- 1Jiangsu Key Laboratory of Zoonosis, Yangzhou University, Yangzhou, China
- 2Jiangsu Co-innovation Center for Prevention and Control of Important Animal Infectious Diseases and Zoonoses, Yangzhou, China
- 3Joint International Research Laboratory of Agriculture and Agri-product Safety of the Ministry of Education, Yangzhou, China
- 4Key Laboratory of Prevention and Control of Biological Hazard Factors (Animal Origin) for Agrifood Safety and Quality, Ministry of Agriculture of China, Yangzhou University, Yangzhou, China
Salmonella enterica serovars Enteritidis, Pullorum/Gallinarum, and Dublin are infectious pathogens causing serious problems for pig, chicken, and cattle production, respectively. Traditional serotyping for Salmonella is costly and labor-intensive. Here, we established a rapid multiplex PCR method to simultaneously identify three prevalent Salmonella serovars Enteritidis, Pullorum/Gallinarum, and Dublin individually for the first time. The multiplex PCR-based assay focuses on three genes tcpS, lygD, and flhB. Gene tcpS exists only in the three Salmonella serovars, and lygD exists only in S. Enteritidis, while a truncated region of flhB gene is only found in S. Pullorum/Gallinarum. The sensitivity and specificity of the multiplex PCR assay using three pairs of specific primers for these genes were evaluated. The results showed that this multiplex PCR method could accurately identify Salmonella Enteritidis, Pullorum/Gallinarum, and Dublin from eight non-Salmonella species and 27 Salmonella serovars. The least concentration of genomic DNA that could be detected was 58.5 pg/μL and the least number of cells was 100 CFU. Subsequently, this developed method was used to analyze clinical Salmonella isolates from one pig farm, one chicken farm, and one cattle farm. The results showed that blinded PCR testing of Salmonella isolates from the three farms were in concordance with the traditional serotyping tests, indicating the newly developed multiplex PCR system could be used as a novel tool to accurately distinguish the three specific Salmonella serovars individually, which is useful, especially in high-throughput screening.
Introduction
Salmonella is a prominent food-borne pathogen, capable of causing serious illness in humans, including gastroenteritis, typhoid fever, septicemia, and sometimes even death (Tatavarthy and Cannons, 2010). It is reported almost 75% of Salmonella infections in human cases are caused by contaminated food products, including pork, poultry, and beef (Hald et al., 2004).
Although more than 2,600 Salmonella serovars exist (Ranieri et al., 2013; Xiong et al., 2016), S. Enteritidis, S. Pullorum/Gallinarum, and S. Dublin are the main serovars causing animal diseases (Nielsen, 2013; Saeki et al., 2013; Zhu et al., 2015). S. Enteritidis could cause severe infection in humans (Rodrigue et al., 1990; Nesbitt et al., 2012), and was the main serovar in the contaminated food products and infected individuals in southern Brazil between 1999 and 2008 (Paião et al., 2013). S. Gallinarum only infects birds and has two biovars Gallinarum and Pullorum, causing fowl typhoid and “white diarrhea,” respectively (Soria et al., 2012; Xiong et al., 2016). Particularly, S. Gallinarum could transmit to the reproductive system and result in salmonellosis (Keller et al., 1997). S. Dublin causes widespread losses in cattle husbandry, mainly as a result of increased levels of abortion, mortality, and morbidity, and a reduced milk yield, and has attracted considerable attention from cattle industries worldwide (Carrique-Mas et al., 2010; Nielsen and Dohoo, 2013). Human infections are most caused by the consumption of milk or beef products (Nielsen, 2013). Thus, timely detection of the three prominent Salmonella serovars, S. Enteritidis, S. Pullorum/Gallinarum, and S. Dublin, is very essential and urgent.
Traditional serotyping for Salmonella is based on the identification of the somatic (O) and flagellar (H) antigens by using specific sera following the White-Kauffmann-Le Minor scheme (Majchrzak et al., 2014). Many useful data could be obtained by Salmonella serotyping. Thus, an accurate diagnostic method for Salmonella serovars is highly important for public health. Despite its wide use, traditional Salmonella serotyping has many disadvantages, which is expensive, time-consuming and labor-intensive (Ranieri et al., 2013). Recent studies showed that polymerase chain reaction (PCR) can be a useful method to detect pathogens for its high specificity and sensitivity (Abdissa et al., 2006; Moyo et al., 2007). PCR-based method for Salmonella serotyping is a rapid and economical tool (Karns et al., 2015). Gene lygD in Sdf locus has been found only in S. Enteritidis and could be used to distinguish this serovar specifically (Zhu et al., 2015). Previously, we have proved that flhB gene can be used to detect S. Pullorum/Gallinarum because a unique region was deficient only in this serovar (Xiong et al., 2016).
In the present study, we established a rapid multiplex PCR method to distinguish the three prevalent Salmonella serovars Enteritidis, Pullorum/Gallinarum, and Dublin individually for the first time. The approach was based on designing three pairs of primers targeting tcpS, lygD, and flhB genes. The sensitivity and specificity of the multiplex PCR assay were determined, and the PCR assay was used to detect three sets of Salmonella isolates from one pig farm, one chicken farm, and one cattle farm. The newly developed multiplex PCR with three pairs of primers could be used as a novel tool to timely identify the three specific Salmonella serovars, and reinforce the traditional Salmonella serotyping method, particularly in high-throughput screening.
