- 1Institute of Technology, University of Tartu, Tartu, Estonia
- 2Department of Molecular Biology, Umeå University, Umeå, Sweden
- 3The Laboratory for Molecular Infection Medicine Sweden (MIMS), Umeå University, Umeå, Sweden
To appropriately switch between sessile and motile lifestyles, bacteria control expression of biofilm-associated genes through multiple regulatory elements. In Pseudomonas aeruginosa, the post-transcriptional regulator RsmA has been implicated in the control of various genes including those related to biofilms, but much of the evidence for these links is limited to transcriptomic and phenotypic studies. RsmA binds to target mRNAs to modulate translation by affecting ribosomal access and/or mRNA stability. Here, we trace a global regulatory role of RsmA to inhibition of the expression of Vfr—a transcription factor that inhibits transcriptional regulator FleQ. FleQ directly controls biofilm-associated genes that encode the PEL polysaccharide biosynthesis machinery. Furthermore, we show that RsmA alone cannot bind vfr mRNA but requires the assistance of RNA chaperone protein Hfq. This is the first example where a RsmA protein family member requires another protein for binding to its target RNA.
Introduction
The opportunistic pathogen Pseudomonas aeruginosa can be isolated from a wide range of environmental niches (Crone et al., 2020), in large part owing to its versatile metabolic capabilities (Arai, 2011). It is proficient in colonizing various eukaryotic organisms (Rahme et al., 1995) and can cause both acute and chronic infections, with the latter often associated with biofilm-like modes of growth at the sites of infection (Gellatly and Hancock, 2013). P. aeruginosa is highly competitive against other microbial species, with its adaptability and competitive fitness being aided by the production of various secondary metabolites and virulence factors (Tashiro et al., 2013; Khare and Tavazoie, 2015). Many of these cellular processes are inter-regulated by multiple regulatory pathways, presumably to provide appropriate response mechanisms to a wide range of environmental cues. One of the central regulators that determine P. aeruginosa behavior is the post-transcription factor RsmA (Regulator of Secondary Metabolites) as evidenced by its null mutant phenotypes linked to motility (Heurlier et al., 2004), virulence (Mulcahy et al., 2006, 2008; O’Grady et al., 2006), biofilm (Mulcahy et al., 2006; Irie et al., 2010), growth (Irie et al., 2010), and secreted products (Pessi et al., 2001; Heurlier et al., 2004).
RsmA belongs to the RsmA/CsrA family of dimeric RNA-binding proteins (Gutiérrez et al., 2005; Rife et al., 2005; Schubert et al., 2007). RsmA/CsrA homologs are found across a wide range of both Gram-negative and Gram-positive bacterial species: in some species including Escherichia coli, they are called CsrA, while in other species including P. aeruginosa, they are named RsmA (White et al., 1996). RsmA/CsrA proteins inhibit translation of target mRNAs by binding to the ribosome binding sites, typically overlapping either the Shine–Dalgarno sequence or the start codon (Romeo et al., 2013). However, in a few cases identified thus far, they can also serve as positive regulators of translation by altering the secondary structures of bound RNAs (Patterson-Fortin et al., 2013; Ren et al., 2014) or protecting RNAs from degradation (Yakhnin et al., 2013).
Many RsmA/CsrA-targeted mRNAs are reported to have altered RNA decay rates. Several studies attempted to exploit this property to determine the global P. aeruginosa RsmA regulon using microarrays (Lawhon et al., 2003; Burrowes et al., 2006; Brencic and Lory, 2009). However, this approach cannot distinguish directly regulated genes from indirectly regulated gene sets that result from cascade regulation, nor can it identify target RNAs that do not exhibit changes in their stability (Baker et al., 2007; Pannuri et al., 2012). In P. aeruginosa, direct RsmA-mediated translation inhibition has been demonstrated for the P. aeruginosa-specific biofilm polysaccharide operon psl without any effect on the level of transcription (Irie et al., 2010). Hence, the psl operon was not prominently featured in transcriptomic studies despite a strong biofilm phenotype (Burrowes et al., 2006; Brencic and Lory, 2009), raising concerns of what is currently considered to be “the RsmA regulon” of P. aeruginosa. To address this concern, more direct global analyses of RsmA/CsrA using techniques such as ChIP-Seq, CLIP-Seq, RIP-Seq, and ribosome profiling have recently been performed (Holmqvist et al., 2016; Potts et al., 2017; Sahr et al., 2017; Gebhardt et al., 2020).
Here, we present genetic and biochemical evidence that RsmA serves a master regulatory role through several intermediate transcription factors that are known to control different cellular processes. Through cascade regulation, originating from a direct effect on vfr mRNA, RsmA indirectly affects biofilm polysaccharide expression. Furthermore, we discovered an unexpected regulatory interplay between RsmA and the RNA chaperone protein Hfq on the target mRNA, whereby RsmA could only bind in the presence of Hfq. To our knowledge, this is the first example of RsmA/CsrA being unable to bind to its RNA target unless assisted by another protein.
Results
RsmA Is a Post-transcriptional Regulator of vfr Through Which fleQ and pel Transcripts Are Indirectly Regulated
Pseudomonas aeruginosa produces at least three different extracellular biofilm polysaccharides: alginate, PEL, and PSL (Ryder et al., 2007). PEL and PSL are co-regulated by several factors including the secondary messenger molecule c-di-GMP via the c-di-GMP-binding transcriptional regulator FleQ (Hickman and Harwood, 2008) and possibly also quorum sensing (Sakuragi and Kolter, 2007; Gilbert et al., 2009). Similar to PSL, PEL has long been considered to be regulated by RsmA (Gooderham and Hancock, 2009) based on a model originally proposed from a microarray study (Goodman et al., 2004). However, unlike the psl operon transcript, which was genetically and biochemically demonstrated to be directly regulated by RsmA (Irie et al., 2010), no direct evidence for the pel operon has been presented.
Consistent with a previous microarray study (Brencic and Lory, 2009), using a chromosomal transcriptional reporter, we found that pel is up-regulated in the ΔrsmA mutant as compared with the wild-type (WT) background (Figure 1A). However, the PSL polysaccharides are also over-produced in a ΔrsmA strain (Irie et al., 2010), resulting in elevated intracellular levels of c-di-GMP due to PSL signaling (Irie et al., 2012). Given that the pel genes are transcriptionally up-regulated by c-di-GMP (Hickman and Harwood, 2008; Baraquet et al., 2012) and the ΔrsmA strain has elevated c-di-GMP levels (Irie et al., 2012), it is plausible that the up-regulation of pel could be a consequence of increased c-di-GMP levels. To uncouple pel expression from elevated c-di-GMP, we also measured pel transcriptional reporter activities when introduced into a low c-di-GMP ΔrsmA Δpel Δpsl triple mutant background (Irie et al., 2012). In this strain, pel is still up-regulated (Figure 1A), indicating that the up-regulation is caused by lack of RsmA rather than changes in intracellular c-di-GMP concentration imposed by RsmA regulation of diguanylate cyclases and/or phosphodiesterases.
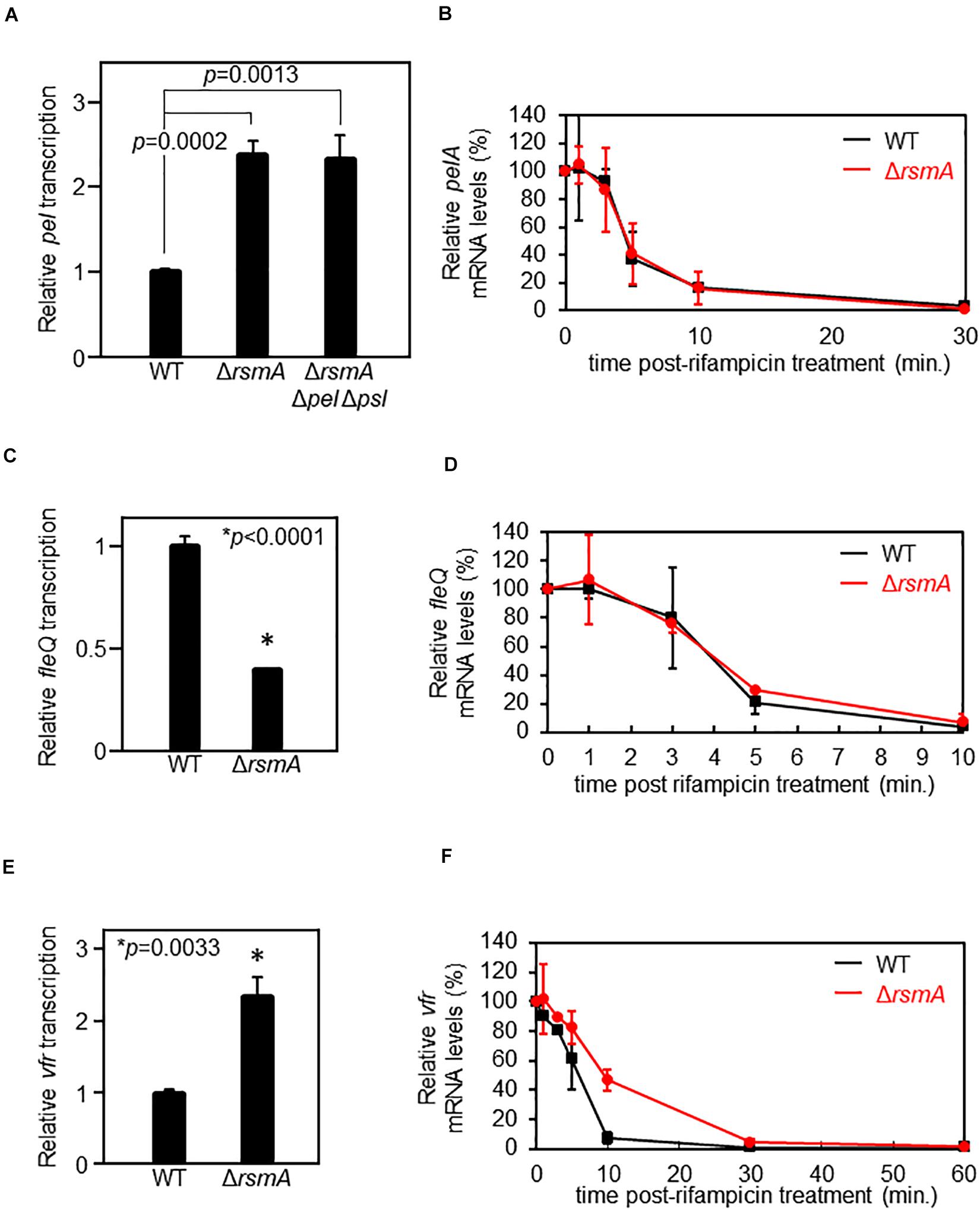
Figure 1. RsmA does not directly regulate pel nor fleQ but represses vfr by altering mRNA stability. (A) Single-copy transcriptional lacZ fusion constructs reveal up-regulation of pel transcripts in the ΔrsmA background as compared with wild type (WT). The ΔrsmA strain overexpresses the PSL polysaccharides, which causes an elevation of intercellular c-di-GMP (Irie et al., 2012). Thus, we also tested a ΔrsmA Δpel Δpsl strain, which has low c-di-GMP due to the absence of PSL (Irie et al., 2012). Note that pel is equally up-regulated in the ΔrsmA Δpel Δpsl strain, demonstrating that the pel transcriptional phenotype of ΔrsmA is PEL-, PSL-, and c-di-GMP-independent. The y-axis values are expressed as relative to WT as 1. (B) Real-time quantitative PCR (qPCR) at different time points after rifampicin treatment shows indistinguishable stabilities of the pelA transcript in the WT and ΔrsmA strains. The y-axis values are expressed as relative to t = 0 as 100%. (C) Single-copy transcriptional lacZ fusion constructs reveal down-regulation of fleQ transcripts in the ΔrsmA background as compared with WT. The y-axis values are expressed as relative to WT as 1. (D) qPCR at different time points after rifampicin treatment shows indistinguishable stabilities of the fleQ transcript in the WT and ΔrsmA strains. The y-axis values are expressed as relative to t = 0 as 100%. (E) Single-copy transcriptional lacZ fusion constructs reveal up-regulation of vfr transcripts in the ΔrsmA background as compared with WT. The y-axis values are expressed as relative to WT as 1. (F) qPCR at different time points after rifampicin treatment show that vfr mRNA is less stable in WT as compared with the ΔrsmA strain. The y-axis values are expressed as relative to t = 0 as 100%.
