- 1Department of Molecular Genetics, Groningen Biomolecular Sciences and Biotechnology Institute, University of Groningen, Groningen, Netherlands
- 2Membrane Biochemistry and Biophysics, Bijvoet Centre for Biomolecular Research, Department of Chemistry, Faculty of Science, Utrecht University, Utrecht, Netherlands
- 3NMR Spectroscopy, Bijvoet Centre for Biomolecular Research, Department of Chemistry, Faculty of Science, Utrecht University, Utrecht, Netherlands
Bacterial non-ribosomally produced peptides (NRPs) form a rich source of antibiotics, including more than 20 of these antibiotics that are used in the clinic, such as penicillin G, colistin, vancomycin, and chloramphenicol. Here we report the identification, purification, and characterization of a novel NRP, i.e., brevibacillin 2V (lipo-tridecapeptide), from Brevibacillus laterosporus DSM 25. Brevibacillin 2V has a strong antimicrobial activity against Gram-positive bacterial pathogens (minimum inhibitory concentration = 2 mg/L), including difficult-to-treat antibiotic-resistant Enterococcus faecium, Enterococcus faecalis, and Staphylococcus aureus. Notably, brevibacillin 2V has a much lower hemolytic activity (HC50 > 128 mg/L) and cytotoxicity (CC50 = 45.49 ± 0.24 mg/L) to eukaryotic cells than previously reported NRPs of the lipo-tridecapeptide family, including other brevibacillins, which makes it a promising candidate for antibiotic development. In addition, our results demonstrate that brevibacillins display a synergistic action with established antibiotics against Gram-negative bacterial pathogens. Probably due to the presence of non-canonical amino acids and D-amino acids, brevibacillin 2V showed good stability in human plasma. Thus, we identified and characterized a novel and promising antimicrobial candidate (brevibacillin 2V) with low hemolytic activity and cytotoxicity, which can be used either on its own or as a template for further total synthesis and modification.
Introduction
The non-ribosomally produced peptides (NRPs) have formed a rich source of antimicrobials, including more than 20 marketed antibiotics, such as chloramphenicol, penicillin G, colistin, and vancomycin (Süssmuth and Mainz, 2017). Among these, lipopeptides form a rich class of NRPs that have shown a strong antimicrobial activity against Gram-positive (Hamamoto et al., 2015; Ling et al., 2015) and/or Gram-negative (Cochrane et al., 2014; Cociancich et al., 2015) pathogens.
Large numbers of antimicrobial NRPs have been isolated from different Brevibacillus spp. (Yang and Yousef, 2018), including many linear lipopeptides, such as bogorol A–E (Barsby et al., 2001, 2006), brevibacillin (Yang et al., 2016), and brevibacillin V (Wu et al., 2019). Bogorol A–E, brevibacillin, and brevibacillin V are produced by Brevibacillus laterosporus PNG-276, B. laterosporus OSY-I1, and B. laterosporus fmb70 (Barsby et al., 2001, 2006; Yang et al., 2016; Wu et al., 2019), respectively, and all of them can be grouped into the family of non-ribosomally produced linear lipo-tridecapeptides (Yang and Yousef, 2018). These lipo-tridecapeptides show a strong antimicrobial activity against Gram-positive pathogenic bacteria, including methicillin-resistant Staphylococcus aureus and vancomycin-resistant Enterococcus spp. (Barsby et al., 2006; Yang et al., 2016; Wu et al., 2019). However, many lipo-tridecapeptides have the disadvantage of being highly hemolytic and cytotoxic (Li et al., 2018), which have limited the potential of many antimicrobial peptides to be developed as therapeutics (Mor and Nicolas, 1994; Ciornei et al., 2005; Andraö et al., 2007; Savoia et al., 2008). Due to the presence of non-canonical amino acids, D-amino acids, and several other modifications, NRPs have commonly shown good proteolytic stability. Therefore, it is of interest to discover and further develop lipo-tridecapeptides with low hemolytic activity and cytotoxicity as potential antimicrobial agents.
In this study, four lipo-tridecapeptides were isolated from B. laterosporus DSM 25. Two of these compounds are known peptides, i.e., brevibacillin and brevibacillin V, while the other two compounds are novel lipo-tridecapeptides that we named brevibacillin I and brevibacillin 2V. All these four lipo-tridecapeptides showed a strong antimicrobial activity against Gram-positive pathogenic bacteria and showed synergistic effects with marketed antibiotics against Gram-negative pathogens. In addition, brevibacillin 2V showed good stability in human plasma. Notably, brevibacillin 2V did not exhibit a hemolytic activity when present at 128 mg/L, demonstrating its potency to be further studied and developed as a candidate antimicrobial agent for controlling specific antibiotic-resistant bacterial pathogens.
Materials and Methods
Bacterial Strains Used and Growth Conditions
The strains used in this study are listed in Supplementary Table 1. B. laterosporus DSM 25 cells were inoculated in LB and incubated at 37°C to prepare an overnight culture. For the production of brevibacillins, an overnight culture of B. laterosporus DSM 25 cells was inoculated (50-fold dilution) in minimal expression medium (MEM) and grown at 30°C for 36 h. All indicator strains were inoculated in LB and incubated at 37°C to prepare overnight cultures.
Purification of Cationic Peptides From a B. laterosporus DSM25 Culture
An overnight culture of B. laterosporus DSM 25 cells was inoculated in MEM and grown at 30°C for 36 h. Subsequently, the culture was centrifuged at 15,000 g for 15 min; the supernatant was collected and the pH was adjusted to 7.2 with a saturated sodium hydroxide solution. After that, the culture was applied to a CM SephadexTM C-25 column (GE Healthcare) equilibrated with distilled water. The flow-through was discarded, and the column was subsequently washed with 12 column volumes (CV) of distilled water. The peptide was eluted with 6 CV 2 M NaCl. The eluted peptide was then applied to a SIGMA-ALDRICH C18 Silica gel spherically equilibrated with 10 CV of Milli-Q water containing 5% (v/v) MeCN and 0.1% (v/v) trifluoroacetic acid. After washing with 10 CV of Milli-Q water containing 5% MeCN and 0.1% trifluoroacetic acid, the peptide was eluted from the column using up to 10 CV of Milli-Q water containing 50% MeCN and 0.1% trifluoroacetic acid. Fractions containing the eluted peptide were freeze-dried and dissolved in Milli-Q water. After filtration through a 0.2-μm filter, the cationic peptides were purified on an Agilent 1260 Infinity high-performance liquid chromatography (HPLC) system with a Phenomenex AerisTM C18 column (250 × 4.6 mm, 3.6 μm particle size, 100 Å pore size). Acetonitrile was used as the mobile phase, and a gradient of 30–45% MeCN/Milli-Q water (v/v; containing 0.1% trifluoroacetic acid) over 30 min at 1 ml per min was used for separation. The purified lipo-tridecapeptides were eluted with between 35 and 40% MeCN/Milli-Q water (v/v; containing 0.1% trifluoroacetic acid). The yields for brevibacillin, brevibacillin V, brevibacillin 2V, and brevibacillin I were 5, 3, 1.5, and 0.5 mg/L, respectively.
Mass Spectrometry
Matrix-assisted laser desorption ionization-time-of-flight (MALDI-TOF) mass spectrometer analysis was performed using a 4800 Plus MALDI TOF/TOF Analyzer (Applied Biosystems) in the linear-positive mode as in previous studies (Zhao et al., 2020a,b,c). Briefly, a 1-μl sample was spotted on the target and dried at room temperature. Subsequently, 0.6 μl of matrix solution (5 mg/ml of α-cyano-4-hydroxycinnamic acid) was spotted on each sample. After the samples had dried, MALDI-TOF MS was performed.
Liquid Chromatography–Tandem Mass Spectrometry Analysis
Since liquid chromatography–tandem mass spectrometry (LC–MS/MS) can be used to yield b and y ions for peptides, it is widely used in peptide structure elucidation (McClerren et al., 2006; Zhang et al., 2014; Zhao et al., 2020b). To gain insight into the molecular structures of the peptides, LC–MS/MS was performed using a Q-Exactive mass spectrometer fitted with an Ultimate 3000 UPLC, an ACQUITY BEH C18 column (2.1 × 50 mm, 1.7 μm particle size, 200 Å; Waters), a HESI ion source, and a Orbitrap detector. A gradient of 5–90% MeCN/Milli-Q water (v/v) with 0.1% formic acid (v/v), at a flow rate of 0.35 ml/min over 60 min, was used. MS/MS was performed in a separate run in parallel reaction monitoring mode, selecting the doubly and triply charged ion of the compound of interest.
