- 1Center for Gene Diagnosis, Zhongnan Hospital of Wuhan University, Wuhan, China
- 2Department of Translational Molecular Pathology, The University of Texas MD Anderson Cancer Center, Houston, TX, USA
MicroRNAs (miRNAs) directly and indirectly affect tumorigenesis. To be able to perform their myriad roles, miRNA machinery genes, such as Drosha, DGCR8, Dicer1, XPO5, TRBP, and AGO2, must generate precise miRNAs. These genes have specific expression patterns, protein-binding partners, and biochemical capabilities in different cancers. Our preliminary analysis of data from The Cancer Genome Atlas consortium on multiple types of cancer revealed significant alterations in these miRNA machinery genes. Here, we review their biological structures and functions with an eye toward understanding how they could serve as cancer biomarkers.
Introduction
MicroRNAs (miRNAs) are non-coding RNAs ~22 nt long that bind to target mRNAs, resulting in mRNA degradation or inhibition of mRNA expression (1, 2), and play a key role in post-transcriptional gene regulation in up to 30–60% of all human genes (3). miRNA targets mRNA by specific base-pairing interactions between the seed region of the miRNA and the 5′-untranslated regions of the mRNA (4–6). miRNAs can be grouped into families on the basis of their seed sequences, and members of one family usually effect the same mRNAs. A small number of miRNAs outside the seed sequences have also been reported (7, 8).
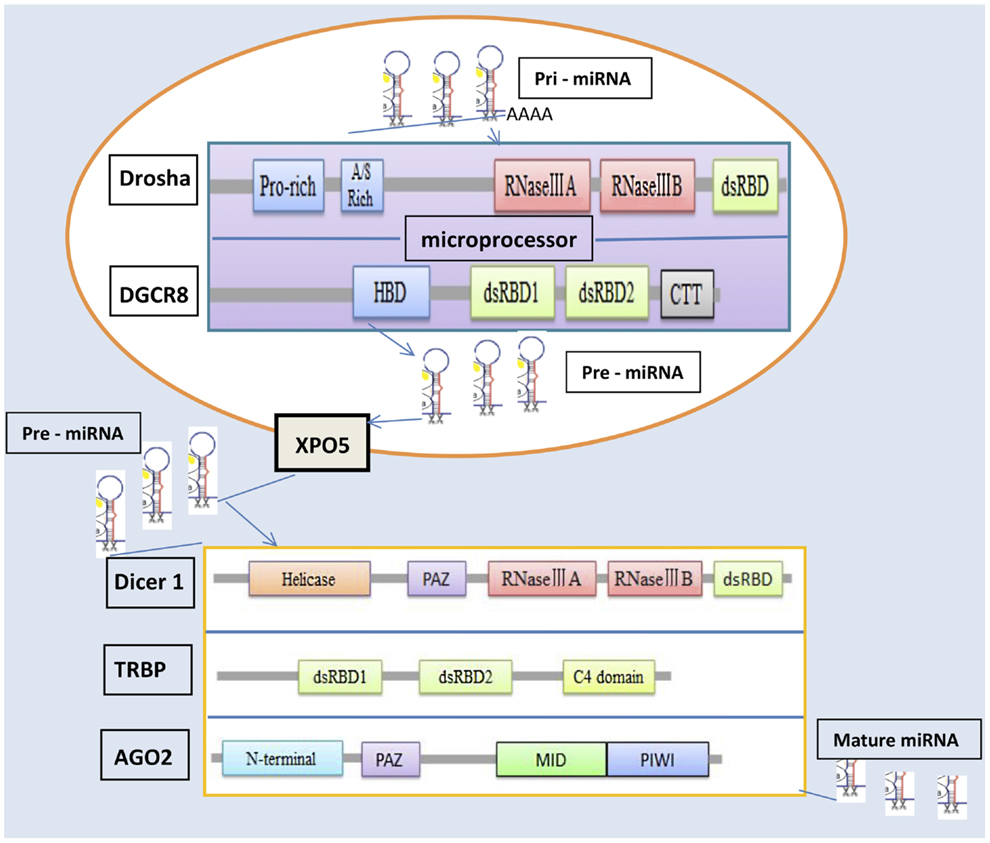
MicroRNAs can be produced from long RNA transcripts. Primary miRNAs (pri-miRNAs), which are 1–2 kb long and contain one or more 70-nt hairpin precursor miRNAs (pre-miRNAs), are excised to pre-miRNAs by ribonuclease III (RNase III) and DiGeorge critical region 8 (DGCR8) in the cell nucleus (9–11). The Drosha–DGCR8 complex, known as a microprocessor, is essential for miRNA maturation. Drosha, as the catalytic subunit, has been shown to cleave pri-miRNA-like hairpins harbored within the 5′-untranslated region of the mRNA encoding the DGCR8 protein (12, 13). Drosha is a member of the RNase III family and can convert pri-miRNAs into pre-miRNAs (11), which are exported from the nucleus into the cytoplasm by an exportin-5 (XPO5)/Ran–GTP complex (14–16). In the cytoplasm, the endoribonuclease Dicer complex catalyzes these pre-RNAs to form miRNAs (17). The mature miRNAs are loaded into an argonaute 2 (AGO2) protein, which associates with a TAR RNA-binding protein (TRBP) and forms an RNA-induced silencing complex (RISC) (18, 19), which plays a crucial role in the repression or degradation of mRNAs.
miRNA Machinery Genes
The miRNA machinery genes include Drosha, DGCR8, Dicer1, XPO5, TRBP, and AGO2, which synthesize proteins to regulate the processing of miRNAs and influence different fields in vivo. Drosha, a nuclear RNase III enzyme, has two RNase III catalytic sites with a double-stranded RNA-binding domain (dsRBD) at the C terminus and a proline-rich domain and arginine/serine-rich domains at the N terminus (11). Drosha recognizes the stem-loop structure and cleaves both arms of the stem-loop through the tandem RNase III domains. The RNase III family of enzymes, which are found in all eubacteria and eukaryotes (20), is divided into three classes based on their structure. Of these classes, Drosha class II and Dicer class III have crucial effects on miRNA processing. The long pri-miRNA, which is typically generated by RNA polymerase II, contains a short stem-loop structure (11). DGCR8 can stabilize the Drosha protein through protein–protein interaction (12) and is an essential miRNA processing factor that includes an N-terminal region for nuclear localization, a heme-binding domain, two dsRBDs, and a C-terminal tail (21, 22). DGCR8 binds to the base of the long primary transcript pri-miRNA hairpin, positioning Drosha to cleave the pri-miRNA stem at a distance of 11 base pairs from the junction between the double-stranded RNA (dsRNA) stem and the flanking single-stranded RNA regions (23).
XPO5 is a nuclear receptor that transports pre-miRNA from the nucleus to the cytoplasm (24, 25). Once in the cytoplasm, pre-miRNA is cleaved by Dicer in complex with another dsRNA-binding protein, TRBP (19, 25, 26). As a key protein in the cleaving process of pri-miRNA, Dicer has two RNase III domains, a less-conserved ATPase/DExD helicase domain and a Piwi–Argonaute–Zwille (PAZ) domain (27, 28). The key regions for miRNA maturation, these domains have different effects. The RNase IIIA domain of Dicer1 is essential for generating small RNAs embedded in the 3′ stem of exogenous hairpin-like RNAs (29). Inactivation of this domain results in complete loss of 3p-derived mature miRNAs but only partial reduction in 5p-derived mature miRNAs (30). In contrast, inactivation of the RNase IIIB domain by mutation of D1709 results in complete loss of 5p-derived mature miRNAs but only partial reduction in 3p-derived mature miRNAs. Mutation of the PAZ domain in Dicer results in global reduction of miRNA processing (30).
Argonaute proteins are core components of RISCs and are highly conserved between species. Many organisms encode multiple members of this protein family, which have essential roles in RNA-mediated gene silencing (31). AGO2 protein contains four major domains, N-terminal, PAZ domain, MID domain, and PIWI domain (32, 33), as well as two structured linker domains, L1 and L2 (34). The PAZ domain, like Dicer, binds to the 3′ end of guide RNA (35). The MID domain of the eukaryotic AGO protein QDE-2 adopts a Rossmann-like fold and recognizes the 50-nt terminal of a guide RNA in a manner similar to its prokaryotic counterparts, for which the 50-nt-binding site shares common residues with a second, adjacent ligand-binding site (36).