Since direct regulation of mRNA levels by RsmA/CsrA often acts via altered mRNA stability (Romeo et al., 2013), we tested whether the apparent pel transcriptional changes between the WT and ΔrsmA strains is due to altered pel mRNA stability. This, however, is not the case (Figure 1B), suggesting that the differences in pel expression are likely due to altered activity of the promoter. Because the RNA-binding protein RsmA is not a transcription factor, we tested the possibility of RsmA affecting expression of a known transcriptional regulator of pel, namely, FleQ. As a well-characterized transcriptional activator of pel (Hickman and Harwood, 2008; Baraquet et al., 2012), potential RsmA-mediated alterations in expression of FleQ would be anticipated to affect transcription of the pel genes. We found that a transcriptional reporter fusion of fleQ gave lower levels in the absence of RsmA (Figure 1C). This is consistent with previous data showing pel transcript levels are up-regulated in ΔfleQ backgrounds (Hickman and Harwood, 2008).
The data above could lend themselves to the interpretation that RsmA serves as a positive regulator of fleQ. While RsmA is more commonly known to be a translational repressor, this possibility is not without precedence. Examples of RsmA/CsrA proteins stimulating translation of target mRNAs include phz2 in P. aeruginosa (Ren et al., 2014) and moaA and flhDC in Escherichia coli (Patterson-Fortin et al., 2013; Yakhnin et al., 2013). While P. aeruginosa FleQ and E. coli FlhD4C2 share no sequence or mechanistic similarities, they are functional counterparts, with both being class I master regulators of flagellar biosynthesis in their respective organisms (Dasgupta et al., 2003). In addition, flagellar motility was previously found to be positively controlled by RsmA in P. aeruginosa (Heurlier et al., 2004), while in E. coli, CsrA has been shown to up-regulate flagellar gene expression through protection of flhDC mRNA from degradation (Yakhnin et al., 2013). It follows that if RsmA regulated fleQ in an analogous manner, fleQ mRNAs would be stabilized by RsmA and, thus, less stable in the null mutant. However, no evidence of altered mRNA turnover rate was found between WT and ΔrsmA strains (Figure 1D). Therefore, we hypothesized that the direct action of RsmA may lie even further upstream in a regulatory cascade that involved control of fleQ by Vfr.
Vfr acts as a transcriptional repressor of the fleQ promoter (Dasgupta et al., 2002). Therefore, to be consistent with the above phenotypes, RsmA would have to serve as a repressor of vfr. In line with this idea, comparison of vfr transcriptional reporter activities in WT and ΔrsmA strains revealed up-regulation of vfr transcripts in the absence of RsmA (Figure 1E and Supplementary Figure S1). In contrast to pel (Figure 1B) and fleQ (Figure 1D), the transcript stability of vfr is significantly different between WT and ΔrsmA strains, such that vfr mRNAs are more stable in the absence of RsmA (Figure 1F). These results indicate that RsmA affects vfr mRNA levels by mediating changes in turnover rates.
RsmA-Mediated Regulation of Vfr Alters Swimming and Twitching Motility
Pseudomonas aeruginosa uses at least two major modes of motility: flagellar-driven swimming motility through liquid and surface “twitching” motility using Type IV pilus (Burrows, 2012; Kazmierczak et al., 2015). Bacterial migration is an important aspect during P. aeruginosa biofilm development, as initial attachment to a surface is thought to require flagella, and mature microcolony formation has been attributed to Type IV pilus functions (O’Toole and Kolter, 1998; Klausen et al., 2003). Thus, fine-tuned regulatory control of motility and biofilm genes is a necessity for effective P. aeruginosa adaptation, particularly when switching between motile and sessile lifestyles.
Because FleQ is a class I master regulator of flagellar genes (Dasgupta et al., 2003) and RsmA and Vfr lie upstream of FleQ in the regulatory cascade, lack of either protein should predictively impact flagellar motility, but in opposite ways. As would be predicted, ΔrsmA and Vfr over-expressing strains have reduced flagellar (swimming) motilities (Figures 2A,B). Swimming motility level of Δvfr is comparable with that of the WT (Figure 2A) probably because de-repression of flagellar genes does not necessarily lead to functional hyper-motility. There was no additive effect of over-expressing Vfr in the ΔrsmA background (Figure 2C), consistent with our model of a direct regulatory cascade from RsmA ⊣ Vfr ⊣ FleQ. Although expression of both the PEL (Figure 1A) and PSL (Irie et al., 2010) polysaccharides are up-regulated in ΔrsmA, expression of these two polysaccharides did not affect flagellar motility significantly (Supplementary Figure S2).
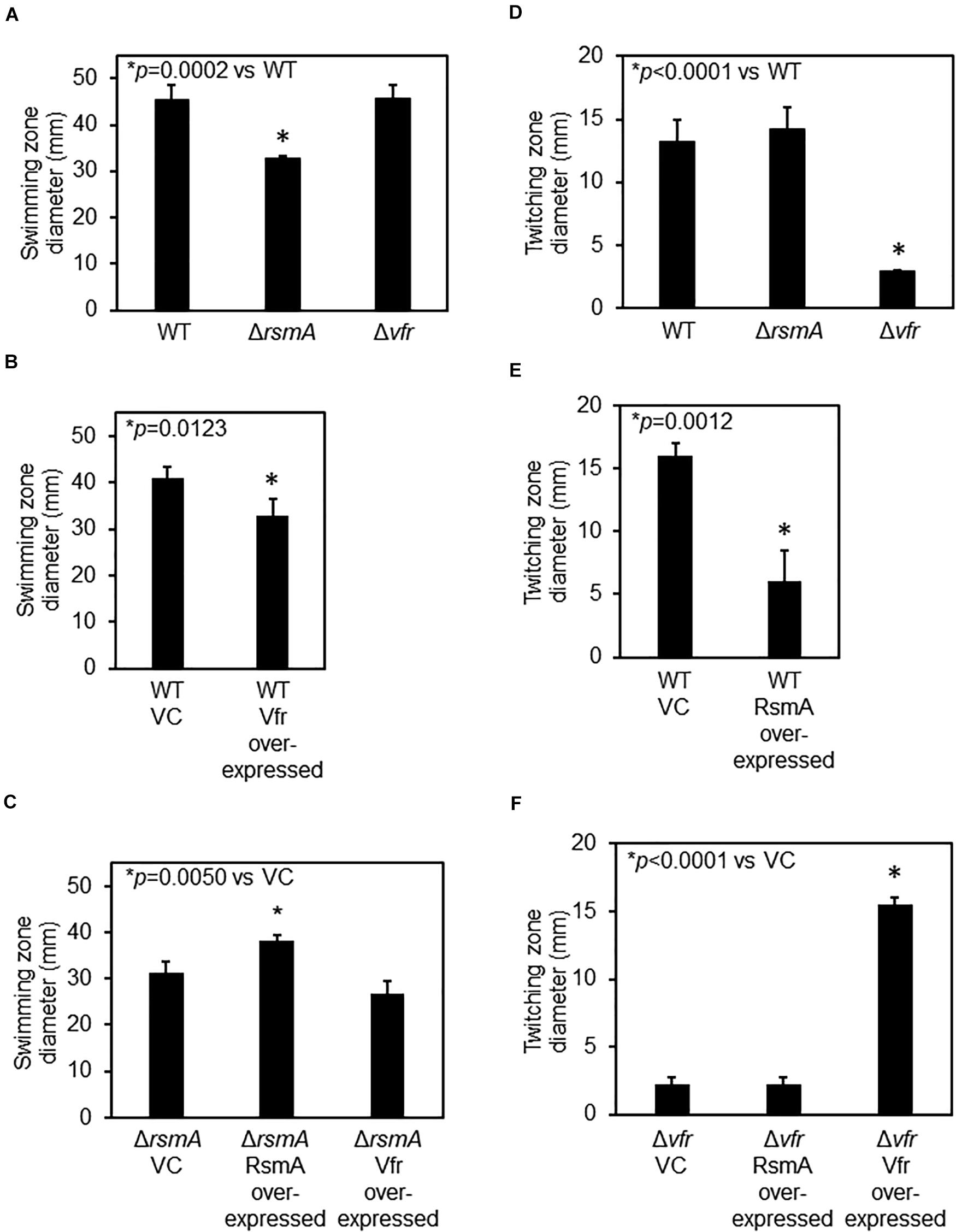
Figure 2. RsmA regulatory cascade affects flagellar and Type IV pilus-mediated motilities. (A) Flagellar-mediated swimming motility is decreased for the ΔrsmA strain but not the Δvfr strain. (B) Vfr over-expression leads to decreased swimming motility. VC, vector control. Motility assays for VC and Vfr over-expressing strains were performed in the presence of 300 μg⋅ ml–1 of carbenicillin to ensure maintenance of the plasmids. (C) There is no statistical significance difference between the swimming motility of ΔrsmA and Vfr over-expression strains, indicating that Vfr is epistatically downstream of and in the same regulatory pathway as RsmA. Motility assays for these plasmid-containing strains were performed in the presence of 300 μg⋅ ml–1 of carbenicillin. (D) Type IV pilus-mediated twitching motility is reduced for Δvfr as compared with wild type (WT). (E) RsmA over-expression leads to decreased twitching motility. Motility assays for VC and the RsmA over-expressing strain were performed in the presence of 300 μg⋅ ml–1 of carbenicillin. (F) RsmA over-expression in Δvfr does not affect the low twitching motility of the parental Δvfr. Motility assays for these plasmid-containing strains were performed in the presence of 300 μg⋅ ml–1 of carbenicillin.
Mutants of vfr have previously been documented to be defective in Type IV pilus-dependent twitching motility (Whitchurch et al., 1996; Beatson et al., 2002). This is due to the positive regulation of the AlgZR two-component system by Vfr (Luo et al., 2015; Pritchett et al., 2015), which is required for the expression of Type IV pilus genes (Lizewski et al., 2004; Belete et al., 2008). As shown in Figures 2D,E, both the Δvfr and RsmA over-expressing strains exhibited defects in twitching motility. Twitching motility of the ΔrsmA strain was comparable with that of WT. The latter is probably because, like de-repression of flagellar genes does not necessarily lead to functional hyper-swimming motility, de-repression of pilin genes also does not necessarily lead to functional hyper-twitching motility. Over-expressing RsmA in the Δvfr background had no effect (Figure 2F) due to RsmA lying upstream of Vfr.
Loss of RsmA Results in an Increased Acute Virulence Phenotype
Vfr is a cAMP-binding transcription factor that positively regulates acute virulence factors, including the Type III secretion system (Wolfgang et al., 2003). Vfr activates expression of the transcription factor ExsA, which in turn is required for the expression of the Type III secretion machinery (Marsden et al., 2016). In light of the data in Figures 1, 2, suggesting that RsmA is a direct negative regulator of vfr expression, it became of interest to test if the ΔrsmA mutant had enhanced virulence against eukaryotic cells due to increased expression of Vfr and therefore activation of virulence factors. To test this idea, we monitored cytotoxicity against eukaryotic cells by measuring lactate dehydrogenase (LDH) activity in the supernatants. LDH is a eukaryotic cytoplasmic protein and is released into the supernatant upon cell death/lysis. After 24-hour infection of human retinal pigment epithelial (RPE) cells, we observed increased cytotoxicity toward the RPE cells that were infected with ΔrsmA as compared with WT (Figure 3A). On the other hand, Δvfr was attenuated in this respect as would be expected (Wolfgang et al., 2003). This pathogenicity phenotype was independent of bacterial load since equal numbers of bacteria were recovered from the same supernatant for WT, ΔrsmA, and Δvfr (Figure 3B). The ΔrsmA cytotoxicity phenotype is consistent with a previous report (Mulcahy et al., 2006) and further supports the idea that effects of RsmA through Vfr are propagated to downstream processes in regulatory cascades.
