a commercial company (Sangon, China).
Bioinformatics Analyses
The identities of SpBark with other Bark-like proteins were revealed using the online Basic Local Alignment Search Tool Program (BLASTP) (http://blast.ncbi.nlm.nih.gov/Blast.cgi). The domain architecture was predicted using simple modular architecture research tool (SMART) (http://smart.embl-heidelberg.de). Multiple alignment was generated through the ClustalX 2.0 program (http://www.ebi.ac.uk/tools/clustalw2) and GENEDOC software. The putative amino acid sequence was deduced and predicted on http://web.expasy.org/translate/. The signal peptide was predicted with SignalP (http://www.cbs.dtu.dk/services/SignalP/). The pI and molecular weight (MW) were calculated on http://web.expasy.org/compute_pi/. MEGA 7.0 was used to construct a neighbor-joining tree, and the bootstrap of 1,000 was selected to assess reliability.
Quantitative Real-Time PCR (qRT-PCR)
qRT-PCR was conducted to investigate the relative expression level of SpBark in a real-time thermal cycler Quantstudio 6 Flex (ABI, USA) according to a previous protocol (35). A pair of primers (SpBarkRF and SpBarkRR) was designed to generate an amplicon of SpBark. Another pair of primers (18SRF and 18SRR) was synthesized to produce a 121-bp fragment of 18S rRNA as reference. qRT-PCR was programed as follows: an initial denaturation at 94°C for 3 min; 40 cycles of 94°C for 10 s and 60°C for 1 min; and finally a melting curve analysis from 65 to 95°C. The total volume of reaction mixture was 20 μL (10 μL of 2 × SYBR Premix Ex Taq, 2 μL of cDNA, and 4 μL of each primer). All tests were performed thrice with individual templates. The algorithm of 2−ΔCT was used to investigate tissue distribution of SpBark. To analyze the time-course expression profiles, the formula of 2−ΔΔCT was used to normalize the data in two steps. First, the expression of SpBark was normalized to the reference gene (18S rRNA) in the same sample. Second, the relative expression of SpBark in the experimental sample was normalized to that in the control sample. Unpaired Student's t-test was used to analyze significant differences (*P < 0.05; **P < 0.01).
Recombinant Expression and Purification
To explore the potential immune function of SpBark, four predicted functional domains (SRCRD1, SRCRD2, SRCRD3, and CTLD) were overexpressed using an E. coli expression system. On the basis of the SpBark cDNA sequence, four primer pairs (SRCRD1EF and SRCRD1RF, SRCRD2EF and SRCRD2ER, SRCRD3EF and SRCRD3ER, and CTLDEF and CTLDER) were designed to produce DNA fragments encoding SRCRD1, SRCRD2, SRCRD3, and CTLD, respectively (Table 1). These DNA fragments were digested and then inserted into the pET32a expression vectors. The constructed plasmids were transformed into competent E. coli cells for overexpression. Recombinant proteins were induced by adding isopropyl-β-d-thiogalactoside (IPTG) to a final concentration of 0.5 mM at 37°C for 6 h and purified using High Affinity Ni-NTA Resin (GenScript, Nanjing) in accordance with the manufacturer's instructions.
Microorganism-Binding Assay
Nine kinds of microorganisms, including four Gram-negative bacteria (Vibrio harveyi, V. parahemolyticus, V. alginolyticus, and E. coli), three Gram-positive bacteria (S. aureus, Bacillus subtilis, and B. megaterium), and two fungi (Candida albicans and Pichia pastoris), were used to investigate the microorganism-binding activities of SRCRD1, SRCRD2, SRCRD3, and CTLD. The experiment was performed following a protocol used in our previous study (36). In brief, the microorganisms (1 × 108 CFU) were incubated in 1 mL of each recombinant protein (200 μg/mL) with gentle rotation for 30 min at room temperature. The microorganisms were pelleted, washed four times with 1 mL of TBS (150 mM NaCl, 10 mM Tris–HCl, pH 7.5), and eluted with 200 μL of 7% SDS solution with mild agitation for 5 min at room temperature. The microorganism pellets were rewashed thrice with 1 mL TBS. Finally, the elution and pellet of each microorganism were sampled and analyzed on 12.5% SDS-PAGE. The recombinant proteins (SRCRD1, SRCRD2, SRCRD3, and CTLD) were sampled as the positive controls. The binding activity to microorganisms was determined by Western blot. After the proteins were transferred onto a PVDF membrane, the membrane was blocked with 4% bovine serum albumin (BSA) in TBS and incubated with peroxidase-conjugated monoclonal antibody against His-tag (GenScript, Nanjing). The target proteins were visualized with an ECL Western blot reagent kit.
Binding Activity of Recombinant Proteins to Microbial Polysaccharides
Enzyme-linked immunosorbent assay (ELISA) was performed to test the binding abilities of four domain proteins (SRCRD1, SRCRD2, SRCRD3, and CTLD) to microbial polysaccharides and acetylated LDL (ac-LDL) following a previously described protocol with slight modifications (37). Microbial polysaccharides, including PGN and LTA from S. aureus, LPS from E. coli 0111:B4, and β-glucan from Laminaria digitata, together with human ac-LDL were used in this study. Each well of the microtiter plate was incubated with 100 μL of ac-LDL (50 μg/mL) at 4°C overnight or microbial polysaccharide (20 μg/mL) at 37°C until the plate came to desiccation. Wells incubated with 100 μL of distilled water were employed as blank control. Each well was blocked with BSA (1 mg/mL, 200 μL) at 37°C for 2 h and washed four times with TBST (0.05% Tween 20 in TBS). Subsequently, a series of diluted recombinant protein SRCRD1, SRCRD2, SRCRD3, CTLD, or TRX (0–5 μM in TBS containing 0.1 mg/mL BSA) was added to polysaccharide-coated plates. TRX was used as negative control. The serially diluted recombinant proteins supplemented with 5 mM of CaCl2 were added to ac-LDL-coated plates. After incubation at room temperature for 3 h, the plates were rinsed with TBST four times. Each well was then incubated with 100 μL of peroxidase-conjugated mouse monoclonal anti-His antibody (1:5,000 dilution in TBS with 0.1 mg/mL BSA) at 37°C for 2 h. The color reaction was developed with 0.01% 3,3′,5,5′-tetramethylbenzidine (Sigma, USA) and stopped by adding 2 M H2SO4. The absorbance was read at 405 nm on a Spark 10 M microplate reader (Tecan, Switzerland). The assay was conducted in triplicate. For a competitive binding assay, purified recombinant CTLD was diluted to a final concentration of 0.25 μM and pre-incubated with different LPS concentrations (0.0005, 0.0025, 0.005, 0.025, 0.05, 0.25, 0.5, and 2.5 μM) by gentle shaking at room temperature for 2 h (38). Subsequently, the pre-incubated CTLD protein (100 μL per well) was added to the plates coated with LPS. The following steps were the same as those in ELISA above. The competitive binding assay was repeated three times.
Agglutination Assay
The microorganisms used in microorganism-binding assay were applied for agglutination assay following a method described by (36). Microorganisms were harvested at the mid-logarithmic growth phase by centrifugation at 5,000 × g for 5 min, re-suspended in TBS after washing three times, and finally adjusted to 2 × 108 cells/mL (for bacteria) or 2 × 107 cells/mL (for fungi). The microorganism suspensions were mixed with equal volume (100 μL) of diluted recombinant protein (SRCRD1, SRCRD2, SRCRD3, CTLD, or TRX) in TBS at the protein concentration range of 0.48–15.2 μM with or without 10 mM CaCl2 at 28°C for 30 min. TRX tag protein served as the negative control. Agglutination was observed under a light microscope.
