Abstract
Conjugal transfer is a major driving force of genetic exchange in eubacteria, and the system in IncP1-type broad-host-range plasmids transfers DNA even to eukaryotes and archaea in a process known as trans-kingdom conjugation (TKC). Although conjugation factors encoded on plasmids have been extensively analyzed, those on the donor chromosome have not. To identify the potential conjugation factor(s), a genome-wide survey on a comprehensive collection of Escherichia coli gene knockout mutants (Keio collection) as donors to Saccharomyces cerevisiae recipients was performed using a conjugal transfer system mediated by the type IV secretion system (T4SS) of the IncP1α plasmid. Out of 3,884 mutants, three mutants (ΔfrmR, ΔsufA, and ΔiscA) were isolated, which showed an increase by one order of magnitude in both E. coli–E. coli and E. coli–yeast conjugations without an increase in the mRNA accumulation level for the conjugation related genes examined. The double-knockout mutants for these genes (ΔfrmRΔsufA and ΔiscAΔfrmR) did not show synergistic effects on the conjugation efficiency, suggesting that these factors affect a common step in the conjugation machinery. The three mutants demonstrated increased conjugation efficiency in IncP1β-type but not in IncN- and IncW-type broad-host-range plasmid transfers, and the homologous gene knockout mutants against the three genes in Agrobacterium tumefaciens also showed increased TKC efficiency. These results suggest the existence of a specific regulatory system in IncP1 plasmids that enables the control of conjugation efficiency in different hosts, which could be utilized for the development of donor strains as gene introduction tools into bacteria, eukaryotes, and archaea.
Introduction
The conjugal transfer mechanism is a major driving force of genetic exchange between bacteria. Besides that, this mechanism can also occur from bacteria to either eukaryotes or archaea, known as TKC (Dodsworth et al., 2010; Moriguchi et al., 2013a; Garushyants et al., 2015; Lacroix and Citovsky, 2016). TKC is a type of horizontal gene transfer (HGT), which differs from vertical gene transfer (VGT), and promotes the transfer of genetic materials between non-related species. Within a prokaryotic population, HGT occurs ubiquitously and permits fast dissemination of new genes, which is essential for species adaptation and survival. This mechanism has been acknowledged as a driving force for the evolution of bacterial species (Richard et al., 2017; Ward et al., 2018).
Conjugation is a mechanism which involves the transfer of genetic material from donor to recipient cells due to the expression and regulation of their responsible genes harbored within the conjugative plasmid [e.g., IncP1 (Pansegrau et al., 1994; Haase et al., 1995), IncN (Winans and Walker, 1985; Yeo et al., 2003), and IncW (Fernández-López et al., 2006) plasmids] in the donor cells. In comparison to other incompatibility group plasmids including IncN and IncW plasmids, the IncP1-type plasmid has been suggested to have a broader host range, as it carries genomic signatures, which are predicted to be derived from various host origins (Suzuki et al., 2010). This IncP1-type plasmid has the ability to be transferred and replicated in hosts belonging to at least three proteobacteria subclasses: Alphaproteobacteria (Schmidhauser and Helinski, 1985; Yano et al., 2013), Betaproteobacteria (Kamachi et al., 2006; Suzuki et al., 2010), and Gammaproteobacteria (Schmidhauser and Helinski, 1985; Adamczyk and Jagura-Burdzy, 2003; Suzuki et al., 2010; Norberg et al., 2011). The ability of this IncP1-type plasmid to be adapted to, and replicated in different hosts confer its potential as gene introduction tool.
In a previous study, the IncP1-type conjugation system was reported to give a detectable DNA transfer to yeast, in addition to proteobacteria (Hayman and Bolen, 1993). IncF1 and IncI1 conjugation systems showed undetectable DNA transfer to yeast, although comparable DNA transfer to proteobacteria as IncP1-type system was observed (Bates et al., 1998). This broader transferability of the IncP1-type plasmid, such as to yeast, employed the usage of this plasmid as a gene introduction tool. Generally, to make convenient to use as the tool, a native self-transmissible plasmid (e.g., RP4 plasmid) is separated into two parts, a helper plasmid and a shuttle vector. The helper plasmid provides genes for the biosynthesis of the conjugative pilus and production of stable mating aggregates (IncP1-T4SS) for the transfer, and genes for mobilization. On the other hand, the shuttle vector comprises of origin of transfer (oriT) derived from the IncP1 plasmid and genes for the plasmid maintenance and propagation within the donor and recipient during the conjugation process. Besides the IncP1-type shuttle vectors, an IncQ-type mobilizable plasmid is alternatively used as a backbone of the shuttle vector, as it is also transferred to the recipient cells, facilitated by helper plasmids derived from IncP1α (Moriguchi et al., 2013b) or IncP1β (Willetts and Crowther, 1981; Moriguchi et al., 2016) subfamilies such as RP4 and R751 plasmids, respectively. This shuttle vector-and-helper system has been used by researchers as gene introduction tool from proteobacteria to various recipient organisms, such as, yeast (Moriguchi et al., 2013a,b; Soltysiak et al., 2019), archaea (Dodsworth et al., 2010), diatoms (Karas et al., 2015), and plant (Regnard et al., 2010).
Recently, publications related to the identification of chromosomal gene(s) within the donor cells that is responsible in promoting HGT of the RP4 plasmid upon the abiotic stress exposures (e.g., antibiotics or heavy metals exposures) have been reported. These studies performed genome-wide expression analysis of the chromosomal genes by using transcriptome analysis in relation to the expression of the selected conjugal-transfer genes in RP4 plasmid and/or genes that are probably responsible for the physiological changes of the donor cell, consequently affecting the HGT (e.g., SOS or/and Reactive Oxygen Species-related genes) (Shun-Mei et al., 2018; Zhang et al., 2019). However, in these studies, the screening approach is based on the expression analysis of the various genes within the stress-exposed donor cells which possibly influence the conjugation mechanism, and no functionality test was performed on the isolated candidate gene(s) for further validation. Thus, it is still not clear whether the genes expressed higher or less are correlated directly to the conjugation mechanism or not.
In our study, we identified and characterized the genetic features of the factor(s) within the E. coli genome that may influence the conjugative transfer mediated by IncP1α-type plasmids from an E. coli single-knockout mutant donor library. We focused on the “up”-mutants that have the ability to accelerate conjugative transfer to both prokaryotes and eukaryotes as they could be potent donor strains applicable to gene introduction tools. The isolated mutants were characterized by examining the possible correlation with the expression of the conjugation-related genes in the IncP1α plasmid. In addition, the generality of the improved characteristics by up-mutants was further characterized by assessing the conjugation efficiency of other broad-host-range plasmids, such as IncP1β, IncN, and IncW, as well as homologous gene mutants in another class of proteobacteria, Agrobacterium tumefaciens (Alphaproteobacteria).
Materials and Methods
Bacterial Strains, Yeast, and Growth Media
The bacterial strains and yeast used in this study are listed in Table 1. A complete set of E. coli non-essential gene deletion clones (Keio collection) was provided by the National BioResource Project (NBRP) of the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan. All E. coli strains and A. tumefaciens were commonly cultured in LB Lennox medium at 37 and 28°C, respectively. In addition, S. cerevisiae was cultured in yeast-extract/peptone/dextrose (YPD) medium. Synthetic defined (SD) medium containing appropriate individual amino acids (leucine, 0.03 mg/mL; histidine, 0.02 mg/mL; and lysine, 0.03 mg/mL) was used as the selection media (SC–Ura) for yeast transconjugants at 28°C. Solid LB Lennox medium was prepared by the addition of 1.5% agar, and solid YPD and SC–Ura media were prepared by the addition of 2% agar. All components used in making these media, except polypeptone, were purchased from Becton, Dickinson, and Company (Franklin Lakes, NJ, United States) and supplied by Wako Pure Chemical Ind., Ltd. (Osaka, Japan). Appropriate antibiotics were added to the media at the following concentrations, which corresponded to the selection of bacteria and plasmids: ampicillin (Ap), 50 μg/mL; chloramphenicol (Cm), 30 μg/mL; gentamicin (Gm), 30 μg/mL; kanamycin (Km), 50 μg/mL; rifampicin (Rf), 30 μg/mL; streptomycin (Sm), 50 μg/mL; tetracycline (Tc), 7.5 μg/mL; and meropenem (Me), 10 μg/mL.