Spot-on-Lawn Assay
Overnight-cultured S. aureus ATCC15975 (MRSA) was added to 0.7% Mueller–Hinton agar (w/v, temperature 42°C) at a final concentration of 0.25% (v/v), and then the mixture was poured onto plates, with 10 ml for each 8.5-cm-diameter circular plate or 20 ml for each 9.5-cm square plate. Subsequently, a spot-on-lawn assay was used to analyze the antimicrobial activity (Zhao et al., 2020c). In short, antibiotics were loaded to the agar plate that contains an indicator strain. After the antibiotic solution drops had dried, the plates were transferred to a 37°C incubator for overnight incubation.
Minimum Inhibitory Concentration Assay
Minimum inhibitory concentration (MIC) values were determined by using broth micro-dilution according to the standard guidelines (Wiegand et al., 2008). Briefly, the test medium was cation-adjusted Mueller–Hinton broth. Cell concentration was adjusted to approximately 5 × 105 cells per milliliter. After 20 h of incubation at 37°C, the MIC was defined as the lowest concentration of antibiotic with no visible growth (Zhao et al., 2020c). Each experiment was performed in triplicate.
Hemolytic Activity
This assay was performed as per the method described in previous studies (Ling et al., 2015; Li et al., 2018). Briefly, erythrocytes were isolated from a healthy human volunteer donor (ordered from Sanquin, Netherlands1) and washed with phosphate-buffered saline (PBS) three times. After that, peptides were added at final concentrations of 128, 64, 32, 16, 8, 4, 2, 1, and 0 mg/L in PBS containing 2% (v/v) erythrocytes. The cells were incubated at 37°C for 1 h and centrifuged for 5 min at 8,000 g. The supernatant was transferred to a 96-well plate, and the absorbance was measured at a wavelength of 450 nm with a Thermo Scientific VarioskanTM LUX multimode microplate reader. The absorbance relative to the positive control, which was treated with 10% Triton X-100, was defined as the percentage of hemolysis. The 50% human blood cell hemolysis concentration (HC50) of peptides was calculated as described in previous studies (Reed and Muench, 1938).
Mammalian Cytotoxicity
The cytotoxicity of brevibacillins was evaluated on HepG2 cells by using the XTT (Cell Proliferation Kit XTT, AppliChem) assay (Roehm et al., 1991). HepG2 cells [in Dulbecco’s modified Eagle’s medium (DMEM) supplemented with 10% fetal bovine serum] were seeded into 96-well plates and incubated at 37°C with 5% CO2. After 24 h, the medium was replaced with a fresh medium (DMEM with 2% FBS, 100 μl per well) containing different concentrations of brevibacillins (Zhao et al., 2017). After 24 h of incubation, the XTT reagent was added to the cultures according to the manufacturer’s instructions, and the plates were incubated at 37°C for 2 h with 5% CO2. Subsequently, the absorbance values were measured by using a VarioskanTM LUX multimode microplate reader (Thermo Fisher Scientific) at 485 nm (reference, 690 nm). The 50% cell toxicity (CC50) of peptides was calculated as described in a previous study (Reed and Muench, 1938).
Synergy Assay
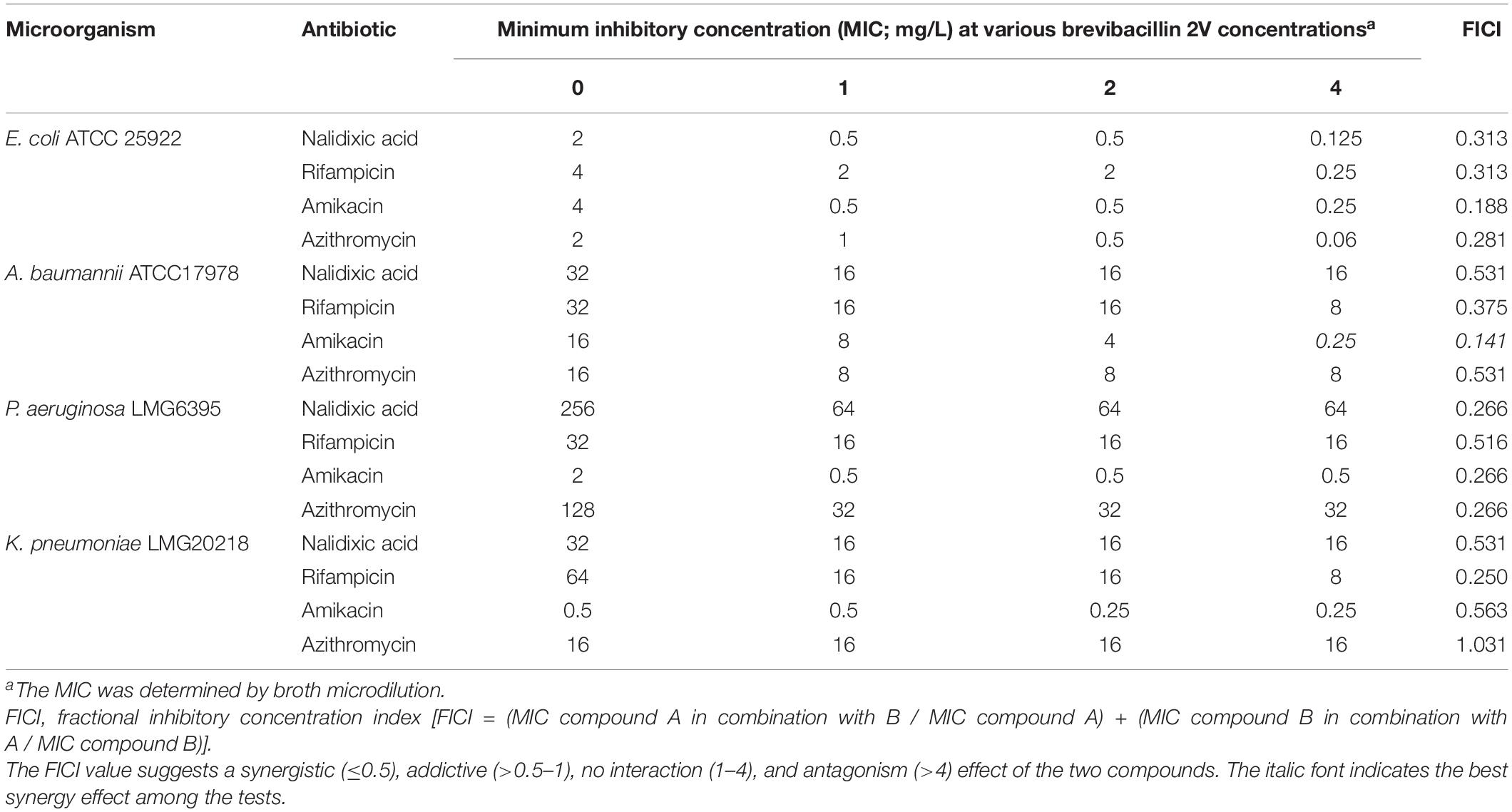
The synergistic effect of brevibacillins with established antibiotics was monitored using the checkerboard method (Doern, 2014). Briefly, antibiotics were diluted in twofold series, while brevibacillins were added at a certain final concentration of 1, 2, or 4 mg/L. Indicator strains were added at a final concentration of 5 × 105 cfu/ml. After incubation at 37°C for 20 h, the OD600 of plates was determined. The fractional inhibitory concentration index (FICI) was calculated using the following formula: FICI = (MIC compound A in combination with B / MIC compound A) + (MIC compound B in combination with A / MIC compound B). The FICI value suggests a synergistic (≤0.5), addictive (>0.5–1), no interaction (1–4), and antagonism (>4) effect of the two compounds (Doern, 2014).
Plasma Stability Assay
For plasma stability tests, brevibacillin and brevibacillin 2V were lyophilized and resuspended in plasma (isolated from a healthy human volunteer donor; ordered from Sanquin, Netherlands; see text footnote 1) at a final concentration of 200 mg/L. After that, the peptides were incubated with shaking (200 rpm) for 8 h (this evaluation time was set based on a continuing study, which showed that brevibacillins killed all S. aureus cells in 4 h) at 37°C. The samples were collected at desired time points (0, 1, 2, 4, and 8 h) and stored at −80°C. After the collection of samples of the last time point, the antimicrobial activity of all samples against S. aureus (MRSA) was investigated by a spot-on-lawn assay as per the method described above. The reduction in the size of the halo was related to the degradation of the peptide (Li et al., 2021).
