- 1Discipline of Paediatrics, Trinity College, The University of Dublin, Dublin, Ireland
- 2Paediatric Research Laboratory, Trinity Translational Medicine Institute (TTMI), St James’ Hospital, Dublin, Ireland
- 3Trinity Health Kidney Centre, TTMI, Trinity College, Dublin, Ireland
- 4Viral Immunology Group, School of Biochemistry and Immunology, Trinity Biomedical Sciences Institute, Dublin, Ireland
- 5Neonatology, Coombe Women and Infant’s University Hospital, Dublin, Ireland
- 6National Children’s Research Centre, Dublin, Ireland
- 7Viral Immunology Group, Royal College of Surgeons in Ireland‐Medical University of Bahrain, Zallaq, Bahrain
- 8Irish Centre for Vascular Biology, Trinity College Dublin, Dublin, Ireland
- 9Neonatology, Children’s Hospital Ireland (CHI) at Crumlin, Dublin, Ireland
- 10Paediatrics, CHI at Tallaght, Tallaght University Hospital, Dublin, Ireland
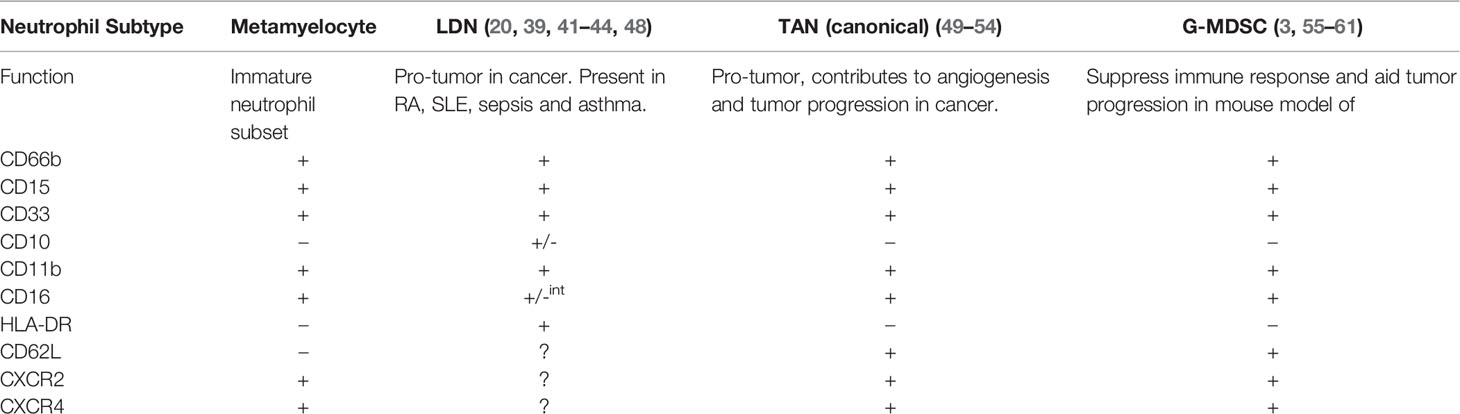
Neutrophils are the most abundant innate immune cell with critical anti-microbial functions. Since the discovery of granulocytes at the end of the nineteenth century, the cells have been given many names including phagocytes, polymorphonuclear neutrophils (PMN), granulocytic myeloid derived suppressor cells (G-MDSC), low density neutrophils (LDN) and tumor associated neutrophils (TANS). This lack of standardized nomenclature for neutrophils suggest that biologically distinct populations of neutrophils exist, particularly in disease, when in fact these may simply be a manifestation of the plasticity of the neutrophil as opposed to unique populations. In this review, we profile the surface markers and granule expression of each stage of granulopoiesis to offer insight into how each stage of maturity may be identified. We also highlight the remarkable surface marker expression profiles between the supposed neutrophil populations.
Introduction
Neutrophils are critical actors in the innate immune system and the body’s first line of defense against pathogens (1, 2). Approximately 100,000,000,000 neutrophils are generated in the bone marrow every day, making them the most common leukocyte in human blood (3). Neutrophils extravasate from the vasculature and are recruited to the site of infection to kill invading pathogens (4). Deficiencies in neutrophils have significant adverse effects on the overall response to infection. For instance, neutropenia, a reduction in the number of circulating neutrophils, is a condition associated with high morbidity and mortality (5). A hallmark of chronic granulomatous disease is impaired nicotinamide adenine dinucleotide phosphate (NADPH) activity, which results in reduced neutrophil bactericidal capacity (6).
The anti-microbial roles of neutrophils also include degranulation, whereby neutrophils release granule-derived mediators, engulfing pathogens by phagocytosis, and the release of neutrophil extracellular traps (NETs) which ensnare invading bacteria, fungi and viruses (7, 8). Neutrophils are highly pro-inflammatory; therefore, excessive neutrophil accumulation and prolonged activation can result in tissue damage and chronic inflammation (4). A vital activity of neutrophils is the release of reactive oxygen species (ROS). During this oxidative burst, the neutrophil consumes oxygen, which is converted to superoxide radicals via the NADPH oxidase 2 (NOX2) complex (4, 9, 10). Through NOX2, neutrophils primarily produce hydrogen peroxide and superoxide (11). NOX2 is highly active during neutrophil-mediated phagocytosis and localized in the phagosomal membrane to guide superoxide into the phagosome.
Neutrophils follow the leukocyte adhesion cascade to move from the bone marrow to sites of infection or inflammation (12). Novel aspects occur in the regulation of the leukocyte adhesion cascade during which time they display different phenotypes which results in various neutrophils with distinct properties (13) and specialized functions. The neutrophil response varies dramatically during this time in a process known as priming, allowing the cell to specifically target the particular site and modulate its anti-microbial action (14). This migration of neutrophils follows a circadian pattern. Circadian rhythms are important regulators of specific immune functions (15) and amongst the three main functions of neutrophils, degranulation, netosis and phagocytosis, neutrophils can produce cytokines in a circadian manner (16) fine-tuning the immune response and playing an essential role in modulating the activity of the innate immune response.
Neutrophils recognize opsonized microorganisms and engulf them via phagocytosis, once internalized the microorganisms are stored in intracellular vacuoles called phagosomes where they are destroyed by an oxidative burst released by active NOX2 (11). However, NETs can be produced independently of NOX2 via vital NETosis using mitochondrial ROS (17). NOX2 is necessary for NET production although it is unknown which oxide is involved, it is speculated to be single oxygen, superoxide, hydrogen peroxide and hypochlorous acid. NOX2 is an electron-transfer complex assembled in the plasma membrane. Gp91phox is the catalytic subunit composed of the FAD-containing cytoplasmic domain, two b-type cytochromes and the p22 subunit. Neutrophil activation triggers phosphorylation events which activates enzyme activity of NOX2 by recruiting several cytoplasmic regulatory subunits to the cytoplasmic domain. Phagocytosis stimulates NOX2 assembly and electrons move through the NOX2 complex from NADPH so that oxygen loses an electron (11). The flow of electrons in NOX2 complex is in one direction, from NADPH in the cytosol to oxygen in the membrane. The pH of NOX2 phagosomes is constantly alkaline (pH 9) when the complex is active, in contrast to macrophage phagosomes.
Neutrophil dysfunction has been associated with adverse prognosis in a variety of diseases including sepsis, rheumatoid arthritis (RA), systemic lupus erythematosus (SLE), human immunodeficiency virus (HIV), mycobacterium tuberculosis infection and antineutrophil cytoplasmic antibody (ANCA)-associated vasculitis (AAV) (18–24). The production of NETs can contribute to autoimmune disease progression due to exposure to autoantigens within NETS, this occurs in several conditions such as RA, SLE and autoimmune small vessel vasculitis (25–27).
Neutrophil dysfunction is also evident in cancer whereby a high volume of intra-tumoral neutrophils is correlated with poor outcome (28). Emergency granulopoiesis typically associated with cancer progression contributes to poor patient survival, likely due to the generation of neutrophils with altered immune functions, namely immunosuppressive TANs and G-MDSC. Neutrophils contribute to the disease progression of cancer by favoring metastasis, angiogenesis and inhibiting anti-tumor immune cells, for example, inhibition of T cells via programmed death ligand 1 (PD-L1) (20).
The stages of neutrophil granulopoiesis are promyelocytes, myelocytes, metamyelocytes, band cells and segmented neutrophils (29). Several studies have identified a higher proportion of immature neutrophils is indicative of infection, particularly neonatal sepsis (30–32). This expansion of immature neutrophils in the bloodstream is known as left shift and can be measured using immature-to-total (I/T) neutrophil ratio (33) in the blood of adult patients with sepsis, the presence of immature band cells may be useful as a diagnostic marker of sepsis while immature myelocytes and metamyelocytes may be predictors of mortality (34). Immature neutrophils have been implicated in lung, breast and ovarian cancer and associated with poor prognosis (35). Therefore, the ability to differentiate neutrophil lineages is of paramount clinical importance in the setting of disease. However, it remains challenging to differentiate between stages of neutrophil granulopoiesis because there are no defined surface markers to identify immature and mature neutrophils.
Neutrophils normally have a short half-life of approximately 6-8 hours, hence studying them is a challenge as they need to be processed rapidly upon sampling (3). Measurement of absolute neutrophil counts (ANC) to detect neutrophilia and neutropenia is one of the most commonly used tests clinically (36). In this review, we propose a guide to aid in identifying the different lineages of neutrophils based on surface marker expression and correlate granule production to neutrophil function. We also explore the remarkable similarities between proposed neutrophil ‘subpopulations’.
Neutrophil “Subset” Nomenclature
In a pre-antibiotic era, Elie Metchnikoff and Paul Ehrlich were awarded the Nobel prize in 1908 for their discovery of phagocytic cells, macrophages and neutrophils (as they were subsequently named) (37). Since this discovery of granulocytes at the end of the nineteenth century, the cells Ehrlich termed “cells with polymorphous nuclei” have been referred to by many names, including phagocytes, polymorphonuclear neutrophils (PMN), myeloid derived suppressor cells (MDSC), low density neutrophils (LDN) and tumor associated neutrophils (TANS) (1). As Shakespeare remarked “A rose by any other name would smell as sweet”, different nomenclature has been used in the literature for neutrophils in the absence of clear biological differences. This had led to the misleading concept that biologically distinct populations of neutrophils exist, particularly in disease, when in fact these are all one adaptable cell type. Neutrophils are an incredibly plastic cell type that allows them to respond and adapt to a variety of stimuli, which in fact may explain the apparent biological differences between these neutrophil “subpopulations” (38).
Low Density Neutrophils (LDN) or Granulocytes (LDG)
Discontinuous density gradients are used to isolate neutrophils from whole blood. Normal-density neutrophils (NDN) usually reside in the high-density fraction but a subpopulation of neutrophils are found in the low-density fraction, which are known as low-density neutrophils (LDNs) or, less commonly, LD-PMN (39, 40). LDNs have been reported in a wide array of diseases including rheumatoid arthritis, systemic lupus erythematous (SLE), cancer, sepsis and asthma (41–43). LDNs display either an immature morphology with banded nuclei or myelocyte features, and are likely released in response to emergency granulopoiesis, or have a mature morphology with segmented nuclei (3, 20). Uí Mhaonaigh et al. found that CD16int/-CD10- LDNs resemble immature neutrophils while CD16+/CD10+ LDNs share morphological features with NDNs (44). Similar to descriptions of TANs, LDNs exhibit pro-tumorigenic and immunosuppressive functions (20).