TABLE 1
| Strains | Relevant characteristics | Source or reference |
| E. coli | ||
| Keio collection | An in-frame single-gene knockout mutant collection derived from BW25113, KmR | NBRP Japan |
| BW25113ΔfrmRΔsufA | frmR and sufA double-gene knockout mutant, constructed from ΔfrmR derived from Keio collection, KmR | This study |
| BW25113ΔiscAΔfrmR | iscA and frmR double-gene knockout mutant, constructed from ΔiscA derived from Keio collection, KmR | This study |
| BW25113ΔfrmAΔfrmR | frmA and frmR double-gene knockout mutant, constructed from ΔfrmA derived from Keio collection, KmR | This study |
| BW25113ΔfrmBΔfrmR | frmB and frmR double-gene knockout mutant, constructed from ΔfrmB derived from Keio collection, KmR | This study |
| BW25113 | F–Δ(araD-araB)567 ΔlacZ4787(::rrnB-3) λ-rph-1 Δ(rhaD-rhaB)568 hsdR514 | NBRP Japan |
| SY327 (λpir) | Δ(lac pro) argE(Am) recA56 λpir RifR NalR | NBRP Japan |
| S17-1 (λpir) | F– RP4-2(KmR::Tn7,TcR::Mu-1) pro-82λpir recA1 endA1 thiE1 hsdR17 creC510 | NBRP Japan |
| A. tumefaciens | ||
| C58C1 | pTiC58-cured and RifR derivative of C58 | Yamamoto et al., 2007 |
| C58C1ΔATU_RS04380 | ATU_RS04380 (atu0890) single-gene knockout derived from C58C1, RifR | This study |
| C58C1ΔATU_RS08905 | ATU_RS08905 (atu1819) single-gene knockout derived from C58C1, RifR | This study |
| C58C1ΔATU_RS08390 | ATU_RS08390 (atu1713) single-gene knockout derived from C58C1, RifR | This study |
| S. cerevisiae | ||
| BY4742 | MATα SSD1-V his3Δ1 leu2Δ0 lys2Δ0 ura3Δ0 | Invitrogen |
Strains used in this study.
Construction of TKC E. coli Donor Library From Single-Knockout Keio Collection
The TKC E. coli donor library was constructed by introducing a helper plasmid, pRH220 (Nishikawa and Yoshida, 1998), and a TKC vector, pRS316:oriTP (Moriguchi et al., 2013b), into the E. coli comprehensive gene knockout (KO) mutant collection [Keio collection (Baba et al., 2006)]. Both helper and vector plasmids were introduced into each KO mutant by conjugation as described in our previous report (Moriguchi et al., 2020). The conjugation reaction was performed under liquid conditions, followed by the selection of conjugants by inoculation to fresh selection LB Lennox medium containing Ap, Cm, and Km by 100-fold dilution at a 100 μL culture scale. The KO mutant lines that were successfully introduced with both helper and TKC vector were stored at −70°C as glycerol stocks. For the slow-growth mutant lines, which were anticipated to come from either slow growth, low conjugation efficiency, or antibiotic-susceptible phenotypes, the culture scale was increased up to 300 μL. The cultures were concentrated threefold before being stored as glycerol stocks. For the extremely slow-growth mutant lines, the conjugation reaction mixtures were directly spread onto solid selection plates and were incubated for up to 48 h at either 30°C or 37°C. Several colonies for each mutant were isolated and resuspended into the selection media, and then stored as glycerol stocks. In total, 3,884 mutant donor lines were constructed.
Donor and Recipient Cell Cultures
The details of the genotypes for both donor and recipient cells as well as plasmids used in this study are described in Tables 1, 2, respectively. For the genome-wide screening analysis, donor E. coli Keio mutants and control BW25113 (pBBR122ΔCmR) (IncP1α-pRH220, pRS316:oriTP) were inoculated from 96-well frozen stock plates using a 96-pinner tool and cultured in 100 μL medium supplemented with Ap, Cm, and Km in 96-well flat-bottom plates at 37°C for 15 to 18 h. For double-KO conjugation efficiency assessment, BW25113 single- and double-KO mutants and control (IncP1α-pRH220, pRS316:oriTP) were cultured in media supplemented with Ap and Cm. Details for the construction of E. coli double-knockout mutant strains are described in Supplementary Materials and Methods.
TABLE 2
| Plasmids | Relevant characteristics | Source or References |
| pK18mobsacB | Mobilizable plasmid; sacB oriT KmR used for the construction of A. tumefaciens knock-out mutant strains | Schäfer et al., 1994 |
| pK18mobsacB-ATU_RS04380 | Partial ATU_RS04365, ATU_RS04370, ATU_RS04375, ATU_RS04380, and ATU_RS04385 integrated within pK18mobsacB; KmR | This study |
| pK18mobsacB-ATU_RS08905 | Partial ATU_RS08895, ATU_RS08900, ATU_RS08905, ATU_RS08910 and partial nifS integrated within pK18mobsacB; KmR | This study |
| pK18mobsacB-ATU_RS08390 | dgt, ATU_RS08390, and partial ATU_RS08395 integrated within pK18mobsacB; KmR | This study |
| pK18mobsacBΔATU_RS04380 | ATU_RS04380 single-gene knockout within pK18mobsacB-ATU_RS04380; KmR | This study |
| pK18mobsacBΔATU_RS08905 | ATU_RS08905 single-gene knockout within pK18mobsacB-ATU_RS08905; KmR | This study |
| pK18mobsacBΔATU_RS08390 | ATU_RS08390 single-gene knockout within pK18mobsacB-ATU_RS08390; KmR | This study |
| pJP5603sacBGmR | Mobilizable plasmid; sacB oriT GmR Used for the construction of E. coli complementation strains | This study *LC599391 |
| pJP5603sacBGmR_sufA | Partial menI, ydiH, RydB, sufA, and sufB integrated within pJP5603sacBGmR | This study |
| pJP5603sacBGmR_iscA | iscS, iscU, iscA, hscB, and hscA integrated within pJP5603sacBGmR | This study |
| pJP5603sacBGmR_frmR | Partial yaiX, yaiO, frmR, frmA, and frmB integrated within pJP5603sacBGmR | This study |
| pBBR122ΔCmR | Derivative of a commercially provided plasmid vector pBBR122; ReppBBR’ (non-transmissible) KmR ΔCmR | Moriguchi et al., 2020 |
| RP4 | IncP1α-type conjugative broad host range plasmid; KmRTcRApR | Pansegrau et al., 1994 |
| pSa | IncW-type conjugative broad host range plasmid; CmRSuRSpRSmRKmRGmRTbR | Tait et al., 1982 |
| R46 | IncN-type conjugative broad host range plasmid; TcRSmRSuRApR | Brown and Willetts, 1981. |
| pRH220 | Helper plasmid; traP1αtrbP1αoriTP1αori-pSC101 CmR | *AB526840 |
| pDPT51 | Helper plasmid; traP1βtrbP1βori-CoIE1 TpRApR | Taylor et al., 1983. |
| pRS316:oriTP | Mobilizable plasmid; URA3 CEN6/ARSH4 ori-pMB1 ApRoriTRP4 | Moriguchi et al., 2013a. |
| pAY205 | Mobilizable plasmid; oriVQoriTQmobQURA3 TRP1 ARS1 KmRTcR | *AB526841 |
| pYN402 | Mobilizable plasmid; oriVQoriTQmobQURA3 2 μ-ori GmR | *AB531984 |
Plasmids used in this study.
*DDBJ/EMBL/GenBank accession number.
Donor E. coli BW25113 ΔsufA, ΔiscA, or ΔfrmR and control (IncW-pSa or IncN-R46) were cultured in media supplemented with Gm or Ap, respectively. Donor E. coli BW25113 ΔsufA, ΔiscA, or ΔfrmR and control (IncP1β-pDPT51, pAY205) were cultured in media supplemented with Ap and Tc.
For the conjugation efficiency assessment in Agrobacterium, donor A. tumefaciens C58C1; ΔATU_RS04380, ΔATU_RS08905, or ΔATU_RS08390 (RP4, pYN402) corresponding to frmR as well as sufA and iscA in E. coli, respectively, were cultured in media supplemented with Gm, Km, and Rf at 28°C for 16 to 18 h. C58C1 was used as a control. Details for the construction of these strains are described in Supplementary Materials and Methods.
The recipient cells of the S. cerevisiae BY4742 strain or the E. coli SY327 strain were cultured in media supplemented without or with Rf, respectively, in 5 mL glass tubes for 18 to 22 h at 28°C or 16 to 18 h at 37°C, respectively, following the inoculation from the pre-cultured plates. Both donor and recipient cultures were pre-cultured with agitation to allow aeration.
Trans-Kingdom Conjugation
Two screening strategies were performed in order to isolate the up-mutant candidates. The first screening was performed using rich medium (Supplementary Figure 1A), followed by the exclusion of antibiotic and nutrient reaction in the second screening (Supplementary Figure 2A) to create stringent conditions for transconjugant selection during the mating reaction. Fifty microliters of each donor overnight culture and 50 μL of yeast recipient suspension containing (2.0 × 106 cfu/50 μL) were mixed in the first screening and was substituted from the cultured medium to TNB (80 mM Tris-HCl [pH 7.5], and 0.05% NaCl) in the second screening. Both donor and recipient were mixed and incubated at 28°C for 1 h, followed by the selection of transconjugants by spotting 15 μL of the conjugation reaction on SC–Ura supplemented with Tc. The culture plate was incubated for 48 to 72 h at 28°C. The TKC conjugation efficiency was calculated as the log2 value (number of transconjugants per median number of transconjugants of the control strain). The turbidity of the donor cells was measured using a microtiter-plate reader MTP-310 (Corona, Ibaraki, JAPAN).
