- 1State Key Laboratory of Biocatalysis and Enzyme Engineering, Environmental Microbial Technology Center of Hubei Province, College of Life Sciences, Hubei University, Wuhan, China
- 2Mechanical Engineering and Materials Science, Washington University, St. Louis, MO, United States
- 3State Key Laboratory of Agricultural Microbiology, Huazhong Agricultural University, Wuhan, China
Poly-γ-glutamic acid (γ-PGA) is an anionic polymer with various applications. Teichoic acid (TA) is a special component of cell wall in gram-positive bacteria, and its D-alanylation modification can change the net negative charge of cell surface, autolysin activity and cationic binding efficiency, and might further affect metabolic production. In this research, four genes (dltA, dltB, dltC, and dltD) of dlt operon were, respectively, deleted and overexpressed in the γ-PGA producing strain Bacillus licheniformis WX-02. Our results implied that overexpression of these genes could all significantly increase γ-PGA synthetic capabilities, among these strains, the dltB overexpression strain WX-02/pHY-dltB owned the highest γ-PGA yield (2.54 g/L), which was 93.42% higher than that of the control strain WX-02/pHY300 (1.31 g/L). While, the gene deletion strains produced lower γ-PGA titers. Furthermore, 13C-Metabolic flux analysis was conducted to investigate the influence of dltB overexpression on metabolic flux redistribution during γ-PGA synthesis. The simulation data demonstrated that fluxes of pentose phosphate pathway and tricarboxylic acid cycle in WX-02/pHY-dltB were 36.41 and 19.18 mmol/g DCW/h, increased by 7.82 and 38.38% compared to WX-02/pHY300 (33.77 and 13.86 mmol/g DCW/h), respectively. The synthetic capabilities of ATP and NADPH were also increased slightly. Meanwhile, the fluxes of glycolytic and by-product synthetic pathways were all reduced in WX-02/pHY-dltB. All these above phenomenons were beneficial for γ-PGA synthesis. Collectively, this study clarified that overexpression of dltB strengthened the fluxes of PPP pathway, TCA cycle and energy metabolism for γ-PGA synthesis, and provided an effective strategy for enhanced production of γ-PGA.
Introduction
Poly-γ-glutamic acid (γ-PGA) is an important anionic polypeptide consisting of D-glutamic acid and/or L-glutamic acid residues, which linked together via amide bonds between α-amino and γ-carboxyl (Feng et al., 2015; Sirisansaneeyakul et al., 2017). Since γ-PGA owns the excellent features of biocompatibility, biodegradability, water solubility, edibility, non-toxicity, environmentally friendliness, etc. (Cao et al., 2018), it has the wide-range applications in the areas of food, medicine, cosmetics industries, etc. (Cai et al., 2017).
Bacillus species have been proven as the efficient γ-PGA producers, and a number of metabolic engineering strategies have been developed to improve γ-PGA yield. For example, knocking out glutamate dehydrogenase genes rocG and gudB improved glutamic acid accumulation, which led to a 38% increase of γ-PGA yield in Bacillus amyloliquefaciens (Zhang et al., 2015). The synthesis of extracellular polysaccharide and lipopolysaccharide were blocked to decrease by-product yields for γ-PGA production in B. amyloliquefaciens LL3 (Feng et al., 2015). In addition, strengthening of NADPH and ATP supplies all benefited γ-PGA production (Cai et al., 2017, 2018). Recently, cell surface engineering was proven to be an effective strategy for enhancement production of metabolics. Overexpression of phosphatidylserine synthase gene pssA could enhance the cell membrane integrity and hydrophilicity, and further improved the cell tolerance and biorenewable yields (short-chain fatty acids, organic alcohols, organic acids and other aromatic compounds, etc) in E. coli (Tan et al., 2017). Elevation of membrane cardiolipin levels via overexpressing cardiolipin synthase gene clsA significantly increased hyaluronic acid titer by 204% in Bacillus subtilis (Westbrook et al., 2018). However, no research has been focused on the relationship between cell surface engineering and γ-PGA synthesis.
Teichoic acids (TAs) is an anionic polymer composed of ribitol and glycerol residues, and it was linked by phosphodiester bonds in the cell wall. The dlt operon, which consisted of four genes dltA, dltB, dltC, and dltD, owns the function of incorporating D-alanine to TAs in Bacillus (Koprivnjak et al., 2006). Previously, removal of D-alanyl esters from TAs of Staphylococcus aureus increased the amounts of Mg2+ bound to cell wall, which might affect the activities of several membrane proteins or enzymes (Lambert et al., 1975). Previous researches implied that the expression of dlt operon could regulate the cell surface net negative charge, and the changes on surface charge could mediate the recruitment of signaling molecules (lignin) (Hyyrylainen et al., 2000; Heit et al., 2011). γ-PGA synthesis is regulated by two component systems ComP∼ComA and DegS∼DegU, the regulators DegQ and SwrA (Tran et al., 2000). The recruitment of ComP might be affected by the reduction of cell surface negative charge, and further activated these regulators for γ-PGA synthesis. Furthermore, overexpression of dltA triggered the electrostatic repulsion between S. aureus and daptomycin, which represented the keystone of DAP resistance (Cafiso et al., 2014). γ-PGA and TAs are all anionic polymer, and the electrostatic repulsion generated between γ-PGA and TAs was not conducive to γ-PGA secretion. Thus, reducing cell surface negative charge might be an effective strategy for enhancement production of γ-PGA.
The biochemical reactions in cells were difficult to be profoundly described with the existing techniques. While, acting as a new method, 13C metabolic flux analysis (13C-MFA) is a promising tool to quantify cellular metabolism reaction rates in a network via isotopomer tracer experiments (Long et al., 2016). Moreover, 13C-MFA is a powerful tool to quantify the energy generation and consumption rates in the metabolic pathways (He et al., 2014; Yao et al., 2016), and it also applied to comprehend and analyze the changes in central carbon metabolism under aerobic and anaerobic conditions. For instance, the optimal tracers [1,2-13C] glucose, [1,6-13C] glucose, [1,2-13C] xylose, and [5-13C] xylose were applied in Escherichia coli under aerobic and anaerobic conditions, and their results demonstrated that the fluxes of EMP pathway and TCA cycle were all increased under anaerobic conditions, which generated more energy, formate, alcohol and succinate to adapt the adverse environments (Gonzalez et al., 2017).
In this study, the genes dltA, dltB, dltC, and dltD of dlt operon were, respectively, deleted and overexpressed in B. licheniformis WX-02, and 13C-MFA was performed to expound the effect of dltB overexpression on metabolic flux redistribution. In addition, the transcriptional level, by-products contents were also measured during γ-PGA synthesis. The aim of this study is to illustrate the relationship between dlt operon overexpression and γ-PGA synthesis by 13C-MFA, and provides an efficient strategy of strain improvement for γ-PGA production.
Materials and Methods
Strains and Culture Conditions
Strains and plasmids used in this study were provided in Table 1. B. licheniformis WX-02 was acted as the original strain for constructing recombinants, and E. coli DH5α was served as the host strain for plasmid construction. B. licheniformis and E. coli were grown in LB medium with responsible antibiotic (20 μg/mL kanamycin, 100 μg/mL ampicillin, or 20 μg/mL tetracycline), when required.
The γ-PGA fermentation medium contains 20 g/L glucose, 5 g/L NH4NO3, 11.35 g/L Na2HPO4, 8.15 g/L KH2PO4, 0.20 g/L MgSO4⋅7H2O, 0.01 g/L MnSO4⋅H2O, 0.0008 g/L CaCl2, and 0.0045 g/L Na2-EDTA. B. licheniformis were grown in twenty-four well plates with 2 mL working volume, and cultivated at 220 rpm and 37°C for 14 h. For 13C-MFA, glucose was replaced by [1, 2-13C] glucose (Sigma-Aldrich, CAS#138079-87-5) in γ-PGA production medium.