RNA Interference
A partial SpBark DNA fragment, which covered the SpBark CTLD, was amplified by PCR with specific primers linked to the T7 promoter (Table 1) and used as the template to generate dsSpBark (SpBark dsRNA) with T7 RiboMAX™ Express RNAi System (Promega, USA). The dsGFP (EGFP dsRNA) was synthesized as control with primers listed in Table 1. To obtain better RNA interference efficiency, the crabs with an average weight of ~25 g were selected and divided into two groups (four crabs in each group). Each crab in the experimental group was injected with 25 μg of dsSpBark into the base of the fourth leg. The crabs in the control group were injected with an equal amount of dsGFP. A second dsRNA injection was performed 24 h later in the same way. Hemocytes were collected from each group at 40 and 48 h after the first dsRNA injection. RNAi efficiency was assessed by qRT-PCR using the total RNAs from hemocytes with a pair of primers located outside the above DNA fragment (Table 1). Unpaired t-test was used to analyze significant differences (*P < 0.05; **P < 0.01).
Bacterial Clearance Assay
V. parahemolyticus was incubated with recombinant CTLD protein and then injected into crabs to determine whether coating bacteria with CTLD could influence bacterial clearance. Approximately 500 μL of CTLD protein in PBS (400 μg/mL) was mixed with an equal volume of V. parahemolyticus (1 × 108 CFU) with gentle rotation at 28°C for 60 min. The same number of V. parahemolyticus incubated with TRX tag protein (400 μg/mL) and equal volume of PBS were also prepared to serve as controls. Mud crabs were divided into three groups (four crabs in each group). After incubation, the mixtures (200 μL) were injected into crabs. At each time point post-injection (5, 15, and 30 min), hemolymph (200 μL) was collected from crabs and mixed with an equal volume of anticoagulant buffer. After serial dilution with PBS, the samples (100 μL) were plated onto the LB plates. These plates were then incubated at 37°C overnight, and the number of bacterial clones on the plates was counted, which was used to determine the number of residual bacteria in hemolymph. After validating that SpBark expression could be silenced by injection of dsSpBark, we investigated whether SpBark knockdown could affect bacterial clearance. Each crab was injected with 200 μL of V. parahemolyticus suspension (2 × 107 cells) at 40 h after injection with dsSpBark or dsGFP. After mock PBS injection, the crabs were treated with an equal amount of bacteria in the same way. The number of residual bacteria in hemolymph was calculated using the method described above. The experiments were performed three times, and the results presented the mean of three individual experiments. Unpaired t-test was used to analyze significant differences (*P < 0.05; **P < 0.01).
Fluorescent Labeling of Bacteria and Phagocytosis Assay
The labeling of V. parahemolyticus with fluorescein isothiocyanate isomer I (FITC, Sigma) was conducted as described earlier (32). Overnight cultures of V. parahemolyticus were washed twice with PBS, re-suspended in carbonate buffered saline (pH 9.5) containing 0.2 mg/mL FITC, and then incubated at 25°C for 1 h. Subsequently, the FITC-labeled bacteria were rewashed four times to remove dissociated FITC and then re-suspended in PBS (~1 × 109 CFU/mL). Approximately 200 μL of FITC-labeled bacteria was injected into the mud crabs at 40 h after injections with dsSpBark and dsGFP. Hemolymph was collected at 30 min after bacterial injection using a syringe preloaded with anticoagulant containing 4% paraformaldehyde. After incubation at 4°C for 15 min, the hemocytes were centrifuged and re-suspended in anticoagulant. Trypan blue (2 mg/mL, Sigma) was added into the hemocyte suspension and incubated for 5 min to quench the fluorescence of non-phagocytized bacteria. After washing with PBS twice, the fixed hemocytes were dropped onto poly-L-lysine-coated glass slides, and the phagocytized bacteria were evidently detected under a fluorescence microscope (Nikon Eclipse Ti-E, Japan). A total of 600 cells (each group) were counted to determine the phagocytic percentage, which was defined here as [hemocytes ingesting bacteria/all cells observed or tested] × 100%. The phagocytosis assay was conducted three times. The data were subjected to Student's t-test, and differences with P < 0.05 were considered statistically significant.
Results
cDNA Cloning of SpBark
The full-length cDNA sequence of SpBark is 10,053 bp long, including a 24-bp 5′ untranslated region (UTR), a 1,437-bp 3′ UTR containing a poly(A) tail, and an 8,592-bp open reading frame encoding a polypeptide of 2,863 amino acids (GenBank Accession No. MH595537) (Figure 1S). The deduced protein had an 18-amino-acid signal peptide at the N-terminus and a transmembrane region close to the C-terminus. Its mature peptide had an estimated MW of 320.8 kDa and a theoretical pI of 5.58. Three putative SRCR domains, including SRCRD1 (Leu106 to Asp210), SRCRD2 (Val976 to Tyr1082), and SRCRD3 (Val1797 to Gly1914), and a following CTLD (Cys2512 to Glu2608) were found in this deduced protein (Figure 1A). The pIs of SRCRD1, SRCRD2, SRCRD3, and CTLD were 5.12, 4.58, 5.48, and 4.95, respectively.
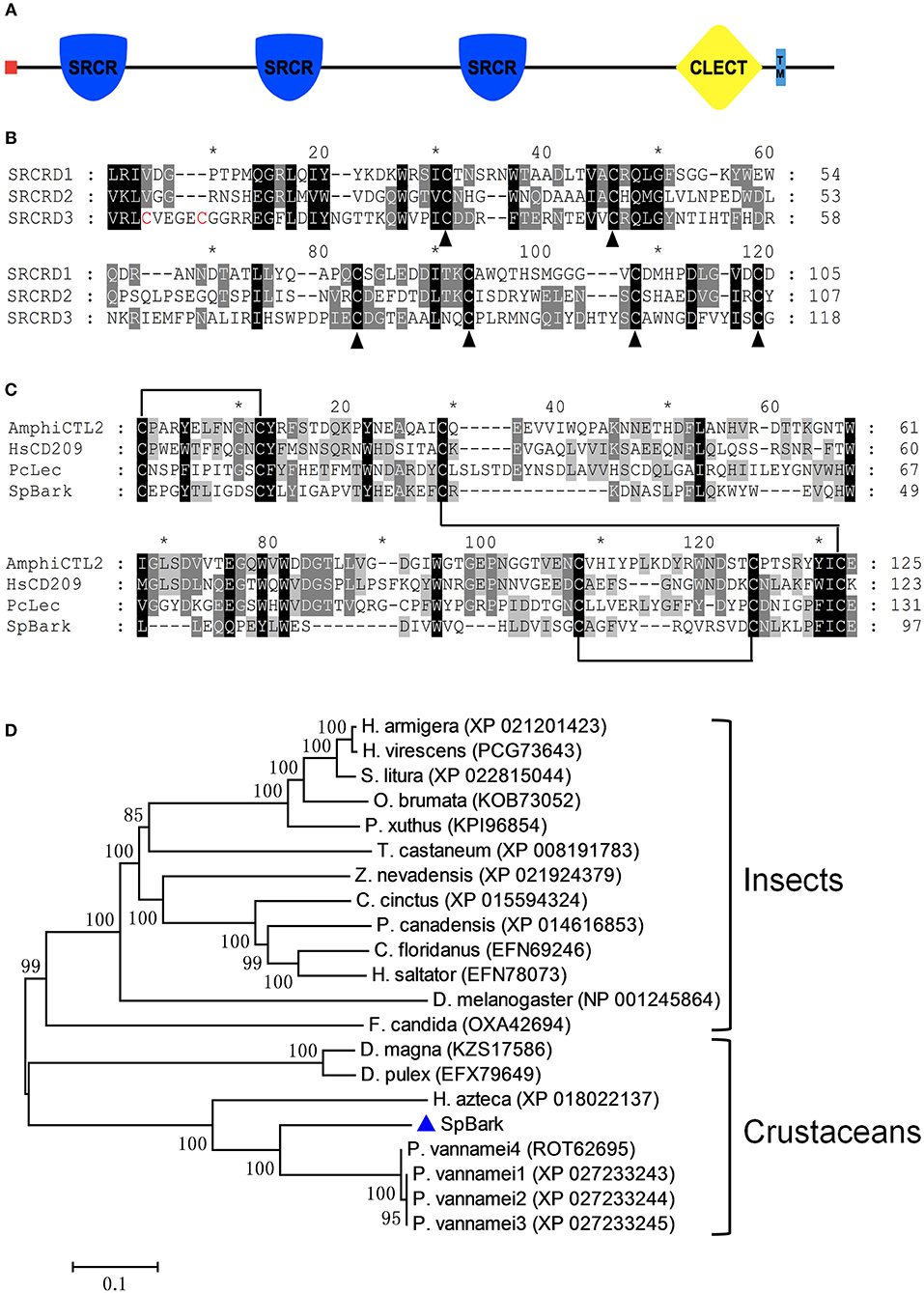
Figure 1. Architecture and alignments of SpBark domains and phylogenetic analysis. (A) Schematic of SpBark domains predicted with online software SMART. A signal peptide, three SRCRDs (from N-terminus to C-terminus: SRCRD1, SRCRD2, and SRCRD3), a CTLD, and a transmembrane region were predicted and marked with different shapes and colors. (B) Alignment of three SRCRDs. These three domains show low similarity, but six cysteine residues in these sequences were conserved except the additional two cysteine residues at the N-terminus of SRCRD3. The alignment was marked with an asterisk every 10 amino acids. (C) Alignment of CTLDs. The alignment was conducted with CTLDs from SpBark, Pclec2, human CD209 (AAI10616), and amphioxus C-type lectin (ABY54815). (D) Phylogenetic analysis of retrieved Bark or Bark-like proteins from mud crab and other species on the basis of BLASTP results. MEGA 7.0 software was used to generate the neighbor-joining tree with a bootstrap of 1,000. The bar shows the relative distance of genetic variation. SpBark was marked with a blue triangle.
