- Key Laboratory of National Forestry and Grassland Administration for Control of Diseases and Pests of South Plantation, Hunan Provincial Key Laboratory for Control of Forest Diseases and Pests, Key Laboratory for Non-wood Forest Cultivation and Conservation of Ministry of Education, Central South University of Forestry and Technology, Changsha, China
Oil tea (Camellia oleifera), mainly used to produce high-quality edible oil, is an important cash crop in China. Anthracnose of oil tea is a considerable factor that limits the yield of tea oil. In order to effectively control the anthracnose of oil tea, researchers have worked hard for many years, and great progress has been made in the research of oil tea anthracnose. For instance, researchers isolated a variety of Colletotrichum spp. from oil tea and found that Colletotrichum fructicola was the most popular pathogen in oil tea. At the same time, a variety of control methods have been explored, such as cultivating resistant varieties, pesticides, and biological control, etc. Furthermore, the research on the molecular pathogenesis of Colletotrichum spp. has also made good progress, such as the elaboration of the transcription factors and effector functions of Colletotrichum spp. The authors summarized the research status of the harm, pathogen types, control, and pathogenic molecular mechanism of oil tea anthracnose in order to provide theoretical support and new technical means for the green prevention and control of oil tea anthracnose.
Introduction
Cultivation and application of oil tea
Oil tea generally refers to the Camellia genus, which has rich seed oil content that is produced and highly valuable (Chen, 2008). The history of extracting oil from oil teaseed in China can be traced back to 2,300 years ago (Zhuang, 2008). The main varieties of oil tea cultivated in China include Camellia yuhsienensis Hu, Camellia oleifera var monosperma, Camellia vietnamensis, and Camellia oleifera, among which the planting range of Ca. oleifera is the widest (Qin et al., 2018; Chen et al., 2021b). Tea oil extracted from the seed of oil tea is rich in unsaturated fatty acids and vitamin E and has unique nutritional value (Shi et al., 2020). Thus, oil tea is as famous as coconut, palm, and olive, and is also known as one of the four major woody oil plants in the world (Yang et al., 2016, 2020a). Moreover, the United Nations Food and Agriculture Organization (FAO) recommended tea oil as a high-quality and healthy vegetable oil, owing to its nutritional value and excellent storage quality (Chen et al., 2020b). In 2020, the area in China planted oil tea reached 45,333.3 km2; the output of tea oil reached 627,000 tons, and the output value of the tea oil industry reached 18 billion U.S. dollars, indicating that tea oil is highly valuable (Chen et al., 2021a).
Anthracnose of oil tea
Anthracnose of oil tea is a considerable factor that limits the yield of tea oil (Chen et al., 2020a; Li et al., 2021b). Anthracnose of oil tea is a considerable factor that limits the yield of tea oil (Chen et al., 2020a; Li et al., 2021b). The conidia of Colletotrichum spp. mainly infect the oil tea from the wound but also through the natural pore, such as stoma. Attached conidia germinate and differentiate dome-shaped appressoria on plant surfaces, underneath which penetration pegs form and penetrate epidermal cells. The pathogen then differentiates bulbous biotrophic hyphae, which are enveloped by an intact host plasma membrane; biotrophic hyphae spread across living host cells before differentiating thin necrotrophichyphae that kill and destroy host tissues (Münch et al., 2008; O'Connell et al., 2012).
Colletotrichum spp. primarily infect the leaves and fruits of the oil tea, leading to a 20–40% fruit drop and up to 40% seed loss (Jin et al., 2009; Zhu et al., 2015). In addition, the oil content of oil tea seeds infected by Colletotrichum spp. can be reduced by 50%. Moreover, the anthracnose of oil tea can also cause the germination of infected seeds, which facilitates the long-distance transmission of oil tea anthracnose (Yang, 2009). Therefore, under the conditions of appropriate humidity and ambient temperature, anthracnose of oil tea spreads rapidly and is difficult to control, causing substantial economic losses and seriously damaging the safety of edible oil in China (Liu et al., 2009; Deng, 2011). Colletotrichum spp. are also regarded as among the top 10 plant pathogenic fungi in the field of molecular plant pathology because of their strong pathogenicity and wide spread (Dean et al., 2012).
Colletotrichum spp. was first discovered by Tode in 1790 and named Vermicularia Tode (Tode, 1790). Then, it was further subdivided according to other morphological characteristics and named Colletotrichum Corda (Sturm, 1832; Sutton, 1992). With the development of molecular biology, more and more scholars used multi-gene lineage to identify Colletotrichum spp., which not only improved the accuracy of Colletotrichum spp. identification but also identified more species of Colletotrichum spp. Talkin has identified Colletotrichum spp. according to internal transcribed spacer (ITS), histone 4 (HIS4), and β-tubulin 2 (TUB2) gene polygenic sequences for the first time (Talhinhas et al., 2002).
Nowadays, more and more Colletotrichum spp. has been identified, and the Colletotrichum spp. that can infect oil tea mainly include Colletotrichum fructicola, Colletotrichum gloeosporioides, Colletotrichum horii, Colletotrichum siamense, Colletotrichum camelliae, and Colletotrichum boninense (Li et al., 2014, 2017). Approximately, 406 strains of Colletotrichum spp. were isolated from oil tea in 10 provinces of China by Li. The results showed that Co. fructicola was the most widely distributed in oil tea, so the prevention and the control of Co. fructicola were one of the key points of oil tea anthracnose control (Li, 2018). Co. fructicola is widely distributed and has many hosts, such as oil tea, apple, strawberry, mango, banana, coffee, and other plants of more than 50 species, among which oil tea is one of its main hosts (Prihastuti et al., 2009; Weir et al., 2012; Huang et al., 2013; Li et al., 2013; Diao et al., 2017). Although anthracnose of oil tea has attracted more and more attention, there are few effective control methods. The reason is that the pathogenic mechanism of Colletotrichum spp. and the immune mechanism of the host are not well understood.