TAR RNA-binding protein is a dsRNA-binding protein that includes two dsRBDs and a C4 domain (37). The two dsRBDs together express a much higher affinity for binding dsRNA than either one alone, confirming that the two domains cooperate for dsRNA binding (38, 39). However, a KR-helix motif in dsRBD2 gives it a stronger dsRNA-binding efficacy than dsRBD1 has (38). The C-terminal domain in TRBP binds to the tumor suppressor Merlin, the RNase III Dicer, and PKR activator (PACT) to create the Medipal domain (40). The C4 domain has a major influence on the reactions of TRBP–PACT and TRBP–Dicer.
Figure 1 clarifies the molecular mechanisms underlying the miRNA processing machinery and the three-dimensional structures of the relevant proteins. Two RNase III domains, IIIA and IIIB are, common to Drosha and Dicer1. The PAZ domain is common to Dicer1 and AGO2 (23).
Functions of miRNA Machinery Genes
Because of their core functions in miRNA processing, the genes Drosha, DGCR8, Dicer1, XPO5, AGO2, and TRBP are important in several aspects. Drosha can recognize and cleave the stem-loop structures in mRNAs, leading to their dysfunction, which occurs mostly in stem or progenitor cell populations (41–44). Due to the fact that the transcription factor Neurogenin 2 has a conserved hairpin-like pri-miRNAs do, Drosha can regulate the expression of this transcription factor (41). Drosha knockout mice are infertile due to oligoteratozoospermia or azoospermia, which suggests that Drosha-mediated miRNA production is important in male fertility (45). Drosha can affect the proliferation of human mesenchymal stem cells by regulating rRNA processing (46). Inhibition of Drosha also affects rRNA processing in HeLa cells (47).
DGCR8 is also a part of a microprocessor, the Drosha–DGCR8 complex, with an important function in miRNA maturation. This complex cleaves the hairpin structures in DGCR8 mRNA (12). Deficiency of DGCR8 results in altered short-term plasticity in the prefrontal cortex, affects dendritic spines and complexity, and alters brain miRNA biogenesis (48, 49). Moreover, inactivation of DGRC8 in cardiac neural crest cells results in malformations and increased apoptosis (50). In addition, the loss of DGCR8 in vascular smooth muscle cells results in liver hemorrhage, dilated blood vessels, and disarrayed vascular architecture in murine models, implying that the DGCR8 gene plays an important role in vascular development by regulating the apoptosis and differentiation of these cells (51).
As an RNase III endonuclease, Dicer is a core enzyme which cleaves pre-miRNAs into 21- to 25-nt species in miRNA processing. Dicer has many important roles in the morphogenesis of developing tissues. For example, it plays an essential role in neuron polarity and neuronal development (52) and represses neuronal genes during endocrine cell maturation (53). Loss of Dicer results in gross abnormalities in cell number and function in the cortex and hippocampus (54), and deletion of Dicer in the early pancreatic lineage in Pdx1–Cre mouse models results in early pancreatic bud development and pancreas agenesis (55). On the other hand, Dicer is required for maintaining adult pancreas, and morphologic abnormalities in Dicer1-hypomorphic mice can be detected after 4 weeks of age (56). Deregulation of Dicer1 in β-cells leads to progressive reduction in insulin secretion, glucose tolerance, and development of diabetes and impaired islet architecture (57, 58). Moreover, loss of Dicer results in significant reductions of testis mass and sperm number in germ cell knockout mouse as well as impaired meiotic progression (59). Finally, inactivation of Dicer in developing mouse lymphocytes can impair cell proliferation and survival and alter the repertoires of antigen receptors (60).
Mature RISC consists of a single-stranded small RNA bound to an AGO protein. AGO proteins can bind small interfering RNAs as well as miRNAs and mediate the repression of specific target RNAs either by degrading RNA or by inhibiting translation. Members of the AGO protein family have been implicated in both transcriptional and post-transcriptional gene silencing (31, 61). As a highly specialized member of the AGO family, AGO2 has an essential non-redundant Slicer-independent function within the mammalian miRNA pathway. AGO2 is also a key regulator of B lymphoid and erythroid development and function. However, deficiency in AGO2 impairs miRNA biogenesis from pre-miRNAs and reduces miRNA expression levels (62). On the other hand, AGO protein-associated small RNAs repress mitogen-induced transcripts, and stabilized and stored mature miRNAs can be activated to regulate the mitogenic responses (63). Interestingly, AGO2 in dopamine 2 receptor-expressing neurons regulates cocaine addiction (64), and nuclear AGO2 has been reported to regulate voltage-gated potassium channels in adipose tissue-derived stromal cells with crucial functions in the self-renewal and cell de-aging processes (65). Furthermore, the interaction between the epidermal growth factor receptor gene (EGFR), a novel upstream regulator of the RISC-loading complex, and AGO2 increases under hypoxia stress, which leads to elevated AGO2-Y393 phosphorylation and inhibition of the transition of pre-miRNAs into mature miRNAs (66).
As a steroid receptor RNA activator-binding nuclear receptor coregulator, TRBP targets steroid-responsive promoters and regulates nuclear receptor activity and downstream gene expression (67). TRBP contributes to HIV-1 gene expression by inhibiting the activation of the dsRNA-dependent protein kinase R (PKR) (68). Knockdown of TRBP can reduce the accumulation of hepatitis C virus RNA (69) and TRBP has been proposed as a target for antiviral therapies (68, 70, 71). The structures of TRBP and the PKR activator (PACT) are highly homologous (72). TRBP can control the PACT activation of PKR and the expression of the HIV-1 gene (73). The interaction between TRBP and PACT may influence other cellular processes as well. TRBP can bind to the small-molecule enoxacin and express tumor suppressors in human cell cultures and mouse cancer models (74).
XPO5 protein directly binds and mediates the nuclear export of dsRNA, including pre-miRNAs, viral hairpin RNAs, and tRNAs (16, 75). Inhibition of XPO5 results in down-regulation of Dicer (76), global miRNA elevation disorder, and delayed G1/S transition (77), indicating that XPO5 is a critical component in miRNA biogenesis, regulates global miRNA expression, and is associated with cell-cycle control. Because aberrant expression of XPO5 increases the risk of cancer (78), it is a potential target for drug intervention.
miRNA Machinery Genes in Cancer
Alterations in the miRNA machinery play important roles in the carcinogenesis of a variety of tumors (79). Preliminary analysis of data from The Cancer Genome Atlas consortium of multiple types of cancer through cBioPortal (80, 81) has shown a significant incidence of alterations in miRNA machinery genes (Table 1), especially the AGO2 gene, which has a high incidence of gene alterations across cancer types, including breast invasive carcinoma (23.30%), colon and rectum adenocarcinoma (12.3%), bladder urothelial carcinoma (20.8%), and prostate adenocarcinoma (20.7%). This evidence supports prior reports linking miRNA-related alterations to cancers. The incidence of alterations mutation, copy number variation, and/or deregulated mRNA expression for these cancer types was 80.6, 95.4, 96.0, and 80.5%, respectively.