Similarity and Phylogenetic Analyses
The BLASTP search analysis demonstrated that SpBark shared 71–73% identity with four Bark beetle-like proteins from Penaeus vannamei (four isoforms: ROT62695, XP_027233243, XP_027233244, and XP_027233245). SpBark also shared 59% identity with Hyalella azteca Bark-like protein (XP_018022137) but no more than 42% identity with other putative Bark-like proteins. The alignment of three SRCR domains revealed that six cysteine residues in these SRCR domains, which might be responsible for forming three disulfide bonds to stabilize the dimensional structure of SRCR domains, were well-conserved even though these three amino acid sequences shared a low identity of 29.4% (Figure 1B). Another two cysteine residues at the N-terminus of SRCRD3 were also found, suggesting that SRCRD3 may have a different structure from SRCRD1 and SRCRD2. Alignment of the CTLDs of lectins from mud crab, shrimp, human, and amphioxus showed that four cysteine residues (Cys49, Cys127, Cys142, and Cys150), which can form two disulfide bonds to stabilize the structure of classic CTLDs, were well-conserved in SpBark CTLD. Two additional cysteine residues, which may form another disulfide bond, were found at the N-terminus of each domain. Moreover, some remarkable amino acid motifs, such as EPN (Glu-Pro-Asn), QPD (Gln-Pro-Asp), and WND (Trp-Asn-Asp), responsible for carbohydrate binding were not found in SpBark CTLD. These results suggested that SpBark CTLD was unique with different sequence information from the classic CTLDs (Figure 1C).
On the basis of BLASTP results, a phylogenetic tree was constructed using Bark-like proteins to analyze the evolutionary relationships among them. In this tree, Bark-like proteins were divided into two large clusters (Figure 1D). SpBark together with other crustacean Bark-like proteins was grouped into the crustacean cluster, and the other Bark-like proteins were clustered into the insect group. SpBark and four shrimp Bark-like proteins formed a meaningful subcluster with the node value of 99, indicating that Bark-like proteins may widely exist in decapod crustaceans possessing special biological function different from the ones in insects.
SpBark Was Widely Distributed in Mud Crabs and Upregulated by Bacterial Challenges
qRT-PCR analysis revealed that SpBark was highly expressed in the hepatopancreas, hemocytes, gills, intestine, and stomach, and it was detected in the heart and muscle at a low expression level (Figure 2A). The temporal expression profiles of SpBark in hemocytes after bacterial challenges were further investigated to explore whether this gene participated in immune defense. SpBark was remarkably increased 2–6 h after challenge with V. parahemolyticus and reached the highest level (~7 fold increase) at 6 h post-injection. With regard to the challenge with S. aureus, significant induction (~2 fold increase) was observed at 12 h post-injection (Figure 2B). These results suggested that SpBark was involved in antibacterial responses of mud crabs.
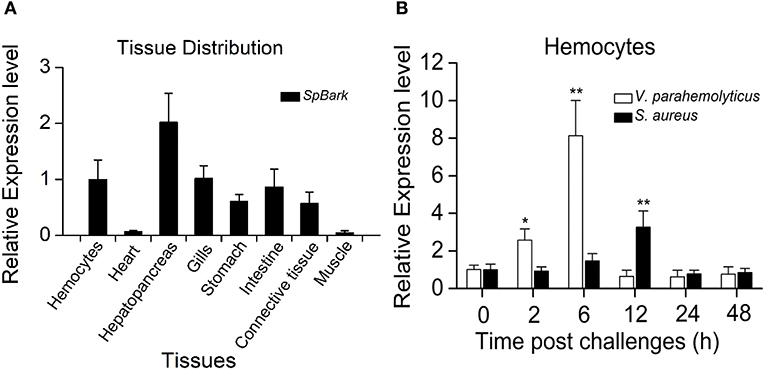
Figure 2. Tissue distribution and expression profiles of SpBark. (A) Tissue distribution of SpBark was analyzed by qRT-PCR using 18S rRNA as the internal control. (B) Expression profiles of SpBark in hemocytes at different time points after challenges with Staphylococcus aureus and Vibrio anguillarum. Significant differences are indicated with asterisks (*P < 0.05; **P < 0.01).
Four Structural Domain Proteins Were Recombinantly Expressed and Purified
The recombinant proteins of the four structural domains of SpBark were overexpressed in soluble form and then purified by Ni-NTA His Bind Resin. The predicted MWs of recombinant SRCRD1, SRCRD2, SRCRD3, and CTLD (each fused with a ~16 kDa Trx tag) were approximately 27.8, 28.0, 29.7, and 27.4 kDa, respectively. The position of each purified protein is roughly in agreement with the size of the corresponding recombinant protein (Figure 3A).