There are several theories about the origin of LDNs. Firstly, LDNs are a mixture of mature and immature neutrophils, which may be immunomodulatory (44). Secondly, LDNs could be neutrophils after undergoing degranulation and hence display a lower density which could explain the presence of neutrophils in the low density fraction (3). Interestingly, after TGF-β stimulation in a mouse model, LDNs can be derived from mature neutrophils and in human studies, LDNs can be generated by incubation with Mycobacterium tuberculosis (45–47). As shown in Table 1, G-MDSC, TANs and LDN show almost the exact same surface marker profiles. TANs and LDN also display similar pro-tumor properties, making it biologically difficult to distinguish between these subpopulations. It is possible that the plasticity of the neutrophil and the influence of the tumor microenvironment may explain how all these neutrophil “subpopulations” are in fact a normal neutrophil under the influence of a distinct local environment. There is not enough scientific evidence to confirm that G-MDSC, TAN and LDN are unique cell populations.
Cassetta et al. suggest that information in the literature regarding neutrophil subsets, such as G-MDSC, LDN, Tan, is varied and contradictory due to the use of different models and isolation techniques. Surface markers on neutrophils isolated from murine models and non-human primates correlate poorly with corresponding human markers. Not only are cell surface markers different between mice and humans but some neutrophil subtypes, such as MDSC, are isolated from blood in humans but are studied at tissue level in mice (58). Therefore, standardized protocols are essential to gain further insight into the biological significance of neutrophil subtypes.
Myeloid Derived Suppressor Cells (MDSC)
MDSC were identified as myeloid cells that suppress immune responses and aid tumor progression in mouse models of cancer but not in humans (55, 62). The cells were named MDSC by Gabrilovich et al. (63). There are believed to be two subpopulations of MDSC: monocytic (Mo-MDSC) and granulocytic (G-MDSC or polymorphonuclear (PMN)-MDSC) (64, 65). Pillay et al. suggest that G-MDSC are a bona fide phenotype of neutrophils, which (unlike conventional neutrophils) are found in the low density fraction of peripheral blood (3, 64). Mo-MDSC are also found in the low density layer and are morphologically similar to monocytes (20). Interestingly, G-MDSC show similar morphology to mature neutrophils (66). MDSC display similar surface marker expression patterns to neutrophils but it is their suppressive functions which define this population: cluster of differentiation molecule (CD)66b+, CD16+, CD15+ and CD14- (56). As reviewed by Rosales in 2018, both Mo-MDSC and G-MDSC are low density, CD11b+, CD33+ and CD66b+, G-MDSC are human leukocyte antigen-DR isotype (HLA-DR)- and Mo-MDSC are HLA-DR-/low (3). Mo-MDSC differ from G-MDSC in being CD14+ and CD15-, whereas G-MDSC are CD14- and CD15+ (3, 57). G-MDSC are often further subdivided based on whether they are CD16- or CD16+/INT (56). G-MDSC are most likely a mixed low-density population of immature and mature neutrophils. A general consensus has been established for identification of G-MDSC whereby there must be, at a minimum, the following surface marker profile: CD15+, CD11b+, CD14-, HLA-DR-, and CD33mild (58).
Tumor-Associated Neutrophils (TAN)/N2
Tumor-associated neutrophils (TAN) may be polarized towards two potential phenotypes similar to macrophages: N1, which are anti-tumor, and N2, which are pro-tumor, however, these are limited to mouse models and not yet identified in humans. Each subpopulation has distinct functions, cytokines and gene expression profiles (50, 67). N2 have circular nuclei while N1 show hypersegmented nuclei (49). N2 are promoted by transforming growth factor-β (TGF-β) and N1 are recruited by interferon-β (IFN-β), N1 neutrophils are likely stimulated by the tumor microenvironment (49, 50, 68). Tumor cells use chemokines to attract TANs to the tumor site, such as the potent neutrophil chemoattractant CXCL8, which entrains the CXCR1 and CXCR2 expression on neutrophils. CXCR1 has been shown to contribute to angiogenesis and tumor progression (50, 69). TANs are believed to be distinct from normal density neutrophils (NDN) and G-MDSC as TANS exhibit high chemokine production, few granules and low ROS production (70). TANs may be evidence of the plasticity of neutrophils in response to factors in the tumor microenvironments of specific cancers, rather than a novel neutrophil subpopulation (20).
Neutrophil Granulopoiesis Stages Defined by Density, Morphology and Maturity
Maturity
Neutrophils are a heterogenous population comprised of phenotypically distinct subtypes during granulopoiesis (71). To date, two contrasting models have been described for the generation of blood cells from hematopoietic stem cells (HSC). These are the ‘classical model’ that has been used for generations and describes a cells ability to determine its cellular fate prior to single lineage commitment and is subsequently defined by its inability to differentiate into other progenitor cells. This model describes HSCs giving rise to either Common myeloid progenitors (CMPs) or common lymphoid progenitors (CLPs). The CMP then further differentiates into either a granulocyte monocyte progenitor (GMP) or a megakaryocyte erythroid progenitor (MEP). However, with the recent advancements in single cell sequencing, an ‘alternative model’ has been proposed which highlights that both CMPs and CLPs have mixed lineage potential defined by their transcription heterogeneity and their cellular fate is determined by external differentiation factors (72).
Transcription factors have long been known to regulate the commitment and subsequent activation of various myeloid derived cells from HSCs (73). This complex process involves the upregulation of and silencing of various developmental genes under the control of certain transcription factors (TF). The process that governs the development of CMPs to GMPs is dependent on the following transcription factors including CCAAT/enhancer-binding proteins (C/EBPs), GATA-1, and PU.1 (74). C/EBP-α, -β, and –ϵ have long been known to regulate neutrophil development in which mutations in either result in the development of myeloid leukemia (75, 76). Additional transcription factors including PU.1, and Irf8 induce CMPs to differentiate into monocytes whereas neutrophil differentiation involves a complex interplay of transcription factors Gfi-1, PU.1 and C/EBPs (77).
In the bone marrow, hematopoietic stem cells (HSCs) differentiate into myeloblasts, which in turn become promyelocytes, myelocytes, metamyelocytes, band cells and, lastly, segmented neutrophils (29) (Figure 1). Inside the bone marrow there are three compartments where neutrophils reside: stem cell pool, mitotic pool and post-mitotic pool (78). Undifferentiated progenitor cells such as HSCs, are found in the stem cell pool, the mitotic pool holds myeloblasts, promyelocytes and myelocytes and, finally, metamyelocytes, band cells and segmented neutrophils are localized within the post-mitotic pool (33). At the metamyelocyte stage, the neutrophil can no longer proliferate, signaling the start of terminal differentiation (79). In the absence of infection, there are more neutrophils stored in the bone marrow than in the circulation. In response to signals at the site of infection, only mature segmented neutrophils migrate out of the bone marrow in large numbers to the site of infection (80).
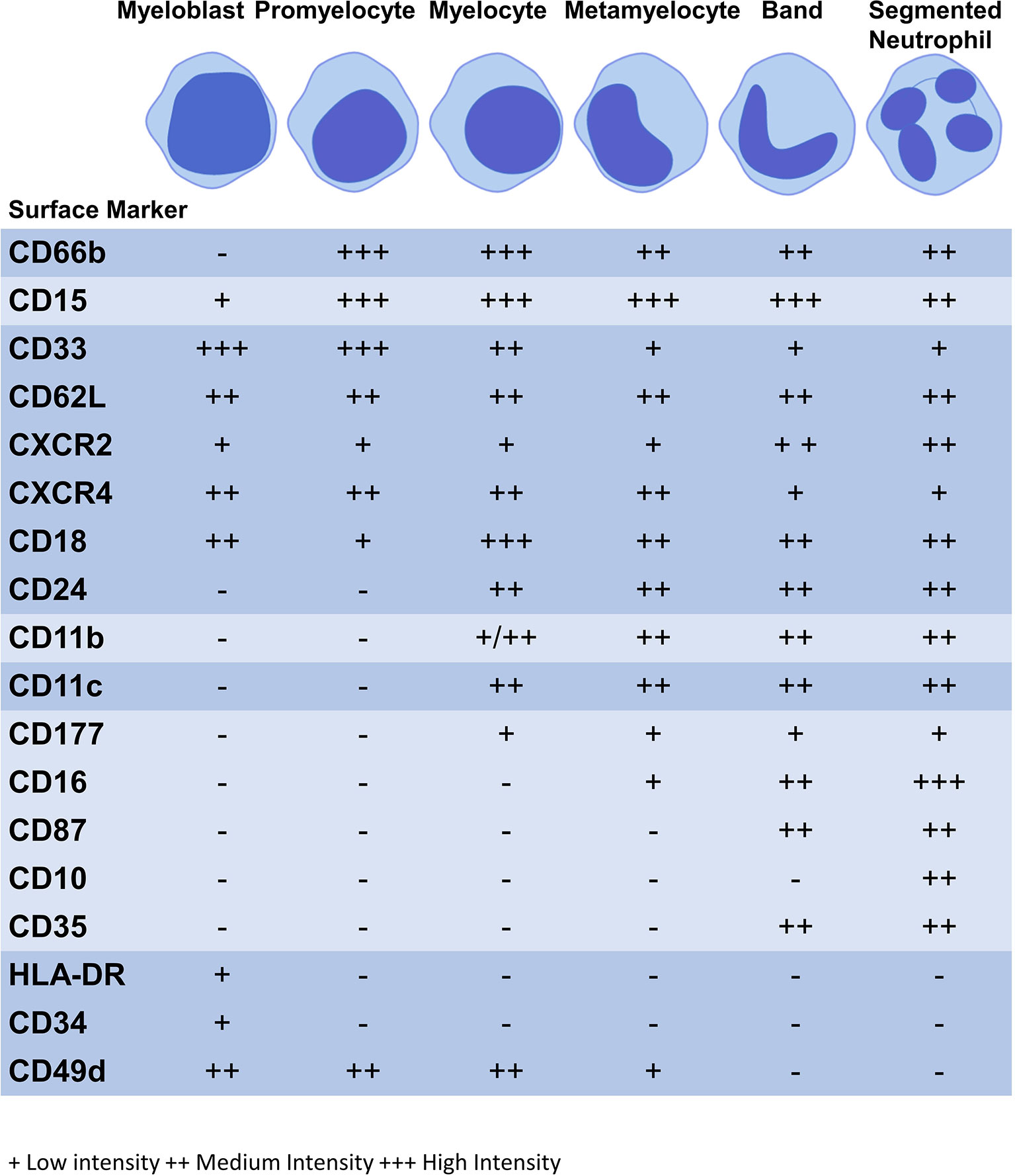
Figure 1 Expression of surface markers during granulopoiesis. Figure illustrates the surface marker expression at each stage of granulopoiesis; myeloblast, promyelocyte, myelocyte, metamyelocyte, band cell and segmented neutrophil. The intensity of the surface marker is shown whereby; low intensity (+), medium intensity (++) and high intensity (+++).
Terminal differentiation of GMPs into neutrophils involves the acquisition of neutrophil specific granule components at various stages of neutrophil maturation. This includes the storing of primary granules such as MPO and NE (at early stages), and, subsequently, Cathelicidin and lactoferrin (81). An important TF in this process is lymphoid enhancer factor-1 (Lef-1), deficiency of which results in impaired neutrophil maturation through the expression C/EBPα (82). Moreover TF Erg, Myb, Fos-like antigen (Fosl) 1, Fosl2, JunB proto-oncogene, B-cell lymphoma (Bcl) 6, Kruppel like factor (Klf) 6, and interferon regulatory factor 1 (Irf) 1 are known to regulate late stage neutrophil differentiation, revealing a complex network of TF involved in the process of granulopoiesis (83).