For the standard TKC reaction, the suspension of donor E. coli strains in LB Lennox medium or A. tumefaciens C58C1 and recipient yeast in TNB containing 1.8 × 107 or 5.0 × 107 and 4.0 × 106 cfu/300 μL of donors and recipient, respectively, were mixed. The donor overnight cultures of low living-cell ratio for E. coli double- and single-KO mutants (ΔfrmA and ΔfrmB) were concentrated to four-times their original concentration to adjust the number of living cells to an input cell number that was comparable to that of the wild-type control. The conjugation reaction was performed for up to 6 h for the assessment of IncP1α conjugation by the E. coli up-mutants. The conjugation reactions for the other TKC experiments were performed for 1 h. For the A. tumefaciens with yeast reaction, the scale of the reaction was increased to sevenfold to detect the transconjugant. TKC efficiency was determined based on the recovery of uracil prototrophic transconjugants supplemented with Tc or Me to inhibit the growth of donor E. coli and A. tumefaciens, respectively, calculated as the log10 value (number of transconjugants per recipient cell), and compared with the control.
Bacterial Conjugation
We used a protocol identical to the one used for the TKC, where 300 μL of donor and SY327 recipient suspensions in LB Lennox medium were used during the conjugation reaction of up to 6 h co-cultivation for IncP1α conjugation assessment. The conjugation reaction for the other bacterial conjugation assessments was performed for 1 h. The conjugation reaction containing 1.8 × 107 and 7.14 × 107 cfu/300 μL of donor and recipient, respectively, was mixed. The transconjugants were selected on LB Lennox solid medium supplemented with Rf and the appropriate antibiotics for the selection of the transferred plasmid. Conjugation efficiency was calculated as the log10 value (number of transconjugants per recipient cell).
Formaldehyde Treatment Assay
Prior to treating the donor cultures of ΔfrmR and the control with formaldehyde, the overnight cultures of both donor and E. coli recipient were transferred into fresh LB Lennox medium (1:10 dilution) containing the appropriate antibiotics and then incubated at 37°C for 2 h with agitation. Then, 250 μM of formaldehyde was added to the donor cultures, and the incubation was continued for a further 2 h. For the non-treated controls, nothing was added to the donor cultures. After the incubation, donor (treated or non-treated with formaldehyde) and recipient cultures were subjected to conjugation using the procedure described in the bacterial conjugation methodology section (above).
RNA Isolation and Quantitative RT-PCR
For the preparation of RNA, identical cultural conditions for E. coli donor cells as for the conjugation experiments were used. RNA was isolated using a NucleoSpin® RNA kit, purchased from Macherey-Nagel GmbH & Co. KG (Dueren, Germany). For DNA removal and cDNA conversion, 2 μg of the total RNA was used as a template according to the manufacturer’s instructions using a PrimeScriptTM RT reagent kit with gDNA Eraser purchased from TaKaRa Bio Inc. (Shiga, Japan). The cDNA was subjected to RT-qPCR on a LightCycler® 96 Instrument purchased from Roche Diagnostics Corporation (Indianapolis, IN, United States) using FastStart Essential DNA Green Master (Roche, Indianapolis, IN, United States). The expression levels of the target genes were normalized to the expression of the internal reference genes, cysG and rrsA (Zhou et al., 2011). All primers used in this experiment are listed in Supplementary Table 1.
Statistical Analysis
Data were expressed as the mean ± standard error of the mean (SEM) of at least three independent biological experiments. The differences between groups were analyzed using Student’s t-test when two groups were compared by two-tailed, and one-way ANOVA (Tukey HSD analysis) for multiple group comparison. Analyses were performed using SPSS IBM Software for Windows, Version 17.0 (SPSS Inc., Chicago, IL, United States). Tests were considered statistically significant when p < 0.05, p < 0.01, or p < 0.001.
Results
Identification of High-TKC-Transfer E. coli Mutants by Genome-Wide Screening
In order to identify the mutants with high TKC ability, genome-wide screenings of donor E. coli single-KO gene mutations of Keio collection on plasmid transfer to yeast recipients were performed. Mutants that showed a log2 substituted relative TKC value equal to or greater than three (eightfold compared to parental strain) during the first screening were isolated prior to the second screening step. During this screening, the relative TKC efficiency of the mutant strains was normalized based on two different median value of parental control strains (median value of all control strains used in this screening, and median value of seven control strains in every experiment consisting of approximately 160 mutant strains) (Supplementary Figures 1B,C, respectively). Data of TKC efficiency in mutant strains using both normalizations were then integrated in order to avoid the exclusion of the possible up-mutant strains in every independent experiment. Mutant strains with log2 value equal to or greater than three in both or either of these two normalization methods were selected for the second screening. In total, 233 out of 3,884 mutants were isolated.
After performing the second screening (Supplementary Figure 2B), we selected the top three mutants out of the 233: ΔfrmR, ΔsufA, and ΔiscA, which stably showed high TKC efficiency compared to the parental strain within triplicate experiments (sum log2 value ≥ 2.48). The frmR gene encodes FrmR transcriptional repressor protein on formaldehyde-sensing (frm) operon (Higgins and Giedroc, 2014; Denby et al., 2016), while the sufA and iscA genes encode the proteins within the iron–sulfur cluster assembly machinery (Lu et al., 2008).
The Abilities of Both E. coli–E. coli and E. coli–Yeast Conjugations Were Improved in the Three Up-Mutants
The increases in TKC efficiency by ΔfrmR, ΔsufA, and ΔiscA mutant donors compared to the parental strain were confirmed at different co-cultivation times (Figure 1A). At 1 h reaction, at least 17-fold increases in conjugation efficiency were observed in these mutants compared to the parental strain (Figure 1A). At 6 h reaction, at least ninefold increases in TKC efficiency were observed in these mutants compared to the parental strain.
FIGURE 1

Effect of frmR, sufA, and iscA mutations in Escherichia coli on IncP1α conjugations. (A) TKC efficiency of IncP1α transfer from E. coli to yeast within four experimental replicates (n = 4). (B) Conjugation efficiency of IncP1α transfer from E. coli to E. coli within seven experimental replicates (n = 7). Both conjugation reactions were performed for 1 h (white bar) and 6 h (black bar). Data are presented as mean ± standard error of the mean (SEM). Different letters indicate significant differences between mutants and wild-type control at p < 0.05 using Tukey HSD multiple comparison analysis. BY4742 and SY327 were used as the recipients. BW25113 parental strain was used as the control.
In order to assess the effect of these mutations on the efficiency of bacterial conjugation, the corresponding conjugation reaction was performed with E. coli SY327 recipient cells. At 1 h co-cultivation, at least ninefold increases in conjugation efficiency were observed in these mutants compared to the parental strain (Figure 1B). At 6 h co-cultivation, at least fourfold increases in conjugation efficiency were observed in these mutants compared to the parental strain (Figure 1B). On the basis of these results, we conclude that these three up-mutants (ΔfrmR, ΔsufA, and ΔiscA) increased E. coli–yeast TKC as well as E. coli–E. coli conjugation. The E. coli–yeast and E. coli–E. coli conjugation efficiencies of IncP1α between the up-mutants and the parental strain consistently showed significant difference at both 1 and 6 h co-cultivation (Figure 1). Thus, we integrated the conjugation reaction at 1 h co-cultivation after these experiments.
To confirm the repressing effect by the frmR, sufA, and iscA on the TKC of IncP1α plasmid, complementation analysis was performed by integrating the wild-type genes into the ΔfrmR, ΔsufA, and ΔiscA donor mutant strains, respectively (Supplementary Figure 3). As a result, the repressing effect on TKC was restored within the complemented donor strains.
Deficiency of frmR, sufA, and iscA Genes Can Affect Independently, but Not Synergistically to Activate IncP1α Plasmid Transfer
In order to assess the correlation between KO gene interaction and conjugation efficiency, double-KO mutant strains were constructed by introducing the second gene mutation, located within the same or different operons.
frmR, within the frm operon of E. coli K-12 derivatives, encodes a transcriptional repressor protein, FrmR (as a negative regulator), that specifically inactivates the expression of this operon in the absence of formaldehyde (Higgins and Giedroc, 2014; Denby et al., 2016; Osman et al., 2016). In the presence of formaldehyde, the expression of this operon is activated when the formaldehyde binds to the FrmR repressor, allowing induction of formaldehyde detoxification machinery catalyzed by FrmA and FrmB proteins, which are encoded by the downstream genes frmA and frmB, respectively (Denby et al., 2016; Osman et al., 2016). In this experiment, we hypothesized that the FrmR protein might be related to the increase in conjugation efficiency, due to its absence or inactivation, as the result of a deletion mutation or the binding of formaldehyde, respectively. In order to determine the effect of formaldehyde-dependent inactivation of FrmR repressor on the conjugation efficiency, both parental and ΔfrmR donor strains were treated or non-treated with formaldehyde prior to the conjugation reaction. The effect on the conjugation efficiency of these treated strains were then compared with the non-treated strains. Based on the results obtained, an effect on conjugation efficiency of the parental strain with native frmR upon the addition of formaldehyde can be observed (about fourfold increase compared to the non-treated parental strain). In addition, no significant difference in conjugation efficiency between treated and non-treated ΔfrmR with formaldehyde, although a significant increase in conjugation efficiency was observed (sevenfold) in comparison to the non-treated parental strain (Figure 2A). The result supported our hypothesis.
