- 1School of Laboratory Medicine and Life Science/Institute of Biomedical Informatics, Wenzhou Medical University, Wenzhou, China
- 2Tongji University School of Medicine, Shanghai, China
- 3Institute of Translational Medicine, Baotou Central Hospital, Baotou, China
- 4School of Medical and Health, Lishui University, Lishui, China
The diversity of class D β-lactamases mediating resistance to β-lactams has been increasingly reported recently. In this study, a novel class D oxacillinase named OXA-830 was identified in a fully sequenced Aeromonas simiae strain, which was isolated from sewage discharged from a farm in southern China. OXA-830 shares the highest amino acid identity of 79.3% with an OXA-12-like variant named OXA-725. When expressed in E. coli DH5α, OXA-830 conferred resistance to penicillins and selected β-lactamase inhibitors but not to cephalosporins and carbapenems. Kinetic analysis of OXA-830 revealed a broad substrate profile including penicillins, cefazolin, cefoxitin, and ceftazidime but not carbapenems. The hydrolytic activity was significantly inhibited by sulbactam, followed by tazobactam, but was less effectively inhibited by clavulanic acid. The blaOXA–830 gene was located on the A. simiae A6 chromosome and the blaOXA–830-related region was bracketed by a pair of perfect inverted repeats.
Introduction
The genus Aeromonas is a distinct group of oxidase-positive, facultatively anaerobic, Gram-negative bacilli of the family Aeromonadaceae (Colwell et al., 1986). Members of Aeromonas can be isolated from every environmental niche where bacterial ecosystems exist, including aquatic habitats and fish as well as food products, and are implicated in human and animal infections (Janda and Abbott, 2010). The species Aeromonas simiae was first described in two strains (CIP 107798 and CIP 107797) isolated from feces of healthy monkeys from Mauritius (Harf-Monteil et al., 2004). Further study revealed that these two strains may originate from the same clone because they share entirely identical 16S rRNA, gyrB, and rpoD genes with each other (Saavedra et al., 2006). To date, there have been a large number of reports about the presence of β-lactamase genes among Aeromonas strains (Carvalho et al., 2012; Chen et al., 2012).
Oxacillin-hydrolyzing (OXA)-type β-lactamases (OXAs) constitute most of the members of Ambler class D active-serine-site β-lactamases (Bush et al., 1995) and are widely identified among clinically relevant Gram-negative bacteria, such as Pseudomonas spp., Acinetobacter spp., Aeromonas spp. and Enterobacteriaceae (Poirel et al., 2010). Most of the OXAs possess the ability to hydrolyze cloxacillin or oxacillin at a rate of >50% that for benzylpenicillin and are typically not inhibited by commercially available β-lactamase inhibitors such as clavulanic acid, tazobactam, and sulbactam (Payne et al., 1994; Bush et al., 1995). According to the Bush functional classification scheme for β-lactamases (Bush et al., 1995), OXAs are classified into group 2d. Although most of the early identified OXAs exhibited a substrate profile strictly restricted to penicillin, oxacillin, cloxacillin and nitrocefin, several OXA members have been demonstrated to be active against extended-spectrum cephalosporins which is typically due to a small number of point mutations occurring in parental narrow-spectrum class D β-lactamases (DBLs), such as the derivatives of OXA-10 (Poirel et al., 2010; Leonard et al., 2013). For instance, compared to OXA-10, OXA-11 (N146S and G167D) exhibits the ability to hydrolyze ceftazidime (Hall et al., 1993); OXA-17 (N76S) has an increased hydrolytic ability for cefotaxime as well as a decreased capacity for ceftazidime (Danel et al., 1999); and OXA-19, which differed from OXA-10 by nine amino acids, can hydrolyze ceftazidime with a low activity (Mugnier et al., 1998). Some OXAs have evolved to exhibit hydrolytic activity toward β-lactams of “last resort,” i.e., carbapenems (El Garch et al., 2011; Antonelli et al., 2015).
Similar to other antimicrobial resistance genes, many OXA β-lactamase genes have been identified on both plasmids and chromosomes with diverse mobile genetic elements (MGEs), such as integrons, insertion sequences and transposons (Poirel et al., 2010). For example, the blaOXA–1-like, blaOXA–2-like, and blaOXA–10-like genes were commonly captured as gene cassettes by integrons in plasmids (Naas and Nordmann, 1999); the blaOXA–23 gene from the chromosome of Acinetobacter radioresistens may be transferred onto plasmids diffusing into Acinetobacter baumannii through the ISAba1-based composite transposon Tn2006 or transposon-like structure named Tn2008, or a single copy of ISAba4 upstream of the gene (Corvec et al., 2007; Adams-Haduch et al., 2008); and the blaOXA–58 gene-encoding regions in A. baumannii isolated from different countries (France, Spain, Romania, and Turkey) were bracketed by ISAba3 on one side and ISAba3, ISAba1 or IS18 on the other side (Poirel and Nordmann, 2006). However, according to the literature, there are intrinsic chromosomally encoded OXAs in many bacterial species (Poirel et al., 2010). The first such gene, identified in 1994, originated from the chromosome of Aeromonas jandaei (formerly Aeromonas sobria), which was not associated with an integron or transposon and was named blaOXA–12 (Rasmussen et al., 1994).
In this study, for the first time, we determined the complete genome sequence of A. simiae, i.e., a sewage-derived A. simiae strain A6. Based on sequence analysis, we identified and characterized a novel chromosomally encoded DBL named OXA-830, which is quite divergent from the other OXA β-lactamases.
Materials and Methods
Bacterial Strains
The host strain A. simiae A6 carrying the blaOXA–830 gene was isolated in November 2017 from sewage discharged from a farm in Wenzhou, China. Species identification was conducted using the Vitek-60 microorganism autoanalysis system (BioMérieux Corporate, Craponne, France). Further verification was performed based on the 16S rRNA sequencing method. Moreover, considering limited resolution the 16S rRNA gene provides to discriminate among closely related species of the genus Aeromonas (Alperi et al., 2008), a multilocus phylogenetic analysis (MLPA) of the concatenated sequences of 6 housekeeping genes (gyrB, rpoD, recA, dnaJ, gyrA, and atpD) as previously reported (Beaz-Hidalgo et al., 2015) as well as whole-genome sequence-based phylogenetic analysis using kSNP3.0 (Gardner et al., 2015) were conducted to determine the evolutionary relationship of A. simiae A6 with 32 other Aeromonas sp. strains of different species. Two neighbor-joining phylogenetic trees were generated by using MEGA7 with 1,000 bootstrap replications (Kumar et al., 2016). The bacteria and plasmids used in this work are listed in Supplementary Table S1.
Antimicrobial Susceptibility Testing
The minimum inhibitory concentrations (MICs) were determined using the agar dilution method following the guidelines of the Clinical and Laboratory Standards Institute (CLSI). Susceptibility patterns were interpreted according to the CLSI breakpoint criteria (CLSI, 2019) and the guidelines of the European Committee on Antimicrobial Susceptibility Testing [EUCAST] (2019) for Enterobacteriaceae. No interpretation criteria were available for benzylpenicillin, benzylpenicillin/clavulanic acid, benzylpenicillin/sulbactam, oxacillin, cloxacillin and polymyxin B from CLSI (2019) and European Committee on Antimicrobial Susceptibility Testing [EUCAST] (2019). E. coli ATCC 25922 was used as a reference strain for quality control.