Strain Construction
The construction procedures of dltB deletion and overexpression strains were served as the examples. For dltB deletion strain, the upstream and downstream arms of gene dltB were, respectively, amplified by the primers ΔdltB-F1/R1 and ΔdltB-F2/R2 (Supplementary Table S1), based on the genomic DNA of B. licheniformis WX-02, and fused by Splicing Overlapping Extension PCR (SOE-PCR) with primers ΔdltB-F1/R2. The fused fragment was inserted into the plasmid T2(2)-Ori at SacI/XbaI, colony PCR and DNA sequence confirmed that the dltB deletion vector was constructed successfully, named as T2-ΔdltB. Then, T2-ΔdltB was electro-transferred into B. licheniformis WX-02, and the positive transformant was cultivated in LB medium with 20 μg/mL kanamycin at 220 rpm and 45°C, and sub-cultured for three times to obtain the single-crossover recombinants. The recombinants were grown in LB medium at 37°C with six subcultures, and the kanamycin sensitive colonies were further confirmed by colony PCR and DNA sequence, and the dltB deletion strain was named WX-02ΔdltB.
For gene overexpression strain, P43 promoter from B. subtilis 168, gene dltB and amyL terminator from B. licheniformis WX-02 were, respectively, amplified (Supplementary Table S1), and fused by SOE-PCR. The fused fragment was inserted into pHY300PLK at the restriction sites EcoRI/XbaI, resulting in the plasmid pHY-dltB. Then, pHY-dltB was introduced into WX-02 by electro-transformation, resulting in the dltB overexpression strain, named WX-02/pHY-dltB.
Analytic Methods
The cell biomass was monitored by measuring OD600 using a UV-spectrophotometer-752 N (Shanghai Instrument Analysis Instrument Co., Ltd., Shanghai, China), glucose concentrations were determined via using the Enzyme Electrode Analyzer SBA-40E (Shandong Academy of Sciences, Shandong, China). The γ-PGA yield was measured according to our previously reported method (Cai et al., 2018). Briefly, the volume of 2 mL fermentation broth was mixed with 4 mL distilled water, and cells were separated by centrifugation at 12000 rpm for 6 min after adjusting the pH to 2.5∼3.0. While, three volumes of absolute ethanol were added into the supernatant after adjusting pH to 7.0, and the precipitate was dried at 80°C to a constant weight for measuring γ-PGA yield. The net negative charge was measured by determining the cation binding rate of cell surface in LB medium, according to the previously described method (Cao et al., 2017; Chen et al., 2018).
The concentrations of by-products, acetoin, 2,3-butanediol and acetic acid, were measured by GC/MS (Gas chromatograph Trace, Thermo; Triple Quadrupole Mass Spectrometer, Thermo; column: DB-WAXMS, 30 m × 0.25 mm × 0.25 μm, Thermo; United States). Briefly, the volume of 0.5 mL fermentation broth was mixed with 1.5 mL absolute ethanol, and the supernatant was separated by centrifugation at 12000 rpm for 10 min. Equal volume of ethyl acetate was added into the supernatant for extracting acetoin, 2,3-butanediol and acetate. The parameters of GC-MS were set as follow: Gas chromatographic inlet temperature was 220°C, the injection volume was 1 μL, carrier gas flow rate was 1 mL/min and the split ratio was 20:1. The temperature program: hold at 40°C for 1 min, and increased to 160°C at 6°C/min, hold for 1.5 min, and then raised to 220°C at 30°C/min, hold for 2 min. Solvent delay was set as 4 min. The range of mass to charge ratio (m/z) in MS was set between 35 and 500.
The gene transcriptional level were measured with TRIzol® Reagent (Invitrogen, United States) and PrimeScriptTM II 1st Strand cDNA Synthesis Kit (TaKaRa, Japan) according to our previously reported method (Cai et al., 2017). The primers used for RT-qPCR were listed in Supplementary Table S2, and the housekeeping gene 16S rRNA was used to normalize the gene expression data. The data were averaged and presented as the mean ± SD.
Mass Isotopomer Distribution Analysis of Amino Acids
To analyze the mass isotopomer distributions (MIDs) of amino acids, [1,2-13C] glucose was served as the sole carbon source for cell growth and γ-PGA synthesis. The cell pellets at the mid-logarithmic phase (7 h) were collected and hydrolyzed by 6 M HCl at 100°C for 24 h. The supernatant of hydrolyzate was air-dried, and the precipitate was subsequently derivatized with N-tert-butyldimethylsilyl-N-methyltrifluoroacetamide (TBDMS). Then, the derivatization products were quantified by GC/MS (Gas chromatograph Trace, Thermo; Triple Quadrupole Mass Spectrometer, Thermo; column: TG-5MS, 30 m × 0.25 mm × 0.25 μm, Thermo; United States). The injection volume and injection split ratio were 1 μL and 1:10, respectively, and carrier gas helium was 1.2 mL/min. GC temperature program was set as follows: hold at 150°C for 2 min, and increased to 280°C at 3°C/min, and increased to 300°C at 20°C/min, and then hold for 5 min. Solvent delay was set as 5 min, and the range of mass to charge ratio (m/z) in MS was set between 60 and 500 (You et al., 2012). The software WuFlux-Ms Tool was used to analyze and correct amino acid MS data (fragments of [M-57]+, [M-159]+, or [M-85]+, and [f302]) (Wahl et al., 2004). Isotopomer labeling fractions (M0, M1, M2, etc.) represent fragments containing unlabeled, singly labeled, doubly labeled amino acids, etc (Wan et al., 2017).
13C-Metabolic Flux Analysis With INCA
13C-MFA was performed by using the INCA software (Young, 2014), which is based on the elementary metabolite units (EMU) framework (Antoniewicz et al., 2007). The central metabolic network was modified according to the previous research (He et al., 2017) and KEGG pathway database. As the target product and main by-products, several steps for γ-PGA, acetoin and 2,3-butanediol syntheses, such as 2 Pyruvate → α-acetolactate + CO2, α-acetolactate → Acetoin + CO2, Acetoin+ NADH ↔ 2,3-Butanediol + NAD+, Acetoin+ NAD+ → Acetyl-CoA + NADH, Acetoin + ATP → Acetoin_ex, 2,3-Butanediol → 2,3-Butanediol_ex, Glutamate + ATP → γ-PGA, and γ-PGA → γ-PGA_ex, were added into the metabolic network (seeing in the Supplementary Table S4). The information of labeling amino acids (alanine, glycine, valine, leucine, serine, theronine, phenylalanine, aspartic acid, glutamic acid, and histidine) (Supplementary Table S3) were substituted into the model for calculating metabolic flux of central metabolism (Wan et al., 2017). The standard deviation of 13C-enrichment was set as 0.0107 for statistical analysis (Nanchen et al., 2007), and the flux estimations were performed by using MATLAB R2014a (The Mathworks Inc.).
Statistical Analyses
All samples were analyzed in triplicate, and the data were presented as the mean ± SD for each sample point. Significant differences were determined by one-way analysis of variance (ANOVA). Statistical significance was defined as P < 0.05.
Results
Effects of Deletion and Overexpression of dlt Operon on γ-PGA Production
To evaluate the relationship between dlt operon expression and γ-PGA synthesis, the genes dltA, dltB, dltC and dltD of dlt operon were deleted and overexpressed in the γ-PGA production strain B. licheniformis WX-02, resulting in the recombinant strains WX-02ΔdltA, WX-02ΔdltB, WX-02ΔdltC, WX-02ΔdltD and WX-02/pHY-dltA, WX-02/pHY-dltB, WX-02/pHY-dltC, WX-02/pHY-dltD, respectively. Based on our results of Figure 1, deletions of dltA, dltB, dltC, and dltD all reduced γ-PGA yield, which decreased by 19.81, 41.55, 22.22, and 33.33% compared with that of control strain WX-02 (2.07 g/L), respectively (Figure 1A). And deletion of one gene has no effects on the transcriptional levels of other genes involved in dlt operon (Supplementary Figure S2). Furthermore, γ-PGA yields produced by the dlt overexpression strains were 2.13, 2.55, 2.05, and 2.25 g/L, increased by 62.21, 94.66, 56.49, and 71.76% compared to WX-02/pHY300 (1.31 g/L), respectively (Figure 1B). Collectively, these above results indicated that overexpression of dlt operon benefited γ-PGA synthesis.