FIGURE 2

Confirmation analysis of the high conjugation efficiency in frmR, sufA, and iscA mutants. (A) Effect of formaldehyde (250 μM) on the conjugation efficiency of IncP1α plasmid transfer by ΔfrmR mutants and wild-type control to E. coli recipient within five experimental replicates (n = 5). (B) TKC efficiency of IncP1α transfer by genes-deficient E. coli donor, belonging to the same operon (frm operon) within five experimental replicates (n = 5). (C) TKC efficiency of IncP1α transfer by genes-deficient E. coli donor, belonging to the different operons. This experiment was performed within 12 experimental replicates (n = 12) for single-KO mutants and wild-type control, while five experimental replicates were performed for double-KO mutants (n = 5). (B,C) BY4742 was used as the recipient. Data are presented as mean ± standard error of the mean (SEM). Asterisks (∗∗∗) indicate statistically significant difference at p < 0.001 (two-tailed t-test) compared to wild-type control. No significant difference is indicated as “n.s.” between treated and non-treated ΔfrmR with formaldehyde. Different letters indicate significant differences between mutants and wild-type control at p < 0.05 using Tukey HSD multiple comparison analysis. BW25113 parental strain was used as the control. All conjugation reactions were performed for 1 h.
The TKC efficiency of the double-KO mutants (ΔfrmAΔfrmR and ΔfrmBΔfrmR) was not significantly different to those of single-KO mutants and not significantly different from each other but was significantly higher (at least sevenfold) compared to the parental strain (Figure 2B). This can possibly be attributed to the accumulation of endogenous ligands, including formaldehyde, which may inactivate the FrmR protein. This result indicates that neither frmA nor frmB alone directly affect the conjugation efficiency of IncP1α plasmids, so the effect is probably solely due to frmR.
The construction of the double-KO mutant, ΔiscAΔsufA, was unsuccessful probably because of its synthetic lethality (Vinella et al., 2009). The KO of either of these genes with ΔfrmR was constructed to discover the genetic interaction. It was observed that the TKC efficiency of the double-KO mutants, ΔiscAΔfrmR and ΔfrmRΔsufA, showed significant higher TKC efficiency compared to the parental strain by at least 11-fold, but showed no significant difference compared to the single-KO ΔfrmR, ΔsufA, and ΔiscA (Figure 2C). These results indicate that the frmR, sufA, and iscA genes probably act on an identical step of the conjugation machinery of the IncP1α plasmid.
In order to validate the correlation of the conjugation efficiency of these up-mutants with tra and trb genes expression, the basal expression levels of the selected tra and trb genes (traI, traJ, traK, and trbL) harbored within the pRH220 helper plasmid in the donor cells were compared between mutant and parental strains. These genes were selected as representatives of the three major operons within the RP4 IncP1α plasmid under the regulation of three major promoters: PtraJ, PtraK, and PtrbB (Pansegrau et al., 1994). As shown in Figure 3A, the expression of traI, traJ, traK, and trbL in the donor mutant strains of ΔfrmR, ΔsufA, and ΔiscA showed no significant increase in comparison with the parental strain.
FIGURE 3
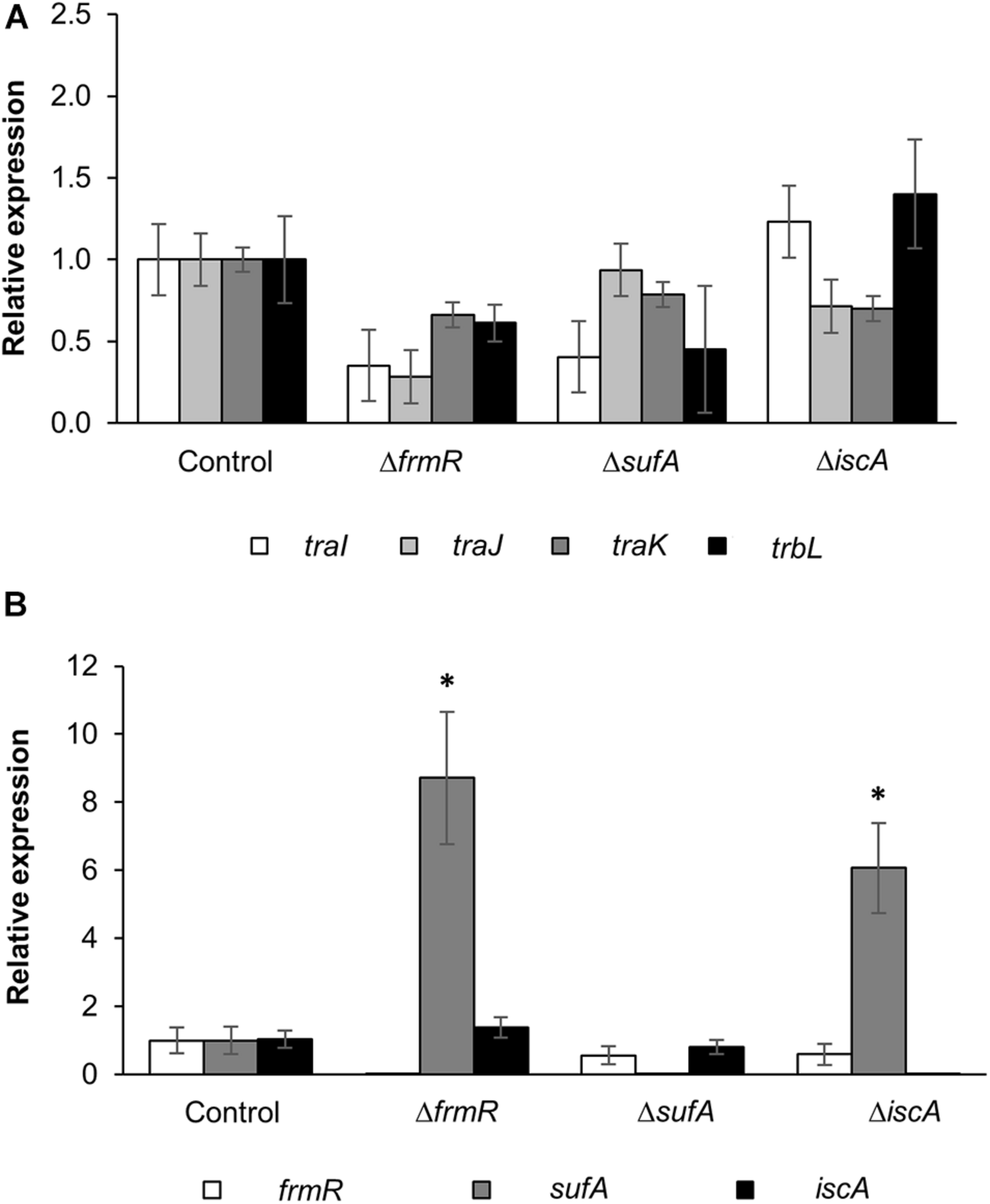
Expression analysis of conjugation-related and up-mutant genes by qRT-PCR. (A) Expression of traI, traJ, traK, and trbL genes within the helper plasmid, IncP1α-pRH220, harbored in the up-mutants and wild-type control donor strains, within triplicate experiments (n = 3). (B) Expression of frmR, sufA, and iscA genes within the up-mutant and wild-type control donor strains, within six experimental replicates (n = 6). Data are presented as mean ± standard error of the mean (SEM). Asterisk (∗) indicates statistically significant differences at p < 0.05 (two-tailed t-test) compared to wild-type control. BW25113 parental strain was used as the control.
In order to assess the correlation between the basal gene expression of sufA, iscA, and frmR with conjugation efficiency, the expression of these genes within the up-mutant donor strains was evaluated (Figure 3B). Real-time PCR analysis revealed that iscA expression in both ΔsufA and ΔfrmR mutants, as well as frmR expression in ΔsufA and ΔiscA, were not significantly different and comparable to the expression levels of the parental strain. These results indicate that iscA and frmR in the mutants expressed at the same level as in the E. coli parental strain. In addition, sufA gene expression in both ΔiscA and ΔfrmR was significantly high (approximately six- and ninefold, respectively) compared to the parental strain. However, no complementary effect on the repression of conjugation efficiency in these two mutants was observed (Figure 2C). Therefore, no clear transcriptional interaction among the three genes, which links conjugation efficiency, was observed.
On the basis of the results from single- and double-KO mutant analyses, it may be suggested that defects of FrmR, SufA, and IscA can affect independently to terminate their repression of IncP1α plasmid conjugation, but probably act on the identical step(s) of conjugation machinery.
The Enhancement of Conjugation Efficiency by Up-Mutants Specifically Affects IncP1-Type Plasmids Transfer
In order to observe the generality of the effect of these E. coli up-mutant strains on conjugation efficiency of the broad-host-range plasmids, the conjugation efficiency of IncN, IncW, and IncP1β to recipient cells was assessed and compared with that of the parental strain.