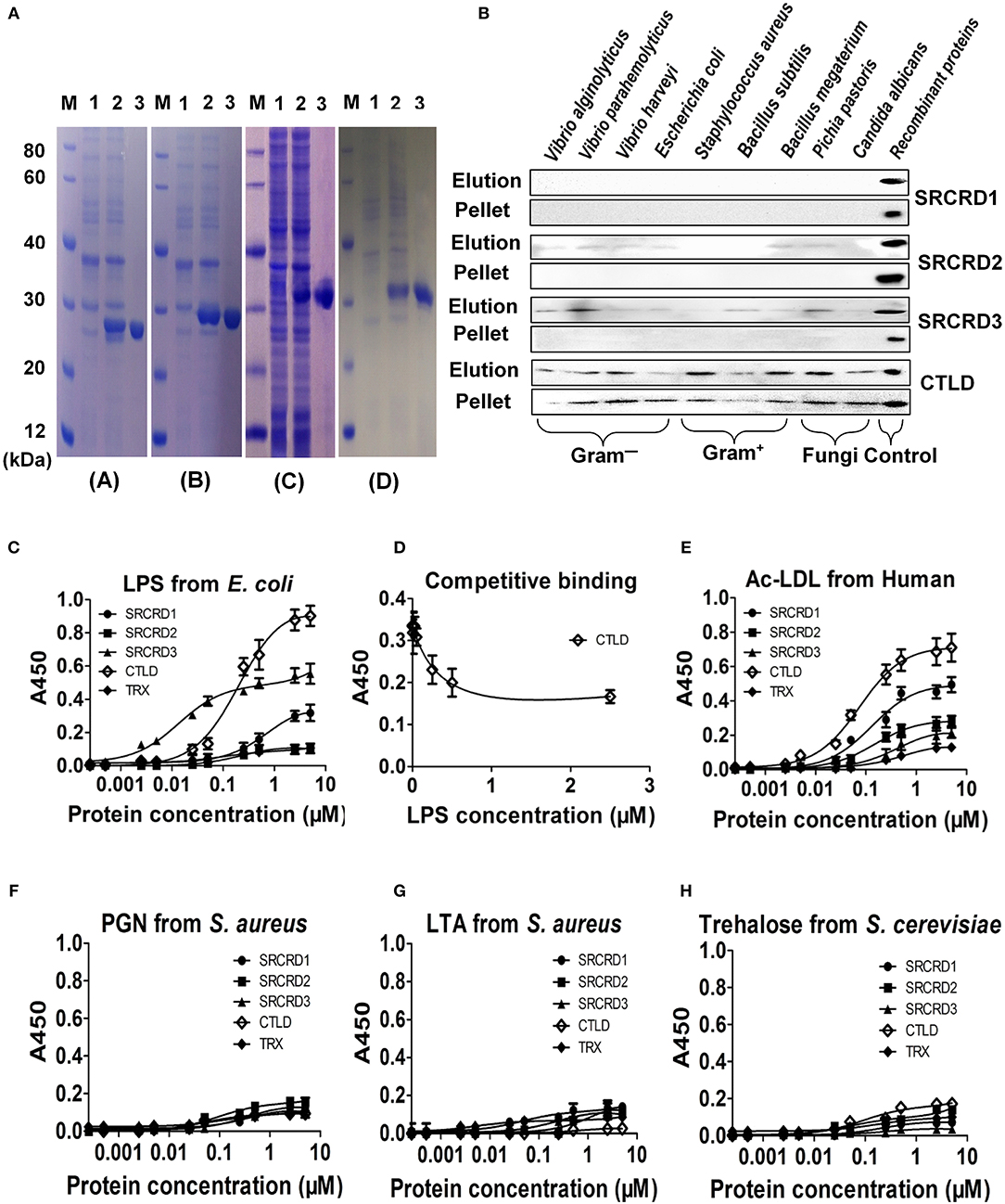
Figure 3. Purification and binding activities of four functional domains of SpBark. (A) All four functional domains (SRCRD1, SRCRD2, SRCRD3, and CTLD) were expressed in E. coli and then purified. Lane M, protein marker; lane 1, total proteins of E. coli without IPTG induction; lane 2, total proteins of E. coli with IPTG induction; lane 3, the purified recombinant proteins. (B) Binding activity of recombinant SRCRD1, SRCRD2, SRCRD3, or CTLD to different microorganisms. The binding activities of these four proteins were confirmed by Western blot, and recombinant proteins SRCRD1, SRCRD2, SRCRD3, and CTLD were sampled as the corresponding positive controls. Elution, the supernatant was harvested after centrifugation of microorganisms treated with 7% SDS solution; pellet, the eluted microorganism was obtained after three washes with PBS. (C–H) Microbial polysaccharide-binding activities were investigated using ELISA. LPS from E. coli (C and D), acetylated LDL from humans (E), PGN from S. aureus (F), LTA from S. aureus (G), and trehalose from S. cerevisiae (H) were used to coat plates. Recombinant proteins SRCRD1, SRCRD2, SRCRD3, CTLD, and TRX (negative control) were serially diluted and added into the coated plates. (D) Recombinant CTLD pre-incubated with different amounts of LPS was added for the competitive binding activity assay. Results were obtained on the basis of three independent repeats.
SpBark CTLD Exhibited the Strongest Binding Activity to Microorganisms
The microbial cell-binding activities of the four structural domain proteins were tested by Western blot. The recombinant protein was detected in pellets representing that it possessed strong binding activity, and the one in elution meant that it exhibited weak binding toward microorganisms. On the basis of these criteria, CTLD exhibited strong binding activity to all tested microorganisms, including four Gram-negative bacteria (V. harveyi, V. parahemolyticus, V. alginolyticus, and E. coli), three Gram-positive bacteria (S. aureus, B. subtilis, and B. megaterium), and two fungi (C. albicans and P. pastoris). By contrast, SRCRD2 and SRCRD3 only exhibited weak binding activities to different kinds of microorganisms, and SRCRD1 possessed no evident binding activity to microorganisms (Figure 3B). These results implied that SpBark CTLD was the major functional domain involved in bacterial recognition.
SpBark CTLD Displayed the Most Potent Binding Activity to LPS and ac-LDL
The binding activities of SpBark SRCRD1, SRCRD2, SRCRD3, and CTLD to ac-LDL and several kinds of polysaccharides were investigated by performing ELISA. In the ac-LDL binding assay, only three of the four domain proteins (SRCRD1, SRCRD2, and CTLD) exhibited ac-LDL-binding activities in varying degrees (Figure 3E). Among them, SpBark CTLD possessed the strongest binding activity, and SRCRD1 displayed more potent binding activity than SRCRD2. Given that the binding activity to mLDL is a marked property of scavenge receptors, SpBark possessing ac-LDL binding activity suggested that it might function as a scavenger receptor in mud crab. In addition, SRCRD1, SRCRD3, and CTLD exhibited notable LPS-binding activities, but none of the four domain proteins displayed apparent binding activities to the other tested polysaccharides, such as LTA, PGN, and trehalose; this finding indicated that the microorganism-binding activity exhibited by CTLD might be through binding to LPS or other untested components on the cell surface (Figures 3C,F–H). This study also revealed that CTLD displayed a much stronger LPS-binding activity than SRCRD1 and SRCRD3. The specific LPS-binding activity exhibited by CTLD was further determined by a competitive binding assay. The results demonstrated that the binding ability of CTLD to immobilized LPS on the microplates was significantly reduced by pre-incubation of CTLD with LPS, and the binding activity to immobilized LPS became weak with increasing amount of LPS to pre-incubate CTLD (Figure 3D). These results suggested that the SpBark binding activity to Gram-negative bacteria was mainly through binding LPS with its CTLD.
SpBark CTLD Agglutinated Bacteria in the Presence of Ca2+
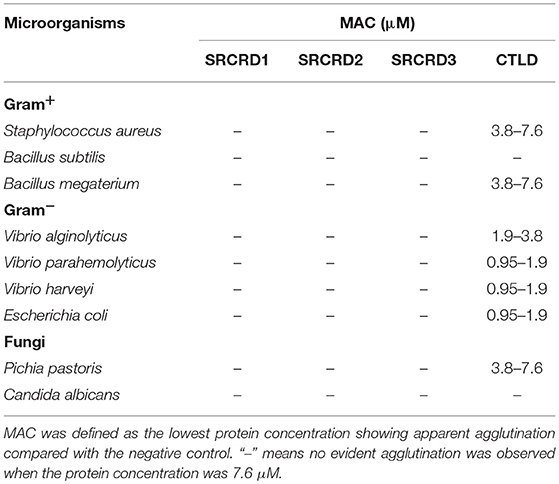
The agglutination activities of SRCRD1, SRCRD2, SRCRD3, and CTLD were investigated to explore whether they were involved in bacterial agglutination. Results demonstrated that only SpBark CTLD of these four proteins exhibited apparent agglutination activity in the presence of Ca2+ (Figure 4). The agglutination activities of CTLD to different microorganisms greatly varied. Table 2 shows that the minimal agglutinating concentration (MAC) of CTLD to Gram-positive bacteria S. aureus and B. megaterium was the same as that to fungus P. pastoris, which was lower than the MACs to Gram-negative bacteria. This finding indicates that CTLD possessed stronger agglutinating abilities to Gram-negative bacteria than the other tested microorganisms. SpBark CTLD could agglutinate bacteria, suggesting that it can function like many common CTLDs acting as a binding site of PRR.
