- Biochemical Process Engineering, Division of Chemical Engineering, Department of Civil, Environmental and Natural Resources Engineering, Luleå University of Technology, Luleå, Sweden
Cellulases are a group of enzymes responsible for the degradation of cellulose, which is one of the most abundant polymers on Earth. The three main groups of cellulases are endoglucosidases, exoglucosidases, and β-glucosidases; however, the mechanism of induction of these enzymes remains poorly characterized. Cellooligosaccharides are among the main inducers of these enzymes in filamentous fungi, yet it is not clear how their degree of polymerization may affect the strength of induction. In the present study, we investigated the effect of different carbohydrate-based inducers, such as lactose, sophorose, cellooligosaccharides, and xylooligosacharides, characterized by different concentrations and degree of polymerization, on cellulases production by the fungus Fusarium oxysporum f. sp. lycopersici, which is one of the most studied lignocellulose degrading fungi with the ability to consume both cellulose and hemicellulose. Moreover, the effect of carbon source on cellulase induction was assessed by growing the biomass on sucrose or glycerol. Results showed a correlation between induction efficiency and the cellooligosaccharides’ concentration and size, as well as the carbon source available. Specifically, cellotetraose was a better inducer when sucrose was the carbon source, while cellobiose yielded a better result on glycerol. These findings can help optimize industrial cellulase production.
Introduction
An increasing human population and limited natural resources have forced a search for new solutions. The strong dependency of industrial and technological advances on energy has led to an increase in fossil fuel consumption. Data show that crude oil is the most consumed fuel (39%), followed by coal (33%) and natural gas (33%) (Ritchie and Roser, 2020). As this trend leads to the accumulation of hazardous pollutants and emission of greenhouse gases, which result in global warming, alternative resources should be investigated. Studies show bio-based energy and products can decrease net CO2 emission to zero (Moshkin et al., 2005; Zhang et al., 2006).
Agriculture and wood industry residues are an abundant and cheap substrate for the production of chemicals and biofuel. Cellulose, which is a homopolymer composed of repeating units of anhydrocellobiose (Zhang et al., 2006), is the main component of plant biomass (Park et al., 2011; Obeng et al., 2017). It is produced by photosynthesis either by higher plants and algae or by non-photosynthetic organisms, including some bacteria and fungi, marine invertebrates, slime molds, and amoebae (Zhang and Lynd, 2004). Long chains and sheets of cellulose are linked by hydrogen bonds and van der Waal’s forces (Zhang et al., 2006). Cellulose has a recalcitrant structure and its β-glucosidic bonds have a half-life of 5–8 million years at 25°C (Wolfenden and Snider, 2001; Zhang et al., 2006). This rigidity is an obstacle for efficient industrial use of cellulose.
Cellulose can be hydrolyzed either chemically or enzymatically. Whereas chemical hydrolysis requires harsh conditions and dilute mineral acids (Panagiotou et al., 2005), enzymatic hydrolysis is a milder process with higher glucose yield (Mielenz, 2001). Enzymatic conversion of cellulose to fermentable sugars is the most complicated part of the process (Donohoe and Resch, 2015). Cellulases, which are responsible for cellulose degradation, can be either cell-bound or free enzymes (Zhang and Zhang, 2013). They are classified in 13 families of glycoside hydrolases (1, 3, 5, 6, 7, 8, 9, 12, 26, 44, 45, 51, and 48) based on similarities between amino acid sequences and crystal structures (Schu, 2000; Zhang and Zhang, 2013). Exoglucanase (EC 3.2.1.91), endoglucanase (EC 3.2.1.4), and β-glucosidase (EC 3.2.1.21) are the three main groups of hydrolytic enzymes that collaborate with auxiliary enzymes, such as lytic polysaccharide mono-oxygenases and phosphorolytic enzymes (in some bacteria), to disintegrate cellulose (Eriksson et al., 1990; Zhang et al., 2006). Endoglucanases randomly hydrolyze β-1,4-glucosidic bonds of cellulose chains and produce chain ends, where exoglucanase can act to release glucose or cellobiose. This rate-limiting step results in the release of variously polymerized soluble sugars from the surface of solid substrate into the liquid phase. Finally, β-glucosidases hydrolyze the released cellobiose and prevent end-product inhibition (Zhang et al., 2006).
Cellulases account for around 20% of the enzyme market (Bhat, 2000; Singhania et al., 2010). They are the third most consumed enzymes in industry, and the growing demand for bioenergy and bio-based materials promises a bright future for cellulase production (Rani et al., 2010; Chandel et al., 2012; Yoon et al., 2014). One of the main hindrances to the use of cellulose in biorefinery is the costly large-scale production of cellulases (Zhang et al., 2006). Specifically, it is difficult to increase the volume of produced enzyme using cheap substrates or generate enzymes with higher stability and/or specific activity on solid substrates (Zhang et al., 2006). Based on estimates, nearly 25 cents of every dollar used to produce bioethanol are spent on enzyme production (Humbird et al., 2011; Huberman et al., 2016), with around 100 g of cellulase being consumed per gallon of ethanol (Zhang et al., 2006; Zhu and Zhang, 2009).
Cellulases exist in a variety of microorganisms, from yeast, bacteria, and fungi to protozoa, plants, and animals (Zhang and Zhang, 2013); however, their industrial production relies on bacterial or fungal fermentation (Yoon et al., 2014). Compared to bacteria, fungi possess more advanced cellulase systems (Cen and Xia, 1999). Moreover, owing to their exceptional ecological, biological, and morphological plasticity, they can adapt to different environments and resources (Selbmann et al., 2013). As the complete saccharification of lignocellulosic biomass is time-consuming and needs a high load of degrading enzymes (around 30–50 mg per gram of crystalline cellulose), only fungi can meet such requirements (Ali et al., 2016).
Fusarium oxysporum is a soil-born plant pathogen responsible for vascular wilt disease in various crops, as well as mycotoxin contamination of human and animal food (Pathology, 2003). It is capable of producing several cell-wall degrading enzymes and is the most studied species in the Fusarium genus regarding lignocellulose degradation (Agrios, 2005). Its importance derives from its applicability in consolidated bioprocessing, whereby enzyme production, biomass hydrolysis, and fermentation of hexoses and pentoses to ethanol can be achieved in a single step (Panagiotou et al., 2003; Xu et al., 2009; Ali et al., 2016). Several endoglucanase genes (located on chromosomes 3, 12, 11, 4, 8, 7), β-glucosidase genes (located on chromosomes 5, 11, 1, 8, 12, 9, 7, 13), and one exoglucanase gene (located on chromosome 1) are deposited with NCBI for Fusarium oxysporum f. sp. lycopersici. Three endoglucanase enzymes from F. oxysporum have been characterized; one of them has a molecular weight of only 23.2 kDa, which facilitates penetration through cellulosic fibers (Kekos et al., 1995). Characterization of a β-glucosidase (Christakopoulos et al., 1991, 1995a), has revealed high β-glucosidase activity in this species that prevents end-product inhibition by cellobiose (Panagiotou et al., 2005). Studies on F. oxysporum strain 11C have indicated a highly efficient carbohydrate metabolism and ethanol production that depends on energy, protein, and sugar transport (Ali et al., 2014).
Despite numerous studies on the mechanism of cellulose degradation in this fungus, the machinery of cellulose sensing and enzyme induction/inhibition remains vague. This knowledge would allow researchers to modify and optimize biomass degradation by fungi, producing the desired product at high yields and with low byproducts. Most of the earlier investigations were based on the application of complex plant material, such as wheat straw and wheat bran, as carbon source with only few studies to use pure inducers. Trichoderma reesei, which is the most well known industrial cellulase producer, is influenced by substrate type (Els and Reese, 1956; Mandels and Reese, 1959; Mandels et al., 1961; Castro et al., 2014). Insoluble complex substrates, including plant biomass or crystalline cellulose, are good inducers of CAZymes (Alazi and Ram, 2018); however, they cannot be absorbed by mycelia and, therefore, they first need to be partially degraded to di- or monosaccharides such as cellooligosaccharides or their positional isomers such as sophorose (Alazi and Ram, 2018) prior to inducing cellulase genes. Sophorose, produced via transglycosylation during cellulose hydrolysis by an extracellular β-glucosidase, is the strongest cellulase inducer of T. reesei (Leite and Eveleight, 1989), but has no effect on Aspergillus niger (Gielkens et al., 1999) or Phanerochaete chrysosporium (Ulmer et al., 1984). Cellobiose induces cellulolytic enzymes in Neurospora crassa, T. reesei, and Aspergillus species but not in P. chrysosporium (Mandels and Reese, 1959; Vaheri et al., 1979; Chikamatsu et al., 1999; Znameroski et al., 2012), in which, instead, cellotriose and cellotetraose are excellent stimulants (Suzuki et al., 2010). In Polyporus arcularius, glucose, cellobiose, cellotriose, and cellotetraose repress enzyme production, while cellopentaose has the opposite effect (Ohnishi and Nagase, 2007). This variability shows the absence of a general rule and each species relies on specific inducers. Lactose, a cheap byproduct of some industries, has been used for cellulase production by T. reesei (Morikawa et al., 1995) and Acremonium cellulolyticus (Fang et al., 2008). Xylose can induce some cellulase genes in A. niger (Gielkens et al., 1999). In contrast, not many inducers have been documented in F. oxysporum and the mechanism of induction remains vague.
In this study, we aimed to identify effective inducers in F. oxysporum by examining some well-known inducers from other species, as well as cellooligosaccharides and xylooligosacharides with different degrees of polymerization and concentration. The obtained result facilitate toward better understanding of the mechanisms of induction mechanisms and the effect of the type of lignocellulosic biomass compounds on the induction of cellulolytic enzymes.
Materials and Methods
Microorganism and Preculture
All chemicals used in the current work were purchased from Sigma-Aldrich (St. Louis, MO, United States) unless otherwise stated. Sophorose, cellooligosaccharides, and xylooligosacharides were purchased from Megazyme, Bray, Ireland. Fusarium oxysporum f. sp. lycopersici (CBS 123668) was obtained from Centraalbureau voor Schimmelcultures (CBS; Utrecht, The Netherlands) and maintained on potato dextrose agar plates (39 g/L). The inoculum was prepared by cultivating the fungus for 2 days at 29°C and 190 rpm in 500 mL flasks containing 100 mL medium, which consisted of 20 g/L malt extract (MERCK; Kenilworth, NJ, United States) and 5 g/L yeast extract (MERCK).
Enzyme Induction Experiment
For the induction experiment, the fungus was first grown on medium containing 0.2 M sodium phosphate buffer, 1 g/L KH2PO4, 0.3 g/L MgSO4⋅7H2O, 10 g/L (NH4)2HPO4, and 1% w/v sucrose as carbon source, with pH adjusted to 6.3. This medium was inoculated with 6% v/v of the preculture and then incubated at 29°C and 190 rpm for 2 days. Biomass was harvested through a sterile vacuum filter (0.45 μm; Sarstedt, Nümbrecht, Germany), aseptically washed twice with 0.9% saline solution, and inoculated in the same medium without the carbon source. The fungus was incubated for 1 day to deplete any residual sugar from the mycelium. Then, the inducers were added to the medium: 0.2 and 0.3% w/v lactose; 0.1, 0.2, and 0.3% w/v cellobiose; 0.1 and 0.2% cellotetraose; 0.1 and 0.2% cellohexaose; 0.1% w/v xylobiose; 0.1% w/v xylotetraose; 0.1% w/v xylohexaose; and 0,1% w/v sophorose. Induction was followed over 3 days, in which each sampling point was a separate flask and the whole culture was harvested to minimize sampling errors. The samples were filtered through glass-microfiber discs (grade MGA; Sartorius, Gottingen, Germany) and both the filtrate and the biomass were collected to determine secreted and cell-bound enzyme activity, respectively. To determine the influence of carbon source on enzyme induction, the same experiment was repeated using glycerol as carbon source and cellooligosaccharides as inducers.
Enzymatic Activity Measurements
The activities of excreted enzymes including endoglucanase, exoglucanase, and extracellular β-glucosidase were measured using cell filtrates as a source of enzyme. For the endoglucanase assay, 200 μL of 1% w/v CMC solution and 50 μL of supernatant were incubated at 40°C for 15 min. The released reducing sugars were measured by the dinitrosalicylic acid method and compared to a glucose standard curve. The exoglucanase assay measured both the amount of released reducing sugars using the dinitrosalicylic method and 1% w/v microcrystalline cellulose solution (Avicel) as substrate, as well as the amount of p-nitrophenyl (pNP) released from 1 mM pNPC. Finally, β-glucosidase activity was measured using 1 mM solutions of p-nitrophenyl β-D-glucopyranoside (pNPG). When assaying exoglucanase and β-glucosidase activities with the pNP method, 100 μL of the enzyme solution was incubated with 900 μL of substrate at 40°C for 30 min. The reaction was stopped by the addition of 200 mL of 30% Na2CO3. All substrates were dissolved in phosphate-citrate buffer (pH 5). Cell-bound β-glucosidase activity was determined using 1.5 mg of air-dried intact cells as substrate and the pNPG assay. The activities are expressed as U/mL and U/mg for extracellular and cell-bound enzymes, respectively. All experiments were performed at least in three replicates, the results are presented as average and standard deviation.
