- 1Department of Respiratory and Infectious Disease of Geriatrics, The First Hospital of China Medical University, Shenyang, China
- 2Department of Pulmonary and Critical Care Medicine, Center of Respiratory Medicine, China-Japan Friendship Hospital, Beijing, China
- 3Graduate School of Peking Union Medical College, Chinese Academy of Medical Science and Peking Union Medical College, Beijing, China
- 4Department of Pathology, Shengjing Hospital of China Medical University, Shenyang, China
- 5Department of Medical Oncology, The First Hospital of China Medical University, Shenyang, China
- 6Department of Medical Oncology, Sir Run Run Shaw Hospital, College of Medicine, Zhejiang University, Hangzhou, China
- 7The First Laboratory of Cancer Institute, The First Hospital of China Medical University, Shenyang, China
Purpose: GPSM2 (G protein signaling modulator 2) was reported to be involved in the cell division of breast cancer cells. Additionally, cytoplasmic dynein may mediate the transport process of GPSM2. DYNC1I1 (Cytoplasmic dynein 1 intermediate chain 1) is the most common cargo-binding subunit of dynein. However, the relationship between GPSM2 and DYNC1I1 and its clinical value is unclear.
Methods: Immunohistochemical staining was performed for assessment of GPSM2 and DYNC1I1 expression. Immunoprecipitation analysis was used to assess the interaction between GPSM2 and DYNC1I1.
Results: GPSM2 was correlated with clinical characteristics of breast cancer patients and is an unfavorable independent prognostic factor. In addition, nuclear expression of GPSM2 is an unfavorable independent prognostic factor (HR = 2.658, 95% CI = 1.490–4.741, p = 0.001). GPSM2 and DYNC1I1 are known to form a complex in breast cancer cells. Patients who were positive for expression of both DYNC1I1 and GPSM2 presented with shorter recurrence-free survival than other patients. Importantly, patients with GPSM2 nuclear expression showed higher DYNC1I1 expression.
Conclusion: GPSM2 was an independent prognostic factor in breast cancer and nuclear expression of GPSM2 was significantly associated with poor prognosis, which was related to the positive expression of DYNC1I1. Examination of both GPSM2 and DYNC1I1 is necessary to establish a prognosis in breast cancer patients.
Introduction
Breast cancer is the most common malignant tumor affecting women worldwide, with ~1.38 million new-onset patients each year and is responsible for 46 million deaths to date (1). Despite advancements in chemotherapy and targeted therapy (2), a significant proportion of patients are at risk of tumor recurrence and metastasis. Therefore, in-depth study of the molecular mechanisms of breast cancer and prognosis biomarkers is essential to improve patient outcomes.
G-protein-coupled receptors (GPCRs) regulate numerous essential biological functions (3, 4). The abnormal expression of GPCRs is associated with occurrence and development of diverse types of cancer, including breast cancer (5, 6). GPSM2 (G protein signaling modulator 2) belongs to a protein family that regulates activation of G proteins (7), which then inhibit GPCR signaling by inhibiting GDP release from Gα subunits (8). GPSM2 is also widely recognized as a determinant of mitotic spindle orientation (9, 10). There are several reports of the role of GPSM2 in cancer. For example, in hepatocellular carcinoma (11) and pancreatic cancer (12), overexpression of GPSM2 promotes tumor progression and is related to prognosis. In addition, one study has observed aberrant expression of GPSM2 in breast cancer (13), but, its clinical value needs to be further explored.
Cytoplasmic dynein is a large protein complex, and has a wide range of cellular functions, including cargo transport (including vesicles, growth factors, and transcription factors) from the cytoplasm to the nucleus (14). DYNC1I1 (cytoplasmic dynein 1 intermediate chain 1) is the most common cargo binding subunit of dynein (15). The role of DYNC1I1 in cancer is controversial. One study reported a tumor-suppressive function of DYNC1I1 in GBM via transport of SK2 (16). Another study indicated, DYNC1I1 promoted gastric cancer progression by increasing NF-kB nuclear translocation (17). Moreover, several studies showed dynein could regulate mitotic spindle localization by mediation of the transport process of GPSM2 (9, 18). However, the clinical value of DYNC1I1 and the relationship between GPSM2 and DYNC1I1 in breast cancer is unclear.
In this study, we identified GPSM2 as an indicator of poor prognosis in breast cancer. Importantly, nuclear expression of GPSM2 was significantly associated with even worse prognosis. In addition, subcellular localization of GPSM2 was correlated with the expression of DYNC1I1, which also indicated a poor prognosis. In summary, examination of both GPSM2 and DYNC1I1 is necessary for accurate prognosis in breast cancer patients.
Materials and Methods
Patients and Tissue Samples
This study was approved by The Human Ethics Review Committee of the China Medical University. A total of 219 primary breast cancer tissue samples were obtained for the study. These 219 patients were diagnosed with invasive breast cancer and had undergone surgery at the Shengjing hospital of China medical university. None of these patients had undergone chemotherapy or radiotherapy before surgery. The clinical criteria of patient recruitment are: (1) All patients recruited had unilateral BC and were histologically diagnosed. (2) Any patient who had distant metastasis or received preoperative radiotherapy, chemotherapy, hormonal therapy, or any other anticancer therapy before surgery was excluded. (3) Patients with serious complications, such as heart disease, cerebrovascular disease, diabetes, or other malignant diseases, were excluded. (4) Complete clinicopathological data for further analysis were available. (5) All patients were followed up through medical appointments or by telephone.
About the follow-up period of the patients, the median follow-up time of patients in this study was 34 months (range 4–73). The treatment information of all patients was obtained from medical archives. As follows, among the 176 patients with stage II–III, all patients received standard adjuvant chemotherapy; a total of 124 patients with hormone receptor-positive were treated with endocrine therapies; Trastuzumab was used in 31 of 50 HER2-positive cases. Clinical characteristics of the tissue donors, such as age, sex, age at initial diagnosis, and stage at diagnosis (tumor, node status, metastasis, and TNM classification), were obtained from medical records and pathological reports.
Immunohistochemistry (IHC) Analysis
IHC staining was performed as described previously (19). Anti-human GPSM2 rabbit antibody was used at a dilution of 1:100 (ab84571; Abcam, Cambridge, UK); Anti-human DYNC1I1 mouse antibody was used at a dilution of 1:100 (ab23905; Abcam, Cambridge, UK); phosphate-buffered saline was used as a negative control. Each section was evaluated and scored independently by two pathologists. A semi-quantitative scoring system was used in this assay. Intensity was scored as “0” (negative), “1” (weak), “2” (moderate), and “3” (strong), and the percentages of tumor cells within each category were calculated. The percentage score was multiplied by the staining intensity score to generate the IHC score. The histological score range was from zero (minimum) to 300 (maximum). Positive expression was defined as detectable immunoreactions with an IHC score > 10.
