Abstract
The black rot pathogen Xanthomonas campestris pv. campestris (Xcc) is a model organism for the study of plant bacterial pathogenesis mechanisms. In bacteria, σ factors serve as important regulatory elements that respond to various environmental signals and cues. Though Xcc encodes 15 putative σ factors little is known about their roles. As an approach to identify the potential role of each σ factor, we constructed mutations in each of the σ-factor genes as well as generating mutants deficient in multiple σ factors to assess these regulators potential additive functions. The work identified two σ70 factors essential for growth. Furthermore, the work discovered a third σ70 factor, RpoE1, important for virulence. Further studies revealed that RpoE1 positively regulates the expression of the hrp gene cluster that encodes the type III secretion system (T3SS) which determines the pathogenicity and hypersensitive response of Xcc on plants. In vivo and in vitro studies demonstrated that RpoE1 could bind to the promoter region and promote transcription of hrpX, a gene encoding a key regulator of the hrp genes. Overall, this systematic analysis reveals important roles in Xcc survival and virulence for previously uncharacterized σ70 factors that may become important targets for disease control.
Introduction
Bacteria have developed various mechanisms to accurately modify their behavior in response to changes in their environmental soundings to facilitate growth, survival, and infection. One important class of global regulators/signaling systems are the sigma (σ) factors which are typically master transcriptional regulators of gene expression. They are subunits of RNA polymerase (RNAP) and are required for transcription initiation, including the recognition and opening of promoters as well as the initial steps in RNA synthesis (Paget, 2015). These regulators are responsible for RNA synthesis in exponentially growing cells, with bacteria possessing multiple sigma factors that are used coordinately to regulate the expression of genes involved in diverse functions, including stress responses, iron uptake, morphological development, and chemotaxis (Saecker et al., 2011). The σ factors can be classified into two structurally and evolutionarily distinct families: the σ54 family and the σ70 family (Kazmierczak et al., 2005; Feklístov et al., 2014; Zhang and Buck, 2015; Davis et al., 2017). Most bacteria harbor a single number of σ54 family factors, which regulate the expression of genes involved in various cellular processes such as nitrogen metabolism, phage shock, flagellar motility, virulence, and cellular differentiation (Studholme and Buck, 2000; Studholme, 2002; Davis et al., 2017). However, many bacterial genomes encode a large number of σ70 family proteins, which can be subdivided into four groups: group I consists of housekeeping σ70 factors and groups II–IV are comprised of alternative σ70 factors with specialized functions. Normally, groups II and III members regulate the expression of genes involved in general stress responses, motility, and chemotaxis (Paget and Helmann, 2003). Group IV members, also known as ECF (extracytoplasmic function) σ70 factors and are numerically the largest group of σ70 family factors regulating the expression of genes involved in sensing and responding to stressful and adverse environmental conditions (Kazmierczak et al., 2005; Davis et al., 2017).
The genus Xanthomonas is a group of Gram-negative plant-associated bacteria that belong to the Gamma subdivision of Proteobacteria (Ryan et al., 2011; Rodriguez et al., 2012). Xanthomonas comprises of a large number of species and some of which include multiple pathovars (pv.) or subspecies (subsp.) (Ryan et al., 2011; Rodriguez et al., 2012). These plant pathogens cause severe crop diseases in important economic crops, such as rice, pepper, tomato, cassava, and citrus. Arguably the most well studied of xanthomonad plant pathogens is Xanthomonas campestris pv. campestris (Xcc) (hereafter Xcc), the causative agent of black rot disease in vegetable brassica crops worldwide (Mansfield et al., 2012; Vicente and Holub, 2013). This is due in part to the substantial amount of genome data now available which has added to our understanding of how Xcc causes disease (Ryan et al., 2011). It is thought that Xcc persists as epiphytes on the plant surface before they enter the plant via natural openings such as hydathodes, stomata, or wounds (Ryan et al., 2011). Once inside the plant tissue they multiply either locally in the intercellular space or colonize the xylem vessels and spread systemically within the plant to cause disease. A number of virulence determinants have been identified and characterized, such as effector proteins secreted by the type III secretion system (T3SS), extracellular polysaccharides (EPS), and extracellular enzymes (Ryan et al., 2011, 2015; Vicente and Holub, 2013; Zhou et al., 2017). The reason for the remarkable success of Xcc can be attributed to its large versatility and environment-driven flexible changes in its transcriptional profile.
To study the signaling systems that underpin the complex interactions between bacterial pathogens and their hosts will facilitate our understanding of the mechanisms by which the pathogens regulate stress and virulence processes during bacterial disease. In order to achieve this establishment of the specific roles of signaling elements and their contribution to the disease process is required. Here we have addressed this issue through a study of σ factors and their involvement in the virulence of Xcc. Despite σ factors being studied extensively in other Gram-negative bacteria, only limited work has been carried out in Xcc with the σ54 factor RpoN2 shown to be required for the regulation of motility (Yang et al., 2009) and one σ70 factor being demonstrated to be involved in stress response regulation (Bordes et al., 2011). Recently, it has been demonstrated that the σ54 factors are required for full virulence of Xanthomonas citri subsp. citri and Xanthomonas oryzae pv. oryzae (Tian et al., 2015; Gicharu et al., 2016). However, no σ factor has been shown to be involved in the virulence or pathogenesis of Xcc although the sequenced genomes of Xcc strains possess fifteen predicted σ factors, two being σ54 family members and the rest being σ70 family members (da Silva et al., 2002; Qian et al., 2005; Vorhölter et al., 2008).
Using the sequenced Xcc strain 8004 we attempted to characterize the 15 predicted σ factor-encoding genes (Qian et al., 2005). As an approach to identify their potential role in Xcc, we constructed mutations in each of these genes as well as generating collective mutations to assess their potential additive functions. These strains were tested in various in vitro and in planta assays. Mutants defective in each of 13 predicted σ factors were obtained, but mutants of two σ factor genes XC_3806 (rpoD) and XC_3843 (rpoH) could not be generated, which appeared essential for viability and survival. This was confirmed when an extra copy of the target gene was introduced into Xcc cells and expressed in trans allowing its chromosomal homolog to be deleted. The in planta assays revealed that one of the mutants had significantly reduced virulence. The mutant was generated by erasing one of the group IV σ70 factors (encoded by XC_2974), which is homologous to the RpoE of Escherichia coli and named RpoE1 in this work. Further studies revealed that RpoE1 positively affects the expression of the hrp genes that encode the T3SS via directly regulating the transcription of hrpX, a gene encoding one of the key hrp regulators. Ultimately, each sigma factor mutant was evaluated for pathogenicity but only RpoE1 appeared to be important for overall virulence under the conditions tested.
Materials and methods
Bacterial strains, plasmids, and growth conditions
The bacterial strains and plasmids used in this study are listed in Supplementary Table S1. Xcc strains were grown at 28°C in the nutrient rich medium NYG (5 g of peptone, 3 g of yeast extract and 20 g of glycerol per liter, pH 7.0) (Daniels et al., 1984) and the minimal medium XCM1 (1.0 g of (NH4)2SO4, 10.5 g of K2HPO4, 4.5 g of KH2PO4, 0.246 g of MgSO4•7H2O, 2.362 g of succinic acid, 0.15 g of casamino acids per liter, pH 6.6) (Jiang et al., 2013). E. coli strains were grown in LB medium (10 g of bactotryptone, 5 g of yeast extract, 5 g of sodium chloride, and 1 g of D-glucose per liter) at 37°C. Antibiotics were added at the following concentrations as required: Ampicillin (Amp) 100 μg/ml; Kanamycin (Kan) 25 μg/ml; Rifampicin (Rif) 50 μg/ml; Spectinomycin (Spc) 50 μg/ml; Tetracycline (Tc) 5 μg/ml for Xcc and 15 μg/ml for E. coli. Sucrose and isopropyl-β-D-thiogalactopyranoside (IPTG) were added as required at 10% (w/v) and 1 mM, respectively.
DNA and RNA manipulations and conjugation between Xcc and E. coli strains
The total RNA of Xcc strains was extracted with a total-RNA extraction kit (Promega, Madison, Wisconsin, USA) and reverse transcription was performed using a cDNA synthesis kit (Takara, Dalian, China), according to the manufacturer's instructions. To assay the transcription level of the genes studied, qRT-PCR and sqRT-PCR were performed using the total RNA extracted from Xcc strains. The synergy brand (SYBR) green-labeled PCR fragments were amplified using the primer sets listed in Supplementary Table S2. The relative expression of genes was determined using the 2−ΔΔCt method (Livak and Schmittgen, 2001). The 16S rRNA gene was used as an internal standard. All sqRT-PCR and qRT-PCR tests were performed in triplicate.
Plasmids were introduced from E. coli strain into Xcc strain by triparental conjugation using pRK2073 (Supplementary Table S1) as the helper plasmid as described by Turner et al. (1985). Xcc transconjugants were selected in NYG medium supplemented with appropriate antibiotics.
Deletion mutant construction and complementation
Xcc gene deletion mutants were constructed by double-crossover homologous recombination using the suicide plasmid pK18mobsacB (Schäfer et al., 1994). For the construction of the rpoE1 deletion mutant, 700 bp upstream and 791 bp downstream fragments flanking the rpoE1 gene (i.e., XC_2974) were amplified from the genome of Xcc strain 8004 with the primer sets D2974LF/LR and D2974RF/RR (Supplementary Table S2). The fragments were cloned into the suicide plasmid pK18mobsacB, yielding pK2974D, which was transferred into strain 8004 by triparental conjugation and the transconjugants were screened on NYG plate supplemented with Rif and 10% sucrose. The obtained rpoE1 deletion mutant was further confirmed by PCR and named ΔrpoE1 (Supplementary Table S1). Other mutants with a deletion in individual σ factor-encoding gene were constructed by the same strategy and listed in Supplementary Table S1. The double deletion mutant of the two σ54 family factor-encoding genes (i.e., XC_1311 and XC_2251), named ΔrpoN1rpoN2 (Supplementary Table S1), and the mutants deficient in 9, 10, or 11 σ70 family factor-encoding genes, named Δ9, Δ10, or Δ11 (Supplementary Table S1), were constructed by deleting the genes one by one using the same method. The primers used for the construction of these mutants are listed in Supplementary Table S2.
For complementation of the rpoE1 deletion mutant ΔrpoE1 in cis, a 691 bp fragment upstream and a 739 bp fragment downstream of the intergenic region between the ORFs XC_0742 and XC_0743 were amplified using the total DNA of Xcc strain 8004 as template and the primer sets 0742LF/LR and 0742RF/RR (Supplementary Table S2), respectively. Simultaneously, an 1121 bp fragment containing the rpoE1 coding region and promoter was amplified with the primer set CC2974F/R (Supplementary Table S2). The 3 fragments were cloned together into the suicide plasmid pK18mobsacB, resulting pK2974CC, which was then introduced into the rpoE1 deletion mutant ΔrpoE1 by triparental conjugation and transconjugants were screened on NYG plate supplemented with Rif and 10% sucrose. The obtained cis-complemented mutant strain was confirmed by PCR and named CΔrpoE1 (Supplementary Table S1).