On the basis of the results obtained regarding the conjugation efficiency of IncN (R46) and IncW (pSa) to E. coli recipient cells, no significant difference was observed compared to the parental strain (Figures 4A,B). By contrast, an increase in conjugation efficiency was observed in both E. coli – E. coli and E. coli – yeast compared to the parental strain (seven–ninefold and three–fivefold, respectively), by measuring the transfer of an IncQ plasmid derived E. coli–yeast shuttle vector (pAY205) facilitated by IncP1β (pDPT51) (Figures 4C,D). No significant difference were observed among the up-mutant strains (Figures 4C,D). These results suggest that the enhancing effect of these mutants on conjugation efficiency in both prokaryotes and eukaryotes is probably specific to IncP1-type T4SS.
FIGURE 4
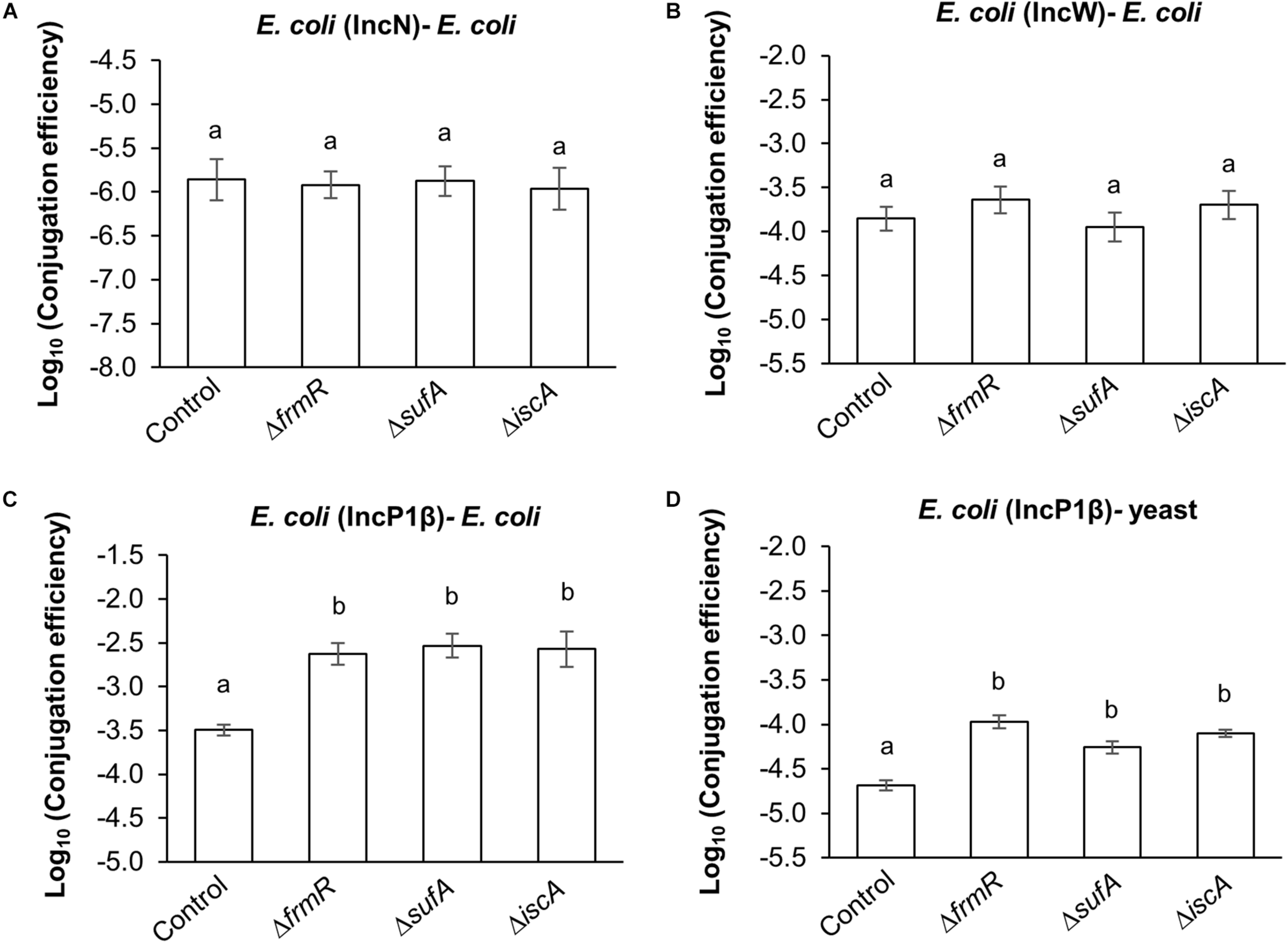
Generality assessment of frmR, sufA, and iscA mutations on the conjugation of broad host range plasmids. (A) Conjugation efficiency of IncN (R46) plasmid transfer to E. coli recipient cells within five experimental replicates (n = 5). (B) Conjugation efficiency of IncW (pSa) plasmid transfer to E. coli recipient cells within seven experimental replicates (n = 7). (C) Conjugation efficiency of IncP1β (pDPT51)-mediated shuttle vector (pAY205) transfer to E. coli and (D) yeast within triplicate experiments (n = 3). BY4742 and SY327 were used as the recipients. All conjugation reactions were performed for 1 h. Data are presented as mean ± standard error of the mean (SEM). Different letters indicate significant differences between mutants and wild-type control at p < 0.05 using Tukey HSD multiple comparison analysis. BW25113 parental strain was used as the control.
ΔATU_RS04380, ΔATU_RS08390, and ΔATU_RS08905 Mutants for Agrobacteria Homologs of the Up-Mutant Genes Promote Trans-Kingdom Conjugation
In order to assess the generality effect of up-mutant genes on IncP1-type T4SS-mediated plasmid transfer in other bacterial species, TKC analysis was performed by combining an IncQ mobilizable plasmid (pYN402) transfer system mediated by an IncP1α-type plasmid (RP4) and mutants of A. tumefaciens.
The homologous E. coli up-mutant genes in A. tumefaciens strain C58 were selected based on BlastP score analysis, phylogenetic trees, and previously reported study (Higgins and Giedroc, 2014; Chen et al., 2016; Heindl et al., 2016). According to the BlastP analysis, the homologous gene of frmR (ATU_RS04380) had the highest similarity (46.03% amino acid identity and 2e–15 e-value). In addition, the shared homologous genes of sufA and iscA: ATU_RS08390 and ATU_RS08905 (sufA:37.14% and 43.69% amino acid identity; 6e–23 and 9e–26 e-value, respectively) as well as (iscA: 37.14% and 39.62% amino acid identity; 3e–23 and 4e–24 e-value, respectively) carry out the same function in E. coli and are representative members of the iron–sulfur cluster assembly.
All examined mutants showed significantly higher TKC efficiency compared to the parental strain (Figure 5). The ΔATU_RS08390 mutant showed a fourfold increase in TKC efficiency compared to the parental strain. In addition, threefold increases in conjugation efficiency were observed in ΔATU_RS04380 and ΔATU_RS08905 mutants (Figure 5). These results suggest that these homologous mutant genes (ATU_RS04380, ATU_RS08390, and ATU_RS08905) have similar characteristics in terms of enhancing the TKC efficiency facilitated by IncP1-type T4SS machinery to the recipient cell.