Cell Lines and Cultures
The human breast cancer cell lines MDA-MB-231 was purchased from the Cell Bank of Type Culture Collection of Chinese Academy (Shanghai, China). MDA-MB-231 cells were grown in L15 (Invitrogen), and supplemented with 10% fetal bovine serum and maintained at 37°C with 5% CO2.
Bioinformatics
The GEPIA website (20) was used to predict gene correlation in cancer. GEPIA is an online tool that provides expression analysis functions for TCGA and GTEx data.
Western Blot (WB) and Immunoprecipitation (IP) Analyses
WB and IP analyses were performed as previously described (21). The following primary antibodies were used: GPSM2 rabbit polyclonal antibody (1:1,000; Abcam, Cambridge, UK, ab84571); DYNC1I1 mouse polyclonal antibody (1:1,000; Abcam, Cambridge, UK, ab23905); actin mouse polyclonal antibody (1:250; Santa Cruz Biotechnology, sc-47778).
Statistical Analysis
Statistical analysis was performed using SPSS 13.0 statistical software (SPSS Inc., Chicago, IL, USA). Spearman's correlation coefficient was calculated to examine the association between continuous variables. The chi-square test was performed to analyses relationships between categorical variables. For the continuous variables, differences between three or more groups were assessed using one-way analysis of variance (ANOVA) with the post-hoc Tukey multiple comparison test (for normally distributed data) or Kruskal-Wallis test (for non-normal distribution). Differences between two groups were assessed using the t-test (normally distributed data) or Mann-Whitney test (non-normal distribution). Survival curves were analyzed by the Kaplan-Meier and log rank test. Cox's proportional hazard method was performed for multivariate analysis to identify the independent prognostic factors. p < 0.05 was considered significant. Relapse-free survival (RFS) was defined as the time from initial surgery to local recurrence, regional recurrence, or distant metastasis but not including disease-related death.
Results
The Expression of GPSM2 Correlates With Clinical Characteristics and Prognosis in Invasive Breast Cancer Patients
To investigate the clinical value of GPSM2 in invasive breast cancer, we performed IHC analysis of 189 breast cancer tissue samples with available long-term follow-up medical records. Representative images of each level of staining are shown in Figure 1A. The IHC score of GPSM2 staining was significantly increased along with advanced T stage (Figure 1B), lymph node metastasis (Figure 1C) and advanced TNM stage (Figure 1D). Based on IHC scores, patients were divided into two group: GPSM2-positive and GPSM2-negative. Furthermore, a chi-square test showed that high GPSM2 expression was significantly correlated with increased T stage (p = 0.029), lymph node metastasis (p = 0.002), and higher TNM stage (p < 0.001) (Table 1). GPSM2-positive patients presented with a shorter relapse-free survival (RFS) time than GPSM2-negative patients (p = 0.008) (Figure 1E). The results demonstrated that GPSM2 could be a prognosis biomarker of invasive breast cancer patients.
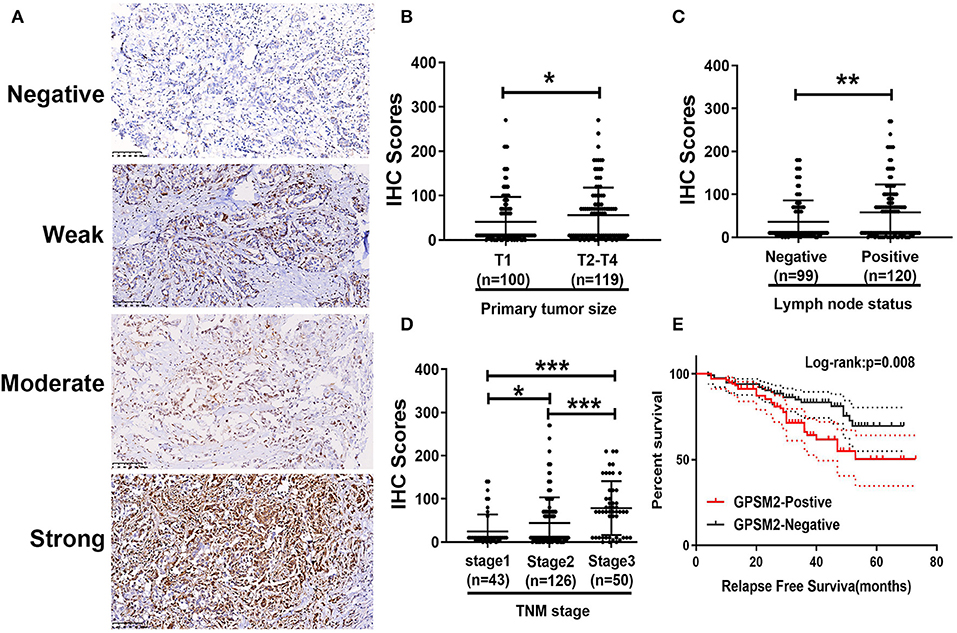
Figure 1. The expression of GPSM2 correlates with clinical features and indicates prognosis in breast patients. (A) Representative images of GPSM2 staining in human breast cancer tissue samples: “–” (negative staining), “+” (weak staining), “++” (moderate staining), and “+++” (strong staining); (B) GPSM2 IHC scores of samples with advanced tumor sizes were significantly increased; (C) GPSM2 IHC scores of samples related to positive lymph node status were significantly increased; (D) GPSM2 IHC staining scores were significantly increased with advanced TNM stage; (E) Kaplan-Meier survival analysis and log-rank test indicated that positive of GPSM2 expression was associated with poor overall survival (p = 0.008); *p < 0.05, **p < 0.01, and ***p < 0.001 (vs. control group).
