- 1Department of Pathology, Zhejiang Rongjun Hospital, Jiaxing, China
- 2Department of Tumor Molecular Laboratory, Zhejiang Rongjun Hospital, Jiaxing, China
- 3Department of Medical Thoracic Oncology, Fujian Cancer Hospital, Fujian Medical University Cancer Hospital, Fuzhou, China
- 4Department of Chemotherapy, Zhejiang Cancer Hospital, Hangzhou, China
- 5Department of Thoracic Disease Diagnosis and Treatment Center, Zhejiang Rongjun Hospital, Jiaxing, China
- 6Department of Pathology, Fujian Cancer Hospital, Fujian Medical University Cancer Hospital, Fuzhou, China
Introduction: Oncogenic mutations in the epidermal growth factor receptor (EGFR) occur frequently in patients with lung cancer. These mutations may serve as critical predictive biomarkers in patients with non-small cell lung cancer (NSCLC). Among them, EGFR exon 18–25 kinase domain duplication (EGFR-KDD) mutations have been identified as a novel EGFR gene subtype in NSCLC.
Case Presentation: We reported a rare case of a 59-year-old male diagnosed with adenocarcinoma. A biopsy revealed an EGFR-KDD identified by the next generation sequencing (NGS). Effective treatment outcome has been observed after administration with afatinib.
Conclusion: This case highlights that comprehensive NGS technique is valuable in detecting novel genetic mutations in tumors.
Introduction
Non-small cell lung cancer (NSCLC) is the leading cause of cancer-related death, with adenocarcinoma being one of the most common forms (1). Epidermal growth factor receptor (EGFR) mutations could be detected in 30–60% of Asian patients and 10–20% of Caucasian patients with lung cancer (2). Being as a driver oncogene, double-blinded randomized clinical trials have indicated that application of EGFR tyrosine kinase inhibitors (TKIs) are effective against NSCLC cases harboring EGFR mutations (3).
EGFR mutations most commonly occur in exon 19 or exon 21 within the EGFR tyrosine kinase domain. The rare EGFR mutations are usually not detected by the first-generation testing techniques. However, advanced precise detection techniques (e.g., next generation sequencing, NGS) enable the discovery of more rare EGFR mutations, including exons 18–25 kinase domain duplications (KDDs) (4). Herein, we reported a first case of an oncogenic EGFR-KDD in lung adenocarcinoma who were responsive to treatment with afatinib in Chinese populations.
Case Presentation
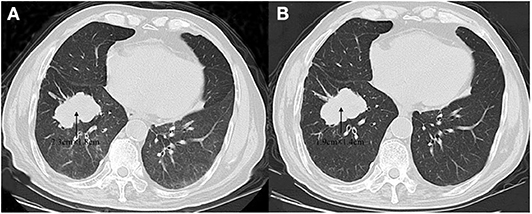
A 59-year-old male was referred to our hospital for detection of a right lung mass on physical examination. He had no history of pulmonary disease or smoking. A mass (2.3 × 1.8 cm) in the right lower lung was observed by computed tomography (CT) scan (Figure 1A). F-18 fluorodeoxyglucose hypermetabolic speckles in fourth vertebral body; no hypermetabolic lesions were demonstrated in other sites, and a MRI of the brain or CT of the head with IV contrast was not performed. We detected a typical morphology for adenocarcinoma cells by hematoxylin and eosin (H&E) staining (Figure 2). Immunohistochemical staining showed positive for the expression of NapsinA, TTF-1, and CK7. The patient was classified as stage IV (T1N0M1), in accordance to the 7th edition of TNM staging. The ARMS assay, the first-generation sequencing technique, revealed wild-type for sensitive EGFR mutations, including EGFR 18-21, and negative for ALK rearrangement or ROS1 rearrangement. Then, a NGS analysis of the tumor biopsy identified a EGFR-KDD mutation (copy number 2.0) accompanied TP53 p.R282W (frequency 13.0%) and CTNNB1 p.S37Y (frequency 5.1%) in the tumor (Geneplus, Beijing, China) (Figure 3), and PD-L1 staining was not done. Therefore, he underwent oral afatinib treatment (30 mg qd). Afterwards, the patient showed a stable tumor response to afatinib (1.9 × 1.4 cm) (Figure 1B). Besides, there were no adverse events, including gastrointestinal reactions, hepatic and renal function, and cardiac damage. Currently, the disease is stable and treatment with afatinib continues for 10 months.