Cellooligosaccharides Uptake Experiment
Cellooligosaccharides separation and detection in the medium and intracellular compartment was done by high-performance anion-exchange chromatography. The induction experiment was repeated using cellooligosaccharides as inducers and sampling was carried out after 1, 3, 5, and 7 h. For intracellular detection, the biomass was collected after 1 h, dried in a towel, grounded by liquid nitrogen, and washed with 5 mL water. Then, it was boiled for 10 min, followed by centrifugation (10,000 rpm, 10 min) and the supernatant was labeled as fraction A. The pellet was resuspended in 3 mL water, boiled for 4 h, and centrifuged. The resulting supernatant was labeled as fraction B. All samples were analyzed by Dionex ICS-5000 (Thermo Fisher Scientific, Waltham, MA, United States) with a pulsed amperometric detector equipped with a disposable electrochemical gold electrode, using a CarboPac PA1 4 × 250 mm analytical column and a CarboPac PA1 4 × 50 mm guard column, at 30°C.
Results
Sucrose as Carbon Source
Our preliminary results using sucrose as carbon source during the production of fungal biomass show that xylooligosacharides and lactose have either no influence or cause a negligible increase in enzymatic activity compared to the control. In addition, sophorose, which is known as a good inducer for some fungi, was ineffective for the induction of most enzymes, except exoglucanase. Based on these findings, inefficient inducers were excluded from the rest of the study, which instead focused on cellooligosaccharides. Moreover, because no exoglucanase activity was detected using Avicel as a substrate, p-nitrophenyl-β-D-cellobioside (pNPC) was employed in its place.
The highest endoglucanase activity was induced by 0.2% cellotetraose (Figure 1). The concentration of inducer appeared to be an important factor, because endoglucanase inductionwas nearly double at 0.2% compared to 0.1% cellotetraose. The same result was observed with cellobiose, whereby induction was highest at 0.3%, followed by 0.2 and 0.1%. Similarly, doubling the cellohexaose concentration from 0.1 to 0.2% induced activity as early as on day 1. A comparison of the results shows that, besides concentration, the degree of polymerization of the inducer was essential for the induction process. The lower activity achieved by adding cellohexaose compared to cellotetraose and cellobiose indicates that the optimum degree of polymerization for an inducer was four.
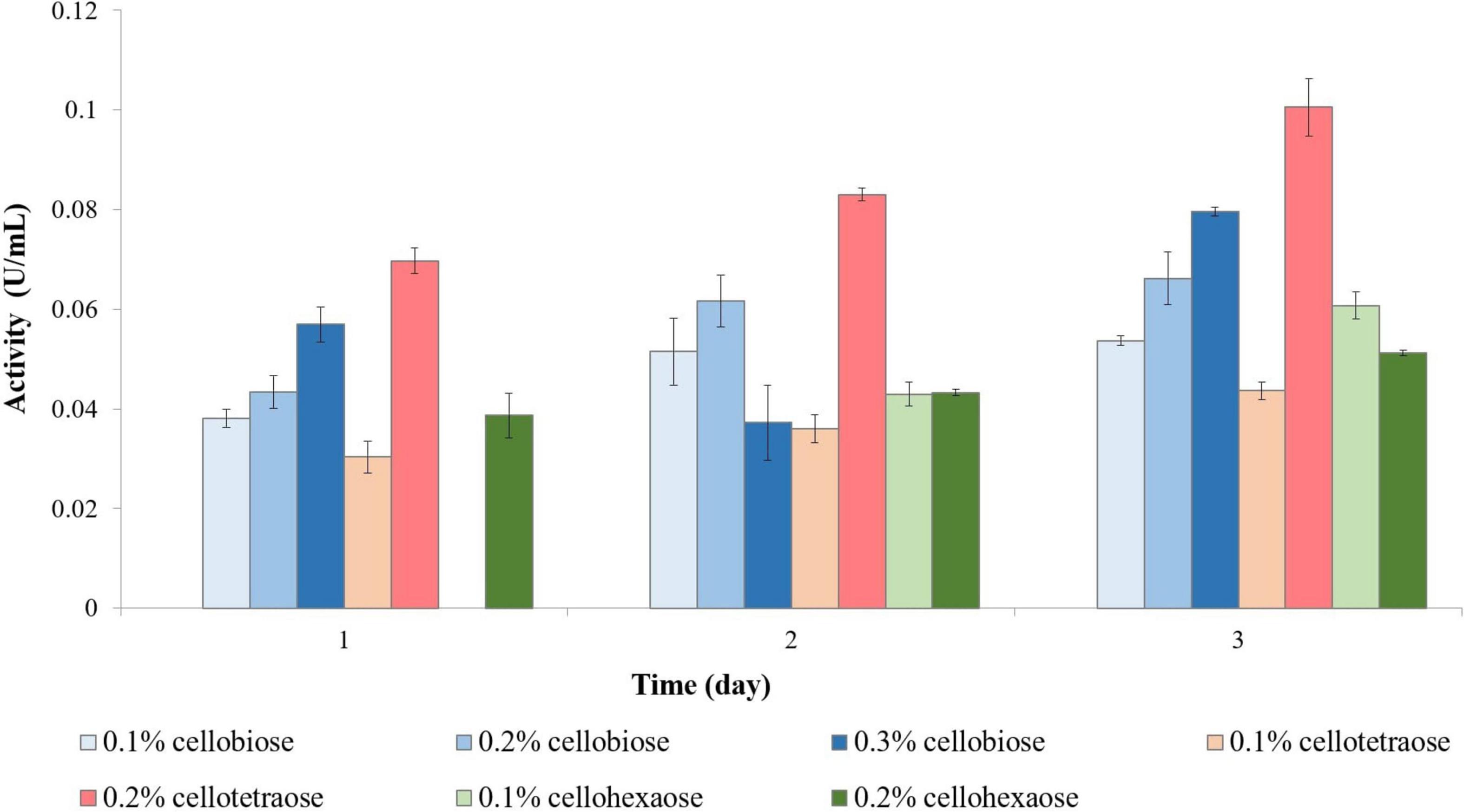
Figure 1. Endoglucanase activity of sucrose-grown biomass in the presence of different inducers and using CMC as substrate. No activity was detected in the presence of other inducers or the control sample.
Exoglucanase activity was highest with 0.2% cellotetraose, followed by 0.1% cellobiose and 0.1% sophorose. Inducer concentration was important for exoglucanase activity, albeit the effect differed between candidates. On the one hand, 0.2% was better than 0.1% cellotetraose; on the other hand, cellobiose and cellohexaose led to better outcomes at lower concentrations (Figure 2). For example, 0.1% cellobiose was effective already on day 1 and continued to be stronger than 0.2 or 0.3% cellobiose even in the following days, with 0.3% showing some induction only on day 3. Unlike endoglucanase, 0.1% sophorose caused a considerable increase in exoglucanase activity; whereas lactose had a negligible effect.
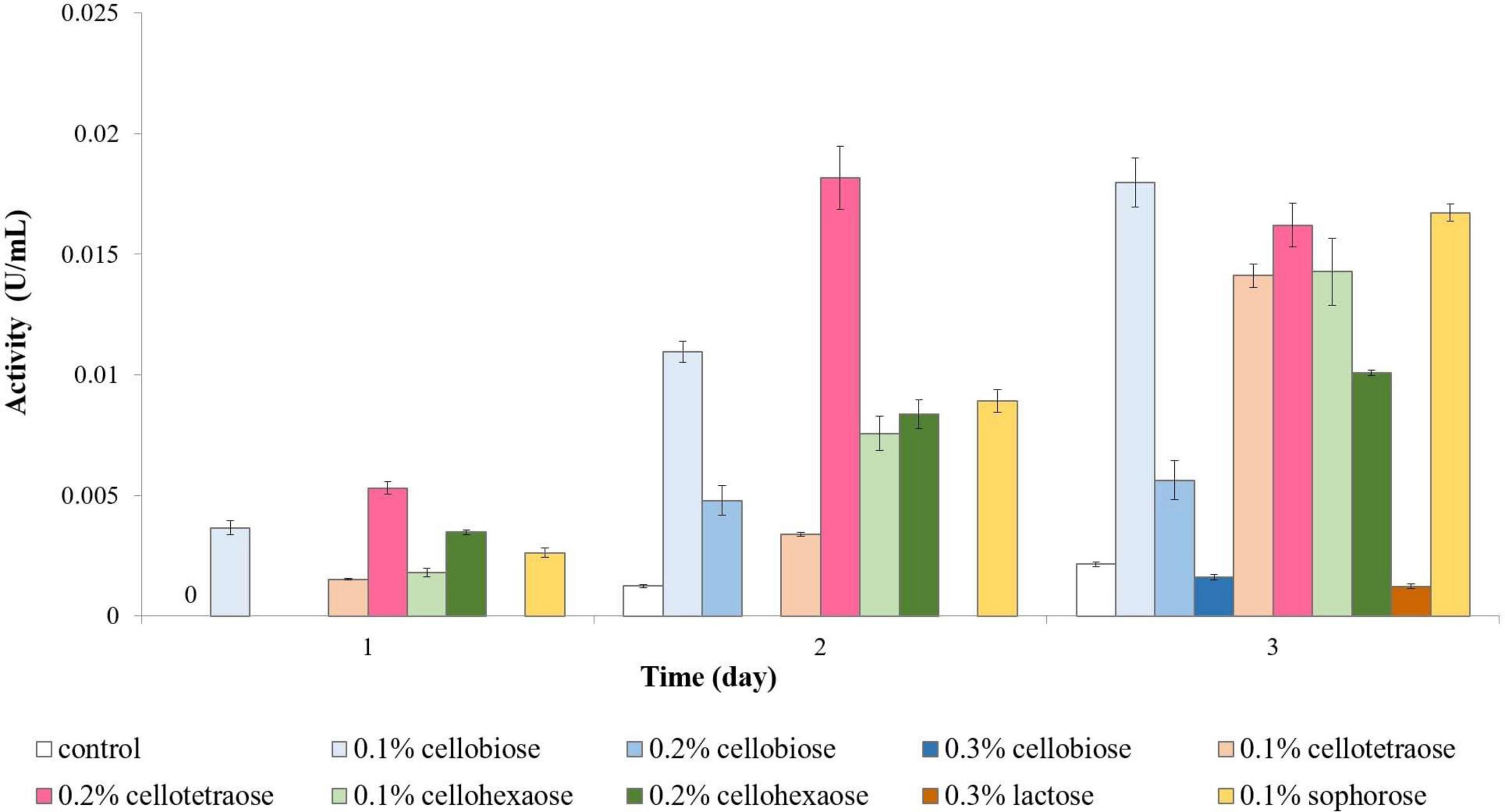
Figure 2. Exoglucanase activity of sucrose-grown biomass in the presence of different inducers and using pNPC as substrate. The sample containing 0.2% lactose failed to induce any activity.
Both extracellular and cell-bound β-glucosidase exhibited the highest activity in the presence of 0.2% cellotetraose, followed by 0.2% cellohexaose and 0.1% cellobiose (Figures 3A,B). As with other enzymes, concentration and degree of polymerization proved to be important factors defining the efficiency of each inducer in stimulating β-glucosidase activity. Confirming previous studies on other fungi, there is elevated cell wall-bound β-glucosidase activity in this study as well (Figure 3B; Umile and Kubicek, 1986; Tiwari et al., 2013).