Quantification and Statistical Analyses
GraphPad Prism 7.0 was used to fit the data of hemolytic assay and cytotoxicity assay in Figure 3. The experiments were conducted in triplicate, and data are presented as the mean value of triplicate experiments. Data Explorer Software was used to analyze the MALDI-TOF data; Thermo Scientific Xcalibur software was used to analyze the LC–MS/MS data. The statistical significance of the data was assessed using Duncan’s multiple-range test with the software SPSS Statistics 19.0. P-values less than 0.05 were considered to be statistically significant.
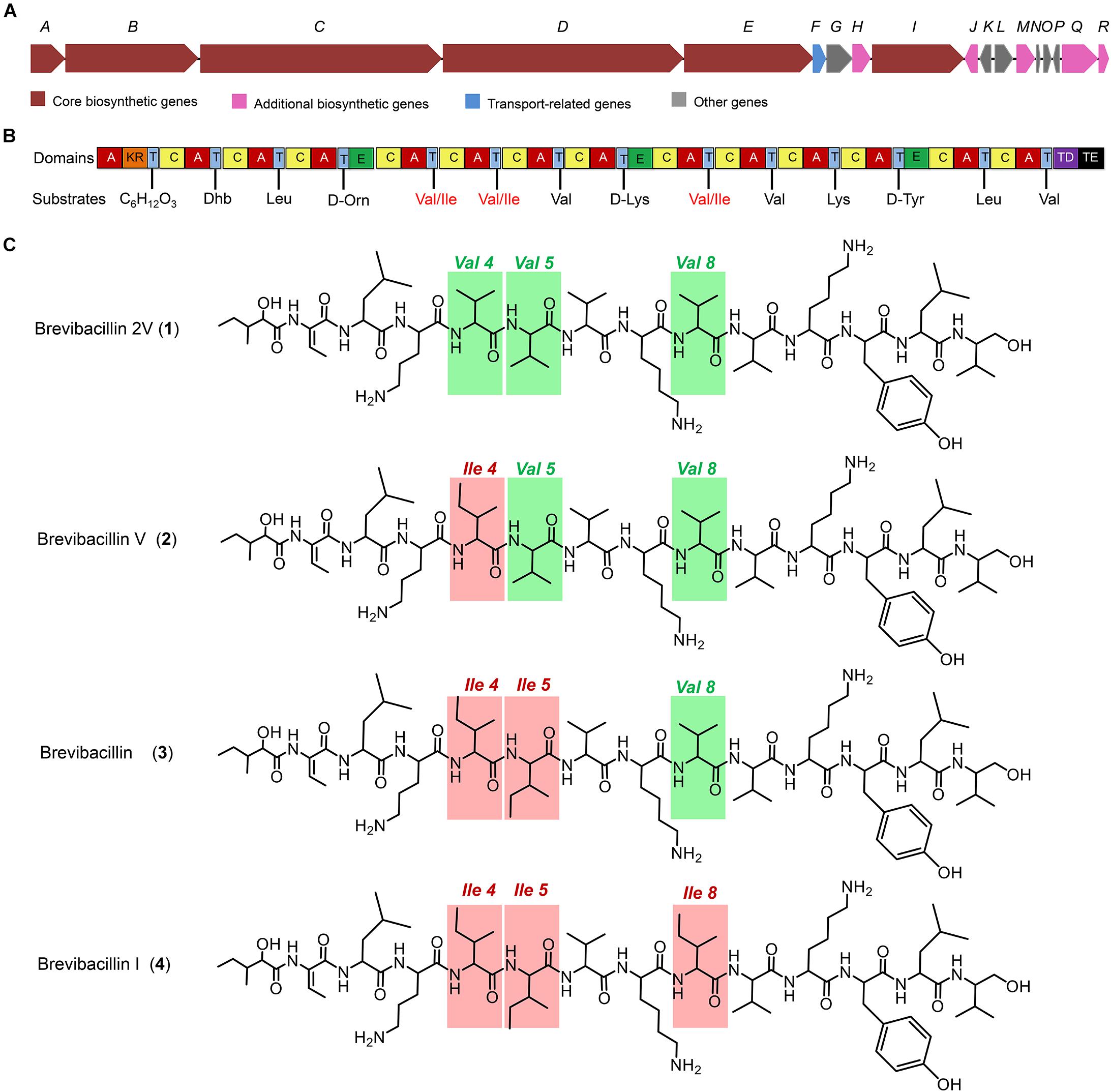
Figure 1. The structures of brevibacillins and the predicted biosynthetic gene cluster. (A) The non-ribosomal peptide synthetase genes harbored by the Brevibacillus laterosporus DSM 25 genome. (B) The catalytic domains encoded by the gene cluster and the substrates incorporated by the respective modules. Domains: A, adenylation; KR, ketoreductase; T, thiolation; C, condensation; E, epimerization; TD, terminal reductase; TE, thioesterase. (C) Structures of brevibacillins. The red rectangles indicate the more hydrophobic amino acid residues of brevibacillin V, brevibacillin, and brevibacillin I relative to brevibacillin 2V.
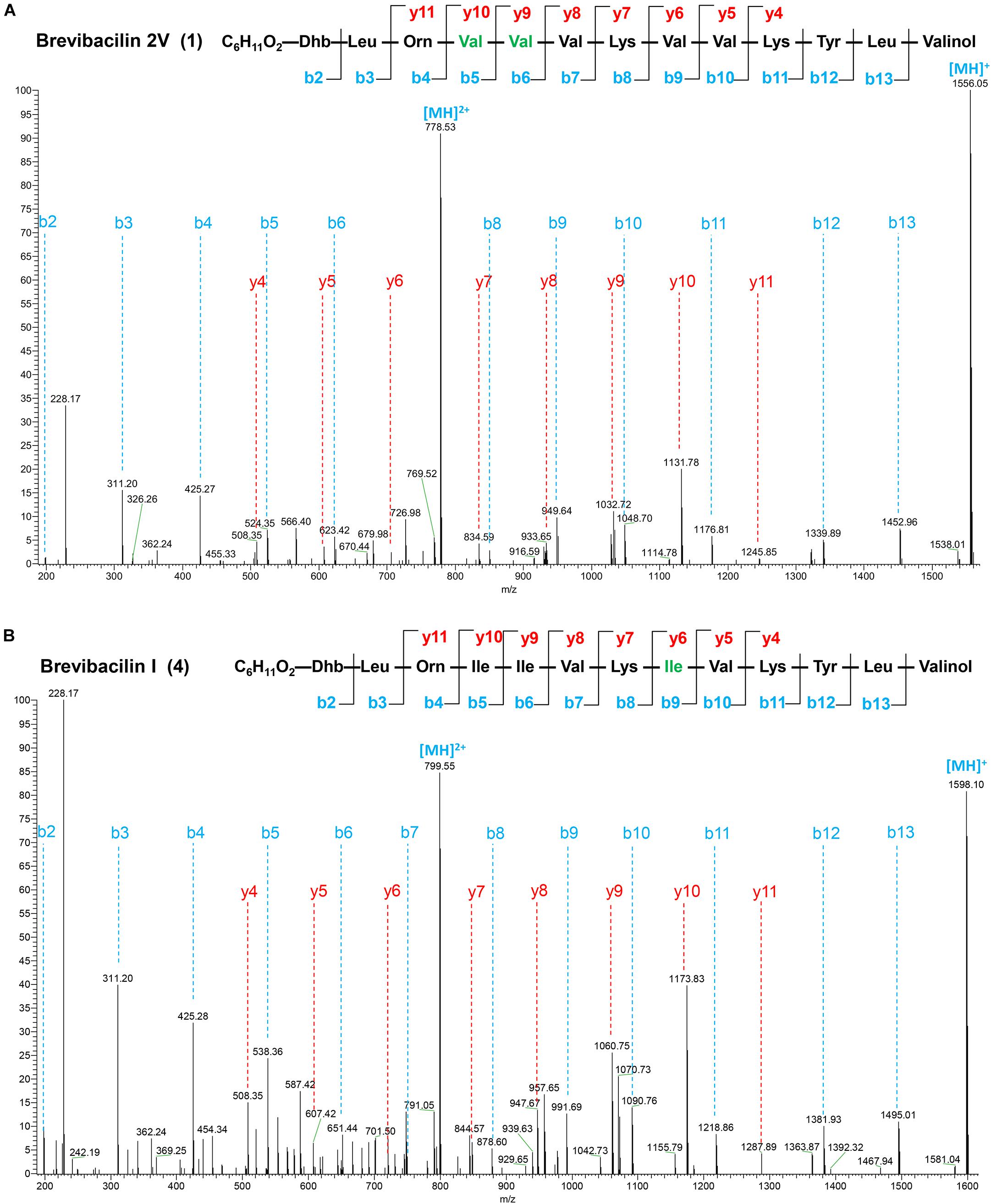
Figure 2. Liquid chromatography-tandem mass spectrometry (LC–MS/MS) spectra and the proposed structures of brevibacillin 2V (A) and brevibacillin I (B). Fragment ions are indicated. LC–MS/MS analysis was performed using a Q-Exactive mass spectrometer fitted with an Ultimate 3000 UPLC, an ACQUITY BEH C18 column (2.1 × 50 mm, 1.7 μm particle size, 200 Å; Waters), a HESI ion source, and a Orbitrap detector. MS/MS was performed in a separate run in parallel reaction monitoring mode, selecting the doubly and triply charged ion of the peptides. The structures of brevibacillin 2V and brevibacillin I were elucidated by using known lipo-tridecapeptides as reference templates.