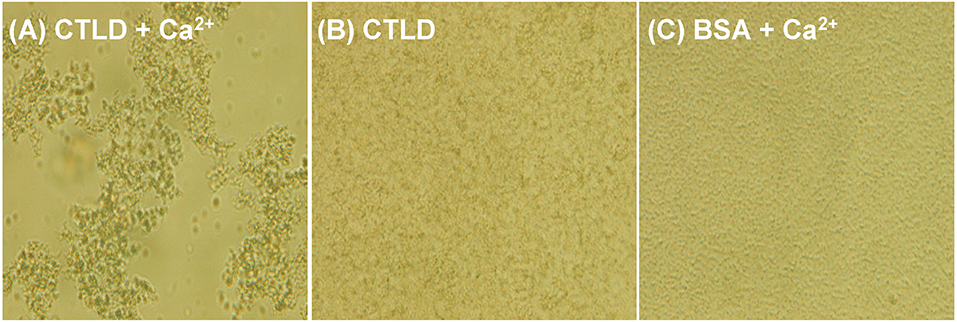

Figure 4. Agglutination of V. parahemolyticus induced by SpBark CTLD in the presence of Ca2+. V. parahemolyticus was incubated with SpBark CTLD with (A) or without Ca2+ (B). BSA plus Ca2+ was used as the negative control (C). Agglutination was observed through light microscopy.
SpBark CTLD Promoted V. parahemolyticus Clearance in Hemolymph
Given that SpBark CTLD exhibited specific binding abilities to LPS and Gram-negative bacteria, we investigated whether this protein facilitated bacterial clearance in crabs through its binding activity. V. parahemolyticus pre-incubated with CTLD, Trx protein, or PBS was injected into healthy crabs. The number of bacteria in vivo was significantly decreased at 15 min post-injection by pre-incubating V. parahemolyticus with CTLD compared with that treated with Trx or PBS; however, no significant differences in bacterial numbers were found at 5 min post-injection among these three groups. At 30 min after the injection of bacteria treated with CTLD, an extremely significant decrease in the number of bacteria (P < 0.01) in crabs was observed compared with that in control crabs (Figure 5). These results revealed that pre-incubation of V. parahemolyticus with CTLD would facilitate bacterial clearance in vivo.
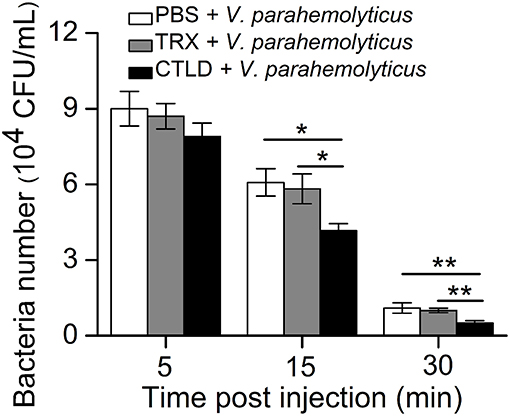
Figure 5. Promotion of bacterial clearance by injection with SpBark CTLD protein. The ability to clear bacteria (V. parahemolyticus) in hemolymph was increased by the “overexpression” of CTLD protein with TRX tag protein and PBS as controls. The results presented the mean of three individual experiments. Asterisks indicate the significant differences (*P < 0.05; **P < 0.01).
Knockdown of SpBark Reduced Bacterial Clearance in Mud Crab
To investigate the in vivo function of SpBark, RNAi of SpBark and bacterial clearance assays were performed. qRT-PCR analysis indicated that the transcripts of SpBark in hemocytes significantly declined at 40 and 48 h after the first injection of dsSpBark (Figure 6A). This result indicated that the injection of SpBark dsRNA into crabs could remarkably suppress SpBark expression. After SpBark knockdown, V. parahemolyticus was injected into crabs, and the residual number in hemolymph was counted to assess the bacterial clearance ability. As shown in Figure 6B, the residual number of V. parahemolyticus in vivo significantly increased at 15 and 30 min post-injection compared with that in dsGFP or PBS group. This result suggested that the bacterial clearance ability in hemolymph of SpBark-silenced crabs was severely impaired.
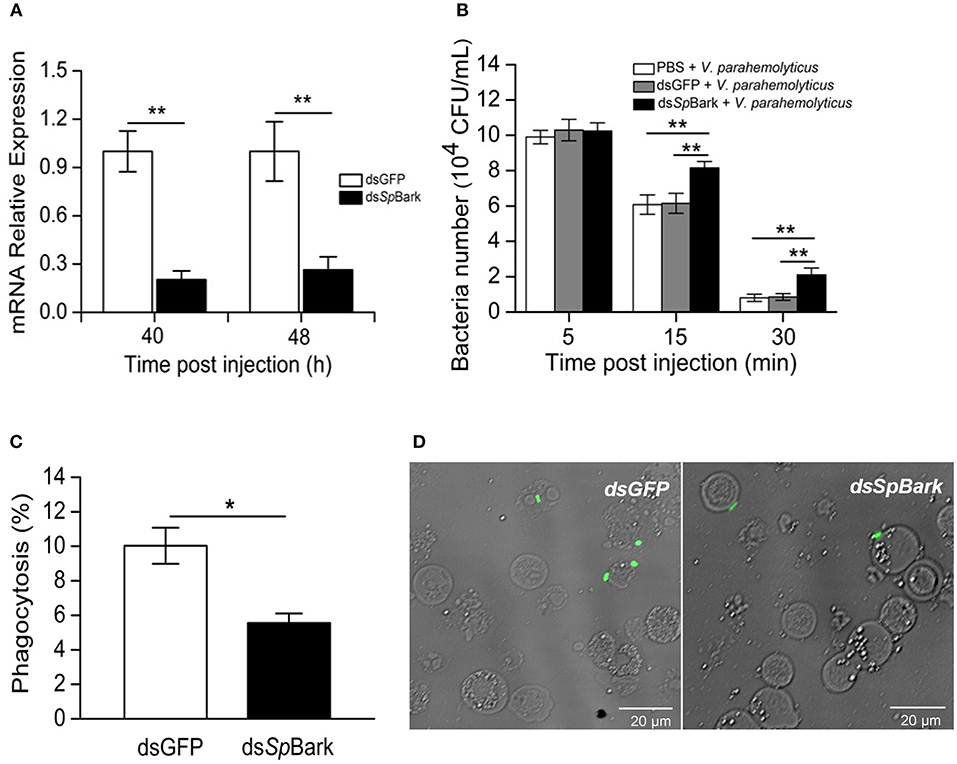
Figure 6. RNAi of SpBark suppresses bacterial clearance in hemolymph by modulating hemocyte phagocytosis. (A) Knockdown of SpBark. The mRNA expression level of SpBark in hemocytes after injection with dsSpBark or dsGFP was analyzed by qRT-PCR. (B) Suppression of bacterial clearance by SpBark dsRNA injection. Bacterial clearance ability in hemolymph after SpBark knockdown was evaluated. Either dsGFP or PBS was injected and served as control. (C) Phagocytosis of FITC-labeled V. parahemolyticus by hemocytes in crabs. At 40 h after the first treatment with dsSpBark, FITC-labeled V. parahemolyticus was injected into crabs. Hemocytes were collected 30 min later, and then hemocytes and V. parahemolyticus cells (green) were observed and counted under a fluorescence microscope. The dsGFP-treated crabs were used as control. (D) The phagocytosis rate was calculated using images captured by the fluorescence microscope, and a total of 600 cells in each group were counted. The results were shown as the means of three individual experiments. Asterisks indicate the significant differences (*P < 0.05; **P < 0.01).
SpBark Promoted Phagocytosis of Bacteria
To reveal the mechanism by which SpBark facilitates bacterial clearance, bacterial phagocytosis assay was conducted. After SpBark was silenced, FITC-labeled bacteria were injected into crabs. Hemocytes were collected to determine the phagocytosis rate. The results showed that the phagocytic rate of hemocytes in SpBark-silenced crabs was significantly lower than that in control crabs: the phagocytosis rate was decreased by ~45% by knockdown of SpBark (from 9.7 to 5.3%) (Figures 6C,D). This result suggests that SpBark modulated hemocyte phagocytosis of V. parahemolyticus.