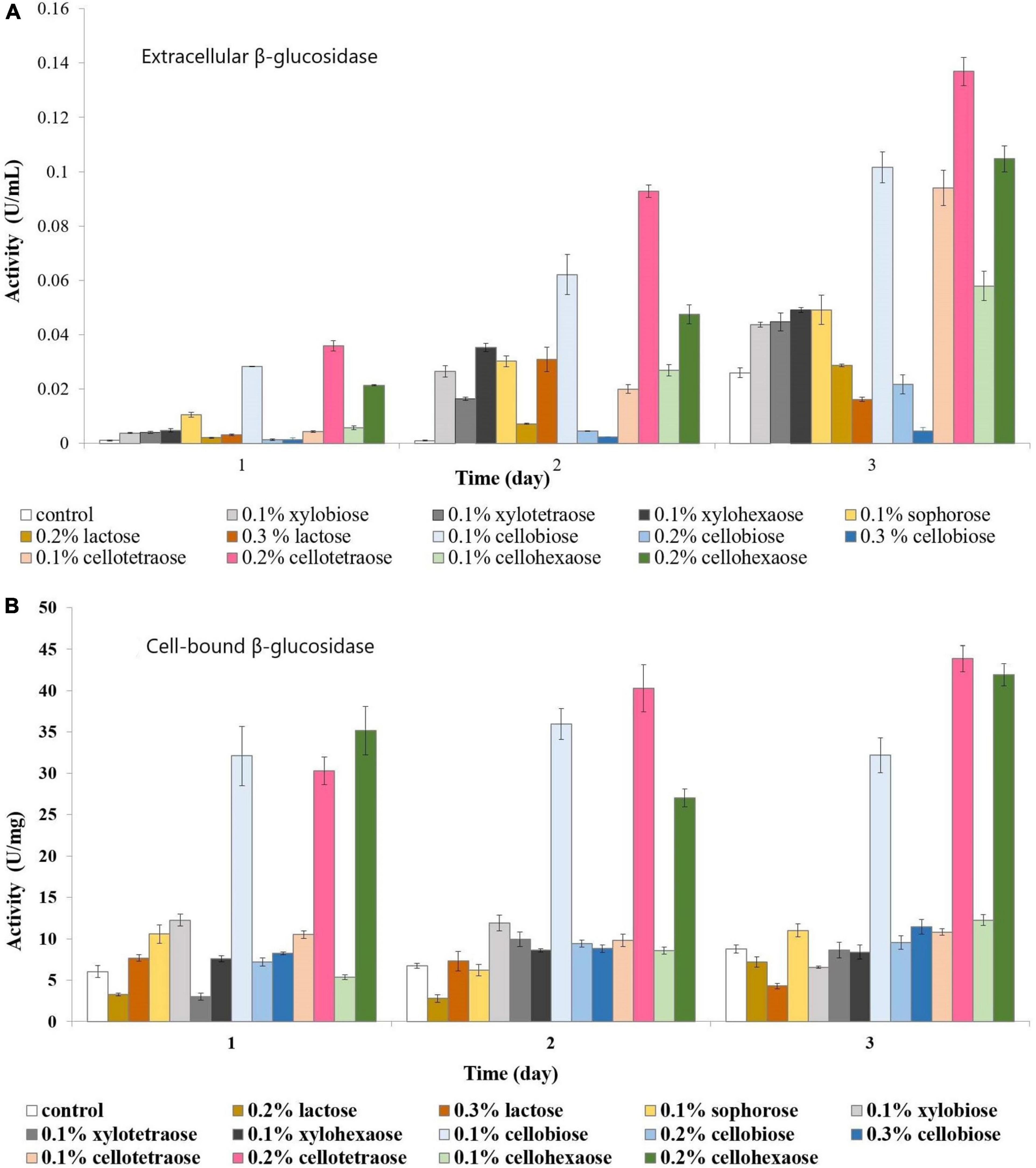
Figure 3. β-glucosidase activity of sucrose-grown biomass in the presence of different inducers and using pNPG as substrate. (A) Extracellular β-glucosidase activity; (B) Cell-bound β-glucosidase activity.
Uptake Experiment Using Sucrose as Carbon Source
Analysis of culture medium provides information about the degradation of all cellooligosaccharides starting with the first hour of induction. Figure 4 shows that the concentration of cellobiose gradually decreased as it was degraded to glucose (Figure 4 and Supplementary Figure S1). Figures 5, 6 and Supplementary Figures S2, S3) indicate that both cellotetraose and cellohexaose were immediately degraded to glucose and cellobiose. As the latter disappeared after 3 h when cellohexaose was added, it appears that cellohexaose was degraded or was uptaken faster than cellotetraose. After 1 h of induction, glucose was detected in all intracellular samples, whereas the cellobiose peak was minimal (Figure 7). There is no sign of cellotetraose and cellohexaose inside the cell that shows either they are first degraded to smaller units and imported to the cell, or are imported and degraded quickly to cellobiose and glucose. Christakopoulos et al., showed that the initial velocity of cello-oligosaccharides degraded by F. oxysporum endoglucanases increased with degree of polymerization (Christakopoulos et al., 1995b,c). Supplementary Figure S4 represents peaks of pure inducers.
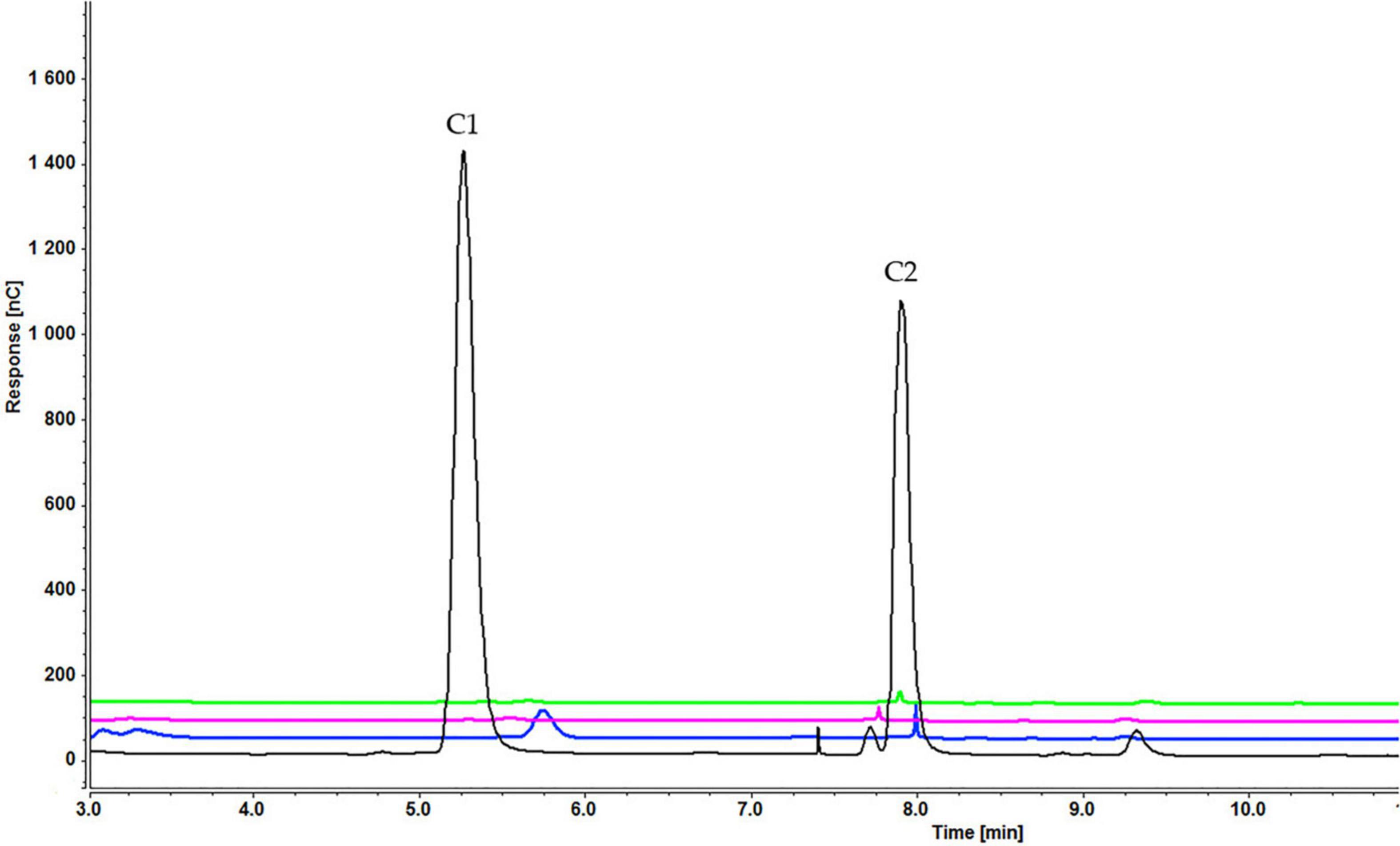
Figure 4. Cellooligosaccharides concentration in the medium after 1 h (black), 3 h (blue), 5 h (pink), and 7 h (green) of induction with 0.3% cellobiose (C1 = glucose, C2 = cellobiose). Figures for 0.1% (Supplementary Figure S1A) and 0.2% cellobiose (Supplementary Figure S1B) is presented in Supplementary Material.
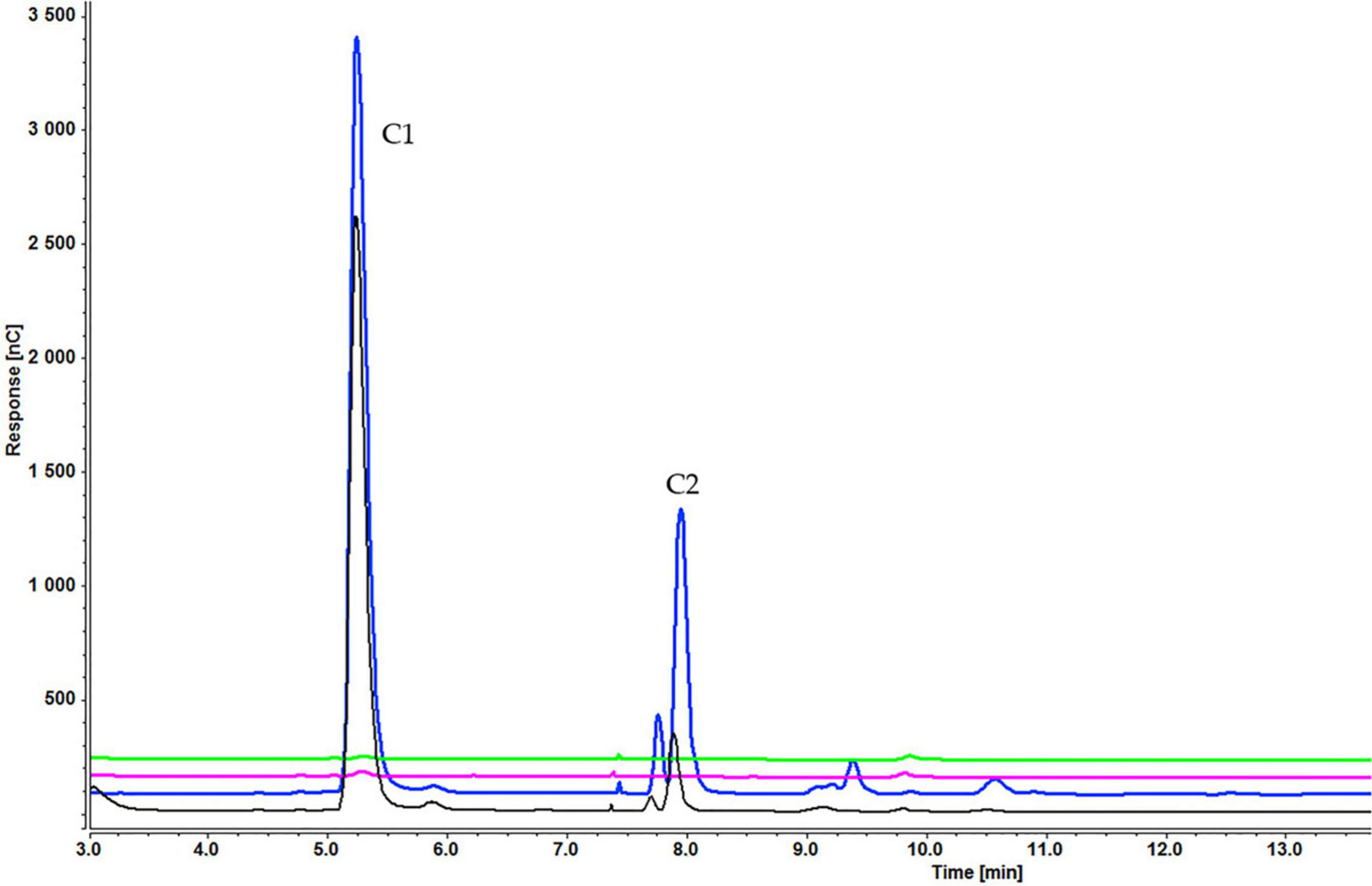
Figure 5. Cellooligosaccharides concentration in the medium after 1 h (black), 3 h (blue), 5 h (pink), and 7 h (green) of induction with 0.2% cellotetraose (C1 = glucose, C2 = cellobiose). The figure of 0.1% cellotetraose (Supplementary Figure S2) is presented in Supplementary Material.