Materials and Methods
Bacterial Strains
Strains of Salmonella and non-Salmonella organisms, including S. Enteritidis, S. Gallinarum, S. Pullorum, S. Dublin, S. Meleagridis, S. Uganda, S. Anatis, S. London, S. Rissen, S. Typhimurium, S. Derby, S. Choleraesuis, S. Sinstorf, S. Indiana, S. Newlands, S. Dumfries, S. Muenster, S. Yoruba, S. Kentucky, S. Agona, S. Senftenberg, S. Thompson, S. Blockley, S. Inchpark, S. Farsta, S. Dabou, S. Virchow, Mycobacterium tuberculosis, Brucella abortus, Listeria monocytogenes, Campylobacter jejuni, and Escherichia coli, were commercially available or previously isolated in our routine monitoring (Table 1).
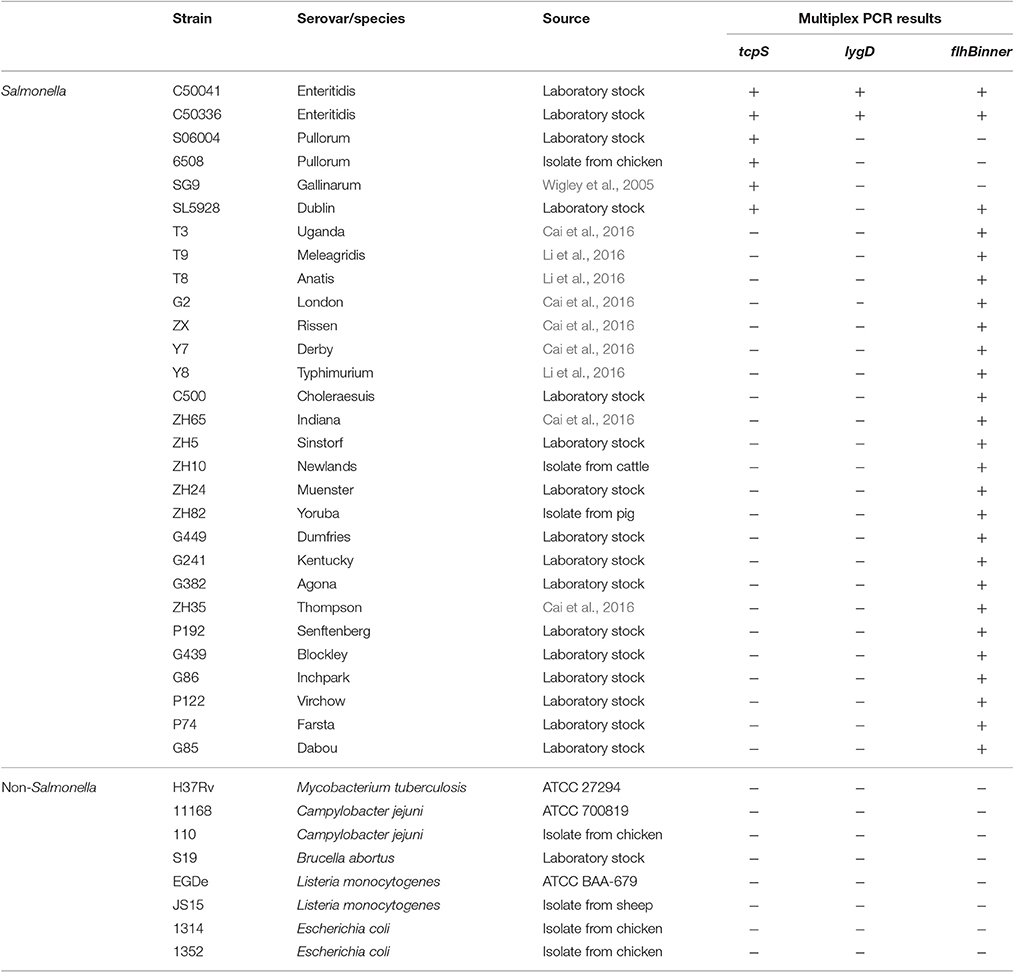
Table 1. Salmonella and non-Salmonella strains used to evaluate the specificity and sensitivity of the developed multiplex PCR method.
Bacterial Growth and Genomic DNA Isolation
The bacterial culture and DNA isolation were performed as previously described (Xiong et al., 2016). Briefly, all strains used in the study were inoculated in Brain Heart Infusion broth (Becton, Dickinson and Company, Sparks, MD, USA) or Luria-Bertani broth (Oxoid, Basingstoke, Hampshire, England) at 37°C overnight with an agitation speed of 180 rpm. Bacterial DNA was harvested with a TIANamp Bacterial DNA kit (TianGen, Beijing, China). The purity and concentration of the extracted DNA were determined using a NanoDrop ND-1000 (Thermo Scientific, Wilmington, DE, USA), and DNA samples were stored at −20°C until use.
In silico Analysis
To establish a sequence- and PCR-based Salmonella serotyping method for discrimination of S. Enteritidis, S. Pullorum/Gallinarum, and S. Dublin individually, the basic local alignment search tool (BLAST) algorithm (NCBI, non-redundant nucleotide collection) was applied. The tcpS, lygD, and flhB nucleotide sequences were used against the nucleotide collection (nr/nt) database, respectively. The aligned sequence number was set to the maximal value 20,000 with other parameters set to the default values. The three pairs of primers for the targets were designed using Primer Premier 5 (Premier, Palo Alto, CA, USA).