Neutrophil Density and Its Association With Neutrophil Development
The increased granularity and cell size that occurs with neutrophil maturity is directly proportional to a change in density. Therefore, neutrophil lineages can be separated using density gradient centrifugation (35). Mature neutrophils localize in normal to high density fractions, while immature neutrophils reside in low density fractions, although this is probably preparation dependent and the low density fraction is likely a mixed population of neutrophil maturity (35, 46).
Morphology
Neutrophil subtypes differ in their nuclear morphology. The most immature neutrophils, the myeloblasts, have large, sphere-shaped nuclei containing few nucleoli. Promyelocytes and myelocytes lack nucleoli and exhibit elevated chromatin condensation compared to myeloblasts. An indented nucleus is characteristic of a metamyelocyte. The band cell nucleus is shaped like a horseshoe and it constricts to form nuclear lobes, while segmented neutrophils are distinguishable by their segmented nucleus, with three to five lobes (84). The thin filaments which connect the lobes contribute to the migratory capability of mature neutrophils (85).
Neutrophil Surface Marker Expression During Granulopoiesis
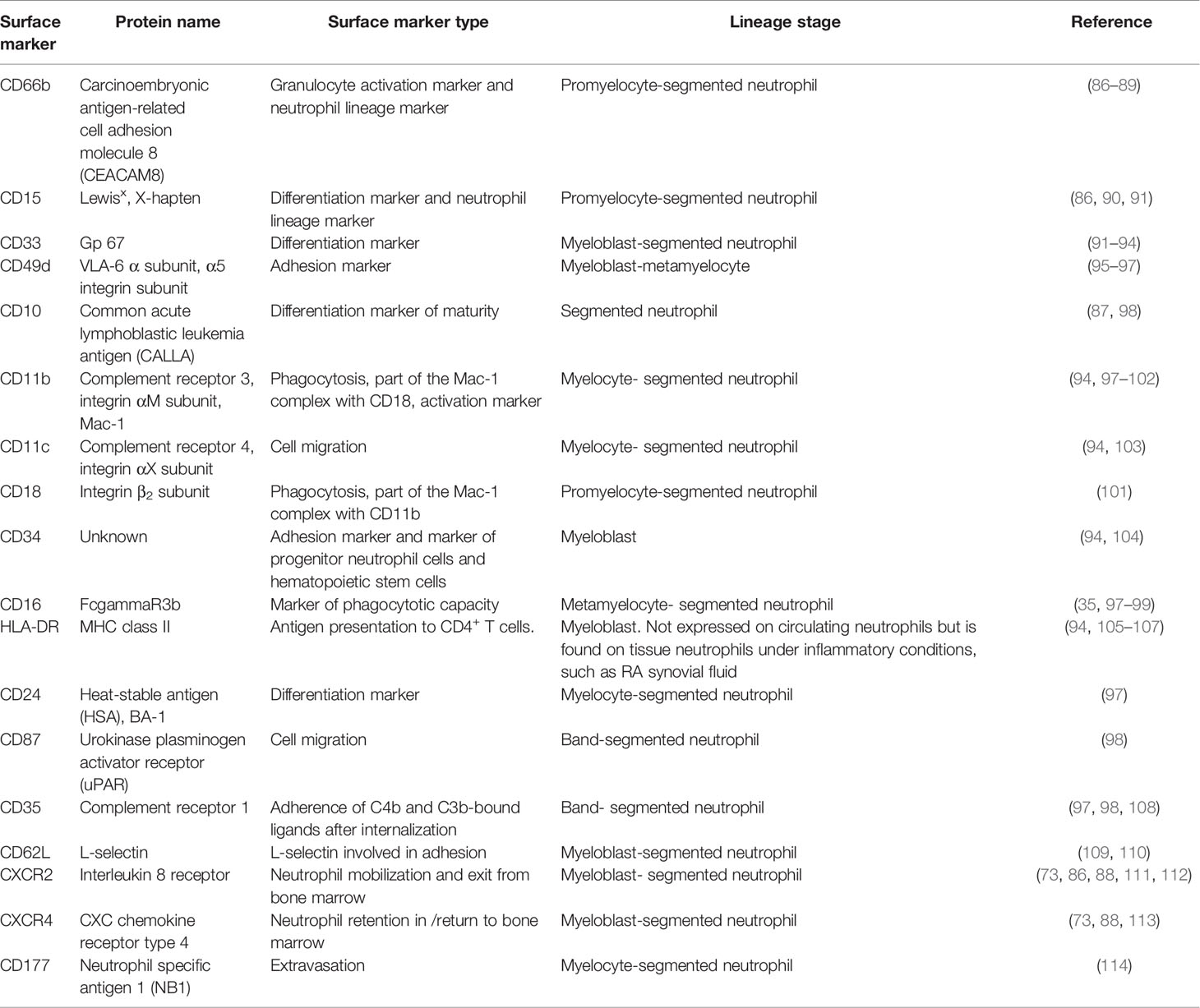
Neutrophil surface markers change to facilitate altered functions as the neutrophil matures (Table 2). During this process, immature surface markers are no longer be expressed (e.g. CD49d) and markers of maturity appear (e.g. CD10) (86). Although there is no consensus on human neutrophil phenotypic markers on flow cytometry the following are commonly used: CD11b+CD66b+CD15+CD14- (20). CD16 is a marker of phagocytic capacity and may be used to exclude CD16- eosinophils. CD14+ cells are considered to be mostly macrophages and monocytes, although some studies indicate that neutrophils express CD14 at low levels (115, 116). CD16 is exclusively expressed at the metamyelocyte stage and is highly expressed by banded and segmented neutrophils, while activation marker CD11b is only found at the myelocyte stage onwards (94, 97–100). CD11b and CD18 form the Mac-1 complex which plays a role in phagocytosis and migration with CD18 expression commencing at the myeloblast stage of granulopoiesis (86). Differentiation marker CD15 and activation marker is found on all neutrophil subpopulations and CD66b from the promyelocyte stage; these are also core markers of neutrophil lineage (86–88). Surface markers may be expressed at low, medium or high intensities, see Figure 1 (86, 88).
Markers of Maturity and Immaturity
HLA-DR (major histocompatibility complex II, MHC class II), which is involved in antigen presentation to CD4+ T cells, and CD34, an adhesion marker, are two markers exclusively expressed on the most immature neutrophil, the myeloblast (94, 106). HLA-DR is not present on circulating neutrophils but is expressed on the surface of tissue neutrophils under specific inflammatory conditions, such as RA synovial fluid (105). CD33 is a differentiation marker found on myeloid blast cells in acute myeloid leukemia (AML) (93). CD33 surface expression is gradually downregulated from the myeloblast stage to segmented neutrophil, with a low level expressed on the latter (91, 92).
CD10, a marker of differentiation and maturity is found only on mature segmented neutrophils, and absent on immature neutrophils. Immature CD10- neutrophils may be important drivers of inflammatory disease and neutropenia (87, 88, 98). Reduced surface expression of CD10 and CD16 on granulocytes predicts poor outcome in sepsis patients (117). CD10dim immature neutrophils have been implicated in the immunosuppression observed in sepsis (118).
Conflicting results exist for the marker of maturation CD24. Elghetany et al. found that CD24 expression begins at the myelocyte stage and is a marker of maturation (97). In contrast, Hernández-Campo et al. found that CD24 is present on CD34-/low myeloblasts, is highly expressed by myelocytes and decreases from metamyelocyte to segmented neutrophil (119).
Markers of Adhesion and Extravasation
The integrin very late antigen complex 4 (VLA-4) is involved in the adhesion of hematopoietic progenitor cells and leukemic blast cells in AML (120). VLA-4 is composed of CD49d and CD29, CD49d being a commonly used marker of neutrophil immaturity (120, 121). Promyelocytes, myelocytes and metamyelocytes are CD49d+ while more mature bands and segmented neutrophils are CD49d- (97). Given the role of CD49d in the recruitment of progenitor cells leukemic blast cells and their migration from the bone marrow, it is likely that CD49d must be expressed on neutrophil blast cells (95, 96). Interestingly, CD49d expression reappears on the surface of aged neutrophils (122).
High CD162, or P-selectin glycoprotein ligand-1 (PSGL1), is an L-selectin molecule involved in adhesion that is found on the surface of myeloblasts to segmented neutrophils (109, 123). CD35, involved in the adherence of C4b and C3b bound ligands after internalization, and CD87, involved in cell migration, are solely expressed by bands and segmented neutrophils (97, 98). CD11c is a another marker of cell migration and is expressed at the myelocyte stage (94, 103).
Surface expression of CD177, a receptor involved in extravasation and surface expression of proteinase-3, begins at the myelocyte stage (114). Interestingly, there is an increase in CD177+ circulating neutrophils in diseases such as sepsis, vasculitis and SLE (124, 125).
Chemokines and Chemokine Receptors
Immature neutrophil retention in the bone marrow is achieved by the cross-talk between CXCR4 expressed on neutrophils and CXCL12 by bone marrow stromal cells. CXCR4 is involved in neutrophil retention in the bone marrow and return of aged neutrophils to the bone marrow (73, 126). Upregulated expression of CXCR4 on segmented neutrophils ready to undergo senescence or apoptosis triggers their return to the bone marrow where they are engulfed by macrophages (113, 127). While both immature and mature neutrophils express CXCR4, immature neutrophils may express a higher level of CXCR4 than mature cells (73, 88). The chemokine receptor CXCR2 plays a role in neutrophil mobilization and exit from the bone marrow (86, 111). Immature neutrophils before the band stage exhibit reduced CXCR2 surface expression compared to bands and segmented neutrophils (73, 88). However, CXCR4 and CXCR2 may not be useful in differentiating between neutrophil subtypes due to similar expression levels in each population.
Thus, surface markers on neutrophils demonstrates that flow cytometry, or other techniques, could be employed to separate neutrophil lineages based on surface marker expression. This information highlights that banded neutrophils and segmented neutrophils are the most similar in their surface expression and may be the most difficult to differentiate due to a lack of unique markers on either cell type (110, 128). Some surface markers undergo changes following migration, e.g. CD62L and activation, e.g. CD62L/CD11b/CD18, which makes it difficult to identify highly specific markers of maturity (35).