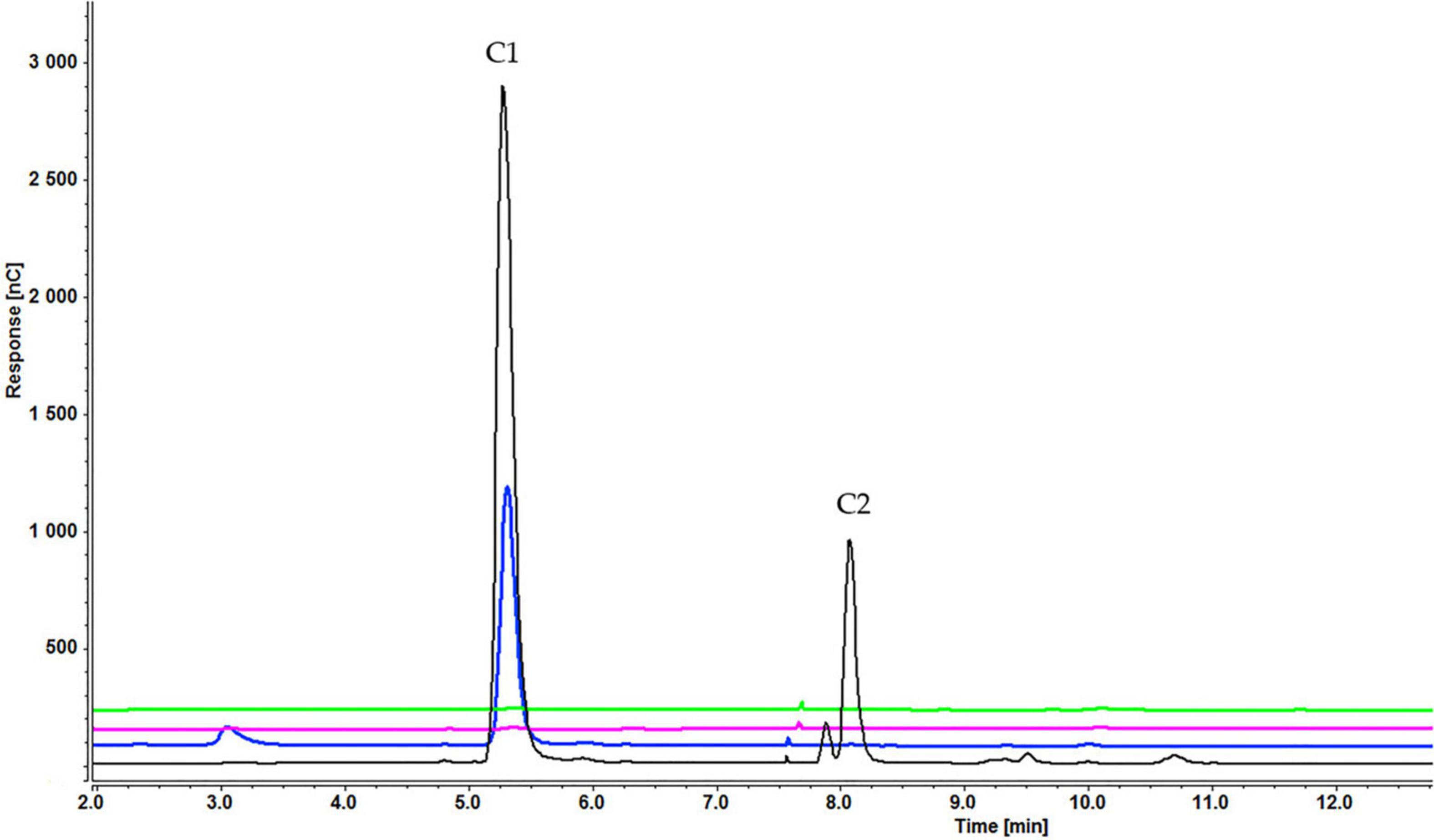
Figure 6. Cellooligosaccharides concentration in the medium after 1 h (black), 3 h (blue), 5 h (pink), and 7 h (green) of induction with 0.2% cellohexaose (C1 = glucose, C2 = cellobiose). The figure of 0.1% cellohexaose (Supplementary Figure S3) is presented in Supplementary Material.
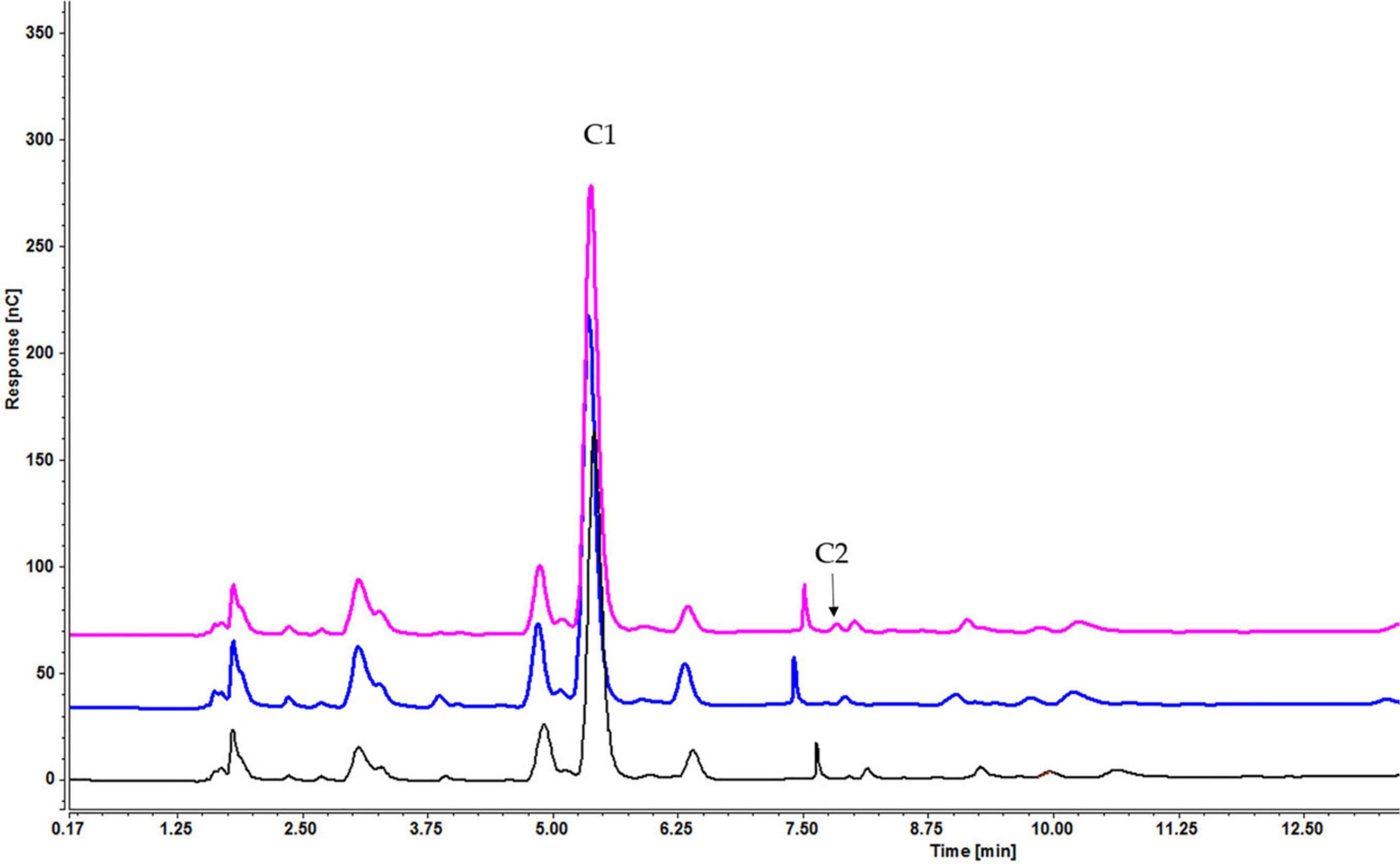
Figure 7. Intracellular concentration of cellooligosaccharides after 1 h of induction by cellobiose (pink), cellotetraose (black), and cellohexaose (blue) (C1 = glucose, C2 = cellobiose).
Enzyme Induction Using Glycerol as a Carbon Source and Cellooligosaccharides as Inducers
To investigate the effect of a non-carbohydrate carbon source on enzyme activity, we assessed enzymatic activity in glycerol-grown F. oxysporum. Growth analysis revealed that sucrose and glycerol led to a comparable amount of biomass. Unlike with sucrose-grown cells, short-chain cellooligosaccharides were better cellulase inducers when cells were grown on glycerol. For example, endoglucanase activity was the highest when cellobiose was used as inducer, followed by cellotetraose and then cellohexaose (Figure 8). Again, cellobiose concentration determined the efficiency of induction, with 0.2% cellobiose leading to the highest activity on day 2 and 0.3% cellobiose on day 3.
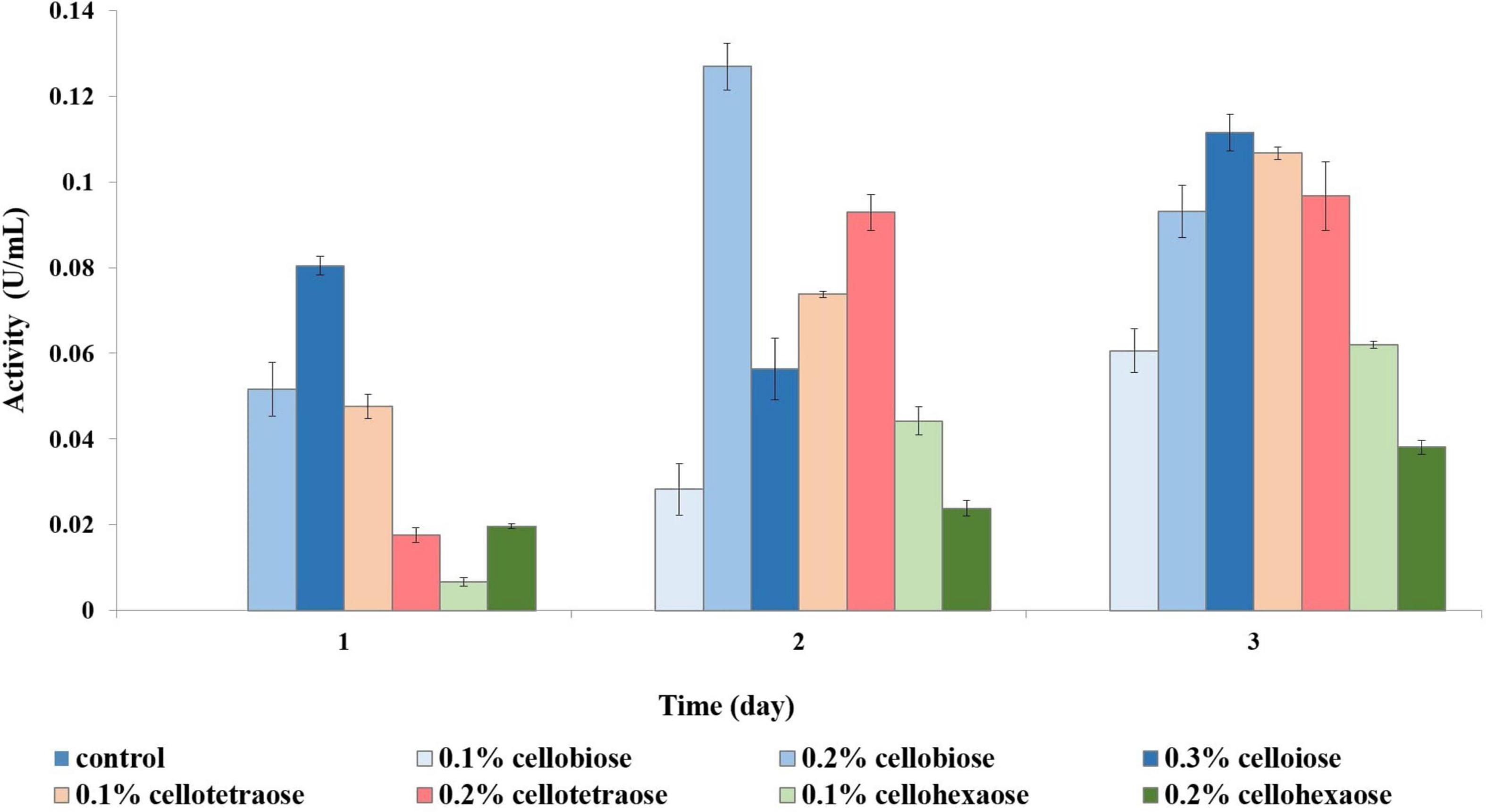
Figure 8. Endoglucanase activity of glycerol-grown biomass in the presence of different inducers and using CMC as substrate.
There appeared to be a direct inverse relationship between the degree of polymerization of an inducer and its potential for exoglucanase induction. Inducers with a lower degree of polymerization were more effective, with cellobiose resulting in the highest activity, followed by cellotetraose and cellohexaose. Similarly, a higher concentration of inducer stimulated more exoglucanase activity. For example, in the case of cellobiose, 0.1% was insufficient to elicit a response, and depending on the sampling time, either 0.2 or 0.3% produced the best results (Figure 9).