Construction of strains 8004/pJC3806, 8004/pJC3843 and 8004RpoE1Flag
To construct strains 8004/pJC3806 and 8004/pJC3843, 2471 and 1476-bp fragments containing the XC_3806 and XC_3843 coding regions and promoters were amplified with the primer sets C3806F/R and C3843F/R (Supplementary Table S2), respectively. The obtained fragments were cloned into the plasmid pLAFRJ (Supplementary Table S1) and the resulting plasmids, named pJC3806 and pJC3843, were then introduced into the Xcc strain 8004 by triparental conjugation. Transconjugants were selected in NYG medium supplemented with Rif and Tc, confirmed by PCR and named 8004/pJC3806 and 8004/pJC3843 (Supplementary Table S1), respectively.
For ChIP (Chromatin Immunoprecipitation) assay, a strain producing a 3 × Flag-tag fused RpoE1 protein (RpoE1-Flag3) was constructed. A 700 bp fragment upstream and a 682 bp fragment downstream of the ORF XC_2974 (rpoE1) stop codon were amplified using the total DNA of Xcc strain 8004 as template and the primer sets Flag2974LF/LR and Flag2974RF/RR (Supplementary Table S2), respectively. The primers were modified to give EcoRI- and BamHI- or XbaI- and HindIII-compatible ends. Simultaneously, a DNA fragment encoding 3 × Flag-tag with BamHI- and XbaI-compatible ends was synthesized. The three fragments were ligated and cloned into the EcoRI and HindIII sites of the suicide plasmid pK18mobsacB, resulting a recombinant plasmid named pKrpoE1Flag. This plasmid was introduced into Xcc strain 8004 by triparental conjugation and transconjugants were screened on selective agar plates containing 10% sucrose. The obtained recombinant strain was further confirmed by PCR and named 8004RpoE1Flag (Supplementary Table S1).
Construction of promoter reporter plasmid
A promoter-gusA transcriptional fusion reporter of rpoE1 was constructed. A 526 bp DNA fragment from 400 bp upstream to 126 bp downstream of the translational start codon of the RpoE1-encoding ORF XC_2974 was amplified using the total DNA of Xcc strain 8004 as template and the primer set GUS2974F/R (Supplementary Table S2) and fused to the promoterless gusA gene with its ribosome binding site in the plasmid pLgus (Supplementary Table S1) in an orientation to allow the gusA gene to be driven by the rpoE1 promoter. The obtained reporter plasmid was confirmed by sequencing and named pLgusrpoE1 (Supplementary Table S1).
Overproduction and purification of RpoE1-His6 protein
To overproduce 6×His-tagged RpoE1 protein, a 621 bp full length rpoE1 gene was amplified from the genome of Xcc strain 8004 using the primer sets E2974F/R (Supplementary Table S2) and cloned into the expression vector pET30a, generating the recombinant plasmid named pET2974 (Supplementary Table S1), in which rpoE1 was fused N-terminally and in frame to the 6×His tag-coding region of pET30a. The recombinant plasmid was transformed into E. coli strain BL21(DE3), resulting strain BL21(DE3)/pET2974 (Supplementary Table S1). After induction by IPTG, the cells were harvested and the 6×His-tagged RpoE1 protein (RpoE1-His6) was extracted and purified by Nickel-NTA resin.
Test of enzyme activity and EPS production
GUS activity was determined by measurement of the absorbance of OD415 using ρ-nitrophenyl-β-D-glucuronide as the substrate, as described by Jefferson et al. (1986), after growth of Xcc strains in minimal medium for 24 h. Three independent experiments were performed and three replicates were used in each experiment. Extracellular amylase, endoglucanase, and protease activity was examined using starch, carboxymethylcellulose, and skimmed milk as substrates, respectively, as described previously (Zang et al., 2007). EPS production was detected by growing bacterial cells on NYG plates containing 2% glucose at 28°C for 2 days.
Analysis of RNA-Seq data
RNA-seq data were analyzed as described by Liu et al. (2013). Differentially expressed genes were determined based on the DESeq package (Anders and Huber, 2010), in which false discovery rate (FDR) was used to determine differentially expressed genes. In this study, genes having FDR ≤ 0.001 and |log2FC| (log2 of the fold changes) ≥1 (equivalent to two times of fold change) were considered as differentially expressed.
ChIP assay
Xcc cells were grown in 1000 ml of the minimal medium XCM1 for 24 h and cross-linked by adding formaldehyde to a final concentration of 1%. After incubation for 20 min at room temperature with slow shaking, glycine was added at a final concentration of 0.125 M to quench the cross-linking reaction. Bacterial cells were collected by centrifugation at 4°C for 5 min at 8,000 g and washed twice in PBS. To lyse the cells, 10 ml of RIPA buffer (50 mM Tris-HCl pH 7.4, 150 mM NaCl, 1 mM EDTA, 1% NP-40, 0.5% sodium deoxycholate) was added and thoroughly mixed by vortexing, and then disrupted by sonication (30 min, 5 s each, with 2 s cooling between each pulse). For each ChIP sample, 50 μl of ANTI-FLAG (agarose conjugated) were added to the bacterial lysates, and incubated with gentle shaking at 4°C overnight. Unbound DNA fragments were washed using RIPA buffer, the bound DNA fragments and proteins were eluted by 0.25 M glycine (pH 2.5).
Western blotting
Bacterial proteins were separated by SDS-PAGE and were transferred onto a PVDF (polyvinylidene difluoride) membrane. After blocking with 1% milk, the proteins in the membrane were incubated with the 1:1,000 diluted anti-Flag-tag mouse monoclonal antibody as the primary antibody, followed by washing with TBST buffer [Tris 20 mM, NaCl 0.3 M, Tween 20 0.08% (V/V)] for six times. The diluted 1:1,000 horseradish peroxidase (HRP) conjugated goat anti-mouse IgG was used as the secondary antibody. The membrane was washed for six times and luminescent signal was then detected according to the manufacturer's instructions.
Electrophoresis mobility shift assay (EMSA)
The purified RpoE1-His6 protein was mixed with the 354-bp DNA fragment containing the promoter region of hrpX, which was amplified from the genome of Xcc strain 8004 by PCR using the FAM-labeled primer set PhrpX-F(FAM)/PhrpX-R(FAM) (Supplementary Table S2), in 20 μl (total volume) of binding buffer (10 mM Tris, 50 mM KCl, 1 mM DTT, pH 7.5). The mixture was incubated for 20 min at room temperature. The samples were then run on a 4% polyacrylamide gel in 0.5 × TBE electrophoresis buffer, and visualized by laser scanning after electrophoresis. A 353-bp DNA fragment containing hrpG promoter was also amplified from the genome of strain 8004 using the primer set PhrpG-F(FAM)/PhrpG-R(FAM) (Supplementary Table S2). Whether the DNA fragment interacts with RpoE1-His6 protein was also determined by EMSA in the same way.
In vitro transcription assay
An in vitro transcription assay was employed to study the effect of RpoE1 on the expression of hrpX. A 647-bp template DNA fragment extending from −438 to +209 relative to the transcriptional initiation site (TIS) of the hrpX promoter was amplified by PCR from the genome of Xcc strain 8004 using the primers ivt3076F/R (Supplementary Table S2). For the in vitro transcription, 2 nM template DNA was incubated with certain amount of RpoE1-His6 protein in the transcription buffer [40 mM Tris-HCl (pH 7.9), 6 mM MgCl2, 2 mM spermidine, 10 mM NaCl, 5 mM DTT, 5% glyceol, 50 mM KCl and 1 U RNase inhibitor]. Then, an NTP mixture (250 μM ATP, 250 μM CTP, 250 μM GTP, 20 μM UTP and 250 μM Biotin-16-UTP) and certain amount of E. coli holo RNA polymerase or core RNA polymerase were added to start the transcription. After incubation at 37°C for 30 min, the reactions were terminated by the addition of one volume of 2 × RNA Loading Dye Solution and incubation at 70°C for 10 min, and then chilled on ice for 1 min. The transcription products were run on a 4.5% polyacrylamide gel containing 7 M urea in 0.5 × TBE electrophoresis buffer. The transcripts obtained were analyzed by a phosphorimager screen (Typhoon 9410; Amersham Biosciences, Piscataway, NJ, USA). A 582-bp DNA fragment containing hrpG promoter, extending from −344 to +238 relative to the hrpG TIS, was amplified by PCR from the genome of Xcc strain 8004 using the primers ivt3077F/R (Supplementary Table S2) and used in the assay.
Plant assay
The virulence of Xcc strains was tested on the host plant Chinese radish (Raphanus sativus) by the leaf-clipping method (An et al., 2017). Briefly, two fully expanded leaves per seedling were cut with scissors dipped in the bacterial suspensions of an OD600 of 0.001 (1 × 106 CFU/ml). Thirty leaves were inoculated for each strain in each independent experiment. After being maintained at 100% humidity for 24 h, the inoculated plants were maintained in a greenhouse with 12 h day/night cycle illumination by fluorescent lamps at 25–28°C. Lesion length was measured 10 days after inoculation, and data were analyzed by t-test.
To determine the bacterial growth of Xcc strains in planta, Chinese radish leaves were inoculated by the same method used for the virulence test as described above, and five inoculated leaves for each sampling were homogenized in 9 ml of sterile distilled water. Diluted homogenates were plated on NYG plates supplemented with appropriate antibiotics. Bacterial CFU were counted after incubation at 28°C for 3 days.
The HR was tested on the pepper plant ECW-10R (Capsicum annuum cv. ECW-10R), a non-hosts plant commonly used to test the HR of Xcc. Bacterial cells from overnight culture were resuspended in sterile distilled water to an OD600 of 0.01 (1 × 107 CFU/ml). The pepper leaves were inoculated by infiltrating an ~5-μl bacterial resuspension into the leaf mesophyll tissue by using a blunt-end plastic syringe. The inoculated plants were maintained in a greenhouse with 12 h day/night cycle illumination with fluorescent lamps and a constant temperature of 28°C, and HR symptoms were observed and photographed. At least three plants were inoculated in each experiment, and each experiment was repeated at least three times. For the electrolyte leakage assay, four 0.6 cm2 leaf disks for each sample were collected from the bacteria-infiltrated area and incubated in 10 ml of ultrapure water. Conductivity was measured with a DDS-307A conductometer. Three samples were taken for each measurement in each experiment, and each experiment was repeated at least three times.
Bioinformatic and statistical analysis
The secondary structure of the sigma factors was bioinformatically determined with SMART (Simple Modular Architecture Research Tool) program (http://smart.embl-heidelberg.de). The homologs of sigma factors were obtained from the KEGG database (https://www.kegg.jp/) and by a BLAST against the NCBI database (https://blast.ncbi.nlm.nih.gov/). Student's t-test was used to determine the statistical significance of differences between the means of at least three technical replicates in an experiment. A P-value ≤ 0.05 was considered statistically significant.