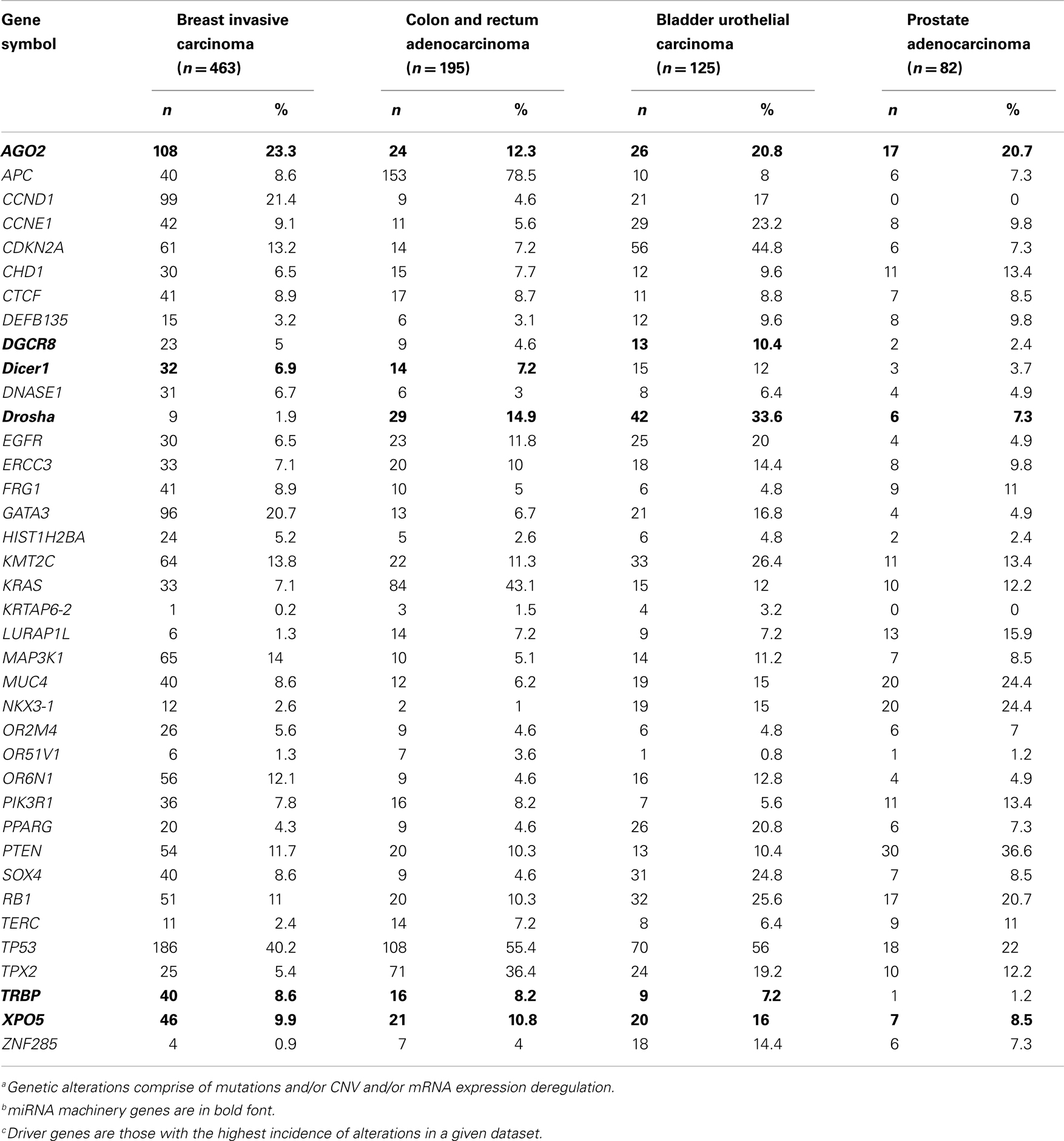
Table 1. The Cancer Genome Atlas consortium data on the incidence of genetic alterationsa in microRNA machinery genesb and driver genesc, by cancer type.
Since the incidence of alteration of the AGO2 gene was highest in breast invasive carcinoma, we analyzed the miRNA machinery genes in breast invasive carcinoma datasets to identify patterns of mutual genetic alterations and driver genes. A strong tendency of mutual exclusivity was noted for genetic alterations in the miRNA machinery gene TRBP with the driver genes PIK3R1 (p = 0.03) and KMT2C (p = 0.0019) (Table 2). We also noted several incidences of co-occurrences in Table 2. Our analysis suggested that alterations in miRNA machinery genes interact with driver genes in at least a subset of tumors. Considering the regulatory role of miRNAs, the underlying mechanisms and cellular consequences of these interactions may be critical for understanding cancer pathology.
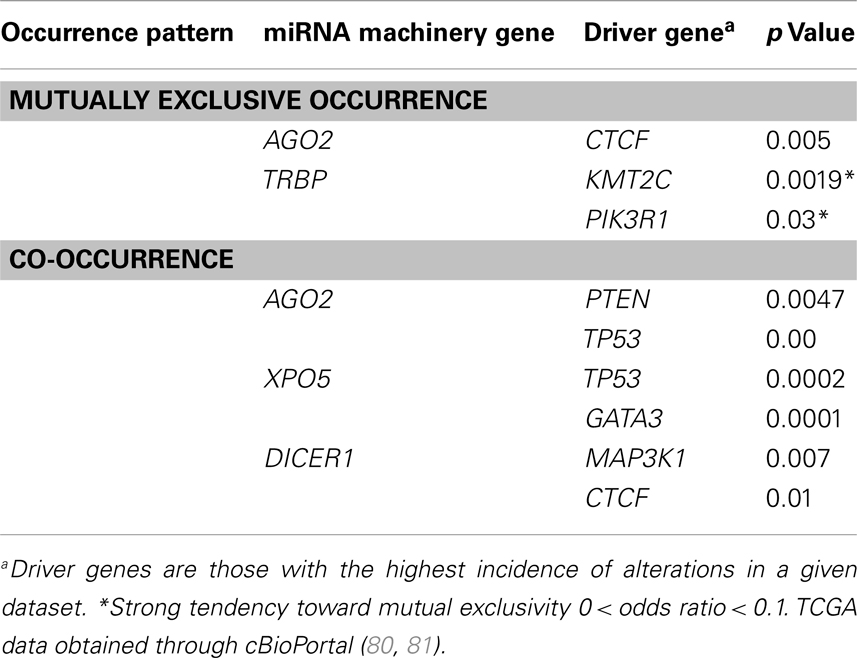
Table 2. Patterns of mutual exclusivity and co-occurrence of driver genes and microRNA machinery gene alteration in The Cancer Genome Atlas consortium data for the breast invasive carcinoma dataset (N = 463).
miRNA Machinery Genes as Biomarkers for Cancers
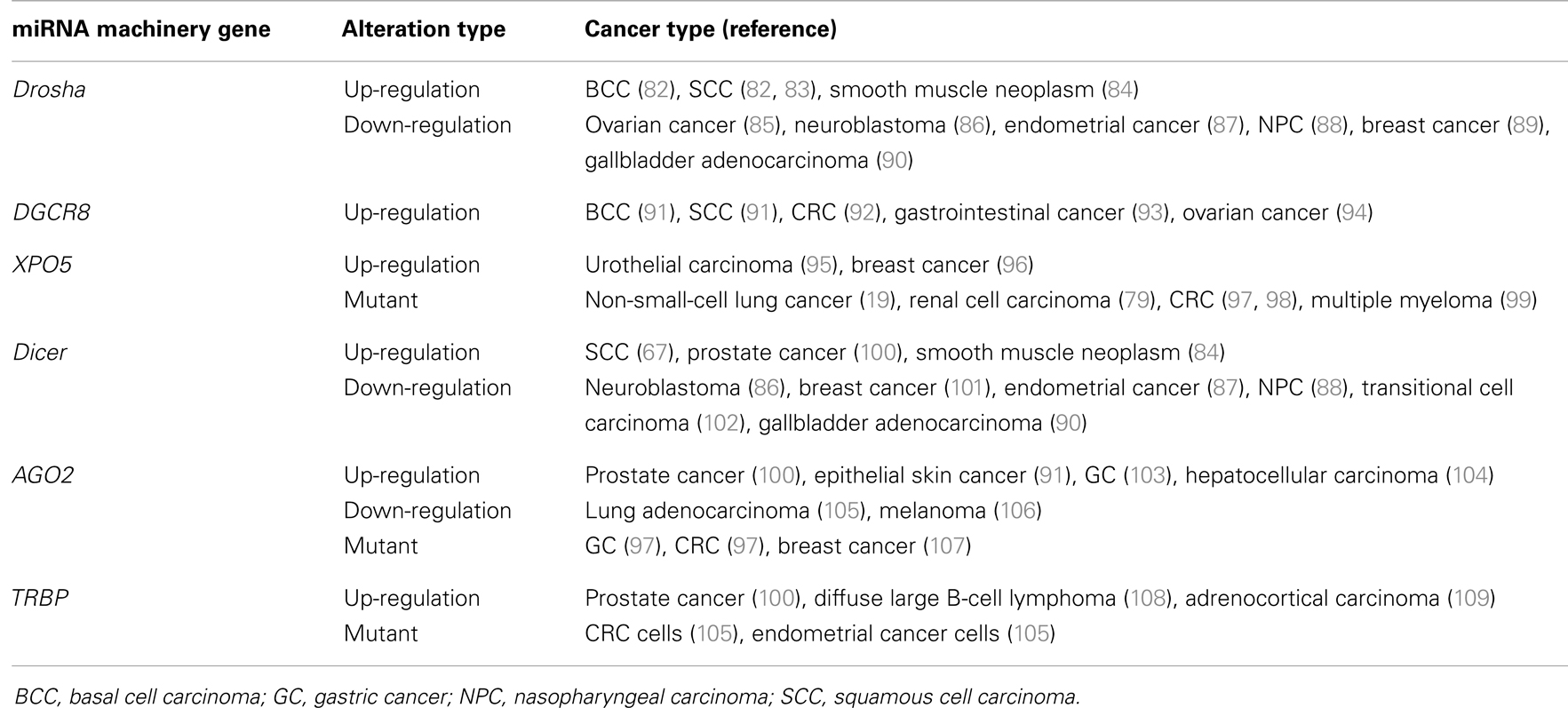
Although the mechanism of microprocessor activity has been intensively investigated and dysregulation of miRNA machinery genes plays a pivotal role in the initiation and progression of malignancies, it remains largely unknown how miRNA machinery genes are regulated and whether they can serve as biomarkers for cancers. Abnormal expression of miRNA machinery genes has been found in a variety of human tumors (Table 3). The expression levels of Drosha, DGCR8, Dicer, XPO5, AGO2, and TRBP have all been associated with several cancers.