Control of anthracnose of oil tea
The control of oil tea anthracnose can be divided into prevention and treatment. Breeding and planting resistant plants are important measures to prevent plant disease (Savchenko, 2017). Ca.yuhsienensis Hu, a species of oil tea, was once widely cultivated in central China because of its high quality, yield, and high resistance to anthracnose (Denton-Giles et al., 2013; Denton-Giles, 2014; Cao et al., 2017; Nie et al., 2020). In contrast to Ca. oleifera, which has the largest planting area, Ca. yuhsienensis is not generally infected by Colletotrichum spp. Yang et al. (2004) and Duan et al. (2005) found that Ca. yuhsienensis, Camellia octopetala Hu, Ca. oleifera Abel var. Huizhou-xiaohong and Ca. oleifera Abel var. Huizhou-dahong have resistance to Colletotrichum spp. (Yang et al., 2004; Duan et al., 2005). Moreover, Ca. yuhsienensis also showed strong resistance to other pathogens, such as Ciborinia camelliae (Denton-Giles et al., 2013; Denton-Giles, 2014; Saracchi et al., 2019; Li et al., 2020). Consequently, Ca. yuhsienensis, as a wild relative of Ca. oleifera, is widely used to breed varieties of oil tea (Nie et al., 2020).
After selecting suitable oil tea varieties, the seedling quarantine should be strictly controlled. When selecting seedlings and other reproductive materials, quality inspection must be carried out in accordance with national and regional standards to ensure the safety of various reproductive materials (Shan et al., 2019). After planting oil tea, the cultivation management should be strengthened to create environmental conditions that are not conducive to the survival, reproduction, and transmission of pathogen and are suitable for the growth of oil tea (Shu and Zhang, 2009).
When oil tea was infected with Colletotrichum spp., the treatment of anthracnose is primarily based on the use of chemical pesticides. For oil tea forest in the early stage of the anthracnose, Bordeaux mixture can be used for treatment. For oil tea forest in the late stage of the anthracnose, chlorothalonil, carbendazim, and thiophanate methyl can be used for treatment (Yu, 2019).
However, the abuse of pesticides not only easily causes environmental pollution but also leads to the emergence of pathogens resistant to pesticides (Holtappels et al., 2021). Therefore, biological control has been a hot spot in the research of oil tea anthracnose in recent years. Bacillus subtilis Y13 was isolated from healthy leaves of Ca. oleifera by Zhou. The results showed that its inhibitory rate on Colletotrichum spp. was 88.5% (Zhou et al., 2008). Bacillus velezensis HBMC–B05 was isolated from healthy leaves of Ca. oleifera by Shang. The results showed that its inhibitory rate on Colletotrichum spp. was 88.5% (Shang et al., 2021). Yu. (2019) further found that lipopeptides, various metabolites of antagonistic bacteria, had antagonistic effect on Colletotrichum spp. (Yu, 2019). Moreover, many strains were isolated, such as Bacillus subtilis R6, Paenibcillus kribbens Z17, Brevibacillus brevis Z26, Streptomvces globisporus subsp Globisporus F10, and Streptomyces albulus cf17. It was found that these strains could inhibit the growth of Colletotrichum spp. (Song et al., 2012).
In addition to antagonizing microorganisms, some elicitors can also improve the resistance of oil tea to Colletotrichum spp., such as salicylic acid (SA), methyl jasmonate (MeJA), and so on (Wang et al., 2022). Although the research on the control of oil tea anthracnose has made good progress, the anthracnose still puzzles the oil tea industry. The main reason is that the pathogenic mechanism of oil tea anthracnose has not been fully proved, which is difficult to provide theoretical guidance for the prevention and control of oil tea anthracnose. Therefore, to understand the pathogenic mechanism of oil tea anthracnose, the molecular mechanism of the interaction between plants and pathogens needs to be understood.
Pathogenic mechanism of anthracnose
The plant immune system includes pathogen-associated molecular pattern-triggered immunity (PTI) and effector-triggered immunity (ETI) (Jones and Dangl, 2006). PTI is triggered by plasma membrane-localized pattern recognition receptors (PRRs) specific recognition of pathogen-associated molecular pattern (PAMP) or damage-associated molecular patterns (DAMPs) (Dangl et al., 2013; Lo Presti et al., 2015; Ranf, 2017). Pathogens secrete effectors to inhibit PTI to promote infection. The resistance gene (R gene) of the host that can be activated by effectors induces hypersensitive reaction (HR) or programmed cell death (PCD) to inhibit the growth of pathogens (Jones and Dangl, 2006). PTI and ETI jointly limit the invasion of pathogens, and pathogens secrete new effectors again to promote infection (Ngou et al., 2021; Yuan et al., 2021). This is the “zigzag” plant immune model proposed by Jones, which expounds the molecular mechanism of the interaction between pathogens and plants (Jones and Dangl, 2006).
These inducible defenses are also associated with wide-ranging transcriptional and hormonal reprogramming in plants (Pieterse et al., 2012; Chen and Ding, 2020). For instance, hormones also play an important role in plant immune regulation. Among them, SA and jasmonic acid (JA) are considered as the main defense hormones, while others such as gibberellin (GAs), ethylene (ET), abscisic acid (ABA), brassinosteroids (BRS), auxin [indole-3-acetic acid (IAA)], cytokinin (CK), and nitric oxide (NO) are also the regulators of plant immune signal network (Browse, 2009; Pieterse et al., 2012; Wang et al., 2020b; Yang et al., 2020b; Zheng et al., 2020). These pathways and signals together construct the plant immune system.
