Abstract
The potato late blight pathogen Phytophthora infestans has both an asexual and a sexual mode of reproduction. In Scandinavia, the pathogen is reproducing sexually on a regular basis, whereas clonal lineages dominate in other geographical regions. This study aimed at elucidating events or key genes underlying this difference in sexual behavior. First, the transcriptomes of eight strains, known as either clonal or sexual, were compared during early stages of mating. Principal component analysis (PCA) divided the samples in two clusters A and B and a clear grouping of the mating samples together with the A1 mating type parents was observed. Induction of genes encoding DNA adenine N6-methylation (6mA) methyl-transferases clearly showed a bias toward the cluster A. In contrast, the Avrblb2 effector gene family was highly induced in most of the mating samples and was associated with cluster B in the PCA, similarly to genes coding for acetyl-transferases, which play an important role in RXLR modification prior to secretion. Avrblb2 knock-down strains displayed a reduction in virulence and oospore formation, suggesting a role during the mating process. In conclusion, a number of gene candidates important for the reproductive processes were revealed. The results suggest a possible epigenetic influence and involvement of specific RXLR effectors in mating-related processes.
Introduction
Phytophthora infestans is one of the most devastating plant pathogens, infecting many species in the Solanaceae family, particularly potato and tomato. It is a fungal-like, filamentous pathogen, which belongs to Oomycota. Its leading role as one of the most notorious plant pathogens worldwide is associated with its rapid adaptation to fungicides and newly introduced plant resistance genes (Fry, 2008). This high speed of adaptation can be explained by an efficient and mixed reproduction system, and the large amount of effector genes and transposable elements in the P. infestans genome (Fry, 2008; Haas et al., 2009). Phytophthora infestans is characterized by asexual, parasexual and sexual reproduction systems that together generate a high evolutionary potential (Smoot et al., 1958; Galindo and Gallegly, 1960; Judelson and Yang, 1998). Mating in P. infestans requires the presence of A1 and A2 mating types, leading to zygote formation within the developing oospore (Goodwin and Drenth, 1997). Oospores can serve as survival structures, due to their thick cell walls, being dormant in soil for several years and also under harsh environmental conditions (Fay and Fry, 1997; Turkensteen et al., 2000), and they are an efficient source to the new cycle of infection for the coming growing seasons (Fernandez-Pavia et al., 2004; Grünwald and Flier, 2005). In Sweden and Finland, oospores can develop on surviving tubers in the soil and on volunteer plants in the next season (Andersson et al., 1998; Lehtinen and Hannukkala, 2004). In the late 20th century, the A1 mating type was dominating and A2 strains were present only in Mexico (Gallegly and Galindo, 1958). In Europe, the first report of A2 strains came in 1984 (Hohl and Iselin, 1984). In the United States, the dominating US-1 lineage (A1) was replaced by the US-8 lineage (A2) at about the same time-point (Fry et al., 1993). Migration and shipment of potato tubers have now resulted in a spread of the two mating types worldwide, making the P. infestans sexual reproduction possible in many regions (Grünwald and Flier, 2005). However, despite the presence of the two mating types, replacement of different virulent lineages is commonly leading to dominance for one genotype (Cooke et al., 2012). Clonal linages have therefore dominated the P. infestans populations in many geographical regions and over long time periods (Goodwin and Fry, 1994; Goodwin, 1997; Fry, 2008).
Still many molecular aspects of the sexual reproduction and emergence of clonal lineages of P. infestans remain unclear. In Europe, the A1 mating type has been dominating in the United Kingdom, France and Switzerland for many years. A shift to A2 dominance has recently been observed. For example, the displacement of other strains by the highly aggressive Blue-13 (also named 3928A) strain, which has acquired tolerance to phenylamide fungicides, has rapidly occurred during the past years (Cooke et al., 2012). A1 populations are present alongside Blue-13 under field conditions but no hybridization between them has been reported. In the Nordic countries, the situation is strikingly different. High genetic variability is present in the P. infestans populations, explained by a high frequency of mating, but no dominating clonal linage has been reported so far (Brurberg et al., 2011; Sjöholm et al., 2013). The mating process of P. infestans is more complicated compared to heterothallic fungal species and many aspects of it are still unknown. It has been demonstrated that the inheritance of the mating-type (MAT) locus does not follow Mendelian segregation patterns (Fabritius and Judelson, 1997). Rearrangements are common in this chromosomal region and the MAT locus is suggested to be hemizygous (Judelson, 1996; Randall et al., 2003; van der Lee et al., 2004).
To expand our knowledge on events explaining the outcome of different activities at the early stages of mating, total RNA from contrasting P. infestans strains collected in Sweden, the Netherlands, and the United Kingdom were prepared from in vitro cultures and sequenced. A clear bias in gene expression toward the A1 parental strains was observed among the samples. Several interesting genes, such as those coding for adenine methyl-transferases and histone modifiers, followed the same expression patterns. Members of the Avrblb2 effector gene family were highly induced, particularly in the Dutch mating sample. Avrblb2 knock-down lines showed reduction in oospore production, indicating a role in sexual reproduction. The impact of epigenetic factors influencing mating process is further discussed.
Materials and Methods
Phytophthora infestans Strains, Mating and Culture Conditions
All P. infestans strains were grown on rye-pea agar (RPA) medium (Caten and Jinks, 1968) supplemented with 2% sucrose and rifampicin (30 μg/ml) and maintained at 20°C in darkness. Eight P. infestans strains from potato were included in the mating experiments; four from Sweden (Sw1_A1, Sw2_A2, Sw3_A1, and Sw4_A2), two from the Netherlands (F80029_A1 and IPO82001_A2) and two from the United Kingdom (Pink 6_A1 and Blue 13/3928A_A2). The mating type of the above strains was confirmed by crossing them with tester strains, and self-fertilization was also investigated. In total, 10 mating samples were prepared; four Swedish one Dutch, one British and four “mixed” samples between Swedish and Dutch strains used to investigate whether there is any specific impact of Swedish strains during mating. Mating was promoted on plates where agar strips of A1 and A2 strains were placed in parallel, 2 cm apart, on RPA media, during the same growth conditions as described above (Figure 1A). After 7 days, material was collected from a 1 cm wide mating zone along the meeting point of the different combinations and used for total RNA extraction. The mating zone predominately contained swollen structures of hyphae, along with a mix of few antheridia and oogonia, grown on polycarbonate membrane. Parental strains and the 88069 strain were grown separately and treated exactly as the mating samples. For oospore formation, mating and parental samples were grown as described above for RNA extraction. Materials from the mating zones were harvested 14 days post inoculation, suspended in 20 ml H20 and oospores were counted using hemocytometer.