Neutrophil Granules
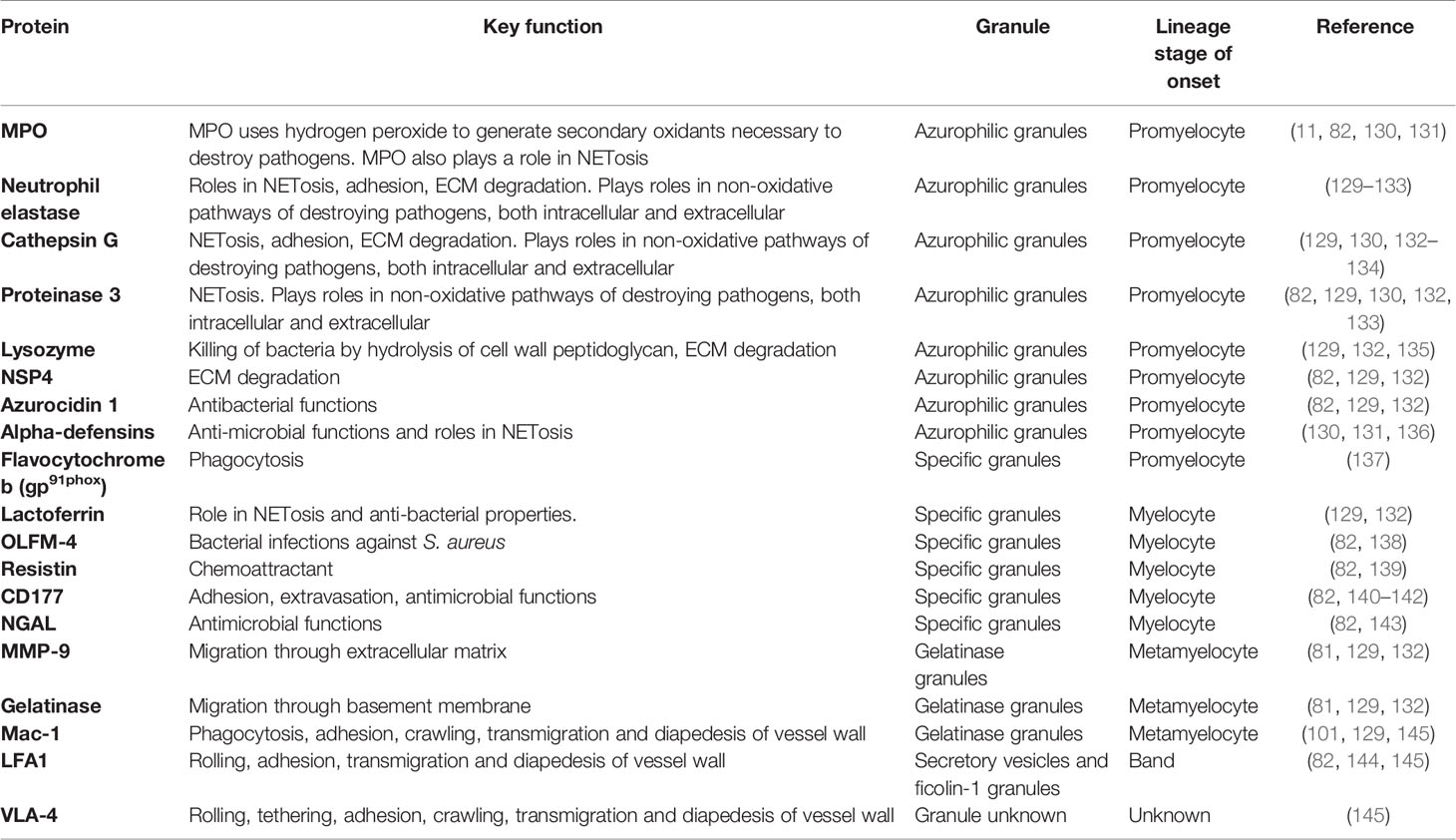
The production of neutrophil granules begins as the immature neutrophil transitions from a myeloblast to a promyelocyte and continue to be produced up to the segmented neutrophil stage (129). There are four main groups of granules: azurophilic (primary) granules, specific (secondary) granules, gelatinase (tertiary) granules and (most recently discovered) ficolin-1 granules (Figure 2, Table 3). Neutrophils also contain secretory vesicles, which are not defined as granules (101, 144). A comprehensive review of granulopoiesis granule production and associated transcription factors is provided by Lawrence et al. (82).
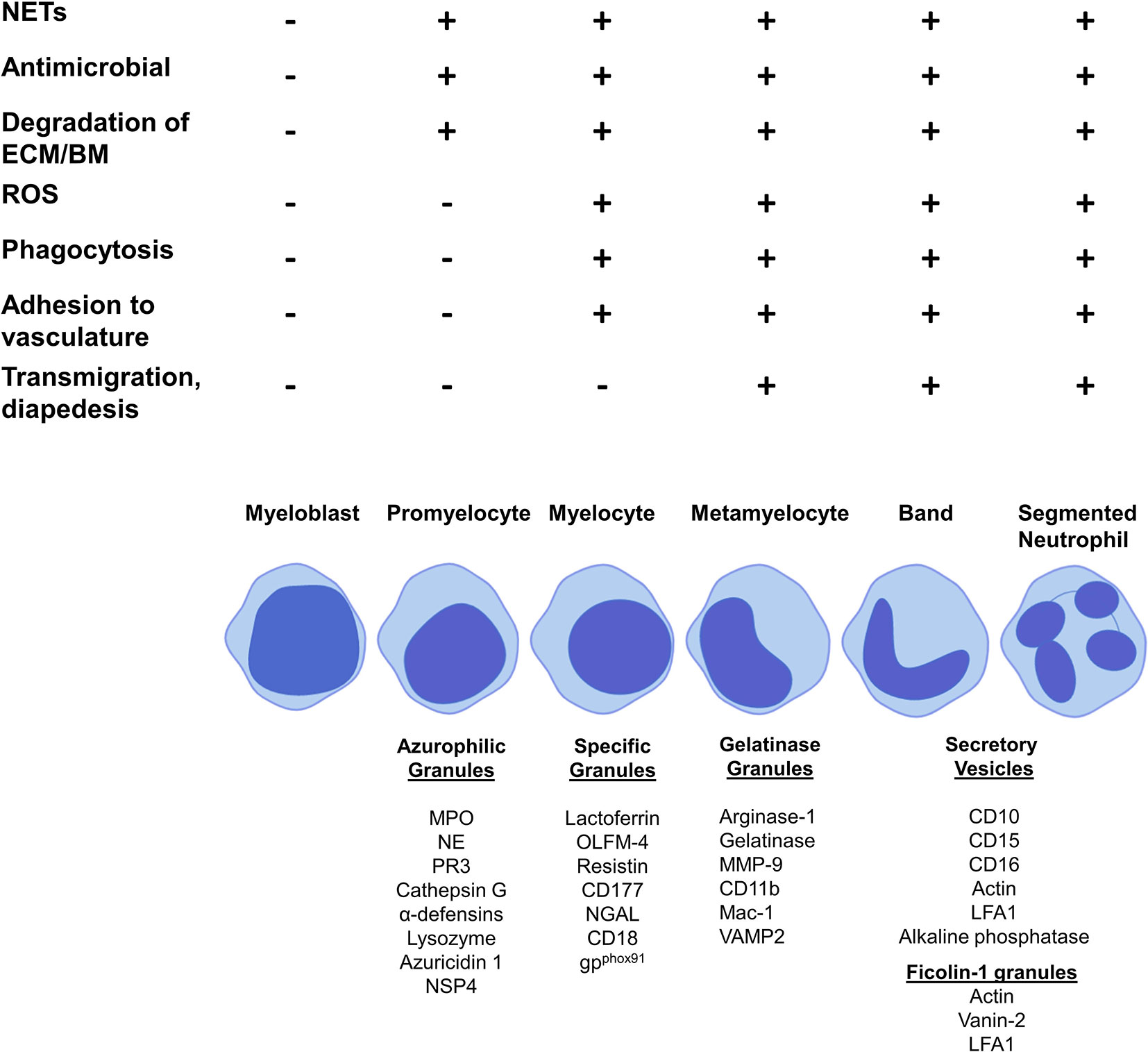
Figure 2 Granule production and functionality during granulopoiesis. Figure shows granule production during the stages of granulopoiesis. Granule formation begins with azurophilic granules at the promyelocyte stage, specific granules are first formed by myelocytes, gelatinase granules are formed at the metamyelocyte stage and only band cells and segmented neutrophils can form ficolin-1 granules and secretory vesicles. NET formation, antimicrobial functions and degradation of the extracellular membrane and basement membrane begin at the promyelocytes stage with azurophilic granule formation. ROS production, phagocytosis and adhesion to vasculature starts at the myelocytes stage with the formation of specific granules. Metamyelocytes are the first stage to perform transmigration and diapedesis through a vessel wall, which correlates with gelatinase granule formation.
Prominent production of the heme enzyme myeloperoxidase (MPO) is the main characteristic feature of azurophilic granules. MPO makes up 5% of the weight of a neutrophil (82). Azurophilic granules contain MPO, neutrophil elastase (NE), proteinase 3 (PR3), lysozyme, azurocidin 1, cathepsin G and neutrophil serine protease 4 (NSP4) (129, 132, 133). The main neutrophil antimicrobial peptides, α-defensins, are also stored in azurophilic granules (136). The formation of azurophilic granules begins at the promyelocyte phase (132) (Figure 2). Cathepsin G and NE may contribute to downregulation of neutrophil adhesion by proteolysis of CD62L, preventing its binding to P-selectin on the endothelium (146). Lysozyme mediates killing of bacteria by hydrolyzing the peptidoglycan cell wall (135). NE, PR3, MPO, α-defensins, and cathepsin G are all involved in NETosis (130, 131, 147–149). PR3, cathepsin G, NE and NSP4 contribute to extracellular matrix (ECM) degradation by mediating proteolysis of components such as laminin, elastin and type IV collagen (82).
Specific granules contain large amounts of lactoferrin, which plays a key role in NETosis (129, 132). Other components are olfactomedin-4 (OLFM-4), which helps fight bacterial infections such as Staphylococcus aureus, resistin, a pro-inflammatory cytokine and T-cell chemoattractant, the receptor CD177 and antimicrobial neutrophil gelatinase-associated protein lipocalin (NGAL) (82, 138–140, 143). Specific granules are formed at the myelocyte stage (82) (Figure 2).
Gelatinase granules store arginase 1, matrix metallopeptidase (MMP)-9, gelatinase and surface receptors including CD11b/CD18 (Mac-1) (129, 132). The main function of gelatinase granules such as MMP-9 and gelatinase is to facilitate the breakdown of the ECM and basement membrane, aiding migration into tissue (81). Metamyelocytes begin to form gelatinase granules (101) (Figure 2).
Segmented neutrophils form ficolin-1 granules, the contents of which play roles in migration and adhesion, for example, actin and vanin-2 (82, 144). Segmented neutrophils also produce secretory vesicles, these contain actin, alkaline phosphatase and receptors including CD11b, CD10 and CD16 (82, 101, 144) (Figure 2).
Neutrophil Granules Involved in Neutrophil Extracellular Traps
Active neutrophils undergo NETosis, a form of cell death that involves releasing decondensed chromatin in response to stimuli, such as bacteria (7). NETs contribute to a wide range of inflammatory diseases including as rheumatoid arthritis, systemic lupus erythematosus and autoimmune small vessel vasculitis (25–27). NETs exhibit anti-microbial functions by trapping extracellular microbes (128). The formation of NETs in response to pathogens involves key granular components: NE, PR3, MPO, α-defensins, cathepsin G, lactoferrin, and reactive oxygen species (ROS) (130, 131, 147–149). NETs are induced by the translocation of NE from granules to the nucleus where it cleaves histones, resulting in chromatin decondensation and breakdown of the plasma membrane (150). With the exception of lactoferrin, which is located in specific granules, all other major proteins involved in NET formation are located in azurophilic granules whose formation begins at the promyelocyte stage. Therefore, NET formation likely begins at this stage of granulopoiesis (129, 132) (Figure 2). To our knowledge, no study has identified any surface markers of NETosis.
Neutrophil Granules Involved in ROS Production and Phagocytosis
Neutrophils are the greatest producers of ROS in response to infection. The NADPH oxidase complex is responsible for the production of ROS, which drives the antimicrobial function of neutrophils by inducing degranulation, NETosis and release of pro-inflammatory cytokines (151). Phagocytosis is the process of engulfing and destroying pathogens (152). The NADPH oxidase complex localizes in the phagocytic vacuole and causes a ‘respiratory burst’ of oxygen, producing superoxide (O2-) during phagocytosis. A flavocytochrome b, gp91phox, enables electron transfer and interacts with NADPH in the phagocytic vacuole (153). CD11b/CD18 (Mac-1) and flavocytochrome b are formed and stored in gelatinase and specific granules, indicating that these granules play a role in phagocytosis (101, 129, 137). This implies that phagocytosis begins at the myelocyte stage when these specific granules are formed (129, 132) (Figure 2).