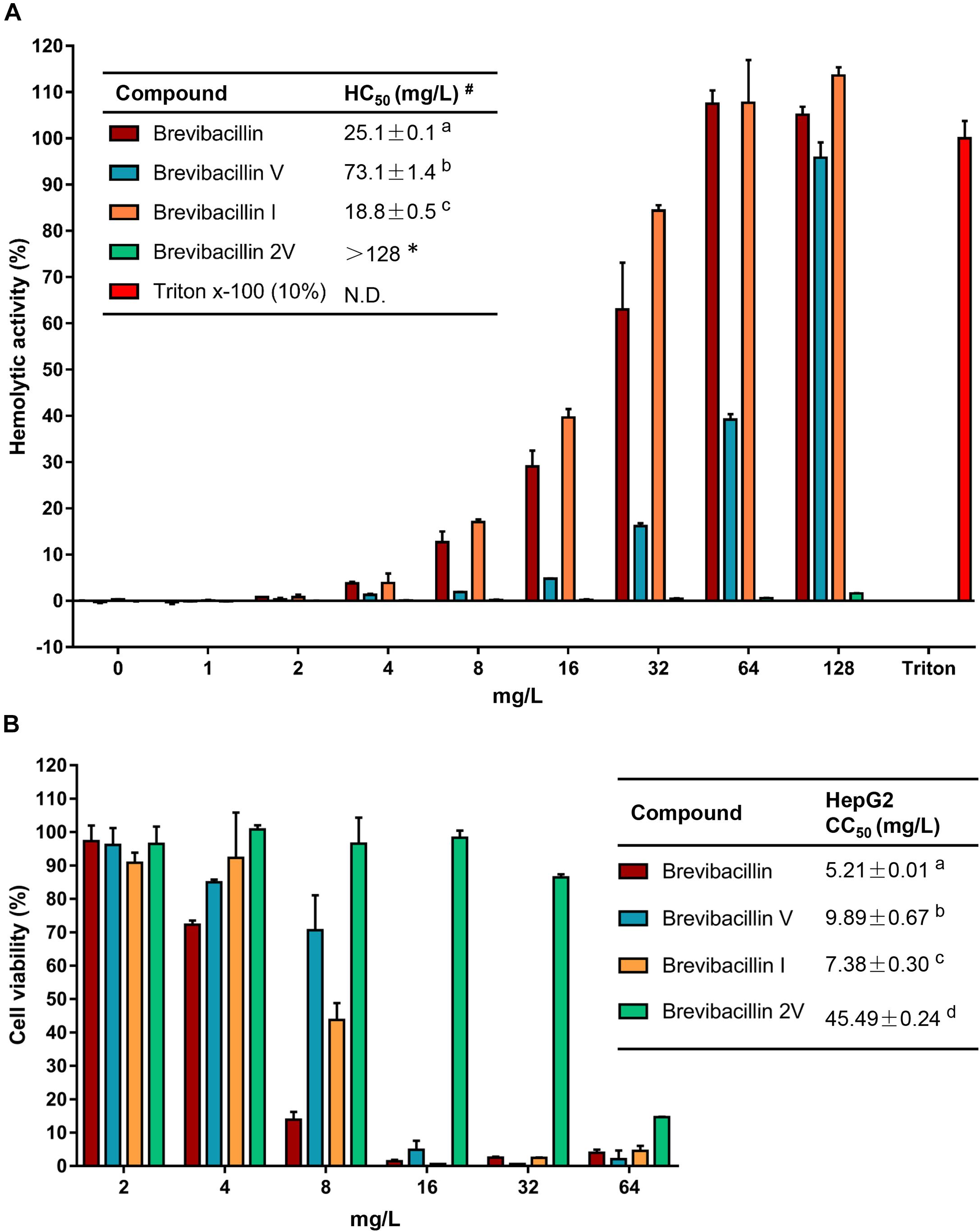
Figure 3. Brevibacillins activities against eukaryotic cells. (A) Human erythrocytes were incubated with brevibacillins at concentrations ranging from 1 to 128 mg/L. Their hemolytic activity was assessed by the release of hemoglobin. Cells treated without a tested compound were used as no lysis control. Cells treated with 10% Triton X-100 were used as complete lysis control. The data are representative of three independent experiments. #HC50 is the concentration that causes 50% hemolysis of human red blood cells. N.D., not determined. a,b,cSignificant differences exist when not the same letters are indicated in the HC50 column (p < 0.05). *No hemolytic activity was observed for brevibacillin 2V at 128 mg/L and its HC50> 128 mg/L. (B) Cytotoxicity of brevibacillins against HepG2 cells. HepG2 cells were treated with brevibacillins at concentrations ranging from 2 to 64 mg/L. Cells treated without a tested compound were used as untreated control. After 24 h of treatment, cell viability was determined by an XTT kit. The data are representative of three independent experiments. a,b,c,dSignificant differences exist when not the same letters are indicated in the CC50 column (p < 0.05).
Results
Characterization of Peptides Discovered by Genome Mining
Many lipopeptide antibiotics that originate from Brevibacillus spp. (Barsby et al., 2001, 2006; Yang et al., 2016; Yang and Yousef, 2018; Wu et al., 2019) have been discovered. In a previous study, a cyclic lipopeptide, brevicidine, was discovered from B. laterosporus DSM 25 by genome mining, which showed an antimicrobial activity against Gram-negative pathogenic bacteria (Li et al., 2018; Zhao et al., 2020b). More recently, brevicidineB, a natural analog of brevicidine, was discovered from B. laterosporus DSM 25, which showed an extended target specificity, showing an antimicrobial activity against both Gram-negative and Gram-positive pathogens (Zhao and Kuipers, 2021). In the present study, we found, by genome mining with the assistance of antiSMASH (Medema et al., 2011; Blin et al., 2019), that the genome (NCBI GenBank accession: GCA_002706795.1) of B. laterosporus DSM 25 contains a gene cluster (NCBI GenBank accession: KY810814.1.) for the synthesis of lipo-tridecapeptides (Figures 1A,B). B. laterosporus DSM 25 produced several cationic peptides that were purified by a CM SephadexTM C-25 column (GE Healthcare). After HPLC purification (Supplementary Figure 1), MALDI-TOF MS was used to measure the molecular weight of the purified peptides. Four compounds with a mass between 1,550 and 1,600 Da were found, i.e., compound 1, 1,555 Da; compound 2, 1,569 Da; compound 3, 1,583 Da; and compound 4, 1,597 Da (Supplementary Figures 1, 2), which potentially belong to the lipo-tridecapeptide family. Further characterization of these peptides was performed by LC–MS/MS analysis, and the results are shown in Figure 2 and Supplementary Figures 3, 4. The structures of the four compounds were elucidated by using known lipo-tridecapeptides as reference templates (Barsby et al., 2001, 2006; Yang et al., 2016; Yang and Yousef, 2018; Wu et al., 2019; Li et al., 2020). Two of these compounds are known peptides, i.e., brevibacillin (compound 3) and brevibacillin V (compound 2) (Yang et al., 2016; Wu et al., 2019), while the other two compounds are novel lipo-tridecapeptides that we named brevibacillin I (compound 4) and brevibacillin 2V (compound 1) (Figure 1C). In previous studies, brevibacillin and brevibacillin V showed a strong antimicrobial activity against Gram-positive bacteria (Yang et al., 2016; Wu et al., 2019). To investigate if the newly discovered brevibacillin I and brevibacillin 2V have similar or different antimicrobial activity with the known brevibacillins, we performed a spot-on-lawn assay using S. aureus ATCC15975 (MRSA) as an indicator strain. Nisin was used as a positive control. The results show that all brevibacillins have a comparable antimicrobial activity to each other against S. aureus (MRSA). In addition, they have a stronger antimicrobial activity against S. aureus (MRSA) than the well-known lantibiotic nisin (Supplementary Figure 5).