The expression level of Drosha is up-regulated in basal cell carcinoma and squamous cell carcinoma (SCC) (82, 83), and elevated levels of Drosha are observed in smooth muscle neoplasms compared with smooth muscle, indicating that this enzyme is involved in smooth muscle neoplasms (85). Down-regulation of Drosha is associated with patient outcome in ovarian cancer (85), outcomes and risk groups in neuroblastoma (86), occurs in endometrial cancer (87), correlates with nasopharyngeal carcinoma and the patient outcomes (88), is associated with the specific subgroups of breast cancer (89), and is associated with metastasis, invasion, and poor prognosis in gallbladder adenocarcinoma (90).
DGCR8 expression levels are over-expressed in basal cell carcinoma (110), SCC (110), colorectal cancer (CRC) (91), gastrointestinal cancer (92), and ovarian cancer (93). Knockdown of DGCR8 in ovarian cancer cells disturbs their proliferation, migration, and invasion and increases their sensitivity to the chemotherapeutic drug cisplatin (93), which suggests that an elevated level of DGCR8 is associated with carcinogenesis.
Dicer is down-regulated in many tumors, such as transitional cell carcinoma of the urinary bladder (94), neuroblastoma (86), nasopharyngeal carcinoma (88), endometrial cancer (87), breast cancer (102), lung cancer (101, 111), gastric cancer (GC) (112), ovarian cancer (113), and gallbladder adenocarcinoma (90). Repression of Dicer is associated with poor prognosis for patients with lung cancer (101), ovarian cancer (114), chronic lymphocytic leukemia (115), or colorectal CRC (116), and it promotes cell proliferation in A2780 and SKOV3 ovarian cancer cells (117). Conversely, compared with normal tissue, the expression of Dicer is higher in cutaneous SCC (82), salivary gland pleomorphic adenoma (118), acute myeloid leukemia (119), smooth muscle neoplasm (85), and prostate cancer (100). Overexpression of Dicer has been shown to lead to poor survival in patients with soft tissue sarcoma (84). Loss of Dicer expression suppresses the growth and oncogenicity of human prostate cancer cell lines but enhances migratory capacity in some prostate cancer cell lines (120). Dicer is increased in human prostate cancer specimens, but lower Dicer expression predicts faster cancer recurrence (120). Complete ablation and hemizygous loss of Dicer reduced tumor growth. Hemizygous loss also resulted in an invasive phenotype and causes seminal vesicle obstruction, which indicated that the regulation of Dicer depends on dosage and context (120).
The expression of AGO2 is up-regulated in GC (103), epithelial skin cancer (110), prostate cancer (100), and hepatocellular carcinoma (104). AGO2 binds to the tumor metastasis factor focal adhesion kinase promoter and triggers its transcription, which suggests a new function of AGO2 in tumor progression (104). Repression of AGO2 protein has been found in human lung adenocarcinomas (105) and in melanoma, for which the mRNA level of AGO2 did not change (106). Overexpression of AGO2 has been shown to inhibit cancer cell proliferation and migration in mice models (105). The stability of AGO2 protein is essential, as is the frame shift mutation of the AGO2 gene in GC and CRC with high microsatellite instability (MSI-H), which suggests that these alterations are risk factors for GC and CRC (97). Single-nucleotide polymorphisms of AGO2 have been associated with the outcome of breast cancer patients (107).
Compared with in lymph nodes, TRBP is over-expressed in prostate cancer (116). Similarly, TRBP is over-expressed in diffuse large B-cell lymphoma and is associated with a poor chemotherapy response (108). Both TRBP mRNA and TRBP protein levels are higher in adrenocortical carcinomas than in adenomas or adrenal cortices (109). Knockdown of TRBP decreases cell proliferation and induces cell apoptosis in diffuse large B-cell lymphoma cells (108) and adrenocortical carcinomas cells (109). However, the expression levels of TRBP are not significantly different between patients with epithelial skin cancer and persons who do not (110). Melo et al. found that the presence of inactivating mutations in TRBP gene in human cancer cell lines and primary tumors with MSI-H impaired miRNA processing and enhanced cellular transformation and the loss of TRBP led to a secondary defect in Dicer1 activity. These results further confirmed the role of loss of function events in the regulation of miRNA processing machinery during tumorigenesis (121).
Dysfunction of XPO5 can also result in carcinogenesis. The expression level of XPO5 is up-regulated in urothelial carcinoma of the bladder (95) and breast cancer (96) and is positively correlated with tumor development and invasion (95). The XPO5 mutant rs11077 increases the risk of renal cell carcinoma (79), is associated with chemotherapy response and survival of patients with advanced non-small-cell lung cancer (24), and is associated with the outcomes of patients with multiple myeloma undergoing autologous stem cell transplantation (99). The discoveries of a mutation in a CRC patient (97) and two CRC cell lines, HCT-15 and DLD-1 (98), with MSI-H imply that the XPO5-inactivating mutant results in pre-miRNA accumulating in the nucleus. The restoration of XPO5 repairs the impaired export and expresses tumor suppressor features (98).
Additional analysis of the expression levels of these miRNA machinery genes and alterations and their interactions with their driver genes in tumors could discriminate cancer patients from healthy controls and be associated with the outcomes of cancer patients.
Future Perspectives
Along with conducting intensive studies of tumor-associated miRNAs and miRNA machinery genes, which play crucial roles in tumorigenesis, scientists are focusing on the miRNA machinery genes Drosha, DGCR8, XPO5, Dicer, AGO2, and TRBP for their potential as cancer biomarkers. The mechanisms involved in miRNA maturation still need to be explored, and new functions of some known genes in miRNA maturation need be uncovered, such as the EGFR gene was induced miRNAs mature as a regulator of AGO2 (71) and ADAR1 formed a complex with Dicer through direct interaction and regulated miRNA processing (122). The dysregulation of miRNA machinery genes (mutation, up-regulation, or down-regulation) can result in oncogenicity and poor patient outcomes. The functions of miRNA machinery genes will be difficult to comprehend because the same gene can have different functions in different types of cancers, and these functions may be not only dosage-dependent but also tissue-dependent (118). Finally, scientists need to explore the different roles of miRNA machinery genes in the physiology and pathology of tumorigenesis. Understanding these roles will help us to use miRNA to develop cancer biomarkers, experimental tools, and antitumor therapy.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by the National Basic Research Program of China 973 Program 2012CB720600, 2012CB720605, the National Natural Science Foundation of China 81271919, and the United States National Cancer Institute grant UO1CA111302.