Whole-Genome Sequencing and Sequence Analysis
Whole-cell DNA of A.simiae A6 was extracted using the AxyPrep Bacterial Genomic DNA Miniprep kit (Axygen Scientific, Union City, CA, United States) and sequenced on a PacBio RS II instrument (Pacific Biosciences). The PacBio long reads were initially assembled by Canu v1.6 (Koren et al., 2017), and then two FASTQ sequence files generated using the Illumina HiSeq 2500 platform were mapped onto the primary assembly to control assembly quality and to correct possible misidentified bases by using Bwa and the Genome Analysis Toolkit (McKenna et al., 2010). The consensus sequence was obtained by a custom-derived script written in Python1. Potential open reading frames (ORFs) were predicted using Glimmer software (Delcher et al., 2007) and annotated against the UniProt/Swiss-Prot and non-redundant protein databases using the BLASTX program with an e-value threshold of 1e-5. GView was used to construct basic genomic features (Petkau et al., 2010). Annotation of MGEs and resistance genes was performed using ISfinder (Siguier et al., 2006), INTEGRALL (Moura et al., 2009) and ResFinder (Zankari et al., 2012) with default parameters. The molecular weight and pI value of OXA-830 was predicted using ProtParam2. The putative signal peptide cleavage site of OXA-830 was identified by SignalP 5.0 (Almagro Armenteros et al., 2019). Amino acid alignment and the neighbor-joining phylogenetic tree construction of OXA-830 with other DBLs were performed using the MAFFT program and MEGA7 with a bootstrap of 1,000 replicates, respectively (Katoh and Standley, 2013; Kumar et al., 2016). Comparisons of the nucleotide sequences were performed using BLASTN. Other bioinformatics tools were written using Python (see text footnote 1) and Biopython (Cock et al., 2009).
Cloning of the blaOXA–830 Gene and Expression and Purification of OXA-830
The gene encoding OXA-830 along with its promoter was amplified by PCR using forward (5′-CGGAATTCAGACACAGATTGGCACAGCA-3′) and reverse (5′-CCCAAGCTTGCCCGGTGAAGAAGAAGTGA-3′) primers with a pair of flanking restriction endonuclease adapters. Then, the PCR product was eluted from an agarose gel, digested with the EcoRI and HindIII, and ligated into the pUCP24 vector with a T4 DNA ligase cloning kit (Takara Bio, Inc., Dalian, China). The recombinant plasmids were transformed into competent E. coli DH5α by the calcium chloride method, and bacterial colonies were cultured on Luria-Bertani agar plates supplemented with 20 μg/mL gentamicin. The recombinant plasmids were verified by both restriction enzyme digestion and Sanger sequencing (Shanghai Sunny Biotechnology Co., Ltd., Shanghai, China). The similar procedure was also applied to cloned complete ORF of blaOXA–830 into pET-28b. The recombinant plasmid (pET-OXA-830) was transformed into competent E. coli BL21 cells by the calcium chloride method, and transformants were selected on Luria-Bertani agar plates supplemented with 50 μg/mL kanamycin. The authenticity of cloned fragments was confirmed by Sanger sequencing (Shanghai Sunny Biotechnology Co., Ltd., Shanghai, China). The above-mentioned E. coli BL21 transformant was grown in Luria-Bertani medium with 50 μg/mL kanamycin at 37°C. Overnight cultures were diluted 100-fold in 200 mL of Luria-Bertani medium and incubated for hours at 37°C with orbital shaking. Isopropyl-β-d-thiogalactopyranoside (IPTG) (Sigma Chemicals Co., St. Louis, MO, United States) was added to a final concentration of 1 mM until the cultures reached an OD600 between 0.6 and 0.8, and incubation was continued for an additional 4 h. OXA-830 was isolated from the periplasm and purified by affinity chromatography based on the instructions of the His-tag Protein Purification Kit (P2226, Beyotime, China).
Determination of Kinetic Parameters
Kinetic parameters for hydrolysis of β-lactams by the purified OXA-830 β-lactamase were examined using a UV-VIS spectrophotometer (U-3900, HITACHI, Japan) at 30°C in 10 mM phosphate buffer (pH 7.0) in a final reaction volume of 300 μL. The steady-state kinetic parameters (kcat and KM) were determined by non-linear regression of the initial reaction rates with the Michaelis–Menten equation in Prism (version 7) software (GraphPad Software, San Jose, CA, United States).
β-lactamase inhibition was studied with benzylpenicillin (500 μM) as the substrate. The β-lactamase inhibitors sulbactam, tazobactam and clavulanic acid at various concentrations were preincubated with the purified OXA-830 β-lactamase for 3 min at 30°C before addition of substrate. The inhibitor concentration required to reduce the hydrolysis of 500 μM benzylpenicillin by 50% was determined by non-linear regression with the log(inhibitor) vs. response – Variable slope equation in Prism (version 7) software (GraphPad Software, San Jose, CA, United States).
Nucleotide Sequence Accession Number
The complete nucleotide sequences of the chromosome of A. simiae A6 and the blaOXA–830 gene in this work have been submitted to DDBJ/EMBL/GenBank under accession numbers CP040449 and MK926981, respectively.
Results and Discussion
Identification and Characterization of the OXA-830-Producing Isolate, A. simiae A6
Aeromonas simiae A6 was isolated in 2017 from sewage discharged from a farm in Wenzhou, southern China. A 16S ribosomal RNA gene homology analysis showed that A. simiae A6 had the closest relationship with one A. simiae strain (A. simiae IBS S6874 [NR_025585.1]), at 99% identity and 99% coverage. The MLPA tree and whole-genome-based phylogeny tree congruously showed that A. simiae A6 was phylogenetically closest to A. simiae CIP 107798, which constituted a robust phylogenetic branch with considerable reliability (Figures 1A,B). We finally grouped the strain into the species A. simiae and named it A. simiae A6.
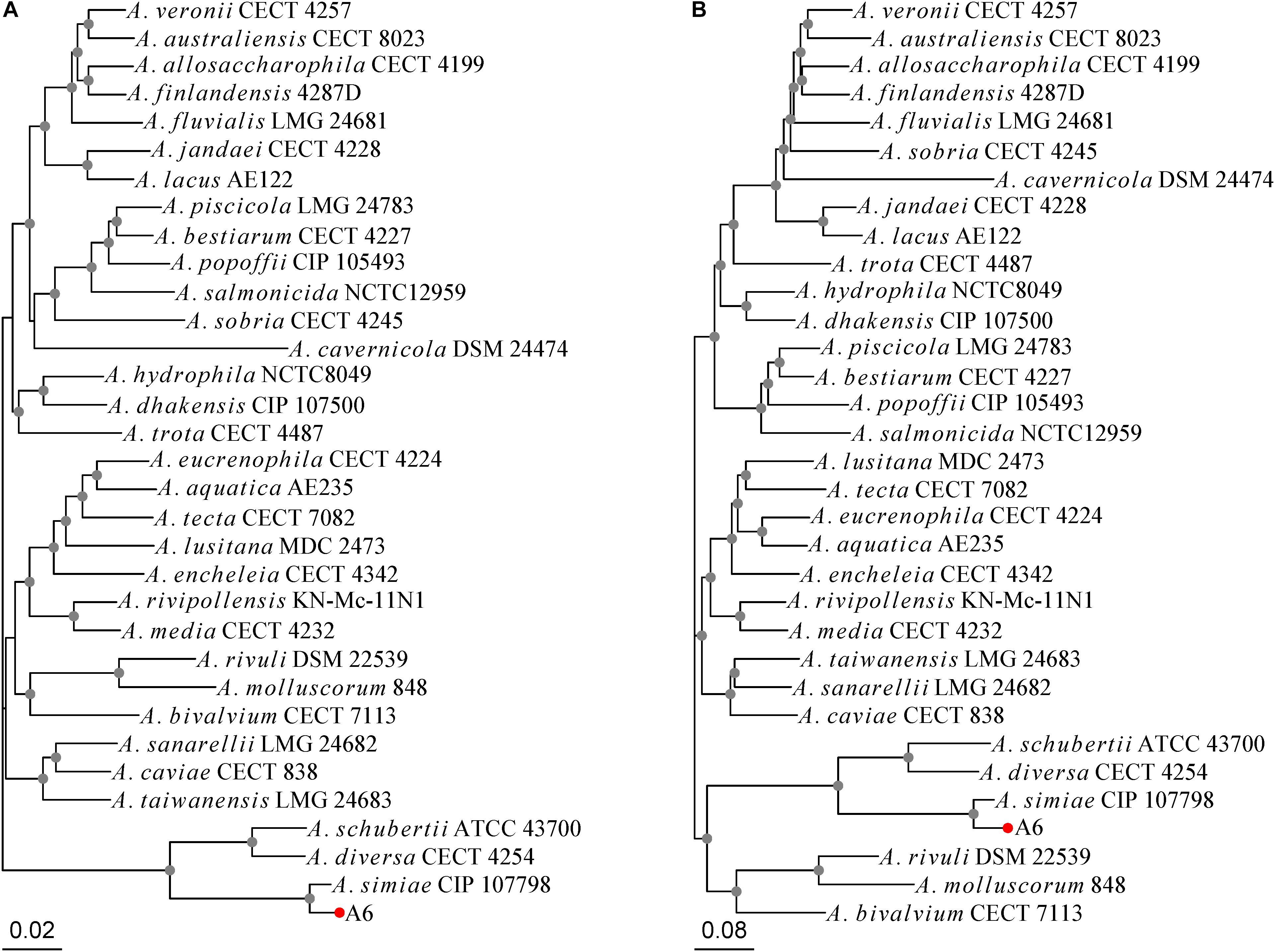
Figure 1. Phylogenetic relationships of A. simiae A6 with all other Aeromonas species inferred on the (A) MLPA tree and (B) whole-genome-based phylogeny tree. Nodes supported with bootstrap values over 50% are marked by gray dots. A. simiae A6 is indicated with a red dot.