Brevibacillins Show a Strong Antimicrobial Activity Against Gram-Positive Bacterial Pathogens
After the spot-on-lawn assays have shown that brevibacillins have a good antimicrobial activity against S. aureus (MRSA), the antimicrobial activity of these lipo-tridecapeptides against pathogenic bacteria was further evaluated by MIC assays. All of the brevibacillins showed a strong antimicrobial activity against tested Gram-positive pathogenic bacteria, with a MIC value of 1–2 mg/L, including difficult-to-treat antibiotic-resistant Enterococcus faecium, Enterococcus faecalis, and S. aureus (Table 1). The similar antimicrobial activities indicate that the varying amino acid residues in the different lipo-tridecapeptides have no significant influence on their antimicrobial activity. In addition, brevibacillins showed an antimicrobial activity against Gram-negative pathogenic bacteria, yet these activities were much lower compared to the antimicrobial activity against Gram-positive bacteria (Table 1). These results are consistent with previous studies which show that lipo-tridecapeptides have an antimicrobial activity against Gram-positive bacteria but have an insufficient antimicrobial activity against Gram-negative bacteria (Barsby et al., 2001, 2006; Yang et al., 2016; Wu et al., 2019).
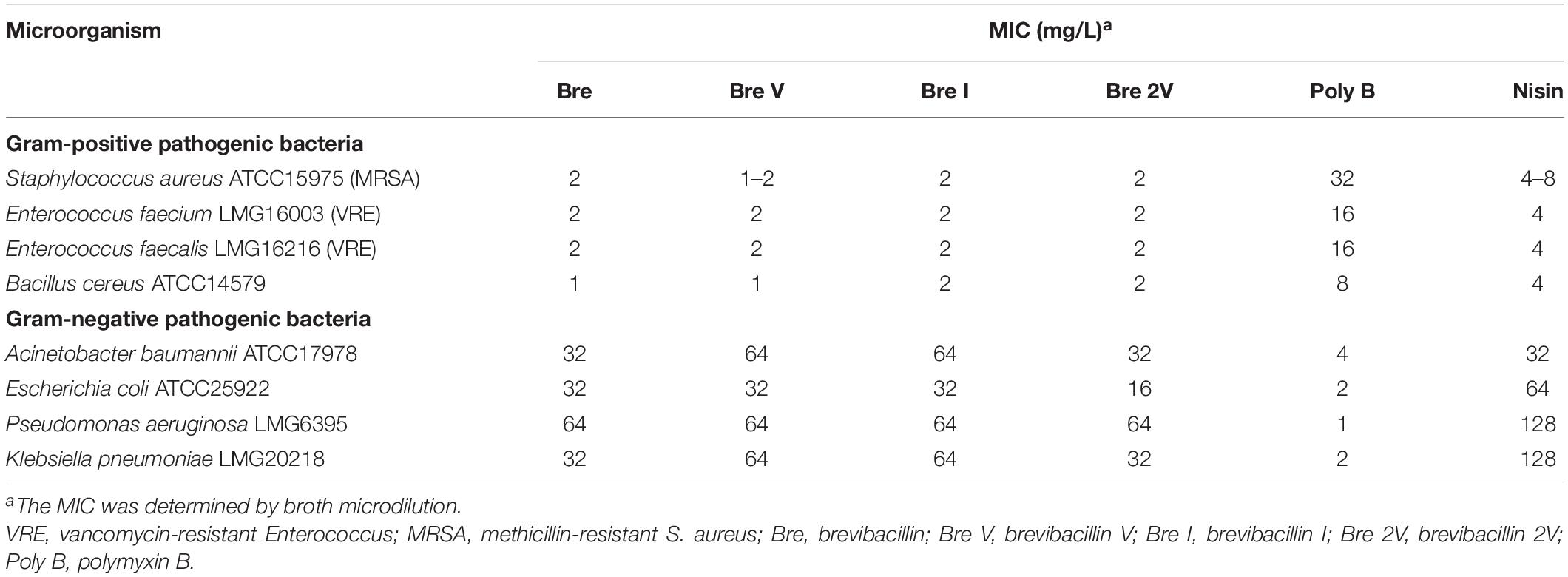
Table 1. Minimum inhibitory concentration (MIC) values of brevibacillins against pathogenic bacteria.
Brevibacillin 2V Does Not Exhibit Hemolytic Activity When Present at 128 mg/L
To assess in an initial test the safety of brevibacillins to human beings or animals, the hemolytic activity of brevibacillins to human blood cells and the cytotoxicity of brevibacillins to a human liver cell line (HepG2) were monitored. For the hemolytic activity assay, human blood cells were incubated in the presence of brevibacillin concentrations ranging from 1 to 128 mg/L. After incubation at 37°C for 1 h, the OD450 of the supernatants was measured, and the hemolytic activities of the brevibacillins were calculated as described in previous studies (Ling et al., 2015; Li et al., 2018). Brevibacillin, brevibacillin V, and brevibacillin I showed a significant hemolytic activity at concentrations that are two- to fourfold higher than their MIC values (Figure 3A and Table 1), with a half red blood cell hemolysis (HC50) concentration of 18.8 ± 0.5, 73.1 ± 1.4, and 25.1 ± 3.9, respectively. In addition, in the presence of either 64 mg/L of brevibacillin/brevibacillin I or 128 mg/L of brevibacillin V, human blood cells were completely lysed (Figure 3A). In contrast, brevibacillin 2V showed no hemolytic activity against human red blood cells at a high concentration of 128 mg/L (Figure 3A).
For cytotoxicity assays, HepG2 cells were incubated in the presence of brevibacillin concentrations ranging from 2 to 64 mg/L. After 24 h of incubation, cell viability was determined using an XTT kit. Under the experimental conditions used, brevibacillin, brevibacillin V, and brevibacillin I showed a high cytotoxicity to HepG2 cells with a half cell toxicity (CC50) value of 5–10 mg/L (Figure 3B), which is only 2.5- to fivefold of their MIC values against the bacterial pathogens (Figure 3B and Table 1). However, brevibacillin 2V showed much lower cytotoxicity than the other brevibacillins with a CC50 value of 45.5 mg/L against HepG2 cells, which is 23-fold higher than its MIC values against bacterial pathogens (Figure 3B and Table 1). In the presence of brevibacillin, brevibacillin V, or brevibacillin I at a concentration of 16 mg/L, all of the HepG2 cells were killed. These results are consistent with a previous study which shows that bogorol has high cytotoxicity with a CC50 of below 8 mg/L (Li et al., 2018). Together these results, with their high hemolytic activity, indicate that brevibacillin, brevibacillin V, brevibacillin I, and bogorol are too toxic to be developed as antibiotics for in vivo therapy. In contrast, brevibacillin 2V showed no cytotoxicity at the relatively high concentration of 32 mg/L (Figure 3B). Together with its non-hemolytic activity and low cytotoxicity, brevibacillin 2V shows a high potential, justifying its further study and development as an alternative antibiotic for controlling antibiotic-resistant bacterial pathogens.
Brevibacillins Show a Synergistic Activity With Other Antibiotics Against Gram-Negative Pathogenic Bacteria
Brevibacillins show a strong antimicrobial activity against Gram-positive bacterial pathogens but have insufficient antimicrobial activity against Gram-negative bacterial pathogens (Table 1). To expand the antimicrobial potential of brevibacillins, we investigated the synergistic activity of brevibacillins with some antibiotics (nalidixic acid, azithromycin, rifampicin, and amikacin) against Gram-negative pathogens. The synergistic effect of peptides was evaluated by determining a FICI value, which can indicate either a synergistic (≤0.5), addictive (>0.5–1), no interaction (1–4), or antagonistic (>4) effect of the two compounds (Doern, 2014). The results are shown in Table 2 and Supplementary Tables 2–4. Brevibacillin, brevibacillin V, brevibacillin I, and brevibacillin 2V showed a synergistic activity with the tested antibiotics against corresponding Gram-negative pathogens. Most importantly, brevibacillins showed the highest synergistic activity with amikacin against Acinetobacter baumannii in all tests (Table 2 and Supplementary Tables 2–4), with A. baumannii being one of the three critical priority pathogens for R&D of new antibiotics (World Health Organization, 2017). The MIC value of amikacin against A. baumannii decreased by 32–64 folds in the presence of 4 mg/L brevibacillins (Table 2 and Supplementary Tables 2–4). Together with its non-hemolytic activity, low cytotoxicity, and the effective low MIC values, brevibacillin 2V shows a high potential to be developed as an adjuvant for other antibiotics to treat Gram-negative bacterial pathogen infections.