FIGURE 1
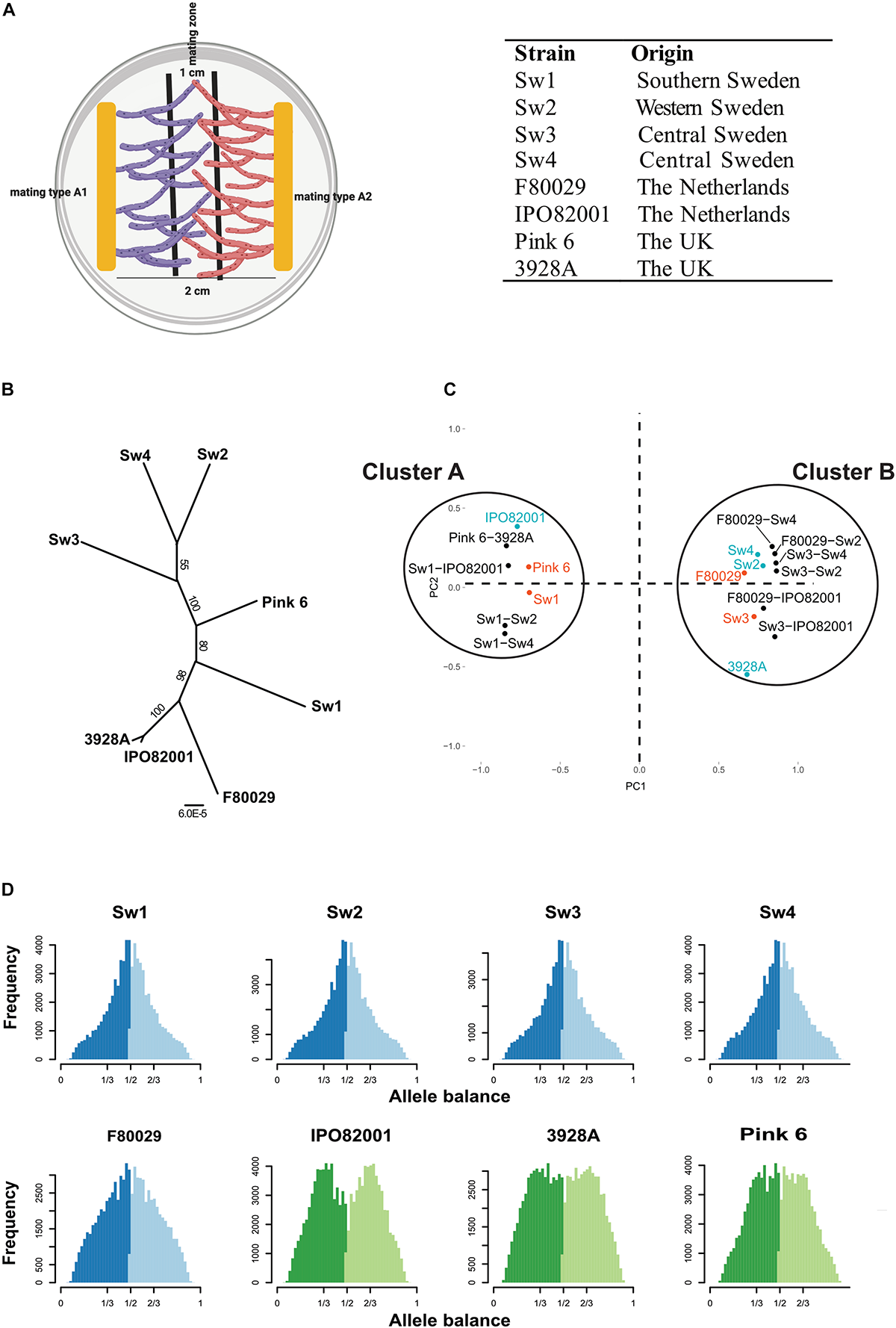
Characterization of P. infestans strains with the following mating types; A1: Sw1, Sw3, F80029, Pink 6. A2: Sw2, Sw4, IPO82001, 3928A. (A) Illustration on how material for RNA-seq analysis was collected. Mycelia for the mating zone (1 cm) between strains with opposite mating types (A1 or A2) were harvested 7 days post inoculation. (B) Phylogeny of P. infestans isolates based on SNP data, implemented with the RAxML v8.2X software, using the GTR + Γ model and automatic rapid bootstrapping. Numbers at the nodes indicate bootstrap values. Bar indicates number (6 × 10– 5) of substitutions per site. (C) Principal component analysis based on significantly differentially expressed genes during mating and in parental strains, grown individually, and normalized against P. infestans strain 88069. Red = mating type A1 strains. Blue = mating type A2 strains. Black = mating samples. (D) Ploidy analysis of P. infestans isolates, based on SNP data. Allele balance of 1/2 = diploid strain. Allele balance of 1/3 and 2/3 = triploid strain. Sw1, Sw2, Sw3, and Sw4 are Swedish isolates, F80029 and IPO82001 are Dutch, Pink6 and 3928A are British. Blue color show diploid strains and green color triploid strains.
Bioinformatic Analyses
Total RNA was extracted from the collected material using the Qiagen Plant mini kit (Qiagen), according to the manufacturer’s instructions. Three biological replicates were taken for each mating sample, and two for each parental sample. In total, 48 RNA strand-specific libraries were generated and sequenced using Illumina HiSeq 2500 at the SNP&SEQ Technology Platform, Science for Life Laboratory at Uppsala University, Sweden. The work flow describing major steps in the analysis is visualized in Supplementary Figure S1. All RNA-seq reads were cleaned for adaptor sequences and low quality reads using Trimmomatic v0.32 (Bolger et al., 2014). The filtered datasets were mapped to the P. infestans reference genome (Haas et al., 2009, NCBI accession number PRJNA17665) using Tophat v2.0.10 (Kim et al., 2013). Novel transcripts were discovered using Cufflinks v2.2.1 (Trapnell et al., 2010) and transcript mapping was assessed using HTSeq count v0.6.1 (Anders et al., 2015) with default settings, allowing no mismatches (Supplementary Table S1). Differential expression analysis was performed using DESeq2 v1.8.1 (Love et al., 2014). Two different normalization procedures were applied for the further analysis. Data from strain 88069 was used for the first data normalization, on which the principal component analysis was based. To analyze the mating specific variation, a second normalization was done, where each mating sample was normalized to both its parental strains, grown separately. Significant genes, showing differential expression between at least three of the Swedish matings and the British or Dutch ones, were selected as the main dataset in the Gene Ontology (GO) enrichment analysis (R-package topGO v2.24.0, Alexa and Rahnenführer, 2016). As background dataset, for the GO enrichment analysis, 1,720 genes with similar expression patterns as the main dataset were selected with Genefinder, R-package Genefilter v1.54.2 (Gentleman et al., 2016). GO-terms were inferred from FungiDB v3.0 (Stajich et al., 2012). For single nucleotide polymorphism (SNPs) analysis, duplicated reads were marked using Picard v1.137, GATK v3.3.0 SplitNCigarReads and Indel Realignment (McKenna et al., 2010). Variants were discovered using GATK HaplotypeCaller (de Pristo et al., 2011) and filtered using GATK VariantFiltration. All GATK tools were used with recommended settings for RNA sequencing standard workflow (van der Auwera et al., 2013). Variants were annotated using SnpEff v4.0 (Cingolani et al., 2012). For each protein-coding gene, a nucleotide sequence alignment with the eight samples was made with the discovered SNPs inferred. Genes lacking more than 50% sequence information, for more than four samples, were omitted from further analysis. A phylogenomic analysis was made on the concatenation of the remaining genes using RAxML v8.2X (GTR + Γ, automatic rapid bootstrap) (Stamatakis, 2014). A ploidy analysis was run based on the SNP data and displayed using the vcfR package (Knaus and Grünwald, 2018), filtering away the 10% quantile. The SNP variants, unique to each parent, were used to estimate the parental RNA content of each mating sample. To calculate the ratio of reads in the mating sample in relation to the parental sample, a linear equation was used (r = m/p, wherer = ratio, p = number of reads in the parental sample, m = number of reads in the mating sample) implemented in the R package matlib tool (Friendly et al., 2018). The distribution of the ratio for all used SNP variants was thereafter estimated for each mating sample, excluding SNP variants with read depth less than 100 and SNP variants with missing data for the mating sample. To filter for outliers, the 10 and 90% quantiles were removed.
Validation of RNA-Seq Data
The RNA-seq data were validated using qRT-PCR assay. 1000 ng of total RNA, treated with DNase I (Thermo Fisher Scientific), was reverse transcribed using iScript cDNA synthesis kit (Bio-Rad). qRT-PCR analysis was conducted as described previously (Tzelepis et al., 2012). Primers were designed from predicted exons and listed in Supplementary Table S2, and expression of genes was normalized by actin (actA) (Vetukuri et al., 2012). SNP analysis on actA gene (PITG_15117) (Supplementary Figure S2) showed that primers were located at regions identical to all the studied strains. Relative expression values were calculated from the threshold cycle (Ct) values by using the 2–ΔΔCT method (Livak and Schmittgen, 2001). Transcription patterns of the Avrblb2 genes were investigated on the same samples as used for RNA-seq analysis. Primers are listed in Supplementary Table S2. Normalization and expression analysis were performed as described above for RNA-seq validation.
Plasmids Construction, P. infestans Transformation and Oospore Production
For hairpin vector construction, the Avrblb2 family genes were cloned with sense and antisense copies, separated by a 71-nt intron from the Ste20-like gene also derived from P. infestans (Ah-Fong et al., 2008). Both copies were amplified from the P. infestans F80029 genomic DNA, using the primers listed in Supplementary Table S2 and Phusion DNA polymerase (Thermo Fisher Scientific). PCR products were digested (FseI and SbfI for sense; SacII and AscI for antisense) and ligated into the pFTORA vector (Vetukuri et al., 2012), driven by the ham34 promoter (Judelson et al., 1992). The orientation and integrity of insertions were confirmed by DNA sequencing (Macrogen Inc.). Transformation of P. infestans (F80029 strain) was performed as described by Vetukuri et al. (2011). Transcription levels of the Avrblb2 genes in P. infestans knock-down (KD) transformants were investigated using qRT-PCR analysis, as described above. Three independent KD transformants, showing similar growth rate and morphology, were used for functional studies.