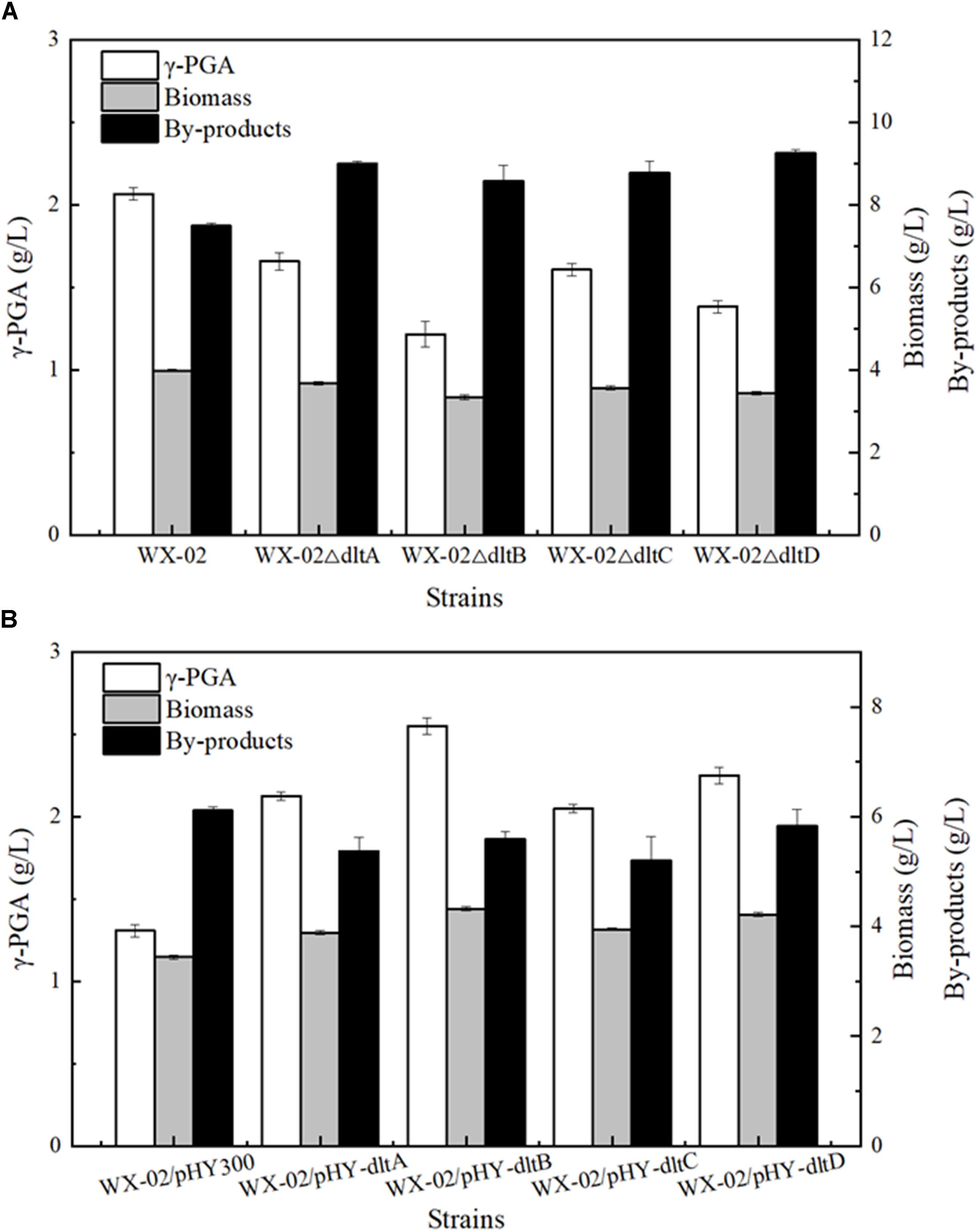
Figure 1. Effects of deletion and overexpression of dlt operon on γ-PGA and by-products production. (A) The γ-PGA yields, biomass formation and by-products yields of WX-02, WX-02ΔdltA, WX-02ΔdltB, WX-02ΔdltC, and WX-02ΔdltD. (B) The γ-PGA yields, biomass formation and by-products yields of WX-02/pHY300, WX-02/pHY- dltA, WX-02/pHY-dltB, WX-02/pHY-dltC, and WX-02/pHY-dltD.
Previously, the dlt operon was proved to play the important role on the net negative charge of cell surface. As shown in Figure 2, the negative charges of cell surface were all increased in the dlt deletion strains, and decreased in the dlt overexpression strains. The negative charge was reduced by 48.93% in the dltB overexpression strain WX-02/pHY-dltB.
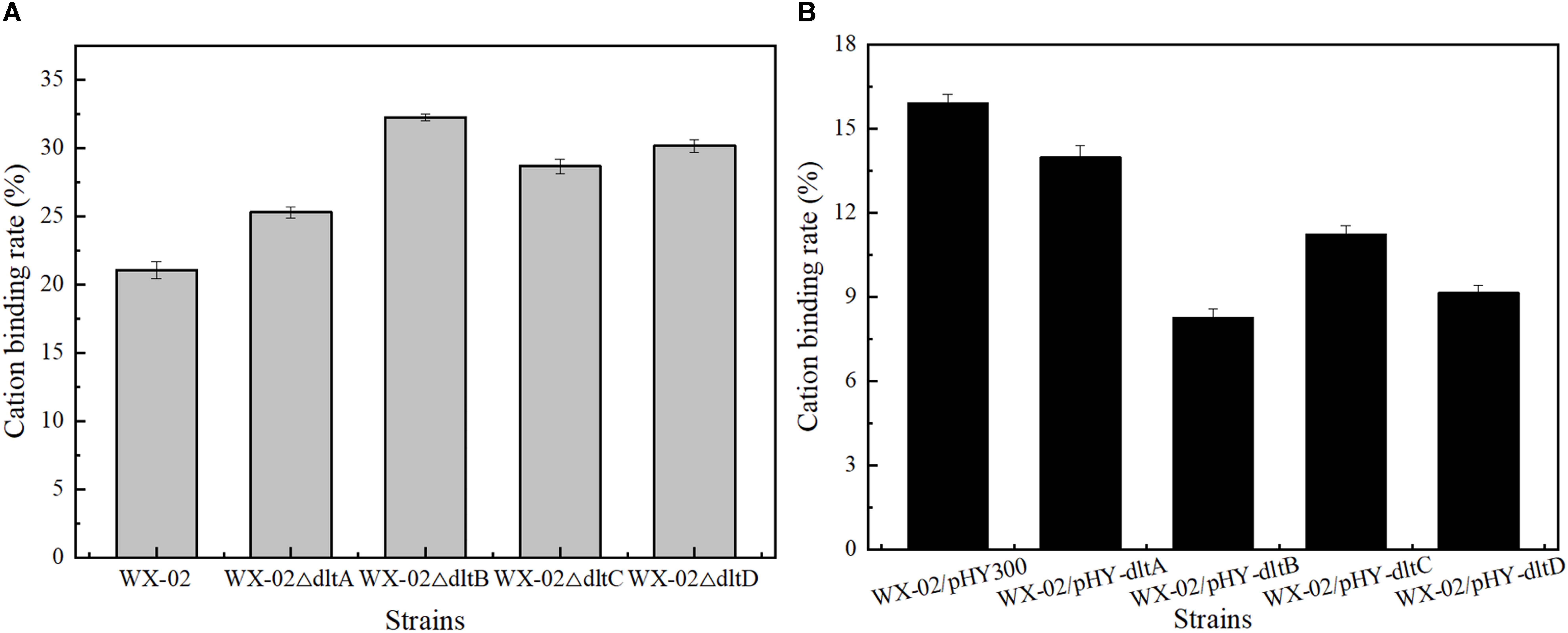
Figure 2. Effects of deletion and overexpression of dlt operon on the cation binding rate. (A) The cation binding rates of WX-02, WX-02ΔdltA, WX-02ΔdltB, WX-02ΔdltC, and WX-02ΔdltD. (B) The cation binding rates of WX-02/pHY300, WX-02/pHY-dltA, WX-02/pHY-dltB, WX-02/pHY-dltC, and WX-02/pHY-dltD.