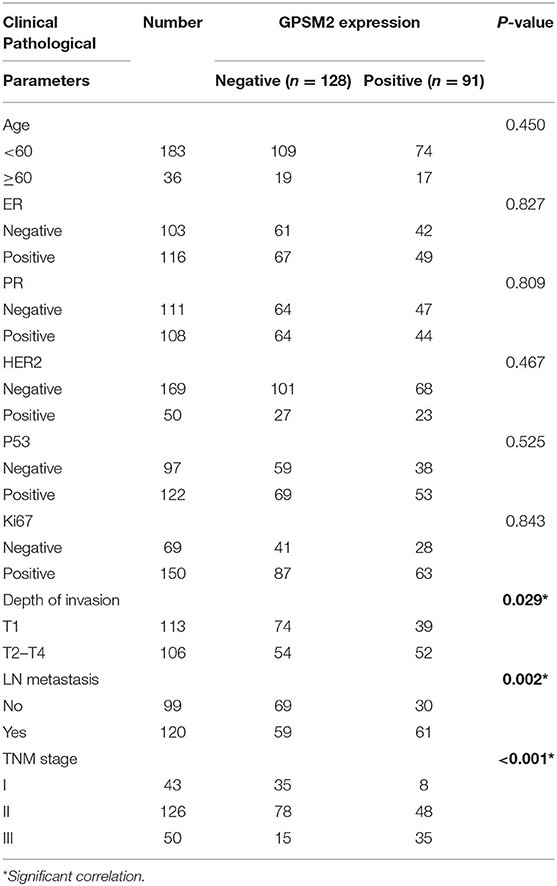
Table 1. Correlation between GPSM2 expression and clinical characteristics in breast cancer patients (n = 219).
Nucleus Localization of GPSM2 Is Significantly Correlated With Poor Clinical Outcome
We examined the subcellular localization of GPSM2 in breast cancer tissues. As shown in Figure 2A, the subcellular distribution of GPSM2 contained three forms: cytoplasm-positive, cytoplasm and nucleus-positive, and nucleus-positive. Comparing the staining intensity between nuclear and cytoplasmic 91 tumors were divided into two groups: 28 tumors were categorized as having high nuclear expression and 63 tumors were categorized as exhibiting high cytoplasmic expression. In Figure 2B, patients with GPSM2-nucleus localization presented with a shorter RFS time than GPSM2-cytoplasm patients (p = 0.001). There was no significant difference between the RFS time of the GPSM2-cytoplasm and GPSM2-negative groups (p = 0.222) (Figure 2C). In the total population, RFS of patients with GPSM2-nucleus was significantly reduced compared with that in other patients (p < 0.001) (Figure 2D). Furthermore, a chi-square test showed that compared with GPSM2-cytoplasm group, GPSM2-nucleus group was significantly correlated with HER2 receptor-positive (p = 0.010), Ki67-positive (p = 0.001), increased T stage (p = 0.001), lymph node metastasis (p = 0.015), and higher TNM stage (p = 0.029) (Table 2).
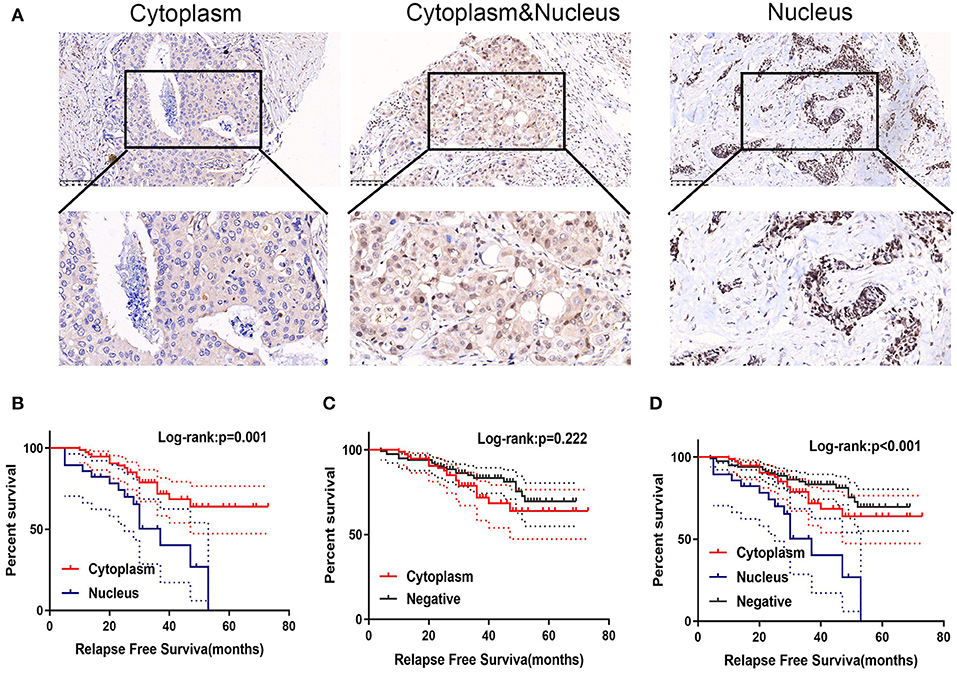
Figure 2. Positive of GPSM2 nucleus expression is significantly related with worse prognosis. (A) GPSM2 staining in different subcellular regions. (B) Kaplan-Meier survival analysis and log-rank test indicated that the GPSM2-nucleus patients presented with a shorter RFS time than GPSM2-cytoplasm patients (p = 0.001). (C) Kaplan-Meier survival analysis and log-rank test showed no significant difference between the RFS time of the GPSM2-cytoplasm group and GPSM2-negative group (p = 0.22). (D) Kaplan-Meier survival analysis and log-rank test demonstrated RFS of patients in the GPSM2-nucleus group was significantly reduced compared with that in other patients (p < 0.001).
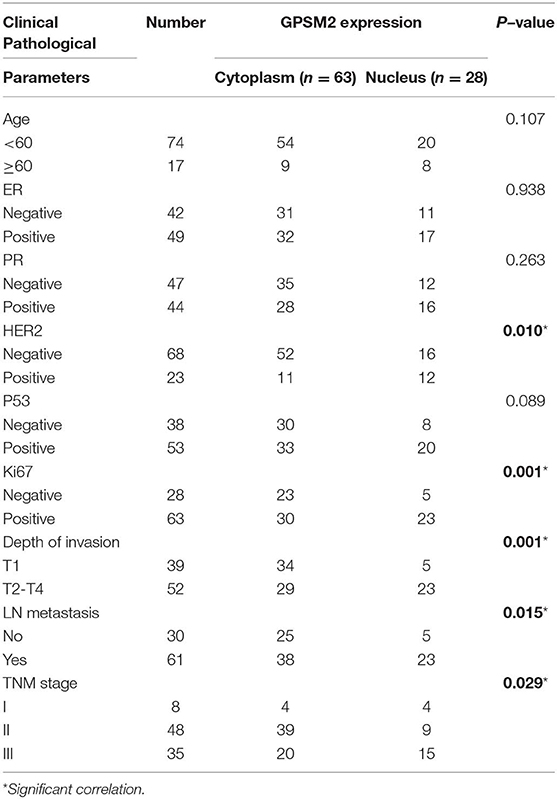
Table 2. Correlation between GPSM2 subcellular localization and clinical characteristics in breast cancer patients (n = 91).