Phytophthora infestans wild type F80029 strain (WT) and KD transformants in F80029 background were used for leaf inoculations. Leaves, derived from 3 to 4 weeks old Bintje and Bionika potato plants, were inoculated with 100,000 sporangia, while leaves, inoculated with water, were used as control samples. P. infestans DNA was extracted using the DNesay Plant Mini kit (Qiagen), and pathogen biomass was quantified in infected potato plants by qPCR, as described above for the qRT-RCR, using the P. infestans actin (actA) gene, normalized with the elongation factor gene (elf-1) from the plant. Primers are listed in Supplementary Table S2. The oospore production was evaluated in crossings in vitro on RPA media between the Avrblb2 KD transformants and the IPO82001 isolate (A2), compared to the WT (F80029) × IPO82001. Microscope slides, containing a thin layer of RPA, were used, and oospore production was counted at the meeting points, 5 and 10 days after mating. Oospores were calculated in the same area in all samples using a microscope (Zeiss Axioplan) and numbers were converted to oospores per mm2. For detached leaf and oospore production assays, at least three biological replicates and three independent KD transformants were used. The experiments were repeated twice. For the statistical analysis, the ANOVA (one-way) was conducted, using a General Linear model implemented in SPSS 20 (IBM), performing the Tukey’s or Student’s T tests at the 95% significance level.
Results
The A1 Genotype Displays Dominance Over the A2 During Mating
In this study we used strains from three European countries; four Swedish (Sw1–Sw4) from different regions (Supplementary Figure S3), two Dutch (IPO82001, F80029), and two British (Pink 6, 3928A). These eight strains had contrasting mating types (A1: Sw1, Sw3, F80029, Pink 6 and A2: Sw2, Sw4, IPO82001, and 3928A), enabling comparisons of events associated to the mating processes. The Dutch mating pair showed reduced oospore production as compared to the Swedish Sw1 × Sw2 and Sw3 × Sw2 samples (Supplementary Figure S4). No oospore production was observed in any parental samples. Total RNA was prepared from mycelia in the mating zones of 10 mating combinations: four Swedish (Sw1 × Sw2, Sw1 × Sw4, Sw3 × Sw2, and Sw3 × Sw4), one Dutch (F80029 × IPO82001), one British (pink 6 × 3928A) and four “mixed” (Sw1 × IPO82001, Sw3 × IPO82001, Sw2 × F80029, and Sw4 × F80029) during early stages of mating and prior to oospore formation. Parental strains were grown separately and treated similar to mating ones.
From Illumina RNAseq data, we first retrieved single nucleotide polymorphism (SNP) information. In phylogenomic analysis three Swedish strains (Sw2, Sw3, and Sw4) grouped together, whereas Sw1 was located closer among the British and Dutch isolates (Figure 1B). A principle component analysis (PCA) based on 10,341 significantly differentially expressed genes (DEGs) normalized against the 88069 strain generated two clusters (Figure 1C). Cluster A included the Sw1, the Pink 6, and the IPO82001 parental strains, and cluster B contained the five remaining parental strains (Sw2, Sw3, Sw4, F80029, and 3928A). All mating samples clustered together with their A1 parental strain. This observation suggests that mating type A1-related gene regulation is dominant during mating. Among the 5,183 DEGs between cluster A and cluster B, 1,602 genes displayed more than doubled expression levels (log2 > 1) between the clusters. This strongly divergent gene subset was slightly biased (868 genes), toward cluster A (Supplementary Figure S5A), whereas 734 genes, showed higher expression for cluster B (Supplementary Figure S5B). Among the 1,602 genes, six (PITG_18355, PITG_07745, PITG_21504, PITG_14044, PITG_03978, and PITG_17089) out of 25 were annotated as basal transcription factors in KEGG1. To check for possible biases in the data, an estimation of RNA content in the mating samples was made based on the SNP data (Supplementary Figure S6). No significant biases in RNA content were seen toward any of the two mating types. For example, the Sw1 × Sw2, Sw1 × Sw4, Sw3 × IPO82001 samples were estimated to have similar or slightly higher A2 RNA content, still displaying A1 grouping in the PCA plot (Figure 1C).
It is known that ploidy levels in P. infestans strains can vary (Goss et al., 2014; Knaus et al., 2016). Here we inferred the ploidy status of the P. infestans strains by exploiting the distribution of read counts at biallelic SNPs. All four Swedish strains and the Dutch F80029 strain displayed a diploid pattern, having an allele balance of 1/2 (Figure 1D). The IPO82001, 3928A, and Pink 6 strains were all triploid (Figure 1D). The results confirm earlier findings that isolates from sexual populations commonly are diploids, whereas clonal linages have a triploid genome constitution (Li et al., 2017). However, the triploids have the ability to change to a diploid genome constitution upon stress, which facilitates sexual reproduction and promotes genetic variation.
Acetylation and Epigenetic Activity During Mating
Transcriptome datasets of the ten different P. infestans isolate combinations were further dissected. The gene ontology (GO) analysis on all differentially expressed genes showed that only the transferase activity term (GO:0016747) displayed a significant enrichment during mating (Supplementary Table S3). Eight acetyl transferase homologs in P. infestans were induced in strains and mating samples (Figure 2A), and followed the B clustering (Figure 1C). Recently, it was shown that DNA methylation in P. infestans is regulated by three adenine N6-methylation (6mA) methyltransferase proteins (Chen et al., 2018). The transcription patterns of these genes (PITG_04679, PITG_07551, and PITG_07552) were investigated in our dataset. Interestingly, all three genes showed a bias toward cluster A (Figure 2B). To follow up on this finding, the transcription profile of genes, coding for putative histone modification proteins (PITG_09927, PITG_08042, and PITG_16916), were checked and found to display similar activity patterns, favoring cluster A (Figure 2C). The PITG_09927 contains multiple Tudor domains, which are recognized methylated histones (Huang et al., 2006; Lee et al., 2008). The PITG_08042 and PITG_16916 are putative Tyrosine kinases containing a colony-stimulating factor-1 (CSF-1), which is associated with chromatin (Tagoh et al., 2002). In contrast to the other gene families displayed, three subgroups were observed among the genes coding for histones; genes that were biased toward either cluster A or B, or those being constitutively expressed among the datasets (Figure 2D).
FIGURE 2
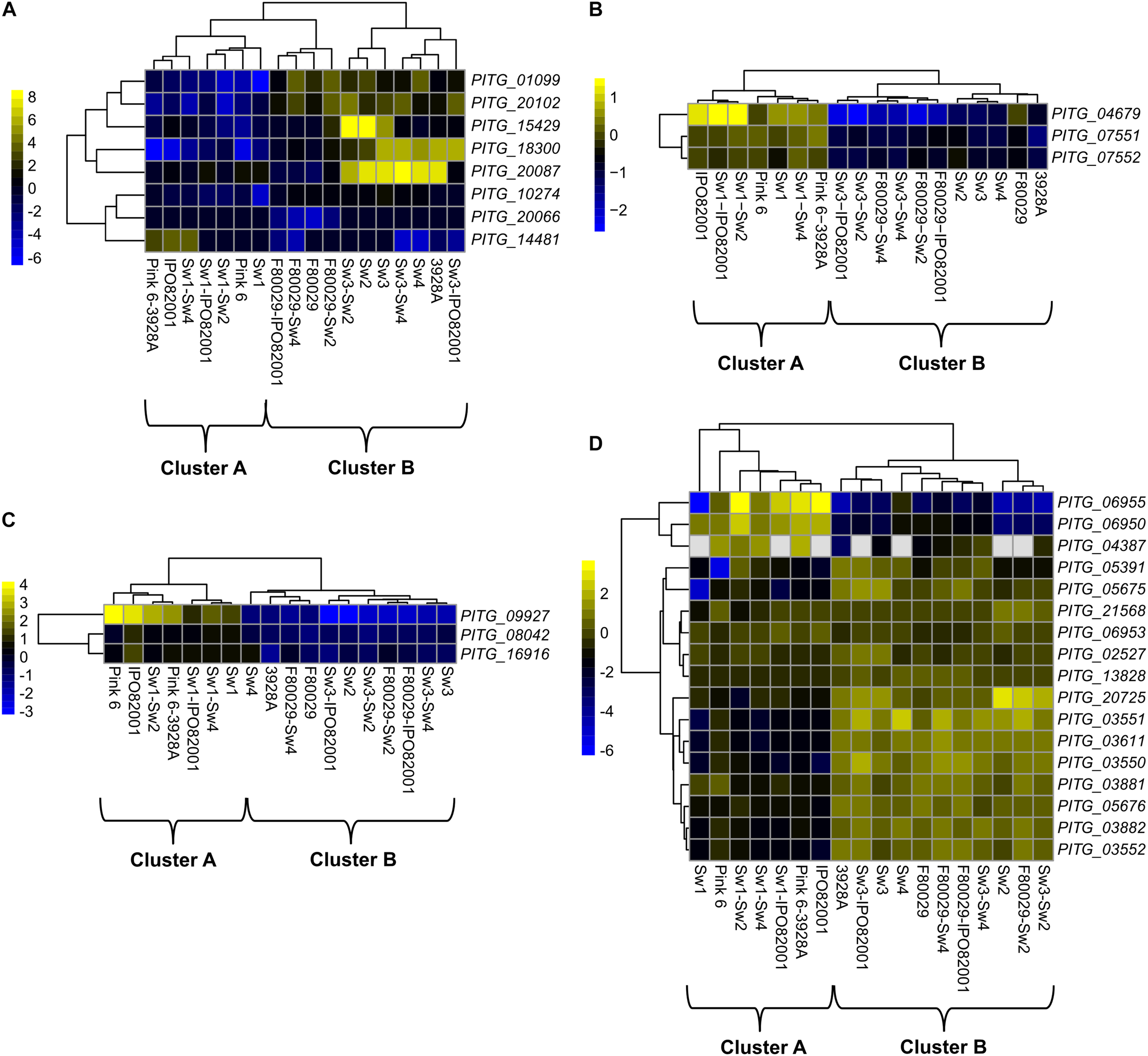
Transcription profiles of selected P. infestans genes. (A) acetyl-transferase genes, (B) adenine N6-methylation (6mA) methyltransferase (DAMT) genes, (C) histone modification genes and (D) genes encoding histone proteins. Data were normalized to the transcriptome of the P. infestans 88069 strain (adjusted p-value < 0.05, absolute log2 fold change > 2). Yellow and blue colors represent up-regulated or down-regulated genes.