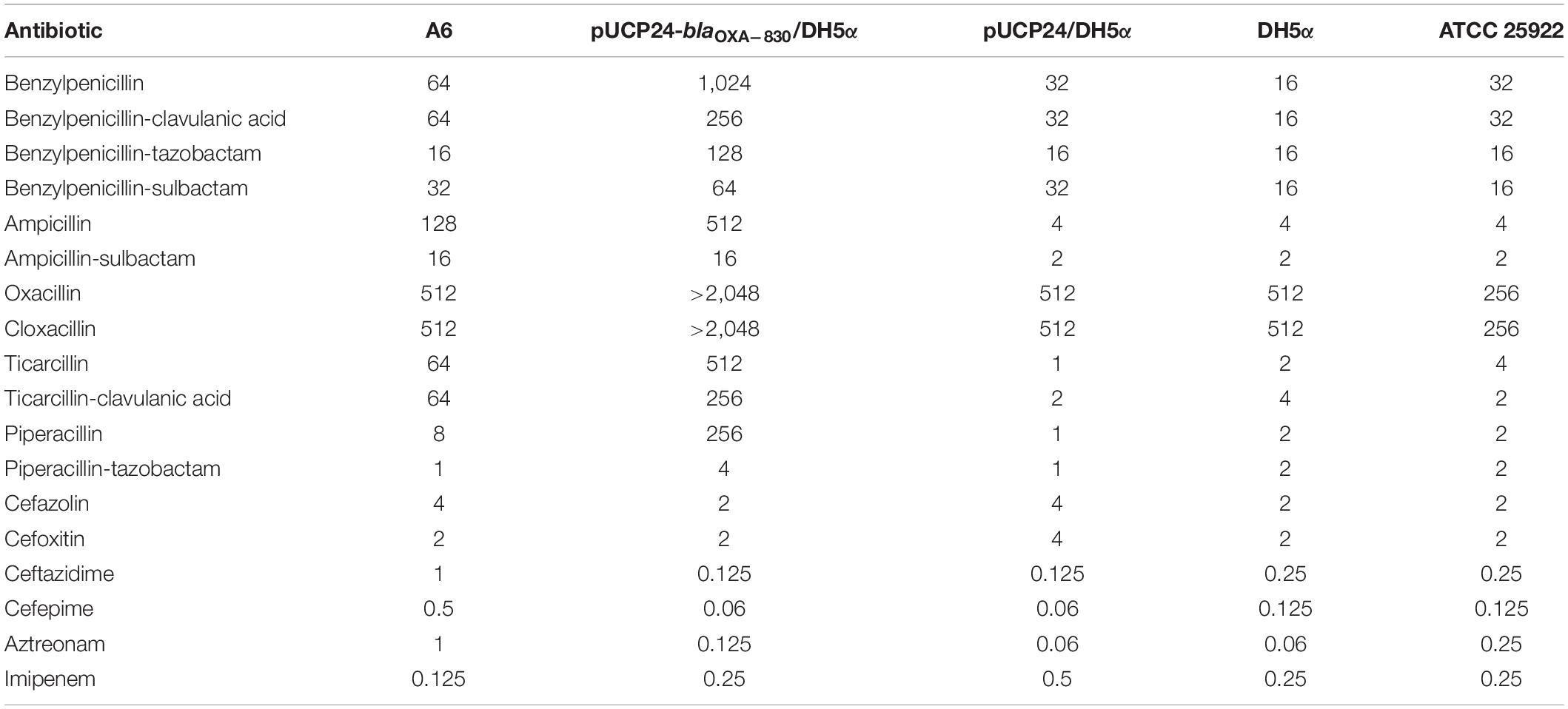
In vitro susceptibility testing showed that wild-type A. simiae A6 exhibited resistance to ampicillin and ticarcillin (Table 1). Moreover, the MICs of oxacillin, cloxacillin, benzylpenicillin, benzylpenicillin/clavulanic acid and benzylpenicillin/sulbactam against A. simiae A6 were 512, 512, 64, 64, and 32 μg/mL, respectively, which were significantly higher than or equal to the resistance breakpoint for ampicillin (≥32 μg/mL), though no interpretation criteria for these antimicrobials were available.
Since the high MICs of the aforementioned antimicrobials and no complete genome sequence of species A. simiae is currently available in the public database, the complete genome of A. simiae A6 was determined to unveil the potential factors associated with the resistance profiles. The results showed that A. simiae A6 has a circular chromosome (without plasmids) of 3.97 Mb in size that contains 3,633 ORFs with an average GC content of 60.56% (Table 2 and Figure 2). Comparative genomics analysis revealed the genome of A. simiae A6 shared the highest sequence similarities with that of A. schubertii WL1483 (accession number CP013067, at 91.62% identity and 65% coverage) and A. schubertii LF1708 (accession number CP039611, at 91.56% identity and 65% coverage) (Figure 2). Notably, A. simiae A6 encodes two predicted DBL-encoding genes. One is blaOXA–10 located in a truncated class I integron with a gene array of intI1-qnrVC4-cmlA5-blaOXA–10-aac(6’)-Ib-cr-aadA1-dfrA14-mobC-IS6100 (ranging from 3261 to 3269 kb), and the other belongs to a novel OXA β-lactamase named blaOXA–830, sharing the top amino acid identity (79.3%) with a function-known β-lactamase gene blaOXA–12 variant called blaOXA–725 (X80276.1) in the database.