Abbreviations
AGO2, argonaute 2; CRC, colorectal cancer; DGCR8, diGeorge critical region 8; dsRBD, double-stranded RNA-binding domain; dsRNA, double-stranded RNA; GC, gastric cancer; miRNA, microRNA; MSI-H, high microsatellite instability; PACT, PKR activator; PKR, protein kinase R; pre-miRNA, precursor miRNA; pri-miRNA, primary miRNA; RISC, RNA-induced silencing complex; RNase III, ribonuclease III; SCC, squamous cell carcinoma; TRBP, TAR RNA-binding protein; XPO5, exportin-5.
References
1. Bartel DP. MicroRNAs: target recognition and regulatory functions. Cell (2009) 136:215–33. doi: 10.1016/j.cell.2009.01.002
2. Bartel DP. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell (2004) 116:281–97. doi:10.1016/S0092-8674(04)00045-5
3. Sand M, Skrygan M, Georgas D, Sand D, Gambichler T, Altmeyer P, et al. The miRNA machinery in primary cutaneous malignant melanoma, cutaneous malignant melanoma metastases and benign melanocytic nevi. Cell Tissue Res (2012) 350:119–26. doi:10.1007/s00441-012-1446-0
4. Friedman RC, Farh KK, Burge CB, Bartel DP. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res (2009) 19:92–105. doi:10.1101/gr.082701.108
5. John B, Enright AJ, Aravin A, Tuschl T, Sander C, Marks DS. Human microRNA targets. PLoS Biol (2004) 2:e363. doi:10.1371/journal.pbio.0020363
6. Pasquinelli AE, Reinhart BJ, Slack F, Martindale MQ, Kuroda MI, Maller B, et al. Conservation of the sequence and temporal expression of let-7 heterochronic regulatory RNA. Nature (2000) 408:86–9. doi:10.1038/35040556
7. Shin C, Nam JW, Farh KK, Chiang HR, Shkumatava A, Bartel DP. Expanding the microRNA targeting code: functional sites with centered pairing. Mol Cell (2010) 38:789–802. doi:10.1016/j.molcel.2010.06.005
8. Lal A, Navarro F, Maher CA, Maliszewski LE, Yan N, O’Day E, et al. miR-24 Inhibits cell proliferation by targeting E2F2, MYC, and other cell-cycle genes via binding to “seedless” 3’UTR microRNA recognition elements. Mol Cell (2009) 35:610–25. doi:10.1016/j.molcel.2009.08.020
9. Denli AM, Tops BB, Plasterk RH, Ketting RF, Hannon GJ. Processing of primary microRNAs by the microprocessor complex. Nature (2004) 432:231–5. doi:10.1038/nature03049
10. Gregory RI, Yan KP, Amuthan G, Chendrimada T, Doratotaj B, Cooch N, et al. The microprocessor complex mediates the genesis of microRNAs. Nature (2004) 432:235–40. doi:10.1038/nature03120
11. Lee Y, Ahn C, Han J, Choi H, Kim J, Yim J, et al. The nuclear RNase III Drosha initiates microRNA processing. Nature (2003) 425:415–9. doi:10.1038/nature01957
12. Han J, Pedersen JS, Kwon SC, Belair CD, Kim YK, Yeom KH, et al. Posttranscriptional crossregulation between Drosha and DGCR8. Cell (2009) 136:75–84. doi:10.1016/j.cell.2008.10.053
13. Triboulet R, Chang HM, Lapierre RJ, Gregory RI. Post-transcriptional control of DGCR8 expression by the microprocessor. RNA (2009) 15:1005–11. doi:10.1261/rna.1591709
14. Lund E, Guttinger S, Calado A, Dahlberg JE, Kutay U. Nuclear export of microRNA precursors. Science (2004) 303:95–8. doi:10.1126/science.1090599
15. Yi R, Qin Y, Macara IG, Cullen BR. Exportin-5 mediates the nuclear export of pre-microRNAs and short hairpin RNAs. Genes Dev (2003) 17:3011–6. doi:10.1101/gad.1158803
16. Bohnsack MT, Czaplinski K, Gorlich D. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA (2004) 10:185–91. doi:10.1261/rna.5167604
17. Lee HY, Doudna JA. TRBP alters human precursor microRNA processing in vitro. RNA (2012) 18:2012–9. doi:10.1261/rna.035501.112
18. Haase AD, Jaskiewicz L, Zhang H, Laine S, Sack R, Gatignol A, et al. TRBP, a regulator of cellular PKR and HIV-1 virus expression, interacts with Dicer and functions in RNA silencing. EMBO Rep (2005) 6:961–7. doi:10.1038/sj.embor.7400509
19. Chendrimada TP, Gregory RI, Kumaraswamy E, Norman J, Cooch N, Nishikura K, et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature (2005) 436:740–4. doi:10.1038/nature03868
20. Lamontagne B, Larose S, Boulanger J, Elela SA. The RNase III family: a conserved structure and expanding functions in eukaryotic dsRNA metabolism. Curr Issues Mol Biol (2001) 3:71–8.
21. Senturia R, Laganowsky A, Barr I, Scheidemantle BD, Guo F. Dimerization and heme binding are conserved in amphibian and starfish homologues of the microRNA processing protein DGCR8. PLoS One (2012) 7:e39688. doi:10.1371/journal.pone.0039688
22. Sohn SY, Bae WJ, Kim JJ, Yeom KH, Kim VN, Cho Y. Crystal structure of human DGCR8 core. Nat Struct Mol Biol (2007) 14:847–53. doi:10.1038/nsmb1294
23. Tsunetsugu-Yokota Y, Yamamoto T. Mammalian microRNAs: post-transcriptional gene regulation in RNA virus infection and therapeutic applications. Front Microbiol (2010) 1:108. doi:10.3389/fmicb.2010.00108
24. Ding C, Li C, Wang H, Li B, Guo Z. A miR-SNP of the XPO5 gene is associated with advanced non-small-cell lung cancer. Onco Targets Ther (2013) 6:877–81. doi:10.2147/OTT.S48284
25. Grosshans H, Bussing I. MicroRNA biogenesis takes another single hit from microsatellite instability. Cancer Cell (2010) 18:295–7. doi:10.1016/j.ccr.2010.10.005
26. Bernstein E, Caudy AA, Hammond SM, Hannon GJ. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature (2001) 409:363–6. doi:10.1038/35053110
27. Park JE, Heo I, Tian Y, Simanshu DK, Chang H, Jee D, et al. Dicer recognizes the 5’ end of RNA for efficient and accurate processing. Nature (2011) 475:201–5. doi:10.1038/nature10198
28. Carmell MA, Hannon GJ. RNase III enzymes and the initiation of gene silencing. Nat Struct Mol Biol (2004) 11:214–8. doi:10.1038/nsmb729
29. Ohishi K, Nakano T. A forward genetic screen to study mammalian RNA interference: essential role of RNase IIIa domain of Dicer1 in 3’ strand cleavage of dsRNA in vivo. FEBS J (2012) 279:832–43. doi:10.1111/j.1742-4658.2012.08474.x
30. Gurtan AM, Lu V, Bhutkar A, Sharp PA. In vivo structure-function analysis of human Dicer reveals directional processing of precursor miRNAs. RNA (2012) 18:1116–22. doi:10.1261/rna.032680.112
31. Hock J, Meister G. The argonaute protein family. Genome Biol (2008) 9:210. doi:10.1186/gb-2008-9-2-210
32. Elkayam E, Kuhn CD, Tocilj A, Haase AD, Greene EM, Hannon GJ, et al. The structure of human argonaute-2 in complex with miR-20a. Cell (2012) 150:100–10. doi:10.1016/j.cell.2012.05.017
33. Gaynor JW, Campbell BJ, Cosstick R. RNA interference: a chemist’s perspective. Chem Soc Rev (2010) 39:4169–84. doi:10.1039/b920362c
34. Song JJ, Smith SK, Hannon GJ, Joshua-Tor L. Crystal structure of argonaute and its implications for RISC slicer activity. Science (2004) 305:1434–7. doi:10.1126/science.1102514
35. Wang Y, Sheng G, Juranek S, Tuschl T, Patel DJ. Structure of the guide-strand-containing argonaute silencing complex. Nature (2008) 456:209–13. doi:10.1038/nature07315