Fermentation Process Curves of WX-02/pHY300 and WX-02/pHY-dltB
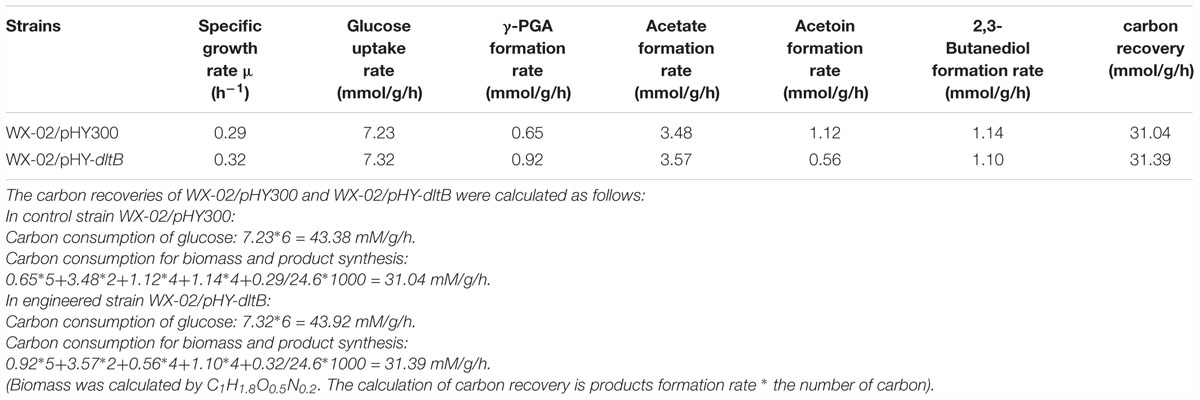
Furthermore, the process curves of WX-02/pHY-300 and WX-02/pHY-dltB were determined, and the cell growth, glucose uptake, γ-PGA yield, by-product (acetoin, 2,3-butanediol and acetic acid) concentrations were measured during γ-PGA production. As shown in Figure 3A, the cell biomass of WX-02/pHY-dltB were slightly lower than those of control strain WX-02/pHY-300 before 9 h, and then increased faster subsequently. Besides, the logarithmic growth period is delayed by 2 or 3 h, and there is no obvious decline phase in the dltB overexpression strain WX-02/pHY-dltB. The specific cell growth rate of WX-02/pHY-dltB was 0.32 h-1, increased by 7.58% compared with that of WX-02/pHY-300 (Table 2). The glucose uptake rate of WX-02/pHY-dltB was 7.32 mmol/g DCW/h, similar with that of WX-02/pHY-300 (7.23 mmol/g DCW/h), and the γ-PGA synthetic rate of WX-02/pHY-dltB (0.92 mmol/g DCW/h) was increased by 41.54% compared to WX-02/pHY-300 (0.65 mmol/g DCW/h). The concentrations of by-products (acetoin, 2,3-butanediol and acetic acid) was significantly lower in WX-02/pHY-dltB than that of WX-02/pHY-300 (Figure 3B). Moreover, the formation rates of acetic acid and 2,3-butanediol showed no significant difference between these strains, while the acetoin formation rate was decreased by 50.00% in WX-02/pHY-dltB (0.56 mmol/g DCW/h) (Table 2). Assuming that there was no other significant by-products and CO2 produced, and the carbon recoveries of WX-02/pHY300 and WX-02/pHY-dltB were roughly consistent for each strain (Table 2).
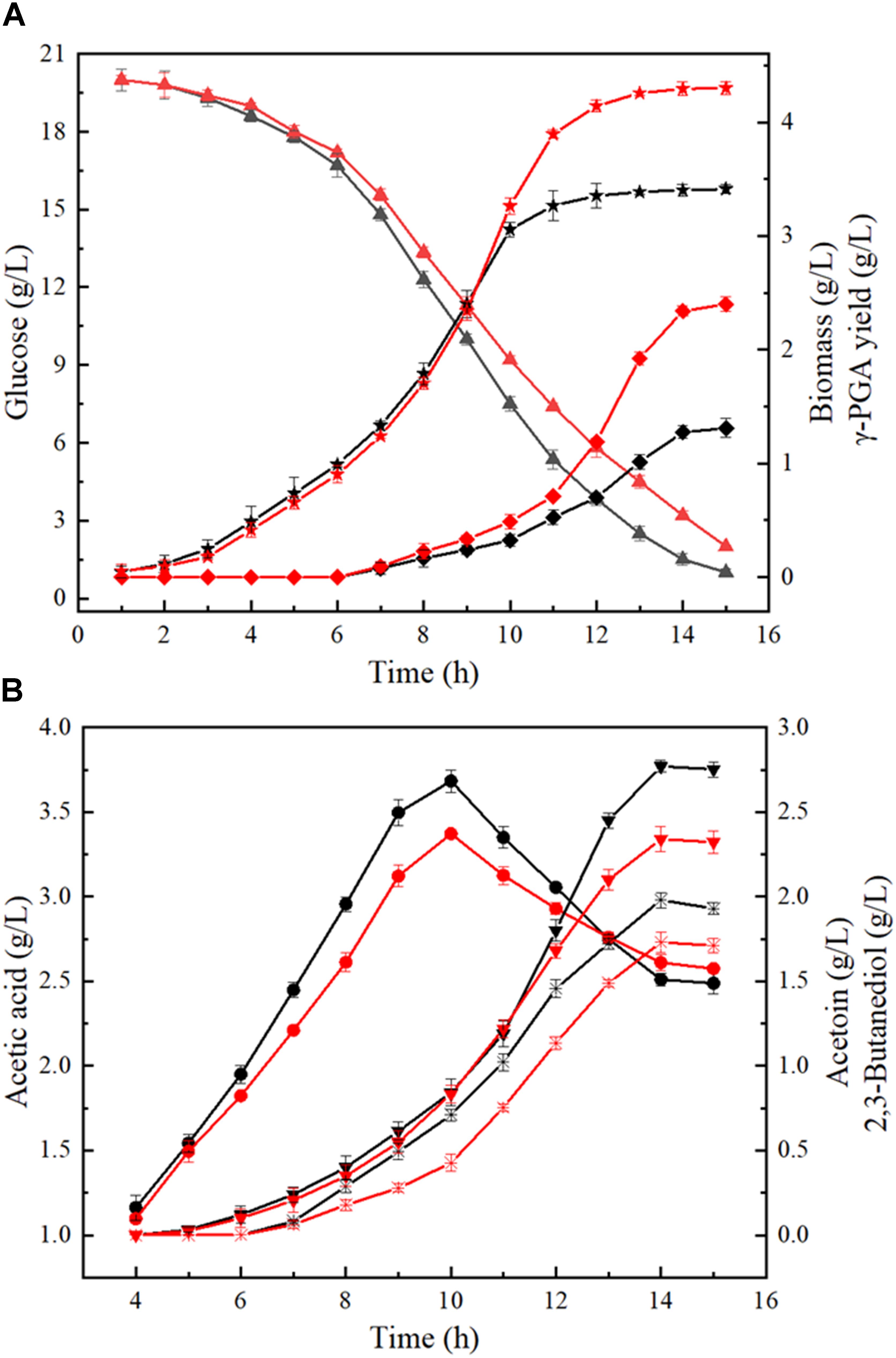
Figure 3. Fermentation process curves of WX-02/pHY300 and WX-02/pHY-dltB. (A) Biomass, glucose concentration, and γ-PGA yield. (B) Acetic acid, acetion, and 2,3-butanediol yields. The solid black line indicates the control strain WX-02/pHY300, and the red solid line represents the recombinant strain WX-02/pHY-dltB. Triangle, glucose; pentagram, biomass; diamond, γ-PGA; circle, acetic acid; inverted triangle, 2,3-butanediol; fork, acetoin.
13C-Metabolic Flux Analysis
The metabolic flux distributions were determined based on the measured mass isotopomer distributions of proteinogenic amino acids and synthetic rates of γ-PGA, acetic acid, acetoin and 2,3-butanediol. The measured mass isotopomer distributions (MIDs) of proteinogenic amino acids coincided well with the simulated MIDs (Supplementary Table S3), indicating the good fit and high flux precision between the measured and simulated data (Yao et al., 2016). The measured biomass formation rates were not employed as the constraints of 13C-MFA model (Table 2). The flux values and exchange coefficients were estimated with the 13C MFA model listed in Supplementary Table S4.