Brevibacillins Show Good Stability in Plasma
In view of the therapeutic development potential of brevibacillin 2V, the stability of brevibacillin 2V and brevibacillin (highest yield among brevibacillins; to be used as a control to compare with brevibacillin 2V) in human plasma was investigated, using the method described in a previous study (Li et al., 2021). Brevibacillins were incubated in human plasma, and the reduction in antimicrobial activity was measured by spot-on-lawn assays against S. aureus (MRSA). Brevibacillins showed good stability in plasma during 4 h after exposure since no reduction in the sizes of the halos was observed (Figure 4). After 8 h of exposure in plasma, the sizes of the halos for brevibacillin 2V and brevibacillin only decreased a little bit (halo diameter decreased from 11 to 10 mm for brevibacillin 2V and from 11 to 10.5 mm for brevibacillin) (Figure 4). The good plasma stability of brevibacillin 2V might be due to the presence of non-canonical amino acids and D-amino acids, which are highly resistant to proteases (Li et al., 2021). These results demonstrate that brevibacillin 2V has good stability in plasma, which makes it more attractive for development as an antibiotic candidate.
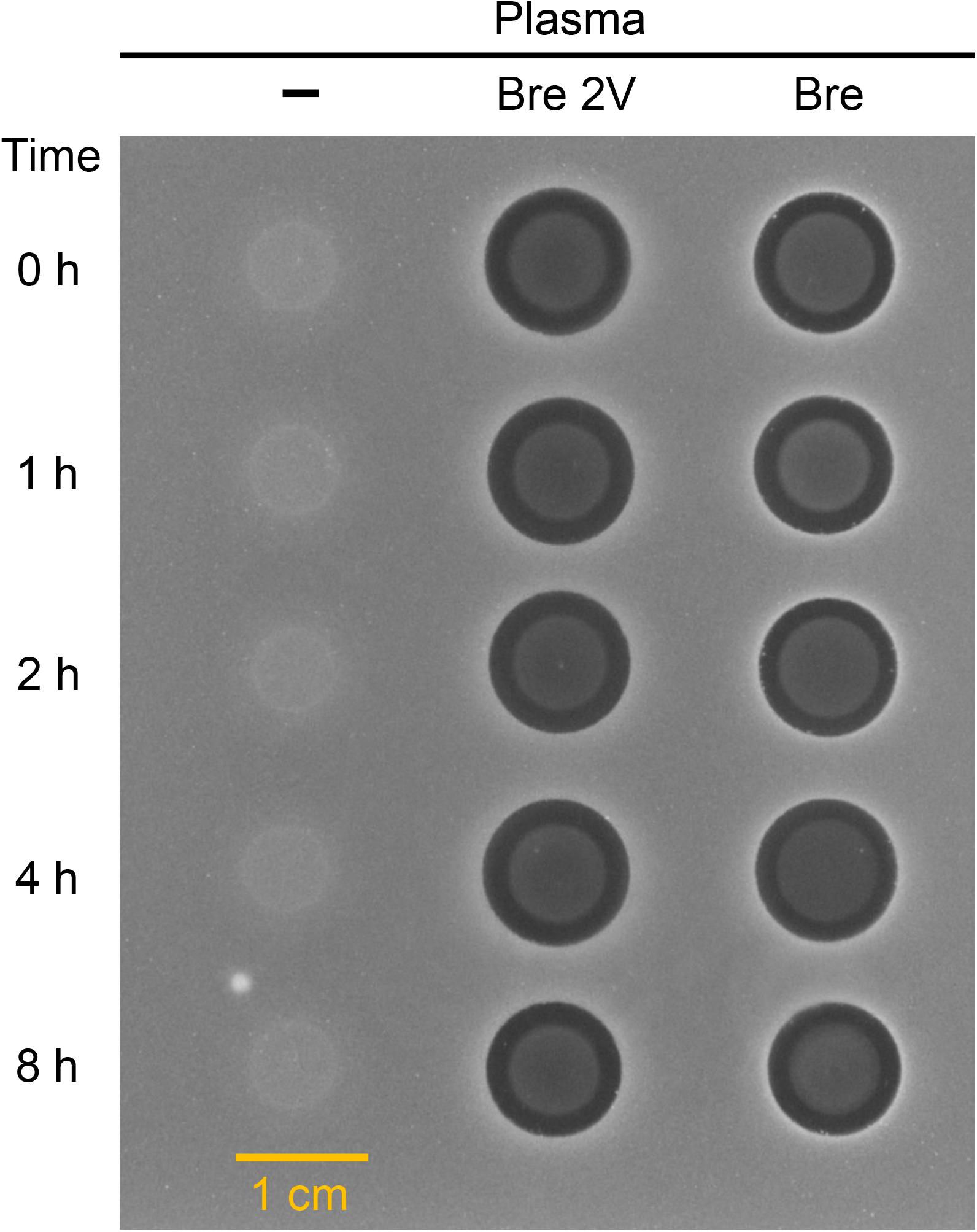

Figure 4. Antimicrobial activity of brevibacillin 2V brevibacillin against Staphylococcus aureus (MRSA) after exposure to human plasma. The starting concentration was 200 mg/L for both brevibacillin 2V and brevibacillin. Samples were collected at 0, 1, 2, 4, and 8 h, and 6 μl of each sample was loaded to a square plate containing S. aureus (MRSA). Bre 2V, brevibacillin 2V; Bre, brevibacillin.
Discussion
Genome mining is a practical approach for discovering bioactive natural products from microorganisms (Montalbán-López et al., 2021). Genome mining tools with different specific uses have been developed, including BAGEL4 (van Heel et al., 2018), antiSMASH (Blin et al., 2019), and RiPP-PRISM (Skinnider et al., 2015). AntiSMASH is a well-known genome mining tool for discovering NRPs with antimicrobial activity (Blin et al., 2019). For instance, two non-ribosomally produced cyclic lipopeptides, i.e., brevicidine and laterocidine, were discovered by genome mining with the assistance of antiSMASH (Li et al., 2018). In this study, a lipo-tridecapeptide synthetic gene cluster was discovered from B. laterosporus DSM 25 by genome mining with the assistance of antiSMASH (Medema et al., 2011; Blin et al., 2019). Subsequently, antimicrobial lipo-tridecapeptides were isolated and identified by following the pipeline in Figure 5.
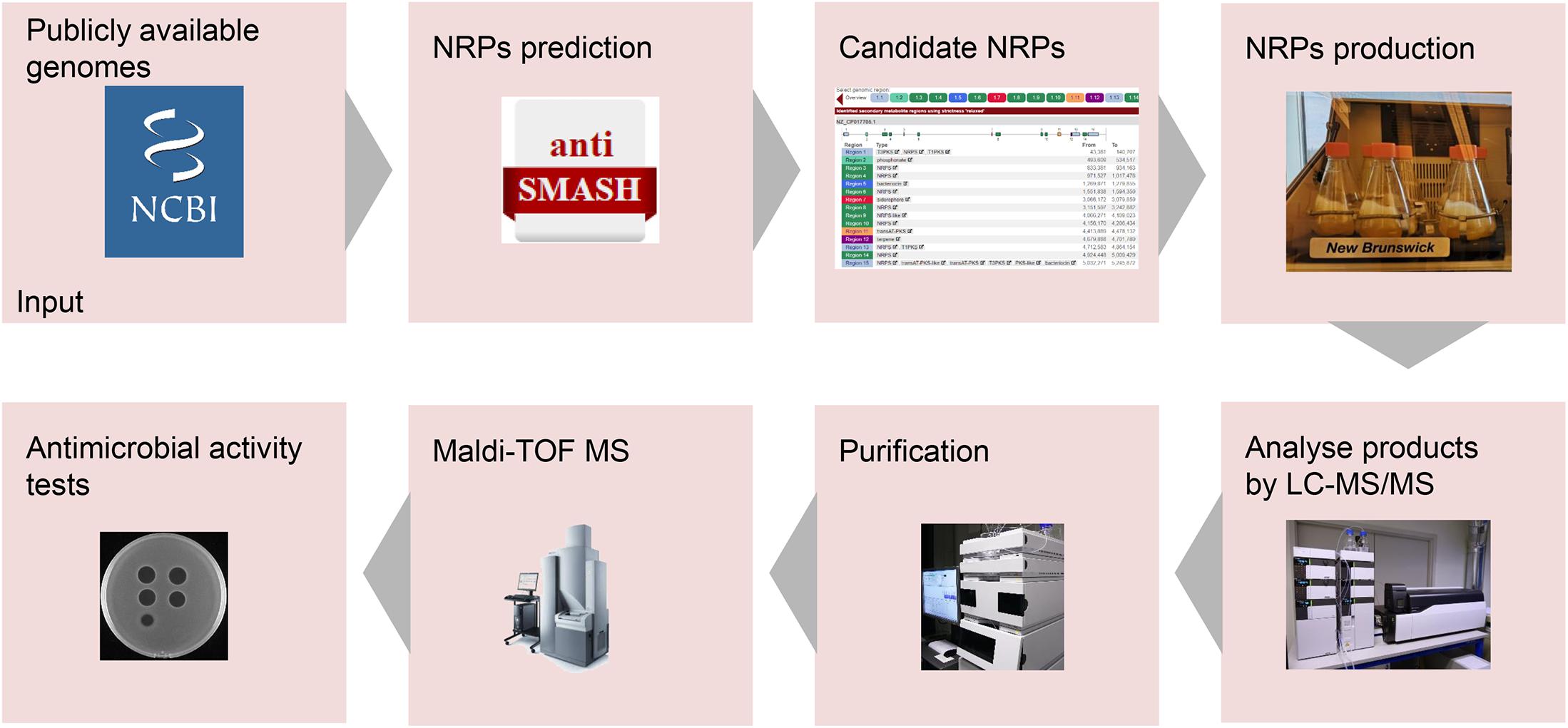
Figure 5. Pipeline for the development of novel non-ribosomally produced peptide (NRP)-based antimicrobials. Following the arrows, the process starts with the identification of NRP clusters. Next, the predicted NRPs have to be produced, isolated, and characterized. Finally, the antimicrobial activity of the purified NRPs will be determined.