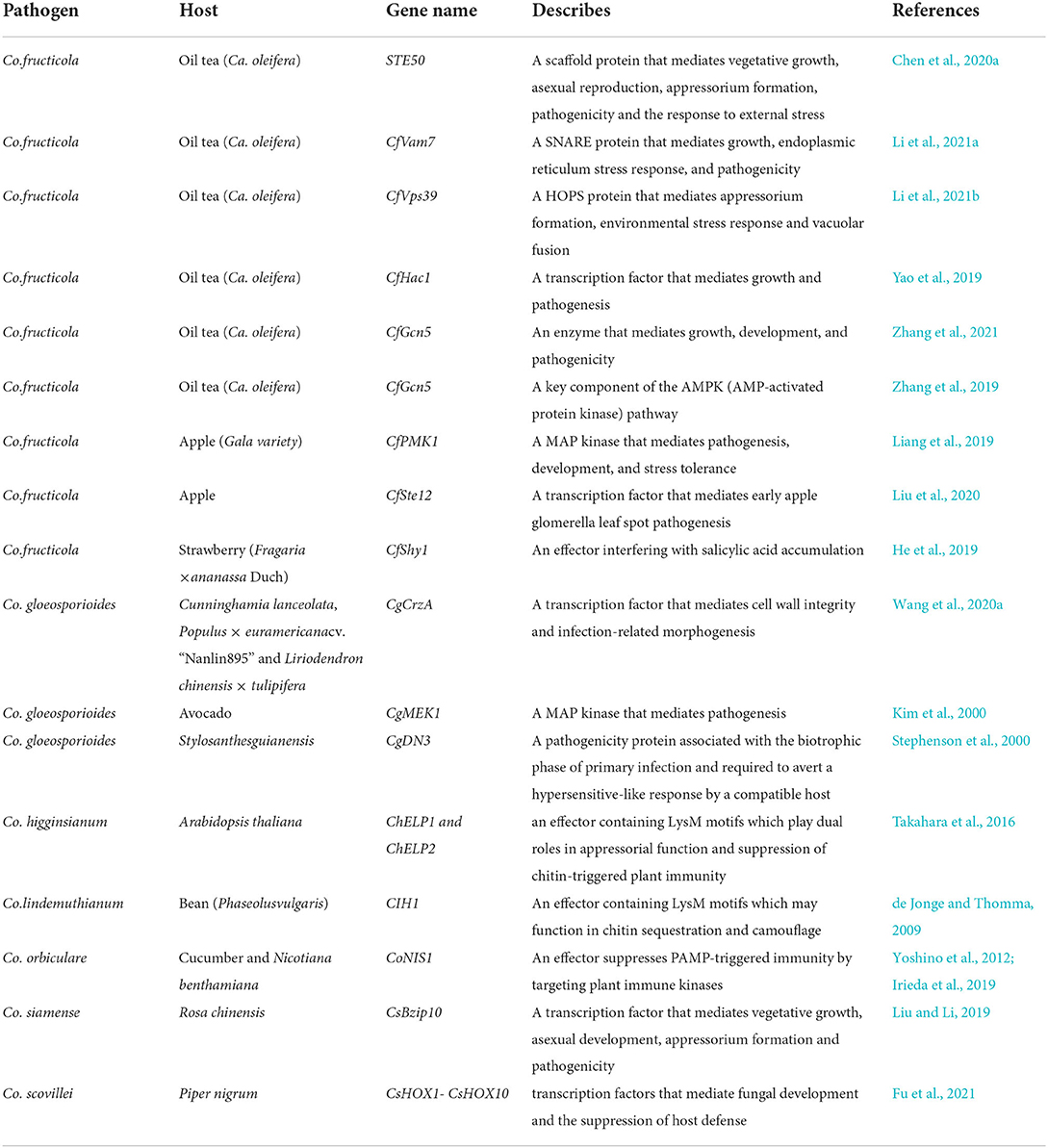
Colletotrichum spp. is widely distributed and extremely destructive, and can infect almost all plants. Therefore, the pathogenic mechanism of anthracnose has always been a research hotspot (Hyde et al., 2009; Cannon et al., 2012). Great progress has been made in the characterization of a single pathogenic gene (Table 1). As early as 1999, Geffroy et al. (1999) found that the R gene of Phaseolus vulgaris has different resistance to Colletotrichum lindemuthianum from different sources. This result further expounded the relationship between Colletotrichum spp. and plant immunity (Geffroy et al., 1999). Subsequently, new R genes were found one after another. These results deepened the understanding of “gene to gene” theory (Melotto and Kelly, 2001; López et al., 2003; Zou et al., 2018). With the deepening of research, great progress has been made in the exploration of Colletotrichum spp., such as the elucidation of protein phosphorylation and dephosphorylation, mitogen-activated protein kinase (MAPK), and calmodulin signal, which is helpful to understand the growth and pathogenesis of Colletotrichum spp. (Dickman and Yarden, 1999; Kim et al., 2000; Takano et al., 2000; Chen and Dickman, 2002, 2004; Ha et al., 2003; Liang et al., 2019; Zhang et al., 2019).
Transcription factors also play a very important role in the growth and pathogenesis of Colletotrichum spp. Wang et al. (2020a) found that calcineurin-resonsive transcription factor CgCrzA was not only involved in regulating cell wall integrity but also in morphogenesis and virulence in Co. gloeosporioides, which proved the importance of the calmodulin signal to Colletotrichum spp. (Wang et al., 2020a). Liu found that the transcription factor CsBzip10 controls vegetative growth, asexual development, appressorium formation, and pathogenicity in Co. siamense (Liu and Li, 2019). Fu also found 10 transcription factors that affect growth and inhibit host immunity from Colletotrichum scovillei (Fu et al., 2021). Moreover, many important transcription factors have been found in Co. fructicola. Yao found that transcription factor CfHac1 plays critical roles in growth, conidiation, appressorium formation, and pathogenicity, and respond to osmotic stress in Co. fructicola (Yao et al., 2019). Li found that transcription factor CfVam7 is required for growth, endoplasmic reticulum stress response, and pathogenicity of Co. fructicola (Li et al., 2021a). In addition to the transcription factors introduced above, the functions of transcription factors, such as CfSte12 and CfSte50 of Co. fructicola, have also been explored, which lays a foundation for understanding the pathogenesis of anthracnose (Chen et al., 2020a; Liu et al., 2020).
Colletotrichum spp. can also secrete effectors to promote infection. As early as 1994, the effector CIH1, an effector containing tandem chitin-binding lysin motifs (LysM), which may function in chitin sequestration and camouflage, was found in Colletotrichum lindemuthianum (Pain et al., 1994; Perfect et al., 1998; de Jonge and Thomma, 2009; Stergiopoulos and Wit, 2009). Subsequently, de Queiroz systematically predicted and obtained several effectors of Co. lindemuthianum (de Queiroz et al., 2019). Takahara et al. (2009, 2016) also found effectors with LysM from Colletotrichum higginsianum; results suggested a dual role for these LysM proteins as effectors for suppressing chitin-triggered immunity and as proteins required for appressorium function (Takahara et al., 2009, 2016). Subsequently, Kleemann et al. (2008, 2012) also identified multiple effectors from Co. higginsianum, and the results showed that most effectors are host induced and expressed in consecutive waves associated with pathogenic transitions, indicating distinct effector suites are deployed at each stage (Kleemann et al., 2008, 2012). Yoshino et al. (2012) and Irieda et al. (2019) found that the effector NIS1 of Colletotrichum orbiculare can interact with PRRs of plants to inhibit immunity (Yoshino et al., 2012; Irieda et al., 2019). Further studies found that the CgDN3 gene, which can inhibit the function of NIS1, is an important gene to maintain the pathogenicity of Co. gloeosporoides (Stephenson et al., 2000). Eisermann et al. (2019) also found two important effectors from Colletotrichum graminicola (Eisermann et al., 2019). Schmidt et al. (2020) Andree found that reactive oxygen species (ROS) can increase the resistance of Arabidopsis to Co. higginsianum; the results further proved the important role of ROS in plant immunity (Schmidt et al., 2020). On the contrary, the CfShy1 and CfGcn5 effectors of Co. fruticola will affect the homeostasis of host SA and inhibit plant immunity (He et al., 2019; Zhang et al., 2021).