As shown in Figure 4, the fluxes in the central metabolism of B. licheniformis were redistributed in WX-02/pHY-dltB. Firstly, the flux from pyruvate to acetyl-CoA was increased by 11.12%, and the flux distributed into tricarboxylic acid cycle was enhanced by 38.38% in WX-02/pHY-dltB. The flux for acetic acid production was increased slightly, whereas, the flux of overflow metabolism (from pyruvate to acetoin and 2,3-butanediol) was decreased by 26.93%. Secondly, the flux of PP pathway was increased by 7.82% in the dltB overexpression strain, which might generate more NADPH for γ-PGA synthesis. Thirdly, the flux through pyruvate carboxylation (PYR + CO2 → OAA), the major anaplerotic flux into TCA cycle, was increased by 25.40% in WX-02/pHY-dltB. The increase of pyruvate carboxylation could enable more carbon flow from glucose to TCA cycle, and further increased the demand for oxaloacetate based biomass synthesis (Table 2; He et al., 2014). Fourthly, the fluxes from α-ketoglutaric acid to glutamic acid and γ-PGA were increased by 19.47 and 57.92%, respectively. Fifthly, the flux of EMP pathway was decreased by 4.13%, which further reduced the NADH generation and by-products (acetoin, 2,3-butanediol and acetic acid) syntheses. Lastly, the estimated biomass biosynthesis flux was slightly higher (about 6.40%) in WX-02/pHY-dltB (Figure 4), which positively correlated with the results of cell biomass (Table 1).
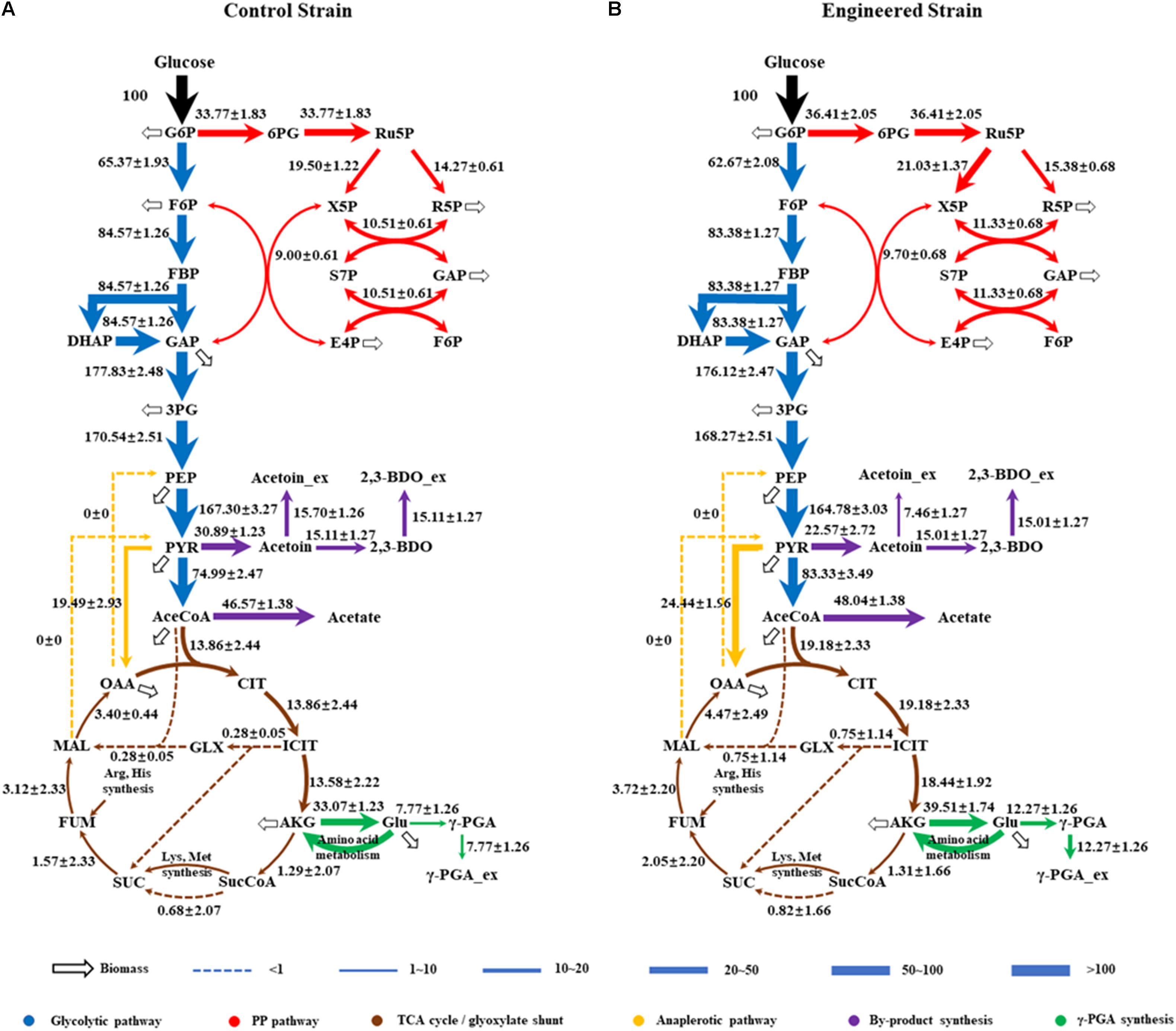
Figure 4. Metabolic flux distributions in the central metabolic pathways of WX-02/pHY300 and WX-02/pHY-dltB during exponential growth on [1,2-13C]glucose. Fluxes shown are normalized to glucose uptake rate of 100 for each strain (estimated flux ± SD). (A) WX-02/pHY300. (B) WX-02/pHY-dltB.
The cofactors (such as NADH, NADPH, etc.) and ATP play the important roles in metabolic production (Kind et al., 2013), as well as for γ-PGA synthesis (Cai et al., 2017, 2018). Based on our metabolic flux model in B. licheniformis, the NADPH and ATP formation rates in WX-02/pHY-dltB were increased by 12.50 and 3.54%, which were beneficial for γ-PGA synthesis (Table 3).
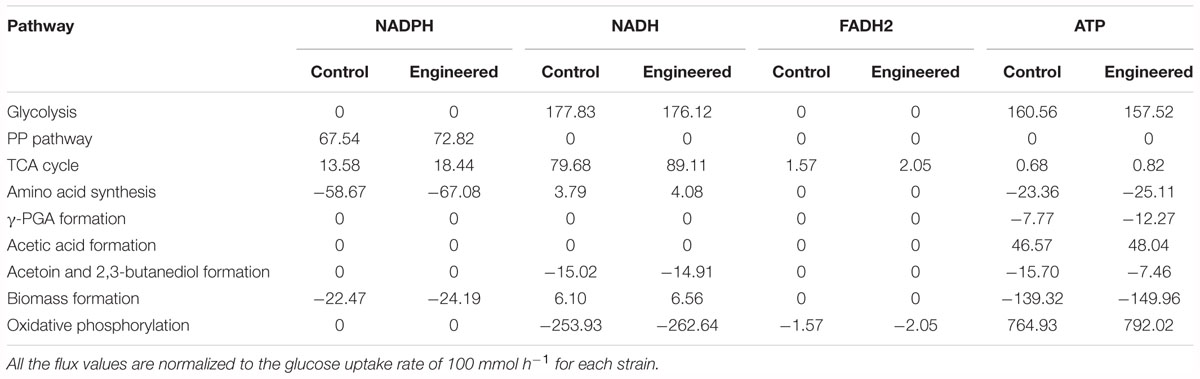
Table 3. The synthesis and consumption contents of NADPH, NADH, ATP, and FADH2 of WX-02/pHY300 and WX-02/pHY-dltB (mmol/g CDW/h).
Transcriptional Level Analysis
The transcriptional levels of genes related to glucose metabolism and γ-PGA synthesis were measured in WX-02/pHY300 and WX-02/pHY-dltB (Figure 5). Compared to WX-02/pHY300, the transcription levels of glucose transporter genes ptsG, glcU, and glcP were reduced obviously in dltB overexpression strain. The glucose-6-phosphate dehydrogenase gene zwf in PP pathway showed higher expression levels, while the transcription level of glucose-6-phosphate isomerase gene pgi in EMP pathway was decreased by 25.01%, indicating the elevated PP pathway and depressed EMP pathway fluxes in WX-02/pHY-dltB. In addition, the transcription levels of genes citB (encoding citrate synthase) and icd (encoding isocitrate dehydrogenase) in TCA cycle were enhanced by 90.03 and 105.12%, which strengthened the metabolic flux in TCA cycle.