The Dutch Mating Shows a Different Transcriptome Than the Swedish Ones
In order to investigate the transcriptome differences in Swedish mating samples versus samples from the Netherlands and the United Kingdom, a second normalization was performed where each mating sample was normalized to both parental strains, grown separately. Data from this analysis revealed that the Dutch (F80029 × IPO82001) and the Sw1 × IPO82001 pairs generated the most divergent patterns compared to the other eight combinations (Figures 3A,B). The expression profiles of selected genes were investigated using qRT-PCR analyses and were in agreement with the RNA-seq data (Figure 3 and Supplementary Figure S7), In total, 527 genes were differentially up-regulated in Swedish vs. Dutch strains, while only 18 in Swedish vs. British mating samples (Figure 3A). Among the genes that showed elevated transcripts in the Dutch pair compared to the Swedish pairs (Sw3 × Sw2, Sw1 × Sw4, Sw3 × Sw4, and Sw1 × Sw2) were genes encoding enzymes that are involved in energy production and metabolism, such as FAD-dependent oxidoreductase (PITG_03815) and dehydrogenases (PITG_00908, PITG_10290) (Supplementary Table S4). Genes that encoded ATP-binding cassette proteins (ABC transporters; PITG_07717, PITG_07716, PITG_04787) were also up-regulated in the Swedish mating samples, whereas PITG_17956 (the croquemort mating protein M82) was induced in the Dutch mating pair (Supplementary Table S4). Induction of PITG_17956 has been reported during the early stages of mating in P. infestans (Fabritius et al., 2002). Increased transcripts of ABC transporter genes might be related to higher needs of energy and mating hormones during potential mating in the Dutch strains. Elevated levels were observed also for genes that encode hydrolytic enzymes, such as endo-glucanase (PITG_08944) and beta-glucosidases (PITG_01398, PITG_01399, and PITG_01484) in the Dutch strains (Supplementary Table S4).
FIGURE 3
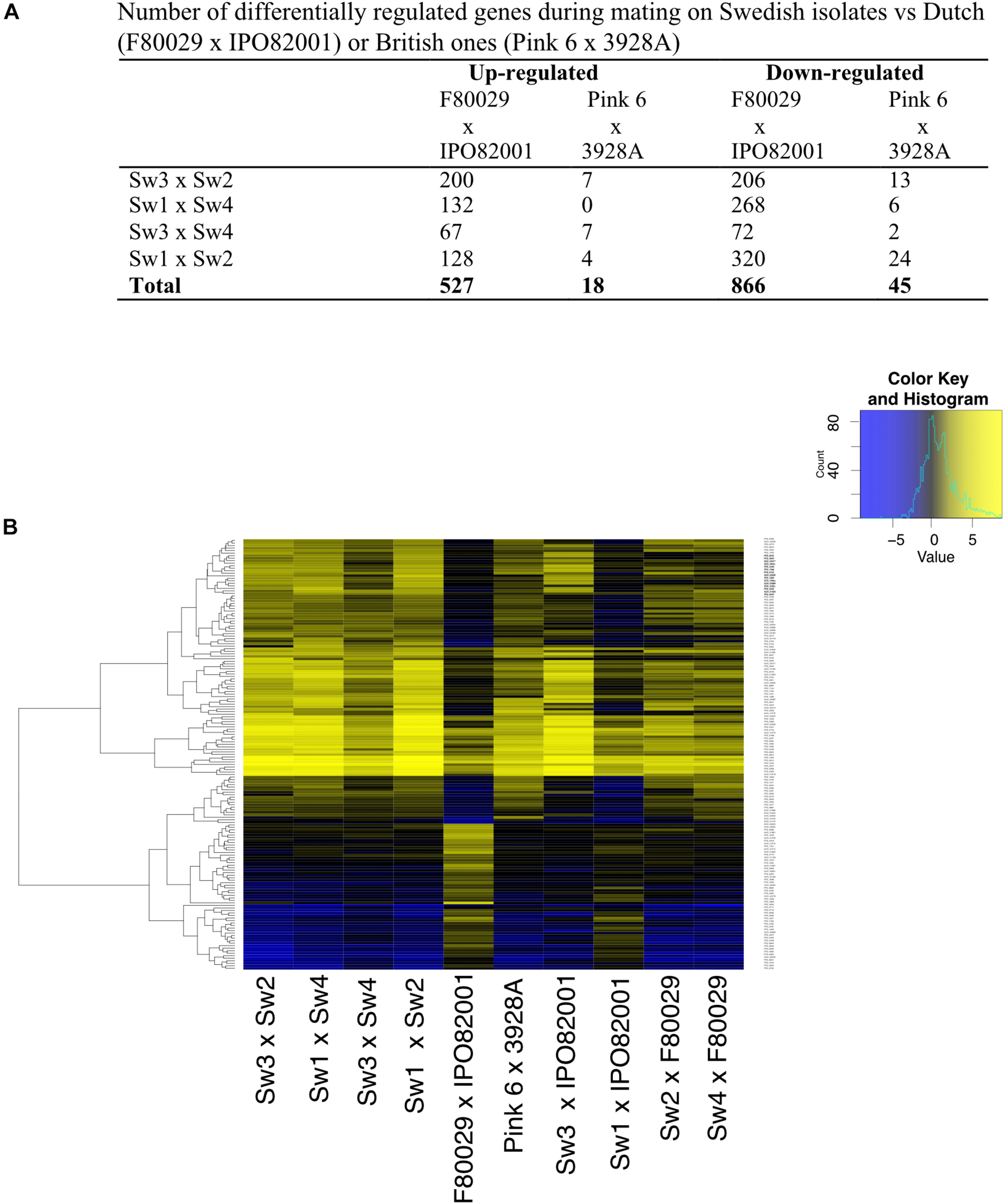
Transcription profiles of P. infestans strains during early stages of mating. (A) Number of differentially regulated genes in Swedish strains vs. Dutch (F80029 × IPO82001), or British ones (Pink 6 × 3928A). (B) Differentially expressed genes in the Dutch, Swedish or British mating samples. Data were normalized to the average expression of both parental strains grown individually (adjusted p-value < 0.05, absolute log2 fold change > 2). Yellow and blue colors represent up-regulated or down-regulated genes.
Finally, when comparing the repressed genes, 866 were differentially regulated between Swedish and Dutch mating samples and only 45 in Swedish vs. British ones (Figures 3A,B and Supplementary Table S5). We observed that genes, encoding the mating-induced protein 96 (M96) (PITG_02519, PITG_09067, PITG_22707, PITG_05905), had reduced transcript levels in the Dutch mating pair (F80029 × IPO82001). Similar patterns were observed for genes coding for Crinkler effector proteins (PITG_09043 and PITG_09053).