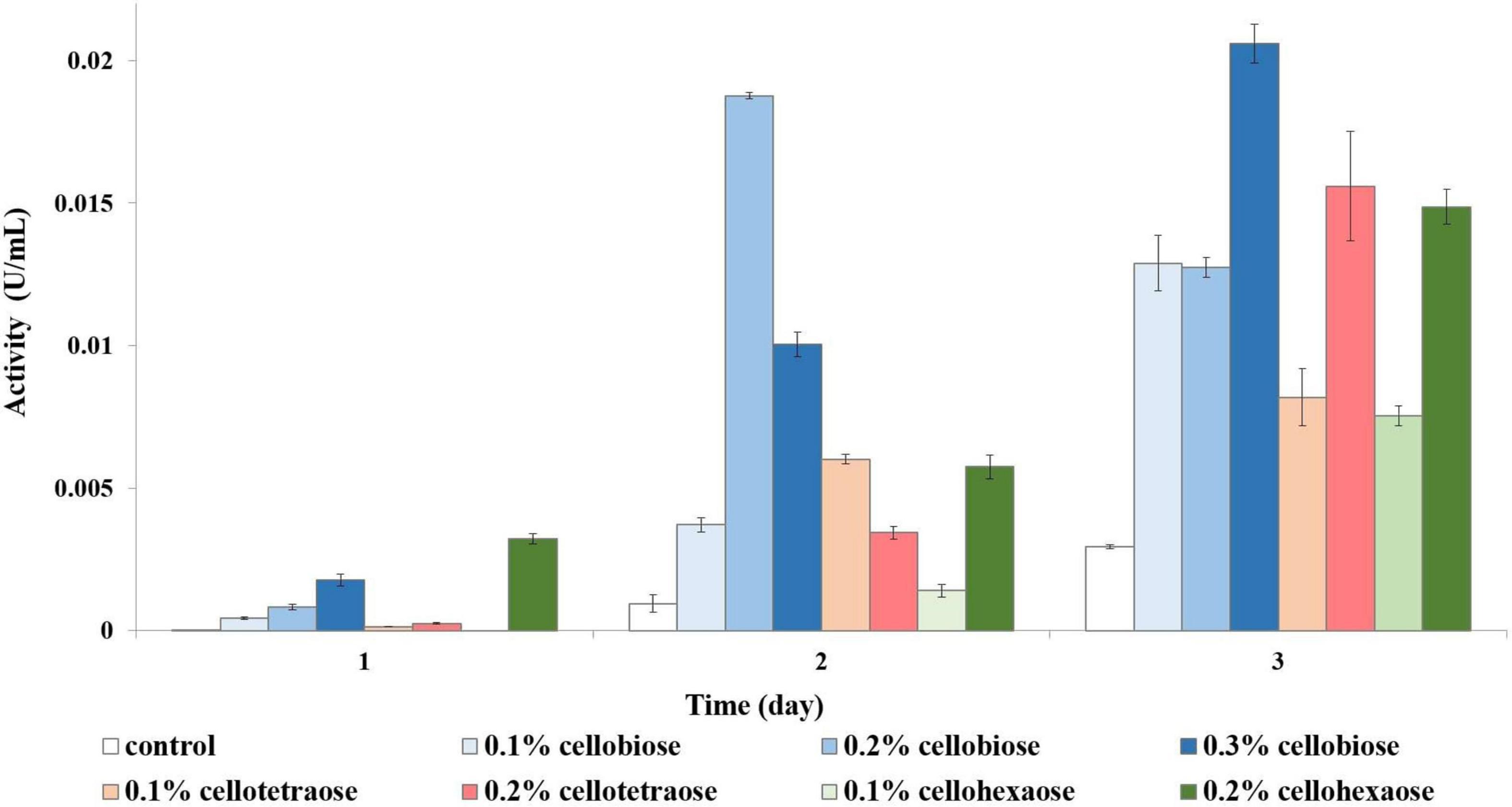
Figure 9. Exoglucanase activity of glycerol-grown biomass in the presence of different inducers and using pNPC as substrate.
In the case of cell-bound β-glucosidase, most inducers except 0.2% cellohexaose and 0.3% cellobiose did not contribute to a considerable increase in activity compared to the control (Figure 10A). Even though cell-bound β-glucosidase exhibited elevated activity in all samples, including the control, it was nevertheless lower than in sucrose-grown samples, where it was clearly influenced by the added effect of invertase.
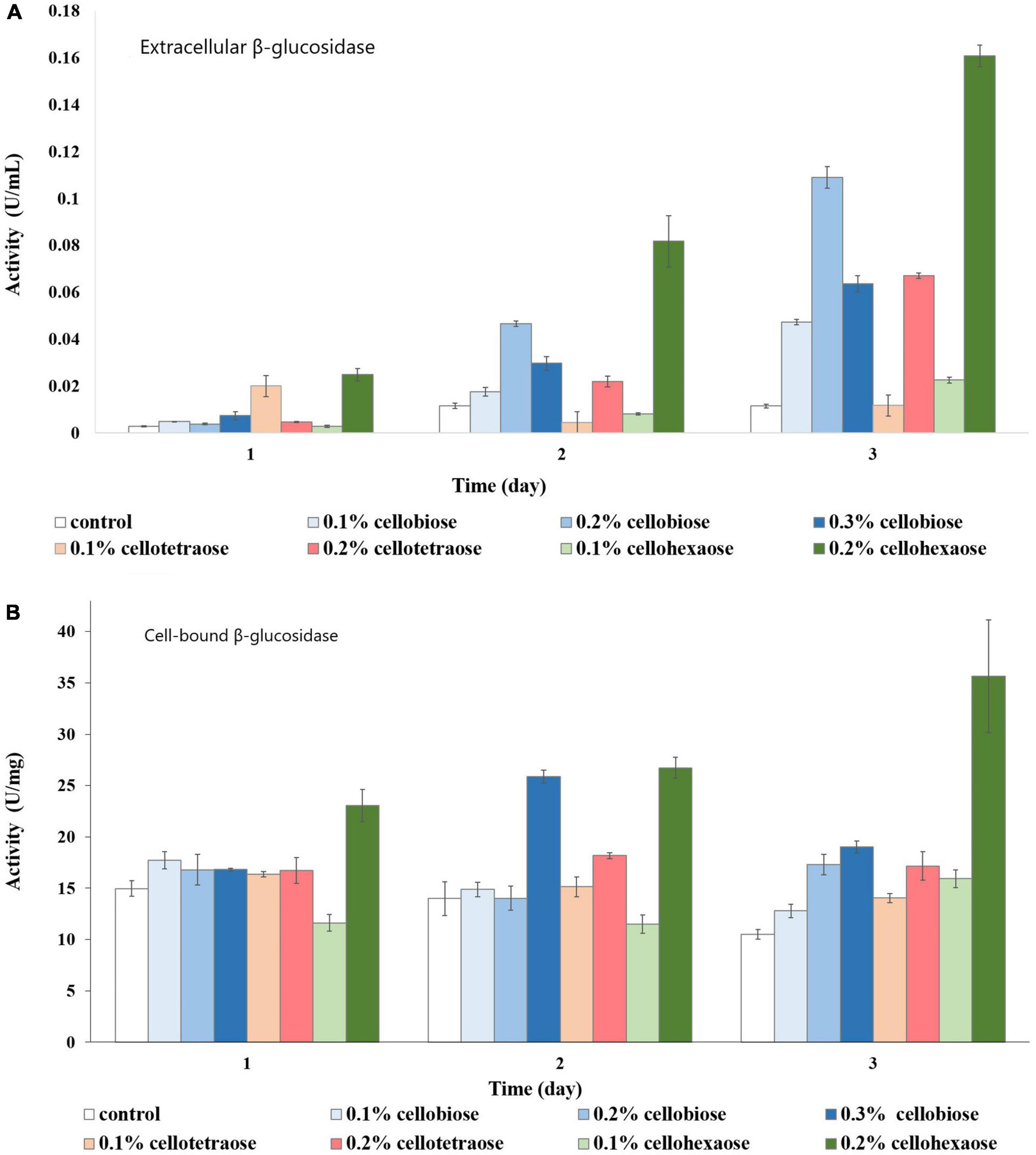
Figure 10. β-glucosidase activity of glycerol-grown biomass in the presence of different inducers and using pNPG as substrate. (A) Extracellular β-glucosidase activity; (B) Cell-bound β-glucosidase activity.
As with the cell-bound form, cellohexaose, and cellobiose proved to be better inducers of extracellular β-glucosidase activity. Here, induction efficiency depended on the concentration of the final product acting as inducer, which in turn was related to its degree of polymerization and initial concentration. Again, the best result was obtained with 0.2% cellohexaose, while the effect of 0.1% cellohexaose was negligible (Figure 10B). A concentration of 0.2% was optimal also for cellobiose. In line with previous studies (Christakopoulos et al., 1994; Olajuyigbe et al., 2016) our result confirms the elevated β-glucosidase activity of F. oxysporum, which prevents end-product inhibition. In addition, activity was slightly higher in glycerol-grown cells compared to sucrose-grown cultures.
Uptake Experiment Using Glycerol as Carbon Source
As with sucrose-grown cells, the cellooligosaccharides uptake result for cells grown on glycerol showed the gradual degradation and consumption of inducers. Particularly, cell-bound β-glucosidases induced by glycerol can be involved in disappearance of lower concentrations of inducers. In a sample containing cellobiose, both glucose and cellobiose disappeared faster (within 1 h) when 0.1% (Supplementary Figure S5A) rather than 0.2% cellobiose (Supplementary Figure S5B) (within 3 h) and 0.3% cellobiose (5 h) was added (Figure 11). Cellotetraose was rapidly degraded to cellobiose, which was then completely consumed after 1 (0.1% cellotetraose) (Supplementary Figure S6) and 3 h (0.2% cellotetraose) of induction (Figure 12). In the case of 0.2% cellotetraose, cellotetraose was detected until the first hour of induction. Finally, when cellohexaose was used as inducer, it was degraded to cellotetraose and cellobiose within the first hour of induction, as indicated by the minuscule amount of cellotetraose detected in the 0.2% cellohexaose sample (Figure 13 and Supplementary Figure S7). Cellotetraose itself was further degraded to cellobiose, which acted as cellulase inducer. Finally, as with the sucrose experiment, intracellular cellooligosaccharides analysis revealed the presence of glucose and probably a small amount of cellobiose (Figure 14).
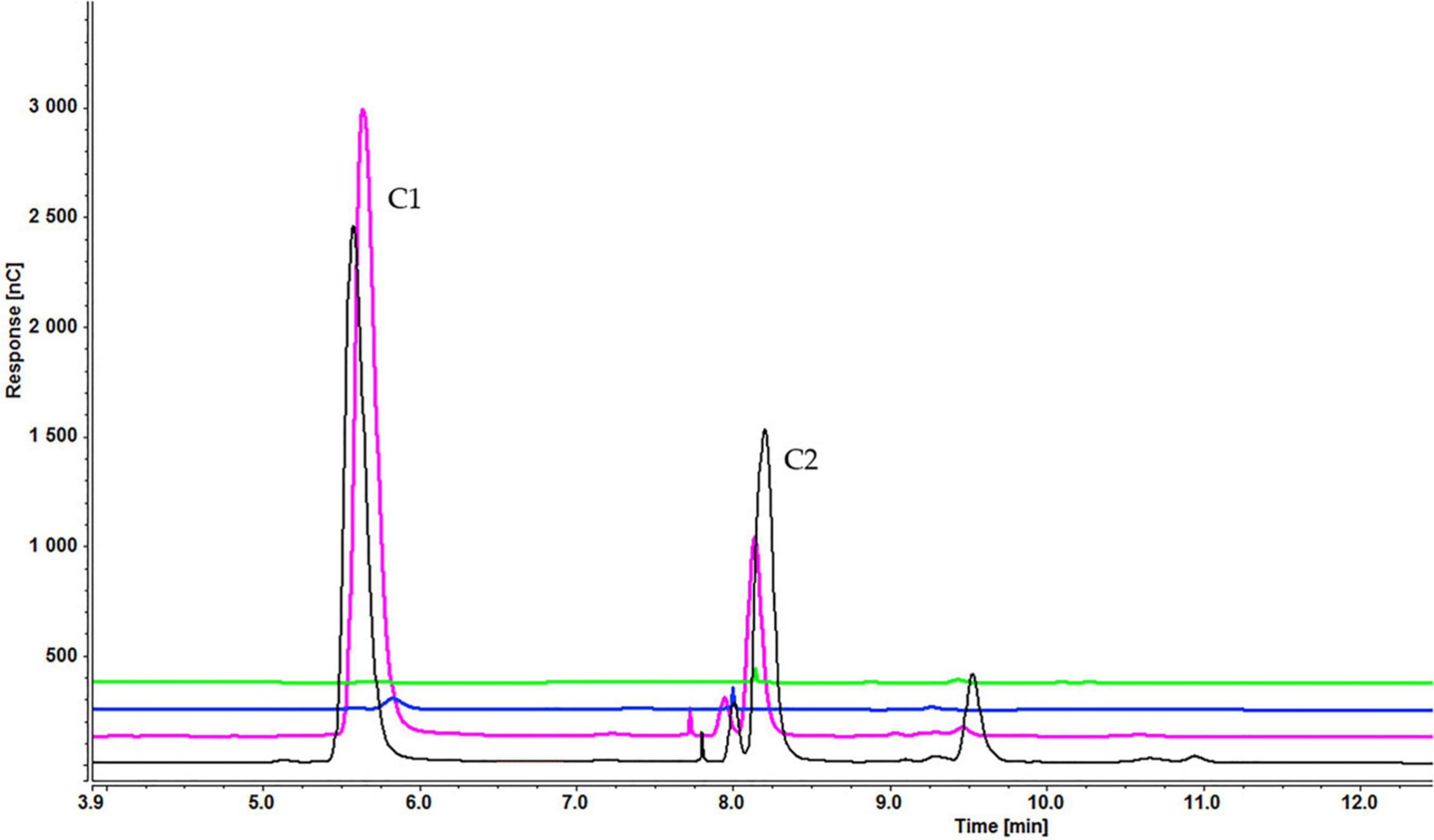
Figure 11. Cellooligosaccharides concentration in the medium after 1 h (black), 3 h (pink), 5 h (blue), and 7 h (green) of induction with 0.3% cellobiose (C1 = glucose, C2 = cellobiose). Figures for 0.1% (Supplementary Figure S5A) and 0.2% cellobiose (Supplementary Figure S5B) is presented in Supplementary Material.