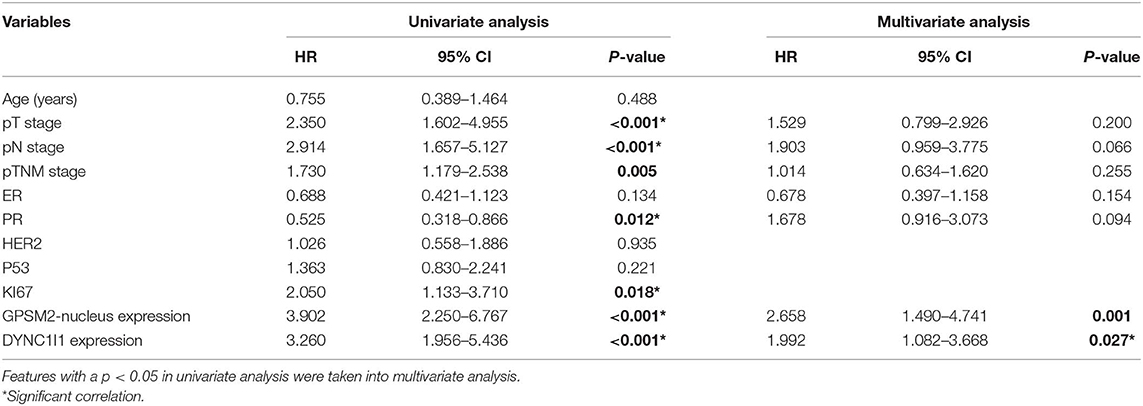
In addition, Cox proportional hazards model was used to analyze the impact of clinical and pathological parameters on the prognosis of patients. Univariate analysis showed that T stage (HR = 2.350, 95% confidence interval [CI] = 1.602–4.955, P < 0.001), N stage (HR = 2.914, 95% CI = 1.657–5.127, P < 0.001), TNM stage (HR = 1.730, 95% CI = 1.179–2.538, P = 0.005), PR status (HR = 0.525, 95% CI = 0.318–0.866, P = 0.012), KI67 status (HR = 2.050, 95% CI = 1.133–3.710, P = 0.018) and GPSM2-nucleus expression (HR = 3.902, 95% CI = 2.250–6.767, P < 0.001) were risk factors for prognosis. In additional, Age(HR = 0.755, 95% CI = 0.389–1.464, P = 0.488), ER status (HR = 0.688, 95% CI = 0.421–1.123, P = 0.488), HER2 status (HR = 1.026, 95% CI = 0.558–1.886, P = 0.935), P53 status (HR = 1.363, 95% CI = 0.830–2.241, P = 0.221) were not showed significant effect on prognosis. Furthermore, multivariate analysis showed that GPSM2-nucleus expression (HR = 2.658, 95% CI = 1.490–4.741, P = 0.001) were independent risk factors for prognosis. Meanwhile, T stage (HR = 1.529, 95% CI =0.799–2.926, P = 0.200), N stage (HR = 1.903, 95% CI = 0.959–3.775, P = 0.066), TNM stage (HR = 1.014, 95% CI = 0.634–1.620, P = 0.255), PR status (HR = 0.678, 95% CI = 0.397–1.158, P = 0.154), KI67 status (HR =1.678, 95% CI =0.916–3.073, P = 0.094) are not independent risk factors for prognosis (Table 3). These results indicated the prognosis value of nuclear localization of GPSM2 in breast cancer patients.
GPSM2 and DYNC1I1 Form a Complex in Breast Cancer Cells
Next, we examined the mechanisms of GPSM2 subcellular localization. DYNC1I1 is an important cargo binding subunit of cytoplasmic dynein. So, we explored the relationship of DYNC1I1 and GPSM2. First, we used the GEPIA Website to analyze the correlation between GPSM2 and DYNC1I1. In Figure 3A, GPSM2 was positively correlated with DYNC1I1 in several cancers, such as bladder urothelial carcinoma (BLCA, R = 0.16, p = 0.0016), breast invasive carcinoma (BRCA, R = 0.28, p = 0), glioblastoma multiforme (GBM, R = 0.27, p < 0.001) and prostate adenocarcinoma (PRAD, R = 0.33, p < 0.001). Next, we performed immunoprecipitation analysis using the breast cancer cell line MDA-MB-231. As shown in Figure 3B, anti-GPSM2 antibodies efficiently immunoprecipitated DYNC1I1. Further, anti-DYNC1I1 antibodies also efficiently immunoprecipitated GPSM2. These results indicated that GPSM2 and DYNC1I1 formed a complex in breast cancer cells and may play an important role in BRCA patients.
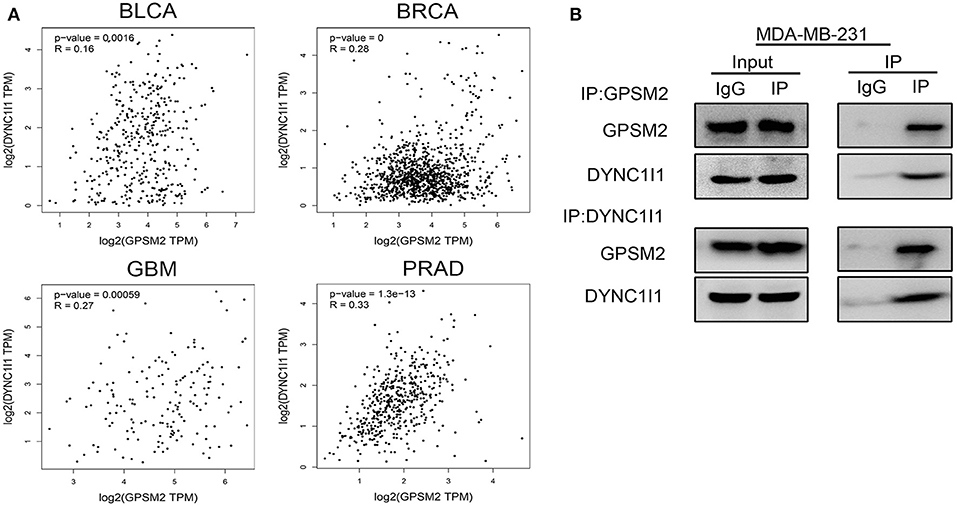
Figure 3. GPSM2 interacts with DYNC1I1 in breast cancer cells. (A) GEPIA website analysis the correlation between GPSM2 and DYNC1I1; (B) Immunoprecipitation assay was performed to detect the interaction between GPSM2 and DYNC1I1 in breast cancer MDA-MB-231 cells.