Neutrophil Granules Involved in Degranulation
The steps involved in neutrophil degranulation are, firstly, translocation of granules to the target membrane, which is achieved by the assembly of microtubules and actin rearrangement. G protein, Rab, and snap receptor (SNARE) membrane trafficking proteins facilitate docking of granules which can then be released from the neutrophil by exocytosis (154). SNARE protein vesicle-associated membrane protein 2 (VAMP2) aids the exocytosis of granules from the neutrophil and is formed in gelatinase granules (8, 101). Granules produced at more advanced stages of neutrophil granulopoiesis are more likely to be released by exocytosis than those formed in early stages. Degranulation appears to occur at a low level with the formation of azurophilic granules by promyelocytes and increases with each subsequent stage of granulopoiesis, with secretory vesicles produced by bands and segmented neutrophils accounting for the highest degree of degranulation (154, 155) (Figure 2).
Neutrophil Granules Involved in Extravasation
Neutrophil extravasation is the process by which circulating neutrophils pass through the blood vessel endothelium to reach the site of infection. The neutrophil extravasation cascade is initiated by leukocytes at the site of infection, which release inflammatory signals (e.g. histamine) to induce changes in endothelial cells, for instance the upregulation of P-selectin. Extravasation is facilitated by receptor-ligand interactions between the neutrophil and endothelial cells. The steps involved include neutrophil tethering to endothelial cells via receptor-ligand interactions, adhesion to the endothelium, crawling, rolling and, finally, transmigration through gaps between endothelial cells (145).
CD177 has a high affinity for platelet endothelial cell adhesion molecule (PECAM-1), allowing the neutrophil to pass through the endothelium (141, 142). CD177 and PR3 are co-expressed on the neutrophil surface and may co-operate to promote extravasation, although a recent study has indicated that CD177 may have an inhibitory effect on PR3 (141, 156). CD177 is stored in specific granules starting at the myelocyte stage and is expressed on the surface of myelocytes through to segmented neutrophils (114, 140).
Mac-1 (CD11b/CD18) plays a vital role in neutrophil extravasation. It binds to endothelial markers ICAM1 and ICAM2 and facilitates neutrophil adhesion, crawling along the blood vessel wall, transmigration and diapedesis (145). The Mac-1 complex can only be formed when both CD11b and CD18 have been produced; CD18 is produced by promyelocytes in specific granules and CD11b is produced by myelocytes in gelatinase granules. Therefore, Mac-1 is first formed at the myelocyte stage (101) (Figure 2).
Leucocyte function antigen 1 (LFA1) is composed of CD11a and CD18 and is involved in neutrophil rolling and adhesion by binding to ICAM1 and transmigration and diapedesis via binding to junctional adhesion molecule A (JAM-A) on the endothelium (145). LFA1 is stored in secretory vesicles and ficolin-1 granules and, therefore is only produced by bands and segmented neutrophils (82, 144).
There are several other adhesion molecules involved in neutrophil extravasation with unclear granule location; these are β2-integrin, L-selectin (CD62L), CD44 and CD49. In terms of extravasation, VLA-4 (CD49D/CD29) aids neutrophil tethering, rolling, adhesion, transmigration and diapedesis via VCAM1 or JAM-B. Interestingly, VLA-4 is produced by myelocytes but the specific granule that produces VLA-4 is unknown (145).
Conclusions
The neutrophil is an adaptable cell type capable of rapidly responding to changes in its environment. However, the neutrophil has been given many names in the literature, suggesting that multiple unique populations exist despite biological similarities. There is a lack of standard techniques for isolating neutrophils, which may account for varied results in the literature, and differences between these neutrophil populations. Neutrophils from murine models and humans have distinct patterns of surface markers that make it difficult to draw significant conclusions about the idiosyncratic nature of neutrophil populations such as MDSC, LDN and Tan. Therefore, standardized protocols are necessary to gain further insight into the biological significance of neutrophil populations and determine whether they are genuine, distinct populations or a result of activation during isolation or differences in the biology of mice and humans.
Different neutrophil nomenclature is often necessary, for instance LDN and NDN to differentiate between neutrophils of different densities and since LDNs are often present at higher levels in disease. Minimal nomenclature based on biological properties such as density should be maintained. We suggest using the longstanding nomenclature describing mature and immature neutrophils with immature types classified as band forms, myelocytes, promyelocytes and metamyelocytes with clarification of the surface markers for each on flow cytometry. This can then standardize the cell types across all disease states rather than the plethora of names such as MDSC, LDN and TAN often describing similar cells. The detailed function of each cell stage of maturity, granules and surface markers could be clarified. This would allow targeted interventions in distinct cell types that could be used in areas from cancer to other inflammatory disorders. In the future which would allow standardized classification and allow more extensive collaboration to characterize all functional and phenotypic variation among all the cells with this classification system.
Author Contributions
EM, AU, and RW conducted the research and wrote the manuscript. EJM supervised the work and co-wrote and edited the manuscript. AD, TH, LK, NS, and ML assisted in writing and revision of the manuscript. All authors contributed to the article and approved the submitted version.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Kay AB. Paul Ehrlich and the Early History of Granulocytes. Microbiol Spectr (2016) 4(4):1–2. doi: 10.1128/microbiolspec.MCHD-0032-2016
2. Kobayashi SD, DeLeo FR. Role of neutrophils in innate immunity: a systems biology-level approach. Wiley Interdiscip Rev Syst Biol Med (2009) 1(3):309–33. doi: 10.1002/wsbm.32
3. Rosales C. Neutrophil: A Cell with Many Roles in Inflammation or Several Cell Types? Front Physiol (2018) 9:113. doi: 10.3389/fphys.2018.00113
4. Amulic B, Cazalet C, Hayes GL, Metzler KD, Zychlinsky A. Neutrophil function: from mechanisms to disease. Annu Rev Immunol (2012) 30:459–89. doi: 10.1146/annurev-immunol-020711-074942
6. Segal BH, Veys P, Malech H, Cowan MJ. Chronic granulomatous disease: lessons from a rare disorder. Biol Blood Marrow Transplant (2011) 17(1 Suppl):S123–31. doi: 10.1016/j.bbmt.2010.09.008
7. Brinkmann V, Reichard U, Goosmann C, Fauler B, Uhlemann Y, Weiss DS, et al. Neutrophil extracellular traps kill bacteria. Science (2004) 303(5663):1532–5. doi: 10.1126/science.1092385
8. Lacy P. Mechanisms of degranulation in neutrophils. Allergy Asthma Clin Immunol (2006) 2(3):98–108. doi: 10.1186/1710-1492-2-3-98
9. Decoursey TE, Ligeti E. Regulation and termination of NADPH oxidase activity. Cell Mol Life Sci (2005) 62(19-20):2173–93. doi: 10.1007/s00018-005-5177-1
10. Nunes P, Demaurex N, Dinauer MC. Regulation of the NADPH oxidase and associated ion fluxes during phagocytosis. Traffic (2013) 14(11):1118–31. doi: 10.1111/tra.12115
11. Winterbourn CC, Kettle AJ, Hampton MB. Reactive Oxygen Species and Neutrophil Function. Annu Rev Biochem (2016) 85:765–92. doi: 10.1146/annurev-biochem-060815-014442
12. Chavakis E, Choi EY, Chavakis T. Novel aspects in the regulation of the leukocyte adhesion cascade. Thromb Haemost (2009) 102(2):191–7. doi: 10.1160/TH08-12-0844
13. Adrover JM, Nicolas-Avila JA, Hidalgo A. Aging: A Temporal Dimension for Neutrophils. Trends Immunol (2016) 37(5):334–45. doi: 10.1016/j.it.2016.03.005
14. Vogt KL, Summers C, Chilvers ER, Condliffe AM. Priming and de-priming of neutrophil responses in vitro and in vivo. Eur J Clin Invest (2018) 48(Suppl 2):e12967. doi: 10.1111/eci.12967
15. Scheiermann C, Kunisaki Y, Frenette PS. Circadian control of the immune system. Nat Rev Immunol (2013) 13(3):190–8. doi: 10.1038/nri3386
16. Tecchio C, Cassatella MA. Neutrophil-derived chemokines on the road to immunity. Semin Immunol (2016) 28(2):119–28. doi: 10.1016/j.smim.2016.04.003
17. Petretto A, Bruschi M, Pratesi F, Croia C, Candiano G, Ghiggeri G, et al. Neutrophil extracellular traps (NET) induced by different stimuli: A comparative proteomic analysis. PLoS One (2019) 14(7):e0218946. doi: 10.1371/journal.pone.0218946
18. Zhang F, Liu AL, Gao S, Ma S, Guo SB. Neutrophil Dysfunction in Sepsis. Chin Med J (Engl) (2016) 129(22):2741–4. doi: 10.4103/0366-6999.193447
19. Chen W, Wang Q, Ke Y, Lin J. Neutrophil Function in an Inflammatory Milieu of Rheumatoid Arthritis. J Immunol Res (2018) 2018:8549329. doi: 10.1155/2018/8549329
20. Grecian R, Whyte MKB, Walmsley SR. The role of neutrophils in cancer. Br Med Bull (2018) 128(1):5–14. doi: 10.1093/bmb/ldy029
21. Hensley-McBain T, Klatt NR. The Dual Role of Neutrophils in HIV Infection. Curr HIV/AIDS Rep (2018) 15(1):1–10. doi: 10.1007/s11904-018-0370-7
22. Jones BE, Yang J, Muthigi A, Hogan SL, Hu Y, Starmer J, et al. Gene-Specific DNA Methylation Changes Predict Remission in Patients with ANCA-Associated Vasculitis. J Am Soc Nephrol (2017) 28(4):1175–87. doi: 10.1681/ASN.2016050548
23. Muefong CN, Sutherland JS. Neutrophils in Tuberculosis-Associated Inflammation and Lung Pathology. Front Immunol (2020) 11:962. doi: 10.3389/fimmu.2020.00962
24. Wirestam L, Arve S, Linge P, Bengtsson AA. Neutrophils-Important Communicators in Systemic Lupus Erythematosus and Antiphospholipid Syndrome. Front Immunol (2019) 10:2734. doi: 10.3389/fimmu.2019.02734
25. Kessenbrock K, Krumbholz M, Schonermarck U, Back W, Gross WL, Werb Z, et al. Netting neutrophils in autoimmune small-vessel vasculitis. Nat Med (2009) 15(6):623–5. doi: 10.1038/nm.1959
26. Khandpur R, Carmona-Rivera C, Vivekanandan-Giri A, Gizinski A, Yalavarthi S, Knight JS, et al. NETs are a source of citrullinated autoantigens and stimulate inflammatory responses in rheumatoid arthritis. Sci Transl Med (2013) 5(178):178ra40. doi: 10.1126/scitranslmed.3005580
27. Leffler J, Martin M, Gullstrand B, Tyden H, Lood C, Truedsson L, et al. Neutrophil extracellular traps that are not degraded in systemic lupus erythematosus activate complement exacerbating the disease. J Immunol (2012) 188(7):3522–31. doi: 10.4049/jimmunol.1102404