Results
Characterization of σ factor signaling systems in Xcc
Fifteen putative σ factors were identified within the proteome of Xcc strain 8004 using the simplified modular architecture research tool (SMART) program and blast analysis (http://smart.embl-heidelberg.de) (Table 1). SMART analysis revealed two proteins (XC_1311 and XC_2251) as members of σ54 family and the rest are σ70 family (Table 1). Of the σ70 family members, one (XC_3806), two (XC_3843 and XC_2281), and 10 (XC_0556, XC_1193, XC_1474, XC_2566, XC_2905, XC_2934, XC_2974, XC_3099, XC_3383 and XC_3864) belong to group I, group III, and group IV, respectively (Table 1). Notably, only 14 σ factors in the genome of the strain were annotated and XC_3864 was predicted as a hypothetical protein (Qian et al., 2005, Table 2). The σ70 group I member XC_3806 is highly homologous to the housekeeping σ factor RpoD, while the group III members XC_3843 and XC_2281 are highly homologous to the housekeeping σ factor RpoH and the flagellar σ factor FliA of E. coli (Table 2). Apart from XC_2974 which was annotated as RpoE, all other group IV members were only annotated as sigma factors (Qian et al., 2005, Table 2). Given that all the group IV members have a motif similar to RpoE, we named XC_2974 RpoE1 and the others RpoE2 to RpoE10 based on the annotation of Xcc strain B100 and X. campestris pv. vesicatoria strain 85-10 (Table 2). Importantly, homologs of these predicted σ factors were found in other bacterial plant pathogens. All were highly conserved among sequenced Xcc strains (Table 2). The homologs of the two predicted σ54 factors XC_1311 (RpoN1) and XC_2251 (RpoN2) are found in all species of Xanthomonas and Ralstonia solanacearum, but only XC_1311 homolog is found in other plant associated bacterial species of Pseudomonas and Erwinia (Table 2).
Table 1
| ID (gene name) | Motif analysis | |
|---|---|---|
| σ70FAMILY | ||
| Group 1 | XC_3806 (rpoD) |  |
| Group 3 | XC_3843 (rpoH) |  |
| XC_2281 (fliA) |  | |
| Group 4 | XC_2974 (rpoE1) |  |
| XC_3383 (rpoE2) | 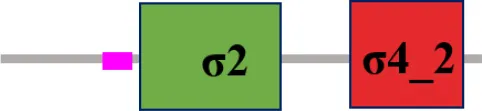 | |
| XC_2905 (rpoE3) | 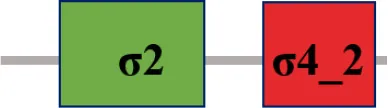 | |
| XC_2566 (rpoE4) | 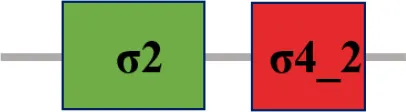 | |
| XC_1474 (rpoE5) | 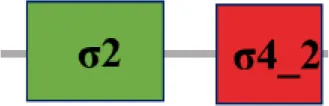 | |
| XC_1193 (rpoE6) | 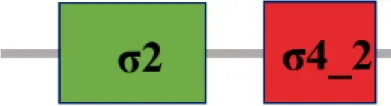 | |
| XC_2934 (rpoE7) | 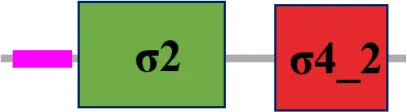 | |
| XC_3864 (rpoE8) |  | |
| XC_0556 (rpoE9) | 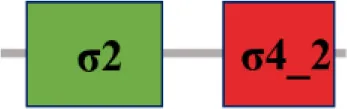 | |
| XC_3099 (rpoE10) | 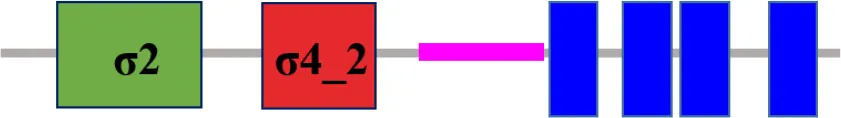 | |
| σ54 Family | XC_1311 (rpoN1) |  |
| XC_2251 (rpoN2) |  | |
The ORFs annotated as sigma factors in Xcc strain 8004*.
The protein sequence of σ70 family members can be divided into four regions (σ1, σ2, σ3, σ4) which are further divided into subregions on the basis of sequence conservation. A σ54 protein is divided into three regions (AID, CBD, DBD). Ner, non-essential region;  , Low complexity region;
, Low complexity region;  : Transmembrane domain.
: Transmembrane domain.
Table 2
| Rename in Xcc strain 8004 | Xanthomonas | Pseudomonas | Ralstonia | Erwinia | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation inXcc** | Xcv | Xac | Xoo | Xoc | Pst | Pa | Rs | Ea | ||||
| 8004 | ATCC 33913 | B100 | 85–10 | 306 | PXO99 | BLS256 | DC3000 | PAO1 | GMI1000 | CFBP1430 | ||
| σ70FAMILY | ||||||||||||
| Group 1 | XC_3806 rpoD | XC_3806 rpoD | XCC3736 (100) rpoD | xcc-b100_3917 (98.4) rpoD | XCV3912 (97.6) rpoD | XAC3788 (89.2) rpoD | PXO_04069 (97.1) rpoD | XOC_4120 (97.6) rpoD | PSPTO_0537 (62.2) rpoD | PA0576 (62.9) rpoD | RSc2215 (56.2) rpoD | EAMY_0415 (60.6) rpoD |
| Group 3 | XC_3843 rpoH | XC_3843 rpoH | XCC3771 (100) rpoH | xcc-b100_3953 (100) rpoH | XCV3949 (98.3) rpoH | XAC3824 (98.3) rpoH | PXO_03711 (98.9) rpoH | XOC_0605 (98.6) rpoH | PSPTO_0430 (58.7) rpoH | PA0376 (58.0) rpoH | RSc0374 (56.1) rpoH | EAMY_3492 (58.8) rpoH |
| XC_2281 fliA | XC_2281 fliA | XCC1906 (100) fliA | xcc-b100_2201 (100) fliA | XCV1977 (98.8) fliA | XAC1933 (99.2) fliA | PXO_00957 (97.2) fliA | XOC_2329 (98.8) fliA | PSPTO_1979 (54.3) fliA | PA1455 (50.8) fliA | RSp1390 (47.2) fliA | EAMY_2139 (49.6) fliA | |
| Group 4 | XC_2974 rpoE1 | XC_2974 rpoE | XCC1267 (100) rpoE | xcc-b100_3036 (100) rpoE | XCV1370 (97.6) rpoE2 | XAC1319 (97.6) algU | PXO_01711 (95.6) rpoE | XOC_3188 (97.6) | PSPTO_4224 (61.7) rpoE | PA0762 (62.6) algU | RSc1055 (58.4) rpoE | EAMY_2631 (55.8) rpoE |
| XC_3383 rpoE2 | XC_3383 σ70 | XCC0847 (100) σ70 | xcc-b100_3503 (100) σ70 | XCV0954 (86.9) rpoE1 | XAC0922 (87.3) | PXO_04572 (88.4) | XOC_0976 (88.5) | N | N | RSc2078 (40.2) | N | |
| XC_2905 rpoE3 | XC_2905 σ70 | XCC1334 (100) rpoE | xcc-b100_2963 (100) σ70 | XCV1436 (95.2) rpoE3 | XAC1380 (95.2) rpoE | PXO_01782 (95.7) | XOC_3114 (94.1) | PSPTO_5176 (34.8) | N | RSc2361 (40.0) | EAMY_0593 (28.0) | |
| XC_2566 rpoE4 | XC_2566 σ70 | XCC1665 (99.0) rpoE | xcc-b100_2594 (99.5) rpoE4 | XCV1718 (93.3) rpoE4 | XAC1682 (93.3) rpoE | PXO_00212 (93.3) | XOC_2834 (93.8) | N | N | N | N | |
| XC_1474 rpoE5 | XC_1474 σ70 | XCC2643 (100) rfaY | xcc-b100_2995 (26.9) rpoE5 | XCV2975 (75.2) rpoE5 | XAC2814 (77.8) rfaY | PXO_04771 (69.9) | XOC_1631 (77.2) | N | N | N | N | |
| XC_1193 rpoE6 | XC_1193 σ70 | XCC2916 (100) σ70 | xcc-b100_1237 (97.9) σ70 | XCV3224 (89.7) rpoE6 | N | N | N | N | PA2896 (38.2) | N | N | |
| XC_2934 rpoE7 | XC_2934 σ70 | XCC1306 (100) algU | xcc-b100_1518 (19) σ70 | N | N | N | XOC_3146 (77.2) | N | N | N | N | |
| XC_3864 rpoE8 | XC_3864 hypothetical protein | XCC3792 (97.6) hypothetical protein | xcc-b100_3976 (97.9) σ70 | N | N | N | N | N | PA1351 (45.6) | RSp0636 (61.8) | N | |
| XC_0556 rpoE9 | XC_0556 σ70 | XCC3593 (100) fecI | xcc-b100_0572 (100) σ70 | XCV4222 (31.2) rpoE9 | XAC4129 (30.8) rpoE | N | N | PSPTO_1209 (47.9) | PA3899 (52.4) | RSp0849 (42.4) prhI | N | |
| XC_3099 rpoE10 | XC_3099 σ70 | XCC1143 (100) rfaY | xcc-b100_3196 (99.8) σ70 | XCV1276 (70.8) rfaY | N | PXO_04856 (76.8) | XOC_3475 (76.1) | N | N | N | N | |
| σ54 Family | XC_1311 rpoN1 | XC_1311 σ54 | XCC2802 (100) rpoN | xcc-b100_1358 (100) rpoN1 | XCV3118 (93.1) rpoN2 | XAC2972 (93.5) rpoN2 | PXO_02227 (93.8) rpoN1 | XOC_1439 (93.7) rpoN | PSPTO_4453 (42.5) rpoN | PA4462 (43.2) rpoN | RSc0408 (42.4) rpoN1 | EAMY_0316 (42.4) rpoN |
| XC_2251 rpoN2 | XC_2251 σ54 | XCC1935 (100) rpoN | xcc-b100_2232 (100) rpoN2 | XCV2016 (91.6) rpoN1 | XAC1969 (92.3) rpoN1 | PXO_00995 (93.1) rpoN2 | XOC_2368 (92.9) rpoN | N | N | Rsp1671 (37.8) rpoN2 | N | |
Distribution of the 15 putative σ factors among other phytopathogenic bacteria*.
Numbers in parentheses represent percentages of amino-acid identity; N denotes no homologous (or similar) protein was found.
Xcc, X. campestris pv. campestris; Xcv, X. campestris pv. vesicatoria; Xac, X. axonopodis pv. citri; Xoo, X. oryzae pv. oryzae; Xoc, X. oryzae pv. oryzicola; Pst, P. syringae pv. tomato; Pa, P. aeruginosa; Rs, R. solanacearum; Ea, E. amylovora.