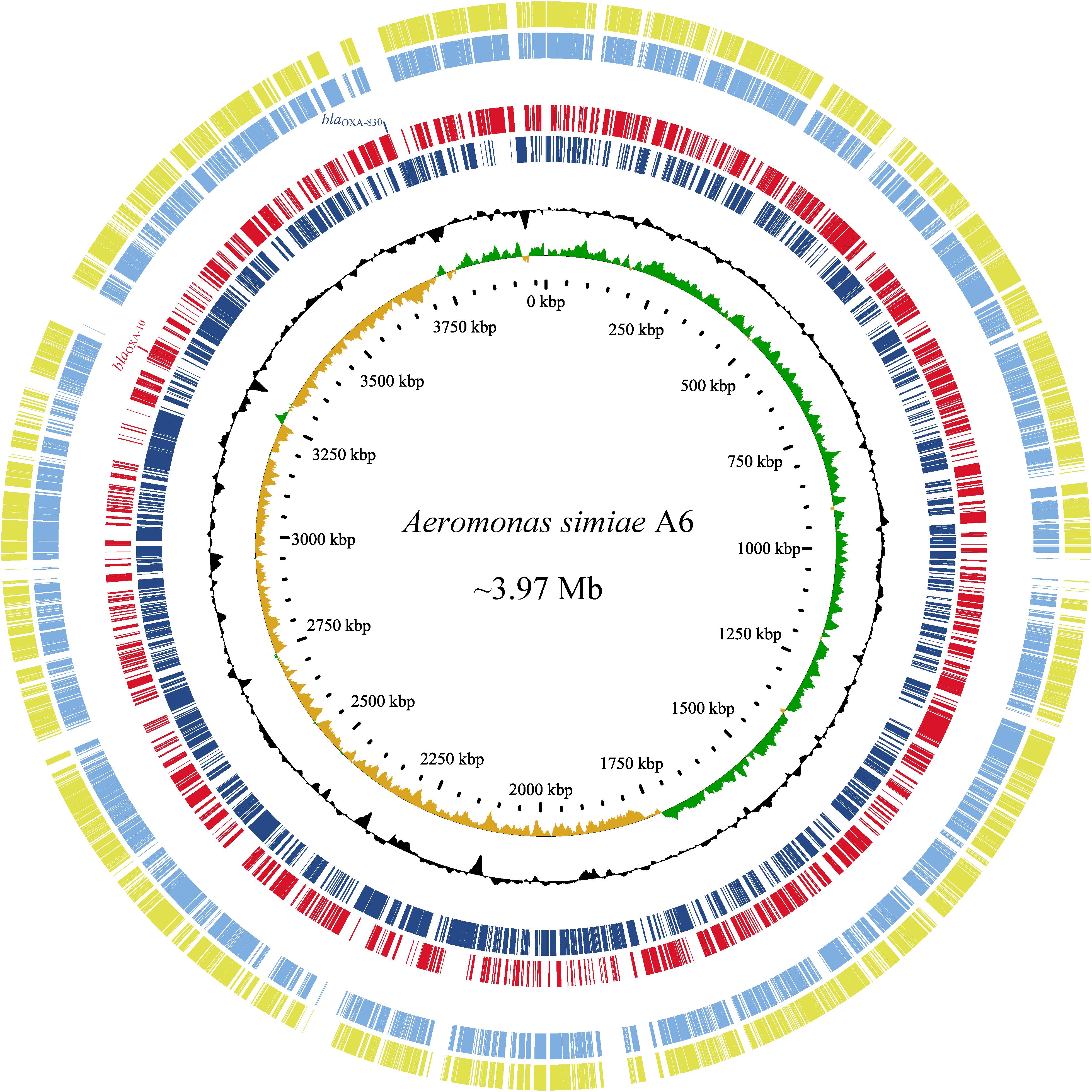
Figure 2. Genomic comparison of A. simiae A6 with its close relatives. From outside to inside: circles 1 and 2 are homologous regions of A. schubertii WL1483 (accession number CP013067) and A. schubertii LF1708 (CP039611) compared to A. simiae A6, while unmatched regions are left blank; circles 3 and 4 display predicted ORFs encoded in the forward strand and reverse strand, respectively; circles 5 and 6 represent the GC content and GC skew maps, respectively; and circle 7 shows the scale in kb.
Possible Origin of the New Class D β-Lactamase OXA-830
blaOXA–830 gene was 801 bp in length and encoded a 266 amino acid preprotein of ca. 29.4 kDa. A signal peptide cleavage site was predicted to be between an alanine and asparagine at amino acid residues 22 and 23. In addition, the pI value of OXA-830 was predicted to be 9.18. The closest relative of OXA-830 was OXA-725, a class D β-lactamase with detectable hydrolytic activity against oxacillin, penicillin and ampicillin as well as carbenicillin, which was previously described in a clinical A. jandaei (formerly A. sobria) isolate from the Hammersmith Hospital, London (Walsh et al., 1995). Screening for blaOXA–830-homologous genes (>70% amino acid sequence identity) in the NCBI nucleotide database showed that most of the close relatives (89.19%, 66/74) were derived from the genus Aeromonas, except 4 genes from Enterobacteriaceae and 4 of undetermined origin, suggesting the importance of Aeromonas as the reservoir for blaOXA–830-like genes. Additionally, OXA-830 was distantly related to other class D β-lactamases in amino acid sequence identity, i.e., OXA-12 and its close variants (77.1% with OXA-12; 78.9% with OXA-726; and 78.6% with OXA-724). Moreover, it shared identities of respective 72.2, 71.1, and 70.7% with OXA-427, OXA-780, and OXA-504 (Figure 3), and possessed less than 50% identities to all of the other DBLs.
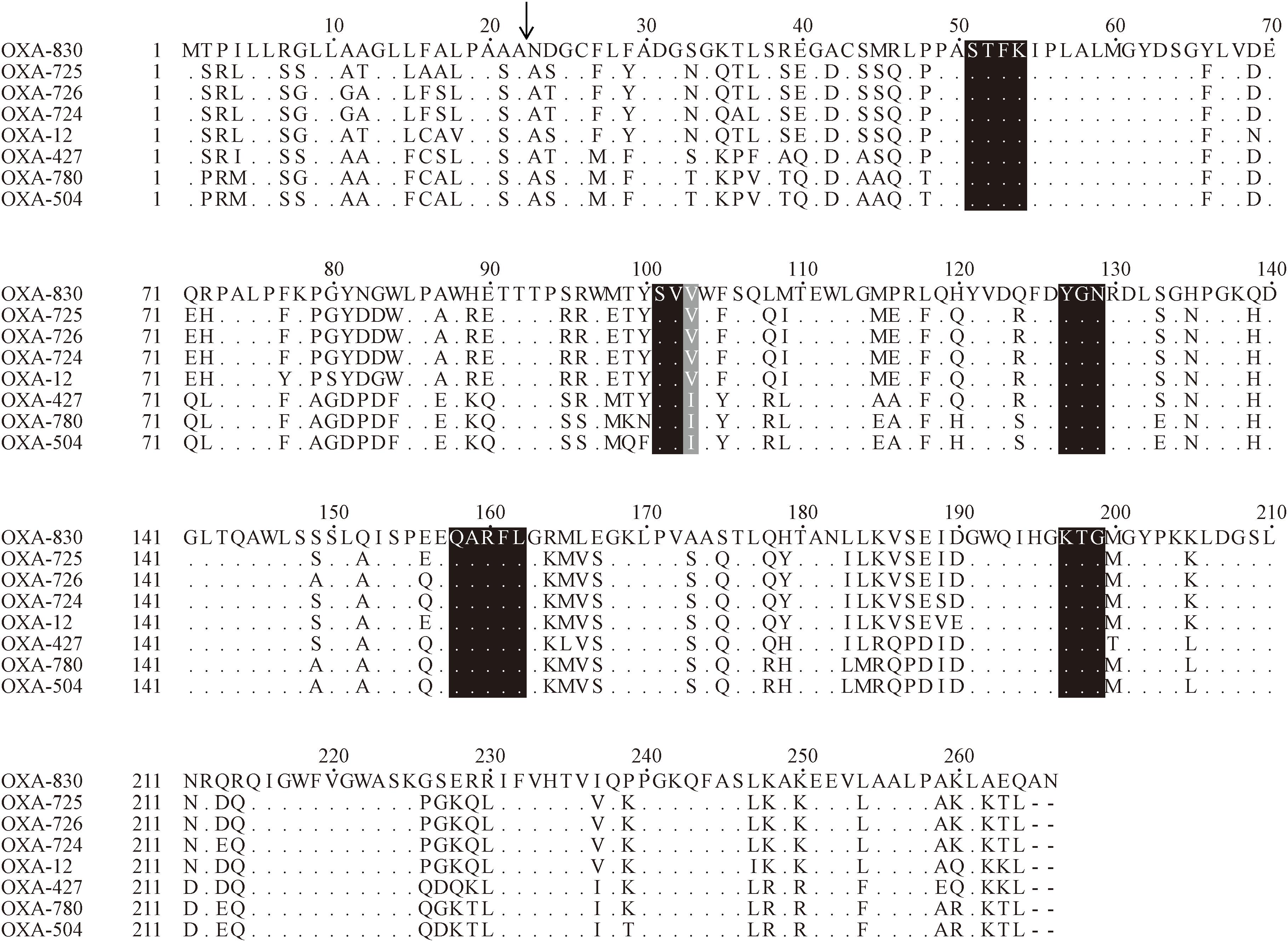
Figure 3. Amino acid alignment of OXA-830 with other selected class D β-lactamases. Dots indicate amino acid residues identical to those of OXA-830. Conserved motifs are shaded in black, while the poorly conserved site at residue 120 (DBL numbering) is shaded in gray. The predicted signal sequence cleavage site for OXA-830 is indicated by a vertical arrow.