28. Shen M, Hu P, Donskov F, Wang G, Liu Q, Du J. Tumor-associated neutrophils as a new prognostic factor in cancer: a systematic review and meta-analysis. PLoS One (2014) 9(6):e98259. doi: 10.1371/journal.pone.0098259
29. Borregaard N. Neutrophils, from marrow to microbes. Immunity (2010) 33(5):657–70. doi: 10.1016/j.immuni.2010.11.011
30. Greenberg DN, Yoder BA. Changes in the differential white blood cell count in screening for group B streptococcal sepsis. Pediatr Infect Dis J (1990) 9(12):886–9. doi: 10.1097/00006454-199012000-00006
31. Hornik CP, Benjamin DK, Becker KC, Benjamin DK Jr, Li J, Clark RH, et al. Use of the complete blood cell count in early-onset neonatal sepsis. Pediatr Infect Dis J (2012) 31(8):799–802. doi: 10.1097/INF.0b013e318256905c
32. Newman TB, Puopolo KM, Wi S, Draper D, Escobar GJ. Interpreting complete blood counts soon after birth in newborns at risk for sepsis. Pediatrics (2010) 126(5):903–9. doi: 10.1542/peds.2010-0935
34. Mare TA, Treacher DF, Shankar-Hari M, Beale R, Lewis SM, Chambers DJ, et al. The diagnostic and prognostic significance of monitoring blood levels of immature neutrophils in patients with systemic inflammation. Crit Care (2015) 19:57. doi: 10.1186/s13054-015-0778-z
35. Mackey JBG, Coffelt SB, Carlin LM. Neutrophil Maturity in Cancer. Front Immunol (2019) 10:1912. doi: 10.3389/fimmu.2019.01912
36. Hoang AN, Jones CN, Dimisko L, Hamza B, Martel J, Kojic N, et al. Measuring neutrophil speed and directionality during chemotaxis, directly from a droplet of whole blood. Technology (Singap World Sci) (2013) 1(1):49. doi: 10.1142/S2339547813500040
37. Gordon S. Elie Metchnikoff, the Man and the Myth. J Innate Immun (2016) 8(3):223–7. doi: 10.1159/000443331
38. Cuartero MI, Ballesteros I, Moraga A, Nombela F, Vivancos J, Hamilton JA, et al. N2 neutrophils, novel players in brain inflammation after stroke: modulation by the PPARgamma agonist rosiglitazone. Stroke (2013) 44(12):3498–508. doi: 10.1161/STROKEAHA.113.002470
39. Davis RE, Sharma S, Conceicao J, Carneiro P, Novais F, Scott P, et al. Phenotypic and functional characteristics of HLA-DR(+) neutrophils in Brazilians with cutaneous leishmaniasis. J Leukoc Biol (2017) 101(3):739–49. doi: 10.1189/jlb.4A0915-442RR
40. Galdiero MR, Varricchi G, Loffredo S, Mantovani A, Marone G. Roles of neutrophils in cancer growth and progression. J Leukoc Biol (2018) 103(3):457–64. doi: 10.1002/JLB.3MR0717-292R
41. Fu J, Tobin MC, Thomas LL. Neutrophil-like low-density granulocytes are elevated in patients with moderate to severe persistent asthma. Ann Allergy Asthma Immunol (2014) 113(6):635–40.e2. doi: 10.1016/j.anai.2014.08.024
42. Hacbarth E, Kajdacsy-Balla A. Low density neutrophils in patients with systemic lupus erythematosus, rheumatoid arthritis, and acute rheumatic fever. Arthritis Rheumatol (1986) 29(11):1334–42. doi: 10.1002/art.1780291105
43. Sagiv JY, Voels S, Granot Z. Isolation and Characterization of Low- vs. High-Density Neutrophils in Cancer. Methods Mol Biol (2016) 1458:179–93. doi: 10.1007/978-1-4939-3801-8_13
44. Ui Mhaonaigh A, Coughlan AM, Dwivedi A, Hartnett J, Cabral J, Moran B, et al. Low Density Granulocytes in ANCA Vasculitis Are Heterogenous and Hypo-Responsive to Anti-Myeloperoxidase Antibodies. Front Immunol (2019) 10:2603. doi: 10.3389/fimmu.2019.02603
45. Deng Y, Ye J, Luo Q, Huang Z, Peng Y, Xiong G, et al. Low-Density Granulocytes Are Elevated in Mycobacterial Infection and Associated with the Severity of Tuberculosis. PLoS One (2016) 11(4):e0153567. doi: 10.1371/journal.pone.0153567
46. Sagiv JY, Michaeli J, Assi S, Mishalian I, Kisos H, Levy L, et al. Phenotypic diversity and plasticity in circulating neutrophil subpopulations in cancer. Cell Rep (2015) 10(4):562–73. doi: 10.1016/j.celrep.2014.12.039
47. Su R, Peng YP, Deng Z, Deng YT, Ye JQ, Guo Y, et al. Mycobacterium tuberculosis Infection Induces Low-Density Granulocyte Generation by Promoting Neutrophil Extracellular Trap Formation via ROS Pathway. Front Microbiol (2019) 10:1468. doi: 10.3389/fmicb.2019.01468
48. Ssemaganda A, Kindinger L, Bergin P, Nielsen L, Mpendo J, Ssetaala A, et al. Characterization of neutrophil subsets in healthy human pregnancies. PloS One (2014) 9(2):e85696. doi: 10.1371/journal.pone.0085696
49. Fridlender ZG, Sun J, Kim S, Kapoor V, Cheng G, Ling L, et al. Polarization of tumor-associated neutrophil phenotype by TGF-beta: “N1” versus “N2” TAN. Cancer Cell (2009) 16(3):183–94. doi: 10.1016/j.ccr.2009.06.017
50. Sionov RV, Fridlender ZG, Granot Z. The Multifaceted Roles Neutrophils Play in the Tumor Microenvironment. Cancer Microenviron (2015) 8(3):125–58. doi: 10.1007/s12307-014-0147-5
51. Fortunati E, Kazemier KM, Grutters JC, Koenderman L, Van den Bosch v J. Human neutrophils switch to an activated phenotype after homing to the lung irrespective of inflammatory disease. Clin Exp Immunol (2009) 155(3):559–66. doi: 10.1111/j.1365-2249.2008.03791.x
52. Eruslanov EB, Bhojnagarwala PS, Quatromoni JG, Stephen TL, Ranganathan A, Deshpande C, et al. Tumor-associated neutrophils stimulate T cell responses in early-stage human lung cancer. J Clin Invest (2014) 124(12):5466–80. doi: 10.1172/JCI77053
53. Uribe-Querol E, Rosales C. Neutrophils in Cancer: Two Sides of the Same Coin. J Immunol Res (2015) 2015:983698. doi: 10.1155/2015/983698
54. Eruslanov EB. Phenotype and function of tumor-associated neutrophils and their subsets in early-stage human lung cancer. Cancer Immunol Immunother (2017) 66(8):997–1006. doi: 10.1007/s00262-017-1976-0
55. Kusmartsev SA, Li Y, Chen SH. Gr-1+ myeloid cells derived from tumor-bearing mice inhibit primary T cell activation induced through CD3/CD28 costimulation. J Immunol (2000) 165(2):779–85. doi: 10.4049/jimmunol.165.2.779
56. Dumitru CA, Moses K, Trellakis S, Lang S, Brandau S. Neutrophils and granulocytic myeloid-derived suppressor cells: immunophenotyping, cell biology and clinical relevance in human oncology. Cancer Immunol Immunother (2012) 61(8):1155–67. doi: 10.1007/s00262-012-1294-5
57. Favaloro J, Liyadipitiya T, Brown R, Yang S, Suen H, Woodland N, et al. Myeloid derived suppressor cells are numerically, functionally and phenotypically different in patients with multiple myeloma. Leuk Lymphoma (2014) 55(12):2893–900. doi: 10.3109/10428194.2014.904511
58. Cassetta L, Baekkevold ES, Brandau S, Bujko A, Cassatella MA, Dorhoi A, et al. Deciphering myeloid-derived suppressor cells: isolation and markers in humans, mice and non-human primates. Cancer Immunol Immunother (2019) 68(4):687–97. doi: 10.1007/s00262-019-02302-2
59. Khaled YS, Ammori BJ, Elkord E. Increased levels of granulocytic myeloid-derived suppressor cells in peripheral blood and tumour tissue of pancreatic cancer patients. J Immunol Res (2014) 2014:879897. doi: 10.1155/2014/879897
60. Kang X, Zhang X, Liu Z, Xu H, Wang T, He L, et al. CXCR2-Mediated Granulocytic Myeloid-Derived Suppressor Cells’ Functional Characterization and Their Role in Maternal Fetal Interface. DNA Cell Biol (2016) 35(7):358–65. doi: 10.1089/dna.2015.2962
61. Benedicto A, Romayor I, Arteta B. CXCR4 receptor blockage reduces the contribution of tumor and stromal cells to the metastatic growth in the liver. Oncol Rep (2018) 39(4):2022–30. doi: 10.3892/or.2018.6254
62. Keskinov AA, Shurin MR. Myeloid regulatory cells in tumor spreading and metastasis. Immunobiology (2015) 220(2):236–42. doi: 10.1016/j.imbio.2014.07.017
63. Gabrilovich DI, Bronte V, Chen SH, Colombo MP, Ochoa A, Ostrand-Rosenberg S, et al. The terminology issue for myeloid-derived suppressor cells. Cancer Res (2007) 67(1):425. doi: 10.1158/0008-5472.CAN-06-3037
64. Pillay J, Tak T, Kamp VM, Koenderman L. Immune suppression by neutrophils and granulocytic myeloid-derived suppressor cells: similarities and differences. Cell Mol Life Sci (2013) 70(20):3813–27. doi: 10.1007/s00018-013-1286-4
65. Raber PL, Thevenot P, Sierra R, Wyczechowska D, Halle D, Ramirez ME, et al. Subpopulations of myeloid-derived suppressor cells impair T cell responses through independent nitric oxide-related pathways. Int J Cancer (2014) 134(12):2853–64. doi: 10.1002/ijc.28622
66. Wu L, Saxena S, Awaji M, Singh RK. Tumor-Associated Neutrophils in Cancer: Going Pro. Cancers (Basel) (2019) 11(4):3. doi: 10.3390/cancers11040564
67. Shaul ME, Levy L, Sun J, Mishalian I, Singhal S, Kapoor V, et al. Tumor-associated neutrophils display a distinct N1 profile following TGFbeta modulation: A transcriptomics analysis of pro- vs. antitumor TANs. Oncoimmunology (2016) 5(11):e1232221. doi: 10.1080/2162402X.2016.1232221
68. Jablonska J, Leschner S, Westphal K, Lienenklaus S, Weiss S. Neutrophils responsive to endogenous IFN-beta regulate tumor angiogenesis and growth in a mouse tumor model. J Clin Invest (2010) 120(4):1151–64. doi: 10.1172/JCI37223
69. Hietbrink F, Koenderman L, Althuizen M, Pillay J, Kamp V, Leenen LP. Kinetics of the innate immune response after trauma: implications for the development of late onset sepsis. Shock (2013) 40(1):21–7. doi: 10.1097/SHK.0b013e318295a40a
70. Fridlender ZG, Albelda SM. Tumor-associated neutrophils: friend or foe? Carcinogenesis (2012) 33(5):949–55. doi: 10.1093/carcin/bgs123
71. Scapini P, Cassatella MA. Social networking of human neutrophils within the immune system. Blood (2014) 124(5):710–9. doi: 10.1182/blood-2014-03-453217
72. Cvejic A. Mechanisms of fate decision and lineage commitment during haematopoiesis. Immunol Cell Biol (2016) 94(3):230–5. doi: 10.1038/icb.2015.96
73. Evrard M, Kwok IWH, Chong SZ, Teng KWW, Becht E, Chen J, et al. Developmental Analysis of Bone Marrow Neutrophils Reveals Populations Specialized in Expansion, Trafficking, and Effector Functions. Immunity (2018) 48(2):364–79.e8. doi: 10.1016/j.immuni.2018.02.002
74. Bartels M, Govers AM, Fleskens V, Lourenco AR, Pals CE, Vervoort SJ, et al. Acetylation of C/EBPepsilon is a prerequisite for terminal neutrophil differentiation. Blood (2015) 125(11):1782–92. doi: 10.1182/blood-2013-12-543850
75. Cirilli M, Bereshchenko O, Ermakova O, Nerlov C. Insights into specificity, redundancy and new cellular functions of C/EBPa and C/EBPb transcription factors through interactome network analysis. Biochim Biophys Acta Gen Subj (2017) 1861(2):467–76. doi: 10.1016/j.bbagen.2016.10.002
76. Gade P, Kimball AS, DiNardo AC, Gangwal P, Ross DD, Boswell HS, et al. Death-associated Protein Kinase-1 Expression and Autophagy in Chronic Lymphocytic Leukemia Are Dependent on Activating Transcription Factor-6 and CCAAT/Enhancer-binding Protein-beta. J Biol Chem (2016) 291(42):22030–42. doi: 10.1074/jbc.M116.725796
77. Fiedler K, Brunner C. The role of transcription factors in the guidance of granulopoiesis. Am J Blood Res (2012) 2(1):57–65.