In recent years, in addition to exploring the function of a single gene of Colletotrichum spp., some scholars have also used omics methods to explore Colletotrichum spp. RNA-seq has been used to study Colletotrichum–host interactions. Previous studies have explored the transcriptional profile of Colletotrichum spp. after being infected by the host; the results showed that small secreted proteins (SSPs), cytochrome P450s, carbohydrate-active enzymes (CAZYs), and secondary metabolite (SM) synthetases were enriched (Liang et al., 2018). There are also studies to explore the transcriptional profile of a host after Colletotrichum spp. infection revealed that many genes were mainly related to immune response, plant hormone signal transduction, and secondary metabolites (Fang et al., 2021; Mehmood et al., 2021). Some studies have discussed the simultaneous response between Colletotrichum spp. and hosts, which provides a new perspective for understanding the pathogenesis of anthracnose and the immune mechanism of the hosts (Alkan et al., 2015; Zhang et al., 2018).
Conclusions and future perspective
Colletotrichum spp.is one of the important pathogenic fungi with many species, hosts, and wide distribution. In recent years, the research on the control and pathogenic mechanism of Colletotrichum spp. has made good progress. In the future, the research on the control and pathogenic molecular mechanism of Colletotrichum spp. is still the focus. However, there is still a lack of safe and effective drugs or biological reagents to control oil tea anthracnose. Although the understanding of Colletotrichum spp. has made significant progress, the current research on molecular mechanism of anthracnose is obviously insufficient. There are only a few species mentioned above, such as Co. gloeosporoides, Co. higginsianum, and Co. fruticola. However, there are more other pathogenic molecular mechanisms of Colletotrichum spp. that have not been explored. Secondly, the research on the pathogenic molecular mechanism of anthracnose should gradually develop from the functional research of a single gene to the analysis of signal network, regulation mechanism, and omics. More importantly, there are few reports on the interaction mechanism between Colletotrichum spp. and hosts, especially the mechanism of oil tea responding to Colletotrichum spp. Exploring the pathogenic mechanism of anthracnose in oil tea is expected to provide reference for the green prevention and control of anthracnose in oil tea, and also helps people better understand the molecular mechanism of plant pathogen growth, development, and pathogenesis. At present, the prevention and control technology of oil tea anthracnose is still based on traditional agricultural control and chemical control. With the continuous innovation of technology and the demand for safe and efficient control technology, the development of biological control and seed selection of disease resistant varieties will be rapidly promoted.
Author contributions
XinggC, GZ, and JL contributed to the conception of the manuscript. XinggC, XingzC, XM, and QT contributed significantly to manuscript preparation. All the authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the National Natural Science Foundation of China (Grant No. 31971661); Postgraduate Scientific Research Innovation Project of Hunan Province (Grant No. CX20200712); Scientific Innovation Fund for Post-graduates of Central South University of Forestry and Technology (Grant No. CX20201008).
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher's note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
Alkan, N., Friedlander, G., Ment, D., Prusky, D., and Fluhr, R. (2015). Simultaneous transcriptome analysis of Colletotrichum gloeosporioides and tomato fruit pathosystem reveals novel fungal pathogenicity and fruit defense strategies. New Phytol. 205, 801–815. doi: 10.1111/nph.13087
Browse, J. (2009). Jasmonate passes muster: a receptor and targets for the defense hormone. Annu. Rev. Plant Biol. 60, 183–205. doi: 10.1146/annurev.arplant.043008.092007
Cannon, P. F., Damm, U., Johnston, P. R., and Weir, B. S. (2012). Colletotrichum – current status and future directions. Stud. Mycol. 73, 181–213. doi: 10.3114/sim0014
Cao, Y. Q., Yao, X. H., Ren, H. D., and Wang, K. L. (2017). Determination of fatty acid composition and metallic element content of four Camellia species used for edible oil extraction in China. J. Consum. Prot. Food Saf. 12, 165–169. doi: 10.1007/s00003-017-1104-2
Chen, C. B., and Dickman, M. B. (2002). Colletotrichum trifolii TB3 kinase, a COT1 homolog, is light inducible and becomes localized in the nucleus during hyphal elongation. Eukaryot. Cell 1, 626–633. doi: 10.1128/EC.1.4.626-633.2002
Chen, C. B., and Dickman, M. B. (2004). Dominant active Rac and dominant negative Rac revert the dominant active Ras phenotype in Colletotrichum trifolii by distinct signalling pathways. Mol. Microbiol. 51, 1493–1507. doi: 10.1111/j.1365-2958.2003.03932.x
Chen, X. G., Jiang, L. Y., Bao, A. H., Liu, C. L., Liu, J. A., and Zhou, G. Y. (2021a). Molecular characterization, pathogenicity and biological characterization of Colletotrichum species associated with anthracnose of Camellia yuhsienensis Hu in China. Forests 12:1712. doi: 10.3390/f12121712
Chen, X. G., Liu, C. L., Liu, J. A., and Zhou, G. Y. (2021b). First report of Colletotrichum fructicola causing anthracnose on Camellia yuhsienensis Hu in China. Plant Dis. 106, 321–321. doi: 10.1094/PDIS-04-21-0772-PDN
Chen, Y. Y., Liu, J. A., Jiang, S. Q., Li, H., and Zhou, G. Y. (2020a). Colletotrichum fructicolaSTE50 is required for vegetative growth, asexual reproduction, appressorium formation, pathogenicity and the response to external stress. J. Plant Pathol. 102, 335–342. doi: 10.1007/s42161-019-00422-3
Chen, Y. Z. (2008). Oil Tea Camellia Superior Germplasm Resources. Beijing: China Forestry Publishing House.