Four conserved motifs were identified from OXA-830 and other OXA family DBLs: serine-threonine-phenylalanine-lysine (S-T-F-K) at positions 70 to 73, tyrosine-glycine-asparagine (Y-G-N) at positions 144 to 146, Q-X-X-X-L at positions 176 to 180, and lysine-threonine-glycine (K-T-G) at positions 216 to 218 (DBL numbering). However, the conserved S-X-V motif at residues 118 to 120 (DBL numbering), one of the active sites of DBLs (Paetzel et al., 2000), was found to be slightly different in OXA-427, OXA-780, and OXA-504, whose valine at residue 120 was replaced by isoleucine. In addition, OXA-830 possessed as many as 40 unique amino acid differences compared to all the other DBLs (Figure 3). A phylogenetic tree containing OXA-830 and other DBLs (>41% amino acid identity) (Figure 4) was constructed and the results showed that OXA-830 clustered closest to OXA-12, OXA-725, OXA-726, and OXA724. Thus, OXA-830 represents a novel lineage of DBLs.
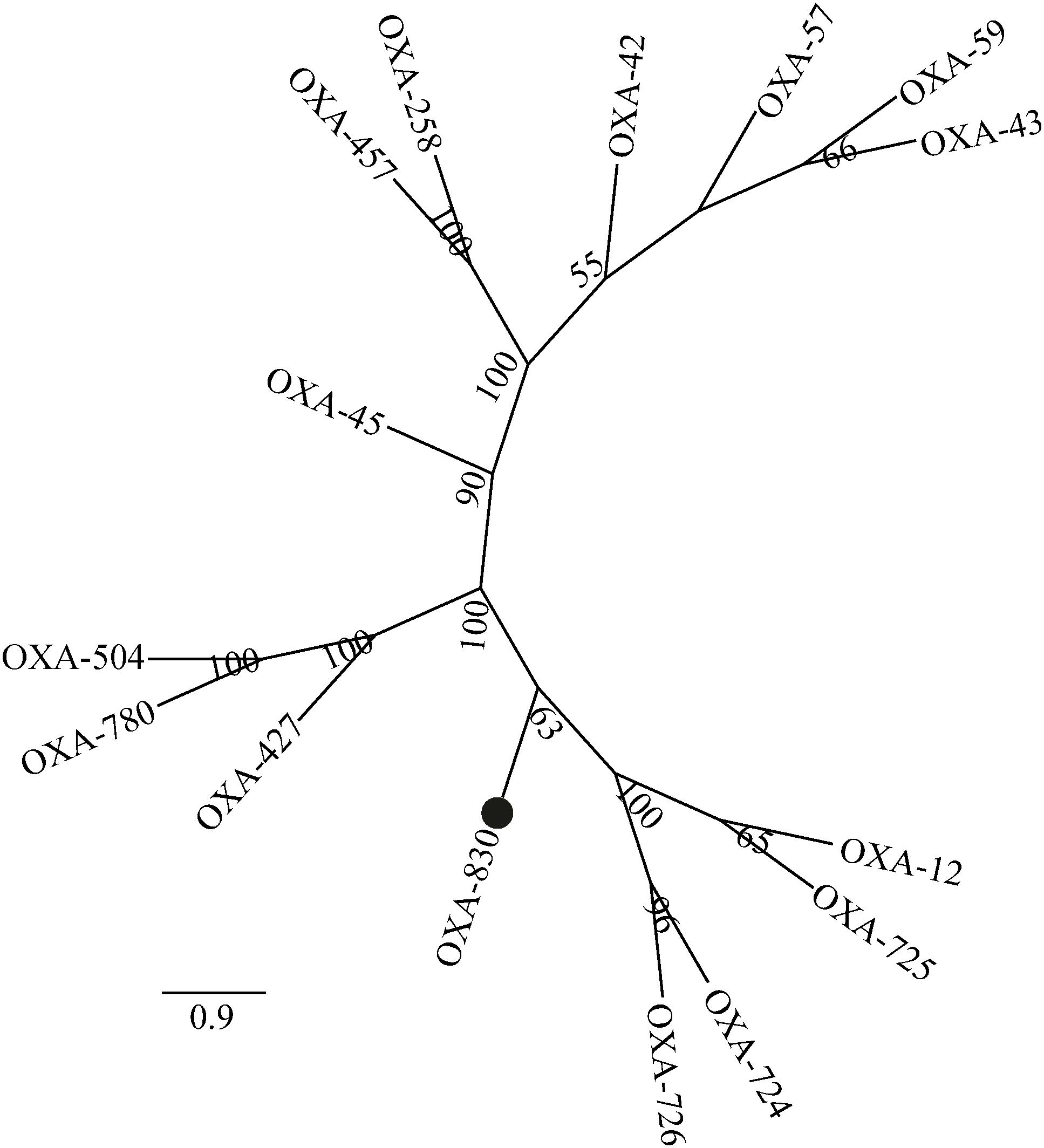
Figure 4. A phylogenetic tree showing the relationship of OXA-830 with other homologous DBLs. Bootstrap values are shown at branch points. OXA-830 from our study is indicated with a filled circle.
Genetic Context of blaOXA–830 Gene
A comparative genomics analysis of the genetic environment of the blaOXA–830 gene in A6 with that of closely related DBL-encoding genes in three other Aeromonas strains showed that glmU (encoding the bifunctional protein GlmU) together with upstream sequences shared a conserved syntenic architecture in gene content and gene order (Figure 5). However, the genes downstream of glmU in the blaOXA–830-related region were highly distant to those of all the other blaOXA genes. In fact, the region from blaOXA–830 to a large hypothetical gene in the blaOXA–830-related region was flanked by a pair of perfect 9-bp inverted repeats (IRs), suggesting these genes might be transferred from the other strain through horizontal gene transfer. Furthermore, an approximately 25-kb genomic island including two phage integrase-encoding genes was found upstream of the large hypothetical gene, which was enwrapped by a pair of perfect 10-bp direct repeats (DRs). This indicated that the mobilization of this genomic island onto the genome of A6 might have resulted from the integrase-mediated site-specific recombination. Altogether, these potential lateral gene transfer events may well explain the discrepancy of gene content between the blaOXA–830-related region and other blaOXA-related regions.