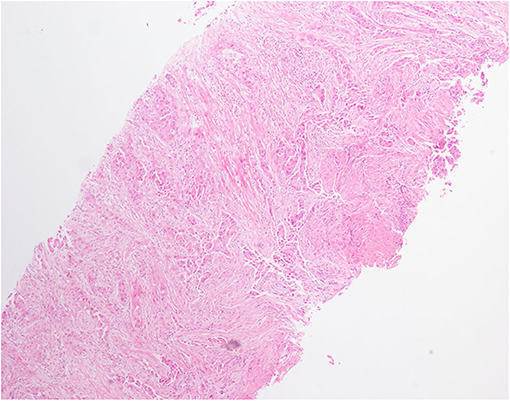
Figure 2. Hematoxylin and eosin (H&E) staining of lung biopsy showed a typical morphology for adenocarcinoma cells (H&E × 100).
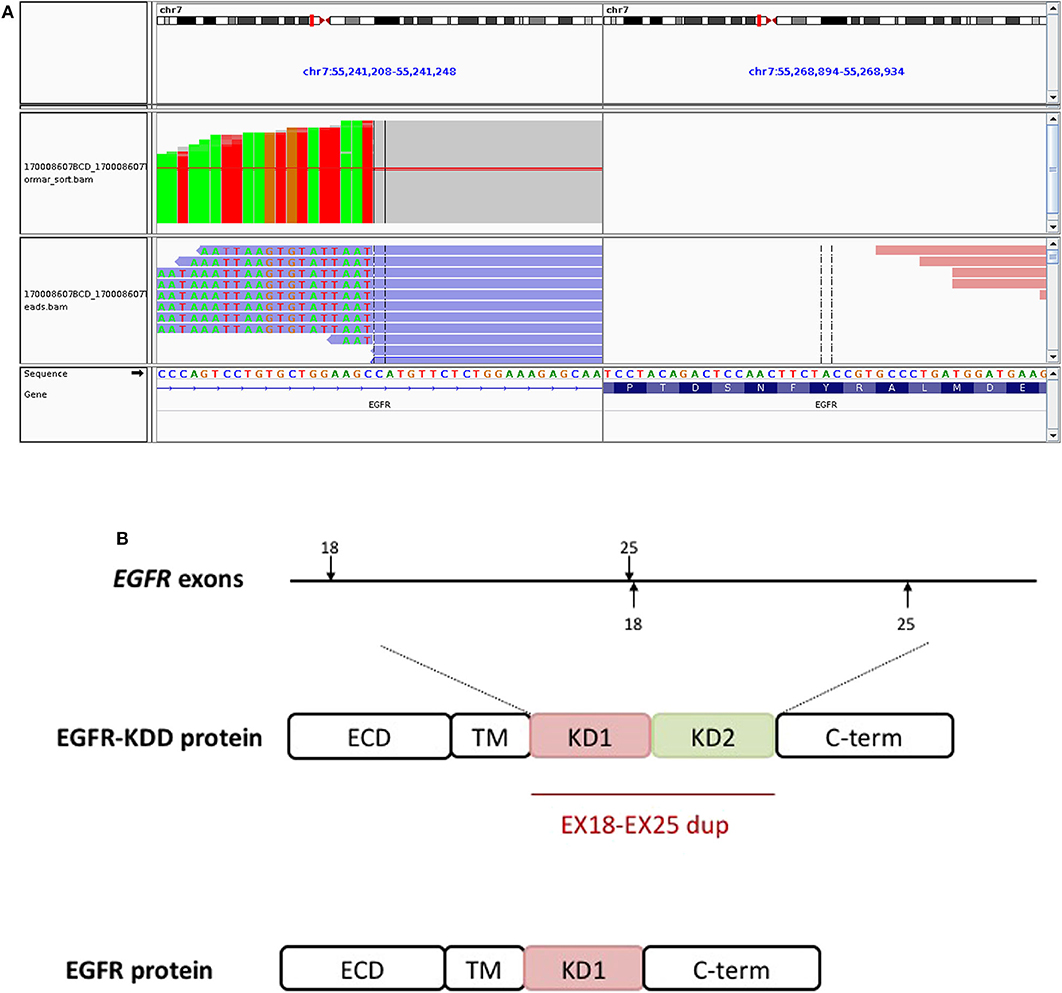
Figure 3. The EGFR-KDD is an oncogenic EGFR alteration. (A) The Integrative Genomics Viewer snapshot of paired NGS reads of tumor and matched blood. (B) Schematic representation of EGFR-KDD depicting the genetic and protein domain structures. ECD, extracellular domain; TM, transmembrane domain; KD1, first kinase domain; KD2, second kinase domain; C-term, carboxyl terminus. Blue, EGFR exons 18–25 #1; orange, EGFR exons 18–25 #2.