PCR Procedure
PCRs were conducted in a 25 μL reaction volume, consisting of 100 ng of genomic DNA template, 1 × polymerase buffer, 1 U of Taq polymerase (Takara Biotechnology Co., Dalian, China), 200 μM of deoxynucleoside triphosphate, and 80 nM of tcpS/lygD/flhBinner primers. PCR were conducted with a T100 Thermal Cycler (Bio-Rad, Hercules, California, USA) as follows: 94°C for 5 min, 30 cycles of 94°C for 45 s, 55°C for 45 s, and 72°C for 1 min, followed by 72°C for 10 min. The amplified PCR products were analyzed on the 1% agarose gel in 1 × TAE buffer.
Specificity of the Multiplex PCR
The specificity of the three pairs of primers in the multiplex PCR system was evaluated by detecting genomic DNA from 29 different Salmonella strains, which included 27 Salmonella serovars and eight non-Salmonella species (Table 1).
Sensitivity of the Multiplex PCR
The genomic DNA of S. Enteritidis strain C50041, S. Pullorum strain S06004 and S. Dublin strain SL5928 were 10-fold continuously diluted from 58.5 ng/μL to 585 fg/μL, respectively. Each dilution (2 μL) was used in the multiplex PCR assay. This assay was to determine the minimum limit of DNA that can be detected by the multiplex PCR method.
S. Enteritidis strain C50041, S. Pullorum strain S06004, and S. Dublin strain SL5928 were cultured overnight and the bacterial concentration was evaluated by the plate count assay. The bacterial culture was washed with phosphate buffered saline (PBS) twice, 10-fold serially diluted to the concentrations 2 × 107 to 2 × 102 CFU/mL, and boiled for 10 min to collect the genomic DNA. In the multiplex PCR method, each dilution (5 μL) was used as templets to determine the minimum cells of Salmonella that can be detected.
Isolation of Salmonella Strains from Different Farms
Additional clinical Salmonella strains with unknown serovars were obtained from three farms, one pig farm, one chicken farm, and one cattle farm in Yangzhou, China. The Salmonella isolates were collected from feces, floors and lairage, and identified as previously described methods (Cai et al., 2016; Li et al., 2016; Xiong et al., 2016). In brief, each sample was suspended in 50 mL buffered peptone water (Difco, BD, Sparks, MD, USA) and followed by incubation at 37°C for 16–18 h. This broth culture (0.1 mL) was subcultured in 10 mL of Rappaport–Vassiliadis enrichment broth (Difco, BD) at 42°C for 24 h. After incubation, the RV broth was streaked onto xylose lysine tergitol 4 (Difco, BD), and cultured at 37°C for 24–48 h. The presumptive Salmonella colonies were picked from all plates and followed by biochemically confirmation using an API-20E test kit (bioMérieux, Marcy l'Etoile, France).
Application of the Multiplex PCR Method on Clinical Samples
The multiplex PCR method was applied to detect the genomic DNA of Salmonella isolates from one pig farm (24 Salmonella isolates), one chicken farm (24 Salmonella isolates), and one cattle farm (11 Salmonella isolates). The obtained results of the developed PCR method were compared with traditional Salmonella serotyping approach.
Traditional Serotyping of Salmonella Isolates from Different Farms
The traditional serotyping for all isolated Salmonella strains from the pig, chicken and cattle farms were conducted by slide agglutination assay using the specific antisera (Tianrun Bio-Pharmaceutical, Ningbo, China) following the White-Kauffmann-LeMinor instructions (Grimont and Weill, 2007).
Results
In silico Analysis and Primer Design
In silico analysis showed that tcpS exists only in S. Enteritidis, S. Pullorum/Gallinarum, and S. Dublin, and has 98–100% sequence similarity across the three Salmonella serovars in the database. Although tcpS in one E. coli strain showed 89% DNA sequence identity to the Salmonella tcpS, it does not contain the C-terminal region of tcpS or a match to the tcpS-R primer site (data not shown). lygD gene exists only in S. Enteritidis, and shares 98–100% sequence similarity among this serovar in the database (data not shown). Our previous study showed that flhB gene of S. Pullorum/Gallinarum lacks a unique region flhBinner compared with other serovars, and could be used to identify S. Pullorum/Gallinarum (Xiong et al., 2016). Therefore, three pairs of oligonucleotide primers distinguishing three specific Salmonella serovars were designed based on the three targets tcpS, lygD and flhBinner (Table 2).
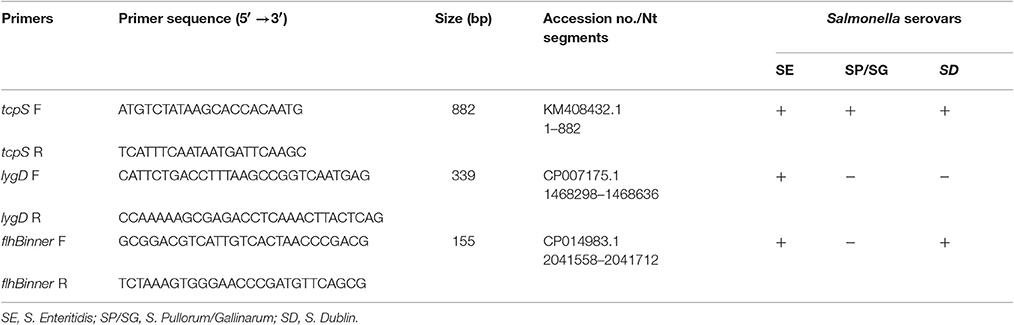
Table 2. Multiplex PCR primers used for identification of Salmonella Enteritidis, S. Pullorum/Gallinarum, and S. Dublin.