The Avrblb2 Effector Family Is Induced During Early Stages of Mating
Four genes, coding for RXLR effector proteins, were highly induced in the Dutch mating pair. These were Avr2 (PITG_22870), Avrblb2 (PITG_04085, PITG_04086), and PITG_12057 (Supplementary Table S4). The Avrblb2 is a family of seven paralogs sharing more than 90% sequence similarity (Supplementary Figure S8), and the PITG_04085 and PITG_04086 are antisense copies oriented in a head-to-head configuration (Oliva et al., 2015). SNP analysis on Avrblb2 genes showed one nucleotide insertion or deletion in some stains (Supplementary Figure S9). Further analysis on Avrblb2 genes, using the transcriptome of the isolate 88069 as a reference, revealed that induction of these genes, in parental and mating samples (Figure 4A), grouped to cluster B in the PCA plot, similar to acetyltransferase expression patterns (Figure 2A). N-terminal acetyltransferases play an important role in acetylation of effectors before secretion and cleavage of the RXLR motif in P. infestans, as exemplified by AVR3aEM (Wawra et al., 2017). Validation by qRT-PCR analysis revealed that Avrblb2 was highly induced, not only in the Dutch strain mating (F80029 × IPO82001), but also in two of the Swedish samples (Sw3 × Sw2 and Sw3 × Sw4) as compared with the parental samples (Figure 4B). No up-regulation of this effector was observed during mating, between the British strains (Figure 4B).
FIGURE 4
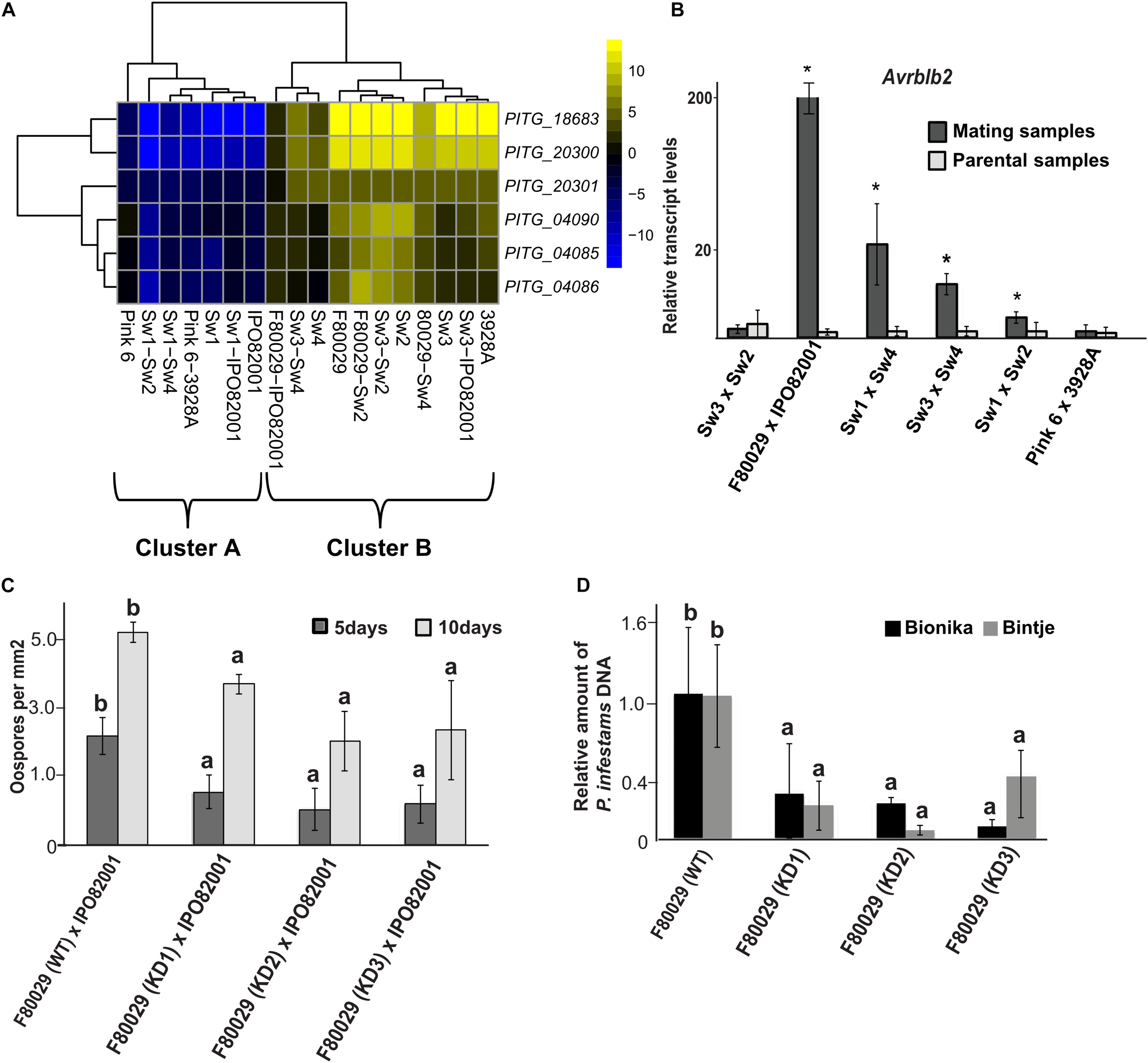
Transcription and phenotypic analysis of P. infestans. Avrblb2 effector family. (A) Heatmap of genes coding for Avrblb2 effectors. Data were normalized to the transcriptome of the 88069 P. infestans isolate (adjusted p-value < 0.05, absolute log2 fold change > 2). Yellow and blue colors represent up- or down-regulated genes. (B) qRT-PCR analysis of the Avrblb2 genes during mating compared to parental strains. Data were normalized to the expression of actin (act) and calculated, using the 2–DDCt method. Average of both parental strains is shown at the graph. Asterisks (*) indicate statistically significant differences between mating and parental samples according to Student’s T-test (p < 0.05). (C) Quantification of oospore production in F80029 (WT) strain and Avrblb2 (KD) transformants, crossed with IPO82001 (A2) strain, 5 and 10 days after mating. Different letters (a,b) indicate statistically significant differences between crossings of WT × IPO82001 vs. KD × IPO82001 according to Tukey’s T test (p < 0.05). (D) Quantification of P. infestans F80029 strain (WT) and Avrblb2 (KD) transformants biomass upon infection of Bintje and Bionika leaves, 3 and 4 days post inoculation, respectively. qPCR was conducted in genomic DNA, extracted from infected leaves using the P. infestans act gene, and normalized to the potato elf-1 gene. Error bars represent standard deviation (SD), based on at least three biological replicates. Different letters (a,b) indicate statistically significant differences according to Tukey’s test (p < 0.05). For detached leaf and oospore production assays, at least three biological replicates and three independent KD transformants were used. The experiments were repeated twice.
The Avrblb2 family genes were knocked down (KD), using a hairpin construct, and the role of this effector was studied in virulence and oospore formation. Three independent transformants in the F80029 background (mating type A1), displaying stable resistance to geneticin, were selected, and all of them showed significantly reduced expression levels of the Avrblb2 genes (Supplementary Figure S10). Any attempt to knock-down the Avrblb2 homolog in the IPO82001 strain continuously failed. The potential impact of the Avrblb2 effector in reproduction was investigated, using crosses of the KD transformants in F80029 genomic background with the IPO82001 (mating type A2) strain. Oospore production was measured at two time points, 5 and 10 days after mating. Reduced oospore production was observed at both time points, supporting a potential role of this effector category in sexual reproduction (Figure 4C). The role of the Avrblb2 effectors in P. infestans virulence was also evaluated on two potato cultivars, with variable resistance levels. Bintje is a highly susceptible cultivar, while Bionika harbors the Rpi-blb-2 gene introduced from Solanum bulbocastanum (Song et al., 2003; van der Vossen et al., 2005). Pathogen biomass was quantified upon infection with F80029 or the three KD transformants applied individually and we observed that the silenced lines displayed reduced biomass on infected plant tissue compared with the wild type in both cultivars (Figure 4D).