The Expression of DYNC1I1 Is Correlated With Clinical Characteristics and Prognosis in Invasive Breast Cancer
Next, in order to investigate the clinical value of DYNC1I1 in invasive breast cancer, we performed IHC analysis. Representative images of each level of DYNC1I1 staining are shown in Figure 4A. There were no significant differences between the expression of DYNC1I1 in different T stages (Figure 4B). Moreover, the IHC score of DYNC1I1 staining was significantly increased along with lymph node metastasis (Figure 4C) and advanced TNM stage (Figure 4D). According to IHC scores, patients were divided into two groups: DYNC1I1-positive and DYNC1I1-negative. Furthermore, a chi-square test showed that high DYNC1I1 expression was significantly correlated with ages (p = 0.042), PR-negative (p = 0.041), increased T stage (p < 0.001), lymph node metastasis (p = 0.007), and higher TNM stage (p < 0.001) (Table 4). Moreover, DYNC1I1-positive patients presented with a shorter RFS time than GPSM2-negative patients (p < 0.001) (Figure 4E). Additionally, the DYNC1I1-positive group exhibited a shorter RFS rate (HR = 1.992, 95% CI = 1.082–3.668, p = 0.027) (Table 3). The results demonstrated that DYNC1I1 is an independent risk factor, and may be a biomarker for the prognosis of invasive breast cancer patients.
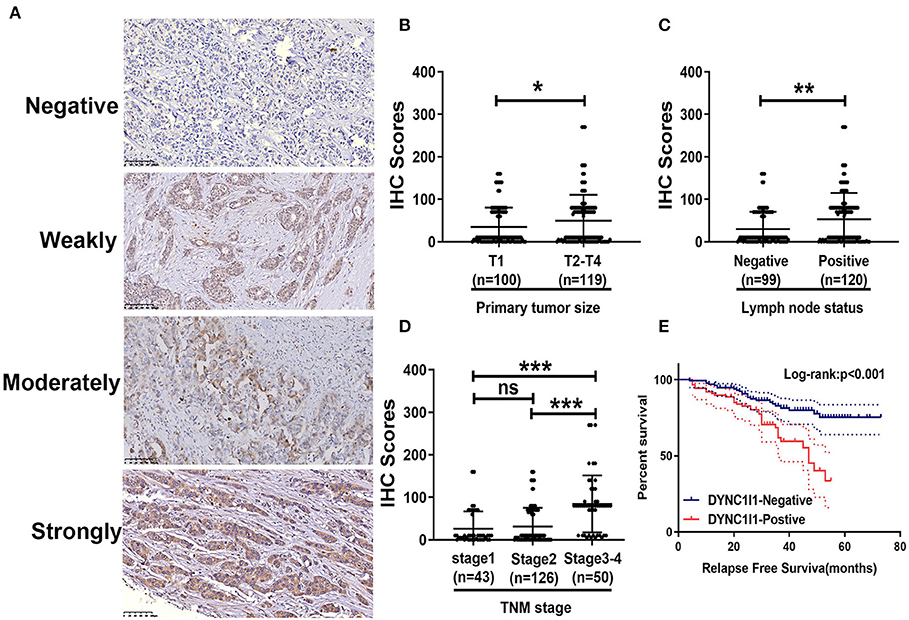
Figure 4. The expression of DYNC1I1 correlates with clinical characteristics and poor prognosis in breast cancer patients. (A) Representative images of DYNC1I1 staining in human breast cancer tissue samples: “–” (negative staining), “+” (weak staining), “++” (moderate staining), and “+++” (strong staining); (B) The IHC score of DYNC1I1 did not differ significantly between different tumor sizes; (C) DYNC1I1 IHC scores of samples related to positive lymph node status were significantly increased; (D) DYNC1I1 IHC staining scores were significantly increased with advanced TNM stage; (E) Kaplan-Meier survival analysis and log-rank test indicated that positive DYNC1I1 expression was associated with poor overall survival (p < 0.001); *p < 0.05, **p < 0.01, and ***p < 0.001 (vs. control group).
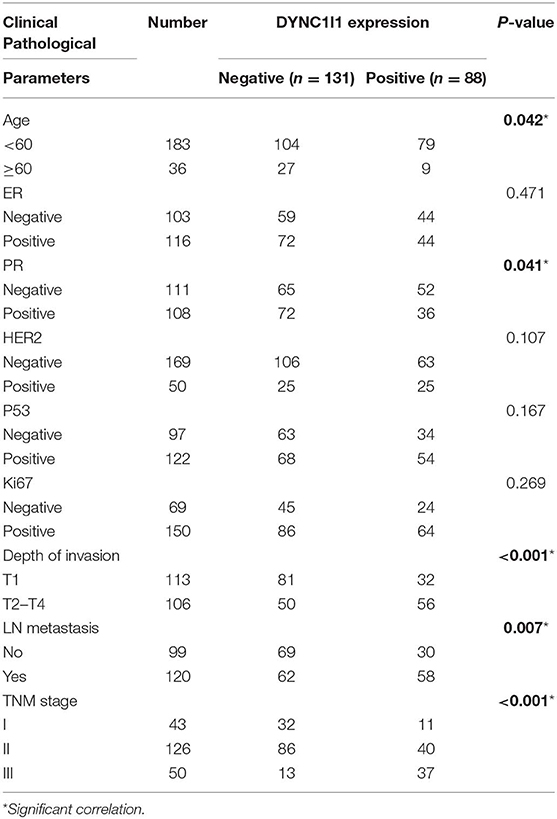
Table 4. Correlation between DYNC1I1 expression and clinical characteristics in breast cancer patients (n = 219).