Specificity of the Multiplex PCR Assay
The specificity of the multiplex PCR method was determined by detecting 29 Salmonella strains and eight non-Salmonella species. The results showed that S. Enteritidis generated three specific bands for tcpS, lygD and flhBinner, and S. Dublin generated two specific bands for tcpS and flhBinner, while S. Pullorum/Gallinarum generated only one specific band for tcpS. In contrast, only one band of flhBinner was generated in the other 23 Salmonella serovars, and no amplification product was observed in all non-Salmonella pathogens (Figure 1).
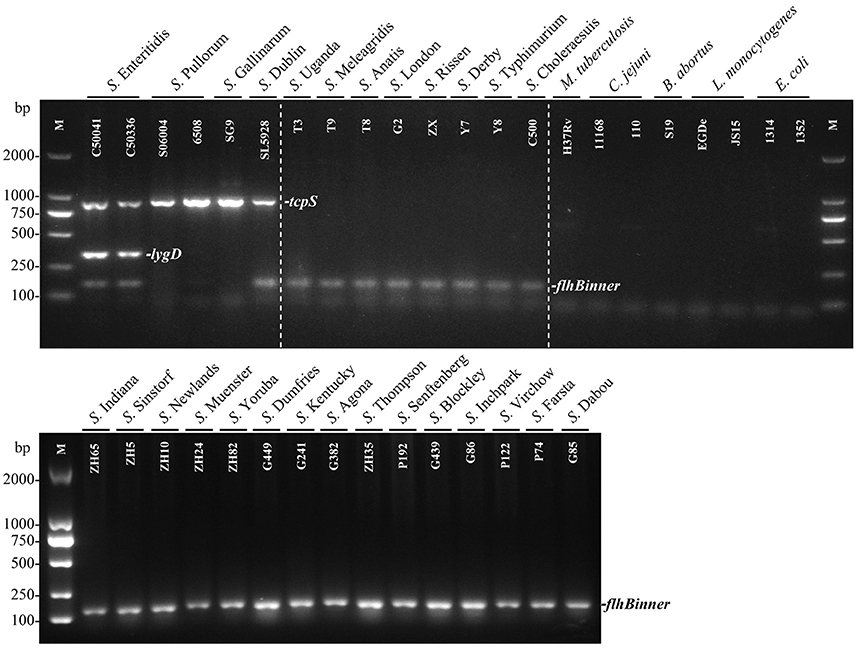
Figure 1. Specificity of the multiplex PCR method for the identification of Salmonella serovars Enteritidis, Pullorum/Gallinarum, and Dublin. The multiplex PCR assays, using genomic DNA from various Salmonella and non-Salmonella strains, were conducted using the designed primers targeting tcpS (882 bp), lygD (339 bp), and flhBinner (155 bp). The three specific PCR products could be amplified in S. Enteritidis. tcpS and flhBinner could be amplified in S. Dublin, while only tcpS gene could be amplified in S. Pullorum/Gallinarum. Detailed strain information is given in Table 1.
Sensitivity of the Multiplex PCR Assay
To evaluate the sensitivity of the multiplex PCR method, genomic DNA of S. Enteritidis, Pullorum, and Dublin were consecutively diluted and used as templates. The targeted fragments could be amplified at the lowest concentration of 58.5 pg/μL (Figure 2A), suggesting 58.5 pg/μL of genomic DNA was needed to detect S. Enteritidis, Pullorum, or Dublin using this method. Besides, the minimum cells of S. Enteritidis, Pullorum, and Dublin that could be detected using this multiplex PCR method were determined. After detecting various dilutions of Salmonella cells, we validated that the least cells that could be detected was 100 CFU (Figure 2B).
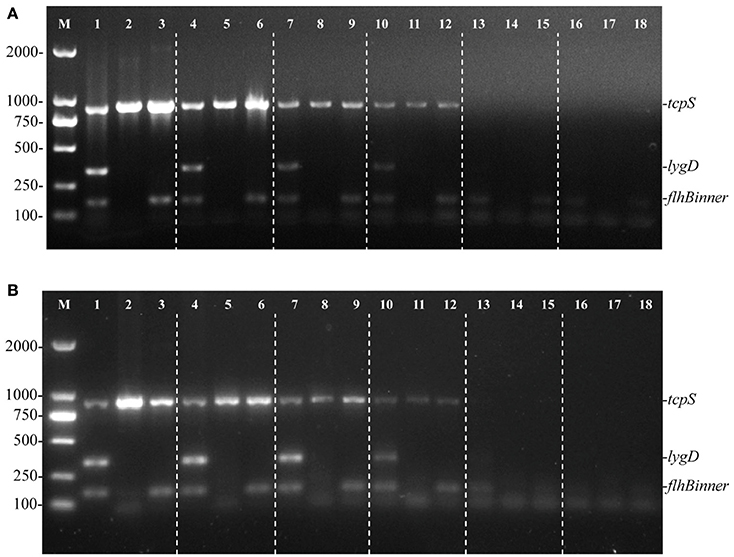
Figure 2. Sensitivity of the multiplex PCR method for the detection of genomic DNA and cells from S. Enteritidis (C50041), S. Pullorum (S06004), and S. Dublin (SL5928). The multiplex PCR amplifies three specific bands of tcpS (882 bp), lygD (339 bp), and flhBinner (155 bp). Lane M: DL2000 DNA marker (Takara Biotechnology Co., Dalian, China). The multiplex PCR for the detection of genomic DNA (A) and Salmonella cells (B), lanes 1, 4, 7, 10, 13, 16 (S. Enteritidis), 2, 5, 8, 11, 14, 17 (S. Pullorum), and 3, 6, 9, 12, 15, 18 (S. Dublin): Genomic DNA used as templates at the following concentrations, respectively: 58.5 ng/μL, 5.85 ng/μL, 585 pg/μL, 58.5 pg/μL, 5.85 pg/μL, 585 fg/μL; the number of cells per PCR assay at the following concentrations, respectively: 105, 104, 103, 102, 101, and 100 CFU.