FIGURE 5
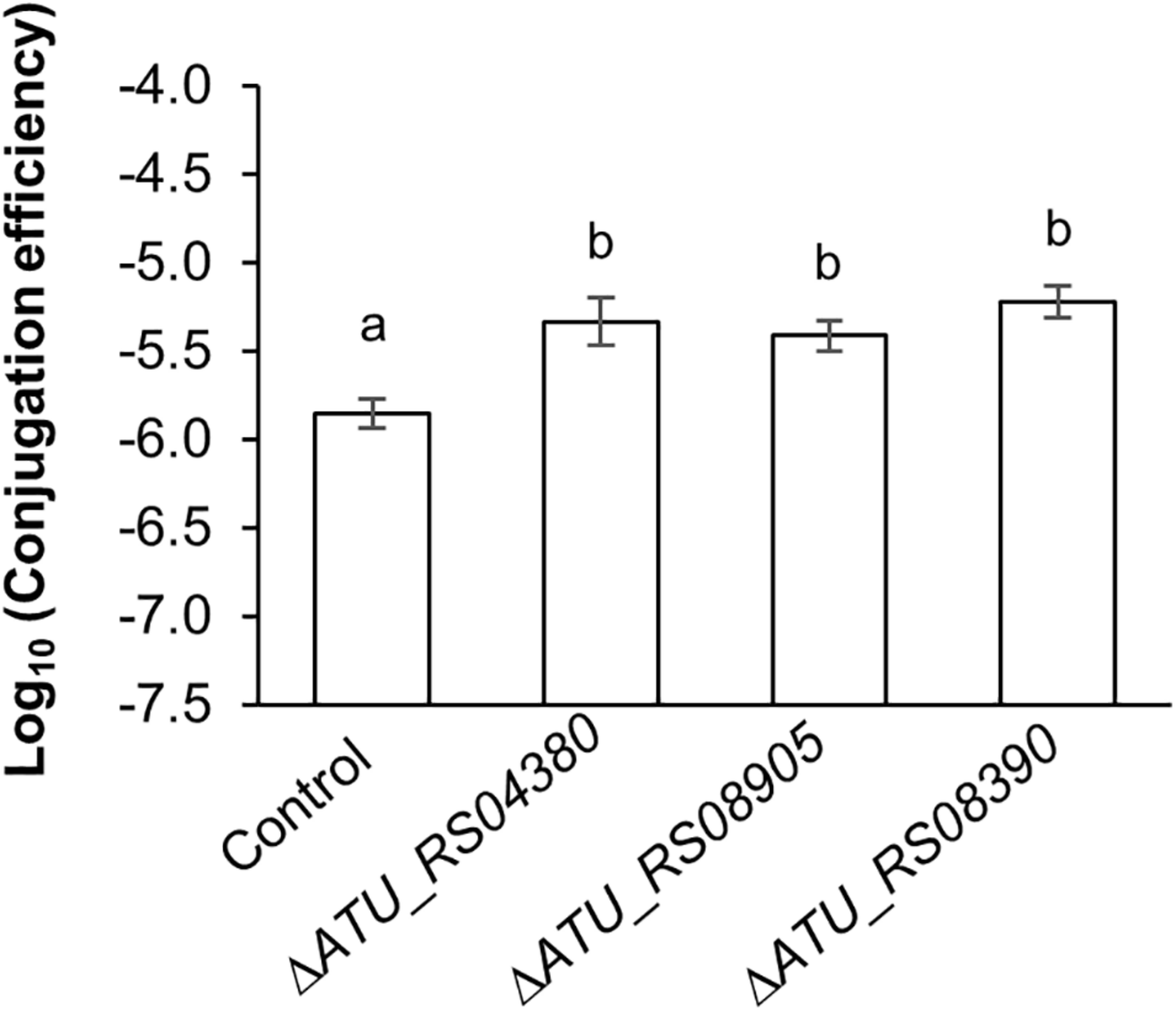
Effect of up-mutant homologs gene-knockout in A. tumefaciens on IncP1-type conjugation. The conjugation reaction was performed for 1 h. Data are presented as mean ± standard error of the mean (SEM) within six experimental replicates (n = 6). Different letters indicate significant differences between mutants and wild-type control at p < 0.05 using Tukey HSD multiple comparison analysis. BY4742 was used as the recipient. C58C1 parental strain was used as the control.
Discussion
The enhancing effect of the three mutants to both prokaryotes and eukaryotes is probably IncP1-type T4SS-dependent, and it was commonly observed in two donor species belonging to different classes (Figures 1, 4C,D, 5).
FrmR is a formaldehyde-sensing transcriptional repressor of the frm operon (Denby et al., 2016; Osman et al., 2016). On the basis of the experimental result shown in Figure 2A, no additional effect on conjugation efficiency was observed in the ΔfrmR mutant due to the addition of formaldehyde. These results suggested that the absence or inactivation of FrmR from the frm operon either due to a gene deletion mutation or the binding of this protein to the excessive formaldehyde, respectively, resulted in an increase in conjugation efficiency. In the case of TKC efficiency by the gene mutation within the frm operon (Figure 2B), single-KO of both ΔfrmA and ΔfrmB conferred no significant difference compared to ΔfrmR. This was probably due to the accumulation of endogenous ligands, including formaldehyde, caused by the failure of the detoxification mechanism by FrmA and FrmB within the cells, which inactivated FrmR, leading to the increase in conjugation efficiency (Figure 6A). A previous study showing that the deletion of frmA causes increase in basal frmR promoter activity as well as improved sensitivity to formaldehyde supports our expectation (Woolston et al., 2018). In the case of the double-KO mutants, ΔfrmAΔfrmR and ΔfrmBΔfrmR, no additional increase in TKC efficiency was observed between these mutants and single-KO mutants. On the basis of this status, we propose that FrmR represses the expression of other target factor(s) within the E. coli donor which may increase the conjugation efficiency (Figure 6 and Supplementary Figure 5).
FIGURE 6

Possible model mechanisms of the FrmR, SufA, and IscA protein interactions within E. coli donor in repressing the conjugation of IncP1α plasmid. (A) Feedback regulation of frm operon. FrmR is a transcriptional repressor of the frm operon. The accumulation of ligands (e.g., formaldehyde) causes the inactivation of FrmR repressor activity by binding to the ligands, consequently activates the transcriptional activity of this operon. This transcriptional activation leads to the expression of the downstream genes, frmA and frmB, which encode FrmA and FrmB for formaldehyde detoxification. (B,C) FrmR is also predicted to be a transcriptional repressor on the operon of another target factor (activator) within the E. coli donor which represents as factor Z. IscA and SufA are predict to work in repressing the activators (factors V and U, respectively) either by directly or indirectly (B) or directly activate the repressors (factors S and T, respectively) (C). At the same time, both of these activators may form a complex with the FrmR target factor (Z) in order to activate conjugation (B) or the repressor may form a complex with other factor (factor R) repressed by factor Z either directly or indirectly, resulting to the repression of conjugation (C), either by directly or indirectly. Based on these model mechanisms, the FrmR, SufA, and IscA proteins are possible to repress the conjugation at the identical step(s) of IncP1 conjugation machinery.
A previous study reported that the autoinducers, N-acyl homoserine lactones (AHLs), such as N-(3-oxododecanoyl)-L-homoserine lactone (OdDHL), produced by Pseudomonas aeruginosa PAO1, are capable to repress the conjugation of an IncP1α plasmid (RP4). The repressing mechanism was explained to be caused by a decrease in the expression of the traI conjugation-related gene in the RP4 plasmid, which results from transcriptional repression by AHLs-SdiA in donor E. coli, by binding to the SdiA-box located at the promoter sequence of traI (Lu et al., 2017). However, E. coli does not produce AHLs, and no significant effect was observed on conjugation efficiency even though the OdDHL was exogenously supplied in the conjugation reaction mixture (Supplementary Figure 4). On the basis of this status, at least under our experimental conditions, it can be concluded that the increase in conjugation efficiency by the up-mutants is not related to the autoinducer-mediated mechanism.
FrmR belongs to the CsoR/RcnR metal ion-sensing transcriptional repressor family, and the family phylogenetically consists of three clades, namely, CsoRs, RcnRs, and FrmRs (Chen et al., 2016). In E. coli, frmR and rcnR genes are encoded on its genome, and only the ATU_RS04380 gene, which is in the CsoR clade, has been found in the A. tumefaciens C58 genome. This suggests that, although it is possible that CsoR/RcnR family proteins might affect IncP1 plasmid conjugation, the screening and results shown in Figures 1A, 5 indicate that FrmR has specificity for the regulation of IncP1 plasmid conjugation in E. coli.
In E. coli, the iscA, sufA, and erpA genes are paralogs and coding members of iron–sulfur cluster carrier proteins having overlapping functions (Loiseau et al., 2007; Roche et al., 2013). The KO mutant of erpA is not included in the Keio library because of its essentiality. In addition, double-KO of ΔiscAΔsufA genes result in synthetic lethality under aerobic condition (Vinella et al., 2009). Thus, this double-KO mutant was excluded from this conjugation assessment, and the KO mutation of either of these genes with ΔfrmR was constructed to confirm the gene interaction. The conjugation efficiency of double-KO mutants ΔiscAΔfrmR and ΔfrmRΔsufA did not exhibit any synergistic increase in conjugation efficiency and was comparable with that of ΔiscA, ΔsufA, and ΔfrmR single-KO mutants (Figure 2C). The loss of expression of any of the three up-mutant genes did not lead to attenuated expression of other two genes (Figure 3B). Based on this status, we predicted that FrmR, SufA, and IscA target to different unknown factor(s) (activator or repressor) within the E. coli donor cells and independently affect the conjugation mechanism. At this status, we predict that the defect of FrmR, SufA, and IscA probably target to the activator(s) which may direct or indirectly activate the conjugation mechanism. In addition, SufA and IscA are also predicted to work in activating or repressing the conjugation mechanism indirectly, either by repressing or activating respective unknown target factor(s). This prediction was made since no decreasing effect in conjugation efficiency (or at comparable level to that of parental strain) was observed in ΔsufA and ΔiscA single- as well as ΔiscAΔfrmR and ΔfrmRΔsufA double-KO mutants, regardless in the presence of ΔfrmR. This probably due to the absence of complementation effect between both SufA and IscA. Thus, we predict that both SufA and IscA are probably necessary in activating or repressing the conjugation mechanism indirectly with unknown target factor of FrmR.
These results suggest that the unknown target factors of these three genes form a complex in order to activate or repress the conjugation, either by directly or indirectly at an identical step(s) of IncP1 conjugation machinery although the exact mechanism beyond this phenomenon remains unknown. Since KO mutants for iscA, sufA, and their agrobacterial homologs have been commonly shown to increase conjugation efficiency (Figures 1, 5), iron–sulfur cluster delivery deficiency probably causes a common physiological status, which specifically promotes the IncP1-type conjugation system (Figures 6B,C and Supplementary Figures 5A,B).