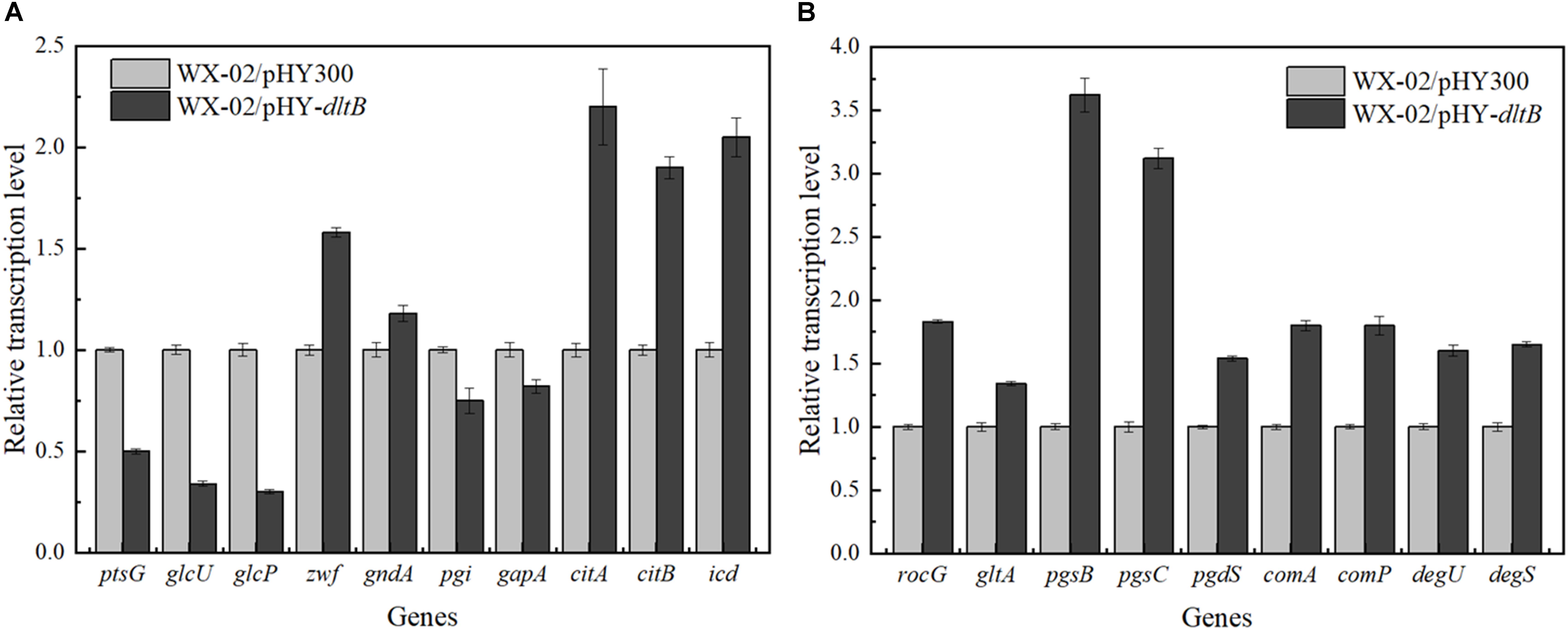
Figure 5. Transcription levels of genes in glucose metabolism and γ-PGA synthesis in WX-02/pHY300 and WX-02/pHY-dltB. (A) Genes in glucose metabolism. (B) Genes in γ-PGA synthesis.
Furthermore, the transcription levels of glutamate dehydrogenase gene rocG, which mainly catalyzes the formation of glutamic acid from α-ketoglutaric acid in B. licheniformis WX-02 (Tian et al., 2017), was improved by 83.25% in WX-02/pHY-dltB, and γ-PGA synthetase genes pgsB, pgsC were, respectively, increased by 2.62- and 2.12-fold, which indicated that more glutamic acid was synthesized and converted to γ-PGA in WX-02/pHY-dltB. Besides, the gene pgdS (encoding γ-DL-glutamyl hydrolase) was also increased in WX-02/pHY-dltB, and the improvement of pgdS could decrease the molecular weight of γ-PGA (Tian et al., 2017), positively correlated with the actual measurement results (Date not shown). Moreover, transcription factor genes comA, comP, degS, and degU, which associated with the synthesis of γ-PGA, were all increased in the dltB overexpression strain.
Discussion
Cell wall and cell membrane are the selective permeation barrier of bacteria, and engineering of cell surface negative charge, membrane phospholipid head and fatty acid hydrophobic tail composition could affect bacterial growth and product production (Hyyrylainen et al., 2007; Ghorbal et al., 2013; Tan et al., 2017). Previous researches demonstrated that the increases of negative charge on the cell surface could improve the secretion efficiency of target protein, and the increase rate of target protein with lower isoelectric points (PI) was higher than that of protein with higher PI (Cao et al., 2017; Chen et al., 2018). While, based on our results, the increases of cell surface negative charge led to the decrease of γ-PGA yield in the dlt operon mutants.
On the one hand, overexpression of dltB slowed down the absorption and utilization rate of glucose (Figure 3A), and further decreased the flux of EMP pathway. Previous research implied that reduction of glycolysis flux in E. coli THRD could reduce the accumulation of acetate and increase NADPH and L-threonine generations (Xie et al., 2014), which was consistent with our results (Figure 3A and Table 2). Moreover, the alanylation of teichoic acids could modulate the negative charge of cell wall to protect secretory or cell wall-associated proteins against degradation during the post-translocational folding (Hyyrylainen et al., 2000), and enhancement of cell surface negative charge could increase the binding capacity of autolysin, and accelerated autolysis of bacteria (Steen et al., 2005). Figure 3A showed that the logarithmic period of WX-02/pHY-dltB was extended by 2∼3 h, and the growing status of WX-02ΔdltB was obviously badness in post-fermentation compared with WX-02 (Supplementary Figure S1). This results indicated that overexpression of dltB could promote the cell growth by reducing the autolysis of cells. Therefore, overexpression of dltB reduced the absorption and utilization rate of glucose, which decreased the flux of EMP pathway and synthesis capabilities of by-products (acetoin, 2,3-butanediol and acetic acid), and extended logarithmic growth phase, and all these phenomenons were beneficial for γ-PGA synthesis.
Then, 13C-labeled isotope tracer and 13C-MFA were applied to further evaluate the metabolic flux redistributions in the dltB overexpression strain. Based on our results, the flux of EMP pathway is weakened, which reduced the NADH generation and by-products synthesis, and our results were positively consisted with the previous research (Lee and Oh, 2016). Secondly, the flux of PPP pathway and NADPH supply were all enhanced in the dltB overexpression strain. Since the conversion of α-ketoglutarate to glutamic acid requires NADPH in B. licheniformis WX-02, enhancement of NADPH supply contributes to γ-PGA synthesis (Cai et al., 2017; Tian et al., 2017). In addition, the flux of TCA cycle was enhanced by 38.38% in the dltB overexpression strain, and the increase of α-ketoglutaric acid synthesis flux was beneficial for cell growth (Wan et al., 2017) and γ-PGA production. Besides, the flux of complement-deficient pathway (PYR → OAA) was enhanced by 25.40%, which is beneficial for oxaloacetate accumulation and cell growth (Sauer et al., 1999). Finally, the flux from α-ketoglutarate to glutamate and further generate γ-PGA were, respectively, increased by 19.47 and 57.92%, which were consistent with fermentation characters (Figure 1).