RpoE1 is a putative σ70 factor which influences Xcc pathogenesis
To investigate whether these putative σ factors are involved in the regulation of pathogenesis we constructed a panel of strains defective for each gene encoding an individual σ factor. These deletion mutants were generated by a double-crossover homologous recombination strategy using the suicide plasmid pK18mobsacB. Only 13 of the 15 predicted σ factor mutants were obtained in Xcc 8004. Despite several attempts strains with XC_3806 or XC_3843 successfully deleted could not be attained. Given that no mutant strain could be generated for XC_3806 or XC_3843 and that the proteins encoded by these two ORFs are highly homologous to the housekeeping σ factors RpoD and RpoH, it is very possible that these genes might play a role in the viability of Xcc. To verify this, we performed the deletion mutagenesis in the recombinant strains 8004/pJC3806 and 8004/pJC3843 respectively. These strains were constructed by introducing the vector pLAFRJ containing a copy of XC_3806 or XC_3843 respectively into the wild-type strain 8004 (Supplementary Table S1). This allows the chromosomal copy of XC_3843 or XC_3806 to be deleted (Supplementary Figure S1). This result provides evidence demonstrating that XC_3843 (rpoD) or XC_3806 (rpoH) are housekeeping genes in Xcc strain 8004.
In parallel, mutants were tested for their impact on virulence in planta using the Chinese radish leaf-clipping assay. Ten days post-inoculation the majority of mutants tested produced disease symptoms similar to the wild-type strain 8004 with mean lesion lengths from 10.78 to 12.17 mm (t-test, P > 0.1) (Figure 1A, Supplementary Table S3). However, in the case of strain ΔrpoE1 (XC_2974) it showed reduced disease symptoms compared to wild-type with a mean lesion length of 8.09 mm which was significant (t-test, P = 0.007) (Figure 1A, Supplementary Table S3). Furthermore, examining the growth rate of strain ΔrpoE1 in the host plant revealed that bacterial cell number (cfu, colony forming units) was lower than that of wild type (Figure 1B). Importantly, phenotypes of the mutant strain ΔrpoE1 for virulence and in planta growth could be restored toward the wild-type phenotype by introduction vector expressing rpoE1 (CΔrpoE1) (Figure 1). These data suggest that the σ70 factor RpoE1 is required for Xcc full virulence. The growth rate of ΔrpoE1 in the nutrient rich medium NYG or the minimal medium XCM1 was similar to the wild type (Supplementary Figure S2), suggesting that mutation of rpoE1 did not have an impact on Xcc growth.
Figure 1
Although RpoE1 had a strong influence on virulence the other individual σ70 factors did not appear to contribute to virulence regulation (Figure 1). However, it could not be excluded that several of these σ70 factors may have redundant functions and/or collectively contribute to virulence. As a result individual mutations would not reveal any insight of their regulation. To evaluate whether any of other σ70 factor-encoding genes may have a collective effect on Xcc virulence, we constructed a mutant strains, named Δ9, Δ10, and Δ11 (Supplementary Table S1). The Δ9 strain has the other nine σ70 factor-encoding genes (except rpoE1, rpoD and rpoH) deleted sequentially, while the Δ10 also had rpoE1 deleted and Δ11 the additional σ70 factor-encoding gene deleted. The Δ9 and Δ10 strains were tested for virulence using the Chinese radish leaf-clipping assay (Figure 1). The Δ9 showed similar disease symptoms and in planta growth rate to the wild-type strain (Figure 1). However, the mutant Δ10 and Δ11 displayed similar disease symptoms and in planta growth rate to the mutant strain ΔrpoE1 but no further additive effect was seen (Figure 1). The double deletion mutant strain (named ΔrpoN1rpoN2) of the σ54 family members RpoN1 and RpoN2 could still induce wild-type disease symptoms (Figure 1A). Taken together, it is clear that RpoE1, RpoD, and RpoH play a significant role in Xcc physiology and pathogenesis. However, no roles in virulence for the other σ54 or σ70 factors were revealed under the tested conditions.
RpoE1 influences T3SS but not extracellular enzymes or extracellular polysaccharides secretion
Xcc is a very adaptable pathogen and employs several virulence mechanisms and factors when causing disease. These include secreted factors like extracellular enzymes (such as protease, endoglucanase, and amylase) and the extracellular polysaccharides (EPS) that contribute collectively to Xcc virulence (Ryan et al., 2011, 2015; Vicente and Holub, 2013; Zhou et al., 2017). The most study of virulence mechanisms in Xcc is the T3SS that translocates effector proteins into host cells and is essential for disease. Xcc effector proteins induce disease symptoms on susceptible host plants and the hypersensitive response (HR) on resistant host or non-host plants (Ryan et al., 2011, 2015; Vicente and Holub, 2013; Zhou et al., 2017). To gain an insight into the specific virulence functions that the σ70 factor RpoE1 may influence in Xcc we examined EPS production, extracellular enzyme activity as well as HR induction. To do this we compared mutant strain ΔrpoE1 with those of the wild-type. The results demonstrated that ΔrpoE1 and the wild-type produced comparable levels of EPS, extracellular protease, endoglucanase, and amylase activities (Supplementary Figure S3), suggesting that RpoE1 is not involved in the production of these virulence factors.
To examine if RpoE1 is involved in the T3SS, the mutant strain ΔrpoE1 was examined for its ability to induce HR in the pepper cultivar ECW-10R (Capsicum annuum cv. ECW-10R). The wild-type strain 8004 carries the T3SS effector AvrBs1 and elicits visible HR symptoms on the leaves 8 h after inoculation (Xu et al., 2008). In this study, after 8 h post inoculation the ΔrpoE1 strain triggered a delayed and weakened HR, compared to the wild type (Figure 2A). Furthermore, the ΔavrBs1 strain, an AvrBs1-defective mutant, was also unable to elicit visible HR symptoms in the pepper cultivar ECW-10R (Supplementary Table S1). Interestingly, ΔrpoE1 strain could elicit visible HR symptoms 16 h post-inoculation. However, the HR symptoms were weaker compared to those elicited by the wild-type (Figure 2A). Importantly, the complemented strain CΔrpoE1 caused similar HR symptoms to the wild-type 8 h after inoculation (Figure 2A), indicating that the HR-induction capability of ΔrpoE1 could be restored by rpoE1 in cis complementation.
Figure 2
These observations were substantiated using an electrolyte leakage assay. Where the ΔrpoE1 mutant displayed significantly lower electrolyte leakages at 8, 12, and 16 h after inoculation compared with the wild-type (Figure 2B). Similar to the leaf clipping virulence assays the mutant Δ9 and Δ10 were tested along with the ΔrpoE1 strain. Like the leaf clipping assays the Δ10 demonstrated similar impact on HR and electrolyte leakage as the ΔrpoE1 strain, while the Δ9 mirrored the wild-type strain phenotypes (Figure 2). Taken together, these data suggest that the σ70 factor RpoE1 is involved in the T3SS but not extracellular enzymes or EPS secretion.
Global transcriptome profiling reveals the extended scope of regulation by σ70 factor RpoE1 in Xcc
One of the important features of bacterial σ70 factor is the activation of selective gene transcription depending on the environmental conditions (Kazmierczak et al., 2005; Davis et al., 2017). To gain an understanding of the impact of RpoE1 on Xcc gene transcription we compared the global transcriptional profile of rpoE1 deletion mutant (ΔrpoE1 strain) with the wild-type. These strains were grown in minimal medium XCM1 as it mimics to an extent the conditions in planta, given that genes involved in virulence such as the hrp cluster (T3SS) are active (Jiang et al., 2013). The total RNA was isolated from planktonic cultures growing in exponential phase. Three biological replicates were analyzed. This transcriptome profile analysis revealed that of the 4273 annotated genes, 131 genes were found to be differentially expressed; of which, 21 and 110 were up- and down-regulated, respectively (Table 3). To confirm the expression changes, semi-quantitative RT-PCR (sqRT-PCR) was employed to analyze selected genes. For the 10 selected genes expression was consistent with changes seen in the global transcriptome analysis (Table 3).
Table 3
| Gene ID | log2 Ratio (ΔrpoE1/WT) | Predicted product | sqRT-PCR results WT ΔrpoE1 |
|---|---|---|---|
| XC_0117 | +1.044 | Hypothetical protein | |
| XC_0118 | +1.007 | Transcriptional regulator | |
| XC_1263 | +1.155 | MFS transporter | |
| XC_1298 | +1.904 | Pectate lyase II | |
| XC_1386 | +1.114 | Oxidoreductase | |
| XC_1514 | +1.01 | Extracellular protease | |
| XC_1515 | +1.309 | Extracellular protease | |
| XC_1789 | +1.251 | Glutathione S-transferase | |
| XC_2580 | +1.462 | Endonuclease | |
| XC_2772 | +1.617 | Serine peptidase | |
| XC_2782 | +1.002 | Hypothetical protein | |
| XC_2838 | +1.487 | Multidrug resistance efflux pump | |
| XC_2839 | +1.409 | Outer membrane efflux protein | |
| XC_3054 | +1.797 | endo-1,3-beta-glucanase | |
| XC_3549 | +2.831 | Hypothetical protein | |
| XC_3550 | +2.599 | Serine protease | |
| XC_3575 | +1.550 | Protease | |
| XC_3576 | +1.599 | Outer membrane protein | |
| XC_3591 | +3.184 | Pectate lyase |  |
| XC_3956 | +2.324 | Hypothetical protein | |
| XC_4318 | +2.021 | Avirulence protein AvrXccA1 |  |
| XC_0052 | −1.233 | Avirulence protein AvrBs2 | |
| XC_0241 | −1.735 | Type III effector XopXccN |  |
| XC_0253 | −1.088 | Dipeptidyl anminopeptidase | |
| XC_0268 | −1.422 | Putative type III effector protein XopR | |
| XC_0334 | −1.297 | Transcriptional regulator, MarR family | |
| XC_0361 | −1.605 | MFS transporter | |
| XC_0419 | −1.365 | Hypothetical protein | |
| XC_0431 | −1.202 | VirK protein | |
| XC_0519 | −1.671 | Ice nucleation protein | |
| XC_0541 | −1.101 | Conserved hypothetical protein | |
| XC_0542 | −1.502 | Conserved hypothetical protein | |
| XC_0563 | −1.409 | Conserved hypothetical protein | |
| XC_0705 | −1.297 | Endopolygalacturonase | |
| XC_0784 | −1.002 | Cellulase S | |
| XC_0817 | −1.741 | Hypothetical protein | |
| XC_0932 | −1.082 | Hypothetical protein | |
| XC_0933 | −1.170 | Hypothetical protein | |
| XC_1084 | −5.523 | Hypothetical protein |  |
| XC_1111 | −1.801 | Cytochrome P450 hydroxylase | |
| XC_1146 | −1.754 | Hypothetical protein | |