Correlation Between GPSM2 and DYNC1I1 and Prognosis in Invasive Breast Cancer Patients
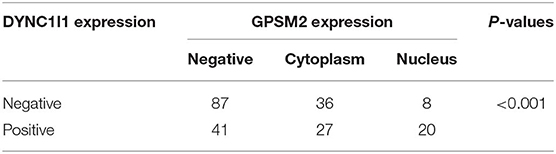
Due to the clinical value of GPSM2 and DYNC1I1 in BRCA and importance of the GPSM2-DYNC1I1 complex, we aimed to explore the relationship between GPSM2 and DYNC1I1 in BRCA patients. As shown in Figure 5A, the expression of GPSM2 was positively correlated with that of DYNC1I1 (R = 0.367, p < 0.001). In addition, the RFS for patients exhibiting positive expression of both DYNC1I1 and GPSM2 was significantly reduced compared to that of the other patients (p < 0.001). GPSM2/DYNC1I1 expression is shown in Figure 5C. Our results showed the prognosis value of the GPSM2-DYNC1I1 complex.
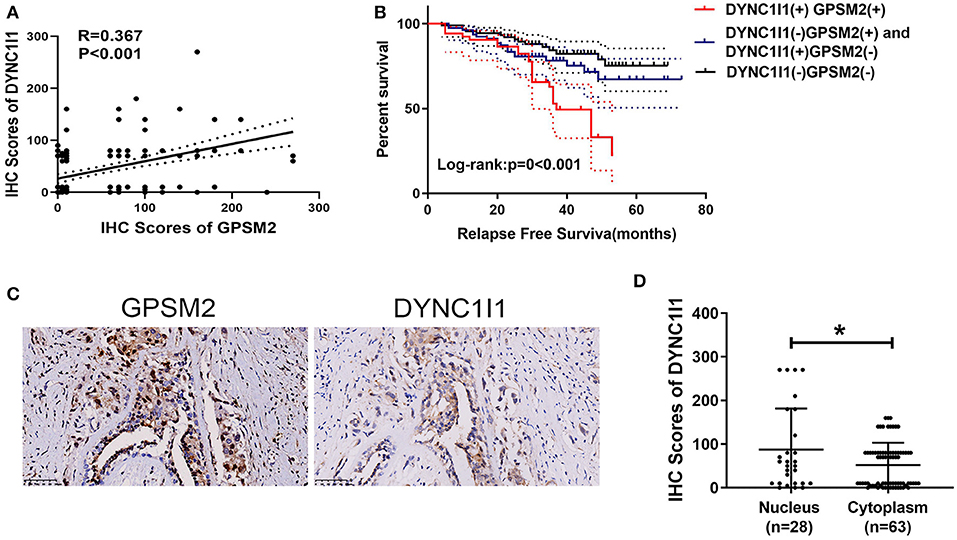
Figure 5. Correlations between GPSM2 expression and DYNC1I1 in breast cancer patients. (A) The IHC scores of GPSM2 is positively correlated with the IHC scores of DYNC1I1 (R = 0.367, p < 0.001); (B) Kaplan-Meier survival analysis and log-rank test indicated that the RFS of patients exhibiting expression of both DYNC1I1 and GPSM2 was significantly reduced compared to other patients (p < 0.001); (C) GPSM2 and DYNC1I1 expression; (D) IHC scores of DYNC1I1 in the GPSM2-nucleus group were significantly higher than the GPSM2-cytoplasm group (p < 0.05); *p < 0.05 (vs. control group).
Nuclear Localization of GPSM2 Was Correlated With Positive Expression of DYNC1I1
Next, we explored the relationship between nuclear localization of GPSM2 and DYNC1I1 in invasive breast cancer. As shown in Figure 5, the IHC scores of DYNC1I1 in the GPSM2-nucleus group were higher than those of the GPSM2-cytoplasm group (p < 0.05). Furthermore, a chi-square test showed that nuclear localization of GPSM2 was correlated with positive expression of DYNC1I1 (Table 5).
Discussion
In the past three decades, the incidence of breast cancer in China has been rising, and has become the primary malignant tumor in females (22, 23). In order to increase both RFP and overall survival in breast cancer patients, investigation of drug targets and prognosis biomarkers is essential. Previous study identified an interaction of GPSM2 with G alpha proteins. Subsequent research showed that GPSM2 was involved in the regulation of asymmetric cell division and spindle orientation. Additionally, several studies indicated that GPSM2 is essential to the development of normal hearing (24). GPSM2 mutations are known to cause brain malformations and hearing loss in Chudley-McCullough syndrome (25–27). Embryonic development and tumorigenesis have similar molecular mechanisms, and thus, GPSM2 may play an important role in both.
A previous study reported that GPSM2 was overexpressed in hepatocellular carcinoma and related to poor prognosis of these patients (11). Another study in pancreatic cancer showed GPSM2 promoted the proliferation and migration ability of CD133+ pancreatic cancer stem cells (12). In addition, one study suggested GPSM2 was involved in the cell division of breast cancer cells and was regulated by PBK/TOPK (13). However, the clinical and prognosis value of GPSM2 in breast cancer has not been elucidated. Based our IHC results, we are the first to report that the expression of GPSM2 was correlated with patient prognosis and clinicopathological factors. Importantly, our results indicated GPSM2 nuclear expression was found in patients with the shortest RFS. In additional, other pathology characteristics (such as, hormone receptor negativity, HER2 status, high Ki-67 proliferation index, pT stage) did not shown affect survival in this study according the result of multivariate COX analysis. Several reasons may exist for this discrepancy, including the number of samples was relatively small, heterogenous of patients' treatments. These issues need to be addressed in future studies evaluating a larger sample population. Taken together, we report here the clinical value of GPSM2, especially GPSM2 nuclear expression, in breast cancer.
Another important finding of our study was that subcellular localization of GPSM2 was correlated with the expression of DYNC1I1, which indicated a poor prognosis in breast cancer. DYNC1I1 is the intermediate chain of cytoplasmic dynein, and is an important cargo binding subunit (15). Previous studies indicated that loss of DYNC1I1 caused split hand/split foot malformation type I (28, 29). However, the role it plays in cancer remains controversial. Depending on its transport cargo, DYNC1I1 can act as a promotion or suppression factor in tumors (16, 17). Several studies showed subcellular distribution of GPSM2 was regulated by dynein (9, 18). However, the relationship between GPSM2 and DYNC1I1 has not been fully elucidated. In our study, we demonstrated by immunoprecipitation that GPSM2 and DYNC1I1 could form a complex in breast cancer cells. Moreover, the expression of DYNC1I1 was correlated with patient prognosis and clinicopathological factors. Importantly, patients with GPSM2 nuclear expression showed higher DYNC1I1 expression. These results suggested DYNC1I1 was an indicator of poor prognosis that may function via transfer of GPSM2 into the nucleus. Examination of both GPSM2 and DYNC1I1 could help to assess patient prognosis more effectively.