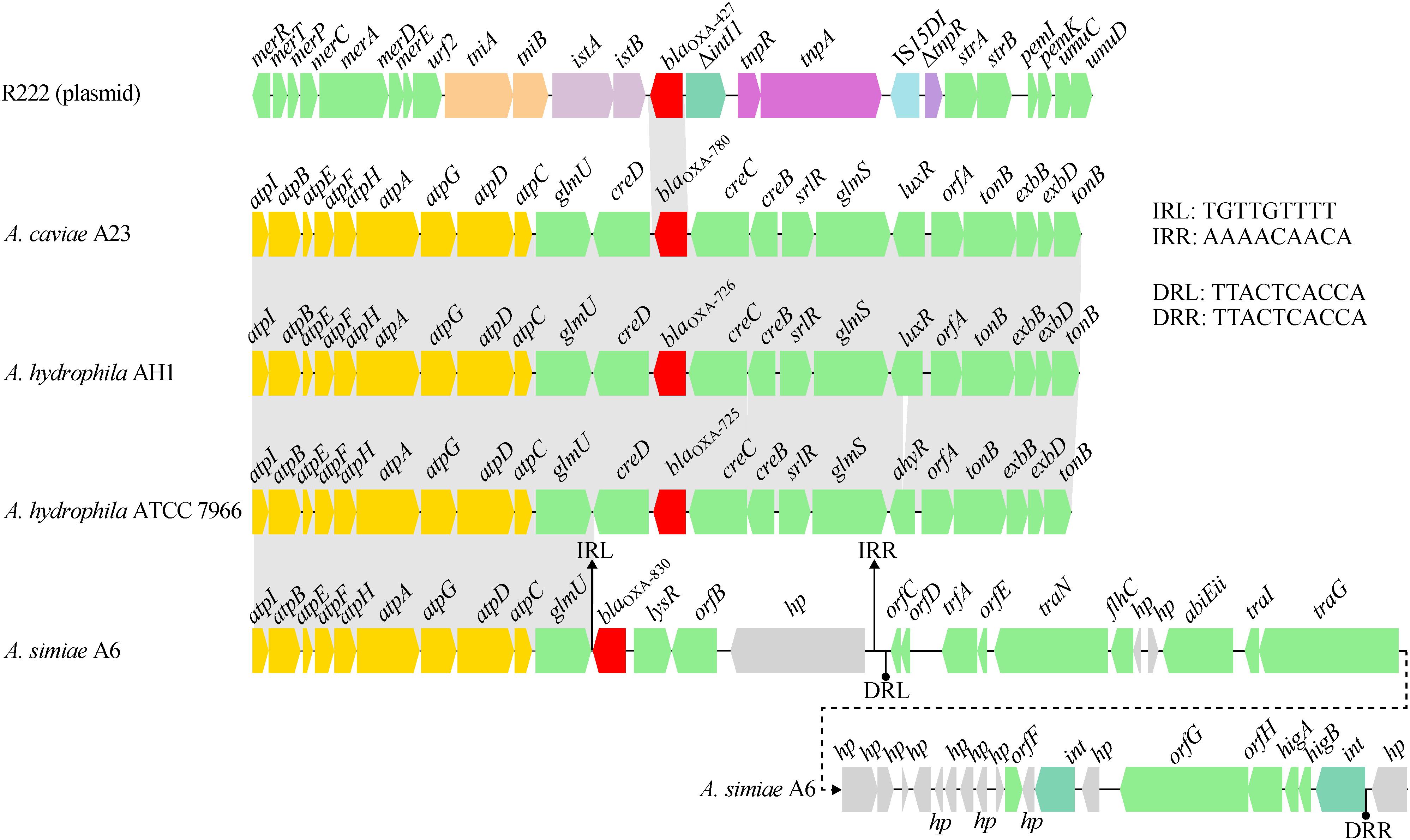
Figure 5. Comparison of the genetic context of the blaOXA–830 gene with that of selected β-lactamase genes. Genes are shown as arrows and colored based on gene function classification. Shading denotes regions of homology (>85% nucleotide sequence identity). Genes without direct gene names are illustrated as orfA, chromosome segregation ATPase; orfB, NADH:flavin oxidoreductase/NADH oxidase; orfC, AlpA family phage regulatory protein; orfD and orfE, DNA-binding proteins; orfF, Permease; orfG, DEAD/DEAH box helicase; and orfH, virulence associated protein. Regions are drawn to scale from accession numbers CP000462.1 (A. hydrophila ATCC 7966 chromosme), NZ_LSZC01000019.1 (A. hydrophila AH1 chromosome), LFXO01000139.1 (A. caviae A23 chromosome), and KX869741.1 (plasmid R222 from E. cloacae 20130723).
Functional Characterization of the OXA-830 β-Lactamase
The cloned blaOXA–830 gene in E. coli DH5α recombinant clones (pUCP24-blaOXA–830/DH5α) conferred resistance to all the penicillins and β-lactamase inhibitors examined in this study except for benzylpenicillin/sulbactam (Table 1). More specifically, it facilitated different increases in MIC levels for ampicillin (128-fold), ampicillin/sulbactam (8-fold), oxacillin (>4-fold), cloxacillin (>4-fold), benzylpenicillin (32-fold), benzylpenicillin/tazobactam (8-fold), benzylpenicillin/clavulanic acid (8-fold), ticarcillin (512-fold), ticarcillin/clavulanic acid (128-fold), piperacillin (256-fold) and piperacillin/tazobactam (4-fold) compared with those of the control strain (pUCP24/DH5α), although the MICs against ampicillin, benzylpenicillin, ticarcillin, and piperacillin were decreased in the presence of inhibitors. In addition, OXA-830 did not confer resistance to cephalosporins, aztreonam or carbapenems (Table 1).
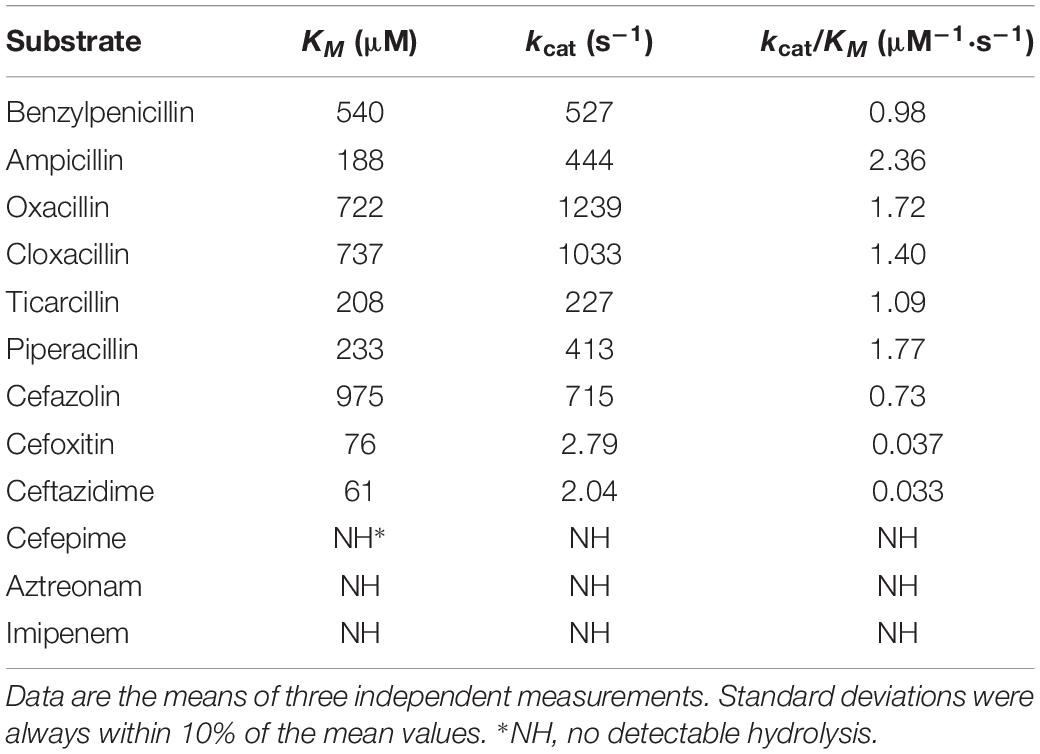
The kinetic parameters of the OXA-830 β-lactamase purified from the extract of E. coli BL21 harboring the recombinant plasmid pET-OXA-830 demonstrated that OXA-830 had a strong hydrolytic activity against penicillins (including oxacillin, cloxacillin, benzylpenicillin, ampicillin, piperacillin, and ticarcillin) and cefazolin (kcat/Km ratios were ≥0.73 μM–1⋅s–1), while very poor hydrolytic activities were detected for cefoxitin and ceftazidime. Moreover, no measurable hydrolytic activities were observed for cefepime, aztreonam and imipenem (Table 3). Nevertheless, this finding was not entirely consistent with the above MIC changes for E. coli DH5α recombinant clones (pUCP24-blaOXA–830/DH5α) in the antibiotic susceptibility assay, as the hydrolytic activities against cefazolin, cefoxitin and ceftazidime did not result in significant changes in the MICs for E. coli DH5α recombinant clones. It may be that the activity in vitro was simply too low to contribute to activity in vivo. A similar phenomenon was observed from OXA-436 which showed hydrolytic activity against ceftazidime but not confer resistance to ceftazidime (Samuelsen et al., 2018). OXA-830 exhibited an approximately 2-fold increase in the turnover rate (kcat) for oxacillin and cloxacillin compared to that for benzylpenicillin. The highest catalytic efficiency was observed with ampicillin (kcat/Km ratio was 2.36 μM–1⋅s–1). Of note, the catalytic efficiencies of OXA-830 against oxacillin and ampicillin matched or surpassed those of several clinically important DBLs in Gram-negative pathogens, such as OXA-48 and OXA-58 (Docquier et al., 2009; Verma et al., 2011). Compared to OXA-12 (Rasmussen et al., 1994), OXA-830 exhibited significantly lower affinity for penicillins used in both studies. Although susceptible to cephalosporins, the extended hydrolysis spectrum of OXA-830 indicated that it could be exceptionally classified into subgroup 2de, a new subgroup of DBLs including the oxacillin- or cloxacillin-hydrolyzing β-lactamases exhibiting hydrolytic activity against oxyimino-cephalosporins but not carbapenems (Bush and Jacoby, 2010). Interestingly, most of the members of this subgroup originated from OXA-10, with a small number of amino acid substitutions described as previously (Bush and Jacoby, 2010). However, OXA-830 exhibited only 22.6% global amino acid identity with OXA-10. In contrast, OXA-830 shared much higher global amino acid identity (72.2%) with OXA-427, which could hydrolyze imipenem and exhibit resistance to extended-spectrum cephalosporins (mostly ceftazidime) (Bogaerts et al., 2017).