On the basis of the results from single- and double-KO conjugation experiment and the relation with basal gene expression, as well as the known functions of FrmR, SufA, and IscA, we propose models for the repression mechanism of the IncP1-type conjugation system (Figure 6). We deduce that SufA and IscA work in repressing other target factors (activators) within the E. coli donor either by directly or indirectly. At the same time, the inactivation of FrmR, which may also be a repressor of other target factor (activator), will derepress the expression of that particular factor. The unknown target factors of FrmR, IscA, and SufA may form a complex to activate the conjugation, either by directly or indirectly at an identical step of IncP1 conjugation machinery (Figure 6B). At present, there are still several models that we deduce how these three gene products interact with other unknown target factors in the process of IncP1α conjugation which fits to our results (Figure 6C and Supplementary Figure 5).
Lastly, IncP1-type T4SS carries high potential for the application of a gene introduction system into various organisms (Dodsworth et al., 2010; Norberg et al., 2011; Moriguchi et al., 2013a). The data regarding the mutants isolated in this study should be an appropriate basis for the breeding of donor strains from various proteobacteria, each of which carries high cytological affinity with target organisms in addition to high conjugation ability. Further characterization in terms of possible gene interaction within the chromosomal mutants based on physiological analysis with the possible regulators within the IncP1 plasmids will lead for better understanding of the isolated genes’ diversity with the TKC mechanism. Additionally, it will be interesting to determine the specificity of T4SS-mediated IncQ conjugal transfer in the isolated mutant strains by using Ti plasmid (VirB/D4-T4SS system) in comparison to IncP1-type system as it has been reported to serve as a delivery system in Agrobacterium (Bohne et al., 1998; Ohmine et al., 2018; Kiyokawa et al., 2020).
Statements
Data availability statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation, to any qualified researcher.
Author contributions
KM, SY, and KS conceived and designed the experiments. FZ, KM, and YC performed the experiments and analyzed the data. SY and KK contributed the reagents and materials. FZ and KM wrote the manuscript. FZ, KM, YC, SY, KK, and KS revised the manuscript. All authors read and approved the manuscript.
Funding
This work was supported in part by the Japan Society for the Promotion of Science KAKENHI (Grant Number JP16K07200).
Acknowledgments
We express our gratitude to the National BioResource Project (NBRP) of the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan, for providing E. coli strains, including the Keio collection and the pK18mobsacB plasmid vector. We also would like to express our sincere gratitude and special thanks to Prof. Maruyama (Hiroshima University) for his kindness in lending the real-time PCR equipment throughout this work. We also would like to thank to Naoki Umei for his excellent assistance for the mutant screening experiment.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2021.620535/full#supplementary-material
Abbreviations
- AHL
acyl homoserine lactones
- HGT
horizontal gene transfer
- NBRP
National BioResource Project
- SD
synthetic defined
- SEM
standard error of the mean
- TKC
trans-kingdom conjugation
- TNB
Tris-HCl with NaCl buffer
- VGT
vertical gene transfer
- YPD
yeast-extract/peptone/dextrose.
References
1
AdamczykM.Jagura-BurdzyG. (2003). Spread and survival of promiscuous IncP-1 plasmids.Acta Biochim. Pol.50425–453. 10.18388/abp.2003_3696
2
BabaT.AraT.HasegawaM.TakaiY.OkumuraY.BabaM.et al (2006). Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection.Mol. Syst. Biol.2:2006.0008.
3
BatesS.CashmoreA. M.WilkinsB. M. (1998). IncP plasmids are unusually effective in mediating conjugation of Escherichia coli and Saccharomyces cerevisiae: involvement of the Tra2 mating system.J. Bacteriol.1806538–6543. 10.1128/jb.180.24.6538-6543.1998
4
BohneJ.YimA.BinnsA. N. (1998). The Ti plasmid increases the efficiency of Agrobacterium tumefaciens as a recipient in virB-mediated conjugal transfer of an IncQ plasmid.Proc. Natl. Acad. Sci. U.S.A.957057–7062. 10.1073/pnas.95.12.7057
5
BrownA. M. C.WillettsN. S. (1981). A physical and genetic map of the IncN plasmid R46.Plasmid5188–201. 10.1016/0147-619x(81)90020-2
6
ChenN. H.DjokoK. Y.VeyrierF. J.McEwanA. G. (2016). Formaldehyde stress responses in bacterial pathogens.Front. Microbiol.7:257. 10.3389/fmicb.2016.00257
7
DenbyK. J.IwigJ.BissonC.WestwoodJ.RolfeM. D.SedelnikovaS. E.et al (2016). The mechanism of a formaldehyde-sensing transcriptional regulator.Sci. Rep.6:38879.
8
DodsworthJ. A.LiL.WeiS.HedlundB. P.LeighJ. A.De FigueiredoP. (2010). Interdomain conjugal transfer of DNA from bacteria to archaea.Appl. Environ. Microbiol.765644–5647. 10.1128/aem.00967-10
9
Fernández-LópezR.Garcillán-BarciaM. P.RevillaC.LázaroM.VielvaL.De La CruzF. (2006). Dynamics of the IncW genetic backbone imply general trends in conjugative plasmid evolution.FEMS. Microbiol. Rev.30942–966. 10.1111/j.1574-6976.2006.00042.x
10
GarushyantsS. K.KazanovM. D.GelfandM. S. (2015). Horizontal gene transfer and genome evolution in Methanosarcina.BMC. Evol. Biol.15:102. 10.1186/s12862-015-0393-2
11
HaaseJ.LurzR.GrahnA. M.BamfordD. H.LankaE. (1995). Bacterial conjugation mediated by plasmid RP4: RSF1010 mobilization, donor-specific phage propagation, and pilus production require the same Tra2 core components of a proposed DNA transport complex.J. Bacteriol.1774779–4791. 10.1128/jb.177.16.4779-4791.1995
12
HaymanG. T.BolenP. L. (1993). Movement of shuttle plasmids from Escherichia coli into yeasts other than Saccharomyces cerevisiae using Trans-kingdom conjugation.Plasmid30251–257. 10.1006/plas.1993.1056
13
HeindlJ. E.HibbingM. E.XuJ.NatarajanR.BuechleinA. M.FuquaC. (2016). Discrete responses to limitation for iron and manganese in Agrobacterium tumefaciens: Influence on attachment and biofilm formation.J. Bacteriol.198816–829. 10.1128/jb.00668-15
14
HigginsK. A.GiedrocD. (2014). Insights into protein allostery in the csor/rcnr family of transcriptional repressors.Chem. Lett.4320–25. 10.1246/cl.130965
15
KamachiK.SotaM.TamaiY.NagataN.KondaT.InoueT. (2006). Plasmid pBP136 from Bordetella pertussis represents an ancestral form of IncP-1β plasmids without accessory mobile elements.Microbiology1523477–3484. 10.1099/mic.0.29056-0
16
KarasB. J.DinerR. E.LefebvreS. C.McQuaidJ.PhillipsA. P. R.NoddingsC. M.et al (2015). Designer diatom episomes delivered by bacterial conjugation.Nat. Commun.6:6925.
17
KiyokawaK.OhmineY.YunokiK.YamamotoS.MoriguchiK.SuzukiK. (2020). Enhanced Agrobacterium-mediated transformation revealed attenuation of exogenous plasmid DNA installation in recipient bacteria by exonuclease VII and SbcCD.Genes Cell.25, 663–674. 10.1111/gtc.12802
18
LacroixB.CitovskyV. (2016). Transfer of DNA from bacteria to eukaryotes.mBio7:e00863-16.