| XC_1147 | −2.794 | Hypothetical protein | |
| XC_1210 | −1.268 | Conserved hypothetical protein | |
| XC_1422 | −1.336 | Cysteine protease | |
| XC_1441 | −1.594 | Hypothetical protein | |
| XC_1442 | −2.238 | Serine protease | |
| XC_1445 | −2.271 | Oxidoreductase | |
| XC_1446 | −1.581 | Oxidoreductase | |
| XC_1447 | −2.795 | Serine protease | |
| XC_1448 | −2.502 | Hypothetical protein | |
| XC_1449 | −2.639 | Serine protease | |
| XC_1450 | −2.678 | Serine protease |  |
| XC_1553 | −1.756 | Avirulence protein AvrAC | |
| XC_1713 | −1.391 | Hypothetical protein | |
| XC_1715 | −1.176 | Peptidase | |
| XC_1740 | −1.728 | Hypothetical protein | |
| XC_1811 | −1.762 | Virulence protein | |
| XC_1849 | −2.003 | Polygalacturonase | |
| XC_2004 | −1.847 | Avirulence protein AvrXccC | |
| XC_2081 | −1.251 | Avirulence protein AvrBs1 | |
| XC_2082 | −1.671 | Tyrosine phosphatase | |
| XC_2405 | −1.341 | Transport transmembrane protein | |
| XC_2406 | −1.785 | Hypothetical protein | |
| XC_2407 | −1.822 | Hypothetical protein | |
| XC_2408 | −2.099 | Hydroxyproline-rich glycoprotein DZ-HRGP | |
| XC_2409 | −1.989 | Hypothetical protein | |
| XC_2410 | −1.982 | Hypothetical protein |  |
| XC_2411 | −1.372 | Hypothetical protein | |
| XC_2412 | −1.383 | Hypothetical protein | |
| XC_2413 | −1.377 | Type IV secretion system NTPase VagA | |
| XC_2414 | −2.028 | Hypothetical protein | |
| XC_2415 | −2.229 | Hypothetical protein | |
| XC_2416 | −2.427 | Hypothetical protein | |
| XC_2417 | −1.289 | Plasmid mobilization protein | |
| XC_2512 | −1.794 | TonB-dependent receptor | |
| XC_2546 | −1.564 | MFS transporter | |
| XC_2547 | −1.671 | ABC transporter ATP-binding protein | |
| XC_2602 | −1.551 | Avirulence protein AvrXccE1 | |
| XC_2691 | −1.554 | Hypothetical protein | |
| XC_2827 | −1.326 | MarR family transcriptional regulator |  |
| XC_2972 | −2.373 | Periplasmic protease | |
| XC_2973 | −2.325 | Regulatory protein | |
| XC_2974 | −18.599 | RNA polymerase sigma factor RpoE2 | |
| XC_2995 | −1.925 | Type III effector XopXccE1 | |
| XC_2996 | −1.738 | Hypothetical protein | |
| XC_2998 | −1.785 | Hypothetical protein | |
| XC_2999 | −2.385 | Hypothetical protein | |
| XC_3000 | −3.919 | Hypothetical protein |  |
| XC_3001 | −3.048 | Hpa2 protein | |
| XC_3002 | −2.958 | Hpa1 protein | |
| XC_3003 | −2.535 | HrcC protein | |
| XC_3004 | −2.437 | HrcT protein | |
| XC_3005 | −2.128 | HrpB7 protein | |
| XC_3006 | −2.536 | HrcN protein | |
| XC_3007 | −2.444 | HrpB5 protein | |
| XC_3008 | −2.407 | HrpB4 protein | |
| XC_3009 | −2.384 | HrcJ protein | |
| XC_3010 | −2.418 | HrpB2 protein | |
| XC_3011 | −2.552 | HrpB1 protein | |
| XC_3012 | −2.205 | HrcU protein | |
| XC_3013 | −2.232 | HrcV protein | |
| XC_3014 | −2.401 | HpaP protein | |
| XC_3015 | −2.164 | HrcQ protein | |
| XC_3016 | −2.533 | HrcR protein | |
| XC_3017 | −2.543 | HrcS protein | |
| XC_3018 | −2.428 | HpaA protein | |
| XC_3019 | −2.466 | HrpD5 protein | |
| XC_3020 | −2.205 | HrpD6 protein | |
| XC_3021 | −2.398 | HrpE protein | |
| XC_3022 | −2.481 | HpaB protein | |
| XC_3023 | −2.196 | HrpW protein | |
| XC_3024 | −2.441 | Hypothetical protein | |
| XC_3025 | −2.583 | HrpF protein | |
| XC_3076 | −1.226 | HrpX protein | |
| XC_3147 | −1.778 | Hypothetical protein |  |
| XC_3148 | −1.557 | DNA polymerase III subunit alpha | |
| XC_3159 | −1.232 | Beta-glucosidase | |
| XC_3160 | −1.858 | Type III effector XopXccR | |
| XC_3176 | −1.369 | Type III effector protein | |
| XC_3177 | −1.810 | Type III effector XopXccQ | |
| XC_3178 | −1.009 | Hypothetical protein | |
| XC_3425 | −1.177 | Transcriptional regulator | |
| XC_3426 | −1.329 | Protocatechuate 4,5-dioxygenase subunit alpha | |
| XC_3427 | −1.331 | Protocatechuate 4,5-dioxygenase subunit beta | |
| XC_3676 | −1.268 | Chorismate mutase | |
| XC_3802 | −1.402 | Avirulence protein AvrXccB | |
| XC_3895 | −1.458 | Disulfide-isomerase | |
| XC_3922 | −2.161 | Hypothetical protein |  |
| XC_4206 | −1.495 | Hypothetical protein | |
| XC_4273 | −1.615 | Type III effector XopXccLR | |
| XC_4326 | −1.226 | Phosphatase | |
| 16S rRNA |  |
Genes whose expression was altered in the rpoE1 deletion mutant ΔrpoE1*.
WT, wild type strain; “+” and “–” represent genes whose expression was increased and decreased in the rpoE1 deletion mutant ΔrpoE1, compared to the wild type strain 8004.
Of the 131 differentially expressed genes, 39 were predicted to encode hypothetical proteins (Table 3) with their functions still remaining to be investigated in Xcc. Interestingly, 51 of the down-regulated genes are grouped in three large clusters, i.e., XC_1441-XC_1450 (except XC_1443 and XC_1444) (9.022 kb, lies between nucleotides 1738592 and 1747613 in the genome), XC_2405-XC_2417 (16.425 kb, lies between nucleotides 2912173 and 2928598 in the genome), and XC_2995-XC_3025 (except XC_2997) (27.626 kb, lies between nucleotides 3591815 and 3619441 in the genome) (Table 3). Cluster A (XC_1441-XC_1450) comprises of four-serine protease, one oxidoreductase and two hypothetical protein genes and one pseudogene (Table 3). It is known that the major extracellular protease PrtA, encoded by XC_3379, responsible for almost all extracellular protease activity of Xcc strain 8004. Inactivation of prtA leads to almost complete loss of extracellular protease activity (Meng et al., 2011). Therefore, it is not surprising that deletion of rpoE1 did not reduce significantly the extracellular protease activity, although RpoE1 regulates four serine protease genes but not prtA. Of the 13 genes in cluster B (XC_2405-XC_2417), nine encode hypothetical proteins and the others encode transport transmembrane protein, hydroxyproline-rich glycoprotein, putative NTPase, and plasmid mobilization protein, respectively (Table 3). Surprisingly, cluster B is absent in other Xcc strains and is probably a recent horizontally acquired DNA segment in strain 8004, as the segment is linked with mobile genetic elements (ORFs XC_2402 and XC_2417) (da Silva et al., 2002; Qian et al., 2005; He et al., 2007; Vorhölter et al., 2008). Importantly for virulence, cluster C (XC_2995-XC_3025) includes the hrp genes which includes 24 well-defined hrp, hrc (hrp-conserved), and hpa (hrp-associated) genes as well as one type III effector gene (XC_2995) (Table 3). Furthermore, the down-regulated effector genes include the avrBs1 (XC_2081) which is shown to contribute to HR in pepper cultivar ECW-10R (Xu et al., 2008). Additionally, XC_3076, which encodes the hrp master regulator HrpX is also down-regulated (Table 3).
Taken together, data reveals that deletion of rpoE1 significantly decreased the expression of hrp/hrc/hpa and type III effector genes, suggesting that RpoE1 plays a positive role in the regulation during conditions that mimic in planta infection.
RpoE1 influences the expression of hrp/hrc/hpa and Type III effector genes via the master regulator HrpX
Global transcriptome profiling described above reveals RpoE1 positively influences the expression of the hrp gene cluster and related genes. The Xcc hrp gene cluster consists of six main operons (hrpA-F) (Huang et al., 2009). To validate the positive regulation of these operons by RpoE1, the expression levels of the operons in the mutant strain ΔrpoE1 were determined by qRT-PCR. As illustrated in Figure 3A, the mRNA levels of all the hrp operons (hrpA-F) in ΔrpoE1 cells were reduced by 76–86% compared to that in the wild-type. Two representative type III effector genes, XC_0241 and XC_1553, were also examined. Their expression was also reduced by 81 and 86% in ΔrpoE1, respectively (Figure 3A).
Figure 3
In order to gain real time promoter activation data on the two representative hrp operons, hrpB and hrpF, and the effector genes XC_0241 and XC_1553 a set of GUS promotor assays were generated. To this end, a promoter-gusA transcriptional fusion was constructed with these target operons (or genes) (Supplementary Table S1). The resulting reporter strains were grown in XCM1 minimal medium for 24 h and GUS activity was recorded. GUS activity produced by each of the promoter-reporters in ΔrpoE1 cells was reduced by 60–78% compared to wild type (Figure 3B). Taken together, these data indicate that the σ factor RpoE1 regulates positively the expression of hrp gene cluster and a large number of type III effector genes.
Previous studies have demonstrated that the expression of all the hrp genes in the Xcc is positively controlled by several key regulators that include HpaS, HrpG, and HrpX (Huang et al., 2009; Li et al., 2014). Global transcriptome analysis revealed that hrpX is one of the down-regulated genes in the ΔrpoE1 cells grown in XCM1 minimal medium (Table 3). HrpX is an AraC-type transcriptional activator, which directly controls the expression the hrp operons and many type III effector genes (Koebnik et al., 2006; Huang et al., 2009). Our promoter reporter assays and qRT-PCR analyses confirmed that the expression of hrpX but not hrpG is indeed regulated positively by RpoE1 (Figure 3). These results imply that RpoE1 may regulate the expression of the hrp operons and the type III effector genes via HrpX. As an approach to demonstrate this idea we constitutively expressed hrpX in ΔrpoE1 strain bypassing the requirement of RpoE1 for the expression of the hrp operons and the effector genes. This was achieved by introducing the recombinant plasmid pR3X constitutively expressing hrpX (Supplementary Table S1) into the rpoE1 deletion mutant ΔrpoE1. The expression levels of the hrp operons and the effector genes XC_0241 and XC_1553 in the obtained recombinant strain ΔrpoE1/pR3X were determined by qRT-PCR. The result showed that the expression levels of all of the tested hrp operons and effector genes in ΔrpoE1/pR3X were significantly higher than those in the rpoE1 deletion mutant ΔrpoE1 and the wild type 8004 (Supplementary Figure S4), indicating that constitutive expression of hrpX could bypass RpoE1 for the expression of the genes tested. HR induction capability of strain ΔrpoE1/pR3X was also tested. The result displayed that the strain could produce wild-type HR symptoms (Figure 4). As expected, constitutively expressing HrpG in the mutant ΔrpoE1 (i.e., strain ΔrpoE1/pR3G) could not restore HR induction (Figure 4). Taken together, the above data demonstrates that the σ70 factor RpoE1 influences the expression of hrp and type III effector genes via HrpX.