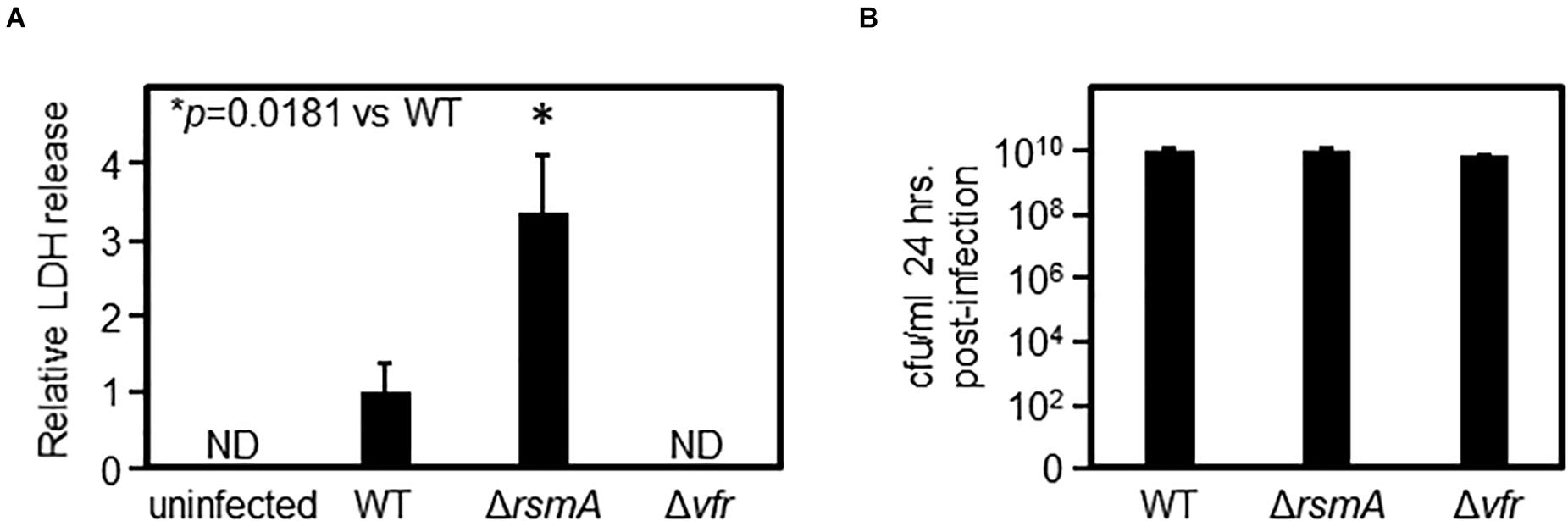
Figure 3. ΔrsmA displays an increased virulence phenotype against tissue culture cells. (A) After retinal pigment epithelial (RPE) cells were infected by Pseudomonas aeruginosa strains for 24 h, the levels of lactate dehydrogenase (LDH) in the media were measured. The increase of LDH released into the media by ΔrsmA-infected cells as compared with WT-infected cells indicates increased RPE cell lysis and, therefore, increased cytotoxicity caused by the ΔrsmA strain. Δvfr is defective in this virulence assay, as no LDH could be detected in the media. The y-axis values are expressed as relative to WT as 1. ND, not detected. (B) Bacteria were retrieved from the wells of infected RPE cells, plated on agar media from a serial dilution series, and subsequently enumerated by counting colonies to determine colony-forming units (cfu) per milliliter. The identical cfu⋅ ml–1 for all three strains demonstrate that the differences in virulence as measured by LDH release in panel (A) is not due to an imbalance in bacterial loads.
Hfq Is Required for RsmA to Bind to vfr mRNA
Given the evidence that RsmA may post-transcriptionally regulate vfr, we next examined the binding properties of RsmA to vfr mRNA. P. aeruginosa RsmA has a binding specificity for the CANGGAYG consensus sequence of its target mRNA (Schulmeyer et al., 2016), which is similar to the E. coli CsrA consensus sequences RUACARGGAUGU generated by SELEX (Dubey et al., 2005) and AUGGAUG generated by CLIP-Seq (Potts et al., 2017). The transcriptional start site for vfr lies 153 bases upstream of the translational start site (Fuchs et al., 2010). Inspection of the vfr leader sequence identified one putative RsmA-binding site overlapping the likely GGGA Shine–Dalgarno sequence (Figure 4A).
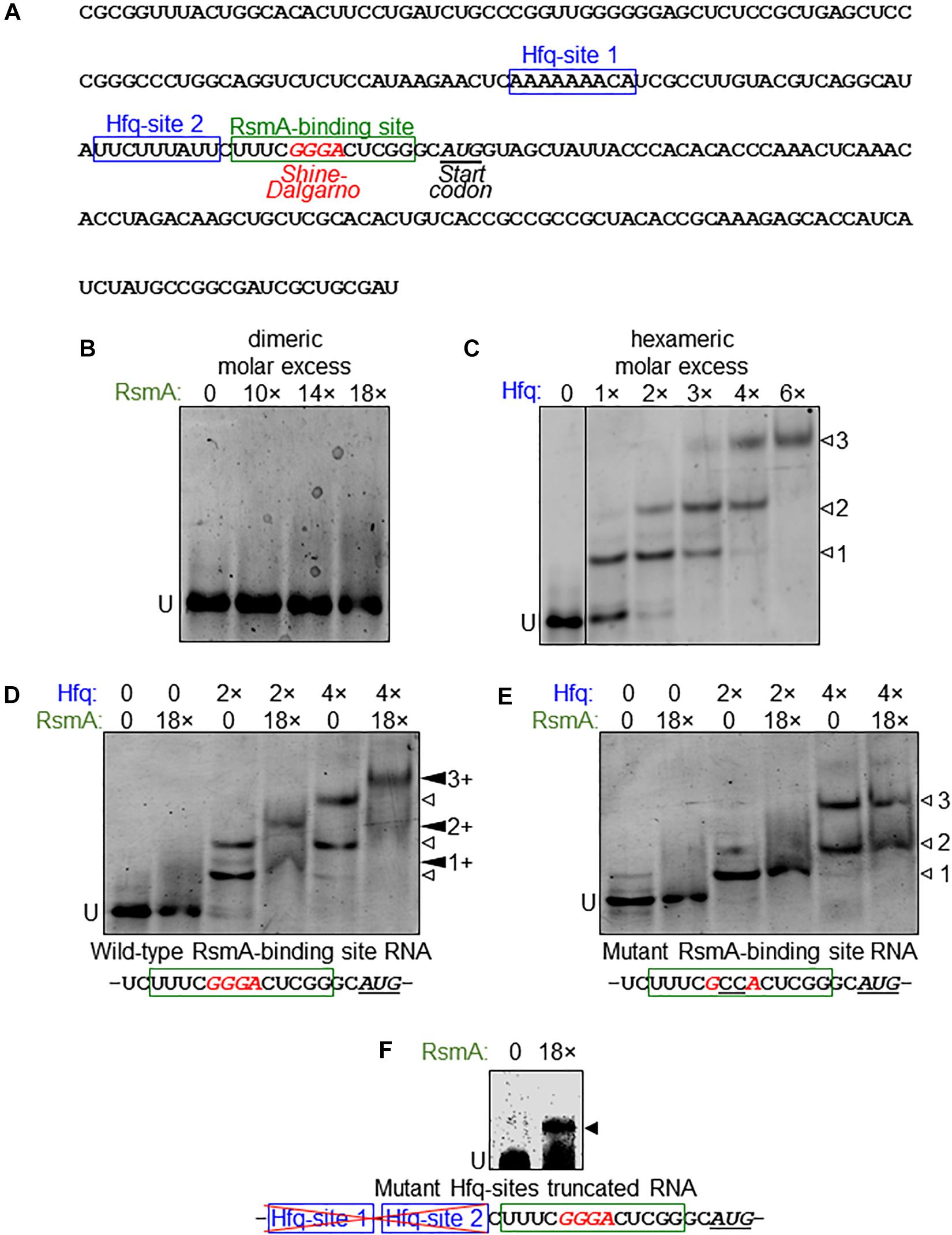
Figure 4. Direct binding of vfr mRNA by RsmA requires Hfq. (A) Sequence of the vfr RNA used in electrophoretic mobility shift assay (EMSA) experiments are shown, spanning from the transcriptional start and partially through the vfr open reading frame (ORF). The leader sequence of the vfr transcript (Fuchs et al., 2010) possesses two putative Hfq-binding sites (blue boxes) and a RsmA-binding site (green box) overlapping the predicted Shine–Dalgarno sequence (red italics) upstream of the translational start site (underlined italics). (B) RNA EMSA shows that RsmA does not bind to the vfr RNA when RsmA is present alone. Dimeric molar excess relative to RNA; U, unbound RNA. (C) The vfr RNA produces three distinct band shifts (open arrowheads 1, 2, and 3) with increasing concentrations of Hfq. Hexameric molar excess relative to RNA. (D) The presence of RsmA causes all Hfq-shifted bands to super-shift (filled arrowheads 1+, 2+, and 3+), indicating that RsmA binds to Hfq-bound RNA. Note that the RsmA concentration used (18 molar excess of RNA) is incapable of producing a shift when present alone (B). (E) A GG→CC substitution within the RsmA-binding site abolishes RsmA super-shifts but does not alter Hfq binding. (F) RsmA can bind to the truncated form of the vfr RNA that lacks the region immediately upstream of the RsmA-binding site that encompasses the two Hfq-binding sites. Note that this indicates removal of an inhibitory structure that otherwise blocks RsmA binding unless counteracted by binding of Hfq.
Purified RsmA protein directly binds psl mRNA (Irie et al., 2010) in RNA electrophoretic mobility shift assays (EMSAs). However, no RsmA binding was observed in similar assays with vfr RNA spanning the leader sequence (Figure 4B). This result led us to consider the possibility that RsmA may require another factor to bind vfr mRNA. The RNA-binding chaperone protein Hfq was raised as a candidate for three reasons. First, the small RNA RsmY, which binds multiple RsmA proteins at high affinity to sequester and relieve RsmA-bound mRNAs (Kay et al., 2006), also has the capacity to be bound by Hfq (Sorger-Domenigg et al., 2007), although it was unclear from that study whether co-binding of both proteins occurred. Second, Hfq was identified to bind to vfr mRNA by global ChIP-Seq analyses (Kambara et al., 2018). Third, in silico analysis of the vfr leader sequence revealed two potential Hfq binding sites upstream of the putative RsmA-binding site (Figure 4A).
Biochemical studies, primarily of E. coli Hfq, have shown that this hexameric protein complex has at least four regions, which can all be involved in RNA-binding: the proximal face, the distal face, the rim, and the C-terminal tail (Updegrove et al., 2016). P. aeruginosa and Pseudomonas putida Hfq have shortened C-termini compared with E. coli Hfq and may lack C-terminal tail-binding altogether (Milojevic et al., 2013). The proximal site preferentially binds to U-rich RNA sequence (Møller et al., 2002), the distal site binds to A-rich (ARN)n triplet repeats (Link et al., 2009), and the rim associates with UA-rich regions (Schu et al., 2015). The two potential Hfq-binding sites found within the vfr leader sequence (Figure 4A) represent one A-rich ARN repeat region and one U-rich region.