Given the appreciation of their role in cancer, the importance of GPCRs for anticancer drug discovery is undisputable, although very few members have been exploited in pursuit of anticancer therapies. In this study, we have confirmed the clinical value of GPSM2.Thus, GPSM2 may be a promising target for new cancer therapy through regulating GPCRs pathway (6, 30). Due the value of GPSM2 in predicting disease recurrence, it would be better to add GPMS2 to the NGS panels like Oncotype Dx (31). Including GPSM2 to Oncotype DX gene panel may have considerable benefit in predicting disease recurrence.
However, the mechanisms of why GPSM2 nucleus-positive patients presented with poorer prognoses are unclear. We speculate that this is mainly because nuclear localization of GPSM2 is unavailable to inhibit activity of Gα protein that is expressed on the membrane, thereby enhancing the activity of the GPCR pathway and promoting tumor progression. Further studies are needed to determine whether the association between GPSM2 and DYNC1I1 is direct or indirect. We plan to further investigate these questions with additional in vitro and in vivo experiments. There are two limitations in this study. Firstly, the samples involved in this study were acquired retrospectively; secondly, the number of samples was relatively small. Thus, it needs further comprehensive and in-depth prospective study with enough samples to demonstrate that nucleus expression of GPSM2 is associated with high rate of recurrence in breast cancer patients.
In conclusion, our results strongly suggested that GPSM2 was an independent prognostic factor in breast cancer. Importantly, nuclear expression of GPSM2 was significantly associated with even worse prognoses, and was related to positive expression of DYNC1I1. We believe that assessment of the combination of GPSM2 and DYNC1I1 expression could be a promising biomarker and drug target for breast cancer.
Data Availability Statement
The datasets generated for this study can be found in the GEPIA website (20).
Ethics Statement
The studies involving human participants were reviewed and approved by the Research Ethics Committee of China Medical University. The patients/participants provided their written informed consent to participate in this study.
Author Contributions
QL and YZ contributed conception and design of the study. KH, XQ, YL, and XC gave the technical support. MD and ZZ performed the experiment and statistical analysis. MD wrote the manuscript. QL provided fund supports. All authors contributed to manuscript revision, read and approved the submitted version.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This work was supported by Shenyang Special Funds of Technology Innovation Plan Projects (No. F14-231-1-46), Liaoning Science and Technology Plan Projects (No. 2013225021), Liaoning Natural Science Foundation of 2018 (No. 20180530050).
References
1. Perez-Garcia J, Cortes J. Breast cancer in 2017: spurring science, marking progress, and influencing history. Nat Rev Clin Oncol. (2018) 15:79–80. doi: 10.1038/nrclinonc.2017.191
2. Pondé NF, Zardavas D, Piccart M. Progress in adjuvant systemic therapy for breast cancer. Nat Rev Clin Oncol. (2019) 16:27–44. doi: 10.1038/s41571-018-0089-9
3. Campbell AP, Smrcka AV. Targeting G protein-coupled receptor signalling by blocking G proteins. Nat Rev Drug Discov. (2018) 17:789–803. doi: 10.1038/nrd.2018.135
4. Dunn HA, Orlandi C, Martemyanov KA. Beyond the ligand: extracellular and transcellular G protein-coupled receptor complexes in physiology and pharmacology. Pharmacol Rev. (2019) 71:503–19. doi: 10.1124/pr.119.018044
5. Arakaki AKS, Pan WA, Trejo J. GPCRs in cancer: protease-activated receptors, endocytic adaptors and signaling. Int J Mol Sci. (2018) 19:1886–910. doi: 10.3390/ijms19071886
6. Lappano R, Jacquot Y, Maggiolini M. GPCR modulation in breast cancer. Int J Mol Sci. (2018) 19:3840–76. doi: 10.3390/ijms19123840
7. Mochizuki N, Cho G, Wen B, Insel PA. Identification and cDNA cloning of a novel human mosaic protein, LGN, based on interaction with G alpha i2. Gene. (1996) 181:39–43. doi: 10.1016/S0378-1119(96)00456-8
8. Jeon JP, Thakur DP, Tian JB, So I, Zhu MX. Regulator of G-protein signalling and GoLoco proteins suppress TRPC4 channel function via acting at Gαi/o. Biochem J. (2016) 473:1379–90. doi: 10.1042/BCJ20160214
9. Woodard GE, Huang NN, Cho H, Miki T, Tall GG, Kehrl JH. Ric-8A and Gi alpha recruit LGN, NuMA, and dynein to the cell cortex to help orient the mitotic spindle. Mol Cell Biol. (2010) 30:3519–30. doi: 10.1128/MCB.00394-10
10. Kiyomitsu T, Cheeseman IM. Chromosome- and spindle-pole-derived signals generate an intrinsic code for spindle position and orientation. Nat Cell Biol. (2012) 14:311–7. doi: 10.1038/ncb2440
11. He XQ, Zhang YF, Yu JJ, Gan YY, Han NN, Zhang MX, et al. High expression of G-protein signaling modulator 2 in hepatocellular carcinoma facilitates tumor growth and metastasis by activating the PI3K/AKT signaling pathway. Tumour Biol. (2017) 39:1010428317695971. doi: 10.1177/1010428317695971
12. Dang SC, Qian XB, Jin W, Cui L, Chen JX, Gu M. G-protein-signaling modulator 2 expression and role in a CD133 pancreatic cancer stem cell subset. Onco Targets Ther. (2019) 12:785–94. doi: 10.2147/OTT.S187670
13. Fukukawa C, Ueda K, Nishidate T, Katagiri T, Nakamura Y. Critical roles of LGN/GPSM2 phosphorylation by PBK/TOPK in cell division of breast cancer cells. Genes Chromosomes Cancer. (2010) 49:861–72. doi: 10.1002/gcc.20795