Application of the Multiplex PCR Method
To determine the effectiveness of the developed multiplex PCR method, samples from one pig farm (24 Salmonella isolates), one chicken farm (24 Salmonella isolates), and one cattle farm (11 Salmonella isolates) were examined. The PCR results showed that three isolates from the pig farm generated three specific bands of tcpS, lygD, and flhBinner, suggesting that the three isolates were S. Enteritidis. Five samples from the chicken farm generated three specific bands of tcpS, lygD, and flhBinner, and 11 samples generated only one specific band of tcpS, suggesting that the five isolates and the other 11 isolates were S. Enteritidis and S. Pullorum/Gallinarum, respectively. Among the isolates from the cattle farm, only one sample generated two specific bands of tcpS and flhBinner, suggesting that this isolate were S. Dublin (Table 3).
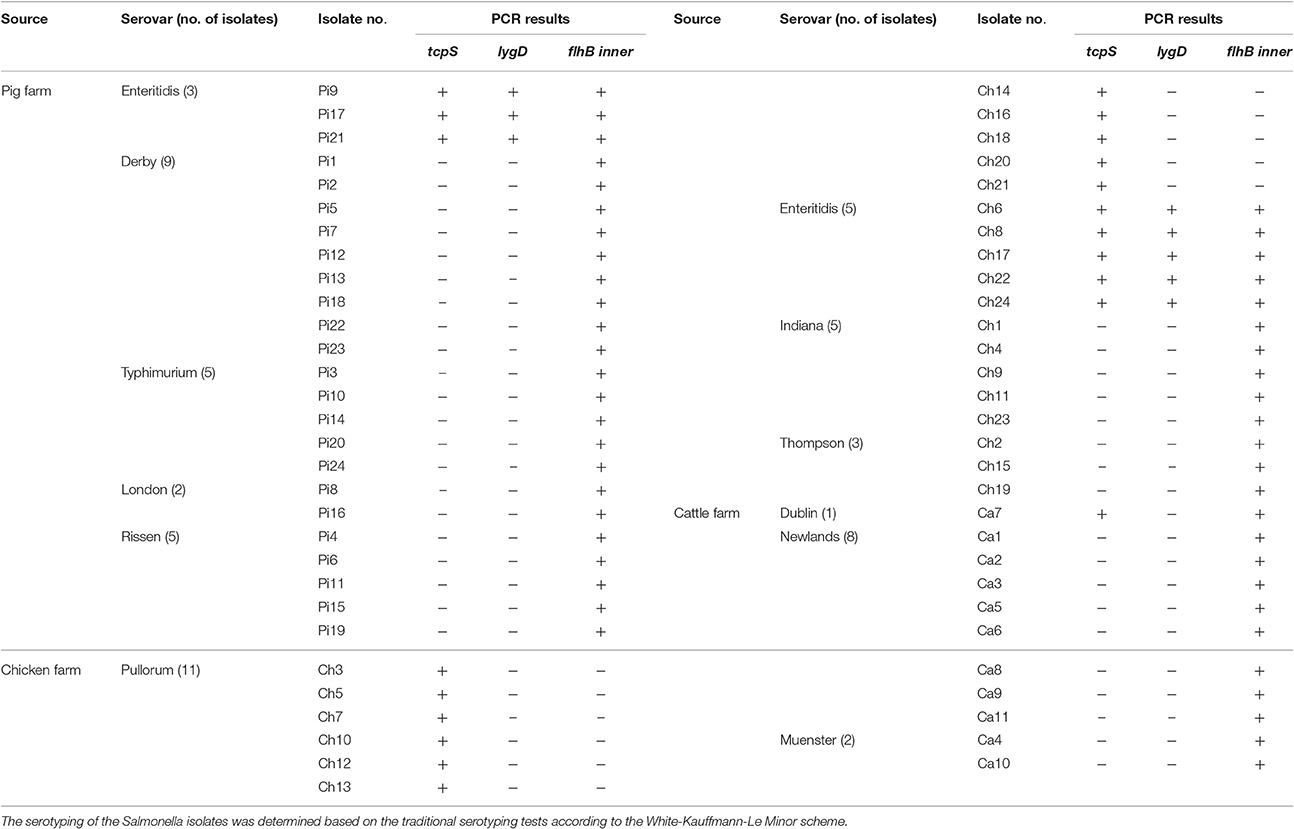
Table 3. Salmonella strains isolated from three different farms to examine the application of the developed multiplex PCR method.
Traditional Serotyping of Salmonella Isolates
The serotypes of Salmonella isolates from the three farms were identified using slide agglutination assays using specific O and H antisera. The isolates from the pig farm were obtained from lairage, floors, and feces, and included three strains of S. Enteritidis, nine strains of S. Derby, five strains of S. Typhimurium, two strains of S. London, and five strains of S. Rissen. The isolates from the chicken farm were obtained from fecal samples and floors, and included 11 strains of S. Pullorum, five strain of S. Enteritidis, five strains of S. Indiana, and three strains of S. Thompson. The isolates from the cattle farm were obtained from fecal samples, and included one strain of S. Dublin, eight strains of S. Newlands, and two strains of S. Muenster. The traditional serotyping results showed complete concordance with the developed multiplex PCR methods for all samples (Table 3).