Furthermore, overexpression of dlt operon led to the decrease of cell surface negative charge (Figure 2), and further changed the extracellular microenvironment (Hyyrylainen et al., 2000), which might promote the recruitment of signal molecular ComP to activate two component system ComP-ComA for γ-PGA synthesis. Based on our results, the transcriptional levels of comP/comA and degU/degS were all increased in the dltB overexpression strian, which further affected the transcription levels of genes involved in γ-PGA synthesis and carbon metabolism (Figure 5). Moreover, the decrease of cell surface negative charge could reduce the binding of cationic (e.g., Mg2+ and Ca2+, etc.) and cationic antimicrobial peptides on cell surface (Neuhaus and Baddiley, 2003), and the low levels of cationic or cationic antimicrobial peptides could activate PhoQ(R)∼PhoP system (Garcia Vescovi et al., 1996; Ren et al., 2017). In this study, the concentration of phosphate in the culture medium was too high to activate PhoQ(R)∼PhoP system (Devine, 2018) in WX-02/pHY300, however, due to the low level of Mg2+ and Ca2+ on cell surface of dltB overexpression strain, the PhoQ(R)∼PhoP might be activated in WX-02/pHY-dltB, which was consistent with our unpublished results that strengthening PhoQ(R)∼PhoP system benefits γ-PGA synthesis. Finally, γ-PGA is an anionic polypeptide, and the decrease of cell surface negative charge might reduce the strength of electrostatic repulsion (Cafiso et al., 2014) between TAs and γ-PGA, which further benefited γ-PGA secretion. To test this hypothesis, the negatively charged products (such as lichenysin, bacitracin, pulcherrimin, et al.) were analyzed, and our results showed that the yields of those products with negative charge were all increased in dlt overexpression strains (Supplementary Figure S3).
Conclusion
Cell surface engineering is a promising tactic for enhanced production of metabolites. This study demonstrated that overexpression of dlt operon could increase γ-PGA production for the first time. Furthermore, 13C-MFA was applied to illustrate the affect of dltB overexpression on γ-PGA synthesis. Our results showed that overexpression of dltB could reduce the negative charge of cell surface, which further reduced the absorption and utilization rate of glucose and the flux of EMP pathway. Meanwhile, the increases of TCA cycle flux, NADPH and ATP supplies, glutamic acid formation benefited γ-PGA synthesis. This work demonstrated that 13C-MFA provided the distinct metabolic insights in engineered microbes for the changes in central carbon metabolism.
Author Contributions
PH and SC designed the study. PH, SH, and YC carried out the molecular biology studies and construction of recombinant strains. PH, DC, SH, and YC carried out the fermentation studies. PH and NW carried out the metabolic flux analysis. PH, DC, SL, and SC analyzed the data and wrote the manuscript. All authors read and approved the final manuscript.
Funding
This work was supported by the National Program on Key Basic Research Project (973 Program, No. 2015CB150505), the Technical Innovation Special Fund of Hubei Province (2018ACA149), and the China Postdoctoral Science Foundation (2018M642802).
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank to Lian He from University of Washington for his help on 13C-metabolic flow analysis.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2019.00105/full#supplementary-material
All the primers sequences for strain construction and RT-qPCR were listed in Supplementary Tables S1 and S2, respectively. The results of labeled amino acids were provided in Supplementary Table S3, and the specific data in metabolic pathway were provided in Supplementary Table S4.
Abbreviations
2,3-BDO, 2,3-butanediol; 3PG, 3-phosphoglyceric acid; 6PG, 6-phosphogluconolactone; AceCoA, acetyl-CoA; AKG, α-ketoglutarate; CIT, citrate; DHAP, dihydroxyacetone phosphate; E4P, erythrose 4-phosphate; EMP pathway, glycolysis pathway; F6P, fructose 6-phosphate; FBP, fructose 1,6-bisphosphate; FUM, fumarate; G6P, glucose-6-phosphate; GAP, glyceraldehyde 3-phosphate; Glu, glutamic acid; GLX, glyoxylate; ICIT, isocitrate; MAL, malate; OAA, oxaloacetate; PEP, phosphoenolpyruvate; PPP pathway, pentose phosphate pathway; PYR, pyruvate; R5P, ribose 5-phosphate; Ru5P, ribulose 5-phosphate; S7P, sedoheptulose 7-phosphate; SUC, succinate; SucCoA, succinyl-CoA; TCA cycle, tricarboxylic acid cycle; X5P, xylulose 5-phosphate.
References
Antoniewicz, M. R., Kelleher, J. K., and Stephanopoulos, G. (2007). Elementary metabolite units (EMU): a novel framework for modeling isotopic distributions. Metab. Eng. 9, 68–86. doi: 10.1016/j.ymben.2006.09.001
Cafiso, V., Bertuccio, T., Purrello, S., Campanile, F., Mammina, C., Sartor, A., et al. (2014). dltA overexpression: a strain-independent keystone of daptomycin resistance in methicillin-resistant Staphylococcus aureus. Int. J. Antimicrob. Agents 43, 26–31. doi: 10.1016/j.ijantimicag.2013.10.001
Cai, D., Chen, Y., He, P., Wang, S., Mo, F., Li, X., et al. (2018). Enhanced production of poly-gamma-glutamic acid by improving ATP supply in metabolically engineered Bacillus licheniformis. Biotechnol. Bioeng. 115, 2541–2553. doi: 10.1002/bit.26774
Cai, D., He, P., Lu, X., Zhu, C., Zhu, J., Zhan, Y., et al. (2017). A novel approach to improve poly-gamma-glutamic acid production by NADPH regeneration in Bacillus licheniformis WX-02. Sci. Rep. 7:43404. doi: 10.1038/srep43404
Cao, H., Van Heel, A. J., Ahmed, H., Mols, M., and Kuipers, O. P. (2017). Cell surface engineering of Bacillus subtilis improves production yields of heterologously expressed alpha-amylases. Microb. Cell Fact. 16:56. doi: 10.1186/s12934-017-0674-0
Cao, M., Feng, J., Sirisansaneeyakul, S., Song, C., and Chisti, Y. (2018). Genetic and metabolic engineering for microbial production of poly-gamma-glutamic acid. Biotechnol. Adv. 36, 1424–1433. doi: 10.1016/j.biotechadv.2018.05.006
Chen, Y., Cai, D., He, P., Mo, F., Zhang, Q., Ma, X., et al. (2018). Enhanced production of heterologous proteins by Bacillus licheniformis with defective D-alanylation of lipoteichoic acid. World J. Microb. Biotechnol. 34:135. doi: 10.1007/s11274-018-2520-x
Devine, K. M. (2018). Activation of the PhoPR-mediated response to phosphate limitation is regulated by wall teichoic acid metabolism in Bacillus subtilis. Front. Microbiol. 9:2678. doi: 10.3389/fmicb.2018.02678
Feng, J., Gu, Y., Quan, Y., Cao, M., Gao, W., Zhang, W., et al. (2015). Improved poly-gamma-glutamic acid production in Bacillus amyloliquefaciens by modular pathway engineering. Metab. Eng. 32, 106–115. doi: 10.1016/j.ymben.2015.09.011
Garcia Vescovi, E., Soncini, F. C., and Groisman, E. A. (1996). Mg2+ as an extracellular signal: environmental regulation of Salmonella virulence. Cell 84, 165–174. doi: 10.1016/S0092-8674(00)81003-X
Ghorbal, S. K., Chatti, A., Sethom, M. M., Maalej, L., Mihoub, M., Kefacha, S., et al. (2013). Changes in membrane fatty acid composition of Pseudomonas aeruginosa in response to UV-C radiations. Curr. Microbiol. 67, 112–117. doi: 10.1007/s00284-013-0342-5
Gonzalez, J. E., Long, C. P., and Antoniewicz, M. R. (2017). Comprehensive analysis of glucose and xylose metabolism in Escherichia coli under aerobic and anaerobic conditions by (13)C metabolic flux analysis. Metab. Eng. 39, 9–18. doi: 10.1016/j.ymben.2016.11.003