In this study, we isolated the predicted lipo-tridecapeptides from the supernatant of B. laterosporus DSM 25 culture. This allows an iron change-based CM SephadexTM C-25 column to be applied to purify the produced products since lipo-tridecapeptides are cationic peptides. In contrast, lipo-tridecapeptides were previously isolated from bacterial cells (Barsby et al., 2001, 2006; Yang et al., 2016), which is more difficult to process. Thus, the here found lipo-tridecapeptides are much more readily purified than the previously reported lipo-tridecapeptides. For elucidation of the purified lipo-tridecapeptides structures, we used the previously reported lipo-tridecapeptides as templates. Such template-based structure elucidation approach has been successfully used in previous studies to elucidate peptide structures (Barsby et al., 2006; Wu et al., 2019). These results suggest that a template-based structure elucidation is a helpful approach for elucidating the structures of peptides that belong to the same type. All structures were confirmed by MS.
Many lipo-tridecapeptides have shown an antimicrobial activity against pathogenic bacteria, including antibiotic-resistant S. aureus and Enterococcus spp. (Barsby et al., 2006; Yang et al., 2016; Wu et al., 2019). However, the high hemolytic activity of lipo-tridecapeptides has limited their potential for being developed as therapeutics (Figure 3; Li et al., 2018). In this study, one of the discovered novel lipo-tridecapeptides, brevibacillin 2V, showed no hemolytic activity at a high concentration of 128 mg/L (Figure 3). Moreover, the hemolytic activity of brevibacillins shows a positive correlation with their calculated hydrophobicity, i.e., brevibacillin I > brevibacillin > brevibacillin V > brevibacillin 2V (Figure 1C and Supplementary Figure 1). These results demonstrate that brevibacillin 2V shows a high potential to be further studied and developed as an alternative antibiotic for controlling antibiotic-resistant bacterial pathogens. In addition, the positive correlation of hemolytic activity and hydrophobicity provides a guideline for the synthesis of lipo-tridecapeptide analogs in future studies. The use of peptides as adjuvants for marketed antibiotics has attracted the attention of several researchers. Cochrane and Vederas (2014) reported that unacylated tridecaptin A1 acts as an effective sensitizer of Gram-negative bacteria to other antibiotics. More recently, Li et al. (2021) reported that outer-membrane-acting peptides showed a synergistic activity with lipid II-targeting antibiotics against Gram-negative pathogens. In the present study, brevibacillins showed a synergistic activity with marketed antibiotics against Gram-negative pathogens. These results suggest that more antibiotics can be tested for developing brevibacillins (at least, brevibacillin 2V) as adjuvants for other antibiotics to treat Gram-negative bacterial pathogen infections.
Conclusion
In conclusion, in this study, we characterized a novel lipo-tridecapeptide, i.e., brevibacillin 2V, which was identified from B. laterosporus DSM 25 by genome mining and subsequently isolated and purified. Brevibacillin 2V has a strong antimicrobial activity against antibiotic-resistant Gram-positive bacterial pathogens, and it shows an effective synergistic activity with other antibiotics against Gram-negative bacterial pathogens. Notably, brevibacillin 2V has a much lower hemolytic activity and cytotoxicity towards eukaryotic cells than previously reported NRPs of the lipo-tridecapeptide family, including all known brevibacillins, which makes it a promising candidate for antibiotic development. In addition, brevibacillin 2V showed good stability in human plasma. This study provides a novel and promising antibiotic candidate (brevibacillin 2V) with low hemolytic activity, which can be used either directly as it is or serves as a template for further total synthesis and modifications.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding author/s.
Author Contributions
OK and XZ conceived the project. XZ designed and carried out the experiments, analyzed the data, and wrote the manuscript. OK and EB supervised the work and corrected the manuscript. XW, RS, RK, and MW did the experimental work. All authors contributed to and commented on the manuscript text and approved its final version.
Funding
XZ was financed by the Netherlands Organization for Scientific Research (NWO), research program TTW (17241). XW was funded by the China Scholarship Council (201508330301). RS was funded by the Dutch Research Council (NWO) (711.018.001).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Marcel P. de Vries (University of Groningen) for support with the LC–MS/MS.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fmicb.2021.693725/full#supplementary-material
Footnotes
References
Andraö, J., Monreal, D., de Tejada, G. M., Olak, C., Brezesinski, G., Gomez, S. S., et al. (2007). Rationale for the design of shortened derivatives of the NK-lysin-derived antimicrobial peptide NK-2 with improved activity against Gram-negative pathogens. J. Biol. Chem. 282, 14719–14728. doi: 10.1074/jbc.m608920200
Barsby, T., Kelly, M. T., Gagné, S. M., and Andersen, R. J. (2001). Bogorol A produced in culture by a marine Bacillus sp. reveals a novel template for cationic peptide antibiotics. Org. Lett. 3, 437–440. doi: 10.1021/ol006942q
Barsby, T., Warabi, K., Sørensen, D., Zimmerman, W. T., Kelly, M. T., and Andersen, R. J. (2006). The bogorol family of antibiotics: template-based structure elucidation and a new approach to positioning enantiomeric pairs of amino acids. J. Org. Chem. 71, 6031–6037. doi: 10.1021/jo060667p
Blin, K., Shaw, S., Steinke, K., Villebro, R., Ziemert, N., Lee, S. Y., et al. (2019). antiSMASH 5.0: updates to the secondary metabolite genome mining pipeline. Nucleic Acids Res. 47, W81–W87.
Ciornei, C. D., Sigurdardóttir, T., Schmidtchen, A., and Bodelsson, M. (2005). Antimicrobial and chemoattractant activity, lipopolysaccharide neutralization, cytotoxicity, and inhibition by serum of analogs of human cathelicidin LL-37. Antimicrob. Agents Chemother. 49, 2845–2850. doi: 10.1128/aac.49.7.2845-2850.2005
Cochrane, S. A., Lohans, C. T., Brandelli, J. R., Mulvey, G., Armstrong, G. D., and Vederas, J. C. (2014). Synthesis and structure–activity relationship studies of N-terminal analogues of the antimicrobial peptide tridecaptin A1. J. Med. Chem. 57, 1127–1131. doi: 10.1021/jm401779d
Cochrane, S. A., and Vederas, J. C. (2014). Unacylated tridecaptin A1 acts as an effective sensitiser of Gram-negative bacteria to other antibiotics. Int. J. Antimicrob. Agents 44, 493–499. doi: 10.1016/j.ijantimicag.2014.08.008
Cociancich, S., Pesic, A., Petras, D., Uhlmann, S., Kretz, J., Schubert, V., et al. (2015). The gyrase inhibitor albicidin consists of p-aminobenzoic acids and cyanoalanine. Nat. Chem. Biol. 11:195. doi: 10.1038/nchembio.1734
Doern, C. D. (2014). When does 2 plus 2 equal 5? A review of antimicrobial synergy testing. J. Clin. Microbiol. 52, 4124–4128. doi: 10.1128/jcm.01121-14
Hamamoto, H., Urai, M., Ishii, K., Yasukawa, J., Paudel, A., Murai, M., et al. (2015). Lysocin E is a new antibiotic that targets menaquinone in the bacterial membrane. Nat. Chem. Biol. 11, 127. doi: 10.1038/nchembio.1710
Li, Q., Cebrián, R., Montalbán-López, M., Ren, H., Wu, W., and Kuipers, O. P. (2021). Outer-membrane-acting peptides and lipid II-targeting antibiotics cooperatively kill Gram-negative pathogens. Commun. Biol. 4, 1–11.