Discussion
Salmonella remains the most frequently isolated bacteria among food-borne pathogens, and over 19,000 cases were reported in the USA in 2013 (Crim et al., 2014). Thus, a simple method to detect and monitor Salmonella serovars in farms is urgently required. Several approaches based on antigens and DNA analysis have been developed to detect Salmonella in foodstuffs, including enzyme-linked immunosorbent assay, PCR analysis, and next generation sequencing (Ricke et al., 2013; Park et al., 2014).
Traditional serotyping could provide subtyping information that allows worldwide comparison. This has promoted the characterization of many international Salmonella outbreaks (Werber et al., 2005; Elviss et al., 2009). Furthermore, comparison with historical data was also available based on serotyping because of its wide use for almost 70 years. Verifying the causative pathogens is usually the essential first step in many important epidemiological investigations. Traditional serotyping could be a tough task because it requires necessary expertise and numerous antisera to interpret the agglutination results (Hong et al., 2008). Traditional serotyping methods are also labor-intensive, complicated, expensive, and time-consuming. More importantly, morphological descriptions and biochemical tests may produce ambiguous results (de Freitas et al., 2010). Although whole genome sequencing is becoming more accessible and has been used as a genoytping method, it could be costly and time-consuming, and not practical for sequencing numerous isolates simultaneously. Therefore, rapid PCR-based detection systems for Salmonella have been developed in recent years (Persson et al., 2012).
Comparative genomic analysis is becoming common to validate novel serovar-specific genes because of the improved BLAST program and continuously supplemented genomic data (Zhai et al., 2014). This approach is more economical, convenient, and effective than traditional methods. For example, serovar-specific sequences (STM4495 and SEN1392) for identifying S. Enteritidis and S. Typhimurium were obtained by comparative genomics (Liu et al., 2012). At present, vagC, located in the Salmonella virulence plasmid, is considered a better target for PCR detection of S. Dublin (Persson et al., 2012). However, false-positive results still occur, such as misidentification of a S. Muenchen serovar as S. Dublin (Zhai et al., 2014). Previously, we have found Salmonella flhB gene could be used to identify S. Pullorum/Gallinarum from others because a unique region was deficient only in this serovar (Xiong et al., 2016). Here, we took advantage of three Salmonella genes, tcpS, lygD, and flhB, which were predicted by comparative genomic analysis, to design primers that can accurately distinguish Salmonella serovars Enteritidis, Pullorum/Gallinarum, and Dublin. This allowed the development of a reliable and rapid multiplex PCR method to screen these three serovars individually. To the best of our knowledge, it is the first one-step multiplex PCR method to detect these three prominent Salmonella serovars individually.
The multiplex PCR method produced positive results in S. Enteritidis, S. Pullorum/Gallinarum, and S. Dublin only, with negative results obtained in other Salmonella serovars and eight non-Salmonella bacteria (Figure 1). Besides, the PCR method is very rapid and takes about 3 h to complete. Thus, the PCR results agreed with the BLAST analysis, and the proposed application of the multiplex PCR method was verified by screening the three prominent Salmonella serovars in samples isolated from pig, chicken, and cattle farms. The results described in this study provide a proof of concept and demonstrate the feasibility of using this PCR method to screen S. Enteritidis, Pullorum/Gallinarum, and Dublin. Future studies will investigate different approaches to isolate DNA directly from infected animals and determine if it can be applied in the field.
This multiplex PCR method could be used for a rapid screening of the three specific Salmonella serovars and simplify the procedures of traditional serotyping. Besides, the combination of traditional serotyping methods and the developed PCR-based approach would promote the efficiency for the serovar identification of Salmonella strains.
Conclusion
In summary, an efficient multiplex PCR method targeting three prominent Salmonella serovars, S. Enteritidis, S. Pullorum/Gallinarum, and S. Dublin, was identified for the first time. The multiplex PCR method was based on three genes of tcpS, lygD, and flhB, and the specificity and sensitivity of the multiplex PCR method were determined. The multiplex PCR system was exploited to test extensive sets of Salmonella strains isolated from three farms, thus validating the effectiveness and specificity of the assay. The results suggest that the developed rapid and efficient multiplex PCR assay could be used as a novel and high-throughput diagnostic tool for simultaneous identification of the three specific Salmonella serovars.
Author Contributions
ZP and XJ designed the experiments; DX and LS performed the PCR assays; DX, JT, HZ, and ZZ isolated the samples from the chicken farm; SG participated in the data analysis and interpretation; DX, ZP, and XJ wrote the paper. All authors read and approved the final manuscript.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by the National Key Research and Development Program Special Project (2016YFD0501607), the Special Fund for Agroscientific Research in the Public Interest (201403054), the National Natural Science Foundation of China (nos. 31320103907 and 31230070), the Project for Agricultural Products Quality and Safety Supervision (Risk Assessment) (18162130109236), the Research and Development Program of Jiangsu (BE2015343), the “Six Talent Peaks Program” of Jiangsu Province (NY-028), the Yangzhou University Science and Technology Innovation Team, and the Priority Academic Program Development of Jiangsu Higher Education Institutions (PAPD).