Figure 4
The expression of rpoE1 is not regulated by HrpG but is induced in minimal medium and by plant extract
The expression of hrpX is positively controlled by HrpG, the OmpR-type response regulator of a two-component signal transduction system (Huang et al., 2009). Although our data suggest that RpoE1 does not directly affect hrpG expression, there is a possibility that HrpG may modulate the expression of rpoE1. To test this hypothesis, we assessed whether removal of HrpG alters the expression of rpoE1 by promoter reporter assay. Using promoter-gusA transcriptional fusion reporter of rpoE1, named pLgusrpoE1 (Supplementary Table S1), which was constructed by fusing a DNA fragment containing the rpoE1 promoter region to the promoterless gusA gene with its ribosome binding site and cloning the fused DNA segment into the vector pLAFR6. The reporter plasmid pLgusrpoE1 was introduced into the hrpG deletion strain (ΔhrpG) and the wild-type strain. These strains (ΔhrpG/pLgusrpoE1 and 8004/pLgusrpoE1) were grown in NYG and XCM1 media and after 24 h the GUS activities were measured. Results showed that the GUS activities produced by the two strains were similar (Figure 5A), indicating that lack of HrpG did not affect the expression of rpoE1. This was substantiated by qRT-PCR analysis (Figure 5B). These data indicate that HrpG does not regulate the expression of rpoE1. As illustrated in Figure 5B, qRT-PCR analysis showed that the expression level of rpoE1 in the wild-type cells grown in XCM1 was almost 4 times as high as grown in NYG. Moreover, the result from the promoter reporter assay also displayed that the GUS activity produced by 8004/pLgusrpoE1 in XCM1 was twice as high as in NYG (Figure 5A). These data reveal that the expression of rpoE1 is induced in the minimal medium XCM1. The expression of rpoE1 was further determined in XCM1 medium supplemented with plant extract from Chinese radish leaves. As shown in Figure 5C, the GUS activity produced by 8004/pLgusrpoE1 in XCM1 supplemented with plant extract was significantly higher than that in XCM1, suggesting that the expression of rpoE1 is stimulated in host plant.
Figure 5
RpoE1 induces and modulates expression of hrpX in vivo and in vitro
Our qRT-PCR and promoter reporter analyses demonstrated that hrpX is still expressed at a certain level in the rpoE1 deletion mutant background (Figure 3) suggesting that RpoE1 is not an essential factor but an enhancer for hrpX expression activity. In order to gain evidence to support this extrapolation, rpoE1 was overexpressed in wild-type strain and the expression level of hrpX gene was assessed. For this purpose, the promoterless rpoE1 was cloned into the vector pLAFRJ in an orientation allowing rpoE1 to be driven by the lac promoter and the resulting recombinant plasmid named pJrpoE1 was introduced into the wild-type. The results from qRT-PCR analysis showed that expression level of hrpX was enhanced approximately six times in the rpoE1 overexpressing strain (8004/pJrpoE1), compared to the wild type (Figure 6A). This indicates that RpoE1 could enhance hrpX expression in vivo.
Figure 6
In vitro evidence was also acquired using a 6 × His-tagged RpoE1 protein (RpoE1-His6). This protein was constructed, overexpressed, and purified from E. coli. Using the RpoE1-His6 protein an in vitro transcription assay was carried out where the 647 bp template DNA fragment extending from −438 to +209 relative to the TIS (transcription initiation site) of Xcc hrpX gene and the E. coli RNAP holoenzyme () or core enzyme () were used. The result showed that for the RNAP holoenzyme a certain amount of hrpX transcripts could be generated without RpoE1-His6 protein; however, the hrpX transcript level was significantly increased when RpoE1-His6 protein was added to the reaction mixture, even in the case with RNAP core enzyme only (Figure 6B). The data suggest that RpoE1-His6 could enhance significantly hrpX transcription in vitro. As expected, RpoE1-His6 did not have an effect on in vitro transcription of hrpG (Figure 6B).
RpoE1 binds to the hrpX promoter region in vivo and in vitro to modulate gene transcription
As discussed above, bacterial σ70 factors activate gene transcription by binding to the promoter region of target genes (Kazmierczak et al., 2005; Davis et al., 2017). To confirm that RpoE1 regulates directly the expression of hrp/hrc/hpa and type III effector genes by binding the promotor of hrpX we carried out several in vivo and in vitro assays.
Initially, an in vivo RpoE1 protein-hrpX promoter DNA complex assay was carried out by ChIP assay. A wild-type strain expressing the RpoE1-Flag3 (8004RpoE1Flag) was generated (Supplementary Table S1), for this a DNA segment encoding 3×Flag tag was fused to the 3′ end of the rpoE1 gene in the genome of the wild-type. The resulting strains were grown in the minimal medium XCM1 for 24 h and used for the ChIP assay. A Western blot assay showed that the 3×Flag fused RpoE1 protein (RpoE1-Flag3) could be eluted from the sample strain 8004RpoE1Flag but not the wild-type strain 8004 (Figure 7A). As illustrated in Figure 7A, the result of ChIP assay showed that using the eluted DNA from RpoE1-Flag3 protein as template, PCR product was obtained by the primer pair designed for amplification of DNA fragment containing hrpX promoter, but no product could be obtained by the primers for hrpG or hrpA-F operons, suggesting that RpoE1 binds directly to hrpX promoter region in vivo.
Figure 7
For in vitro determination of whether RpoE1 interacts with the hrpX promoter an EMSA was carried out. Here, a 354-bp DNA fragment containing the hrpX promoter region from 294 bp upstream to 60 bp downstream of hrpX translational start codon was amplified by the FAM-labeled primers and designated as PhrpX. The binding ability of RpoE1-His6 to PhrpX was then determined by EMSA. A 353 bp FAM-labeled fragment containing hrpG promoter, named PhrpG, was also obtained and included in the EMSA experiment. As shown in Figure 7B, the RpoE1-His6 protein alone did not interact with PhrpX; however, it could bind PhrpX and retarded PhrpX to a defined position when the core RNAP was added. No interaction between RpoE1-His6 and PhrpG was observed in the experiment (Figure 7B). This suggests that the binding of RpoE1-His6 to the hrpX promoter was specific. Taken together, these data suggest that RpoE1 directly interacts with the hrpX promoter region.
Discussion
Here, we report the systematic characterization of σ factors that are involved in virulence regulation in the phyotpathogen Xcc. This work led to the identification of important roles in Xcc survival and pathogenesis for previously uncharacterized Xcc σ factors including the alternative σ70 factor RpoE1 and the role it plays in virulence and the regulation of T3SS.
The Xcc strain 8004 possesses 15 ORFs encoding two putative σ54 and 13 σ70 factors (Table 1). Our bioinformatic analysis revealed that these ORFs are highly conserved in the other two completely sequenced Xcc genomes, i.e., ATCC33913, and B100 (da Silva et al., 2002; Vorhölter et al., 2008). Moreover, the 13 predicted σ70 family members could be assigned to group I (1 identified), III (2 identified), and IV (10 identified). The group I member encoded by XC_3806 was predicted as the housekeeping σ factor RpoD that is indispensable for the expression of essential genes. Consequently, no mutant strain could be generated for XC_3806 (rpoD). Similarly, no mutant strain could be created for XC_3843, a gene encoding a σ70 group III member homologous to RpoH, which was found to be essential σ factor for the growth of E. coli and Francisella tularensis (Zhou et al., 1988; Grall et al., 2009). This work showed that when an extra copy of XC_3843 (rpoH) was introduced into Xcc cells in trans, the chromosomal XC_3843 (rpoH) could be removed from the cells. Similarly, a mutation of XC_3806 (rpoD) could be generated in the same way. These data provide evidence indicating that both RpoD and RpoH are essential σ factors for Xcc growth in the conditions tested (in the nutrient rich medium NYG and the optimum growth temperature 28°C). As shown in Table 1, RpoD possesses four domains named σ1, σ2, σ3, and σ4, while RpoH has only σ2, σ4, and partial σ1 domains. It is believed that a Gram-negative bacterium have over 300 essential genes that are critical for survival and growth (Juhas et al., 2014). Structurally different RpoD and RpoH are central to growth raises a lot of questions that would be interesting to address going forward. These would include asking the questions: Do RpoD and RpoH control all the essential genes in Xcc together or they control subsets of essential genes independently? If they work together, how do they regulate the expression of the target genes? Notably, homologs of these two σ factors are present not only in all other Xanthomonas species but also in most plant associated species of Pseudomonas, Ralstonia, and Erwinia (Table 2). This suggests that this mechanism for viability control is present in these strains but it would be interesting to know whether they are essential for these strains also.
The remaining predicted σ factor-encoding genes could be individually deleted from the genome of Xcc strain 8004. Using these mutants in leaf clipping assays revealed that a strain deficient in the σ70 factor RpoE1, had attenuated virulence toward the host plant Chinese radish but the strains defective for the other σ factors displayed virulence similar to that of the wild-type (Figure 1). RpoE1 is one of the 10 predicted σ70 family group IV members. Deletion of all of the other nine group IV members, i.e., mutant strain Δ9, did not alter the virulence of Xcc (Figure 1). Furthermore, the mutant strain Δ10, in which all of the 10 group IV members were deleted, had similar virulence to the rpoE1 deletion mutant ΔrpoE1 (Figure 1). These data suggest that all of the predicted σ70 family factors except RpoE1 are not involved in Xcc virulence in the tested conditions. Notably, unlike many other bacteria which encode only one σ54 factor, the overwhelming majority of xanthomonads harbor two σ54 factors (RpoN1 and RpoN2). To date, the whole genome sequences of 28 Xanthomonas species or pathovars are available and 26 of them have two σ54 factors (https://www.ncbi.nlm.nih.gov/genome/?term=xanthomonas). In addition to xanthomonads, some other bacteria also contain more than one σ54 factors. For instance, Ralstonia solanacearum and Rhodobacter sphaeroides have two and four σ54 factors, respectively (Domenzain et al., 2012; Ray et al., 2015). It has been shown that RpoN1, but not RpoN2, is required for twitching motility, natural competence, growth on nitrate, and virulence of Ralstonia solanacearum (Ray et al., 2015). In X. oryzae pv. oryzae RpoN2 is required for flagellar motility and full virulence (Tian et al., 2015). Both RpoN1 and RpoN2 contribute to the virulence of X. citri subsp. citri (Gicharu et al., 2016). Interestingly, the rpoN1 mutant of X. citri subsp. citri showed a reduction in cell motility, while the rpoN2 mutant increased cell motility, suggesting that the RpoN1 and RpoN2 play diverse roles in X. citri subsp. citri (Gicharu et al., 2016). Our data presented here showed that the mutant strain ΔrpoN1rpoN2 defective in both σ54 factors still had wild-type virulence (Figure 1), suggesting that the σ54 factors are not involved in Xcc virulence. Similarly, previous work demonstrated that one of the rpoN genes is not required for the pathogenicity of X. campestris pv. vesicatoria (Horns and Bonas, 1996). Yang et al. (2009) showed that the RpoN2 of Xcc is essential for motility and normal flagellar biogenesis. It is worthy to investigate whether the σ54 factors in Xcc are involved in any other cellular processes in addition to flagellar biogenesis. Only the mutation of rpoE1 resulted in a significant reduction in pathogenicity and HR of Xcc in plants. This was mirrored by gene expression analysis revealing that RpoE1 had a role to play in controlling the expression of the hrp gene cluster and therefore regulating positively the T3SS. It is well known that the T3SS is crucial for the pathogenicity of many Gram-negative bacterial pathogens, including a large number of important plant pathogens in the genera Erwinia, Pantoea, Pectobacterium, Pseudomonas, Ralstonia as well as Xanthomonas (He, 1998; Galán and Collmer, 1999; Cornelis and Van Gijsegem, 2000; Galán and Wolf-Watz, 2006). The T3SS systems of these pathogens are encoded by the hrp cluster of genes which is highly conserved and acquired by horizontal gene transfer (Gürlebeck et al., 2006). However, it is believed that due to evolution, the pathogens have formed two groups based on their manner of hrp gene transcriptional regulation (Tang et al., 2006; Mole et al., 2007). The first group (Group A) includes the pathogens in the genera Erwinia, Pantoea, Pectobacterium, and Pseudomonas, while the second group (Group B) is composed of the pathogens of the genera Ralstonia and Xanthomonas. The expression of the hrp gene cluster in Group A is directly activated by the alternative σ70 factor named HrpL, while in Group B is controlled by AraC-type transcriptional regulator named HrpX (for Xanthomonas) or HrpB (for Ralstonia) (Tang et al., 2006; Mole et al., 2007). It has been demonstrated that the expression of hrpX is positively regulated by the two-component signal transduction system composed of the sensor histidine kinase HpaS and the OmpR-type response regulator HrpG (Büttner and Bonas, 2010; Li et al., 2014). Transcriptome, qRT-PCR, and promoter reporter analyses showed that mutation of rpoE1 reduced the expression of hrpX (but not hrpG) (Table 3, Figure 3). Moreover, mutation of hrpG did not alter rpoE1 expression (Figure 5). These data imply that there is no regulatory relationship between HrpG and RpoE1 at transcriptional level. However, RpoE1 and HrpG may regulate the expression of hrpX independently of each other. Notably, a comparison revealed that RpoE1 and the HrpL of the Group A pathogens share only about 23% identity in amino acid sequences. ChIP analysis showed that unlike HrpL which directly regulates the hrp cluster of genes RpoE1 does not target any operons of the hrp gene cluster (Figure 7A). These data suggest that RpoE1 and HrpL inference the expression of hrp genes via different manners. Furthermore, as shown in Table 2, Xcc does not have a σ70 factor highly homologous to HrpL, but the Group A pathogens such as Pseudomonas syringae pv. tomato and Erwinia amylovora harbor a RpoE highly homologous to the RpoE1 of Xcc. It would be interesting to know whether this RpoE is involved in the hrp gene regulation of these pathogens.