The results of β-lactamase activity inhibition measurement, as measured by IC50s (50% inhibitory concentrations), showed that OXA-830 was strongly inhibited by sulbactam (IC50: 0.05 ± 0.01 μM) and tazobactam (IC50: 0.07 ± 0.02 μM), and was less sensitive to clavulanic acid (IC50: 0.26 ± 0.02 μM), which could explain the above MIC changes of β-lactams with or without β-lactamase inhibitors. However, this finding was inconsistent with what was found for OXA-12 (Rasmussen et al., 1994), which was strongly inhibited by clavulanic acid (IC50: 0.009 μM) and tazobactam (IC50: 0.03 μM), and less effectively inhibited by sulbactam (IC50: 0.24 μM).
Conclusion
The complete genome sequence of A. simiae was firstly reported in this work. We identified a novel OXA DBL named OXA-830 that was encoded on the chromosome of A. simiae A6. The new enzyme exhibited high amino acid sequence divergence from all currently available DBLs and showed the top amino acid identities (77.1–79.3%) with OXA-12 and OXA-12-like β-lactamases. Therefore, OXA-830 could be characterized as the first member of a new lineage of DBLs. OXA-830 showed some common functional properties with other OXA β-lactamases such as OXA-12, but significant differences were also observed. It possessed extended-spectrum hydrolytic properties which was strongly inhibited by both sulbactam and tazobactam. These findings revealed the importance of OXA-830 as a new model to study the structure/function relationships among OXA β-lactamases. Paired terminal IRs at both ends of the blaOXA–830-related region suggested that the possibility of dissemination of blaOXA–830 in the environment is still existent, although it is not associated with any known MGE.
Data Availability Statement
The datasets generated for this study can be found in the GenBank database as GenBank: CP040449 for the A. simiae A6 genome sequence, GenBank: MK926981 for the OXA-830 gene.
Author Contributions
QC, KS, XZ, DZ, HL, WL, and KL collected the strains and performed the experiments. WZ, ZS, and QC analyzed the experimental results. WZ, QC, and TX performed the bioinformatics analysis. WZ, JL, and QB co-led the writing of the manuscript. JL and QB designed the work. All authors read and approved the final version of the manuscript.
Funding
This research was supported by grants from the Natural Science Foundation of Zhejiang Province (LY14C060005 and LQ17H190001), the National Natural Science Foundation of China (31500109, 80215049, and 81570013), and Science and Technology Project of Inner Mongolia Autonomous Region, China (201802125).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors would like to acknowledge all study participants and individuals who contributed to this study.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2019.02732/full#supplementary-material
Footnotes
References
Adams-Haduch, J. M., Paterson, D. L., Sidjabat, H. E., Pasculle, A. W., Potoski, B. A., Muto, C. A., et al. (2008). Genetic basis of multidrug resistance in Acinetobacter baumannii clinical isolates at a tertiary medical center in Pennsylvania. Antimicrob. Agents Chemother. 52, 3837–3843. doi: 10.1128/AAC.00570-08
Almagro Armenteros, J. J., Tsirigos, K. D., Sønderby, C. K., Petersen, T. N., Winther, O., Brunak, S., et al. (2019). SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat. Biotechnol. 37, 420–423. doi: 10.1038/s41587-019-0036-z
Alperi, A., Figueras, M. J., Inza, I., and Martínez-Murcia, A. J. (2008). Analysis of 16S rRNA gene mutations in a subset of aeromonas strains and their impact in species delineation. Int. Microbiol. 11, 185–194.
Antonelli, A., D’Andrea, M. M., Vaggelli, G., Docquier, J.-D., and Rossolini, G. M. (2015). OXA-372, a novel carbapenem-hydrolysing class D β-lactamase from a Citrobacter freundii isolated from a hospital wastewater plant. J. Antimicrob. Chemother. 70, 2749–2756. doi: 10.1093/jac/dkv181
Beaz-Hidalgo, R., Latif-Eugenin, F., Hossain, M. J., Berg, K., Niemi, R. M., Rapala, J., et al. (2015). Aeromonas aquatica sp. nov., Aeromonas finlandiensis sp. nov. and Aeromonas lacus sp. nov. isolated from Finnish waters associated with cyanobacterial blooms. Syst. Appl. Microbiol. 38, 161–168. doi: 10.1016/j.syapm.2015.02.005
Bogaerts, P., Naas, T., Saegeman, V., Bonnin, R. A., Schuermans, A., Evrard, S., et al. (2017). OXA-427, a new plasmid-borne carbapenem-hydrolysing class D beta-lactamase in Enterobacteriaceae. J. Antimicrob. Chemother. 72, 2469–2477. doi: 10.1093/jac/dkx184
Bush, K., and Jacoby, G. A. (2010). Updated functional classification of beta-lactamases. Antimicrob. Agents Chemother. 54, 969–976. doi: 10.1128/AAC.01009-09
Bush, K., Jacoby, G. A., and Medeiros, A. A. (1995). A functional classification scheme for beta-lactamases and its correlation with molecular structure. Antimicrob. Agents Chemother. 39:1211. doi: 10.1128/aac.39.6.1211
Carvalho, M. J., Martinez-Murcia, A., Esteves, A. C., Correia, A., and Saavedra, M. J. (2012). Phylogenetic diversity, antibiotic resistance and virulence traits of Aeromonas spp. from untreated waters for human consumption. Int. J. Food Microbiol. 159, 230–239. doi: 10.1016/j.ijfoodmicro.2012.09.008
Chen, P. L., Ko, W. C., and Wu, C. J. (2012). Complexity of beta-lactamases among clinical aeromonas isolates and its clinical implications. J. Microbiol. Immunol. Infect. 45, 398–403. doi: 10.1016/j.jmii.2012.08.008
CLSI (2019). Performance Standards for Antimicrobial Susceptibility Testing, CLSI Supplement M100, 29th Edn. Wayne, PA: Clinical and Laboratory Standards Institute.
Cock, P. J., Antao, T., Chang, J. T., Chapman, B. A., Cox, C. J., Dalke, A., et al. (2009). Biopython: freely available Python tools for computational molecular biology and bioinformatics. Bioinformatics 25, 1422–1423. doi: 10.1093/bioinformatics/btp163
Colwell, R., MacDonell, M., and De Ley, J. (1986). Proposal to recognize the family Aeromonadaceae fam. nov. Int. J. Syst. Evol. Microbiol. 36, 473–477. doi: 10.1099/00207713-36-3-473
Corvec, S., Poirel, L., Naas, T., Drugeon, H., and Nordmann, P. (2007). Genetics and expression of the carbapenem-hydrolyzing oxacillinase gene blaOXA-23 in Acinetobacter baumannii. Antimicrob. Agents Chemother. 51, 1530–1533. doi: 10.1128/AAC.01132-06
Danel, F., Hall, L. M., Duke, B., Gur, D., and Livermore, D. M. (1999). OXA-17, a further extended-spectrum variant of OXA-10 β-lactamase, isolated from Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 43, 1362–1366. doi: 10.1128/aac.43.6.1362
Delcher, A. L., Bratke, K. A., Powers, E. C., and Salzberg, S. L. (2007). Identifying bacterial genes and endosymbiont DNA with Glimmer. Bioinformatics 23, 673–679. doi: 10.1093/bioinformatics/btm009
Docquier, J. D., Calderone, V., De Luca, F., Benvenuti, M., Giuliani, F., Bellucci, L., et al. (2009). Crystal structure of the OXA-48 beta-lactamase reveals mechanistic diversity among class D carbapenemases. Chem. Biol. 16, 540–547. doi: 10.1016/j.chembiol.2009.04.010
El Garch, F., Bogaerts, P., Bebrone, C., Galleni, M., and Glupczynski, Y. (2011). OXA-198, an acquired carbapenem-hydrolyzing class D beta-lactamase from Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 55, 4828–4833. doi: 10.1128/AAC.00522-11
European Committee on Antimicrobial Susceptibility Testing [EUCAST] (2019). Breakpoint Tables for Interpretation of MICs and Zone Diameters, Version 9.0. Available at: http://www.eucast.org.