References
Abdissa, A., Asrat, D., Kronvall, G., Shittu, B., Achiko, D., Zeidan, M., et al. (2006). High diversity of group A streptococcal emm types among healthy schoolchildren in Ethiopia. Clin. Infect. Dis. 42, 1362–1367. doi: 10.1086/503422
Cai, Y., Tao, J., Jiao, Y., Fei, X., Zhou, L., Wang, Y., et al. (2016). Phenotypic characteristics and genotypic correlation between Salmonella isolates from a slaughterhouse and retail markets in Yangzhou, China. Int. J. Food Microbiol. 222, 56–64. doi: 10.1016/j.ijfoodmicro.2016.01.020
Carrique-Mas, J. J., Willmington, J. A., Papadopoulou, C., Watson, E. N., and Davies, R. H. (2010). Salmonella infection in cattle in Great Britain, 2003 to 2008. Vet. Rec. 167, 560–565. doi: 10.1136/vr.c4943
Crim, S. M., Iwamoto, M., Huang, J. Y., Griffin, P. M., Gilliss, D., Cronquist, A. B., et al. (2014). Incidence and trends of infection with pathogens transmitted commonly through food–foodborne diseases active surveillance network, 10 U.S. sites, 2006-2013. MMWR. Morb. Mortal. Wkly. Rep. 63, 328–332. Available online at: https://www.cdc.gov/mmwr/preview/mmwrhtml/mm6315a3.htm
de Freitas, C. G., Santana, A. P., da Silva, P. H., Gonçalves, V. S., Barros Mde, A., Torres, F. A., et al. (2010). PCR multiplex for detection of Salmonella Enteritidis, Typhi and Typhimurium and occurrence in poultry meat. Int. J. Food Microbiol. 139, 15–22. doi: 10.1016/j.ijfoodmicro.2010.02.007
Elviss, N. C., Little, C. L., Hucklesby, L., Sagoo, S., Surman-Lee, S., de Pinna, E., et al. (2009). Microbiological study of fresh herbs from retail premises uncovers an international outbreak of salmonellosis. Int. J. Food Microbiol. 134, 83–88. doi: 10.1016/j.ijfoodmicro.2009.01.015
Grimont, P. A. D., and Weill, F. X. (2007). Antigenic Formulae of the Salmonella Serovars, 9th Edn. Paris: WHO Collaborating Center for Reference and Research on Salmonella; Institut Pasteur.
Hald, T., Vose, D., Wegener, H. C., and Koupeev, T. (2004). A Bayesian approach to quantify the contribution of animal-food sources to human salmonellosis. Risk. Anal. 24, 255–269. doi: 10.1111/j.0272-4332.2004.00427.x
Hong, Y., Liu, T., Lee, M. D., Hofacre, C. L., Maier, M., White, D. G., et al. (2008). Rapid screening of Salmonella enterica serovars Enteritidis, Hadar, Heidelberg and Typhimurium using a serologically-correlative allelotyping PCR targeting the O and H antigen alleles. BMC Microbiol. 8:178. doi: 10.1186/1471-2180-8-178
Karns, J. S., Haley, B. J., and Van Kessel, J. A. (2015). Improvements to a PCR-based serogrouping scheme for Salmonella enterica from dairy farm samples. J. Food Prot. 78, 1182–1185. doi: 10.4315/0362-028X.JFP-14-475
Keller, L. H., Schifferli, D. M., Benson, C. E., Aslam, S., and Eckroade, R. J. (1997). Invasion of chicken reproductive tissues and forming eggs is not unique to Salmonella enteritidis. Avian Dis. 41, 535–539. doi: 10.2307/1592142
Li, Y., Cai, Y., Tao, J., Kang, X., Jiao, Y., Guo, R., et al. (2016). Salmonella isolated from the slaughterhouses and correlation with pork contamination in free market. Food Control 59, 591–600. doi: 10.1016/j.foodcont.2015.06.040
Liu, B., Zhou, X., Zhang, L., Liu, W., Dan, X., Shi, C., et al. (2012). Development of a novel multiplex PCR assay for the identification of Salmonella enterica Typhimurium and Enteritidis. Food Control 27, 87–93. doi: 10.1016/j.foodcont.2012.01.062
Majchrzak, M., Krzyzanowska, A., Kubiak, A. B., Wojtasik, A., Wolkowicz, T., Szych, J., et al. (2014). TRS-based PCR as a potential tool for inter-serovar discrimination of Salmonella Enteritidis, S. Typhimurium, S. Infantis, S. Virchow, S. Hadar, S. Newport and S. Anatum. Mol. Biol. Rep. 41, 7121–7132. doi: 10.1007/s11033-014-3592-9
Moyo, S. J., Maselle, S. Y., Matee, M. I., Langeland, N., and Mylvaganam, H. (2007). Identification of diarrheagenic Escherichia coli isolated from infants and children in Dar es Salaam, Tanzania. BMC Infect. Dis. 7:92. doi: 10.1186/1471-2334-7-92
Nesbitt, A., Ravel, A., Murray, R., McCormick, R., Savelli, C., Finley, R., et al. (2012). Integrated surveillance and potential sources of Salmonella enteritidis in human cases in Canada from 2003 to 2009. Epidemiol. Infect. 140, 1757–1772. doi: 10.1017/S0950268811002548
Nielsen, L. R. (2013). Review of pathogenesis and diagnostic methods of immediate relevance for epidemiology and control of Salmonella Dublin in cattle. Vet. Microbiol. 162, 1–9. doi: 10.1016/j.vetmic.2012.08.003