Chen, Y. Z., Deng, S. H., Chen, L. S., Ma, L., He, H., Wang, X. N., et al. (2020b). A new view on the development of oil tea camellia industry. J. Nanjing For. Univ. 44, 1–10. doi: 10.3969/j.issn.1000-2006.201909033
Chen, Z. Q., and Ding, S. W. (2020). JAcked responses Go Viral: hormonal regulation of antiviral RNAi. Cell Host Microbe 28, 7–9. doi: 10.1016/j.chom.2020.06.007
Dangl, J. L., Horvath, D. M., and Staskawicz, B. J. (2013). Pivoting the plant immune system from dissection to deployment. Science 341, 746–751. doi: 10.1126/science.1236011
de Jonge, R., and Thomma, B. P. H. J. (2009). Fungal LysM effectors: extinguishers of host immunity? Trends Microbiol. 17, 151–157. doi: 10.1016/j.tim.2009.01.002
de Queiroz, C. B., Correia, H. L. N., Santana, M. F., Batista, D. S., Vidigal, P. M. P., Brommonschenkel, S. H., et al. (2019). The repertoire of effector candidates in Colletotrichum lindemuthianum reveals important information about Colletotrichum genus lifestyle. Appl. Microbiol. Biotechnol. 103, 2295–2309. doi: 10.1007/s00253-019-09639-9
Dean, R., Van Kan, J. A., Pretorius, Z. A., Hammond-Kosack, K. E., Di Pietro, A., Spanu, P. D., et al. (2012). The Top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 13, 414–430. doi: 10.1111/j.1364-3703.2011.00783.x
Deng, Z. X. (2011). Present situation and countermeasures to control Camellia Oleifera pest and disease. Trop. Forest. 39, 42–46.
Denton-Giles, M. (2014). Characterization of Incompatible and Compatible Camellia-Ciborinia Camelliae Plant-Pathogen Interactions. [Dissertation]. [Palmerston North, New Zealand]: Massey University.
Denton-Giles, M., Bradshaw, R. E., and Dijkwel, P. P. (2013). Ciborinia camelliae (Sclerotiniaceae) induces variable plant resistance responses in selected species of Camellia. Phytopathology 103, 725–732. doi: 10.1094/PHYTO-11-12-0289-R
Diao, Y. Z., Zhang, C., Liu, F., Wang, W. Z., Liu, L., Cai, L., et al. (2017). Colletotrichum species causing anthracnose disease of chili in China. Persoonia 38, 20–37. doi: 10.3767/003158517X692788
Dickman, M. B., and Yarden, O. (1999). Serine/threonine protein kinases and phosphatases in filamentious fungi. Fungal Genet. Biol. 26, 99–117. doi: 10.1006/fgbi.1999.1118
Duan, L., Yang, G. D., Shu, Q. L., and Zheng, H. B. (2005). Relationship of peel color with resistance to anthracnose in Oiltea Camellia. Nonwood Forest Res. 23, 9–12. doi: 10.14067/j.cnki.1003-8981.2005.02.003
Eisermann, I., Weihmann, F., Krijger, J. J., Kroling, C., Hause, G., Menzel, M., et al. (2019). Two genes in a pathogenicity gene cluster encoding secreted proteins are required for appressorial penetration and infection of the maize anthracnose fungus Colletotrichum graminicola. Environ. Microbiol. 21, 4773–4791. doi: 10.1111/1462-2920.14819
Fang, H. C., Liu, X., Dong, Y. H., Feng, S., Zhou, R., Wang, C. X., et al. (2021). Transcriptome and proteome analysis of walnut (Juglans regia L.) fruit in response to infection by Colletotrichum gloeosporioides. BMC Plant Biol. 21:249. doi: 10.1186/s12870-021-03042-1
Fu, T., Han, J.-H., Shin, J.-H., Song, H., Ko, J., Lee, Y.-H., et al. (2021). Homeobox transcription factors are required for fungal development and the suppression of host defense mechanisms in the Colletotrichum scovillei-pepper pathosystem. mBio 12, e01620–01621. doi: 10.1128/mBio.01620-21
Geffroy, V., Sicard, D., de Oliveira, J. C. F., Sévignac, M., Cohen, S., Gepts, P., et al. (1999). Identification of an ancestral resistance gene cluster Involved in the coevolution process between Phaseolus vulgaris and its fungal pathogen Colletotrichum lindemuthianum. Mol. Plant-Microbe Interact. 12, 774–784. doi: 10.1094/MPMI.1999.12.9.774
Ha, Y. S., Memmott, S. D., and Dickman, M. B. (2003). Functional analysis of Ras in Colletotrichum trifolii. FEMS Microbiol. Lett. 226, 315–321. doi: 10.1016/S0378-1097(03)00589-5
He, C. Y., Duan, K., Zhang, L. Q., Zhang, L., Song, L. L., Yang, J., et al. (2019). Fast quenching the burst of host salicylic acid Is common in early strawberry/Colletotrichum fructicola interaction. Phytopathology 109, 531–541. doi: 10.1094/PHYTO-02-18-0043-R
Holtappels, D., Fortuna, K., Lavigne, R., and Wagemans, J. (2021). The future of phage biocontrol in integrated plant protection for sustainable crop production. Curr. Opin. Biotechnol. 68, 60–71. doi: 10.1016/j.copbio.2020.08.016
Huang, F., Chen, G. Q., Hou, X., Fu, Y. S., Cai, L., Hyde, K. D., et al. (2013). Colletotrichum species associated with cultivated citrus in China. Fungal Divers. 61, 61–74. doi: 10.1007/s13225-013-0232-y
Hyde, K. D., Cai, L., Cannon, P. F., Crouch, J. A., Crous, P. W., Damm, U., et al. (2009). Colletotrichum - names in current use. Fungal Divers. 39, 147–182.
Irieda, H., Inoue, Y., Mori, M., Yamada, K., Oshikawa, Y., Saitoh, H., et al. (2019). Conserved fungal effector suppresses PAMP-triggered immunity by targeting plant immune kinases. Proc. Natl. Acad. Sci. U.S.A. 116, 496–505. doi: 10.1073/pnas.1807297116
Jin, A. X., Zhou, G. Y., and Li, H. (2009). Progress, problem and prospect of oil camelliae anthracnose (Colletotrichum gloeosporioides) research. Forest Pest Dis. 28, 27–31.