36. Boland A, Tritschler F, Heimstadt S, Izaurralde E, Weichenrieder O. Crystal structure and ligand binding of the MID domain of a eukaryotic argonaute protein. EMBO Rep (2010) 11:522–7. doi:10.1038/embor.2010.81
37. Daniels SM, Melendez-Pena CE, Scarborough RJ, Daher A, Christensen HS, El Far M, et al. Characterization of the TRBP domain required for dicer interaction and function in RNA interference. BMC Mol Biol (2009) 10:38. doi:10.1186/1471-2199-10-38
38. Yamashita S, Nagata T, Kawazoe M, Takemoto C, Kigawa T, Guntert P, et al. Structures of the first and second double-stranded RNA-binding domains of human TAR RNA-binding protein. Protein Sci (2011) 20:118–30. doi:10.1002/pro.543
39. Daniels SM, Gatignol A. The multiple functions of TRBP, at the hub of cell responses to viruses, stress, and cancer. Microbiol Mol Biol Rev (2012) 76:652–66. doi:10.1128/MMBR.00012-12
40. Laraki G, Clerzius G, Daher A, Melendez-Pena C, Daniels S, Gatignol A. Interactions between the double-stranded RNA-binding proteins TRBP and PACT define the Medipal domain that mediates protein-protein interactions. RNA Biol (2008) 5:92–103. doi:10.4161/rna.5.2.6069
41. Knuckles P, Vogt MA, Lugert S, Milo M, Chong MM, Hautbergue GM, et al. Drosha regulates neurogenesis by controlling neurogenin 2 expression independent of microRNAs. Nat Neurosci (2012) 15:962–9. doi:10.1038/nn.3139
42. Chong MM, Zhang G, Cheloufi S, Neubert TA, Hannon GJ, Littman DR. Canonical and alternate functions of the microRNA biogenesis machinery. Genes Dev (2010) 24:1951–60. doi:10.1101/gad.1953310
43. Karginov FV, Cheloufi S, Chong MM, Stark A, Smith AD, Hannon GJ. Diverse endonucleolytic cleavage sites in the mammalian transcriptome depend upon microRNAs, Drosha, and additional nucleases. Mol Cell (2010) 38:781–8. doi:10.1016/j.molcel.2010.06.001
44. Shenoy A, Blelloch R. Genomic analysis suggests that mRNA destabilization by the microprocessor is specialized for the auto-regulation of Dgcr8. PLoS One (2009) 4:e6971. doi:10.1371/journal.pone.0006971
45. Wu Q, Song R, Ortogero N, Zheng H, Evanoff R, Small CL, et al. The RNase III enzyme DROSHA is essential for microRNA production and spermatogenesis. J Biol Chem (2012) 287:25173–90. doi:10.1074/jbc.M112.362053
46. Oskowitz AZ, Penfornis P, Tucker A, Prockop DJ, Pochampally R. Drosha regulates hMSCs cell cycle progression through a miRNA independent mechanism. Int J Biochem Cell Biol (2011) 43:1563–72. doi:10.1016/j.biocel.2011.07.005
47. Wu H, Xu H, Miraglia LJ, Crooke ST. Human RNase III is a 160-kDa protein involved in preribosomal RNA processing. J Biol Chem (2000) 275:36957–65. doi:10.1074/jbc.M005494200
48. Fenelon K, Mukai J, Xu B, Hsu PK, Drew LJ, Karayiorgou M, et al. Deficiency of Dgcr8, a gene disrupted by the 22q11.2 microdeletion, results in altered short-term plasticity in the prefrontal cortex. Proc Natl Acad Sci U S A (2011) 108:4447–52. doi:10.1073/pnas.1101219108
49. Stark KL, Xu B, Bagchi A, Lai WS, Liu H, Hsu R, et al. Altered brain microRNA biogenesis contributes to phenotypic deficits in a 22q11-deletion mouse model. Nat Genet (2008) 40:751–60. doi:10.1038/ng.138
50. Chapnik E, Sasson V, Blelloch R, Hornstein E. Dgcr8 controls neural crest cells survival in cardiovascular development. Dev Biol (2012) 362:50–6. doi:10.1016/j.ydbio.2011.11.008
51. Chen Z, Wu J, Yang C, Fan P, Balazs L, Jiao Y, et al. DiGeorge syndrome critical region 8 (DGCR8) protein-mediated microRNA biogenesis is essential for vascular smooth muscle cell development in mice. J Biol Chem (2012) 287:19018–28. doi:10.1074/jbc.M112.351791
52. Li Z, He X, Feng J. Dicer is essential for neuronal polarity. Int J Dev Neurosci (2012) 30:607–11. doi:10.1016/j.ijdevneu.2012.08.002
53. Kanji MS, Martin MG, Bhushan A. Dicer1 is required to repress neuronal fate during endocrine cell maturation. Diabetes (2013) 62:1602–11. doi:10.2337/db12-0841
54. Davis TH, Cuellar TL, Koch SM, Barker AJ, Harfe BD, McManus MT, et al. Conditional loss of Dicer disrupts cellular and tissue morphogenesis in the cortex and hippocampus. J Neurosci (2008) 28:4322–30. doi:10.1523/JNEUROSCI.4815-07.2008
55. Lynn FC, Skewes-Cox P, Kosaka Y, McManus MT, Harfe BD, German MS. MicroRNA expression is required for pancreatic islet cell genesis in the mouse. Diabetes (2007) 56:2938–45. doi:10.2337/db07-0175
56. Morita S, Hara A, Kojima I, Horii T, Kimura M, Kitamura T, et al. Dicer is required for maintaining adult pancreas. PLoS One (2009) 4:e4212. doi:10.1371/journal.pone.0004212
57. Mandelbaum AD, Melkman-Zehavi T, Oren R, Kredo-Russo S, Nir T, Dor Y, et al. Dysregulation of Dicer1 in beta cells impairs islet architecture and glucose metabolism. Exp Diabetes Res (2012) 2012:470302. doi:10.1155/2012/470302
58. Kalis M, Bolmeson C, Esguerra JL, Gupta S, Edlund A, Tormo-Badia N, et al. Beta-cell specific deletion of Dicer1 leads to defective insulin secretion and diabetes mellitus. PLoS One (2011) 6:e29166. doi:10.1371/journal.pone.0029166
59. Greenlee AR, Shiao MS, Snyder E, Buaas FW, Gu T, Stearns TM, et al. Deregulated sex chromosome gene expression with male germ cell-specific loss of Dicer1. PLoS One (2012) 7:e46359. doi:10.1371/journal.pone.0046359
60. Brady BL, Rupp LJ, Bassing CH. Requirement for dicer in survival of proliferating thymocytes experiencing DNA double-strand breaks. J Immunol (2013) 190:3256–66. doi:10.4049/jimmunol.1200957
61. Czech B, Hannon GJ. Small RNA sorting: matchmaking for argonautes. Nat Rev Genet (2011) 12:19–31. doi:10.1038/nrg2916
62. O’Carroll D, Mecklenbrauker I, Das PP, Santana A, Koenig U, Enright AJ, et al. A slicer-independent role for argonaute 2 in hematopoiesis and the microRNA pathway. Genes Dev (2007) 21:1999–2004. doi:10.1101/gad.1565607
63. Olejniczak SH, La Rocca G, Gruber JJ, Thompson CB. Long-lived microRNA-argonaute complexes in quiescent cells can be activated to regulate mitogenic responses. Proc Natl Acad Sci U S A (2013) 110:157–62. doi:10.1073/pnas.1219958110
64. Schaefer A, Im HI, Veno MT, Fowler CD, Min A, Intrator A, et al. Argonaute 2 in dopamine 2 receptor-expressing neurons regulates cocaine addiction. J Exp Med (2010) 207:1843–51. doi:10.1084/jem.20100451
65. Kim BS, Im YB, Jung SJ, Park CH, Kang SK. Argonaute2 regulation for K+ channel-mediated human adipose tissue-derived stromal cells self-renewal and survival in nucleus. Stem Cells Dev (2012) 21:1736–48. doi:10.1089/scd.2011.0388
66. Shen J, Xia W, Khotskaya YB, Huo L, Nakanishi K, Lim SO, et al. EGFR modulates microRNA maturation in response to hypoxia through phosphorylation of AGO2. Nature (2013) 497:383–7. doi:10.1038/nature12080
67. Redfern AD, Colley SM, Beveridge DJ, Ikeda N, Epis MR, Li X, et al. RNA-induced silencing complex (RISC) proteins PACT, TRBP, and Dicer are SRA binding nuclear receptor coregulators. Proc Natl Acad Sci U S A (2013) 110:6536–41. doi:10.1073/pnas.1301620110