19
LoiseauL.GerezC.BekkerM.Ollagnier-De ChoudensS.PyB.SanakisY.et al (2007). ErpA, an iron-sulfur (Fe-S) protein of the a-type essential for respiratory metabolism in Escherichia coli.Proc. Natl. Acad. Sci. U.S.A.10413626–13631. 10.1073/pnas.0705829104
20
LuJ.YangJ.TanG.DingH. (2008). Complementary roles of SufA and IscA in the biogenesis of iron-sulfur clusters in Escherichia coli.Biochem. J.409535–543. 10.1042/bj20071166
21
LuY.ZengJ.WuB.ShunmeiE.WangL.CaiR.et al (2017). Quorum sensing N-acyl homoserine lactones-SdiA suppresses Escherichia coli-Pseudomonas aeruginosa conjugation through inhibiting traI expression.Front. Cell. Infect. Microbiol.7:7. 10.3389/fcimb.2017.00007
22
MoriguchiK.EdahiroN.YamamotoS.TanakaK.KurataN.SuzukiK. (2013a). Transkingdom genetic transfer from Escherichia coli to Saccharomyces cerevisiae as a simple gene introduction tool.Appl. Environ. Microbiol.794393–4400. 10.1128/aem.00770-13
23
MoriguchiK.YamamotoS.OhmineY.SuzukiK. (2016). A fast and practical yeast transformation method mediated by Escherichia coli based on a trans-kingdom conjugal transfer system: just mix two cultures and wait one hour.PLoS One11:e0148989. 10.1371/journal.pone.0148989
24
MoriguchiK.YamamotoS.TanakaK.KurataN.SuzukiK. (2013b). Trans-kingdom horizontal DNA transfer from bacteria to yeast is highly plastic due to natural polymorphisms in auxiliary nonessential recipient genes.PLoS One8:e74590. 10.1371/journal.pone.0074590
25
MoriguchiK.ZoolkefliF. I. R. M.AbeM.KiyokawaK.YamamotoS.SuzukiK. (2020). Targeting antibiotic resistance genes is a better approach to block acquisition of antibiotic resistance than blocking conjugal transfer by recipient cells: a genome-wide screening in Escherichia coli.Front. Microbiol.10:2939. 10.3389/fmicb.2019.02939
26
NishikawaM.YoshidaK. (1998). Trans-kingdom conjugation offers a powerful gene targeting tool in yeast.Genet. Anal. Biomol. Eng.1465–73. 10.1016/s1050-3862(97)10003-1
27
NorbergP.BergströmM.JethavaV.DubhashiD.HermanssonM. (2011). The IncP-1 plasmid backbone adapts to different host bacterial species and evolves through homologous recombination.Nat. Commun.2:268.
28
OhmineY.KiyokawaK.YunokiK.YamamotoS.MoriguchiK.SuzukiK. (2018). Successful transfer of a model T-DNA plasmid to E. coli revealed its dependence on recipient RecA and the preference of VirD2 relaxase for eukaryotes rather than bacteria as recipients.Front. Microbiol.9:895. 10.3389/fmicb.2018.00895
29
OsmanD.PiergentiliC.ChenJ.SayerL. N.UsónI.HugginsT. G. (2016). The effectors and sensory sites of formaldehyde-responsive regulator FrmR and metal-sensing variant.J. Biol. Chem.29119502–19516. 10.1074/jbc.m116.745174
30
PansegrauW.LankaE.BarthP. T.FigurskiD. H.GuineyD. G.HaasD. (1994). Complete nucleotide sequence of Birmingham IncPα plasmids.J. Mol. Biol.239623–663. 10.1006/jmbi.1994.1404
31
RegnardG. L.Halley-StottR. P.TanzerF. L.HitzerothI. I.RybickiE. P. (2010). High level protein expression in plants through the use of a novel autonomously replicating geminivirus shuttle vector.Plant Biotechnol. J.838–46. 10.1111/j.1467-7652.2009.00462.x
32
RichardD.RavignéV.RieuxA.FaconB.BoyerC.BoyerK. (2017). Adaptation of genetically monomorphic bacteria: evolution of copper resistance through multiple horizontal gene transfers of complex and versatile mobile genetic elements.Mol. Ecol.262131–2149. 10.1111/mec.14007
33
RocheB.AusselL.EzratyB.MandinP.PyB.BarrasF. (2013). Iron/sulfur proteins biogenesis in prokaryotes: formation, regulation, and diversity.Biochim. Biophys. Acta1827455–469. 10.1016/j.bbabio.2012.12.010
34
SchäferA.TauchA.JsgerW.KalinowskiJ.ThierbachbG.PiihlerA. (1994). Small mobilizable multi-purpose cloning vectors derived from the Escherichia coli plasmids pK18 and pK19: selection of defined deletions in the chromosome of Corynebacterium glutamicum.Gene14569–73. 10.1016/0378-1119(94)90324-7
35
SchmidhauserT. J.HelinskiD. R. (1985). Regions of broad-host-range plasmid RK2 involved in replication and stable maintenance in nine species of gram-negative bacteria.J. Bacteriol.164446–455. 10.1128/jb.164.1.446-455.1985
36
Shun-MeiE.ZengJ. M.YuanH.LuY.CaiR. X.ChenC. (2018). Sub-inhibitory concentrations of fluoroquinolones increase conjugation frequency.Microb. Pathog.11457–62. 10.1016/j.micpath.2017.11.036
37
SoltysiakM. P. M.MeaneyR. S.HamadacheS.JanakiramaP.EdgellD. R.KarasB. J. (2019). Trans-kingdom conjugation within solid media from Escherichia coli to Saccharomyces cerevisiae.Int. J. Mol. Sci.20:5212. 10.3390/ijms20205212
38
SuzukiH.YanoH.BrownC. J.TopE. M. (2010). Predicting plasmid promiscuity based on genomic signature.J. Bacteriol.1926045–6055. 10.1128/jb.00277-10
39
TaitR. C.LunquistR. C.KadoC. I. (1982). Genetic map of the crown gall suppressive IncW plasmid pSa.Mol. Gen. Genet.18610–15. 10.1007/bf00422905
40
TaylorD. P.CohenS. N.ClarkW. G.MarrsB. L. (1983). Alignment of the genetic and restriction maps of the photosynthetic region of the Rhodopseudomonas capsulata chromosome by a conjugation-mediated marker rescue technique.J. Bacteriol.154580–590. 10.1128/jb.154.2.580-590.1983
41
VinellaD.Brochier-ArmanetC.LoiseauL.TallaE.BarrasF. (2009). Iron-sulfur (Fe/S) protein biogenesis: phylogenomic and genetic studies of a-type carriers.PLoS Genet.5:e1000497. 10.1371/journal.pgen.1000497
42
WardL. M.HempJ.ShihP. M.McGlynnS. E.FischerW. W. (2018). Evolution of phototrophy in the Chloroflexi phylum driven by horizontal gene transfer.Front. Microbiol.9:260. 10.3389/fmicb.2018.00260
43
WillettsN.CrowtherC. (1981). Mobilization of the non-conjugative IncQ plasmid RSF1010.Genet. Res.37311–316. 10.1017/s0016672300020310
44
WinansS. C.WalkerG. C. (1985). Conjugal transfer system of the IncN plasmid pKM101.J. Bacteriol.161402–410. 10.1128/jb.161.1.402-410.1985
45
WoolstonB. M.RothT.KohaleI.LiuD. R.StephanopoulosG. (2018). Development of a formaldehyde biosensor with application to synthetic methylotrophy.Botechnol. Bioeng.115, 206–215. 10.1002/bit.26455
46
YamamotoS.UrajiM.TanakaK.MoriguchiK.SuzukiK. (2007). Identification of pTi-SAKURA DNA region conferring enhancement of plasmid incompatibility and stability.Genes Genet. Syst.82197–206. 10.1266/ggs.82.197
47
YanoH.RogersL. M.KnoxM. G.HeuerH.SmallaK.BrownC. J. (2013). Host range diversification within the IncP-1 plasmid group.Microbiol. (United Kingdom)1592303–2315. 10.1099/mic.0.068387-0
48
YeoH. J.YuanQ.BeckM. R.BaronC.WaksmanG. (2003). Structural and functional characterization of the VirB5 protein from the type IV secretion system encoded by the conjugative plasmid pKM101.Proc. Natl. Acad. Sci. U.S.A.10015947–15952. 10.1073/pnas.2535211100
49
ZhangS.WangY.SongH.LuJ.YuanZ.GuoJ. (2019). Copper nanoparticles and copper ions promote horizontal transfer of plasmid-mediated multi-antibiotic resistance genes across bacterial genera.Environ. Int.129478–487. 10.1016/j.envint.2019.05.054
50
ZhouK.ZhouL.LimQ. ’ZouR.StephanopoulosG.TooH. P. (2011). Novel reference genes for quantifying transcriptional responses of Escherichia coli to protein overexpression by quantitative PCR.BMC. Mol. Biol.12:18. 10.1186/1471-2199-12-18
Summary
Keywords
genome-wide screening, IncP1-type plasmid, trans-kingdom conjugation, type IV secretion system, horizontal gene transfer
Citation
Zoolkefli FIRM, Moriguchi K, Cho Y, Kiyokawa K, Yamamoto S and Suzuki K (2021) Isolation and Analysis of Donor Chromosomal Genes Whose Deficiency Is Responsible for Accelerating Bacterial and Trans-Kingdom Conjugations by IncP1 T4SS Machinery. Front. Microbiol. 12:620535. doi: 10.3389/fmicb.2021.620535
Received
23 October 2020
Accepted
06 April 2021
Published
20 May 2021
Volume
12 - 2021
Edited by
Clay Fuqua, Indiana University Bloomington, United States
Reviewed by
Andrew N. Binns, University of Pennsylvania, United States; Ian S. Barton, The Brody School of Medicine at East Carolina University, United States
Updates
Copyright
© 2021 Zoolkefli, Moriguchi, Cho, Kiyokawa, Yamamoto and Suzuki.
This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Kazuki Moriguchi, kmoriguc@hiroshima-u.ac.jp
†These authors have contributed equally to this work
This article was submitted to Evolutionary and Genomic Microbiology, a section of the journal Frontiers in Microbiology
Disclaimer
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article or claim that may be made by its manufacturer is not guaranteed or endorsed by the publisher.