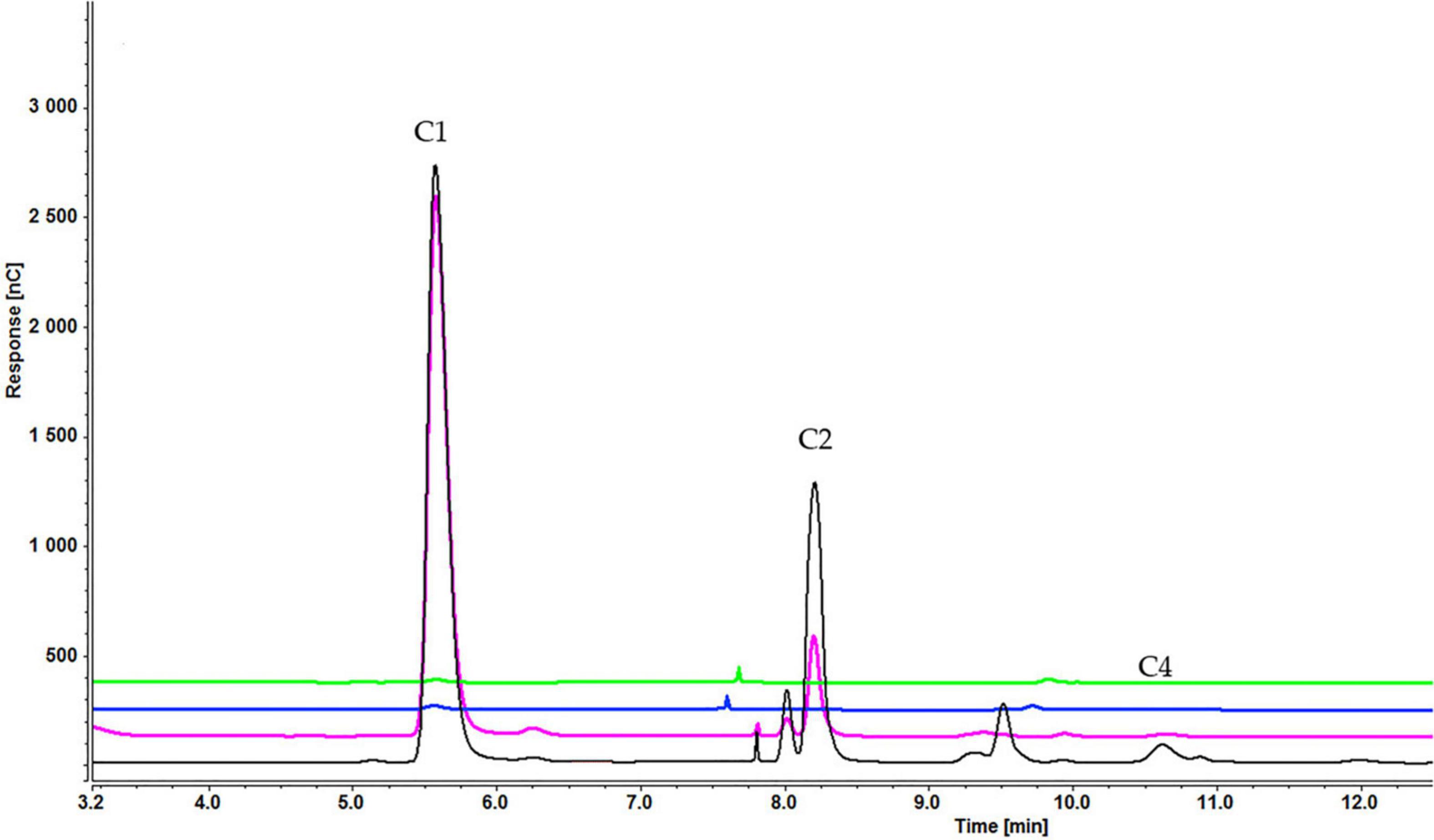
Figure 12. Cellooligosaccharides concentration in the medium after 1 h (black), 3 h (pink), 5 h (blue), and 7 h (green) of induction with) 0.2% cellotetraose (C1 = glucose, C2 = cellobiose, C4 = cellotetraose). The figure of 0.1% cellotetraose (Supplementary Figure S6) is presented in Supplementary Material.
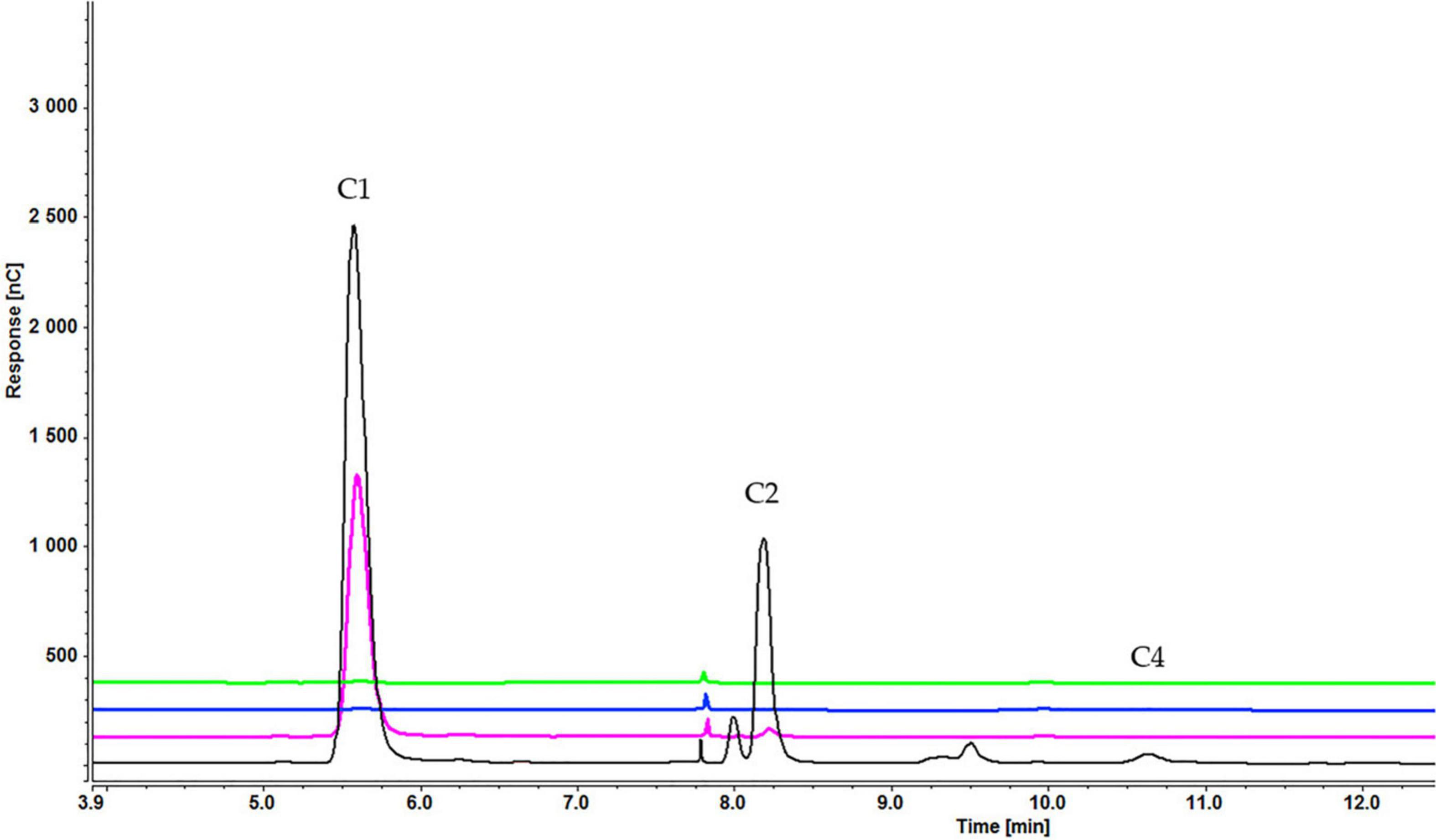
Figure 13. Cellooligosaccharides concentration in the medium after 1 h (black), 3 h (pink), 5 h (blue), and 7 h (green) of induction with 0.2% cellohexaose cellotetraose (C1 = glucose, C2 = cellobiose, C4 = cellotetraose). The figure of 0.1% cellohexaose (Supplementary Figure S7) is presented in Supplementary Material.
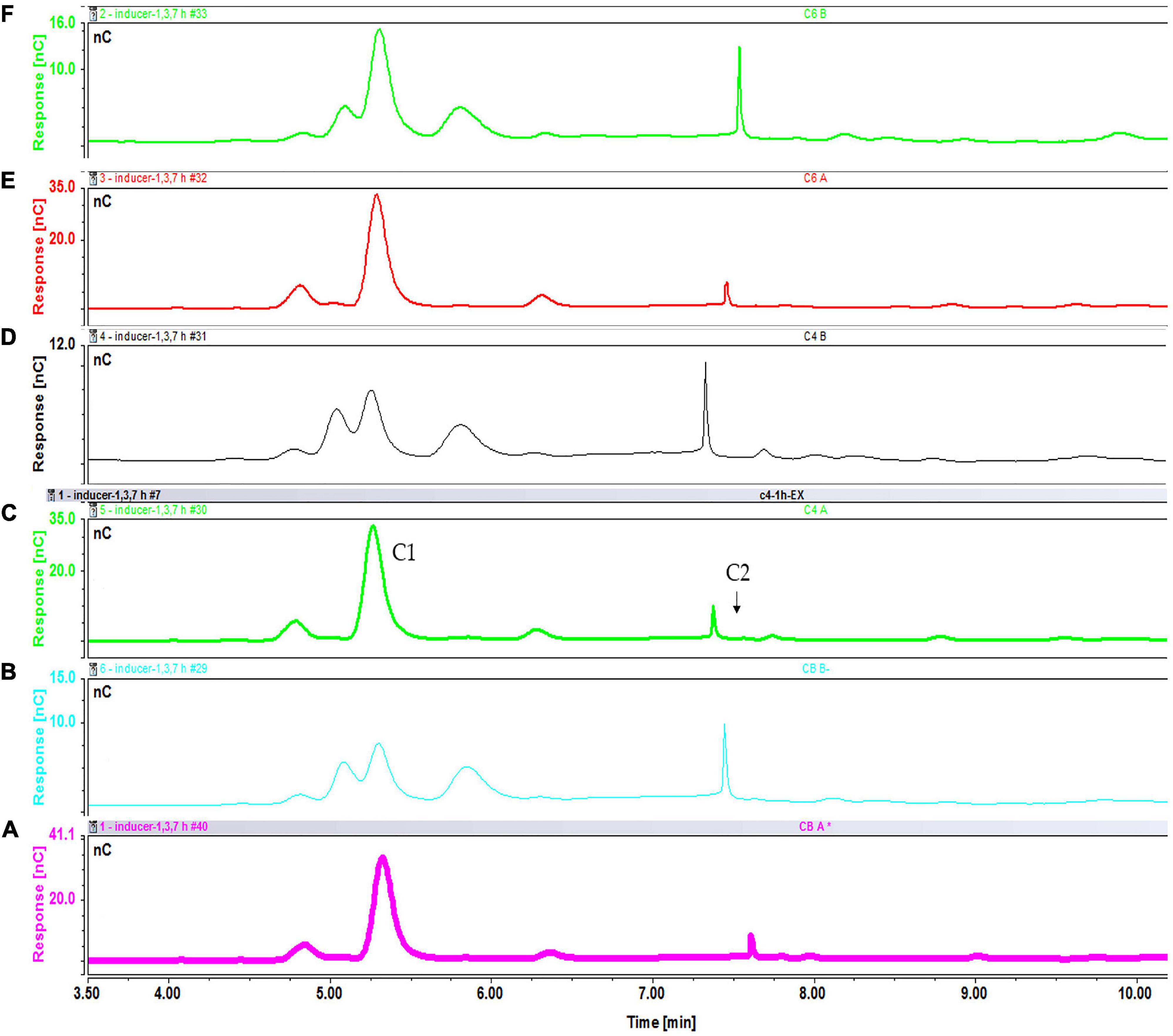
Figure 14. Intracellular concentration of cellooligosaccharides after 1 h of induction with cellobiose fractions A (A) and B (B); cellotetraose fractions A (C) and B (D); and cellohexaose fractions A (E) and B (F).