Our ChIP and EMSA analyses revealed that RpoE1-His6 could bind to the promoter region of hrpX in vivo and in vitro (Figure 7), suggesting that RpoE1 regulates directly the expression of hrpX. In vitro transcription assay displayed that RpoE1-His6 could enhance the E. coli holo RNAP-initiated hrpX transcription (Figure 6A). More importantly, RpoE1-His6 together with the core RNAP could promote the transcription of hrpX (Figure 6B), suggesting that RpoE1 may act as the σ factor that composes the holo RNAP directing hrpX transcription in Xcc. Furthermore, bioinformatics analysis revealed that RpoE1 is a member of ECF σ70 factors that normally respond to environmental stresses. This together with the fact that the expression of rpoE1 is induced in planta suggests that the RpoE1-associated regulation of hrpX may be related to an undefined plant signal. Work to identify the signal and the mechanism by which it controls RpoE1 will be the subject for further studies. Interestingly, the homologs of RpoE1 are widely distributed in other Xanthomonas species as well as Pseudomonas, Ralstonia, and Erwinia strains suggesting a similar mechanism might be broadly conserved in many plant associated bacteria.
Statements
Author contributions
J-LT and B-LJ conceived the study. L-YY, L-CY, and Y-LG carried out the experiments. LW, W-ZZ, Y-QH, and WJ performed bioinformatics analysis. J-LT and B-LJ wrote the manuscript.
Funding
This work was supported by the Guangxi Natural Science Foundation of China (2014GXNSFFA118005), the Ba Gui Scholar Program of Guangxi Zhuang Autonomous Region of China (2014A002), and the National Natural Science Foundation of China (31660505 and 31260261).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2018.01749/full#supplementary-material
Supplementary Figure S1PCR detection showing that the chromosomal XC_3806 or XC_3843 could be deleted from the Xcc cells harboring an extra copy of XC_3806 or XC_3843 in trans. XC_3806 and XC_3843 in the recombinant strains 8004/pJ3806 and 8004/pJ3843 were deleted respectively by double-crossover homologous recombination method using the suicide plasmid pK18mobsacB as a vector. Three mutant colonies for each strain were randomly selected for PCR verification by the primer sets D3806LF/D3806RR and D3843LF/D3843RR complemented respectively to the flanking regions of XC_3806 and XC_3843 in the chromosome of strain 8004 but not any sequence in the plasmids pJ3806 and pJ3843. (A) PCR verification showing that the chromosomal XC_3806 was deleted. Lane 1: PCR product, using the total DNA of 8004 as template, with expected size of 3379 bp; lanes 2-4: PCR products, using the total DNA of selected mutant colonies as templates, with expected size of 1511 bp, suggesting that XC_3806 was removed. (B) PCR verification showing that the chromosomal XC_3843 was deleted. Lane 1: PCR product, using the total DNA of 8004 as template, with expected size of 2272 bp; lanes 2-4: PCR products, using the total DNA of selected mutant colonies as templates, with expected size of 1404 bp, suggesting that XC_3843 was removed.
Supplementary Figure S2The mutant strain ΔrpoE1 and the wild type strain 8004 had a similar growth rate. The Xcc strains were incubated in the nutrient rich medium NYG and the minimal medium XCM1. The optical density at 600 nm (OD600) of the cultures was measured at 4 h intervals.
Supplementary Figure S3The production of the extracellular amylase, endoglucanase, protease, and exopolysaccharide produced by the mutant strains ΔrpoE1 and Δ10 is no significantly different compared to the wild-type strain 8004. Exopolysaccharide production and amylase, endoglucanase, and protease activity were detected by growing bacterial cells at 28°C on NYG plates supplemented with 2% (wt/vol) glucose, 0.1% (wt/vol) starch, 0.25% (wt/vol) carboxymethycellulose, and 0.5% (wt/vol) skimmed milk, respectively. For amylase and endoglucanase, the plates were stained after incubation for 24 h and photographed. The plates for exopolysaccharide production and protease activity were photographed after incubation for 48 h. The clear zones created by the degradation of the substrates and the sticky gum in the colonies reflect the enzyme activities and exopolysaccharide production, respectively. The experiment was repeated three times and the results were similar.
Supplementary Figure S4Constitutive expression of hrpX could bypass the requirement of RpoE1 for the expression of the hrp and type III effector genes, revealed by quantitative real-time PCR (qRT-PCR). RNAs were isolated from the cells of Xcc strains grown in the minimal medium XCM1 for 24 h. The relative mRNA levels of the hrp gene cluster (hrpA-F), the master hrp regulator hrpX, and the type III effector-encoding genes XC_0241 and XC_1553 were analyzed by quantitative real-time PCR and calculated with respect to the transcript level in the wild-type strain 8004. Data are means ± standard deviations of triplicate measurements from a representative experiment; similar results were obtained in two other independent experiments. Wild type, the wild-type strain 8004; ΔrpoE1, the rpoE1 deletion mutant strain; ΔrpoE1/pR3X, ΔrpoE1 constitutively expressing hrpX.
Supplementary Table S1Bacterial strains and plasmids used in this work.
Supplementary Table S2Primers used in this study.
Supplementary Table S3Statistics analysis of the virulence of Xcc strains.
References
1
AnS. Q.TangJ. L.DowJ. M. (2017). Probing the role of cyclic di-GMP signaling systems in disease using Chinese radish. Methods Mol. Biol.1657, 205–212. 10.1007/978-1-4939-7240-1-16
2
AndersS.HuberW. (2010). Differential expression analysis for sequence count data. Genome Biol.11:R106. 10.1186/gb-2010-11-10-r106
3
BordesP.LavatineL.PhokK.BarriotR.BoulangerA.Castanié-CornetM. P.et al. (2011). Insights into the extracytoplasmic stress response of Xanthomonas campestris pv. campestris: role and regulation of σE-dependent activity. J. Bacteriol.193, 246–264. 10.1128/JB.00884-10
4
BüttnerD.BonasU. (2010). Regulation and secretion of Xanthomonas virulence factors. FEMS Microbiol. Rev.34, 107–133. 10.1111/j.1574-6976.2009.00192.x
5
CornelisG. R.Van GijsegemF. (2000). Assembly and function of type III secretory systems. Annu. Rev. Microbiol.54, 735–774. 10.1146/annurev.micro.54.1.735
6
DanielsM. J.BarberC. E.TurnerP. C.SawczycM. K.ByrdeR. J.FieldingA. H. (1984). Cloning of genes involved in pathogenicity of Xanthomonas campestris pv. campestris using the broad host range cosmid pLAFR1. EMBO J.3, 3323–3328.
7
da SilvaA. C.FerroJ. A.ReinachF. C.FarahC. S.FurlanL. R.QuaggioR. B.et al. (2002). Comparison of the genomes of two Xanthomonas pathogens with differing host specificities. Nature417, 459–463. 10.1038/417459a
8
DavisM. C.KesthelyC. A.FranklinE. A.MacLellanS. R. (2017). The essential activities of the bacterial sigma factor. Can J. Microbiol.63, 89–99. 10.1139/cjm-2016-0576
9
DomenzainC.CamarenaL.OsorioA.DreyfusG.PoggioS. (2012). Evolutionary origin of the Rhodobacter sphaeroides specialized RpoN sigma factors. FEMS Microbiol. Lett. 327, 93–102. 10.1111/j.1574-6968.2011.02459.x
10
FeklístovA.SharonB. D.DarstS. A.GrossC. A. (2014). Bacterial σ factors: a historical, structural, and genomic perspective. Annu. Rev. Microbiol.68, 357–376. 10.1146/annurev-micro-092412-155737
11
GalánJ. E.CollmerA. (1999). Type III secretion machines: bacterial devices for protein delivery into host cells. Science284, 1322–1328. 10.1126/science.284.5418.1322
12
GalánJ. E.Wolf-WatzH. (2006). Protein delivery into eukaryotic cells by type III secretion machines. Nature444, 567–573. 10.1038/nature05272
13
GicharuG. K.SunD. L.HuX.FanX. J.ZhuoT.WuC. W.et al. (2016). The sigma 54 genes rpoN1 and rpoN2 of Xanthomonas citri subsp. citri play different roles in virulence, nutrient utilization and cell motility. J. Integr. Agric.15, 2032–2039. 10.1016/S2095-3119(15)61317-X
14
GrallN.LivnyJ.WaldorM.BarelM.CharbitA.MeibomK. L. (2009). Pivotal role of the Francisella tularensis heat-shock sigma factor RpoH. Microbiology155, 2560–2572. 10.1099/mic.0.029058-0
15
GürlebeckD.ThiemeF.BonasU. (2006). Type III effector proteins from the plant pathogen Xanthomonas and their role in the interaction with the host plant. J. Plant Physiol.163, 233–255. 10.1016/j.jplph.2005.11.011
16
HeS. Y. (1998). Type III protein secretion systems in plant and animal pathogenic bacteria. Annu. Rev. Phytopathol.36, 363–392. 10.1146/annurev.phyto.36.1.363
17
HeY. Q.ZhangL.JiangB. L.ZhangZ. C.XuR. Q.TangD. J.et al. (2007). Comparative and functional genomics reveals genetic diversity and determinants of host specificity among reference strains and a large collection of Chinese isolates of the phytopathogen Xanthomonas campestris pv. campestris. Genome Biol.8:R218. 10.1186/gb-2007-8-10-r218
18
HornsT.BonasU. (1996). The rpoN gene of Xanthomonas campestris pv. vesicatoria is not required for pathogenicity. Mol. Plant Microbe Interact. 9, 856–859. 10.1094/MPMI-9-0856
19
HuangD. L.TangD. J.LiaoQ.LiX. Q.HeY. Q.FengJ. X.et al. (2009). The Zur of Xanthomonas campestris is involved in hypersensitive response and positively regulates the expression of the hrp cluster via hrpX but not hrpG. Mol. Plant Microbe Interact.22, 321–329. 10.1094/MPMI-22-3-0321
20
JeffersonR. A.BurgessS. M.HirshD. (1986). β-Glucuronidase from Escherichia coli as a gene-fusion marker. Proc. Natl. Acad. Sci. U.S.A.83, 8447–8451. 10.1073/pnas.83.22.8447
21
JiangG. F.JiangB. L.YangM.LiuS.LiuJ.LiangX. X.et al. (2013). Establishment of an inducing medium for type III effector secretion in Xanthomonas campestris pv. campestris. Braz. J. Microbiol.44, 945–952.