Nielsen, L. R., and Dohoo, I. (2013). Time-to-event analysis of predictors for recovery from Salmonella Dublin infection in Danish dairy herds between 2002 and 2012. Prev. Vet. Med. 110, 370–378. doi: 10.1016/j.prevetmed.2013.02.014
Paião, F. G., Arisitides, L. G., Murate, L. S., Vilas-Bôas, G. T., Vilas-Boas, L. A., and Shimokomaki, M. (2013). Detection of Salmonella spp, Salmonella Enteritidis and Typhimurium in naturally infected broiler chickens by a multiplex PCR-based assay. Braz. J. Microbiol. 44, 37–41. doi: 10.1590/S1517-83822013005000002
Park, S. H., Aydin, M., Khatiwara, A., Dolan, M. C., Gilmore, D. F., Bouldin, J. L., et al. (2014). Current and emerging technologies for rapid detection and characterization of Salmonella in poultry and poultry products. Food Microbiol. 38, 250–262. doi: 10.1016/j.fm.2013.10.002
Persson, S., Jacobsen, T., Olsen, J. E., Olsen, K. E., and Hansen, F. (2012). A new real-time PCR method for the identification of Salmonella Dublin. J. Appl. Microbiol. 113, 615–621. doi: 10.1111/j.1365-2672.2012.05378.x
Ranieri, M. L., Shi, C., Moreno Switt, A. I., den Bakker, H. C., and Wiedmann, M. (2013). Comparison of typing methods with a new procedure based on sequence characterization for Salmonella serovar prediction. J. Clin. Microbiol. 51, 1786–1797. doi: 10.1128/JCM.03201-12
Ricke, S. C., Khatiwara, A., and Kwon, Y. M. (2013). Application of microarray analysis of foodborne Salmonella in poultry production: a review. Poult. Sci. 92, 2243–2250. doi: 10.3382/ps.2012-02740
Rodrigue, D. C., Tauxe, R. V., and Rowe, B. (1990). International increase in Salmonella enteritidis: a new pandemic? Epidemiol. Infect. 10, 21–27. doi: 10.1017/s0950268800047609
Saeki, E. K., Alves, J., Bonfante, R. C., Hirooka, E. Y., and Oliveira, T. C. R. M. (2013). Multiplex PCR (mPCR) for the detection of Salmonella spp. and the differentiation of the Typhimurium and Enteritidis serovars in chicken meat. J. Food Safety 33, 25–29. doi: 10.1111/jfs.12019
Soria, M. C., Soria, M. A., and Bueno, D. J. (2012). Comparison of 2 culture methods and PCR assays for Salmonella detection in poultry feces. Poult. Sci. 91, 616–626. doi: 10.3382/ps.2011-01831
Tatavarthy, A., and Cannons, A. (2010). Real-time PCR detection of Salmonella species using a novel target: the outer membrane porin F gene (ompF). Lett. Appl. Microbiol. 50, 645–652. doi: 10.1111/j.1472-765X.2010.02848.x
Werber, D., Dreesman, J., Feil, F., van Treeck, U., Fell, G., Ethelberg, S., et al. (2005). International outbreak of Salmonella Oranienburg due to German chocolate. BMC Infect. Dis. 5:7. doi: 10.1186/1471-2334-5-7
Wigley, P., Hulme, S., Powers, C., Beal, R., Smith, A., and Barrow, P. (2005). Oral infection with the Salmonella enterica serovar Gallinarum 9R attenuated live vaccine as a model to characterise immunity to fowl typhoid in the chicken. BMC Vet. Res. 1:2. doi: 10.1186/1746-6148-1-2
Xiong, D., Song, L., Geng, S., Tao, J., An, S., Pan, Z., et al. (2016). One-step PCR detection of Salmonella Pullorum/Gallinarum using a novel target: the flagellar biosynthesis gene flhB. Front. Microbiol. 7:1863. doi: 10.3389/fmicb.2016.01863
Zhai, L., Kong, X., Lu, Z., Lv, F., Zhang, C., and Bie, X. (2014). Detection of Salmonella enterica serovar Dublin by polymerase chain reaction in multiplex format. J. Microbiol. Methods 100, 52–57. doi: 10.1016/j.mimet.2014.02.014
Keywords: Salmonella Enteritidis, Salmonella Pullorum/Gallinarum, Salmonella Dublin, multiplex PCR, accurate discrimination
Citation: Xiong D, Song L, Tao J, Zheng H, Zhou Z, Geng S, Pan Z and Jiao X (2017) An Efficient Multiplex PCR-Based Assay as a Novel Tool for Accurate Inter-Serovar Discrimination of Salmonella Enteritidis, S. Pullorum/Gallinarum and S. Dublin. Front. Microbiol. 8:420. doi: 10.3389/fmicb.2017.00420
Received: 12 January 2017; Accepted: 28 February 2017;
Published: 16 March 2017.
Edited by:
David Rodriguez-Lazaro, University of Burgos, SpainReviewed by:
Alberto Quesada, University of Extremadura, SpainXiaohui Zhou, University of Connecticut, USA
Copyright © 2017 Xiong, Song, Tao, Zheng, Zhou, Geng, Pan and Jiao. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Zhiming Pan, em1wYW5AeXp1LmVkdS5jbg==
Xinan Jiao, amlhb0B5enUuZWR1LmNu
 Dan Xiong1,2,3,4
Dan Xiong1,2,3,4 Zhiming Pan
Zhiming Pan