Discussion
Scavenger receptors are widely distributed in invertebrate and vertebrate animals exhibiting diverse biological functions through binding to various ligands (8). In the present study, we characterized a novel scavenger receptor-like protein in S. paramamosain, which shared high identity with Bark or Bark-like protein and thus designated as SpBark. SpBark CTLD displayed the strongest binding activity to ac-LDL and LPS. Knockdown of SpBark remarkably suppressed the clearance of bacteria in vivo and phagocytosis of bacteria. These findings suggested that SpBark might function as a potential PRR playing an important role in immune defense against Gram-negative bacteria.
Recently, a consensus definitive classification of scavenger receptors was proposed, and the scavenger receptors of mammals were categorized into 12 classes on the basis of their sequence structure and biological functions (11). Though SpBark protein contains SRCRD and CTLD, which are characteristic domains present in class A, E, and I scavenger receptors, clustering this protein into any denoted class of mammalian scavenger receptors seems difficult. No consensus nomenclature system for invertebrate scavenger receptors and definitive criteria to define scavenger receptors exist. The scavenger receptor-like protein “Bark” was first characterized in Drosophila, which has the signature structural domains (SRCRD and CTLD) present in the mammalian scavenger receptors. However, it was first identified as a scaffold protein involved in mounting of the core complex of septate junctions (33). Thus, most Bark homologs from other species were displayed as Bark or Bark-like proteins in BLASTP search results because no new biological function was further reported regarding Drosophila Bark and its homologs. Considering that SpBark shared a similar distribution pattern of structural domains (SP at N-terminus, three SRCRDs, one CTLD, and TM region at C-terminus) with Drosophila Bark and had 40% identity at the protein level, we named this scavenger receptor-like protein as a new Bark-like protein in mud crab. Bark homologs were also found in other invertebrate animals. A phylogenetic tree constructed using invertebrate Bark homologs revealed that SpBark together with the other crustacean Bark proteins formed a unique meaningful cluster. This finding suggested that crustacean Bark proteins had a close evolutionary relationship and might possess similar biological functions.
Most C-type lectins possess binding activities to microbial polysaccharides and exhibit agglutination activity with their CTLDs (7, 23). Different from the classic CTLDs with four cysteine residues forming two disulfide bonds, SpBark CTLD was a “long form” CTLD containing six cysteine residues, which could generate three disulfide bonds. This finding suggested that it might form a distinct structure displaying special functions. Our study revealed that SpBark CTLD was different from certain classic CTLDs displaying broad polysaccharide-binding abilities (7, 39, 40), which only exhibited binding activity to LPS but not to other tested polysaccharides. Apart from the possible effect of the distinct structure on its biological function, we speculated that this limited polysaccharide-binding activity partially originated from the lack of some signature motifs (e.g., EPN, QPD, and WND) essential for polysaccharide-binding activity in SpBark CTLD. The specific binding activity to LPS may rely on certain mutant carbohydrate-binding motifs in SpBark CTLD because some mutant or unknown carbohydrate-binding motifs present in reported CTLDs had been shown to be involved in polysaccharide-binding activity (7). SpBark CTLD exhibited notable agglutination activity against bacteria and a fungus in the presence of Ca2+, which was the basic function of classic CTLDs. This finding suggested that it could function similar to the classic CTLDs acting as the binding regions of recognition receptors. Given that the SpBark protein was a macromolecule with three SRCRDs and one CTLD, we speculated that the major biological function of SpBark might be displayed through these functional domains. SpBark CTLD exhibited a much stronger binding activity to LPS than the other SRCRDs, indicating that this CTLD was the key recognition domain of SpBark.
Although Drosophila Bark plays a role in cell adhesion by mounting of the core complex to facilitate the maturation of septate junctions (33), whether this protein and its homologs in other species are involved in immune defenses remains unclear. In this study, SpBark was upregulated in hemocytes by bacterial challenges, suggesting that it might be an important component implicated in immune defense against bacteria. In addition, though many mammalian scavenger receptors are distributed in specific cells or tissues, SpBark was highly expressed in the hemocytes, hepatopancreas, gills, and intestine, which are the major immune tissues or organs in crustaceans. Among them, hemocytes can remove the invading pathogens through different mechanisms, such as phagocytosis, nodulation, and encapsulation, playing key roles in immune defense of circulating hemolymph. The hepatopancreas, equivalent to the liver of mammals or fat body of insects, was considered the key immune tissue. Gills and intestine easily contacted potential pathogens existing in water environment and intestinal contents, which might be involved in the initial defenses against external pathogens. Considering that the functional domains of reported mammalian scavenger receptors and Drosophila Bark were located at the extracellular regions (33), the SRCRDs and CTLD of SpBark might be the extracellular domains because SpBark and Drosophila Bark shared the same distribution pattern of functional domains. SpBark ubiquitously distributed in immune tissues may facilitate the recognition and elimination of bacteria via its extracellular CTLD.
The invading pathogens in hemolymph are routinely removed by cellular immune responses and humoral immune reactions (3, 41). Hemocyte phagocytosis is an important activity that eliminates diverse invading pathogens. Recent reports revealed that certain receptors or receptor-associated proteins are required to modulate hemocyte phagocytosis activity (30, 39, 42, 43). To further confirm whether SpBark was involved in bacterial clearance and modulated hemocyte phagocytosis of bacteria, the number of bacteria in cell-free hemolymph and in hemocytes of SpBark-silenced crabs was counted. Our results showed that knockdown of SpBark significantly suppressed the clearance of V. parahemolyticus in hemolymph and hemocyte phagocytosis of bacteria. These results demonstrated that SpBark participated in the removal of bacteria in hemolymph and modulated the phagocytosis of V. parahemolyticus. Although many immune responses (e.g., the synthesis of AMPs and melanin) are involved in bacterial clearance (41), the fact that the bacterial clearance activity in hemolymph and hemocyte phagocytosis of bacteria were decreased simultaneously by SpBark knockdown clearly suggested that the bacterial clearance in hemolymph is closely related to hemocyte phagocytosis of bacteria. Knockdown of SpBark severely affected hemocyte phagocytosis of bacteria, which finally resulted in the decrease of bacterial clearance. Drosophila Bark was internalized into cells through clathrin-coated vesicles and then sorted back to the plasma membrane via the recycling endosomes (33). This kind of internalization mechanism might apply to SpBark. As many scavenger receptors can bind and internalize pathogens, SpBark may function similar to a scavenger receptor mediating hemocyte phagocytosis through binding and internalizing bacteria. The possible antibacterial mode may be as follows: in hemolymph, the invading bacteria were sensed and bound to by SpBark CTLD, and this binding behavior would modulate hemocyte phagocytosis, which finally led to the internalization and degradation of invading bacteria (Figure 7). Considering that SpBark is ubiquitously distributed in major immune tissues, a similar mechanism might occur in the tissue cells of gills and intestine, which might facilitate the removal of external bacteria from environments and intestinal contents.
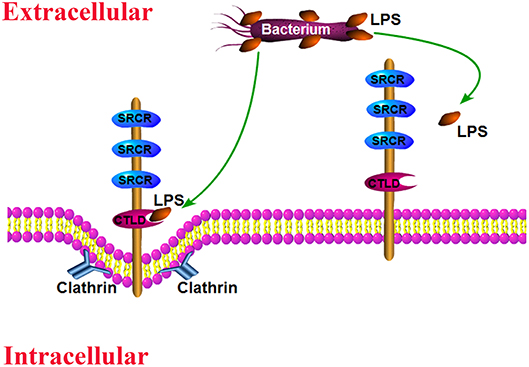
Figure 7. Schematic of the likely antibacterial mode mediated by SpBark. SpBark can sense and bind LPS on the surface of Gram-negative bacteria with its CTLD domain, which would induce phagocytosis of bacteria possibly through a clathrin-dependent mechanism.