RNA EMSA analyses show that P. aeruginosa Hfq binds vfr RNA, resulting in three distinct band shifts (Figure 4C) indicative of two or more binding sites. Because WT P. putida Hfq showed identical binding patterns to P. aeruginosa Hfq on vfr RNA (Supplementary Figure S3A), we took advantage of previously characterized derivatives of P. putida Hfq, namely, a distal site mutant (HfqY25D; A-rich ARN-binding deficient) and a proximal site mutant (HfqK56A; U-rich-binding deficient) (Madhushani et al., 2015) to further analyze Hfq binding to the vfr RNA. As seen in Supplementary Figure S3B, HfqY25D only recapitulates the second band shift, while HfqK56A only recapitulates the first band shift. Notably, the third band shift is absent with both mutant derivatives of Hfq. We conclude that the first shift is caused when the ARN repeats are bound by Hfq (Hfq-site 1 in Figure 4A), the second shift is caused by Hfq binding to the U-rich region of the vfr leader sequence (Hfq-site 2 in Figure 4A), and the third shift occurs when both sites are occupied simultaneously.
Having ascertained that Hfq does bind vfr RNA, we next added RsmA and Hfq simultaneously to the vfr RNA. As seen in Figure 4D, this results in super-shifting of all three Hfq band shifts in the presence of RsmA, suggesting that binding of Hfq to either of its target sites is sufficient to allow RsmA binding. Because the GG doublet within the consensus sequences for RsmA and CsrA binding is essential for their abilities to bind cognate target RNAs (Dubey et al., 2005; Irie et al., 2010), we made a CC substitution of the GG doublet within the predicted RsmA-binding region of vfr RNA. No RsmA-mediated super-shifting of Hfq band shifts was detected with the CC substituted RNA (Figure 4E).
We infer from the findings above that Hfq binding is a prerequisite for RsmA to bind to its target within vfr mRNA. As expanded upon in the Discussion, such a prerequisite is indicative of an inhibitory secondary mRNA structure that blocks RsmA binding in the absence of Hfq binding. To test this idea, we generated a truncated version of the vfr RNA that lacks the upstream region that encompasses the two Hfq sites and thus does not bind Hfq (Supplementary Figure S4). RsmA can bind to the truncated version of vfr RNA independent of Hfq (Figure 4F). Together with the data in preceding sections, these results lead us to conclude that (1) the direct binding of RsmA on vfr RNA is prevented by a mRNA structure that can be counteracted by binding of the RNA chaperone protein Hfq, and (2) RsmA is an indirect regulator of biofilm polysaccharide locus pel and motility, through its Hfq-assisted action on vfr propagated through a regulatory cascade from Vfr to FleQ.
Loss of Hfq Abolishes vfr Expression
Based on our biochemical findings, we rationalized that since Hfq enables RsmA to bind to vfr mRNA, Hfq would serve as a repressor of vfr. The prediction from this model is that vfr expression would be up-regulated in a Δhfq mutant. We therefore constructed an in-frame deletion mutant of hfq. Because Hfq (like RsmA) is a global regulator, its absence can cause pleotropic effects. As previously reported by other laboratories (Sonnleitner et al., 2003; Hill et al., 2019), lack of Hfq in PAO1 causes growth defects (Supplementary Figure S5A). These defects are complemented by WT Hfq (Supplementary Figure S5B). Upon measurement of vfr transcriptional activities of equivalent exponential phase cultures, we found that contrary to initial expectations, lack of Hfq led to a complete abolition of vfr expression (Figure 5A and Supplementary Figure S6). We attribute this lack of vfr expression in the Hfq null strain to be the result of likely indirect effects on translation and RNA stability. We propose that Hfq enables RsmA to bind and inhibit vfr at the post-transcriptional level as depicted in Figures 5B,C. Within this model, Hfq is required to disrupt a large inhibitory stem-loop structure formed through complementary sequences that encompass the A-rich Hfq distal binding site and the U-rich Hfq proximal binding site. This large stem loop also blocks ribosome access to the Shine–Dalgarno sequence due to dsRNA formation at the base of the structure. The lack of Hfq in Δhfq strain would thereby result in an inability of the stem-loop structure to release the ribosome binding site for translation, irrespective of the presence or absence of RsmA.
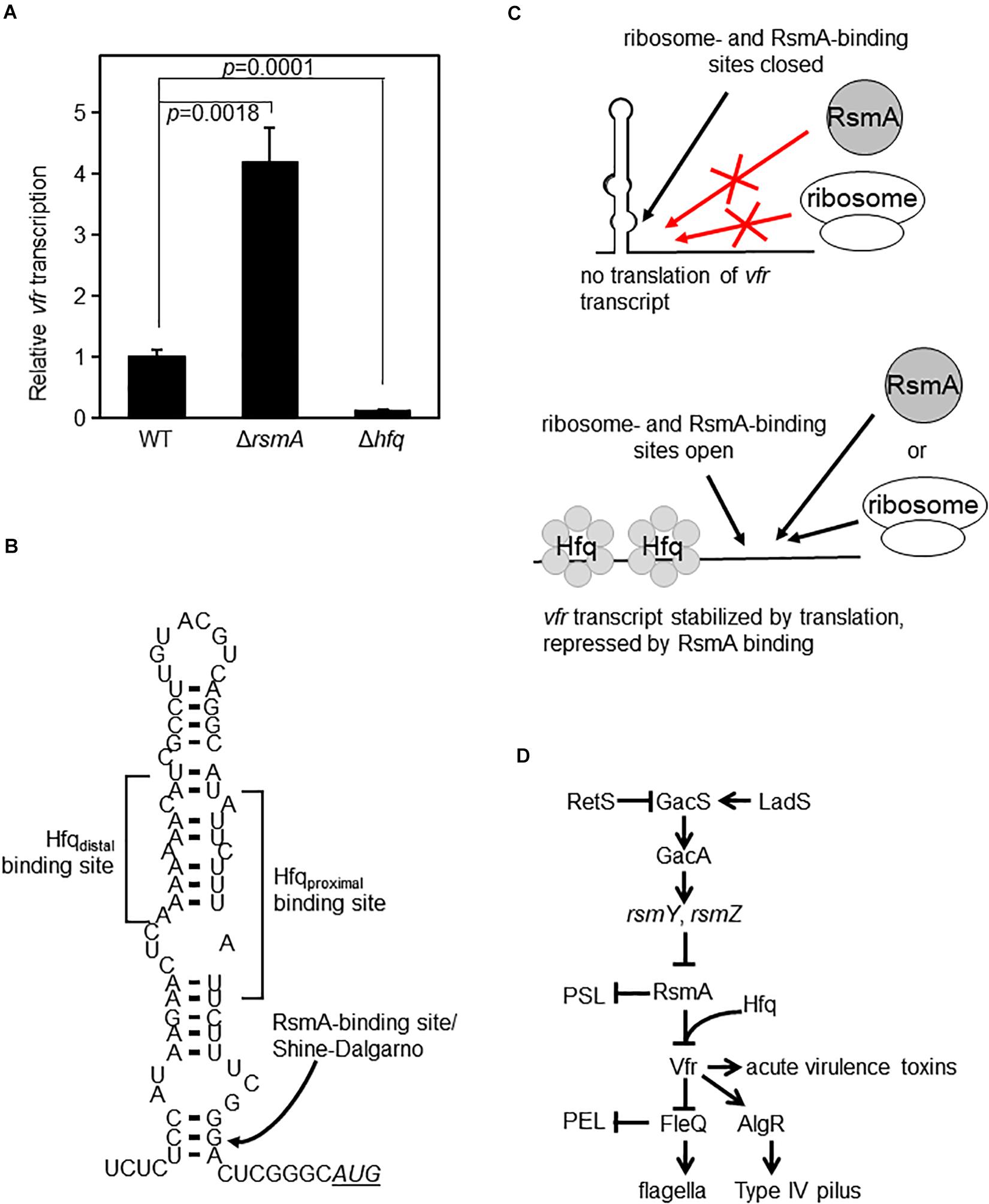
Figure 5. Hfq and RsmA affect vfr expression. (A) Single-copy transcriptional lacZ fusion constructs show that vfr transcript levels are increased in the ΔrsmA background (as in Figure 1C) but are essentially abolished in the Δhfq mutant background. The y-axis values are expressed as relative to WT as 1. (B) RNA secondary structure of the leader sequence region of vfr as predicted by mFOLD (Zuker, 1989, 2003). The locations of in silico-detected Hfq- and RsmA-binding sites are indicated. Note that the RsmA-binding site located at the base of the stem-loop structure overlaps with the Shine–Dalgarno sequence of the ribosome binding site. The AUG start codon is shown in underlined italics. (C) Schematic illustration of a model for Hfq- and RsmA-mediated regulation of vfr. In the absence of Hfq, the stem-loop structure shown in panel (B) blocks binding of both RsmA and ribosomes, resulting in a decrease of vfr expression as seen in panel (A). Upon Hfq binding of vfr RNA, the stem-loop structure shown in panel (B) is disrupted, exposing the ribosome- and RsmA-binding sites. RsmA-binding then directly blocks the Shine–Dalgarno sequence, preventing ribosome access and thereby inhibiting translation. In vivo, the untranslated vfr mRNAs are presumed to be subsequently rapidly processed for degradation. (D) Simplified overview of the Gac/Rsm pathway that controls RsmA levels and pertinent parts of its downstream regulon.
Discussion
In this study, we report that Pseudomonas aeruginosa RsmA requires the RNA chaperone Hfq to assist its binding to vfr mRNA to initiate a regulatory cascade that ultimately impacts pel biofilm polysaccharide genes. The requirement for Hfq was unexpected, since all other biochemical analyses of RsmA/CsrA family members document unaided binding to their target RNAs (Baker et al., 2002; Dubey et al., 2003; Wang et al., 2005; Lapouge et al., 2007; Irie et al., 2010; Patterson-Fortin et al., 2013; Yakhnin et al., 2013; Ren et al., 2014).
We identified two independent Hfq-binding sites upstream of the RsmA-binding motif, which overlaps the ribosome-binding site of vfr (Figure 4A). Based on a mFOLD-predicted secondary structure (Zuker, 1989, 2003), a large hairpin loop base-pairs the identified U-rich and A-rich Hfq-binding sites, resulting in a dsRNA structure that would predictively block both RsmA and the ribosome from accessing the mRNA (Figure 5B). As depicted in Figure 5C, the simplest model that we propose from our findings is that Hfq binding is required to open the stem loop to expose the RsmA-binding site. Subsequent binding of RsmA would directly block the Shine–Dalgarno sequence, preventing ribosome access and thereby inhibiting translation. We also provide evidence that vfr mRNA is more stable in the absence of RsmA (Figure 1F), indicating that Hfq- and RsmA-bound vfr mRNA may undergo more rapid degradation. mRNAs that are not bound by ribosomes are known to be degraded more rapidly (Deneke et al., 2013). Alternatively and not mutually exclusively, it was previously suggested that Hfq may be able to directly recruit RNase E for the degradation of bound RNA in Escherichia coli (Morita et al., 2005), and it is possible that a similar mechanism contributes to changes in vfr mRNA stability.
Due to its histidine-rich C-terminus, E. coli Hfq is a common contaminant of His-tagged proteins purified by nickel affinity chromatography after over-expression in E. coli, and visually undetectable [on sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE)] Hfq levels can critically influence in vitro analyses of RNA binding (Milojevic et al., 2013; Moreno et al., 2015). Because the proteins used in this study were purified after over-expressing them in either P. aeruginosa (RsmA) or a Hfq null strain of E. coli (Hfq proteins), our analyses were not complicated by this issue. However, a large majority of analyses performed in past publications use C-terminal His-tagged RsmA/CsrA proteins derived from over-expression in E. coli, raising the possibility of E. coli native Hfq contamination. Given that Hfq binds vfr RNA with apparent high affinity (Figure 4C and Supplementary Figure S3), it is plausible that Hfq-assisted RsmA binding may be prevalent. Although such a possibility remains to be experimentally verified, it provokes the observation that comparison of P. aeruginosa transcriptomic studies done for RsmA, Hfq, and Vfr regulons identifies numerous overlaps of genes that were differentially expressed between WT and the respective null mutants (Wolfgang et al., 2003; Goodman et al., 2004; Burrowes et al., 2006; Brencic and Lory, 2009; Sonnleitner et al., 2018). Overlaps were also seen between the Hfq and CsrA regulons of E. coli (Potts et al., 2017), suggesting that the interplay between the two RNA-binding regulators is not unique to P. aeruginosa.