Discussion
The present study indicates that the effect of different cellooligosaccharides on induction of cellulases depends on the type of carbon source utilized by the fungal culture. Accordingly, cellotetraose is more effective when F. oxysporum is grown on sucrose; whereas a shorter oligosaccharide such as cellobiose works better in glycerol-grown cells. Moreover, the concentration of the inducer is as important as its chain length. The carbon source used to grow the fungus may change its complement of basal enzymes irrespective of whether cellulosic material is present or not. Sucrose is the main sugar reserve in plants (Hughes and Mitchell, 1995), as well as the main substrate providing UDP-glucose for cellulose biosynthesis (Rende et al., 2017). It is also the preferred sugar consumed by plant pathogenic fungi (Hughes and Mitchell, 1995). Therefore, pathogens have developed mechanisms to optimize sucrose consumption by producing O-glycosyl-degrading enzymes such as invertases. The endo- and exoglucanase activity of samples grown on glycerol is higher than sucrose samples. This can be due to the repression of cellooligosaccharides degrading genes by sucrose, as the preferred carbon source. Investigations on filamentous fungi showed that the carbon catabolic repression by easily metabolized carbon sources such as glucose and sucrose, repress genes coding cellulases and hemicellulases even in presence of the lignocellulosic substrate (Znameroski et al., 2012). Also, there is a possibility that Stl1 (Sugar Transporter Like) transporter be influential in this increase. It is known that glycerol/H+ symporter encoded by stl1 are responsible for active transport of glycerol (Ferreira et al., 2005). Previous studies including structural analysis of Stl1 transporter shows it has characteristics of sugar transporters (twelve transmembrane domains) and participates in sugar transportation, especially hexoses (Zhao et al., 1994). However, based on Ko et al. (1993), sugar transporters are not necessarily contribute in the transportation of the sugars to be metabolized and they may act as a sensor to modulate the activity or expression of the other transporters. So we assume that, in samples grown on glycerol, the higher number of Stl1 transporters can affect the cellooligosaccharides uptake and cellulase induction. Moreover, the applied inducer also influence the profile of membrane transporters and consequently enzyme induction. Carbon sources enter the cell mainly by MFS (major facilitator superfamily) transporters (Dos Reis et al., 2016). There are 17 families of MFS transporters which sugar transportation belong to families 1, 5, and 7 (Saier et al., 1999; Yan, 2013). Despite that not many of these transporters are discovered in fungi, their importance in cellulose and hemicellulose sensing and degradation in T.reesei, N.crassa, and A. nidulans are known (Zhang et al., 2013; Cai et al., 2014; Znameroski et al., 2014; Dos Reis et al., 2016). In N.crassa, CDT-1 and CDT-2 are known as “transceptors” because they are transporting receptors that participate in cellulose sensing, signaling, and cellodextrins transportation (Znameroski et al., 2014). In A. nidulans, CltA transport cellobiose and CltB is mainly responsible for substrate sensing and signaling (Dos Reis et al., 2016). In T. reesei, Stp1 which transports cellobiose and glucose is involved in carbon catabolite repression while Crt1 do not participate in transportation, instead it works for cellulolytic signaling (Zhang et al., 2013).
In this study, exoglucanase assay by using Avicel as enzyme substrate showed no activity therefore, pNPC was replaced for determining the exoglucanase activity. Avicel has a low degree of polymerization and low accessibility, which makes it a good substrate for exoglucanase (Zhang et al., 2006). However, in the absence of activity with Avicel, more sensitive substrates, such as pNPC or p-nitrophenyl-p-D-lactoside, are used. In the presence of these substrates, the exoglucanase breaks the agluconic bond between p-nitrophenyl and the disaccharide moiety to release units of cellobiose (Eriksson et al., 1990). Based on the result sophorose is not an effective inducer of F. oxysporum which is compatible with previous observations reporting that sophorose induced an incomplete set of cellulase enzymes (Bazafkan et al., 2014). Probably, sophorose is not the final inducer of cellulases and its signaling process does not reflect cellulose abundance (Shida et al., 2016). This study helps clarify the mechanism of cellulase production by identifying the biomass products used as preferential inducers by the fungus. Importantly, the study shows the potential of cellobiose as an efficient and cheap inducer compared to cellotetraose and cellohexaose. Understanding the molecular mechanism of cellulase induction in filamentous fungi could allow researchers to manipulate other (e.g., thermophilic) cellulolytic organisms to enable the production, purification, and examination of new lignocellulolytic enzymes (Znameroski et al., 2012). Moreover, this study illustrates that for each specific inducer, the appropriate carbon source should be selected to achieve the best outcome. The uptake experiment yielded a similar result for both carbon sources, indicating that any difference between inducers cannot be ascribed to different transporters and, in both cases, cellobiose is the final inducer. Therefore, the higher endoglucanase and exoglucanase activity of glycerol-grown samples and the different order of inducers is linked to the genes and pathways activated by sucrose or glycerol, which influence inducer sensing and transcription of cellulase-encoding genes.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding author/s.
Author Contributions
UR and PC: conceptualization. NN, LM, UR, and PC: methodology. NN: investigation and writing—original draft preparation. LM, UR, and PC: writing—review and editing. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Swedish Research Council (VR), grant no. 2014-05906.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2021.652655/full#supplementary-material
References
Agrios, G. N. (2005). “Chapter fourteen – plant diseases caused by viruses,” in Plant Pathology, ed. G. N. Agrios (San Diego, CA: Academic Press), 723–824. doi: 10.1016/B978-0-08-047378-9.50020-8
Alazi, E., and Ram, A. F. J. (2018). Modulating transcriptional regulation of plant biomass degrading enzyme networks for rational design of industrial fungal strains. Front. Bioeng. Biotechnol. 6:133. doi: 10.3389/fbioe.2018.00133
Ali, S. S., Khan, M., Mullins, E., and Doohan, F. M. (2014). Identification of fusarium oxysporum genes associated with lignocellulose bioconversion competency. Bioenergy Res. 7, 110–119. doi: 10.1007/s12155-013-9353-0
Ali, S. S., Nugent, B., Mullins, E., and Doohan, F. M. (2016). Fungal-mediated consolidated bioprocessing: the potential of Fusarium oxysporum for the lignocellulosic ethanol industry. AMB Express 6:13. doi: 10.1186/s13568-016-0185-0
Bazafkan, H., Tisch, D., and Schmoll, M. (2014). “Chapter 20 – regulation of glycoside hydrolase expression in trichoderma,” in Biotechnology and Biology of Trichoderma, eds V. K. Gupta, M. Schmoll, A. Herrera-Estrella, R. S. Upadhyay, I. Druzhinina, and M. G. Tuohy (Amsterdam: Elsevier), 291–308. doi: 10.1016/B978-0-444-59576-8.00020-5
Bhat, M. K. (2000). Cellulases and related enzymes in biotechnology. Biotechnol. Adv. 18, 355–383. doi: 10.1016/s0734-9750(00)00041-0
Cai, P., Gu, R., Wang, B., Li, J., Wan, L., Tian, C., et al. (2014). Evidence of a critical role for cellodextrin transporte 2 (CDT-2) in both cellulose and hemicellulose degradation and utilization in Neurospora crassa. PLoS One 9:e89330. doi: 10.1371/journal.pone.0089330
Castro, S., Pedersoli, W. R., Cristina, A., Antoniêto, C., Steindorff, A. S., Silva-rocha, R., et al. (2014). Comparative metabolism of cellulose, sophorose and glucose in Trichoderma reesei using high-throughput genomic and proteomic analyses. Biotechnol. Biofuels. 7:41. doi: 10.1186/1754-6834-7-41
Cen, P., and Xia, L. (1999). “Production of cellulase by solid-state fermentation,” in Recent Progress in Bioconversion of Lignocellulosics, eds G. T. Tsao, A. P. Brainard, H. R. Bungay, N. J. Cao, P. Cen, Z. Chen, et al. (Berlin: Springer Berlin Heidelberg), 69–92. doi: 10.1007/3-540-49194-5_4
Chandel, A. K., Chandrasekhar, G., Silva, M. B., Silvério, S., Chandel, A. K., Chandrasekhar, G., et al. (2012). The realm of cellulases in biorefinery development. Crit. Rev. Biotechnol. 32, 187–202. doi: 10.3109/07388551.2011.595385
Chikamatsu, G., Shirai, K., Kato, M., and Kobayashi, T. (1999). Structure and expression properties of the endo-B-1,4-glucanase a gene from the filamentous fungus Aspergillus nidulans. FEMS Microbiol. Lett. 175, 239–245. doi: 10.1111/j.1574-6968.1999.tb13626.x
Christakopoulos, P., Goodenough, P. W., Kekos, D., Macris, B. J., Claeyssens, M., and Bhat, M. K. (1994). Purification and characterisation of an extracellular β−glucosidase with transglycosylation and exo-glucosidase activities from Fusarium oxysporum. Eur. J. Biochem. 224, 379–385. doi: 10.1111/j.1432-1033.1994.00379.x
Christakopoulos, P., Kekos, D., Kolisis, F. N., and Macris, B. J. (1995a). Controlling simultaneous production of endoglucanase and beta-glucosidase by Fusarium oxysporum in submerged culture. Biotechnol. Lett. 17, 883–888. doi: 10.1007/BF00129023
Christakopoulos, P., Kekos, D., Macris, B. J., Claeyssens, M., and Bhat, M. K. (1995b). Purification and characterization of a less randomly acting endo-1,4-β-D-glucanase from the culture filtrates of Fusarium oxysporum. Arch. Biochem. Biophys. 316, 428–433. doi: 10.1006/abbi.1995.1057
Christakopoulos, P., Kekos, D., Macris, B. J., Claeyssens, M., and Bhat, M. K. (1995c). Purification and mode of action of a low molecular mass endo-1,4-β-d-glucanase from Fusarium oxysporum. J. Biotechnol. 39, 85–93. doi: 10.1016/0168-1656(94)00147-5
Christakopoulos, P., Koullas, D. P., Kekos, D., Koukios, E. G., and Macris, B. J. (1991). Direct conversion of straw to ethanol by Fusarium oxysporum: effect of cellulose crystallinity. Enzyme Microb. Technol. 13, 272–274. doi: 10.1016/0141-0229(91)90141-V
Donohoe, B. S., and Resch, M. G. (2015). Mechanisms employed by cellulase systems to gain access through the complex architecture of lignocellulosic substrates. Curr. Opin. Chem. Biol. 29, 100–107. doi: 10.1016/j.cbpa.2015.08.014
Dos Reis, T. F., De Lima, P. B. A., Parachin, N. S., Mingossi, F. B., De Castro Oliveira, J. V., Ries, L. N. A., et al. (2016). Identification and characterization of putative xylose and cellobiose transporters in Aspergillus nidulans. Biotechnol. Biofuels 9:204. doi: 10.1186/s13068-016-0611-1
Els, M. M., and Reese, E. T. (1956). Induction of cellulase in trichoderma viride as influenced by carbon sources and metals by the dinitrosalicylic glucose. J. Bacteriol. 269–278. doi: 10.1128/JB.73.2.269-278.1957
Eriksson, K.-E. L., Blanchette, R., and Ander, P. (1990). Microbial and Enzymatic Degradation of Wood and Wood Components. Berlin: Springer-Verlag. doi: 10.1007/978-3-642-46687-8
Fang, X., Yano, S., Inoue, H., and Sawayama, S. (2008). Lactose enhances cellulase production by the filamentous fungus Acremonium cellulolyticus. 106, 115–120. doi: 10.1263/jbb.106.115
Ferreira, C., van Voorst, F., Martins, A., Neves, L., Oliveira, R., Kielland-Brandt, M. C., et al. (2005). A member of the sugar transporter family, Stl1p is the glycerol/H+ symporter in Saccharomyces cerevisiae. Mol. Biol. Cell 16, 2068–2076.