Discussion
The overall aim of the current work was to advance the understanding of mechanisms explaining differences of mating capacity at the early stages between different European P. infestans strains. To address this question we compared British, Dutch and Swedish strains on transcriptome levels to elucidate potential differences between mating-associated gene activities. Based on SNP data, we found that the Sw1 genotype was more similar to the Dutch and British strains than to the three other Swedish strains. The comparative transcriptomic analysis between Swedish and Dutch or British mating samples showed also that the Sw1 × IPO82001 mating sample behaves more similar to the Dutch mating rather than to other Swedish ones. This result was not completely unexpected, since the Sw1 isolate was collected in the southernmost part of Sweden, and long-distance wind spreading of sporangia, from Central Europe, would facilitate establishment of new P. infestans genotypes in this region. When comparing the eight parental P. infestans strains, the gene expression patterns differentiated into two groups not linked to mating type or ploidy levels; Sw1, IPO82001, Pink 6 form cluster A and F80029, 3928A, Sw2, Sw3, Sw4 cluster B. We found mating samples in the same groups as the parents, strictly linked to the gene expression of the parental A1 strains, in each combination. The grouping into cluster A and B is stringently seen in 1,602 genes (Supplementary Figure S5). Among those genes, basal transcription factors and epigenetic factors were found, possibly regulating the remaining gene set.
The underlining explanation of the division into frequent sexual reproduction of P. infestans strains, as found in Northern Europe versus the clonal lineages in Western Europe, appears to be regulated by complex events. Epigenetic influences during sexual reproduction, such as DNA methylation, histone modification and small RNAs (sRNAs), have been reported in several organisms including plants, animals and fungi (Quadrana and Colot, 2016; Kronholm et al., 2016; Donkin and Barrès, 2018). So far, no evidence of 5-methylcytosine DNA-methylation, which is prevalent in many organisms, has been reported in P. infestans (van West et al., 2008). Instead, adenine N6-methylation (6mA) methyltransferases (DAMTs) are shown to generate epigenetic marks across its genome (Chen et al., 2018). The three DAMT genes were also found to be up-regulated in our PCA group. A dataset, in contrast to the different developmental stages of P. infestans analyzed by Chen et al. (2018).
Whether epigenetic reprogramming, including transgenerational gene silencing, is a frequently occurring mechanism in the evolution and adaptation to new environments by P. infestans requires new in-depth analysis. The high proportion of transposable elements (TEs) in Phytophthora genomes and the co-localization of effector genes in TE-rich regions (Haas et al., 2009; Whisson et al., 2012) may drive epigenetic events in P. infestans as well. Emerging reports on sexual reproduction in animals and plants have uncovered different active RNA silencing pathways during sporogenesis, gametogenesis and after fertilization, influencing reproductive success. Processes involved are changes in DNA methylation, histone reprogramming, TE reactivation, and involvement of different classes of sRNAs (Martinez and Köhler, 2017; Borges et al., 2018; Martinez et al., 2018). Epigenetic reprogramming in plants also can be triggered by abiotic constrains (Wibowo et al., 2016; Lämke and Bäurle, 2017) and if applicable to P. infestans, such a function could generate increased opportunities to adapt to various changes in climate conditions. The extent of genetic variation in Scandinavian P. infestans populations has been debated (Maurice et al., 2019). Beside different marker systems employed for the analysis by different research groups, variation in the observations may as well reflect epigenetic influences driven by annual climate fluctuations.
Here, we used our dataset to generate some insights into the activity of annotated histone-modifying genes in P. infestans. This strategy demonstrated clear sub-groups among the different samples in our study. We hypothesize that P. infestans has evolved its own epigenetic program, acting on sexual reproduction, eventually leading to either clonal or genetically heterogeneous field populations. Dissection of gametes, zygotes, spores and surrounding somatic nurse tissues or cells, in combination with high-through-put sequencing technologies and transgenic lines with silenced target genes, are required to experimentally demonstrate the proposed transgenerational events in mating between different strains of P. infestans. However, P. infestans is a cumbersome organism to explore this type of advanced molecular functions.
Genes coding for the Avrblb2 effector family were highly induced during mating in most of our analyzed samples. Similarly, induction of RXLR and Crinkler effector genes has also been seen during mating between three other A1/A2 strains (Niu et al., 2018). The function of Avrblb2 effector, in the plant infection process, is to block the plant C14 protease and prevent its secretion into the apoplast (Bozkurt et al., 2011). In a search of proteases in the P. infestans genome, we found a cysteine protease homolog to C14 (PITG_02030), a gene with no differential gene expression in our mating datasets (Supplementary Figure S11). Secreted proteases are also known to play important roles in the mating processes of Saccharomyces cerevisiae, where they influence the mating pheromones in various ways (Barkai et al., 1998; Jin et al., 2011). In the yeast system the aspartyl protease Bar1, functions as a guide to ensure mating with a suitable partner (Ballensiefen and Schmitt, 1997). In the P. infestans genome, we identified a homolog to Bar1, PITG_10524, which displayed similar transcription pattern as Avrblb2 family (Supplementary Figure S11). Whether P. infestans shares some of the gene functions related to mating known in yeast remain to be demonstrated.
In this study, events during mating, between different strains of P. infestans were monitored on transcriptome levels. A clear dominance of mating type A1 and a potential epigenetic impact on the sexual reproduction was observed. Functional analysis of the Avrblb2 effector suggested a possible role in other types of cell-cell interactions rather than only in plant–pathogen interactions, including those affecting the local microbial community (Snelders et al., 2018). The observations reported here on several new potential mating-associated gene candidates and their regulation form the basis of new, research directions on the biology of P. infestans.
Statements
Data availability statement
The RNA-seq data used in this study has been deposited in the National Center for Biotechnology Information (NCBI) database, and Gene Expression Omnibus under the accession number GSE94137.
Author contributions
CD, RV, AÅ, and GT conceived the experiments. GT, AÅ, and RV performed the experiments. JF and KH conducted the bioinformatics analysis and interpreted the data. All authors contribute to writing of the manuscript and text revision.
Funding
This work was funded by Helge Ax:son Johnson’s Foundation, the Swedish Research Council Formas and VR 2015-04259, and the Swedish University of Agricultural Sciences.
Acknowledgments
The authors would like to thank Science for Life Laboratory, the National Genomics Infrastructure (NGI), Sweden, and UPPMAX for providing assistance in RNA sequencing and computational infrastructure. The sequencing platform was supported by the Swedish Research Council (VR-RFI) and the Knut and Alice Wallenberg Foundation.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2020.00252/full#supplementary-material
References
1
Ah-FongA. M. V.Bormann-ChungC. A.JudelsonH. S. (2008). Optimization of transgene-mediated silencing in Phytophthora infestans and its association with small-interfering RNAs.Fungal. Genet. Biol.451197–1205. 10.1016/j.fgb.2008.05.009
2
AlexaA.RahnenführerJ. (2016). topGO: Enrichment Analysis for Gene Ontology. R Package Version 2.32.0.