A diverse range of phenotypes have been associated with RsmA functions in P. aeruginosa (Vakulskas et al., 2015), but phenotypic observations do not distinguish direct regulatory activities from indirect effects through regulatory cascades. Our identification of Hfq and RsmA action on vfr mRNA resulted from an initial aim to determine whether RsmA directly targeted pel RNA, as is the case for psl RNA (Irie et al., 2010). Our backtracking to trace the ultimate cause of RsmA effects on pel expression revealed a regulatory pathway originating from Vfr to the transcriptional regulator FleQ and from there to pel (Figure 5D).
This work places the global post-transcriptional regulators RsmA and Hfq as a hub at the center of upstream and downstream regulatory pathways and thus provides a conceptually new framework to evaluate and interpret past work. The two-component GacS/GacA system lies at the top of the hierarchy that controls RsmA-mediated regulation. Although the environmental cue for activation is still unknown, GacS activity is modulated through interactions with RetS and LadS (Goodman et al., 2009; Kong et al., 2013; Chambonnier et al., 2016). GacA, in turn, controls the production of the small RNAs—rsmY and rsmZ—which possess multiple high-affinity binding sites to titrate RsmA away from target mRNAs (Heurlier et al., 2004; Sonnleitner et al., 2006). In light of a report indicating that Hfq also binds to rsmY (Sorger-Domenigg et al., 2007), it appears plausible that these small RNAs may also require Hfq to assist their functions.
In some previous studies, RsmA was proposed to be a positive regulator of vfr due to decreased cytotoxicity in ΔrsmA mutants (Marden et al., 2013). A subsequent study proposes that Hfq assists sRNAs to inhibit RsmA activity, resulting in increased cytotoxicity in Δhfq mutants since RsmA is de-repressed in this model (Janssen et al., 2020). Our experiments show that RsmA negatively regulates vfr post-transcriptionally. To ensure that our phenotypes further correlate with our model, we performed cytotoxicity assays on tissue culture cells, and in our hands, we found ΔrsmA mutant to have increased cytotoxicity toward tissue culture cells than WT PAO1 (Figure 3), and vfr expression is abolished in Δhfq (Figure 5A). It is unclear why our results are incongruous with some publications, however, it should be noted that in the studies that observe decreased cytotoxicity with a ΔrsmA strain, the P. aeruginosa background is PA103 whereas this study uses PAO1. Our data are consistent with a publication from another group, which reported that ΔrsmA is more cytotoxic in the PAO1 background (Mulcahy et al., 2006). PA103 is frequently used for acute infection studies due to its high toxigenicity (Liu, 1966), and it is possible that its regulatory networks are wired differently from PAO1 due to its high adaptation toward pathogenicity. The precise roles of sRNAs and possible strain-to-strain differential expression patterns in the context of Hfq and RsmA remain to be investigated further.
Numerous processes are associated with RsmA in P. aeruginosa, including biofilm formation, motility, and acute virulence (Mulcahy et al., 2006; Intile et al., 2014). Many of these “RsmA-regulated factors and phenotypes” are likely to be indirectly regulated through Vfr- and/or FleQ-initiated regulatory cascades. Based on our findings, it will be crucial to determine and distinguish direct versus indirect regulatory routes to gain a greater understanding of the RsmA regulon.
Materials and Methods
Bacterial Strains and Growth Conditions
Supplementary Table S1 lists the bacterial strains used in this study. Escherichia coli and Pseudomonas aeruginosa strains were grown in lysogeny broth (LB) Lennox composition at 37°C unless specified otherwise. VBMM citrate medium (Hoang and Schweizer, 1997) was used for selecting P. aeruginosa post-conjugation. For E. coli, the following antibiotics concentrations were used: 50 μg⋅ ml–1 of carbenicillin, 10 μg⋅ ml–1 of gentamicin, and 10 μg⋅ ml–1 of tetracycline. For P. aeruginosa strains, 300 μg⋅ ml–1 of carbenicillin, 100 μg⋅ ml–1 of gentamicin, and 100 μg⋅ ml–1 of tetracycline concentrations were used. Sucrose counter-selection for plasmids carrying the sacB gene used in P. aeruginosa strain constructions was performed by streaking colonies on LB agar (no salt) supplemented with 10% w/v sucrose. Plates were incubated at 30°C for 24 h, after which the counter-selected colonies were confirmed for the loss of antibiotic resistance and mutations confirmed by PCR for double-crossover genomic mutants.
Strain Constructions
Transcriptional Fusion Constructions
β-Galactosidase transcriptional fusion constructs were generated in a single copy on the chromosome of P. aeruginosa strains via integration into the chromosomal attB site as previously published (Irie et al., 2010). In brief, promoter regions were PCR amplified using oligonucleotides in Supplementary Table S2. PCR products were ligated between the EcoRI and BamHI sites of mini-CTX lacZ (Supplementary Figure S7). These plasmids were then introduced into P. aeruginosa by conjugation. After double site recombination, plasmid backbones were removed by FLP recombinase, and then the strains were cured of the pFLP2 plasmid by sucrose counter-selection (Hoang et al., 2000).
In-Frame hfq Mutant Construction
Using the same background strain is important in P. aeruginosa research, since there is great heterogeneity between PAO1 strains of different laboratories (Klockgether et al., 2010; Chandler et al., 2019). Therefore, we generated Δhfq derivatives in the same PAO1 background as used in the corresponding author’s studies for the past 14 years.
To make an in-frame deletion mutant of hfq in different genetic backgrounds, we first attempted to use the suicide plasmid pME3087Δhfq (Sonnleitner et al., 2017). However, even after repeated attempts, we were unable to generate the mutant in a ΔrsmA background strains, which are already growth defective (Irie et al., 2010). Therefore, we constructed a Δhfq construct carried on the suicide vector pEX18Gm that has previously successfully been used in ΔrsmA strains (Irie et al., 2010) and has the additional advantage of the plasmid containing a gentamicin resistance cassette instead of tetracycline (Hoang et al., 1998). We used pME3087Δhfq as the template to create pEX18Gm:Δhfq to keep our resulting Δhfq mutant constructs consistent with the previous publications. In brief, following the circular polymerase extension cloning (CPEC) protocol (Quan and Tian, 2009, 2011), we first PCR amplified the Δhfq fragment using the primer pair CPEC pEX18Gm H71 and CPEC pEX18Gm I71 (Supplementary Table S2) with pME3087Δhfq as template (70°C annealing temperature, 20-second extension time). The plasmid backbone fragment was amplified using the primer pair CPEC pEX18Gm BamHI and CPEC pEX18Gm EcoRI (Supplementary Table S2) with pEX18Gm as template (71.6°C annealing temperature, 60-second extension time). The two fragments were fused and circularized by the second round of PCR that uses a slow ramp anneal of 70 to 55°C at 0.1°C⋅ s–1. The product was transformed into DH5α, and the correct configuration of the plasmid was confirmed prior to introduction into S17-1 λpir for conjugation into P. aeruginosa. Single-site recombinants were selected on gentamicin VBMM plates. Sucrose plates were used for counter-selection of second-site recombinants. The resulting colonies were screened by PCR using H71 and I71 primer pairs (Supplementary Table S2), where half resulted in WT alleles and half were Δhfq. This screening was facilitated by the fact that Δhfq mutants always formed small colonies due to their growth deficiencies (Supplementary Figure S5).
Vfr Over-Expression Construct
Vfr over-expression plasmid pUCP18:vfr (Supplementary Figure S8) was constructed by ligating EcoRI/BamHI double-digested pUCP18 shuttle vector (Schweizer, 1991) and EcoRI/BamHI digested PCR product of primer pairs vfr for1 and vfr rev1 (Supplementary Table S2) generated using PAO1 genomic DNA as template. The verified plasmid was transformed into P. aeruginosa by electroporation (Choi et al., 2006).
Hfq Over-Expression Strains
Hfq over-expression strains were made by introducing pME4510hfqFlag (Sonnleitner et al., 2017) into P. aeruginosa by electroporation (Choi et al., 2006).
Growth Curves
Pseudomonas aeruginosa strains were first grown as shaken LB liquid cultures overnight and then diluted to A600 ≈ 0.02. Diluted bacteria were deposited into the wells of 96-well plates in triplicates. Growths were monitored in a SynergyTM Mx microplate reader (BioTek) with the chamber temperature at 37°C and maximum plate orbital shaking/agitation speed setting. A600 measurements were recorded every 15 min.
β-Galactosidase Assays
Quantitative β-galactosidase activities were assayed using Galacto-Light Plus kit (Thermo-Fisher) as previously published (Lequette et al., 2006). P. aeruginosa cultures were grown at 37°C to exponential phase and lysed using chloroform as previously described (Irie et al., 2010). β-Galactosidase activity units were normalized to total proteins per milliliter as determined using Bradford assay reagents (Bio-Rad). Assays were performed in biological triplicates each in technical triplicates.
Quantitative Real-Time PCR and RNA Stability Analyses
Real-time PCR was performed as previously described (Irie et al., 2012), using the oligonucleotides listed in Supplementary Table S2. For RNA stability experiments, exponential phase P. aeruginosa cultured in VBMM citrate at 37°C were treated with 200 μg⋅ ml–1 of rifampicin (Lory, 1986). RNAs were extracted from 1 ml of the cultures at various time points as previously described (Irie et al., 2010) after the addition of rifampicin. RNA extractions were performed using the RNeasy Mini Kit (Qiagen) after treating the harvested cells with RNAprotect (Qiagen). Genomic DNA was removed using DNase I (Promega), and removal confirmed by PCR using primers designed against the rplU gene (Supplementary Table S2). SuperScript III First-Strand Synthesis (Invitrogen) with random hexamers was used to synthesize cDNA as per manufacturer’s protocol. Quantitative real-time PCR was performed using SYBR Green PCR Master Mix (Thermo/Applied Biosystems). The internal control gene used was ampR. All experiments were done in biological quadruplicates.
Motility Assays
Motility assays were performed essentially as previously described (Shrout et al., 2006). For swimming motility, LB-Lennox plates containing 0.3% Bacto agar were inoculated with overnight cultures and a sterile inoculation needle, ensuring the needle tip was inserted approximately halfway into the agar but not to the plastic petri dish bottom, and incubated for 24 h at 30°C. Swim ring diameters were measured for quantitation. For twitching motility, LB-Lennox plates containing 0.5% Bacto agar were inoculated by inserting the inoculation needle through the agar until it touched the plastic bottom. Plates were incubated for 72 h at 30°C. Subsequently, the agar was peeled off, and the plates were stained with 1% w/v crystal violet to visualize the twitching zone diameters prior to measurement (Déziel et al., 2001). All motility experiments were performed in at least biological triplicates.
Protein Expression and Purification
Proteins were purified by previously established protocols. Briefly, over-expressed RsmA-His6 was purified from a 4 × 1-L cultures of P. aeruginosa PAO1 bearing pUCP18:rsmA-His6 (Irie et al., 2010), while all Hfq-His6 proteins were purified from a 400 ml culture of Hfq null derivative of E. coli BL21(DE3) (Madhushani et al., 2015) carrying PT7 expression plasmids (pVI2344-2346 or pVI2357).
RsmA and Hfq over-expressing strains were grown in 2× YT broth to stationary or exponential phases, respectively. Cell pellets were re-suspended in lysis buffer (buffer A: 0.3 M of NaCl, 20 mM of Tris–HCl pH 8, 5 mM of imidazole) supplemented with 1 tablet⋅ L–1 cOmpleteTM EDTA-free protease inhibitor tablet (Sigma). Cells were lysed by Stansted Fluid Power SPCH ultra high-pressure cell disrupter/homogenizer. Cell debris were removed by centrifugation (35,000 rpm, 40 min, 4°C), and the supernatants loaded onto 1 ml of HisTRAP HP (GE Healthcare) columns equilibrated in lysis buffer (buffer A). The column was washed with high salt buffer (buffer B: 5 M of NaCl, 20 mM of Tris–HCl pH 8, 5 mM of imidazole) to remove RNA contamination, and the proteins subsequently eluted with the linear gradient of imidazole (25 to 750 mM) in buffer C (buffer C: 0.5 M of NaCl, 20 mM of Tris–HCl pH 8). Fractions containing the desired proteins were pooled. For RsmA-His6, an additional ion exchange step using 5 ml of HiTrap SP HP column (GE Healthcare) equilibrated with buffer D (0.1 M of NaCl, 20 mM of Tris–HCl pH 8) was performed, with RsmA subsequently being eluted with 1 M of NaCl.