68. Sanghvi VR, Steel LF. The cellular TAR RNA binding protein, TRBP, promotes HIV-1 replication primarily by inhibiting the activation of double-stranded RNA-dependent kinase PKR. J Virol (2011) 85:12614–21. doi:10.1128/JVI.05240-11
69. Zhang C, Huys A, Thibault PA, Wilson JA. Requirements for human Dicer and TRBP in microRNA-122 regulation of HCV translation and RNA abundance. Virology (2012) 433:479–88. doi:10.1016/j.virol.2012.08.039
70. Christensen HS, Daher A, Soye KJ, Frankel LB, Alexander MR, Laine S, et al. Small interfering RNAs against the TAR RNA binding protein, TRBP, a Dicer cofactor, inhibit human immunodeficiency virus type 1 long terminal repeat expression and viral production. J Virol (2007) 81:5121–31. doi:10.1128/JVI.01511-06
71. Eekels JJ, Geerts D, Jeeninga RE, Berkhout B. Long-term inhibition of HIV-1 replication with RNA interference against cellular co-factors. Antiviral Res (2011) 89:43–53. doi:10.1016/j.antiviral.2010.11.005
72. Kok KH, Ng MH, Ching YP, Jin DY. Human TRBP and PACT directly interact with each other and associate with dicer to facilitate the production of small interfering RNA. J Biol Chem (2007) 282:17649–57. doi:10.1074/jbc.M611768200
73. Daher A, Laraki G, Singh M, Melendez-Pena CE, Bannwarth S, Peters AH, et al. TRBP control of PACT-induced phosphorylation of protein kinase R is reversed by stress. Mol Cell Biol (2009) 29:254–65. doi:10.1128/MCB.01030-08
74. Melo S, Villanueva A, Moutinho C, Davalos V, Spizzo R, Ivan C, et al. Small molecule enoxacin is a cancer-specific growth inhibitor that acts by enhancing TAR RNA-binding protein 2-mediated microRNA processing. Proc Natl Acad Sci U S A (2011) 108:4394–9. doi:10.1073/pnas.1014720108
75. Bohnsack MT, Regener K, Schwappach B, Saffrich R, Paraskeva E, Hartmann E, et al. Exp5 exports eEF1A via tRNA from nuclei and synergizes with other transport pathways to confine translation to the cytoplasm. EMBO J (2002) 21:6205–15. doi:10.1093/emboj/cdf613
76. Bennasser Y, Chable-Bessia C, Triboulet R, Gibbings D, Gwizdek C, Dargemont C, et al. Competition for XPO5 binding between Dicer mRNA, pre-miRNA and viral RNA regulates human Dicer levels. Nat Struct Mol Biol (2011) 18:323–7. doi:10.1038/nsmb.1987
77. Iwasaki YW, Kiga K, Kayo H, Fukuda-Yuzawa Y, Weise J, Inada T, et al. Global microRNA elevation by inducible Exportin 5 regulates cell cycle entry. RNA (2013) 19:490–7. doi:10.1261/rna.036608.112
78. Muqbil I, Bao B, Abou-Samra AB, Mohammad RM, Azmi AS. Nuclear export mediated regulation of microRNAs: potential target for drug intervention. Curr Drug Targets (2013) 14:1094–100. doi:10.2174/1389450111314100002
79. Horikawa Y, Wood CG, Yang H, Zhao H, Ye Y, Gu J, et al. Single nucleotide polymorphisms of microRNA machinery genes modify the risk of renal cell carcinoma. Clin Cancer Res (2008) 14:7956–62. doi:10.1158/1078-0432.CCR-08-1199
80. Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal (2013) 6:l1. doi:10.1126/scisignal.2004088
81. Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA, et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov (2012) 2:401–4. doi:10.1158/2159-8290.CD-12-0095
82. Sand M, Gambichler T, Skrygan M, Sand D, Scola N, Altmeyer P, et al. Expression levels of the microRNA processing enzymes Drosha and dicer in epithelial skin cancer. Cancer Invest (2010) 28:649–53. doi:10.3109/07357901003630918
83. Muralidhar B, Winder D, Murray M, Palmer R, Barbosa-Morais N, Saini H, et al. Functional evidence that Drosha overexpression in cervical squamous cell carcinoma affects cell phenotype and microRNA profiles. J Pathol (2011) 224:496–507. doi:10.1002/path.2898
84. Papachristou DJ, Rao UN, Korpetinou A, Giannopoulou E, Sklirou E, Kontogeorgakos V, et al. Prognostic significance of Dicer cellular levels in soft tissue sarcomas. Cancer Invest (2012) 30:172–9. doi:10.3109/07357907.2011.633293
85. Papachristou DJ, Sklirou E, Corradi D, Grassani C, Kontogeorgakos V, Rao UN. Immunohistochemical analysis of the endoribonucleases Drosha, Dicer and Ago2 in smooth muscle tumours of soft tissues. Histopathology (2012) 60:E28–36. doi:10.1111/j.1365-2559.2012.04192.x
86. Lin RJ, Lin YC, Chen J, Kuo HH, Chen YY, Diccianni MB, et al. microRNA signature and expression of Dicer and Drosha can predict prognosis and delineate risk groups in neuroblastoma. Cancer Res (2010) 70:7841–50. doi:10.1158/0008-5472.CAN-10-0970
87. Torres A, Torres K, Paszkowski T, Jodlowska-Jedrych B, Radomanski T, Ksiazek A, et al. Major regulators of microRNAs biogenesis Dicer and Drosha are down-regulated in endometrial cancer. Tumour Biol (2011) 32:769–76. doi:10.1007/s13277-011-0179-0
88. Guo X, Liao Q, Chen P, Li X, Xiong W, Ma J, et al. The microRNA-processing enzymes: Drosha and Dicer can predict prognosis of nasopharyngeal carcinoma. J Cancer Res Clin Oncol (2012) 138:49–56. doi:10.1007/s00432-011-1058-1
89. Dedes KJ, Natrajan R, Lambros MB, Geyer FC, Lopez-Garcia MA, Savage K, et al. Down-regulation of the miRNA master regulators Drosha and Dicer is associated with specific subgroups of breast cancer. Eur J Cancer (2011) 47:138–50. doi:10.1016/j.ejca.2010.08.007
90. Shu GS, Yang ZL, Liu DC. Immunohistochemical study of Dicer and Drosha expression in the benign and malignant lesions of gallbladder and their clinicopathological significances. Pathol Res Pract (2012) 208:392–7. doi:10.1016/j.prp.2012.05.001
91. Kim B, Lee JH, Park JW, Kwon TK, Baek SK, Hwang I, et al. An essential microRNA maturing microprocessor complex component DGCR8 is up-regulated in colorectal carcinomas. Clin Exp Med (2013). doi:10.1007/s10238-013-0243-8
92. Jafari N, Dogaheh HP, Bohlooli S, Oyong GG, Shirzad Z, Alibeiki F, et al. Expression levels of microRNA machinery components Drosha, Dicer and DGCR8 in human (AGS, HepG2, and KEYSE-30) cancer cell lines. Int J Clin Exp Med (2013) 6:269–74.
93. Guo Y, Tian P, Yang C, Liang Z, Li M, Sims M, et al. Silencing the double-stranded RNA binding protein DGCR8 inhibits ovarian cancer cell proliferation, migration, and invasion. Pharm Res (2013). doi:10.1007/s11095-013-1219-9
94. Wu D, Tao J, Xu B, Li P, Lu Q, Zhang W. Downregulation of Dicer, a component of the microRNA machinery, in bladder cancer. Mol Med Rep (2012) 5:695–9. doi:10.3892/mmr.2011.711
95. Han Y, Liu Y, Gui Y, Cai Z. Inducing cell proliferation inhibition and apoptosis via silencing Dicer, Drosha, and Exportin 5 in urothelial carcinoma of the bladder. J Surg Oncol (2013) 107:201–5. doi:10.1002/jso.23214
96. Leaderer D, Hoffman AE, Zheng T, Fu A, Weidhaas J, Paranjape T, et al. Genetic and epigenetic association studies suggest a role of microRNA biogenesis gene exportin-5 (XPO5) in breast tumorigenesis. Int J Mol Epidemiol Genet (2011) 2:9–18.