3
AndersS.PylP. T.HuberW. (2015). HTSeq–a Python framework to work with high-throughput sequencing data.Bioinformatics31166–169. 10.1093/bioinformatics/btu638
4
AnderssonB.SandstromM.StrombergA. (1998). Indications of soilborne inoculum of Phytophthora infestans.Potato Res.41305–310. 10.1007/BF02358962
5
BallensiefenW.SchmittH. D. (1997). Periplasmic Bar1 protease of Saccharomyces cerevisiae is active before reaching its extracellular destination.Eur. J. Biochem.247142–147. 10.1111/j.1432-1033.1997.00142.x
6
BarkaiN.RoseM. D.WingreenN. S. (1998). Protease helps yeast find mating partners.Nature396422–423. 10.1038/24760
7
BolgerA. M.LohseM.UsadelB. (2014). Trimmomatic: a flexible trimmer for Illumina sequence data.Bioinformatics302114–2120. 10.1093/bioinformatics/btu170
8
BorgesF.ParentJ. S.van ExF.WolffP.MartínezG.KöhlerC.et al (2018). Transposon-derived small RNAs triggered by miR845 mediate genome dosage response in Arabidopsis.Nat. Genet.50186–192. 10.1038/s41588-017-0032-35
9
BozkurtT. O.SchornackS.WinJ.TakayukiS.IlyasM.OlivaR.et al (2011). Phytophthora infestans effector AVRblb2 prevents secretion of a plant immune protease at the haustorial interface.Proc. Natl. Acad. Sci. U.S.A.10820832–20837. 10.1073/pnas.1112708109
10
BrurbergM. B.ElameenA.LeV. H.NaerstadR.HermansenA.LehtinenA.et al (2011). Genetic analysis of Phytophthora infestans populations in the Nordic European countries reveals high genetic variability.Fungal Biol.115335–342. 10.1016/j.funbio.2011.01.003
11
CatenC. E.JinksJ. L. (1968). Spontaneous variability of single isolates of Phytophthora infestans. I. Cultural variation.Can. J. Bot.46329–348. 10.1139/b68-055
12
ChenH.ShuH.WangL.ZhangF.LiX.OcholaS. O.et al (2018). Phytophthora methylomes are modulated by 6mA methyltransferases and associated with adaptive genome regions.BMC Genome Biol.19:181. 10.1186/s13059-018-1564-1564
13
CingolaniP.PlattsA.WangL. L.CoonM.NguyenT.WangetL.et al (2012). A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w(1118); iso-2; iso-3.Fly680–92. 10.4161/fly.19695
14
CookeD. E. L.CanoL. M.RaffaeleS.BainR. A.CookeL. R.EtheringtonG. J.et al (2012). Genome analyses of an aggressive and invasive lineage of the Irish potato famine pathogen.PLoS Pathog.8:e1002940. 10.1371/journal.ppat.1002940
15
de PristoM. A.BanksE.PoplinR.GarimellaK. V.MaguireJ. R.HartlC.et al (2011). A framework for variation discovery and genotyping using next-generation DNA sequencing data.Nat. Genet.43269–276. 10.1038/ng.806
16
DonkinI.BarrèsR. (2018). Sperm epigenetics and influence of environmental factors.Mol. Metab.141–11. 10.1016/j.molmet.2018.02.006
17
FabritiusA. L.JudelsonH. S. (1997). Mating type loci segregate aberrantly in Phytophthora infestans but normally in Phytophthora parasitica: implications for models of mating-type determination.Curr. Genet.3260–65. 10.1007/s002940050248
18
FabritiusA. L.CvitanichC.JudelsonH. S. (2002). Stage-specific gene expression during sexual development in Phytophthora infestans.Mol. Microbiol.451057–1066. 10.1046/j.1365-2958.2002.03073.x
19
FayJ. C.FryW. E. (1997). Effects of hot and cold temperatures on the survival of oospores produced by United States strains of Phytophthora infestans.Am. Potato J.74315–323. 10.1007/BF02851575
20
Fernandez-PaviaS. P.GrünwaldN. J.Diaz-ValasisM.Cadena-HinojosaM.FryW. E. (2004). Soilborne oospores of Phytophthora infestans in central Mexico survive winter fallow and infect potato plants in the field.Plant Dis.8829–33. 10.1094/PDIS.2004.88.1.29
21
FriendlyM.FoxJ.ChalmersP.MonetteG.SanchezG. (2018). Matlib: Matrix Functions for Teaching and Learning Linear Algebra and Multivariate Statistics. R Package Version 0.91.
22
FryW. (2008). Phytophthora infestans: the plant (and R gene) destroyer.Mol. Plant Pathol.9385–402. 10.1111/j.1364-3703.2007.00465.x
23
FryW. E.GoodwinS. B.DyerA. T.MatuszakJ. M.DrenthA.TooleyP. W.et al (1993). Historical and recent migrations of Phytophthora infestans; chronology, pathways, and implications.Plant Dis.77653–661. 10.1094/PD-77-0653
24
GalindoJ.GalleglyM. E. (1960). The nature of sexuality in Phytophthora infestans.Phytopathology50123–128. 10.1094/MPMI-08-16-0156-R
25
GalleglyM. E.GalindoJ. (1958). Mating types and oospores of Phytophthora infestans in nature in Mexico.Phytopathology48274–277.
26
GentlemanR.CareyV.HuberW.HahneF. (2016). Genefilter: Genefilter: Methods for Filtering Genes from High-Throughput Experiments. R Package Version 1.62.0.
27
GoodwinS. B. (1997). The population genetics of Phytophthora.Phytopathology87462–473. 10.1094/PHYTO.1997.87.4.462
28
GoodwinS. B.FryW. E. (1994). Genetic analyses of interspecific hybrids between Phytophthora infestans and Phytophthora mirabilis.Exp. Mycol.1820–32. 10.1006/emyc.1994.1003
29
GoodwinS.B.DrenthA. (1997). Origin of the A2 mating type of Phytophthora infestans outside Mexico.Phytopathol.87992–999. 10.1094/phyto.1997.87.10.992
30
GossE. M.TabimaJ. F.CookeD. E. L.RestrepoS.FryW. E.ForbesG. A.et al (2014). The Irish potato famine pathogen Phytophthora infestans originated in central Mexico rather than in the Andes.Proc. Natl. Acad. Sci. U.S.A.1118791–8796. 10.1073/pnas.1401884111
31
GrünwaldN. J.FlierW. G. (2005). The biology of Phytophthora infestans at its center of origin.Annu. Rev. Phytopathol.43171–190. 10.1146/annurev.phyto.43.040204.135906
32
HaasB. J.KamounS.ZodyM. C.JiangR. H. Y.HandsakerR. E.LilianaM.et al (2009). Genome sequence and analysis of the Irish potato famine pathogen Phytophthora infestans.Nature461393–398. 10.1038/nature08358
33
HohlH. R.IselinK. (1984). Strains of Phytophthora infestans from Switzerland with A2-mating type behavior.Trans. Brit. Mycol. Soc.83529–530. 10.1016/s0007-1536(84)80057-1
34
HuangY.FangJ.BedfordM. T.ZhangY.XuR. M. (2006). Recognition of histone H3 lysine-4 methylation by the double tudor domain of JMJD2A.Science312748–751. 10.1126/science.1125162
35
JinM.ErredeB.BeharM.MatherW.NayakS.HastyJ.et al (2011). Yeast dynamically modify their environment to achieve better mating efficiency.Sci. Signal4:ra54. 10.1126/scisignal.2001763
36
JudelsonH. S. (1996). Chromosomal heteromorphism linked to the mating type locus of the oomycete Phytophthora infestans.Mol. Genet. Genom.252155–161. 10.1007/BF02173215
37
JudelsonH. S.YangG. (1998). Recombination pathways in Phytophthora infestans: polyploidy resulting from aberrant sexual development and zoospore-mediated heterokaryosis.Mycol. Res.1021245–1253. 10.1017/S0953756298006388
38
JudelsonH. S.TylerB. M.MichelmoreR. W. (1992). Regulatory sequences for expressing genes in oomycete fungi.Mol. Gen. Genet.234138–146. 10.1007/BF00272355
39
KimD.PerteaG.TrapnellC.PimentelH.KelleyR.SalzbergS. L. (2013). TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions.Genome Biol.14:R36. 10.1186/gb-2013-14-4-r36
40
KnausB. J.GrünwaldN. J. (2018). Inferring variation in copy number using high throughput sequencing data in R.Front. Genet.9:123. 10.3389/fgene.2018.00123
41
KnausB. J.TabimaJ. F.DavisC. E.JudelsonH. S.GrünwaldN. J. (2016). Genomic analysis of dominant U.S. clonal lineages of Phytophthora infestans reveals a share common ancestry of clonal lineages US11 and US18 and a lack of recently shared ancestry among all other U.S. lineages.Phytopathology1061393–1403. 10.1094/PHYTO-10-15-0279-R
42
KronholmI.JohannessonH.KetolaT. (2016). Epigenetic control of phenotypic plasticity in the filamentous fungus Neurospora crassa.G364009–4022. 10.1534/g3.116.033860
43
LämkeJ.BäurleI. (2017). Epigenetic and chromatin-based mechanisms in environmental stress adaptation and stress memory in plants.Genome Biol.18:124. 10.1186/s13059-017-1263-6
44
LeeJ.ThompsonJ. R.BotuyanM. V.MerG. (2008). Distinct binding modes specify the recognition of methylated histones H3K4 and H4K20 by JMJD2A-tudor.Nat. Struct. Mol. Biol.15109–111. 10.1038/nsmb1326
45
LehtinenA.HannukkalaA. (2004). Oospores of Phytophthora infestans in soil provide an important new source of primary inoculum in Finland.Agr. Food Sci.13399–410. 10.2137/1239099043633332
46
LiY.ShenH.ZhouQ.QianK.van der LeeT.HuangS. (2017). Changing ploidy as a strategy: the Irish potato famine pathogen shifts ploidy in relation to its sexuality.Mol. Plant Microbe Interact.3045–52. 10.1094/MPMI-08-16-0156-R
47
LivakK. J.SchmittgenT. D. (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(T)(-Delta Delta C) method.Methods25402–408. 10.1006/meth.2001.1262
48
LoveM. I.HuberW.AndersS. (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2.Genome Biol.15:550. 10.1186/s13059-014-0550-558
49
MartinezG.KöhlerC. (2017). Role of small RNA in epigenetic reprogramming during pant sexual reproduction.Curr. Opin. Plant. Biol.3622–28. 10.1016/j.pbi.2016.12.006
50
MartinezG.WolffP.WangZ.Moreno-RomeroJ.Santos-GonzálezJ.ConzeL. L.et al (2018). Paternal easiRNAs regulate parental genome dosage in Arabidopsis.Nat. Genet.50193–198. 10.1038/s41588-017-0033-34
51
MauriceS.MontesM. S.NielsenB. J.BødkerL.MichaelD.MartinM.D.et al (2019). Population genomics of an outbreak of the potato late blight pathogen, Phytophthora infestans, reveals both clonality and high genotypic diversity.Mol. Plant Pathol.201134–1146. 10.1111/mpp.12819
52
McKennaA.HannaM.BanksE.SivachenkoA.CibulskisK.KernytskyA.et al (2010). The genome analysis toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data.Genome Res.201297–1303. 10.1101/gr.107524.110
53
NiuX.Ah-FongA. M. V.LopezL. A.JudelsonH. S. (2018). Transcriptomic and proteomic analysis reveals wall-associated and glucan-degrading proteins with potential roles in Phytophthora infestans sexual spore development.PLoS One13:e0198186. 10.1371/journal.pone.0198186
54
OlivaR. F.CanoL. M.RaffaeleS.WinJ.BozkurtT. O.BelhajK.et al (2015). A recent expansion of the RXLR effector gene Avrblb2 is maintained in global populations of Phytophthora infestans indicating different contributions to virulence.Mol. Plant Microbe Interact.28901–912. 10.1094/MPMI-12-14-0393-R
55
QuadranaL.ColotV. (2016). Plant transgenerational epigenetics.Annu. Rev. Genet.51467–491. 10.1146/annurev-genet-120215-135254
56
RandallT. A.Ah-FongA.JudelsonH. S. (2003). Chromosomal heteromorphism and an apparent translocation detected using a BAC contig spanning the mating type locus of Phytophthora infestans.Fungal Genet. Biol.3875–84. 10.1016/S1087-1845(02)00512-511
57
SjöholmL.AnderssonB.HogbergN.WidmarkA. K.YuenJ. (2013). Genotypic diversity and migration patterns of Phytophthora infestans in the Nordic countries.Fungal Biol.117722–730. 10.1016/j.funbio.2013.08.002
58
SmootJ. J.GoughF. J.LameyH. A.EichenmullerJ. J.GalleglyM. E. (1958). Production and germination of oospores of Phytophthora infestans.Phytopathol.48165–171.