All proteins were dialyzed across Slide-a-lyzer 10,000 molecular weight cutoff (MWCO) cassettes (Thermo Fisher) against two lots of 2 L dialysis buffer (40 mM of Tris–HCl pH 8, 600 mM of NaCl, 2 mM of EDTA) at 4°C for 2 days prior to concentration using 10-MWCO Centricon (Amicon). Protein concentrations were determined using the bicinchoninic acid (BCA) Protein Assay Kit (Pierce/Fisher), and purity was confirmed by SDS-PAGE. Proteins were stored at −20°C in final storage buffer (20 mM of Tris–HCl pH 8, 300 mM of NaCl, 1 mM of EDTA, 40% glycerol) until use.
RNA Synthesis
Wild-type and mutant vfr mRNAs were generated using Ambion’s MEGAscript kit as recommended for in vitro transcription from the PT7 promoter of linearized plasmids. Reactions (total 40 μl) containing 1 μg of EcoRV-linearized pVI2358, pVI2359, or pVI2360 were incubated for 6 h prior to 15-min DNase I treatment. Reactions were terminated by adding 230 μl of RNase-free H2O and 30 μl of stop solution (5 M of ammonium acetate; 100 mM of EDTA). The resulting 223-nt RNAs (encompassing coordinates −106 to +116 relative to the A of the initiation codon of vfr) for WT and GG→CC versions of vfr RNAs and 135-nt RNA for the truncated version of vfr RNA (encompassing coordinates −16 to +116 relative to the A of the initiation codon of vfr) were extracted twice with phenol:chloroform:IAA (25:24:1) and once with chloroform prior to precipitation and resuspension in 50 μl of RNase-free H2O.
RNA Electrophoretic Mobility Shift Assays
Reactions (total 10 μl) contained 32 nM of in vitro transcribed RNA with the indicated concentrations of RsmA and/or Hfq. RsmA and Hfq molarities given here are for the dimeric and hexameric complexes, respectively. RNA was first heated at 80°C for 5 min and then immediately chilled in an ice-water bath. Additional final reaction mixture components were as follows: 10 mM of HEPES pH 7.9, 2 mM of MgCl2, 90 ng of yeast total RNA (Fisher), 4U of RNasin, and 35 mM of KCl. Binding reactions were performed at 20°C for 40 min prior to adding 2.5 μl of loading buffer (40% sucrose). Reactions were placed on ice and analyzed on 5% TBE Criterion pre-cast gels (Bio-Rad) with electrophoresis at 100 V for ∼130 min at 4°C. Gels were stained at room temperature with 1:10,000 TBE-diluted SYBR Gold (Fisher) in a light-protected container with agitation. Images were captured using GE Healthcare Typhoon.
Tissue Culture Infections
Human RPE cells were grown in DMEM/F12 (Corning DMEM Hams F-12 50/50 Mix), supplemented with 10% of fetal bovine serum and 3% of sodium bicarbonate solution (Gibco) at 37°C with 5% CO2. Prior to infection, 2.5 × 105 cells⋅ ml–1 of RPE were seeded in 24-well plates to reach ∼80% confluence (microscopically verified) in 24 h. Media were gently aspirated from the top of the cells, and each well was washed using antibiotic-free media. Overnight LB liquid culture-grown bacteria were diluted in DMEM/F12 to achieve the multiplicity of infection (MOI) of 50:1. Infected cells were then incubated at 37°C for 24 h. Bacterial numbers were enumerated by plating serial dilutions onto LB agar and counting colonies at 0 (to verify MOI) and 24 h post-infection.
Lactate Dehydrogenase Release Assay
Supernatants were recovered from each well, and LDH activities were determined using the LDH kit (Sigma) following the manufacturer’s instructions. Signal outputs were read using a 96-well plate reader (BioTek).
Statistics
Presented data are the mean with all error bars representing standard deviations of the data collected. Two-tailed p-values were calculated from Student’s t-test using the GraphPad Prism software. Data are statistically significant if p < 0.025.
Data Availability Statement
The datasets generated for this study are available on request to the corresponding author.
Author Contributions
ALM (infection studies), VS (in vitro RNA synthesis), and YI (all others) designed and performed the experiments and analyzed the data. VM assisted in purification of the RsmA and Hfq proteins. YI and VS wrote the manuscript with input from ALM, VH, VM, and TT. All authors contributed to the article and approved the submitted version.
Funding
This work was supported by the J C Kempe and S M Kempe Foundation (JCK-1523 to VS), the Swedish Research Council (grant numbers 2016-02047 to VS and 2017-03783 to VH), and the University of Tartu Institute of Technology Grant (PLTTIIRIE to YI).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
YI would like to thank Matt Parsek for his more than a decade of scientific and moral support that led to the culmination and conclusion of this project. The authors would like to thank Elisabeth Sonnleitner for her generous gifts of hfq deletion construct and Hfq over-expression plasmids.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2020.482585/full#supplementary-material
References
Arai, H. (2011). Regulation and function of versatile aerobic and anaerobic respiratory metabolism in Pseudomonas aeruginosa. Front. Microbiol. 2:103. doi: 10.3389/fmicb.2011.00103
Baker, C. S., Eöry, L. A., Yakhnin, H., Mercante, J., Romeo, T., and Babitzke, P. (2007). CsrA inhibits translation initiation of Escherichia coli hfq by binding to a single site overlapping the Shine-Dalgarno sequence. J. Bacteriol. 189, 5472–5481. doi: 10.1128/jb.00529-07
Baker, C. S., Morozov, I., Suzuki, K., Romeo, T., and Babitzke, P. (2002). CsrA regulates glycogen biosynthesis by preventing translation of glgC in Escherichia coli. Mol. Microbiol. 44, 1599–1610. doi: 10.1046/j.1365-2958.2002.02982.x
Baraquet, C., Murakami, K., Parsek, M. R., and Harwood, C. S. (2012). The FleQ protein from Pseudomonas aeruginosa functions as both a repressor and an activator to control gene expression from the pel operon promoter in response to c-di-GMP. Nucleic Acids Res. 40, 7207–7218. doi: 10.1093/nar/gks384
Beatson, S. A., Whitchurch, C. B., Sargent, J. L., Levesque, R. C., and Mattick, J. S. (2002). Differential regulation of twitching motility and elastase production by Vfr in Pseudomonas aeruginosa. J. Bacteriol. 184, 3605–3613. doi: 10.1128/jb.184.13.3605-3613.2002
Belete, B., Lu, H., and Wozniak, D. J. (2008). Pseudomonas aeruginosa AlgR regulates type IV pilus biosynthesis by activating transcription of the fimU-pilVWXY1Y2E operon. J. Bacteriol. 190, 2023–2030. doi: 10.1128/jb.01623-07
Brencic, A., and Lory, S. (2009). Determination of the regulon and identification of novel mRNA targets of Pseudomonas aeruginosa RsmA. Mol. Microbiol. 72, 612–632. doi: 10.1111/j.1365-2958.2009.06670.x
Burrowes, E., Baysse, C., Adams, C., and O’Gara, F. (2006). Influence of the regulatory protein RsmA on cellular functions in Pseudomonas aeruginosa PAO1, as revealed by transcriptome analysis. Microbiology 152, 405–418. doi: 10.1099/mic.0.28324-0
Burrows, L. L. (2012). Pseudomonas aeruginosa twitching motility: type IV pili in action. Annu. Rev. Microbiol. 66, 493–520. doi: 10.1146/annurev-micro-092611-150055
Chambonnier, G., Roux, L., Redelberger, D., Fadel, F., Filloux, A., Sivaneson, M., et al. (2016). The hybrid histidine kinase LadS forms a multicomponent signal transduction system with the GacS/GacA two-component system in Pseudomonas aeruginosa. PLoS Genet. 12:e1006032. doi: 10.1371/journal.pgen.1006032
Chandler, C. E., Horspool, A. M., Hill, P. J., Wozniak, D. J., Schertzer, J. W., Rasko, D. A., et al. (2019). Genomic and phenotypic diversity among ten laboratory isolates of Pseudomonas aeruginosa PAO1. J. Bacteriol. 201:e00595-18.
Choi, K. H., Kumar, A., and Schweizer, H. P. (2006). A 10-min method for preparation of highly electrocompetent Pseudomonas aeruginosa cells: application for DNA fragment transfer between chromosomes and plasmid transformation. J. Microbiol. Methods 64, 391–397. doi: 10.1016/j.mimet.2005.06.001
Crone, S., Vives-Flórez, M., Kvich, L., Saunders, A. M., Malone, M., Nicolaisen, M. H., et al. (2020). The environmental occurrence of Pseudomonas aeruginosa. APMIS 128, 220–231.
Dasgupta, N., Ferrell, E. P., Kanack, K. J., West, S. E., and Ramphal, R. (2002). fleQ, the gene encoding the major flagellar regulator of Pseudomonas aeruginosa, is σ70 dependent and is downregulated by Vfr, a homolog of Escherichia coli cyclic AMP receptor protein. J. Bacteriol. 184, 5240–5250. doi: 10.1128/jb.184.19.5240-5250.2002
Dasgupta, N., Wolfgang, M. C., Goodman, A. L., Arora, S. K., Jyot, J., Lory, S., et al. (2003). A four-tiered transcriptional regulatory circuit controls flagellar biogenesis in Pseudomonas aeruginosa. Mol. Microbiol. 50, 809–824. doi: 10.1046/j.1365-2958.2003.03740.x
Deneke, C., Lipowsky, R., and Valleriani, A. (2013). Effect of ribosome shielding on mRNA stability. Phys. Biol. 10:046008. doi: 10.1088/1478-3975/10/4/046008
Déziel, E., Comeau, Y., and Villemur, R. (2001). Initiation of biofilm formation by Pseudomonas aeruginosa 57RP correlates with emergence of hyperpiliated and highly adherent phenotypic variants deficient in swimming, swarming, and twitching motilities. J. Bacteriol. 183, 1195–1204. doi: 10.1128/jb.183.4.1195-1204.2001
Dubey, A. K., Baker, C. S., Romeo, T., and Babitzke, P. (2005). RNA sequence and secondary structure participate in high-affinity CsrA-RNA interaction. RNA 11, 1579–1587. doi: 10.1261/rna.2990205
Dubey, A. K., Baker, C. S., Suzuki, K., Jones, A. D., Pandit, P., Romeo, T., et al. (2003). CsrA regulates translation of the Escherichia coli carbon starvation gene, cstA, by blocking ribosome access to the cstA transcript. J. Bacteriol. 185, 4450–4460. doi: 10.1128/jb.185.15.4450-4460.2003
Fuchs, E. L., Brutinel, E. D., Jones, A. K., Fulcher, N. B., Urbanowski, M. L., Yahr, T. L., et al. (2010). The Pseudomonas aeruginosa Vfr regulator controls global virulence factor expression through cyclic AMP-dependent and -independent mechanisms. J. Bacteriol. 192, 3553–3564. doi: 10.1128/jb.00363-10
Gebhardt, M. J., Kambara, T. K., Ramsey, K. M., and Dove, S. L. (2020). Widespread targeting of nascent transcripts by RsmA in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. U.S.A. 117, 10520–10529. doi: 10.1073/pnas.1917587117
Gellatly, S. L., and Hancock, R. E. (2013). Pseudomonas aeruginosa: new insights into pathogenesis and host defenses. Pathog. Dis. 67, 159–173. doi: 10.1111/2049-632x.12033
Gilbert, K. B., Kim, T. H., Gupta, R., Greenberg, E. P., and Schuster, M. (2009). Global position analysis of the Pseudomonas aeruginosa quorum-sensing transcription factor LasR. Mol. Microbiol. 73, 1072–1085. doi: 10.1111/j.1365-2958.2009.06832.x
Gooderham, W. J., and Hancock, R. E. (2009). Regulation of virulence and antibiotic resistance by two-component regulatory systems in Pseudomonas aeruginosa. FEMS Microbiol. Rev. 33, 279–294. doi: 10.1111/j.1574-6976.2008.00135.x
Goodman, A. L., Kulasekara, B., Rietsch, A., Boyd, D., Smith, R. S., and Lory, S. (2004). A signaling network reciprocally regulates genes associated with acute infection and chronic persistence in Pseudomonas aeruginosa. Dev. Cell 7, 745–754. doi: 10.1016/j.devcel.2004.08.020
Goodman, A. L., Merighi, M., Hyodo, M., Ventre, I., Filloux, A., and Lory, S. (2009). Direct interaction between sensor kinase proteins mediates acute and chronic disease phenotypes in a bacterial pathogen. Genes Dev. 23, 249–259. doi: 10.1101/gad.1739009
Gutiérrez, P., Li, Y., Osborne, M. J., Pomerantseva, E., Liu, Q., and Gehring, K. (2005). Solution structure of the carbon storage regulator protein CsrA from Escherichia coli. J. Bacteriol. 187, 3496–3501. doi: 10.1128/jb.187.10.3496-3501.2005
Heurlier, K., Williams, F., Heeb, S., Dormond, C., Pessi, G., Singer, D., et al. (2004). Positive control of swarming, rhamnolipid synthesis, and lipase production by the posttranscriptional RsmA/RsmZ system in Pseudomonas aeruginosa PAO1. J. Bacteriol. 186, 2936–2945. doi: 10.1128/jb.186.10.2936-2945.2004
Hickman, J. W., and Harwood, C. S. (2008). Identification of FleQ from Pseudomonas aeruginosa as a c-di-GMP-responsive transcription factor. Mol. Microbiol. 69, 376–389. doi: 10.1111/j.1365-2958.2008.06281.x
Hill, I. T., Tallo, T., Dorman, M. J., and Dove, S. L. (2019). Loss of RNA chaperone Hfq unveils a toxic pathway in Pseudomonas aeruginosa. J. Bacteriol. 201:e00232-19.