Jones, J. D. G., and Dangl, J. L. (2006). The plant immune system. Nature 444, 323–329. doi: 10.1038/nature05286
Kim, Y.-K., Kawano, T., Li, D., and Kolattukudy, P. E. (2000). A mitogen-activated protein kinase kinase required for induction of cytokinesis and appressorium formation by host signals in the conidia of Colletotrichum gloeosporioides. Plant Cell 12, 1331–1343. doi: 10.1105/tpc.12.8.1331
Kleemann, J., Rincon-Rivera, L. J., Takahara, H., Neumann, U., van Themaat, E. V. L., van der Does, H. C., et al. (2012). Sequential delivery of host-induced virulence effectors by appressoria and intracellular hyphae of the phytopathogen Colletotrichum higginsianum. PLOS Pathog. 8:e1002643. doi: 10.1371/annotation/0f398a0c-dfda-4277-b172-4ff9cb31aec3
Kleemann, J., Takahara, H., Stüber, K., apos, and Connell, R. (2008). Identification of soluble secreted proteins from appressoria of Colletotrichum higginsianum by analysis of expressed sequence tags. Microbiology 154, 1204–1217. doi: 10.1099/mic.0.2007/014944-0
Li, H. (2018). Population Genetic Analyses of the Fungal Pathogen Colletotrichum on Tea-Oil Trees in China and Characterization of a MAPK gene CfPMK1 in the Pathogen. [Dissertation]. [Changsha, China]: Central South University of Forestry and Technology.
Li, H., Li, Y., Jiang, S. Q., Liu, J. A., and Zhou, G. Y. (2017). Pathogen of oil-tea trees anthracnose caused by Colletotrichum spp. in Hunan Province. Scientia Silvae Sinicae 53, 43–53. doi: 10.11707/j.1001-7488.20170806
Li, H., Zhou, G. Y., Xu, J. P., and Zhu, D. X. (2014). Pathogen identification of a new anthracnose of Camellia oleifera in China based on multiple-gene phylogeny. Acta Phytophylacica Sin 41, 602–607. doi: 10.15889/j.issn.1002-1302.2019.20.016
Li, H. N., Jiang, J. J., Hong, N., Wang, G. P., and Xu, W. X. (2013). First Report of Colletotrichum fructicola causing bitter rot of pear (Pyrus bretschneideri) in China. Plant Dis. 97, 1000–1000. doi: 10.1094/PDIS-01-13-0084-PDN
Li, J., Luo, Z. Q., Zhang, C. H., Qu, X. J., Chen, M., Song, T., et al. (2020). Seasonal variation in the rhizosphere and non-rhizosphere microbial community structures and functions of Camellia yuhsienensis Hu. Microorganisms 8:1385. doi: 10.3390/microorganisms8091385
Li, S. Z., Zhang, S. P., Li, B., and Li, H. (2021a). The SNARE protein CfVam7 is required for growth, endoplasmic reticulum stress response, and pathogenicity of Colletotrichum fructicola. Front. Microbiol. 12, 736066. doi: 10.3389/fmicb.2021.736066
Li, S. Z., Zhang, S. P., and Li, H. (2021b). A HOPS protein, CfVps39, is required for appressorium formation, environmental stress response and vacuolar fusion of Colletotrichum fructicola. For. Pathol. 51:e12692. doi: 10.1111/efp.12692
Liang, X. F., Shang, S. P., Dong, Q. Y., Wang, B., Zhang, R., Gleason, M. L., et al. (2018). Transcriptomic analysis reveals candidate genes regulating development and host interactions of Colletotrichum fructicola. BMC Genomics 19:21. doi: 10.1186/s12864-018-4934-0
Liang, X. F., Wei, T. Y., Cao, M. Y., Zhang, X., Liu, W. K., Kong, Y. Y., et al. (2019). The MAP kinase CfPMK1 is a key regulator of pathogenesis, development, and stress tolerance of Colletotrichum fructicola. Front. Microbiol. 10, 1070. doi: 10.3389/fmicb.2019.01070
Liu, J. A., He, L., and Zhou, G. Y. (2009). Specific and rapid detection of Camellia oleifera anthracnose pathogen by nested-PCR. Afr. J. Biotechnol. 8, 1056-1061.
Liu, R. F., and Li, H. (2019). The transcription factor CsBzip10 controls vegetative growth, asexual development, appressorium formation and pathogenicity in the Rosa chinensis anthracnose fungus Colletotrichum siamense. Austr. Plant Pathol. 48, 595–601. doi: 10.1007/s13313-019-00663-x
Liu, W., Liang, X., Gleason, M. L., Cao, M., Zhang, R., Sun, G., et al. (2020). Transcription factor CfSte12 of Colletotrichum fructicola is a key regulator of early apple Glomerella leaf spot pathogenesis. Appl. Environ. Microbiol. 87, e02212–02220. doi: 10.1128/AEM.02212-20
Lo Presti, L., Lanver, D., Schweizer, G., Tanaka, S., Liang, L., Tollot, M., et al. (2015). Fungal effectors and plant susceptibility. Annu. Rev. Plant Biol. 66, 513–545. doi: 10.1146/annurev-arplant-043014-114623
López, C. E., Acosta, I. F., Jara, C., Pedraza, F., Gaitán-Solís, E., Gallego, G., et al. (2003). Identifying resistance gene analogs associated with resistances to different pathogens in common bean. Phytopathology 93, 88–95. doi: 10.1094/PHYTO.2003.93.1.88
Mehmood, N., Yuan, Y., Ali, M., Ali, M., Iftikhar, J., Cheng, C., et al. (2021). Early transcriptional response of terpenoid metabolism to Colletotrichum gloeosporioides in a resistant wild strawberry Fragaria nilgerrensis. Phytochemistry 181:112590. doi: 10.1016/j.phytochem.2020.112590
Melotto, M., and Kelly, J. D. (2001). Fine mapping of the Co-4 locus of common bean reveals a resistance gene candidate, COK-4, that encodes for a protein kinase. Theor. Appl. Genet. 103, 508–517. doi: 10.1007/s001220100609
Münch, S., Lingner, U., Floss, D. S., Ludwig, N., Sauer, N., and Deising, H. B. (2008). The hemibiotrophic lifestyle of Colletotrichum species. J. Plant Physiol. 165, 41–51. doi: 10.1016/j.jplph.2007.06.008
Ngou, B. P. M., Ahn, H.-K., Ding, P., and Jones, J. D. G. (2021). Mutual potentiation of plant immunity by cell-surface and intracellular receptors. Nature 592, 110–115. doi: 10.1038/s41586-021-03315-7
Nie, Z. Y., Huang, X. Z., Hu, Z. K., Li, X. L., Yin, H. F., and Li, J. Y. (2020). Characterization of the complete chloroplast genome of Camellia yuhsienensis Hu, a resilient shrub with strong floral fragrance. Mitochondrial. DNA. B. 5, 3016–3017. doi: 10.1080/23802359.2020.1797580
O'Connell, R. J., Thon, M. R., Hacquard, S., Amyotte, S. G., Kleemann, J., Torres, M. F., et al. (2012). Lifestyle transitions in plant pathogenic Colletotrichum fungi deciphered by genome and transcriptome analyses. Nat. Genet. 44, 1060–1065. doi: 10.1038/ng.2372
Pain, N. A., O' Connell, R. J., and Mendgen, K. G., Jonathan, R. (1994). Identification of glycoproteins specific to biotrophic intracellular hyphae formed in the Colletotrichum lindemuthianum-bean interaction. New Phytol. 127, 233–242. doi: 10.1111/j.1469-8137.1994.tb04275.x
Perfect, S. E., O'Connell, R. J., Green, E. F., Doering-Saad, C., and Green, J. R. (1998). Expression cloning of a fungal proline-rich glycoprotein specific to the biotrophic interface formed in the Colletotrichum–bean interaction. Plant J. 15, 273–279. doi: 10.1046/j.1365-313X.1998.00196.x
Pieterse, C. M. J., Does, D. V., Zamioudis, C., Leon-Reyes, A., and Wees, S. C. M. V. (2012). Hormonal modulation of plant immunity. Annu. Rev. Cell Dev. Biol. 28, 489–521. doi: 10.1146/annurev-cellbio-092910-154055
Prihastuti, H., Cai, L., Chen, H., McKenzie, E. H. C., and Hyde, K. D. (2009). Characterization of Colletotrichum species associated with coffee berries in northern Thailand. Fungal Divers. 39, 89–109.