Gardner, S. N., Slezak, T., and Hall, B. G. (2015). kSNP3.0: snp detection and phylogenetic analysis of genomes without genome alignment or reference genome. Bioinformatics 31, 2877–2878. doi: 10.1093/bioinformatics/btv271
Hall, L., Livermore, D., Gur, D., Akova, M., and Akalin, H. (1993). OXA-11, an extended-spectrum variant of OXA-10 (PSE-2) beta-lactamase from Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 37, 1637–1644. doi: 10.1128/aac.37.8.1637
Harf-Monteil, C., Fleche, A. L., Riegel, P., Prevost, G., Bermond, D., Grimont, P. A., et al. (2004). Aeromonas simiae sp. nov., isolated from monkey faeces. Int. J. Syst. Evol. Microbiol. 54(Pt 2), 481–485. doi: 10.1099/ijs.0.02786-0
Janda, J. M., and Abbott, S. L. (2010). The genus Aeromonas: taxonomy, pathogenicity, and infection. Clin. Microbiol. Rev. 23, 35–73. doi: 10.1128/CMR.00039-09
Katoh, K., and Standley, D. M. (2013). MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780. doi: 10.1093/molbev/mst010
Koren, S., Walenz, B. P., Berlin, K., Miller, J. R., Bergman, N. H., and Phillippy, A. M. (2017). Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 27, 722–736. doi: 10.1101/gr.215087.116
Kumar, S., Stecher, G., and Tamura, K. (2016). MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874. doi: 10.1093/molbev/msw054
Leonard, D. A., Bonomo, R. A., and Powers, R. A. (2013). Class D beta-lactamases: a reappraisal after five decades. Acc. Chem. Res. 46, 2407–2415. doi: 10.1021/ar300327a
McKenna, A., Hanna, M., Banks, E., Sivachenko, A., Cibulskis, K., Kernytsky, A., et al. (2010). The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303. doi: 10.1101/gr.107524.110
Moura, A., Soares, M., Pereira, C., Leitao, N., Henriques, I., and Correia, A. (2009). INTEGRALL: a database and search engine for integrons, integrases and gene cassettes. Bioinformatics 25, 1096–1098. doi: 10.1093/bioinformatics/btp105
Mugnier, P., Casin, I., Bouthors, A., and Collatz, E. (1998). Novel OXA-10-derived extended-spectrum β-lactamases selected in vivo or in vitro. Antimicrobial. Agents Chemother. 42, 3113–3116. doi: 10.1128/aac.42.12.3113
Paetzel, M., Danel, F., de Castro, L., Mosimann, S. C., Page, M. G., and Strynadka, N. C. (2000). Crystal structure of the class D β-lactamase OXA-10. Nat. Struct. Mol. Biol. 7:918.
Payne, D. J., Cramp, R., Winstanley, D. J., and Knowles, D. (1994). Comparative activities of clavulanic acid, sulbactam, and tazobactam against clinically important beta-lactamases. Antimicrob. Agents Chemother. 38, 767–772. doi: 10.1128/aac.38.4.767
Petkau, A., Stuart-Edwards, M., Stothard, P., and Van Domselaar, G. (2010). Interactive microbial genome visualization with GView. Bioinformatics 26, 3125–3126. doi: 10.1093/bioinformatics/btq588
Poirel, L., Naas, T., and Nordmann, P. (2010). Diversity, epidemiology, and genetics of class D beta-lactamases. Antimicrob. Agents Chemother. 54, 24–38. doi: 10.1128/AAC.01512-08
Poirel, L., and Nordmann, P. (2006). Genetic structures at the origin of acquisition and expression of the carbapenem-hydrolyzing oxacillinase gene blaOXA-58 in Acinetobacter baumannii. Antimicrob. Agents Chemother. 50, 1442–1448. doi: 10.1128/AAC.50.4.1442-1448.2006
Rasmussen, B. A., Keeney, D., Yang, Y., and Bush, K. (1994). Cloning and expression of a cloxacillin-hydrolyzing enzyme and a cephalosporinase from aeromonas sobria AER 14M in Escherichia coli: requirement for an E. coli chromosomal mutation for efficient expression of the class D enzyme. Antimicrob. Agents Chemother. 38, 2078–2085. doi: 10.1128/aac.38.9.2078
Saavedra, M. J., Figueras, M. J., and Martinez-Murcia, A. J. (2006). Updated phylogeny of the genus Aeromonas. Int. J. Syst. Evol. Microbiol. 56(Pt 10), 2481–2487. doi: 10.1099/ijs.0.64351-0
Samuelsen, O., Hansen, F., Aasnaes, B., Hasman, H., Lund, B. A., Leiros, H. S., et al. (2018). Dissemination and characteristics of a novel plasmid-encoded carbapenem-hydrolyzing class D beta-Lactamase, OXA-436, found in isolates from four patients at six different hospitals in denmark. Antimicrob. Agents Chemother. 62:e1260-17. doi: 10.1128/AAC.01260-17
Siguier, P., Perochon, J., Lestrade, L., Mahillon, J., and Chandler, M. (2006). ISfinder: the reference centre for bacterial insertion sequences. Nucleic Acids Res. 34, D32–D36. doi: 10.1093/nar/gkj014
Verma, V., Testero, S. A., Amini, K., Wei, W., Liu, J., Balachandran, N., et al. (2011). Hydrolytic mechanism of OXA-58 enzyme, a carbapenem-hydrolyzing class D beta-lactamase from Acinetobacter baumannii. J. Biol. Chem. 286, 37292–37303. doi: 10.1074/jbc.M111.280115
Walsh, T. R., Hall, L., MacGowan, A. P., and Bennett, P. M. (1995). Sequence analysis of two chromosomally mediated inducible β-lactamases from Aeromonas sobria, strain 163a, one a class D penicillinase, the other an AmpC cephalosporinase. J. Antimicrob. Chemother. 36, 41–52. doi: 10.1093/jac/36.1.41
Keywords: oxacillinase, OXA-830, Aeromonas simiae, resistance, kinetic analysis
Citation: Chen Q, Zhou W, Qian C, Shen K, Zhu X, Zhou D, Sun Z, Lu W, Liu H, Li K, Xu T, Bao Q and Lu J (2019) OXA-830, a Novel Chromosomally Encoded Extended-Spectrum Class D β-Lactamase in Aeromonas simiae. Front. Microbiol. 10:2732. doi: 10.3389/fmicb.2019.02732
Received: 21 August 2019; Accepted: 11 November 2019;
Published: 26 November 2019.
Edited by:
Miklos Fuzi, Semmelweis University, HungaryReviewed by:
Remy A. Bonnin, Université Paris-Saclay, FranceKrisztina M. Papp-Wallace, Louis Stokes Cleveland VA Medical Center, United States
Copyright © 2019 Chen, Zhou, Qian, Shen, Zhu, Zhou, Sun, Lu, Liu, Li, Xu, Bao and Lu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Qiyu Bao, YmFvcXlAZ2Vub21pY3MuY24=; Junwan Lu, MzA0MDQxNDc0QHFxLmNvbQ==
†These authors have contributed equally to this work
 Qianqian Chen
Qianqian Chen Wangxiao Zhou1†
Wangxiao Zhou1† Kai Shen
Kai Shen Kewei Li
Kewei Li Teng Xu
Teng Xu Qiyu Bao
Qiyu Bao Junwan Lu
Junwan Lu