In this study, SpBark SCRCDs exhibited much weaker binding activity to LPS than SpBark CTLD and no binding activity to the other tested polysaccharides, although SRCRD-containing proteins were shown to bind to various ligands participating in immune responses (17, 18, 20, 44, 45). This result suggested that SpBark SRCRDs might play a major role in other biological processes. Our study revealed that SpBark SRCRDs possessed binding ability to ac-LDL, implying that SpBark may be involved in lipoprotein metabolism. Drosophila Bark is shown to mediate cell adhesion and functions as an essential component in a core complex of septate junctions (33). Three SRCRDs of Drosophila Bark, as the major functional domains, might play a role in this biological process. Given that SpBark shared similar structural domains with those of Drosophila Bark, we speculated that SpBark exhibited similar biological function through its SRCRDs. Thus, SpBark might be a multifunctional protein; however, further studies are needed to confirm this speculation.
In summary, we characterized a novel scavenger receptor-like protein, namely, SpBark, in the present study. SpBark CTLD possessed much stronger binding activities to ac-LDL and LPS than three SRCRDs serving as the major ligand-binding site. Knockdown of SpBark significantly suppressed the phagocytosis of bacteria and bacterial clearance in hemolymph. This finding clearly suggested that SpBark modulated hemocyte phagocytosis of V. parahemolyticus through binding to LPS with its CTLD, which was an important activity to eliminate invading Gram-negative bacteria in vivo. This study provided new insights into the biological functions of Bark proteins and antibacterial mechanisms of invertebrates.
Data Availability
The datasets generated for this study can be found in the GeneBank (Accession No. MH595537).
Author Contributions
X-CL and W-HF conceived and designed the experiments. X-CL, W-HF, and JZ wrote the manuscript. JZ and X-CL conducted most of the experiments. YueW, HM, J-FZ, YuanW, and SZ contributed experimental suggestions and revised the manuscript.
Funding
This work was financially supported by the National Natural Science Foundation of China (No. 31772886), the special research grant for the National Non-profit Institute (East China Sea Fisheries Research Institute) (No. 2019T01), the STU Scientific Research Foundation for Talents (No. NTF17006), and the Central Public-interest Scientific Institution Basal Research Foundation, CAFS (No. 2016RC-LX04).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fimmu.2019.01992/full#supplementary-material
References
1. Loker ES, Adema CM, Zhang SM, Kepler TB. Invertebrate immune systems–not homogeneous, not simple, not well understood. Immunol Rev. (2010) 198:10–24. doi: 10.1111/j.0105-2896.2004.0117.x
2. Medzhitov R, Janeway CA Jr. Decoding the patterns of self and nonself by the innate immune system. Science. (2002) 296:298–300. doi: 10.1126/science.1068883
3. Li F, Xiang J. Recent advances in researches on the innate immunity of shrimp in China. Dev Comp Immunol. (2013) 39:11–26. doi: 10.1016/j.dci.2012.03.016
4. Christophides GK, Vlachou D, Kafatos FC. Comparative and functional genomics of the innate immune system in the malaria vector Anopheles gambiae. Immunol Rev. (2004) 198:127–48. doi: 10.1111/j.0105-2896.2004.0127.x
5. Watson FL, Puttmann-Holgado R, Thomas F, Lamar DL, Hughes M, Kondo M, et al. Extensive diversity of Ig-superfamily proteins in the immune system of insects. Science. (2005) 309:1874–8. doi: 10.1126/science.1116887
6. Nakamoto M, Moy RH, Xu J, Bambina S, Yasunaga A, Shelly SS, et al. Virus recognition by Toll-7 activates antiviral autophagy in Drosophila. Immunity. (2012) 36:658–67. doi: 10.1016/j.immuni.2012.03.003
7. Wang XW, Wang JX. Pattern recognition receptors acting in innate immune system of shrimp against pathogen infections. Fish Shellfish Immunol. (2013) 34:981–9. doi: 10.1016/j.fsi.2012.08.008
8. Canton J, Neculai D, Grinstein S. Scavenger receptors in homeostasis and immunity. Nat Rev Immunol. (2013) 13:621–34. doi: 10.1038/nri3515
9. Brown MS, Basu SK, Falck JR, Ho YK, Goldstein JL. The scavenger cell pathway for lipoprotein degradation: specificity of the binding site that mediates the uptake of negatively-charged LDL by macrophages. J Supramol Struct. (1980) 13:67–81. doi: 10.1002/jss.400130107
10. Zani IA, Stephen SL, Mughal NA, Russell D, Homer-Vanniasinkam S, Wheatcroft SB, et al. Scavenger receptor structure and function in health and disease. Cells. (2015) 4:178–201. doi: 10.3390/cells4020178
11. Prabhudas MR, Baldwin CL, Bollyky PL, Dme B, Drickamer K, Febbraio M, et al. A consensus definitive classification of scavenger receptors and their roles in health and disease. J Immunol. (2017) 198:3775–89. doi: 10.4049/jimmunol.1700373
12. Resnick D, Pearson A, Krieger M. The SRCR superfamily: a family reminiscent of the Ig superfamily. Trends Biochem Sci. (1994) 19:5–8. doi: 10.1016/0968-0004(94)90165-1
13. Sarrias MR, Gronlund J, Padilla O, Madsen J, Holmskov U, Lozano F. The Scavenger Receptor Cysteine-Rich (SRCR) domain: an ancient and highly conserved protein module of the innate immune system. Crit Rev Immunol. (2004) 24:1–37. doi: 10.1615/CritRevImmunol.v24.10
14. Martinez VG, Moestrup SK, Holmskov U, Mollenhauer J, Lozano F. The conserved scavenger receptor cysteine-rich superfamily in therapy and diagnosis. Pharmacol Rev. (2011) 63:967–1000. doi: 10.1124/pr.111.004523
15. Franc NC, Dimarcq JL, Lagueux M, Hoffmann J, Ezekowitz RA. Croquemort, a novel drosophila hemocyte/macrophage receptor that recognizes apoptotic cells. Immunity. (1996) 4:431–43. doi: 10.1016/S1074-7613(00)80410-0
16. Liu L, Yang J, Qiu L, Wang L, Zhang H, Wang M, et al. A novel scavenger receptor-cysteine-rich (SRCR) domain containing scavenger receptor identified from mollusk mediated PAMP recognition and binding. Dev Comp Immunol. (2011) 35:227–39. doi: 10.1016/j.dci.2010.09.010
17. Bessa Pereira C, Bockova M, Santos RF, Santos AM, Martins de Araujo M, Oliveira L, et al. The scavenger receptor SSc5D physically interacts with bacteria through the SRCR-containing N-terminal domain. Front Immunol. (2016) 7:416. doi: 10.3389/fimmu.2016.00416
18. Burkard C, Lillico SG, Reid E, Jackson B, Mileham AJ, Ait-Ali T, et al. Precision engineering for PRRSV resistance in pigs: macrophages from genome edited pigs lacking CD163 SRCR5 domain are fully resistant to both PRRSV genotypes while maintaining biological function. PLoS Pathog. (2017) 13:e1006206. doi: 10.1371/journal.ppat.1006206
19. Burkard C, Opriessnig T, Mileham AJ, Stadejek T, Ait-Ali T, Lillico SG, et al. Pigs lacking the scavenger receptor cysteine-rich domain 5 of CD163 are resistant to porcine reproductive and respiratory syndrome virus 1 infection. J Virol. (2018) 92:e00415–18. doi: 10.1128/JVI.00415-18