Qin, S. Y., Rong, J., Zhang, W. J., and Chen, J. K. (2018). Cultivation history of Camellia oleifera and genetic resources in the Yangtze River Basin. Biodivers. Sci. 26, 384–395. doi: 10.17520/biods.2017254
Ranf, S. (2017). Sensing of molecular patterns through cell surface immune receptors. Curr. Opin. Plant Biol. 38, 68–77. doi: 10.1016/j.pbi.2017.04.011
Saracchi, M., Locati, D., Colombo, E. M., and Pasquali, M. (2019). Updates on Ciborinia camelliae, the causal agent of camellia flower blight. J. Plant Pathol. 101, 215–223. doi: 10.1007/s42161-018-0173-0
Savchenko, I. V. (2017). Breeding new varieties and hybrids of agricultural plants. Her. Russ. Acad. Sci. 87, 104–110. doi: 10.1134/S1019331617020150
Schmidt, A., Mächtel, R., Ammon, A., Engelsdorf, T., Schmitz, J., Maurino, V. G., et al. (2020). Reactive oxygen species dosage in Arabidopsis chloroplasts can improve resistance towards Colletotrichum higginsianum by the induction of WRKY33. New Phytol. 226, 189–204. doi: 10.1111/nph.16332
Shan, T. J., Zhang, Y., Xie, Y. Y., Shi, H. Y., Huang, Y. F., and Wu, H. X. (2019). The latest progress of oil tea disease and its control. Agri. Sci. Jiangsu Province 47, 75–80.
Shang, X. N., Liu, J. A., Feng, F. S., and Zhou, G. Y. (2021). Screening, identification and antagonistic effect of endophytic antagonistic bacterial strains from Camellia oleifera. Chin. J. Biol. Control 37, 275–583. doi: 10.16409/j.cnki.2095-039x.2021.04.002
Shi, T., Wu, G. C., Jin, Q. Z., and Wang, X. G. (2020). Camellia oil authentication: A comparative analysis and recent analytical techniques developed for its assessment. A review. Trends Food Sci. Technol. 97, 88–99. doi: 10.1016/j.tifs.2020.01.005
Shu, Q. L., and Zhang, L. F. (2009). Oil Cultivation and Disease Control in China. Beijing: China Forestry Publishing House.
Song, G. T., Zhou, G. Y., Yang, L., and Liu, J. A. (2012). Optimization of fermentation conditions for antagonistic actinomyces CF17. J. Fujian College Forest. 32, 355–359. doi: 10.13324/j.cnki.jfcf.2012.04.001
Stephenson, S.-A., Hatfield, J., Rusu, A. G., Maclean, D. J., and Manners, J. M. (2000). CgDN3: An essential pathogenicity gene of Colletotrichum gloeosporioides necessary to avert a hypersensitive-like response in the host Stylosanthes guianensis. Mol. Plant. Microbe Interact. 13, 929–941. doi: 10.1094/MPMI.2000.13.9.929
Stergiopoulos, I., and Wit, P. J. G. M. (2009). Fungal effector proteins. Annu. Rev. Phytopathol. 47, 233–263. doi: 10.1146/annurev.phyto.112408.132637
Sturm, J. (1832). Deutschlands Flora in Abbildungen nach der Natur mit Beschreibungen. I Abt., Heft 11. Nürnberg.
Sutton, B. C. (1992). The genus Glomerella and its anamorph Colletotrichum. Colletotrichum Biol. Pathol. Control 388, 1–26.
Takahara, H., Dolf, A., Endl, E., and O'Connell, R. (2009). Flow cytometric purification of Colletotrichum higginsianum biotrophic hyphae from Arabidopsis leaves for stage-specific transcriptome analysis. Plant J. 59, 672–683. doi: 10.1111/j.1365-313X.2009.03896.x
Takahara, H., Hacquard, S., Kombrink, A., Hughes, H. B., Halder, V., Robin, G. P., et al. (2016). Colletotrichum higginsianum extracellular LysM proteins play dual roles in appressorial function and suppression of chitin-triggered plant immunity. New Phytol. 211, 1323–1337. doi: 10.1111/nph.13994
Takano, Y., Kikuchi, T., Kubo, Y., Hamer, J. E., Mise, K., and Furusawa, I. (2000). The Colletotrichum lagenarium MAP kinase gene CMK1 regulates diverse aspects of fungal pathogenesis. Mol. Plant-Microbe Interact. 13, 374–383. doi: 10.1094/MPMI.2000.13.4.374
Talhinhas, P., Sreenivasaprasad, S., Neves-Martins, J., and Oliveira, H. (2002). Genetic and morphological characterization of Colletotrichum acutatum causing anthracnose of lupins. Phytopathology 92, 986–996. doi: 10.1094/PHYTO.2002.92.9.986
Wang, C. L., Zuo, L. Y., Zhang, S. Y., and Chen, A. L. (2022). Activities of 7 fungicides and 5 plant disease elicitors on the pathogen of Camellia oleifera. J. Northwest For. Univ. 37, 176–179. doi: 10.3969/j.issn.1001-7461.2022.02.24
Wang, P., Li, B., Pan, Y. T., Zhang, Y. Z., Li, D. W., and Huang, L. (2020a). Calcineurin-responsive transcription factor CgCrzA Is required for cell wall integrity and infection-related morphogenesis in Colletotrichum gloeosporioides. Plant Pathol. J. 36, 385–397. doi: 10.5423/PPJ.OA.04.2020.0071
Wang, W., Withers, J., Li, H., Zwack, P. J., Rusnac, D.-V., Shi, H., et al. (2020b). Structural basis of salicylic acid perception by Arabidopsis NPR proteins. Nature 586, 311–316. doi: 10.1038/s41586-020-2596-y
Weir, B. S., Johnston, P. R., and Damm, U. (2012). The Colletotrichum gloeosporioides species complex. Stud. Mycol. 73, 115–180. doi: 10.3114/sim0011
Yang, C. Y., Liu, X. M., Chen, Z. Y., Lin, Y. S., and Wang, S. Y. (2016). Comparison of oil content and fatty acid profile of ten new Camellia oleifera cultivars. J. Lipids. 2016:3982486. doi: 10.1155/2016/3982486
Yang, G. D. (2009). Resistance Mechanism of Camellia oleifera Cultivars to Colletotrichum gloeosporioides. [Dissertation]. [Hefei, China]: Anhui Agricultural University.