97. Kim MS, Oh JE, Kim YR, Park SW, Kang MR, Kim SS, et al. Somatic mutations and losses of expression of microRNA regulation-related genes AGO2 and TNRC6A in gastric and colorectal cancers. J Pathol (2010) 221:139–46. doi:10.1002/path.2683
98. Melo SA, Moutinho C, Ropero S, Calin GA, Rossi S, Spizzo R, et al. A genetic defect in exportin-5 traps precursor microRNAs in the nucleus of cancer cells. Cancer Cell (2010) 18:303–15. doi:10.1016/j.ccr.2010.09.007
99. de Larrea CF, Navarro A, Tejero R, Tovar N, Diaz T, Cibeira MT, et al. Impact of MiRSNPs on survival and progression in patients with multiple myeloma undergoing autologous stem cell transplantation. Clin Cancer Res (2012) 18:3697–704. doi:10.1158/1078-0432.CCR-12-0191
100. Fu X, Xue C, Huang Y, Xie Y, Li Y. The activity and expression of microRNAs in prostate cancers. Mol Biosyst (2010) 6:2561–72. doi:10.1039/c0mb00100g
101. Karube Y, Tanaka H, Osada H, Tomida S, Tatematsu Y, Yanagisawa K, et al. Reduced expression of Dicer associated with poor prognosis in lung cancer patients. Cancer Sci (2005) 96:111–5. doi:10.1111/j.1349-7006.2005.00015.x
102. Khoshnaw SM, Rakha EA, Abdel-Fatah TM, Nolan CC, Hodi Z, Macmillan DR, et al. Loss of Dicer expression is associated with breast cancer progression and recurrence. Breast Cancer Res Treat (2012) 135:403–13. doi:10.1007/s10549-012-2169-3
103. Zhang J, Fan XS, Wang CX, Liu B, Li Q, Zhou XJ. Up-regulation of Ago2 expression in gastric carcinoma. Med Oncol (2013) 30:628. doi:10.1007/s12032-013-0628-2
104. Cheng N, Li Y, Han ZG. Argonaute2 promotes tumor metastasis by way of up-regulating focal adhesion kinase expression in hepatocellular carcinoma. Hepatology (2013) 57:1906–18. doi:10.1002/hep.26202
105. Zhang X, Graves P, Zeng Y. Overexpression of human argonaute2 inhibits cell and tumor growth. Biochim Biophys Acta (2013) 1830:2553–61. doi:10.1016/j.bbagen.2012.11.013
106. Voller D, Reinders J, Meister G, Bosserhoff AK. Strong reduction of AGO2 expression in melanoma and cellular consequences. Br J Cancer (2013) 109:3116–24. doi:10.1038/bjc.2013.646
107. Sung H, Jeon S, Lee KM, Han S, Song M, Choi JY, et al. Common genetic polymorphisms of microRNA biogenesis pathway genes and breast cancer survival. BMC Cancer (2012) 12:195. doi:10.1186/1471-2407-12-195
108. Caramuta S, Lee L, Ozata DM, Akcakaya P, Georgii-Hemming P, Xie H, et al. Role of microRNAs and microRNA machinery in the pathogenesis of diffuse large B-cell lymphoma. Blood Cancer J (2013) 3:e152. doi:10.1038/bcj.2013.49
109. Caramuta S, Lee L, Ozata DM, Akcakaya P, Xie H, Hoog A, et al. Clinical and functional impact of TARBP2 over-expression in adrenocortical carcinoma. Endocr Relat Cancer (2013) 20:551–64. doi:10.1530/ERC-13-0098
110. Sand M, Skrygan M, Georgas D, Arenz C, Gambichler T, Sand D, et al. Expression levels of the microRNA maturing microprocessor complex component DGCR8 and the RNA-induced silencing complex (RISC) components argonaute-1, argonaute-2, PACT, TARBP1, and TARBP2 in epithelial skin cancer. Mol Carcinog (2012) 51:916–22. doi:10.1002/mc.20861
111. Chiosea S, Jelezcova E, Chandran U, Luo J, Mantha G, Sobol RW, et al. Overexpression of Dicer in precursor lesions of lung adenocarcinoma. Cancer Res (2007) 67:2345–50. doi:10.1158/0008-5472.CAN-06-3533
112. Zheng ZH, Sun XJ, Fu WN, Guan Y, Gao F, Wang Y, et al. Decreased expression of DICER1 in gastric cancer. Chin Med J (Engl) (2007) 120:2099–104.
113. Pampalakis G, Diamandis EP, Katsaros D, Sotiropoulou G. Down-regulation of dicer expression in ovarian cancer tissues. Clin Biochem (2010) 43:324–7. doi:10.1016/j.clinbiochem.2009.09.014
114. Merritt WM, Lin YG, Han LY, Kamat AA, Spannuth WA, Schmandt R, et al. Dicer, Drosha, and outcomes in patients with ovarian cancer. N Engl J Med (2008) 359:2641–50. doi:10.1056/NEJMoa0803785
115. Zhu DX, Fan L, Lu RN, Fang C, Shen WY, Zou ZJ, et al. Downregulated Dicer expression predicts poor prognosis in chronic lymphocytic leukemia. Cancer Sci (2012) 103:875–81. doi:10.1111/j.1349-7006.2012.02234.x
116. Faggad A, Kasajima A, Weichert W, Stenzinger A, Elwali NE, Dietel M, et al. Down-regulation of the microRNA processing enzyme Dicer is a prognostic factor in human colorectal cancer. Histopathology (2012) 61:552–61. doi:10.1111/j.1365-2559.2011.04110.x
117. Kuang Y, Cai J, Li D, Han Q, Cao J, Wang Z. Repression of Dicer is associated with invasive phenotype and chemoresistance in ovarian cancer. Oncol Lett (2013) 5:1149–54. doi:10.3892/ol.2013.1158
118. Zhang X, Cairns M, Rose B, O’Brien C, Shannon K, Clark J, et al. Alterations in miRNA processing and expression in pleomorphic adenomas of the salivary gland. Int J Cancer (2009) 124:2855–63. doi:10.1002/ijc.24298
119. Martin MG, Payton JE, Link DC. Dicer and outcomes in patients with acute myeloid leukemia (AML). Leuk Res (2009) 33:e127. doi:10.1016/j.leukres.2009.02.003
120. Zhang B, Chen H, Zhang L, Dakhova O, Zhang Y, Lewis MT, et al. A dosage-dependent pleiotropic role of Dicer in prostate cancer growth and metastasis. Oncogene (2013). doi:10.1038/onc.2013.281
121. Melo SA, Ropero S, Moutinho C, Aaltonen LA, Yamamoto H, Calin GA, et al. A TARBP2 mutation in human cancer impairs microRNA processing and DICER1 function. Nat Genet (2009) 41:365–70. doi:10.1038/ng.317
Keywords: miRNA, machinery genes, alterations, biomarker, cancer
Citation: Huang J-T, Wang J, Srivastava V, Sen S and Liu S-M (2014) MicroRNA machinery genes as novel biomarkers for cancer. Front. Oncol. 4:113. doi: 10.3389/fonc.2014.00113
Received: 11 March 2014; Accepted: 01 May 2014;
Published online: 19 May 2014.
Edited by:
Yawei Zhang, Yale University, USAReviewed by:
Lifang Hou, Northwestern University, USAManuela Marron, Ernst-Moritz-Arndt University Greifswald, Germany
Copyright: © 2014 Huang, Wang, Srivastava, Sen and Liu. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jin Wang, Department of Translational Molecular Pathology, The University of Texas MD Anderson Cancer Center, Unit 2951, 2130 West Holcombe Boulevard, Houston, TX 77030, USA e-mail:andhbmc0QG1kYW5kZXJzb24ub3Jn;
Song-Mei Liu, Center for Gene Diagnosis, Zhongnan Hospital of Wuhan University, 169 Donghu Road, Wuhan, Hubei 430071, China e-mail:c21saXVAd2h1LmVkdS5jbg==
 Jing-Tao Huang1
Jing-Tao Huang1 Jin Wang
Jin Wang Song-Mei Liu
Song-Mei Liu