Li, Y.-X., Zhong, Z., Zhang, W.-P., and Qian, P.-Y. (2018). Discovery of cationic nonribosomal peptides as Gram-negative antibiotics through global genome mining. Nat. Commun. 9:3273.
Li, Z., de Vries, R. H., Chakraborty, P., Song, C., Zhao, X., Scheffers, D.-J., et al. (2020). Novel modifications of nonribosomal peptides from Brevibacillus laterosporus MG64 and investigation of their mode of action. Appl. Environ. Microbiol. 2020:86.
Ling, L. L., Schneider, T., Peoples, A. J., Spoering, A. L., Engels, I., Conlon, B. P., et al. (2015). A new antibiotic kills pathogens without detectable resistance. Nature 517:455.
McClerren, A. L., Cooper, L. E., Quan, C., Thomas, P. M., Kelleher, N. L., and Van Der Donk, W. A. (2006). Discovery and in vitro biosynthesis of haloduracin, a two-component lantibiotic. Proc. Natl. Acad. Sci. 103, 17243–17248. doi: 10.1073/pnas.0606088103
Medema, M. H., Blin, K., Cimermancic, P., de Jager, V., Zakrzewski, P., Fischbach, M. A., et al. (2011). antiSMASH: rapid identification, annotation and analysis of secondary metabolite biosynthesis gene clusters in bacterial and fungal genome sequences. Nucleic Acids Res. 39, W339–W346.
Montalbán-López, M., Scott, T. A., Ramesh, S., Rahman, I. R., van Heel, A. J., Viel, J. H., et al. (2021). New developments in RiPP discovery, enzymology and engineering. Nat. Prod. Rep. 38, 130–239. doi: 10.1039/d0np00027b
Mor, A., and Nicolas, P. (1994). The NH2-terminal alpha-helical domain 1-18 of dermaseptin is responsible for antimicrobial activity. J. Biol. Chem. 269, 1934–1939. doi: 10.1016/s0021-9258(17)42116-8
World Health Organization (2017). Prioritization of pathogens to guide discovery, research and development of new antibiotics for drug-resistant bacterial infections, including tuberculosis. Geneva: World Health Organization.
Reed, L. J., and Muench, H. (1938). A simple method of estimating fifty per cent endpoints. Am. J. Epidemiol. 1938:27.
Roehm, N. W., Rodgers, G. H., Hatfield, S. M., and Glasebrook, A. L. (1991). An improved colorimetric assay for cell proliferation and viability utilizing the tetrazolium salt XTT. J. Immunol. Methods 142, 257–265. doi: 10.1016/0022-1759(91)90114-u
Savoia, D., Guerrini, R., Marzola, E., and Salvadori, S. (2008). Synthesis and antimicrobial activity of dermaseptin S1 analogues. Bioorg. Med. Chem. 16, 8205–8209. doi: 10.1016/j.bmc.2008.07.032
Skinnider, M. A., Dejong, C. A., Rees, P. N., Johnston, C. W., Li, H., Webster, A. L. H., et al. (2015). Genomes to natural products prediction informatics for secondary metabolomes (PRISM). Nucleic Acids Res. 43, 9645–9662.
Süssmuth, R. D., and Mainz, A. (2017). Nonribosomal peptide synthesis—principles and prospects. Angew. Chemie Int. Ed. 56, 3770–3821. doi: 10.1002/anie.201609079
van Heel, A. J., de Jong, A., Song, C., Viel, J. H., Kok, J., and Kuipers, O. P. (2018). BAGEL4: a user-friendly web server to thoroughly mine RiPPs and bacteriocins. Nucleic Acids Res. 46, W278–W281.
Wiegand, I., Hilpert, K., and Hancock, R. E. W. (2008). Agar and broth dilution methods to determine the minimal inhibitory concentration (MIC) of antimicrobial substances. Nat. Protoc. 3:163. doi: 10.1038/nprot.2007.521
Wu, Y., Zhou, L., Lu, F., Bie, X., Zhao, H., Zhang, C., et al. (2019). Discovery of a Novel Antimicrobial Lipopeptide, Brevibacillin V, from Brevibacillus laterosporus fmb70 and Its Application on the Preservation of Skim Milk. J. Agric. Food Chem. 67, 12452–12460. doi: 10.1021/acs.jafc.9b04113
Yang, X., Huang, E., Yuan, C., Zhang, L., and Yousef, A. E. (2016). Isolation and structural elucidation of brevibacillin, an antimicrobial lipopeptide from Brevibacillus laterosporus that combats drug-resistant Gram-positive bacteria. Appl. Environ. Microbiol. 82, 2763–2772. doi: 10.1128/aem.00315-16
Yang, X., and Yousef, A. E. (2018). Antimicrobial peptides produced by Brevibacillus spp.: structure, classification and bioactivity: a mini review. World J. Microbiol. Biotechnol. 34:57.
Zhang, Q., Ortega, M., Shi, Y., Wang, H., Melby, J. O., Tang, W., et al. (2014). Structural investigation of ribosomally synthesized natural products by hypothetical structure enumeration and evaluation using tandem MS. Proc. Natl. Acad. Sci. 111, 12031–12036. doi: 10.1073/pnas.1406418111
Zhao, X., and Kuipers, O. P. (2021). BrevicidineB, a new member of the brevicidine family, displays an extended target specificity. Front. Microbiol. 12:1482.
Zhao, X., Cebrián, R., Fu, Y., Rink, R., Bosma, T., Moll, G. N., et al. (2020a). High-Throughput Screening for Substrate Specificity-Adapted Mutants of the Nisin Dehydratase NisB. ACS Synth. Biol. 9, 1468–1478. doi: 10.1021/acssynbio.0c00130
Zhao, X., Cui, Q., Fu, Q., Song, X., Jia, R., Yang, Y., et al. (2017). Antiviral properties of resveratrol against pseudorabies virus are associated with the inhibition of IκB kinase activation. Sci. Rep. 7:09365. doi: 10.1038/s41598-017-09365-0
Zhao, X., Li, Z., and Kuipers, O. P. (2020b). Mimicry of a Non-ribosomally Produced Antimicrobial, Brevicidine, by Ribosomal Synthesis and Post-translational Modification. Cell Chem. Biol. 27, 1262.e–1271.e. doi: 10.1016/j.chembiol.2020.07.005
Keywords: antimicrobial activity, lipopeptide, NRPS, brevibacillin, Brevibacillus, Enterococcus, Staphylococcus
Citation: Zhao X, Wang X, Shukla R, Kumar R, Weingarth M, Breukink E and Kuipers OP (2021) Brevibacillin 2V, a Novel Antimicrobial Lipopeptide With an Exceptionally Low Hemolytic Activity. Front. Microbiol. 12:693725. doi: 10.3389/fmicb.2021.693725
Received: 11 April 2021; Accepted: 17 May 2021;
Published: 17 June 2021.
Edited by:
Jianhua Wang, Gene Engineering Laboratory, Feed Research Institute, Chinese Academy of Agricultural Sciences, ChinaReviewed by:
Anders Løbner-Olesen, University of Copenhagen, DenmarkJoseph Boll, University of Texas at Arlington, United States
Copyright © 2021 Zhao, Wang, Shukla, Kumar, Weingarth, Breukink and Kuipers. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Oscar P. Kuipers, by5wLmt1aXBlcnNAcnVnLm5s
 Xinghong Zhao
Xinghong Zhao Xiaoqi Wang
Xiaoqi Wang Rhythm Shukla
Rhythm Shukla Raj Kumar
Raj Kumar Markus Weingarth
Markus Weingarth Eefjan Breukink
Eefjan Breukink Oscar P. Kuipers
Oscar P. Kuipers