78. Hong CW. Current Understanding in Neutrophil Differentiation and Heterogeneity. Immune Netw (2017) 17(5):298–306. doi: 10.4110/in.2017.17.5.298
79. Bjerregaard MD, Jurlander J, Klausen P, Borregaard N, Cowland JB. The in vivo profile of transcription factors during neutrophil differentiation in human bone marrow. Blood (2003) 101(11):4322–32. doi: 10.1182/blood-2002-03-0835
80. Benarafa C, LeCuyer TE, Baumann M, Stolley JM, Cremona TP, Remold-O’Donnell E. SerpinB1 protects the mature neutrophil reserve in the bone marrow. J Leukoc Biol (2011) 90(1):21–9. doi: 10.1189/jlb.0810461
81. Hager M, Cowland JB, Borregaard N. Neutrophil granules in health and disease. J Intern Med (2010) 268(1):25–34. doi: 10.1111/j.1365-2796.2010.02237.x
82. Lawrence SM, Corriden R, Nizet V. The Ontogeny of a Neutrophil: Mechanisms of Granulopoiesis and Homeostasis. Microbiol Mol Biol Rev (2018) 82(1). doi: 10.1128/MMBR.00057-17
83. Grassi L, Pourfarzad F, Ullrich S, Merkel A, Were F, Carrillo-de-Santa-Pau E, et al. Dynamics of Transcription Regulation in Human Bone Marrow Myeloid Differentiation to Mature Blood Neutrophils. Cell Rep (2018) 24(10):2784–94. doi: 10.1016/j.celrep.2018.08.018
84. Veda P. Why are Neutrophils Polymorphonuclear? Eur J Inflammation (2011) 9:85–93. doi: 10.1177/1721727X1100900201
85. Olins AL, Olins DE. Cytoskeletal influences on nuclear shape in granulocytic HL-60 cells. BMC Cell Biol (2004) 5:30. doi: 10.1186/1471-2121-5-30
86. Elghetany MT. Surface antigen changes during normal neutrophilic development: a critical review. Blood Cells Mol Dis (2002) 28(2):260–74. doi: 10.1006/bcmd.2002.0513
87. Marini O, Costa S, Bevilacqua D, Calzetti F, Tamassia N, Spina C, et al. Mature CD10(+) and immature CD10(-) neutrophils present in G-CSF-treated donors display opposite effects on T cells. Blood (2017) 129(10):1343–56. doi: 10.1182/blood-2016-04-713206
88. Grieshaber-Bouyer R, Nigrovic PA. Neutrophil Heterogeneity as Therapeutic Opportunity in Immune-Mediated Disease. Front Immunol (2019) 10:346. doi: 10.3389/fimmu.2019.00346
89. Zhao L, Xu S, Fjaertoft G, Pauksen K, Hakansson L, Venge P. An enzyme-linked immunosorbent assay for human carcinoembryonic antigen-related cell adhesion molecule 8, a biological marker of granulocyte activities in vivo. J Immunol Methods (2004) 293(1-2):207–14. doi: 10.1016/j.jim.2004.08.009
90. Albrechtsen M, Kerr MA. Characterization of human neutrophil glycoproteins expressing the CD15 differentiation antigen (3-fucosyl-N-acetyllactosamine). Br J Haematol (1989) 72(3):312–20. doi: 10.1111/j.1365-2141.1989.tb07710.x
91. Terstappen LW, Hollander Z, Meiners H, Loken MR. Quantitative comparison of myeloid antigens on five lineages of mature peripheral blood cells. J Leukoc Biol (1990) 48(2):138–48. doi: 10.1002/jlb.48.2.138
92. Hoyer JD, Grogg KL, Hanson CA, Gamez JD, Dogan A. CD33 detection by immunohistochemistry in paraffin-embedded tissues: a new antibody shows excellent specificity and sensitivity for cells of myelomonocytic lineage. Am J Clin Pathol (2008) 129(2):316–23. doi: 10.1309/E36008Y2H08Q1AYY
93. Laszlo GS, Estey EH, Walter RB. The past and future of CD33 as therapeutic target in acute myeloid leukemia. Blood Rev (2014) 28(4):143–53. doi: 10.1016/j.blre.2014.04.001
94. Liu M, Weng X, Gong S, Chen H, Ding J, Guo M, et al. Flow cytometric analysis of CD64 expression pattern and density in the diagnosis of acute promyelocytic leukemia: a multi-center study in Shanghai, China. Oncotarget (2017) 8(46):80625–37. doi: 10.18632/oncotarget.20814
95. Oostendorp RA, Dormer P. VLA-4-mediated interactions between normal human hematopoietic progenitors and stromal cells. Leuk Lymphoma (1997) 24(5-6):423–35. doi: 10.3109/10428199709055581
96. Fu S, Liesveld J. Mobilization of hematopoietic stem cells. Blood Rev (2000) 14(4):205–18. doi: 10.1054/blre.2000.0138
97. Elghetany MT, Ge Y, Patel J, Martinez J, Uhrova H. Flow cytometric study of neutrophilic granulopoiesis in normal bone marrow using an expanded panel of antibodies: correlation with morphologic assessments. J Clin Lab Anal (2004) 18(1):36–41. doi: 10.1002/jcla.20001
98. Elghetany MT, Patel J, Martinez J, Schwab H. CD87 as a marker for terminal granulocytic maturation: assessment of its expression during granulopoiesis. Cytometry B Clin Cytom (2003) 51(1):9–13. doi: 10.1002/cyto.b.10008
99. Terstappen LW, Safford M, Loken MR. Flow cytometric analysis of human bone marrow. III. Neutrophil maturation. Leukemia (1990) 4(9):657–63.
100. Weirich E, Rabin RL, Maldonado Y, Benitz W, Modler S, Herzenberg LA, et al. Neutrophil CD11b expression as a diagnostic marker for early-onset neonatal infection. J Pediatr (1998) 132(3 Pt 1):445–51. doi: 10.1016/S0022-3476(98)70018-6
101. Cowland JB, Borregaard N. Granulopoiesis and granules of human neutrophils. Immunol Rev (2016) 273(1):11–28. doi: 10.1111/imr.12440
102. Costantini C, Micheletti A, Calzetti F, Perbellini O, Pizzolo G, Cassatella MA. Neutrophil activation and survival are modulated by interaction with NK cells. Int Immunol (2010) 22(10):827–38. doi: 10.1093/intimm/dxq434
103. Sadhu C, Ting HJ, Lipsky B, Hensley K, Garcia-Martinez LF, Simon SI, et al. CD11c/CD18: novel ligands and a role in delayed-type hypersensitivity. J Leukoc Biol (2007) 81(6):1395–403. doi: 10.1189/jlb.1106680
104. Healy L, May G, Gale K, Grosveld F, Greaves M, Enver T. The stem cell antigen CD34 functions as a regulator of hemopoietic cell adhesion. Proc Natl Acad Sci USA (1995) 92(26):12240–4. doi: 10.1073/pnas.92.26.12240
105. Sandilands GP, McCrae J, Hill K, Perry M, Baxter D. Major histocompatibility complex class II (DR) antigen and costimulatory molecules on in vitro and in vivo activated human polymorphonuclear neutrophils. Immunology (2006) 119(4):562–71. doi: 10.1111/j.1365-2567.2006.02471.x
106. Gorczyca W, Sun ZY, Cronin W, Li X, Mau S, Tugulea S. Immunophenotypic pattern of myeloid populations by flow cytometry analysis. Methods Cell Biol (2011) 103:221–66. doi: 10.1016/B978-0-12-385493-3.00010-3
107. Vono M, Lin A, Norrby-Teglund A, Koup RA, Liang F, Lore K. Neutrophils acquire the capacity for antigen presentation to memory CD4(+) T cells in vitro and ex vivo. Blood (2017) 129(14):1991–2001. doi: 10.1182/blood-2016-10-744441
108. Java A, Liszewski MK, Hourcade DE, Zhang F, Atkinson JP. Role of complement receptor 1 (CR1; CD35) on epithelial cells: A model for understanding complement-mediated damage in the kidney. Mol Immunol (2015) 67(2 Pt B):584–95. doi: 10.1016/j.molimm.2015.07.016
109. Laszik Z, Jansen PJ, Cummings RD, Tedder TF, McEver RP, Moore KL. P-selectin glycoprotein ligand-1 is broadly expressed in cells of myeloid, lymphoid, and dendritic lineage and in some nonhematopoietic cells. Blood (1996) 88(8):3010–21. doi: 10.1182/blood.V88.8.3010.bloodjournal8883010
110. Tak T, Wijten P, Heeres M, Pickkers P, Scholten A, Heck AJR, et al. Human CD62L(dim) neutrophils identified as a separate subset by proteome profiling and in vivo pulse-chase labeling. Blood (2017) 129(26):3476–85. doi: 10.1182/blood-2016-07-727669
111. Eash KJ, Greenbaum AM, Gopalan PK, Link DC. CXCR2 and CXCR4 antagonistically regulate neutrophil trafficking from murine bone marrow. J Clin Invest (2010) 120(7):2423–31. doi: 10.1172/JCI41649
112. Nguyen-Jackson H, Panopoulos AD, Zhang H, Li HS, Watowich SS. STAT3 controls the neutrophil migratory response to CXCR2 ligands by direct activation of G-CSF-induced CXCR2 expression and via modulation of CXCR2 signal transduction. Blood (2010) 115(16):3354–63. doi: 10.1182/blood-2009-08-240317
113. Martin C, Burdon PC, Bridger G, Gutierrez-Ramos JC, Williams TJ, Rankin SM. Chemokines acting via CXCR2 and CXCR4 control the release of neutrophils from the bone marrow and their return following senescence. Immunity (2003) 19(4):583–93. doi: 10.1016/S1074-7613(03)00263-2
114. Stroncek DF, Shankar R, Litz C, Clement L. The expression of the NB1 antigen on myeloid precursors and neutrophils from children and umbilical cords. Transfus Med (1998) 8(2):119–23. doi: 10.1046/j.1365-3148.1998.00136.x
115. Davoine F, Lavigne S, Chakir J, Ferland C, Boulay ME, Laviolette M. Expression of FcgammaRIII (CD16) on human peripheral blood eosinophils increases in allergic conditions. J Allergy Clin Immunol (2002) 109(3):463–9. doi: 10.1067/mai.2002.121952
116. Haziot A, Tsuberi BZ, Goyert SM. Neutrophil CD14: biochemical properties and role in the secretion of tumor necrosis factor-alpha in response to lipopolysaccharide. J Immunol (1993) 150(12):5556–65.