He, L., Xiao, Y., Gebreselassie, N., Zhang, F., Antoniewiez, M. R., Tang, Y. J., et al. (2014). Central metabolic responses to the overproduction of fatty acids in Escherichia coli based on 13C-metabolic flux analysis. Biotechnol. Bioeng. 111, 575–585. doi: 10.1002/bit.25124
He, L., Xiu, Y., Jones, J. A., Baidoo, E. E. K., Keasling, J. D., Tang, Y. J., et al. (2017). Deciphering flux adjustments of engineered E. coli cells during fermentation with changing growth conditions. Metab. Eng. 39, 247–256. doi: 10.1016/j.ymben.2016.12.008
Heit, B., Yeung, T., and Grinstein, S. (2011). Changes in mitochondrial surface charge mediate recruitment of signaling molecules during apoptosis. Am. J. Physiol. Cell Physiol. 300, C33–C41. doi: 10.1152/ajpcell.00139.2010
Hyyrylainen, H. L., Pietiainen, M., Lunden, T., Ekman, A., Gardemeister, M., Murtomaki-Repo, S., et al. (2007). The density of negative charge in the cell wall influences two-component signal transduction in Bacillus subtilis. Microbiology 153, 2126–2136. doi: 10.1099/mic.0.2007/008680-0
Hyyrylainen, H. L., Vitikainen, M., Thwaite, J., Wu, H., Sarvas, M., Harwood, C. R., et al. (2000). D-Alanine substitution of teichoic acids as a modulator of protein folding and stability at the cytoplasmic membrane/cell wall interface of Bacillus subtilis. J. Biol. Chem. 275, 26696–26703. doi: 10.1074/jbc.M003804200
Kind, S., Becker, J., and Wittmann, C. (2013). Increased lysine production by flux coupling of the tricarboxylic acid cycle and the lysine biosynthetic pathway–metabolic engineering of the availability of succinyl-CoA in Corynebacterium glutamicum. Metab. Eng. 15, 184–195. doi: 10.1016/j.ymben.2012.07.005
Koprivnjak, T., Mlakar, V., Swanson, L., Fournier, B., Peschel, A., and Weiss, J. P. (2006). Cation-induced transcriptional regulation of the dlt operon of Staphylococcus aureus. J. Bacteriol. 188, 3622–3630. doi: 10.1128/JB.188.10.3622-3630.2006
Lambert, P. A., Hancock, I. C., and Baddiley, J. (1975). Influence of alanyl ester residues on the binding of magnesium ions to teichoic acids. Biochem. J. 151, 671–676. doi: 10.1042/bj1510671
Lee, S. W., and Oh, M. K. (2016). Improved production of N-acetylglucosamine in Saccharomyces cerevisiae by reducing glycolytic flux. Biotechnol. Bioeng. 113, 2524–2528. doi: 10.1002/bit.26014
Long, C. P., Au, J., Gonzalez, J. E., and Antoniewicz, M. R. (2016). (13)C metabolic flux analysis of microbial and mammalian systems is enhanced with GC-MS measurements of glycogen and RNA labeling. Metab. Eng. 38, 65–72. doi: 10.1016/j.ymben.2016.06.007
Nanchen, A., Fuhrer, T., and Sauer, U. (2007). Determination of metabolic flux ratios from 13C-experiments and gas chromatography-mass spectrometry data: protocol and principles. Methods Mol. Biol. 358, 177–197. doi: 10.1007/978-1-59745-244-1_11
Neuhaus, F. C., and Baddiley, J. (2003). A continuum of anionic charge: structures and functions of D-alanyl-teichoic acids in gram-positive bacteria. Microbiol. Mol. Biol. Rev. 67, 686–723. doi: 10.1128/MMBR.67.4.686-723.2003
Ren, J., Sang, Y., Lu, J., and Yao, Y. F. (2017). Protein acetylation and its role in bacterial virulence. Trends Microbiol. 25, 768–779. doi: 10.1016/j.tim.2017.04.001
Sauer, U., Lasko, D. R., Fiaux, J., Hochuli, M., Glaser, R., Szyperski, T., et al. (1999). Metabolic flux ratio analysis of genetic and environmental modulations of Escherichia coli central carbon metabolism. J. Bacteriol. 181, 6679–6688.
Sirisansaneeyakul, S., Cao, M., Kongklom, N., Chuensangjun, C., Shi, Z., and Chisti, Y. (2017). Microbial production of poly-gamma-glutamic acid. World J. Microbiol. Biotechnol. 33:173. doi: 10.1007/s11274-017-2338-y
Steen, A., Palumbo, E., Deghorain, M., Cocconcelli, P. S., Delcour, J., Kuipers, O. P., et al. (2005). Autolysis of Lactococcus lactis is increased upon D-alanine depletion of peptidoglycan and lipoteichoic acids. J. Bacteriol. 187, 114–124. doi: 10.1128/JB.187.1.114-124.2005
Tan, Z., Khakbaz, P., Chen, Y., Lombardo, J., Yoon, J. M., Shanks, J. V., et al. (2017). Engineering Escherichia coli membrane phospholipid head distribution improves tolerance and production of biorenewables. Metab. Eng. 44, 1–12. doi: 10.1016/j.ymben.2017.08.006
Tian, G., Wang, Q., Wei, X., Ma, X., and Chen, S. (2017). Glutamate dehydrogenase (RocG) in Bacillus licheniformis WX-02: enzymatic properties and specific functions in glutamic acid synthesis for poly-gamma-glutamic acid production. Enzyme Microb. Technol. 99, 9–15. doi: 10.1016/j.enzmictec.2017.01.002
Tran, L. S., Nagai, T., and Itoh, Y. (2000). Divergent structure of the ComQXPA quorum-sensing components: molecular basis of strain-specific communication mechanism in Bacillus subtilis. Mol. Microbiol. 37, 1159–1171. doi: 10.1046/j.1365-2958.2000.02069.x
Wan, N., Delorenzo, D. M., He, L., You, L., Immethun, C. M., Wang, G., et al. (2017). Cyanobacterial carbon metabolism: fluxome plasticity and oxygen dependence. Biotechnol. Bioeng. 114, 1593–1602. doi: 10.1002/bit.26287
Wahl, S. A., Dauner, M., and Wiechert, W. (2004). New tools for mass isotopomer data evaluation in 13C flux analysis: mass isotope correction, data consistency checking, and precursor relationships. Biotechnol. Bioeng. 85, 259–268. doi: 10.1002/bit.10909
Westbrook, A. W., Ren, X., Moo-Young, M., and Chou, C. P. (2018). Engineering of cell membrane to enhance heterologous production of hyaluronic acid in Bacillus subtilis. Biotechnol. Bioeng. 115, 216–231. doi: 10.1002/bit.26459
Xie, X., Liang, Y., Liu, H., Liu, Y., Xu, Q., Zhang, C., et al. (2014). Modification of glycolysis and its effect on the production of L-threonine in Escherichia coli. J. Ind. Microbiol. Biotechnol. 41, 1007–1015. doi: 10.1007/s10295-014-1436-1
Yao, R., Xiong, D., Hu, H., Wakayama, M., Yu, W., Zhang, X., et al. (2016). Elucidation of the co-metabolism of glycerol and glucose in Escherichia coli by genetic engineering, transcription profiling, and (13)C metabolic flux analysis. Biotechnol. Biofuels 9:175. doi: 10.1186/s13068-016-0591-1
You, L., Page, L., Feng, X., Berla, B., Pakrasi, H. B., and Tang, Y. J. (2012). Metabolic pathway confirmation and discovery through (13)C-labeling of proteinogenic amino acids. J. Vis. Exp. 59:e3583. doi: 10.3791/3583
Young, J. D. (2014). INCA: a computational platform for isotopically non-stationary metabolic flux analysis. Bioinformatics 30, 1333–1335. doi: 10.1093/bioinformatics/btu015
Keywords: Bacillus licheniformis, poly-γ-glutamic acid, 13C-metabolic flux analysis, dltB, cell surface negative charge
Citation: He P, Wan N, Cai D, Hu S, Chen Y, Li S and Chen S (2019) 13C-Metabolic Flux Analysis Reveals the Metabolic Flux Redistribution for Enhanced Production of Poly-γ-Glutamic Acid in dlt Over-Expressed Bacillus licheniformis. Front. Microbiol. 10:105. doi: 10.3389/fmicb.2019.00105
Received: 30 October 2018; Accepted: 17 January 2019;
Published: 01 February 2019.
Edited by:
Wei Xiong, National Renewable Energy Laboratory (DOE), United StatesReviewed by:
Deng Liu, Washington University in St. Louis, United StatesJoshua Chan, Colorado State University, United States
Copyright © 2019 He, Wan, Cai, Hu, Chen, Li and Chen. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Shouwen Chen, bWVsMjEyQDEyNi5jb20=
 Penghui He
Penghui He Ni Wan
Ni Wan Dongbo Cai1
Dongbo Cai1 Shouwen Chen
Shouwen Chen