20. Mukhopadhyay S, Gordon S. The role of scavenger receptors in pathogen recognition and innate immunity. Immunobiology. (2004) 209:39–49. doi: 10.1016/j.imbio.2004.02.004
21. Jozefowski S, Arredouani M, Sulahian T, Kobzik L. Disparate regulation and function of the class A scavenger receptors SR-AI/II and MARCO. J Immunol. (2005) 175:8032–41. doi: 10.4049/jimmunol.175.12.8032
22. Mukhopadhyay S, Varin A, Chen Y, Liu B, Tryggvason K, Gordon S. SR-A/MARCO-mediated ligand delivery enhances intracellular TLR and NLR function, but ligand scavenging from cell surface limits TLR4 response to pathogens. Blood. (2011) 117:1319–28. doi: 10.1182/blood-2010-03-276733
23. Vasta GR, Ahmed H, Bianchet MA, Fernandez-Robledo JA, Amzel LM. Diversity in recognition of glycans by F-type lectins and galectins: molecular, structural, and biophysical aspects. Ann N Y Acad Sci. (2012) 1253:E14–26. doi: 10.1111/j.1749-6632.2012.06698.x
24. Zelensky AN, Gready JE. The C-type lectin-like domain superfamily. FEBS J. (2010) 272:6179–17. doi: 10.1111/j.1742-4658.2005.05031.x
25. Herre J, Willment JA, Gordon S, Brown GD. The role of Dectin-1 in antifungal immunity. Crit Rev Immunol. (2004) 24:193–203. doi: 10.1615/CritRevImmunol.v24.i3.30
26. Yoshimoto R, Fujita Y, Kakino A, Iwamoto S, Takaya T, Sawamura T. The discovery of LOX-1, its ligands and clinical significance. Cardiovasc Drugs Ther. (2011) 25:379–91. doi: 10.1007/s10557-011-6324-6
27. Nakamura K, Funakoshi H, Tokunaga F, Nakamura T. Molecular cloning of a mouse scavenger receptor with C-type lectin (SRCL)(1), a novel member of the scavenger receptor family. Biochim Biophys Acta. (2001) 1522:53–58. doi: 10.1016/S0167-4781(01)00284-6
28. Ramet M, Pearson A, Manfruelli P, Li X, Koziel H, Gobel V, et al. Drosophila scavenger receptor CI is a pattern recognition receptor for bacteria. Immunity. (2001) 15:1027–38. doi: 10.1016/S1074-7613(01)00249-7
29. Kim SG, Jo YH, Seong JH, Park KB, Noh MY, Cho JH, et al. TmSR-C, scavenger receptor class C, plays a pivotal role in antifungal and antibacterial immunity in the coleopteran insect Tenebrio molitor. Insect Biochem Mol Biol. (2017) 89:31–42. doi: 10.1016/j.ibmb.2017.08.007
30. Bi WJ, Li DX, Xu YH, Xu S, Li J, Zhao XF, et al. Scavenger receptor B protects shrimp from bacteria by enhancing phagocytosis and regulating expression of antimicrobial peptides. Dev Comp Immunol. (2015) 51:10–21. doi: 10.1016/j.dci.2015.02.001
31. Wu YM, Yang L, Li XJ, Li L, Wang Q, Li WW. A class B scavenger receptor from Eriocheir sinensis (EsSR-B1) restricts bacteria proliferation by promoting phagocytosis. Fish Shellfish Immunol. (2017) 70:426–36. doi: 10.1016/j.fsi.2017.09.034
32. Kong T, Gong Y, Liu Y, Wen X, Tran NT, Aweya JJ, et al. Scavenger receptor B promotes bacteria clearance by enhancing phagocytosis and attenuates white spot syndrome virus proliferation in Scylla paramamosian. Fish Shellfish Immunol. (2018) 78:79–90. doi: 10.1016/j.fsi.2018.04.027
33. Hildebrandt A, Pflanz R, Behr M, Tarp T, Riedel D, Schuh R. Bark beetle controls epithelial morphogenesis by septate junction maturation in Drosophila. Dev Biol. (2015) 400:237–47. doi: 10.1016/j.ydbio.2015.02.008
34. Söderhäll K, Smith VJ. Separation of the haemocyte populations of CarcinusMaenas and other marine decapods, and prophenoloxidase distribution. Dev Comp Immunol. (1983) 7:229–39. doi: 10.1016/0145-305X(83)90004-6
35. Wang Y, Zhang C, Wang H, Ma H, Huang YQ, Lu JX, et al. Involvement of a newly identified atypical type II crustin (SpCrus5) in the antibacterial immunity of mud crab Scylla paramamosain. Fish Shellfish Immunol. (2018) 75:346–56. doi: 10.1016/j.fsi.2018.02.026
36. Wang H, Zhang JX, Wang Y, Fang WH, Wang Y, Zhou JF, et al. Newly identified type II crustin (SpCrus2) in Scylla paramamosain contains a distinct cysteine distribution pattern exhibiting broad antimicrobial activity. Dev Comp Immunol. (2018) 84:1–13. doi: 10.1016/j.dci.2018.01.021
37. Zhou J, Zhao S, Fang WH, Zhou JF, Zhang JX, Ma H, et al. Newly identified invertebrate-type lysozyme (Splys-i) in mud crab (Scylla paramamosain) exhibiting muramidase-deficient antimicrobial activity. Dev Comp Immunol. (2017) 74:154–66. doi: 10.1016/j.dci.2017.04.017
38. Shi XZ, Zhou J, Lan JF, Jia YP, Zhao XF, Wang JX. A Lysin motif (LysM)-containing protein functions in antibacterial responses of red swamp crayfish, Procambarus clarkii. Dev Comp Immunol. (2013) 40:311–9. doi: 10.1016/j.dci.2013.03.011
39. Zhang XW, Wang Y, Wang XW, Wang L, Mu Y, Wang JX. A C-type lectin with an immunoglobulin-like domain promotes phagocytosis of hemocytes in crayfish Procambarus clarkii. Sci Rep. (2016) 6:29924. doi: 10.1038/srep29924
40. Wang XW, Zhang XW, Xu WT, Zhao XF, Wang JX. A novel C-type lectin (FcLec4) facilitates the clearance of Vibrio anguillarum in vivo in Chinese white shrimp. Dev Comp Immunol. (2009) 33:1039–47. doi: 10.1016/j.dci.2009.05.004
41. Tassanakajon A, Rimphanitchayakit V, Visetnan S, Amparyup P, Somboonwiwat K, Charoensapsri W, et al. Shrimp humoral responses against pathogens: antimicrobial peptides and melanization. Dev Comp Immunol. (2018) 80:81–93. doi: 10.1016/j.dci.2017.05.009
42. Wang XW, Zhao XF, Wang JX. C-type lectin binds to beta-integrin to promote hemocytic phagocytosis in an invertebrate. J Biol Chem. (2014) 289:2405–14. doi: 10.1074/jbc.M113.528885
43. Xu JD, Diao MQ, Niu GJ, Wang XW, Zhao XF, Wang JX. A small GTPase, RhoA, inhibits bacterial infection through integrin mediated phagocytosis in invertebrates. Front Immunol. (2018) 9:1928. doi: 10.3389/fimmu.2018.01928
44. Elomaa O, Kangas M, Sahlberg C, Tuukkanen J, Sormunen R, Liakka A, et al. Cloning of a novel bacteria-binding receptor structurally related to scavenger receptors and expressed in a subset of macrophages. Cell. (1995) 80:603–9. doi: 10.1016/0092-8674(95)90514-6
Keywords: scavenger receptor-like protein, C-type lectin domain, LPS-binding activity, bacterial clearance, phagocytosis activity, Scylla paramamosain
Citation: Li X-C, Zhou J, Zhou J-F, Wang Y, Ma H, Wang Y, Zhao S and Fang W-H (2019) SpBark Suppresses Bacterial Infection by Mediating Hemocyte Phagocytosis in an Invertebrate Model, Scylla paramamosain. Front. Immunol. 10:1992. doi: 10.3389/fimmu.2019.01992
Received: 31 May 2019; Accepted: 06 August 2019;
Published: 23 August 2019.
Edited by:
Bok-Luel Lee, Pusan National University, South KoreaReviewed by:
Erjun Ling, Shanghai Institutes for Biological Sciences (CAS), ChinaYeon Soo Han, Chonnam National University, South Korea
Kensuke Shibata, Kyushu University, Japan
Copyright © 2019 Li, Zhou, Zhou, Wang, Ma, Wang, Zhao and Fang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xin-Cang Li, bGl4aW44Njg3QDE2My5jb20=; Wen-Hong Fang, ZndlbmhvbmdAMTYzLmNvbQ==
 Xin-Cang Li
Xin-Cang Li Jian Zhou1
Jian Zhou1 Hongyu Ma
Hongyu Ma Shu Zhao
Shu Zhao