59
SneldersN. C.KettlesG. J.RuddJ. J.ThommaB. P. H. J. (2018). Plant pathogen effector proteins as manipulators of host microbiomes?Mol. Plant Pathol.19257–259. 10.111/mpp.12628
60
SongJ.BradeenJ. M.NaessS. K.RaaschJ. A.WielgusS. M.HaberlachG. T.et al (2003). Gene RB cloned from Solanum bulbocastanum confers broad spectrum resistance to potato late blight.Proc. Natl. Acad. Sci. U.S.A.1009128–913310.1073/pnas.1533501100
61
StajichJ. E.HarrisT.BrunkB. P.BrestelliJ.FischerS.HarbO. S.et al (2012). FungiDB: an integrated functional genomics database for fungi.Nucleic Acids Res.40D675–D681. 10.1093/nar/gkr918
62
StamatakisA. (2014). RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies.Bioinformatics301312–1313. 10.1093/bioinformatics/btu033
63
TagohH.HimesR.ClarkeD.LeenenP. J. M.RiggsA. D.HumeD.et al (2002). Transcription factor complex formation and chromatin fine structure alterations at the murine c-fms (CSF-1 receptor) locus during maturation of myeloid precursor cells.Genes Dev.161721–1737. 10.1101/gad.222002
64
TrapnellC.WilliamsB. A.PerteaG.MortazaviA.KwanG.van BarenM. J.et al (2010). Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation.Nat. Biotechnol.28511–515. 10.1038/nbt.1621
65
TurkensteenL. J.FlierW. G.WanningenR.MulderA. (2000). Production, survival and infectivity of oospores of Phytophthora infestans.Plant Pathol.49688–696. 10.1046/j.1365-3059.2000.00515.x
66
TzelepisG.MelinP.Jensen-FunckD.StenlidJ.KarlssonM. (2012). Functional analysis of glycoside hydrolase family 18 and 20 genes in Neurospora crassa.Fungal Genet. Biol.49717–730. 10.1016/j.fgb.2012.06.013
67
van der AuweraG. A.CarneiroM. O.HartlC.PoplinR.Del AngelG.Levy-MoonshineA.et al (2013). From FastQ data to high confidence variant calls: the genome analysis Toolkit best practices pipeline.Curr. Protoc. Bioinform.4311–33. 10.1002/0471250953.bi1110s43
68
van der LeeT.TestaA.RoboldA.van’t KloosterJ.GoversF. (2004). High-density genetic linkage maps of Phytophthora infestans reveal trisomic progeny and chromosomal rearrangements.Genetics1671643–1661. 10.1534/genetics.104.029652
69
van der VossenE. A.GrosJ.SikkemaA.MuskensM.WoutersD.WoltersP.et al (2005). The Rpi-blb2 gene from Solanum bulbocastanum is an Mi-1 gene homolog conferring broad-spectrum late blight resistance in potato.Plant J.44208–222. 10.1111/j.1365-313X.2005.02527.x
70
van WestP.ShepherdS. J.WalkerC. A.LiS.AppiahA. A.Grenville-BriggsL. J.et al (2008). Internuclear gene silencing in Phytophthora infestans is established through chromatin remodelling.Microbiology1541482–1490. 10.1099/mic.0.2007/015545-0 ?
71
VetukuriR. R.AsmanA. K. M.Tellgren-RothC.JahanS.N.ReimegardJ.FogelqvistJ.et al (2012). Evidence for small RNAs homologous to effector-encoding genes and transposable elements in the oomycete Phytophthora infestans.PLoS One7:e51399. 10.1371/journal.pone.0051399
72
VetukuriR. R.AvrovaA. O.Grenville-BriggsL. J.Van WestP.SöderbomF.SavenkovE. I.et al (2011). Evidence for involvement of dicer-like, argonaute and histone deacetylase proteins in gene silencing in Phytophthora infestans.Mol. Plant Pathol.12772–785. 10.1111/j.1364-3703.2011.00710.x
73
WawraS.TruschF.MatenaA.ApostolakisK.LinneU.ZhukovI.et al (2017). The RxLR motif of the host targeting effector AVR3a of Phytophthora infestans is cleaved before secretion.Plant Cell291184–1195. 10.1105/tpc.16.00552
74
WhissonS.VetukuriR.AvrovaA.DixeliusC. (2012). Can silencing of transposons contribute to variation in effector gene expression in Phytophthora infestans?Mob. Genet. Elem.2110–114. 10.4161/mge.20265
75
WibowoA.BeckerC.MarconiG.DurrJ.PriceJ.HagmannJ.et al (2016). Hyperosmotic stress memory in Arabidopsis is mediated by distinct epigenetic labile sites in the genome and is restricted in the male germline by DNA glycosylase activity.eLife5:13546. 10.7554/eLife.13546
Summary
Keywords
Avrblb2, epigenetics, heterothallism, mating, ploidy
Citation
Tzelepis G, Hodén KP, Fogelqvist J, Åsman AKM, Vetukuri RR and Dixelius C (2020) Dominance of Mating Type A1 and Indication of Epigenetic Effects During Early Stages of Mating in Phytophthora infestans. Front. Microbiol. 11:252. doi: 10.3389/fmicb.2020.00252
Received
16 September 2019
Accepted
03 February 2020
Published
21 February 2020
Volume
11 - 2020
Edited by
Raffaella Balestrini, Institute for Sustainable Plant Protection (CNR), Italy
Reviewed by
Kin-Ming (Clement) Tsui, Weill Cornell Medicine-Qatar, Qatar; Zheng Wang, Yale University, United States; Joseph Heitman, Duke University, United States
Updates
Copyright
© 2020 Tzelepis, Hodén, Fogelqvist, Åsman, Vetukuri and Dixelius.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Georgios Tzelepis, georgios.tzelepis@slu.se
†These authors have contributed equally to this work
‡Present address: Georgios Tzelepis, Department of Forest Mycology and Plant Pathology, Uppsala Biocenter, Swedish University of Agricultural Sciences, Uppsala, Sweden Anna K. M. Åsman, Department of Molecular Sciences, Uppsala Biocenter, Swedish University of Agricultural Sciences, Uppsala, Sweden Ramesh R. Vetukuri, Department of Plant Breeding, Swedish University of Agricultural Sciences, Alnarp, Sweden
This article was submitted to Fungi and Their Interactions, a section of the journal Frontiers in Microbiology
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.