22
JuhasM.ReußD. R.ZhuB.CommichauF. M. (2014). Bacillus subtilis and Escherichia coli essential genes and minimal cell factories after one decade of genome engineering. Microbiology160, 2341–2351. 10.1099/mic.0.079376-0
23
KazmierczakM. J.WiedmannM.BoorK. J. (2005). Alternative sigma factors and their roles in bacterial virulence. Microbiol. Mol. Biol. Rev.69, 527–543. 10.1128/MMBR.69.4.527-543.2005
24
KoebnikR.KrügerA.ThiemeF.UrbanA.BonasU. (2006). Specific binding of the Xanthomonas campestris pv. vesicatoria AraC-type transcriptional activator HrpX to plant-inducible promoter boxes. J. Bacteriol.188, 7652–7660. 10.1128/JB.00795-06
25
LiR. F.LuG. T.LiL.SuH. Z.FengG. F.ChenY.et al. (2014). Identification of a putative cognate sensor kinase for the two-component response regulator HrpG, a key regulator controlling the expression of the hrp genes in Xanthomonas campestris pv. campestris. Environ. Microbiol.16, 2053–2071. 10.1111/1462-2920.12207
26
LiuW.YuY. H.CaoS. Y.NiuX. N.JiangW.LiuG. F.et al. (2013). Transcriptome profiling of Xanthomonas campestris pv. campestris grown in minimal medium MMX and rich medium NYG. Res. Microbiol.164, 466–479. 10.1016/j.resmic.2013.02.005
27
LivakK. J.SchmittgenT. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔC(T) Method. Methods25, 402–408. 10.1006/meth.2001.1262
28
MansfieldJ.GeninS.MagoriS.CitovskyV.SriariyanumM.RonaldP.et al. (2012). Top 10 plant pathogenic bacteria in molecular plant pathology. Mol. Plant Pathol.13, 614–629. 10.1111/j.1364-3703.2012.00804.x
29
MengQ. L.TangD. J.FanY. Y.LiZ. J.ZhangH.HeY. Q.et al. (2011). Effect of interactions between Mip and PrtA on the full extracellular protease activity of Xanthomonas campestris pathovar campestris. FEMS Microbiol. Lett.323, 180–187. 10.1111/j.1574-6968.2011.02377.x
30
MoleB. M.BaltrusD. A.DanglJ. L.GrantS. R. (2007). Global virulence regulation networks in phytopathogenic bacteria. Trends Microbiol.15, 363–371. 10.1016/j.tim.2007.06.005
31
PagetM. S. (2015). Bacterial sigma factors and anti-sigma factors: structure, function and distribution. Biomolecules5, 1245–1265. 10.3390/biom5031245
32
PagetM. S.HelmannJ. D. (2003). The σ70 family of sigma factors. Genome Biol.4:203. 10.1186/gb-2003-4-1-203
33
QianW.JiaY.RenS. X.HeY. Q.FengJ. X.LuL. F.et al. (2005). Comparative and functional genomic analyses of the pathogenicity of phytopathogen Xanthomonas campestris pv. campestris. Genome Res.15, 757–767. 10.1101/gr.3378705
34
RayS. K.KumarR.PeetersN.BoucherC.GeninS. (2015). rpoN1, but not rpoN2, is required for twitching motility, natural competence, growth on nitrate, and virulence of Ralstonia solanacearum. Front. Microbiol. 6:229. 10.3389/fmicb.2015.00229
35
RodriguezR. L. MGrajalesA.Arrieta-OrtizM. L.SalazarC.RestrepoS.BernalA. (2012). Genomes-based phylogeny of the genus Xanthomonas. BMC Microbiol. 12:43. 10.1186/1471-2180-12-43
36
RyanR. P.AnS. Q.AllanJ. H.McCarthyY.DowJ. M. (2015). The DSF family of cell-cell signals: an expanding class of bacterial virulence regulators. PLoS Pathog.11:e1004986. 10.1371/journal.ppat.1004986
37
RyanR. P.VorholterF. J.PotnisN.JonesJ. B.Van SluysM. A.BogdanoveA. J.et al. (2011). Pathogenomics of Xanthomonas: understanding bacterium–plant interactions. Nat. Rev. Microbiol.9, 344–355. 10.1038/nrmicro2558
38
SaeckerR. M.RecordM. T.Jr.DehasethP. L. (2011). Mechanism of bacterial transcription initiation: RNA polymerase - promoter binding, isomerization to initiation-competent open complexes, and initiation of RNA synthesis. J. Mol. Biol.412, 754–771. 10.1016/j.jmb.2011.01.018
39
SchäferA.TauchA.JägerW.KalinowskiJ.ThierbachG.PühlerA. (1994). Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum. Gene145, 69–73. 10.1016/0378-1119(94)90324-7
40
StudholmeD. J. (2002). Enhancer-dependent transcription in Salmonella enterica Typhimurium: new members of the sigmaN regulon inferred from protein sequence homology and predicted promoter sites. J. Mol. Microbiol. Biotechnol.4, 367–374.
41
StudholmeD. J.BuckM. (2000). The biology of enhancer-dependent transcriptional regulation in bacteria: insights from genome sequences. FEMS Microbiol. Lett.186, 1–9. 10.1111/j.1574-6968.2000.tb09074.x
42
TangX.XiaoY.ZhouJ. M. (2006). Regulation of the type III secretion system in phytopathogenic bacteria. Mol. Plant Microbe Interact.19, 1159–1166. 10.1094/MPMI-19-1159
43
TianF.YuC.LiH.WuX.LiB.ChenH.et al. (2015). Alternative sigma factor RpoN2 is required for flagellar motility and full virulence of Xanthomonas oryzae pv. oryzae. Microbiol. Res.170, 177–183. 10.1016/j.micres.2014.07.002
44
TurnerP. L.BarberC. E.DanielsM. J. (1985). Evidence for clustered pathogenicity genes in Xanthomonas campestris pv. campestris. Mol. Gen. Genet.199, 338–343. 10.1007/BF00330277
45
VicenteJ. G.HolubE. B. (2013). Xanthomonas campestris pv. campestris (cause of black rot of crucifers) in the genomic era is still a worldwide threat to brassica crops. Mol. Plant Pathol. 14, 2–18. 10.1111/j.1364-3703.2012.00833.x
46
VorhölterF. J.SchneikerS.GoesmannA.KrauseL.BekelT.KaiserO.et al. (2008). The genome of Xanthomonas campestris pv. campestris B100 and its use for the reconstruction of metabolic pathways involved in xanthan biosynthesis. J. Biotechnol.134, 33–45. 10.1016/j.jbiotec.2007.12.013
47
XuR. Q.BlanvillainS.FengJ. X.JiangB. L.LiX. Z.WeiH. Y.et al. (2008). AvrACXcc8004, a type III effector with a leucine-rich repeat domain from Xanthomonas campestris pathovar campestris confers avirulence in vascular tissues of Arabidopsis thaliana ecotype Col-0. J. Bacteriol.190, 343–355. 10.1128/JB.00978-07
48
YangT. C.LeuY. W.Chang-ChienH. C.HuR. M. (2009). Flagellar biogenesis of Xanthomonas campestris requires the alternative sigma factors RpoN2 and FliA and is temporally regulated by FlhA, FlhB, and FlgM. J. Bacteriol.191, 2266–2275. 10.1128/JB.01152-08
49
ZangN.TangD. J.WeiM. L.HeY. Q.ChenB.FengJ. X.et al. (2007). Requirement of a mip-like gene for virulence in the phytopathogenic bacterium Xanthomonas campestris pv. campestris. Mol. Plant Microbe Interact.20, 21–30. 10.1094/MPMI-20-0021
50
ZhangN.BuckM. (2015). A perspective on the enhancer dependent bacterial RNA polymerase. Biomolecules5, 1012–1019. 10.3390/biom5021012
51
ZhouL.ZhangL. H.CamaraM.HeY. W. (2017). The DSF family of quorum sensing signals: diversity, biosynthesis, and turnover. Trends Microbiol.25, 293–303. 10.1016/j.tim.2016.11.013
52
ZhouY. N.KusukawaN.EricksonJ. W.GrossC. A.YuraT. (1988). Isolation and characterization of Escherichia coli mutants that lack the heat shock sigma factor σ32. J. Bacteriol.170, 3640–3649. 10.1128/jb.170.8.3640-3649.1988
Summary
Keywords
sigma (σ) factor, housekeeping gene, virulence, hypersensitive response, type III secretion, Xanthomonas
Citation
Yang L-Y, Yang L-C, Gan Y-L, Wang L, Zhao W-Z, He Y-Q, Jiang W, Jiang B-L and Tang J-L (2018) Systematic Functional Analysis of Sigma (σ) Factors in the Phytopathogen Xanthomonas campestris Reveals Novel Roles in the Regulation of Virulence and Viability. Front. Microbiol. 9:1749. doi: 10.3389/fmicb.2018.01749
Received
16 April 2018
Accepted
12 July 2018
Published
03 August 2018
Volume
9 - 2018
Edited by
Mari-Anne Newman, University of Copenhagen, Denmark
Reviewed by
Michelle Teresa Hulin, NIAB EMR, United Kingdom; David John Studholme, University of Exeter, United Kingdom
Updates
Copyright
© 2018 Yang, Yang, Gan, Wang, Zhao, He, Jiang, Jiang and Tang.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Bo-Le Jiang jbl1971@gxu.edu.cnJi-Liang Tang jltang@gxu.edu.cn
This article was submitted to Plant Microbe Interactions, a section of the journal Frontiers in Microbiology
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.