117. Guerin E, Orabona M, Raquil MA, Giraudeau B, Bellier R, Gibot S, et al. Circulating immature granulocytes with T-cell killing functions predict sepsis deterioration*. Crit Care Med (2014) 42(9):2007–18. doi: 10.1097/CCM.0000000000000344
118. Demaret J, Venet F, Friggeri A, Cazalis MA, Plassais J, Jallades L, et al. Marked alterations of neutrophil functions during sepsis-induced immunosuppression. J Leukoc Biol (2015) 98(6):1081–90. doi: 10.1189/jlb.4A0415-168RR
119. Hernandez-Campo PM, Almeida J, Matarraz S, de Santiago M, Sanchez ML, Orfao A. Quantitative analysis of the expression of glycosylphosphatidylinositol-anchored proteins during the maturation of different hematopoietic cell compartments of normal bone marrow. Cytometry B Clin Cytom (2007) 72(1):34–42. doi: 10.1002/cyto.b.20143
120. Mahlknecht U, Schonbein C. Histone deacetylase inhibitor treatment downregulates VLA-4 adhesion in hematopoietic stem cells and acute myeloid leukemia blast cells. Haematologica (2008) 93(3):443–6. doi: 10.3324/haematol.11796
121. Sigua JA, Buelow B, Cheung DS, Buell E, Hunter D, Klancnik M, et al. CD49d-expressing neutrophils differentiate atopic from nonatopic individuals. J Allergy Clin Immunol (2014) 133(3):901–4. doi: 10.1016/j.jaci.2013.09.035
122. Zhang D, Chen G, Manwani D, Mortha A, Xu C, Faith JJ, et al. Neutrophil ageing is regulated by the microbiome. Nature (2015) 525(7570):528–32. doi: 10.1038/nature15367
123. Pillay J, Kamp VM, van Hoffen E, Visser T, Tak T, Lammers JW, et al. A subset of neutrophils in human systemic inflammation inhibits T cell responses through Mac-1. J Clin Invest (2012) 122(1):327–36. doi: 10.1172/JCI57990
124. Hu N, Westra J, Huitema MG, Bijl M, Brouwer E, Stegeman CA, et al. Coexpression of CD177 and membrane proteinase 3 on neutrophils in antineutrophil cytoplasmic autoantibody-associated systemic vasculitis: anti-proteinase 3-mediated neutrophil activation is independent of the role of CD177-expressing neutrophils. Arthritis Rheum (2009) 60(5):1548–57. doi: 10.1002/art.24442
125. Stroncek DF, Caruccio L, Bettinotti M. CD177: A member of the Ly-6 gene superfamily involved with neutrophil proliferation and polycythemia vera. J Transl Med (2004) 2(1):8. doi: 10.1186/1479-5876-2-8
126. De Filippo K, Rankin SM. CXCR4, the master regulator of neutrophil trafficking in homeostasis and disease. Eur J Clin Invest (2018) 48 Suppl 2:e12949. doi: 10.1111/eci.12949
127. Nauseef WM, Borregaard N. Neutrophils at work. Nat Immunol (2014) 15(7):602–11. doi: 10.1038/ni.2921
128. Riyapa D, Buddhisa S, Korbsrisate S, Cuccui J, Wren BW, Stevens MP, et al. Neutrophil extracellular traps exhibit antibacterial activity against burkholderia pseudomallei and are influenced by bacterial and host factors. Infect Immun (2012) 80(11):3921–9. doi: 10.1128/IAI.00806-12
129. Borregaard N, Cowland JB. Granules of the human neutrophilic polymorphonuclear leukocyte. Blood (1997) 89(10):3503–21. doi: 10.1182/blood.V89.10.3503
130. Li RHL, Tablin F. A Comparative Review of Neutrophil Extracellular Traps in Sepsis. Front Vet Sci (2018) 5:291. doi: 10.3389/fvets.2018.00291
131. Papayannopoulos V, Metzler KD, Hakkim A, Zychlinsky A. Neutrophil elastase and myeloperoxidase regulate the formation of neutrophil extracellular traps. J Cell Biol (2010) 191(3):677–91. doi: 10.1083/jcb.201006052
132. Borregaard N, Theilgaard-Monch K, Sorensen OE, Cowland JB. Regulation of human neutrophil granule protein expression. Curr Opin Hematol (2001) 8(1):23–7. doi: 10.1097/00062752-200101000-00005
133. Korkmaz B, Moreau T, Gauthier F. Neutrophil elastase, proteinase 3 and cathepsin G: physicochemical properties, activity and physiopathological functions. Biochimie (2008) 90(2):227–42. doi: 10.1016/j.biochi.2007.10.009
134. Korkmaz B, Horwitz MS, Jenne DE, Gauthier F. Neutrophil elastase, proteinase 3, and cathepsin G as therapeutic targets in human diseases. Pharmacol Rev (2010) 62(4):726–59. doi: 10.1124/pr.110.002733
135. Ragland SA, Criss AK. From bacterial killing to immune modulation: Recent insights into the functions of lysozyme. PLoS Pathog (2017) 13(9):e1006512. doi: 10.1371/journal.ppat.1006512
136. Glenthoj A, Nickles K, Cowland J, Borregaard N. Processing of Neutrophil alpha-Defensins Does Not Rely on Serine Proteases In Vivo. PLoS One (2015) 10(5):e0125483. doi: 10.1371/journal.pone.0125483
137. Jesaitis AJ, Buescher ES, Harrison D, Quinn MT, Parkos CA, Livesey S, et al. Ultrastructural localization of cytochrome b in the membranes of resting and phagocytosing human granulocytes. J Clin Invest (1990) 85(3):821–35. doi: 10.1172/JCI114509
138. Liu W, Yan M, Liu Y, McLeish KR, Coleman WG Jr, Rodgers GP. Olfactomedin 4 inhibits cathepsin C-mediated protease activities, thereby modulating neutrophil killing of Staphylococcus aureus and Escherichia coli in mice. J Immunol (2012) 189(5):2460–7. doi: 10.4049/jimmunol.1103179
139. Bostrom EA, Tarkowski A, Bokarewa M. Resistin is stored in neutrophil granules being released upon challenge with inflammatory stimuli. Biochim Biophys Acta (2009) 1793(12):1894–900. doi: 10.1016/j.bbamcr.2009.09.008
140. Stroncek DF, Skubitz KM, McCullough JJ. Biochemical characterization of the neutrophil-specific antigen NB1. Blood (1990) 75(3):744–55. doi: 10.1182/blood.V75.3.744.bloodjournal753744
141. Bauer S, Abdgawad M, Gunnarsson L, Segelmark M, Tapper H, Hellmark T. Proteinase 3 and CD177 are expressed on the plasma membrane of the same subset of neutrophils. J Leukoc Biol (2007) 81(2):458–64. doi: 10.1189/jlb.0806514
142. Kuckleburg CJ, Tilkens SB, Santoso S, Newman PJ. Proteinase 3 contributes to transendothelial migration of NB1-positive neutrophils. J Immunol (2012) 188(5):2419–26. doi: 10.4049/jimmunol.1102540
143. Klausen P, Niemann CU, Cowland JB, Krabbe K, Borregaard N. On mouse and man: neutrophil gelatinase associated lipocalin is not involved in apoptosis or acute response. Eur J Haematol (2005) 75(4):332–40. doi: 10.1111/j.1600-0609.2005.00511.x
144. Rorvig S, Ostergaard O, Heegaard NH, Borregaard N. Proteome profiling of human neutrophil granule subsets, secretory vesicles, and cell membrane: correlation with transcriptome profiling of neutrophil precursors. J Leukoc Biol (2013) 94(4):711–21. doi: 10.1189/jlb.1212619
145. Kolaczkowska E, Kubes P. Neutrophil recruitment and function in health and inflammation. Nat Rev Immunol (2013) 13(3):159–75. doi: 10.1038/nri3399
146. Gardiner EE, De Luca M, McNally T, Michelson AD, Andrews RK, Berndt MC. Regulation of P-selectin binding to the neutrophil P-selectin counter-receptor P-selectin glycoprotein ligand-1 by neutrophil elastase and cathepsin G. Blood (2001) 98(5):1440–7. doi: 10.1182/blood.V98.5.1440
147. Fuchs TA, Abed U, Goosmann C, Hurwitz R, Schulze I, Wahn V, et al. Novel cell death program leads to neutrophil extracellular traps. J Cell Biol (2007) 176(2):231–41. doi: 10.1083/jcb.200606027
148. Sollberger G, Choidas A, Burn GL, Habenberger P, Di Lucrezia R, Kordes S, et al. Gasdermin D plays a vital role in the generation of neutrophil extracellular traps. Sci Immunol (2018) 3(26):7–8. doi: 10.1126/sciimmunol.aar6689
149. Warnatsch A, Ioannou M, Wang Q, Papayannopoulos V. Inflammation. Neutrophil extracellular traps license macrophages for cytokine production in atherosclerosis. Science (2015) 349(6245):316–20. doi: 10.1126/science.aaa8064
150. Knackstedt SL, Georgiadou A, Apel F, Abu-Abed U, Moxon CA, Cunnington AJ, et al. Neutrophil extracellular traps drive inflammatory pathogenesis in malaria. Sci Immunol (2019) 4(40):8. doi: 10.1126/sciimmunol.aaw0336
151. Nguyen GT, Green ER, Mecsas J. Neutrophils to the ROScue: Mechanisms of NADPH Oxidase Activation and Bacterial Resistance. Front Cell Infect Microbiol (2017) 7:373. doi: 10.3389/fcimb.2017.00373
152. May RC, Machesky LM. Phagocytosis and the actin cytoskeleton. J Cell Sci (2001) 114(Pt 6):1061–77.
153. Behe P, Segal AW. The function of the NADPH oxidase of phagocytes, and its relationship to other NOXs. Biochem Soc Trans (2007) 35(Pt 5):1100–3. doi: 10.1042/BST0351100
154. Sheshachalam A, Srivastava N, Mitchell T, Lacy P, Eitzen G. Granule protein processing and regulated secretion in neutrophils. Front Immunol (2014) 5:448. doi: 10.3389/fimmu.2014.00448
155. Rorvig S, Honore C, Larsson LI, Ohlsson S, Pedersen CC, Jacobsen LC, et al. Ficolin-1 is present in a highly mobilizable subset of human neutrophil granules and associates with the cell surface after stimulation with fMLP. J Leukoc Biol (2009) 86(6):1439–49. doi: 10.1189/jlb.1008606
Keywords: neutrophils, granulopoiesis, neutrophil granules, low density neutrophils, nomenclature
Citation: McKenna E, Mhaonaigh AU, Wubben R, Dwivedi A, Hurley T, Kelly LA, Stevenson NJ, Little MA and Molloy EJ (2021) Neutrophils: Need for Standardized Nomenclature. Front. Immunol. 12:602963. doi: 10.3389/fimmu.2021.602963
Received: 04 September 2020; Accepted: 17 March 2021;
Published: 15 April 2021.
Edited by:
Sonja Vermeren, University of Edinburgh, United KingdomReviewed by:
Carsten Krieg, Medical University of South Carolina, United StatesZvi Granot, Hebrew College, United States
Copyright © 2021 McKenna, Mhaonaigh, Wubben, Dwivedi, Hurley, Kelly, Stevenson, Little and Molloy. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Eleanor J. Molloy, RWxlYW5vci5tb2xsb3lAdGNkLmll
 Ellen McKenna
Ellen McKenna Aisling Ui Mhaonaigh
Aisling Ui Mhaonaigh Richard Wubben
Richard Wubben Amrita Dwivedi
Amrita Dwivedi Tim Hurley
Tim Hurley Lynne A. Kelly
Lynne A. Kelly Nigel J. Stevenson
Nigel J. Stevenson Mark A. Little
Mark A. Little Eleanor J. Molloy
Eleanor J. Molloy