Hoang, T. T., Karkhoff-Schweizer, R. R., Kutchma, A. J., and Schweizer, H. P. (1998). A broad-host-range Flp-FRT recombination system for site-specific excision of chromosomally-located DNA sequences: application for isolation of unmarked Pseudomonas aeruginosa mutants. Gene 212, 77–86. doi: 10.1016/s0378-1119(98)00130-9
Hoang, T. T., Kutchma, A. J., Becher, A., and Schweizer, H. P. (2000). Integration-proficient plasmids for Pseudomonas aeruginosa: site-specific integration and use for engineering of reporter and expression strains. Plasmid 43, 59–72. doi: 10.1006/plas.1999.1441
Hoang, T. T., and Schweizer, H. P. (1997). Fatty acid biosynthesis in Pseudomonas aeruginosa: cloning and characterization of the fabAB operon encoding β-hydroxyacyl-acyl carrier protein dehydratase (FabA) and β-ketoacyl-acyl carrier protein synthase I (FabB). J. Bacteriol. 179, 5326–5332. doi: 10.1128/jb.179.17.5326-5332.1997
Holmqvist, E., Wright, P. R., Li, L., Bischler, T., Barquist, L., Reinhardt, R., et al. (2016). Global RNA recognition patterns of post-transcriptional regulators Hfq and CsrA revealed by UV crosslinking in vivo. EMBO J. 35, 991–1011. doi: 10.15252/embj.201593360
Intile, P. J., Diaz, M. R., Urbanowski, M. L., Wolfgang, M. C., and Yahr, T. L. (2014). The AlgZR two-component system recalibrates the RsmAYZ posttranscriptional regulatory system to inhibit expression of the Pseudomonas aeruginosa type III secretion system. J. Bacteriol. 196, 357–366. doi: 10.1128/jb.01199-13
Irie, Y., Borlee, B. R., O’Connor, J. R., Hill, P. J., Harwood, C. S., Wozniak, D. J., et al. (2012). Self-produced exopolysaccharide is a signal that stimulates biofilm formation in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. U.S.A. 109, 20632–20636. doi: 10.1073/pnas.1217993109
Irie, Y., Starkey, M., Edwards, A. N., Wozniak, D. J., Romeo, T., and Parsek, M. R. (2010). Pseudomonas aeruginosa biofilm matrix polysaccharide Psl is regulated transcriptionally by RpoS and post-transcriptionally by RsmA. Mol. Microbiol. 78, 158–172.
Janssen, K. H., Corley, J. M., Djapgne, L., Cribbs, J. T., Voelker, D., Slusher, Z., et al. (2020). Hfq and sRNA 179 Inhibit Expression of the Pseudomonas aeruginosa cAMP-Vfr and Type III Secretion Regulons. mBio 11:e00363-20.
Kambara, T. K., Ramsey, K. M., and Dove, S. L. (2018). Pervasive Targeting of Nascent Transcripts by Hfq. Cell Rep. 23, 1543–1552. doi: 10.1016/j.celrep.2018.03.134
Kay, E., Humair, B., Dénervaud, V., Riedel, K., Spahr, S., Eberl, L., et al. (2006). Two GacA-dependent small RNAs modulate the quorum-sensing response in Pseudomonas aeruginosa. J. Bacteriol. 188, 6026–6033. doi: 10.1128/jb.00409-06
Kazmierczak, B. I., Schniederberend, M., and Jain, R. (2015). Cross-regulation of Pseudomonas motility systems: the intimate relationship between flagella, pili and virulence. Curr. Opin. Microbiol. 28, 78–82. doi: 10.1016/j.mib.2015.07.017
Khare, A., and Tavazoie, S. (2015). Multifactorial competition and resistance in a two-species bacterial system. PLoS Genet. 11:e1005715. doi: 10.1371/journal.pgen.1005715
Klausen, M., Heydorn, A., Ragas, P., Lambertsen, L., Aaes-Jørgensen, A., Molin, S., et al. (2003). Biofilm formation by Pseudomonas aeruginosa wild type, flagella and type IV pili mutants. Mol. Microbiol. 48, 1511–1524. doi: 10.1046/j.1365-2958.2003.03525.x
Klockgether, J., Munder, A., Neugebauer, J., Davenport, C. F., Stanke, F., Larbig, K. D., et al. (2010). Genome diversity of Pseudomonas aeruginosa PAO1 laboratory strains. J. Bacteriol. 192, 1113–1121. doi: 10.1128/jb.01515-09
Kong, W., Chen, L., Zhao, J., Shen, T., Surette, M. G., Shen, L., et al. (2013). Hybrid sensor kinase PA1611 in Pseudomonas aeruginosa regulates transitions between acute and chronic infection through direct interaction with RetS. Mol. Microbiol. 88, 784–797. doi: 10.1111/mmi.12223
Lapouge, K., Sineva, E., Lindell, M., Starke, K., Baker, C. S., Babitzke, P., et al. (2007). Mechanism of hcnA mRNA recognition in the Gac/Rsm signal transduction pathway of Pseudomonas fluorescens. Mol. Microbiol. 66, 341–356. doi: 10.1111/j.1365-2958.2007.05909.x
Lawhon, S. D., Frye, J. G., Suyemoto, M., Porwollik, S., Mcclelland, M., and Altier, C. (2003). Global regulation by CsrA in Salmonella typhimurium. Mol. Microbiol. 48, 1633–1645. doi: 10.1046/j.1365-2958.2003.03535.x
Lequette, Y., Lee, J.-H., Ledgham, F., Lazdunski, A., and Greenberg, E. P. (2006). A distinct QscR regulon in the Pseudomonas aeruginosa quorum-sensing circuit. J. Bacteriol. 188, 3365–3370. doi: 10.1128/jb.188.9.3365-3370.2006
Link, T. M., Valentin-Hansen, P., and Brennan, R. G. (2009). Structure of Escherichia coli Hfq bound to polyriboadenylate RNA. Proc. Natl. Acad. Sci. U.S.A. 106, 19292–19297. doi: 10.1073/pnas.0908744106
Liu, P. V. (1966). The roles of various fractions of Pseudomonas aeruginosa in its pathogenesis. 3. Identity of the lethal toxins produced in vitro and in vivo. J. Infect. Dis. 116, 481–489. doi: 10.1093/infdis/116.4.481
Lizewski, S. E., Schurr, J. R., Jackson, D. W., Frisk, A., Carterson, A. J., and Schurr, M. J. (2004). Identification of AlgR-regulated genes in Pseudomonas aeruginosa by use of microarray analysis. J. Bacteriol. 186, 5672–5684. doi: 10.1128/jb.186.17.5672-5684.2004
Lory, S. (1986). Effect of iron on accumulation of exotoxin A-specific mRNA in Pseudomonas aeruginosa. J. Bacteriol. 168, 1451–1456. doi: 10.1128/jb.168.3.1451-1456.1986
Luo, Y., Zhao, K., Baker, A. E., Kuchma, S. L., Coggan, K. A., Wolfgang, M. C., et al. (2015). A hierarchical cascade of second messengers regulates Pseudomonas aeruginosa surface behaviors. mBio 6:e02456-14.
Madhushani, A., Del Peso-Santos, T., Moreno, R., Rojo, F., and Shingler, V. (2015). Transcriptional and translational control through the 5′-leader region of the dmpR master regulatory gene of phenol metabolism. Environ. Microbiol. 17, 119–133. doi: 10.1111/1462-2920.12511
Marden, J. N., Diaz, M. R., Walton, W. G., Gode, C. J., Betts, L., Urbanowski, M. L., et al. (2013). An unusual CsrA family member operates in series with RsmA to amplify posttranscriptional responses in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. U.S.A. 110, 15055–15060. doi: 10.1073/pnas.1307217110
Marsden, A. E., Intile, P. J., Schulmeyer, K. H., Simmons-Patterson, E. R., Urbanowski, M. L., Wolfgang, M. C., et al. (2016). Vfr Directly activates exsA transcription to regulate expression of the Pseudomonas aeruginosa Type III secretion system. J. Bacteriol. 198, 1442–1450. doi: 10.1128/jb.00049-16
Milojevic, T., Grishkovskaya, I., Sonnleitner, E., Djinovic-Carugo, K., and Bläsi, U. (2013). The Pseudomonas aeruginosa catabolite repression control protein Crc is devoid of RNA binding activity. PLoS One 8:e64609. doi: 10.1371/journal.pone.0064609
Møller, T., Franch, T., Hojrup, P., Keene, D. R., Bachinger, H. P., Brennan, R. G., et al. (2002). Hfq: a bacterial Sm-like protein that mediates RNA-RNA interaction. Mol. Cell 9, 23–30.