Yang, G. D., Shu, Q. L., Duan, L., Chen, C. Y., and Zheng, H. B. (2004). Resistance of main cultivars of oil tea to Colletotrichum gloeosporioides. J. Anhui Agric. Univ. 31, 480–483. doi: 10.13610/j.cnki.1672-352x.2004.04.022
Yang, S. W., Liang, K. H., Wang, A. B., Zhang, M., Qiu, J. M., and Zhang, L. Y. (2020a). Physiological characterization and transcriptome analysis of Camellia oleifera Abel. during leaf senescence. Forests 11:812. doi: 10.3390/f11080812
Yang, Z. R., Huang, Y., Yang, J., 1., Yao, S., z., et al. (2020b). Jasmonate signaling enhances RNA silencing and antiviral defense in rice. Cell Host Microbe 28, 89–103.e108. doi: 10.1016/j.chom.2020.05.001
Yao, Q., Guo, Y., Wei, F. Y., Li, S. Z., Zhang, S. P., and Li, H. (2019). A bZIP-type transcription factor CfHac1 is involved in regulating development and pathogenesis in Colletotrichum fructicola. Mycosystema 38, 1643–1652. doi: 10.13346/j.mycosystema.190228
Yoshino, K., Irieda, H., Sugimoto, F., Yoshioka, H., Okuno, T., and Takano, Y. (2012). Cell death of Nicotiana benthamiana is induced by secreted protein NIS1 of Colletotrichum orbiculare and is suppressed by a homologue of CgDN3. Mol. Plant. Microbe Interact. 25, 625–636. doi: 10.1094/MPMI-12-11-0316
Yu, J. X. (2019). Approach to Biological Control of Oil-Tea Camellia anthracnose. [Dissertation]. Changsha, China: Hunan Agricultural University.
Yuan, M. H., Jiang, Z. Y., Bi, G. Z., Nomura, K., Liu, M. H., Wang, Y. P., et al. (2021). Pattern-recognition receptors are required for NLR-mediated plant immunity. Nature 592, 105–109. doi: 10.1038/s41586-021-03316-6
Zhang, L., Huang, X., He, C., Zhang, Q. Y., Zou, X., Duan, K., et al. (2018). Novel fungal pathogenicity and leaf fefense strategies are revealed by simultaneous transcriptome analysis of Colletotrichum fructicola and strawberry infected by this fungus. Front. Plant Sci. 9, 434. doi: 10.3389/fpls.2018.00434
Zhang, S. P., Guo, Y., Chen, S. Q., and Li, H. (2021). The histone scetyltransferase CfGcn5 regulates growth, development, and pathogenicity in the anthracnose fungus Colletotrichum fructicola on the tea-oil tree. Front. Microbiol. 12, e680415. doi: 10.3389/fmicb.2021.680415
Zhang, S. P., Guo, Y., Li, S. Z., Zhou, G. Y., Liu, J. A., Xu, J. P., et al. (2019). Functional analysis of CfSnf1 in the development and pathogenicity of anthracnose fungus Colletotrichum fructicola on tea-oil tree. BMC Genetics 20:94. doi: 10.1186/s12863-019-0796-y
Zheng, H. Y., Dong, L. L., Han, X. Y., Jin, H. B., Yin, C. C., Han, Y. L., et al. (2020). The TuMYB46L-TuACO3 module regulates ethylene biosynthesis in einkorn wheat defense to powdery mildew. New Phytol. 225, 2526–2541. doi: 10.1111/nph.16305
Zhou, G. Y., Lu, L. L., Liu, J. A., and Li, H. (2008). Screening of beneficial endophytic bacteria to control Colletotrichum gloeosporioides and effects of control diseases. J. Hunan Agri. Univ. 34, 698–700. doi: 10.13331/j.cnki.jhau.2008.06.024
Zhu, Y. Z., Liao, W. J., Zou, D. X., Wu, Y. J., and Deng, Y. (2015). Identification and biological characteristics of the pathogen from Camellia oleifera anthracnose in Guangxi. J. Plant Prot. 42, 382–389. doi: 10.13802/j.cnki.zwbhxb.2015.03.015
Keywords: anthracnose, oil tea, Colletotrichum spp., Camellia oleifera, pathogenic molecular mechanism
Citation: Chen X, Chen X, Tan Q, Mo X, Liu J and Zhou G (2022) Recent progress on harm, pathogen classification, control and pathogenic molecular mechanism of anthracnose of oil-tea. Front. Microbiol. 13:918339. doi: 10.3389/fmicb.2022.918339
Received: 12 April 2022; Accepted: 30 June 2022;
Published: 29 July 2022.
Edited by:
Dong-Qin Dai, Qujing Normal University, ChinaReviewed by:
Artemio Mendoza-Mendoza, Lincoln University, New ZealandChen Zhuo, Guizhou University, China
Copyright © 2022 Chen, Chen, Tan, Mo, Liu and Zhou. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Junang Liu, a2pjOTYyMEAxNjMuY29t; Guoying Zhou, emd5aW5ncXFAMTYzLmNvbQ==
 Xinggang Chen
Xinggang Chen Xingzhou Chen
Xingzhou Chen Qian Tan
Qian Tan