Gielkens, M. M. C., Dekkers, E., Visser, J., and Leo, H. (1999). Two cellobiohydrolase-encoding genes from Aspergillus niger require D -xylose and the xylanolytic transcriptional activator XlnR for their expression. Appl. Environ. Microb. 65, 4340–4345.
Huberman, L. B., Liu, J., Qin, L., and Glass, N. L. (2016). Regulation of the lignocellulolytic response in filamentous fungi. Fungal Biol. Rev. 30, 101–111. doi: 10.1016/j.fbr.2016.06.001
Hughes, E., and Mitchell, D. T. (1995). Utilization of sucrose by Hymenoscyphus ericae (an ericoid endomycorrhizal fungus) and ectomycorrhizal fungi. Mycol. Res. 99, 1233–1238. doi: 10.1016/S0953-7562(09)80284-0
Humbird, D., Davis, R., Tao, L., Kinchin, C., Hsu, D., Aden, A., et al. (2011). Process Design and Economics for Biochemical Conversion of Lignocellulosic Biomass to Ethanol: Dilute-Acid Pretreatment and Enzymatic Hydrolysis of Corn Stover. Golden, CO: National Renewable Energy Laboratory, 1–147. doi: 10.2172/1107470
Kekos, D., Macris, B. J., Claeyssens, M., and Bhat, M. K. (1995). Purification and mode of action of a low molecular mass endo-1,4-B-D-glucanase from Fusarium oxysporum. J. Biotechnol. 39, 85–93.
Ko, C. H., Liang, H., and Gaber, R. F. (1993). Roles of multiple glucose transporters in Saccharomyces cerevisiae. Mol. Cell. Biol. 13, 638–648. doi: 10.1128/mcb.13.1.638
Leite, A., and Eveleight, D. E. (1989). Mechanism by which cellulose triggers cellobiohydrolase I gene expression in Trichoderma reesei. Proc. Natl. Acad. Sci. U. S. A. 86, 6138–6141. doi: 10.1073/pnas.86.16.6138
Mandels, M., Parrish, F. W., Reese, E. T., Parrish, R. W. V., and Reese, E. T. (1961). Sophorose as an inducer of cellulase in trichoderma viride. J. Bacteriol. 400–408. doi: 10.1128/JB.83.2.400-408.1962
Mandels, M., and Reese, E. T. (1959). Induction of cellulase in fungi by cellobiose. J. Bacteriol. 816–826. doi: 10.1128/JB.79.6.816-826.1960
Mielenz, J. R. (2001). Ethanol production from biomass: technology and commercialization status. Curr. Opin. Microbiol. 4, 324–329. doi: 10.1016/S1369-5274(00)00211-3
Morikawa, Y., Ohashi, T., Mantani, O., and Okada, H. (1995). Cellulase induction by lactose in Trichoderma reesei PC-3-7. Appl. Microbiol. Biotechnol. 44, 106–111. doi: 10.1007/BF00164488
Moshkin, N. P., Fomina, A. V., and Chernykh, G. G. (2005). Cellulase, clostridia, and ethanol†. Microbiol. Mol. Biol. Rev. 69, 124–154. doi: 10.1134/S0869864315020043
Obeng, E. M., Nurul, S., Adam, N., Budiman, C., Ongkudon, C. M., Maas, R., et al. (2017). Lignocellulases: a review of emerging and developing enzymes, systems, and practices. Bioresour. Bioprocess. 4, 1–22. doi: 10.1186/s40643-017-0146-8
Ohnishi, Y., and Nagase, M. (2007). Transcriptional regulation of two cellobiohydrolase encoding genes (cel1 and cel2) from the wood-degrading basidiomycete Polyporus arcularius. Appl. Microbiol. Biotechnol. 76, 1069–1078. doi: 10.1007/s00253-007-1090-x
Olajuyigbe, F. M., Nlekerem, C. M., and Ogunyewo, O. A. (2016). Production and characterization of highly thermostable β -glucosidase during the biodegradation of methyl cellulose by Fusarium oxysporum. Biochem. Res. Int. 2016:3978124. doi: 10.1155/2016/3978124
Panagiotou, G., Christakopoulos, P., and Olsson, L. (2005). Simultaneous saccharification and fermentation of cellulose by Fusarium oxysporum F3 – growth characteristics and metabolite profiling. Enzyme Microb. Technol. 36, 693–699. doi: 10.1016/j.enzmictec.2004.12.029
Panagiotou, G., Kekos, D., Macris, B. J., and Christakopoulos, P. (2003). Production of cellulolytic and xylanolytic enzymes by Fusarium oxysporum grown on corn stover in solid state fermentation. Ind. Crops Prod. 18, 37–45. doi: 10.1016/S0926-6690(03)00018-9
Park, S., Ransom, C., Mei, C., Sabzikar, R., Qi, C., Chundawat, S., et al. (2011). The quest for alternatives to microbial cellulase mix production: corn stover-produced heterologous multi-cellulases readily deconstruct lignocellulosic biomass into fermentable sugars. 86, 633–641. doi: 10.1002/jctb.2584
Pathology, M. P. (2003). Fusarium oxysporum: exploring the molecular arsenal of a vascular wilt fungus. 4, 315–325. doi: 10.1046/J.1364-3703.2003.00180.X
Rani, R., Sukumaran, R. K., Kumar, A., Larroche, C., and Pandey, A. (2010). Enzyme and microbial technology advancement and comparative profiles in the production technologies using solid-state and submerged fermentation for microbial cellulases. Enzyme Microb. Technol. 46, 541–549. doi: 10.1016/j.enzmictec.2010.03.010
Rende, U., Wang, W., Gandla, M. L., Jönsson, L. J., and Niittylä, T. (2017). Cytosolic invertase contributes to the supply of substrate for cellulose biosynthesis in developing wood. New Phytol. 214, 796–807. doi: 10.1111/nph.14392
Saier, M. H., Beatty, J. T., Goffeau, A., Harley, K. T., Heijne, W. H. M., Huang, S. C., et al. (1999). The major facilitator superfamily. J. Mol. Microbiol. Biotechnol. 1, 257–279.
Schu, M. (2000). Protein engineering of cellulases. Biochim. Biophys. Acta 1543. doi: 10.1016/S0167-4838(00)00247-8
Selbmann, L., Egidi, E., Isola, D., Onofri, S., Hoog, G. S., De Chinaglia, S., et al. (2013). Biodiversity, evolution and adaptation of fungi in extreme environments. Plant Biol. 147, 237–246. doi: 10.1080/11263504.2012.753134
Shida, Y., Furukawa, T., and Ogasawara, W. (2016). Deciphering the molecular mechanisms behind cellulase production in trichoderma reesei, the hyper-cellulolytic filamentous fungus. Biosci. Biotechnol. Biochem. 80, 1712–1729. doi: 10.1080/09168451.2016.1171701
Singhania, R. R., Sukumaran, R. K., Patel, A. K., Larroche, C., and Pandey, A. (2010). Advancement and comparative profiles in the production technologies using solid-state and submerged fermentation for microbial cellulases. Enzyme Microb. Technol. 46, 541–549. doi: 10.1016/j.enzmictec.2010.03.010
Suzuki, H., Igarashi, K., and Samejima, M. (2010). Cellotriose and cellotetraose as inducers of the genes encoding cellobiohydrolases in the basidiomycete Phanerochaete chrysosporium. Appl. Environ. Microbiol. 76, 6164–6170. doi: 10.1128/AEM.00724-10
Tiwari, P., Misra, B. N., and Sangwan, N. S. (2013). β -Glucosidases from the fungus trichoderma: an efficient cellulase machinery in biotechnological applications. Biomed. Res. Int. 2013:203735. doi: 10.1155/2013/203735
Ulmer, D. C., Leisola, M. S. A., and Fiechter, A. (1984). Possible induction of the ligninolytic system of Phanerochaete chrysosporium. J. Biotechnol. 1, 13–24. doi: 10.1016/S0168-1656(84)90055-5
Umile, C., and Kubicek, C. P. (1986). A constitutive, plasma-membrane bound β-glucosidase in Trichoderma reesei. FEMS Microbiol. Lett. 34, 291–295. doi: 10.1111/j.1574-6968.1986.tb01423.x
Vaheri, M. P., Vaheri, M. E. O., and Kauppinen, V. S. (1979). Formation and release of cellulolytic enzymes during growth of Trichoderma reesei on cellobiose and glycerol. Appl. Microbiol. Biotechnol. 8, 73–80. doi: 10.1007/BF00510268
Wolfenden, R., and Snider, M. J. (2001). The depth of chemical time and the power of enzymes as catalysts. Acc. Chem. Res. 34, 938–945. doi: 10.1021/ar000058i
Xu, Q., Singh, A., and Himmel, M. E. (2009). Perspectives and new directions for the production of bioethanol using consolidated bioprocessing of lignocellulose. Curr. Opin. Biotechnol. 20, 364–371. doi: 10.1016/j.copbio.2009.05.006
Yan, N. (2013). Structural advances for the major facilitator superfamily (MFS) transporters. Trends Biochem. Sci. 38, 151–159. doi: 10.1016/j.tibs.2013.01.003
Yoon, L. W., Ang, T. N., Ngoh, G. C., Seak, A., and Chua, M. (2014). Fungal solid-state fermentation and various methods of enhancement in cellulase production. Biomass and Bioenergy 67, 319–338. doi: 10.1016/j.biombioe.2014.05.013
Zhang, W., Kou, Y., Xu, J., Cao, Y., Zhao, G., Shao, J., et al. (2013). Two major facilitator superfamily sugar transporters from Trichoderma reesei and their roles in induction of cellulase biosynthesis. J. Biol. Chem. 288, 32861–32872. doi: 10.1074/jbc.M113.505826
Zhang, X., and Zhang, Y. P. (2013). “Cellulases: characteristics, sources, production, and applications,” in Bioprocessing Technologies in Biorefi nery for Sustainable Production of Fuels, Chemicals, and Polymers, First Edition, eds S. T. Yang, H. El-Enshasy, and N. Thongchul (New York, NY: John Wiley & Sons Inc.), 131–146. doi: 10.1002/9781118642047
Zhang, Y. H. P., and Lynd, L. R. (2004). Toward an aggregated understanding of enzymatic hydrolysis of cellulose: noncomplexed cellulase systems. Biotechnol. Bioeng. 88, 797–824. doi: 10.1002/bit.20282
Zhang, Y. P., Himmel, M. E., and Mielenz, J. R. (2006). Outlook for cellulase improvement: screening and selection strategies. Biotechnol. Adv. 24, 452–481. doi: 10.1016/j.biotechadv.2006.03.003
Zhao, S., Douglas, N. W., Heine, M. J. S., Williams, G. M., Winther-Larsen, H. C., and Meaden, P. G. (1994). The STL1 gene of Saccharomyces cerevisiae is predicted to encode a sugar transporter-like protein. Gene 146, 215–219. doi: 10.1016/0378-1119(94)90295-X
Zhu, Z., and Zhang, Y. P. (2009). Direct quantitative determination of adsorbed cellulase on lignocellulosic biomass with its application to study cellulase desorption for potential recycling. Analyst 134, 2267–2272. doi: 10.1039/b906065k
Znameroski, E. A., Coradetti, S. T., Roche, C. M., Tsai, J. C., Iavarone, A. T., Cate, J. H. D., et al. (2012). Induction of lignocellulose-degrading enzymes in Neurospora crassa by cellodextrins. Proc. Natl. Acad. Sci. U. S. A. 109, 6012–6017. doi: 10.1073/pnas.1118440109
Keywords: Fusarium, induction, cellulase, cellooligosaccharides, xylooligosacharides, carbon source, degree of polymerization
Citation: Najjarzadeh N, Matsakas L, Rova U and Christakopoulos P (2021) How Carbon Source and Degree of Oligosaccharide Polymerization Affect Production of Cellulase-Degrading Enzymes by Fusarium oxysporum f. sp. lycopersici. Front. Microbiol. 12:652655. doi: 10.3389/fmicb.2021.652655
Received: 12 January 2021; Accepted: 22 February 2021;
Published: 26 March 2021.
Edited by:
Saurabh Dhiman, South Dakota School of Mines and Technology, United StatesReviewed by:
Yosuke Shida, Nagaoka University of Technology, JapanRolf A. Prade, Oklahoma State University, United States
Copyright © 2021 Najjarzadeh, Matsakas, Rova and Christakopoulos. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Paul Christakopoulos, cGF1bC5jaHJpc3Rha29wb3Vsb3NAbHR1LnNl
 Nasim Najjarzadeh
Nasim Najjarzadeh Leonidas Matsakas
Leonidas Matsakas Ulrika Rova
Ulrika Rova Paul Christakopoulos
Paul Christakopoulos