Moreno, R., Hernández-Arranz, S., La Rosa, R., Yuste, L., Madhushani, A., Shingler, V., et al. (2015). The Crc and Hfq proteins of Pseudomonas putida cooperate in catabolite repression and formation of ribonucleic acid complexes with specific target motifs. Environ. Microbiol. 17, 105–118. doi: 10.1111/1462-2920.12499
Morita, T., Maki, K., and Aiba, H. (2005). RNase E-based ribonucleoprotein complexes: mechanical basis of mRNA destabilization mediated by bacterial noncoding RNAs. Genes Dev. 19, 2176–2186. doi: 10.1101/gad.1330405
Mulcahy, H., O’Callaghan, J., O’Grady, E. P., Adams, C., and O’Gara, F. (2006). The posttranscriptional regulator RsmA plays a role in the interaction between Pseudomonas aeruginosa and human airway epithelial cells by positively regulating the type III secretion system. Infect. Immun. 74, 3012–3015. doi: 10.1128/iai.74.5.3012-3015.2006
Mulcahy, H., O’Callaghan, J., O’Grady, E. P., Maciá, M. D., Borrell, N., Gómez, C., et al. (2008). Pseudomonas aeruginosa RsmA plays an important role during murine infection by influencing colonization, virulence, persistence, and pulmonary inflammation. Infect. Immun. 76, 632–638. doi: 10.1128/iai.01132-07
O’Grady, E. P., Mulcahy, H., O’Callaghan, J., Adams, C., and O’Gara, F. (2006). Pseudomonas aeruginosa infection of airway epithelial cells modulates expression of Kruppel-like factors 2 and 6 via RsmA-mediated regulation of type III exoenzymes S and Y. Infect. Immun. 74, 5893–5902. doi: 10.1128/iai.00489-06
O’Toole, G. A., and Kolter, R. (1998). Flagellar and twitching motility are necessary for Pseudomonas aeruginosa biofilm development. Mol. Microbiol. 30, 295–304. doi: 10.1046/j.1365-2958.1998.01062.x
Pannuri, A., Yakhnin, H., Vakulskas, C. A., Edwards, A. N., Babitzke, P., and Romeo, T. (2012). Translational repression of NhaR, a novel pathway for multi-tier regulation of biofilm circuitry by CsrA. J. Bacteriol. 194, 79–89. doi: 10.1128/jb.06209-11
Patterson-Fortin, L. M., Vakulskas, C. A., Yakhnin, H., Babitzke, P., and Romeo, T. (2013). Dual posttranscriptional regulation via a cofactor-responsive mRNA leader. J. Mol. Biol. 425, 3662–3677. doi: 10.1016/j.jmb.2012.12.010
Pessi, G., Williams, F., Hindle, Z., Heurlier, K., Holden, M. T., Cámara, M., et al. (2001). The global posttranscriptional regulator RsmA modulates production of virulence determinants and N-acylhomoserine lactones in Pseudomonas aeruginosa. J. Bacteriol. 183, 6676–6683. doi: 10.1128/jb.183.22.6676-6683.2001
Potts, A. H., Vakulskas, C. A., Pannuri, A., Yakhnin, H., Babitzke, P., and Romeo, T. (2017). Global role of the bacterial post-transcriptional regulator CsrA revealed by integrated transcriptomics. Nat. Commun. 8:1596.
Pritchett, C. L., Little, A. S., Okkotsu, Y., Frisk, A., Cody, W. L., Covey, C. R., et al. (2015). Expression analysis of the Pseudomonas aeruginosa AlgZR two-component regulatory system. J. Bacteriol. 197, 736–748. doi: 10.1128/jb.02290-14
Quan, J., and Tian, J. (2009). Circular polymerase extension cloning of complex gene libraries and pathways. PLoS One 4:e6441. doi: 10.1371/journal.pone.0006441
Quan, J., and Tian, J. (2011). Circular polymerase extension cloning for high-throughput cloning of complex and combinatorial DNA libraries. Nat. Protoc. 6, 242–251. doi: 10.1038/nprot.2010.181
Rahme, L. G., Stevens, E. J., Wolfort, S. F., Shao, J., Tompkins, R. G., and Ausubel, F. M. (1995). Common virulence factors for bacterial pathogenicity in plants and animals. Science 268, 1899–1902. doi: 10.1126/science.7604262
Ren, B., Shen, H., Lu, Z. J., Liu, H., and Xu, Y. (2014). The phzA2-G2 transcript exhibits direct RsmA-mediated activation in Pseudomonas aeruginosa M18. PLoS One 9:e89653. doi: 10.1371/journal.pone.0089653
Rife, C., Schwarzenbacher, R., Mcmullan, D., Abdubek, P., Ambing, E., Axelrod, H., et al. (2005). Crystal structure of the global regulatory protein CsrA from Pseudomonas putida at 2.05 Å resolution reveals a new fold. Proteins 61, 449–453.
Romeo, T., Vakulskas, C. A., and Babitzke, P. (2013). Post-transcriptional regulation on a global scale: form and function of Csr/Rsm systems. Environ. Microbiol. 15, 313–324. doi: 10.1111/j.1462-2920.2012.02794.x
Ryder, C., Byrd, M., and Wozniak, D. J. (2007). Role of polysaccharides in Pseudomonas aeruginosa biofilm development. Curr. Opin. Microbiol. 10, 644–648. doi: 10.1016/j.mib.2007.09.010
Sahr, T., Rusniok, C., Impens, F., Oliva, G., Sismeiro, O., Coppee, J. Y., et al. (2017). The Legionella pneumophila genome evolved to accommodate multiple regulatory mechanisms controlled by the CsrA-system. PLoS Genet. 13:e1006629. doi: 10.1371/journal.pgen.1006629
Sakuragi, Y., and Kolter, R. (2007). Quorum-sensing regulation of the biofilm matrix genes (pel) of Pseudomonas aeruginosa. J. Bacteriol. 189, 5383–5386. doi: 10.1128/jb.00137-07
Schu, D. J., Zhang, A., Gottesman, S., and Storz, G. (2015). Alternative Hfq-sRNA interaction modes dictate alternative mRNA recognition. EMBO J. 34, 2557–2573. doi: 10.15252/embj.201591569
Schubert, M., Lapouge, K., Duss, O., Oberstrass, F. C., Jelesarov, I., Haas, D., et al. (2007). Molecular basis of messenger RNA recognition by the specific bacterial repressing clamp RsmA/CsrA. Nat. Struct. Mol. Biol. 14, 807–813. doi: 10.1038/nsmb1285
Schulmeyer, K. H., Diaz, M. R., Bair, T. B., Sanders, W., Gode, C. J., Laederach, A., et al. (2016). Primary and secondary sequence structure requirements for recognition and discrimination of target RNAs by Pseudomonas aeruginosa RsmA and RsmF. J. Bacteriol. 198, 2458–2469. doi: 10.1128/jb.00343-16
Schweizer, H. P. (1991). Escherichia-Pseudomonas shuttle vectors derived from pUC18/19. Gene 97, 109–121. doi: 10.1016/0378-1119(91)90016-5
Shrout, J. D., Chopp, D. L., Just, C. L., Hentzer, M., Givskov, M., and Parsek, M. R. (2006). The impact of quorum sensing and swarming motility on Pseudomonas aeruginosa biofilm formation is nutritionally conditional. Mol. Microbiol. 62, 1264–1277. doi: 10.1111/j.1365-2958.2006.05421.x
Sonnleitner, E., Hagens, S., Rosenau, F., Wilhelm, S., Habel, A., Jager, K. E., et al. (2003). Reduced virulence of a hfq mutant of Pseudomonas aeruginosa O1. Microb. Pathog. 35, 217–228. doi: 10.1016/s0882-4010(03)00149-9
Sonnleitner, E., Prindl, K., and Bläsi, U. (2017). The Pseudomonas aeruginosa CrcZ RNA interferes with Hfq-mediated riboregulation. PLoS One 12:e0180887. doi: 10.1371/journal.pone.0180887
Sonnleitner, E., Schuster, M., Sorger-Domenigg, T., Greenberg, E. P., and Bläsi, U. (2006). Hfq-dependent alterations of the transcriptome profile and effects on quorum sensing in Pseudomonas aeruginosa. Mol. Microbiol. 59, 1542–1558. doi: 10.1111/j.1365-2958.2006.05032.x
Sonnleitner, E., Wulf, A., Campagne, S., Pei, X. Y., Wolfinger, M. T., Forlani, G., et al. (2018). Interplay between the catabolite repression control protein Crc, Hfq and RNA in Hfq-dependent translational regulation in Pseudomonas aeruginosa. Nucleic Acids Res. 46, 1470–1485. doi: 10.1093/nar/gkx1245
Sorger-Domenigg, T., Sonnleitner, E., Kaberdin, V. R., and Bläsi, U. (2007). Distinct and overlapping binding sites of Pseudomonas aeruginosa Hfq and RsmA proteins on the non-coding RNA RsmY. Biochem. Biophys. Res. Commun. 352, 769–773. doi: 10.1016/j.bbrc.2006.11.084
Tashiro, Y., Yawata, Y., Toyofuku, M., Uchiyama, H., and Nomura, N. (2013). Interspecies interaction between Pseudomonas aeruginosa and other microorganisms. Microbes Environ. 28, 13–24. doi: 10.1264/jsme2.me12167
Updegrove, T. B., Zhang, A., and Storz, G. (2016). Hfq: the flexible RNA matchmaker. Curr. Opin. Microbiol. 30, 133–138. doi: 10.1016/j.mib.2016.02.003
Vakulskas, C. A., Potts, A. H., Babitzke, P., Ahmer, B. M., and Romeo, T. (2015). Regulation of bacterial virulence by Csr (Rsm) systems. Microbiol. Mol. Biol. Rev. 79, 193–224. doi: 10.1128/mmbr.00052-14
Wang, X., Dubey, A. K., Suzuki, K., Baker, C. S., Babitzke, P., and Romeo, T. (2005). CsrA post-transcriptionally represses pgaABCD, responsible for synthesis of a biofilm polysaccharide adhesin of Escherichia coli. Mol. Microbiol. 56, 1648–1663. doi: 10.1111/j.1365-2958.2005.04648.x
Whitchurch, C. B., Alm, R. A., and Mattick, J. S. (1996). The alginate regulator AlgR and an associated sensor FimS are required for twitching motility in Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. U.S.A. 93, 9839–9843. doi: 10.1073/pnas.93.18.9839
White, D., Hart, M. E., and Romeo, T. (1996). Phylogenetic distribution of the global regulatory gene csrA among eubacteria. Gene 182, 221–223. doi: 10.1016/s0378-1119(96)00547-1
Wolfgang, M. C., Lee, V. T., Gilmore, M. E., and Lory, S. (2003). Coordinate regulation of bacterial virulence genes by a novel adenylate cyclase-dependent signaling pathway. Dev. Cell 4, 253–263. doi: 10.1016/s1534-5807(03)00019-4
Yakhnin, A. V., Baker, C. S., Vakulskas, C. A., Yakhnin, H., Berezin, I., Romeo, T., et al. (2013). CsrA activates flhDC expression by protecting flhDC mRNA from RNase E-mediated cleavage. Mol. Microbiol. 87, 851–866. doi: 10.1111/mmi.12136
Zuker, M. (1989). On finding all suboptimal foldings of an RNA molecule. Science 244, 48–52. doi: 10.1126/science.2468181
Keywords: Pseudomonas aeruginosa, RsmA, Vfr, FleQ, Hfq
Citation: Irie Y, La Mensa A, Murina V, Hauryliuk V, Tenson T and Shingler V (2020) Hfq-Assisted RsmA Regulation Is Central to Pseudomonas aeruginosa Biofilm Polysaccharide PEL Expression. Front. Microbiol. 11:482585. doi: 10.3389/fmicb.2020.482585
Received: 02 July 2019; Accepted: 08 October 2020;
Published: 17 November 2020.
Edited by:
Eliane Hajnsdorf, UMR 8261 Expression Génétique Microbienne, FranceReviewed by:
Vladimir Kaberdin, University of the Basque Country, SpainEric Déziel, Université du Québec, Canada
Copyright © 2020 Irie, La Mensa, Murina, Hauryliuk, Tenson and Shingler. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Yasuhiko Irie, eWFzdWhpa28uaXJpZUB1dC5lZQ==
†Present address: Yasuhiko Irie, Institute of Technology, University of Tartu, Tartu, Estonia; Agnese La Mensa, Department of Biomedicine, Neurosciences and Advanced Diagnostics, University of Palermo, Palermo, Italy
‡ORCID: Yasuhiko Irie, orcid.org/0000-0002-7162-4772; Victoriia Murina, orcid.org/0000-0003-1523-1816; Vasili Hauryliuk, orcid.org/0000-0003-2389-5057; Tanel Tenson, orcid.org/0000-0002-0260-3601; Victoria Shingler, orcid.org/0000-0002-7349-1678
 Yasuhiko Irie
Yasuhiko Irie Agnese La Mensa
Agnese La Mensa Victoriia Murina
Victoriia Murina Vasili Hauryliuk
Vasili Hauryliuk Tanel Tenson1‡
Tanel Tenson1‡ Victoria Shingler
Victoria Shingler