14. Reck-Peterson SL, Redwine WB, Vale RD, Carter AP. The cytoplasmic dynein transport machinery and its many cargoes. Nat Rev Mol Cell Biol. (2018) 19:382–98. doi: 10.1038/s41580-018-0004-3
15. Kuta A, Deng W, Morsi El-Kadi A, Banks GT, Hafezparast M, Pfister KK, et al. Mouse cytoplasmic dynein intermediate chains: identification of new isoforms, alternative splicing and tissue distribution of transcripts. PLoS ONE. (2010) 5:e11682. doi: 10.1371/annotation/59badad8-6e55-46f8-8bf1-7a8a957bc68e
16. Neubauer HA, Tea MN, Zebol JR, Gliddon BL, Stefanidis C, Moretti PAB, et al. Cytoplasmic dynein regulates the subcellular localization of sphingosine kinase 2 to elicit tumor-suppressive functions in glioblastoma. Oncogene. (2019) 38:1151–65. doi: 10.1038/s41388-018-0504-9
17. Gong LB, Wen T, Li Z, Xin X, Che XF, Wang J, et al. DYNC1I1 promotes the proliferation and migration of gastric cancer by up-regulating IL-6 expression. Front Oncol. (2019) 9:491. doi: 10.3389/fonc.2019.00491
18. Zheng Z, Wan Q, Liu J, Zhu H, Chu X, Du Q. Evidence for dynein and astral microtubule-mediated cortical release and transport of Gαi/LGN/NuMA complex in mitotic cells. Mol Biol Cell. (2013) 24:901–13. doi: 10.1091/mbc.e12-06-0458
19. Deng M, Yu R, Wang S, Zhang Y, Li Z, Song H, et al. Limb-bud and heart attenuates growth and invasion of human lung adenocarcinoma cells and predicts survival outcome. Cell Physiol Biochem. (2018) 47:223–34. doi: 10.1159/000489801
20. Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. (2017) 45:W98–102. doi: 10.1093/nar/gkx247
21. Deng M, Zhang Y, Liu B, Chen Y, Song H, Yu R, et al. β-Elemene inhibits peritoneal metastasis of gastric cancer cells by modulating FAK/Claudin-1 signaling. Phytother Res. (2019) 33:2448–56. doi: 10.1002/ptr.6436
22. Li J, Zhang BN, Fan JH, Pang Y, Zhang P, Wang SL, et al. A nation-wide multicenter 10-year (1999–2008) retrospective clinical epidemiological study of female breast cancer in China. BMC Cancer. (2011) 11:364. doi: 10.1186/1471-2407-11-364
23. Lee H, Li JY, Fan JH, Li J, Huang R, Zhang BN, et al. Risk factors for breast cancer among Chinese women: a 10-year nationwide multicenter cross-sectional study. J Epidemiol. (2014) 24:67–76. doi: 10.2188/jea.JE20120217
24. Walsh T, Shahin H, Elkan-Miller T, Lee MK, Thornton AM, Roeb W, et al. Whole exome sequencing and homozygosity mapping identify mutation in the cell polarity protein GPSM2 as the cause of non-syndromic hearing loss DFNB82. Am J Hum Genet. (2010) 87:90–4. doi: 10.1016/j.ajhg.2010.05.010
25. Diaz-Horta O, Sirmaci A, Doherty D, Nance W, Arnos K, Pandya A, et al. GPSM2 mutations in Chudley-McCullough syndrome. Am J Med Genet A. (2012) 158:2972–3. doi: 10.1002/ajmg.a.35636
26. Doherty D, Chudley AE, Coghlan G, Ishak GE, Innes AM, Lemire EG, et al. GPSM2 mutations cause the brain malformations and hearing loss in Chudley-McCullough syndrome. Am J Hum Genet. 90:1088–93. doi: 10.1016/j.ajhg.2012.04.008
27. Almomani R, Sun Y, Aten E, Hilhorst-Hofstee Y, Peeters-Scholte CM, van Haeringen A, et al. GPSM2 and Chudley-McCullough syndrome: a Dutch founder variant brought to North America. Am J Med Genet A. (2013) 161:973–6. doi: 10.1002/ajmg.a.35808
28. Lango Allen H, Caswell R, Xie W, Xu X, Wragg C, Turnpenny PD, et al. Next generation sequencing of chromosomal rearrangements in patients with split-hand/split-foot malformation provides evidence for DYNC1I1 exonic enhancers of DLX5/6 expression in humans. J Med Genet. (2014) 51:264–7. doi: 10.1136/jmedgenet-2013-102142
29. Tayebi N, Jamsheer A, Flöttmann R, Sowinska-Seidler A, Doelken SC, Oehl-Jaschkowitz B, et al. Deletions of exons with regulatory activity at the DYNC1I1 locus are associated with split-hand/split-foot malformation: array CGH screening of 134 unrelated families. Orphanet J Rare Dis. (2014) 9:108. doi: 10.1186/s13023-014-0108-6
30. Singh A, Nunes JJ, Ateeq B. Role and therapeutic potential of G-protein coupled receptors in breast cancer progression and metastases. Eur J Pharmacol. (2015) 763:178–83. doi: 10.1016/j.ejphar.2015.05.011
31. Mamounas EP, Tang G, Fisher B, Paik S, Shak S, Costantino JP, et al. Association between the 21-gene recurrence score assay and risk of locoregional recurrence in node-negative, estrogen receptor-positive breast cancer: results from NSABP B-14 and NSABP B-20. J Clin Oncol. (2010) 28:1677–83. doi: 10.1200/JCO.2009.23.7610
Keywords: GPSM2, DYNC1I1, breast cancer, prognosis, subcellular localization
Citation: Deng M, Zhang Z, Liu B, Hou K, Che X, Qu X, Liu Y, Hu X, Zhang Y and Lv Q (2020) Localization of GPSM2 in the Nucleus of Invasive Breast Cancer Cells Indicates a Poor Prognosis. Front. Oncol. 10:227. doi: 10.3389/fonc.2020.00227
Received: 08 November 2019; Accepted: 10 February 2020;
Published: 03 March 2020.
Edited by:
Takayuki Ueno, The Cancer Institute Hospital (JFCR), JapanCopyright © 2020 Deng, Zhang, Liu, Hou, Che, Qu, Liu, Hu, Zhang and Lv. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Ye Zhang, eXpoYW5nMjFAY211LmVkdS5jbg==; Qingjie Lv, Y211X2x2cWluZ2ppZUAxMjYuY29t
†These authors share first authorship
‡These authors share senior authorship
 Mingming Deng1,2,3†
Mingming Deng1,2,3† Xiaofang Che
Xiaofang Che Xiujuan Qu
Xiujuan Qu Yunpeng Liu
Yunpeng Liu Ye Zhang
Ye Zhang Qingjie Lv
Qingjie Lv