Discussion
Oncogenic EGFR mutations are detected in 30–60% of Asian patients and 10–20% of Caucasian patients. Such mutations are most detected as small in-frame deletions in exon 19 or point mutations in exon 21. Uncommon EGFR alterations, including rare point mutations and gene rearrangements, have also been reported previously (5).
EGFR-KDDs could activate EGFR signaling by forming an intra-molecule dimer (6). EGFR-KDD of exons 18–25, firstly discovered in a glioblastoma (7), has been recognized as a driver gene in lung cancer. In 2015, Baik et al. (8) first reported an EGFR-KDD in a female patient with a bronchoalveolar carcinoma and responsive to the first-generation EGFR-TKIs, including erlotinib and gefitinib. Another case reported a male patient with lung adenocarcinoma and an EGFR-KDD mutation detected with NGS, who had a preferable anti-tumor response to afatinib, a second-generation EGFR-TKI (6). Zhu et al. (9) reported the first case involving the presence of oncogenic EGFR-KDD in China, who had stable disease to treatment with an EGFR-TKI icotinib. Another case report of a prolonged multi-year response to gefitinib and then erlotinib has been described for advanced EGFR-KDD mutated lung adenocarcinoma (8). Therefore, it seems these EGFR variants are sensitive to first- and second- generation EGFR-TKIs. Consistently, in vitro study showed that EGFR-KDD is constitutively active, and computational modeling provides potential mechanistic support for its auto-activation (9). Herein, we for the first time detected EGFR-KDD in a Chinese patient who achieved sustained anti-tumor responses from treatment with afatinib.
Polymerase chain reaction (PCR) is frequently applied for detection of common EGFR variants in NSCLC patients, which is unable to identify some rare types of EGFR alterations (10). By contrast, NGS allows for multiplex testing and enables the detection of known as well as uncommon genomic events, as reported in this case (11). Thus, clinical treatment should improve with clinical diagnostics for multiple gene testing to provide personalized cancer therapy.
In summary, the present case increases the evidence supporting afatinib treatment of NSCLC patients harboring EGFR-KDD variants. The NGS assay provides a useful way to identify rare and uncommon EGFR gene mutations in NSCLC patients.
Ethics Statement
This study was approved by the Ethic Committee of Zhejiang Rongjun Hospital. The patient provided written informed consent for the publication of this case report.
Author Contributions
DC, XL, and BW conceptualized and designed the entire study. XZ, HC, and MF carried out patient clinical management and sample collection. WW analyzed the data. WW, YD, and CX wrote the manuscript. WW and CX revised the manuscript. All authors read and approved the final version of manuscript for submission.
Funding
This study was supported in part by grants from the Medical Scientific Research Foundation of Zhejiang Province of China (2019RC027), Science and Technology Bureau Project of Jiaxing (2017BY18050, 2019AD32266), Scientific Research Foundation of Zhejiang Medical Association (2019ZYC-A76), and Xisike-Hanson Cancer Research Foundation (Y-HS2019-20).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. (2019) 69:7–34. doi: 10.3322/caac.21551
2. Ghafoor Q, Baijal S, Taniere P, O'Sullivan B, Evans M, Middleton G. Epidermal growth factor receptor (EGFR) kinase inhibitors and non-small cell lung cancer (NSCLC) - advances in molecular diagnostic techniques to facilitate targeted therapy. Pathol Oncol Res. (2018) 24:723–31. doi: 10.1007/s12253-017-0377-1
3. Le T, Gerber DE. Newer-generation EGFR inhibitors in lung cancer: how are they best used? Cancers. (2019) 3:366. doi: 10.3390/cancers11030366
4. Costa DB. Kinase inhibitor-responsive genotypes in EGFR mutated lung adenocarcinomas: moving past common point mutations or indels into uncommon kinase domain duplications and rearrangements. Transl Lung Cancer Res. (2016) 5:331–7. doi: 10.21037/tlcr.2016.06.04
5. Milano GA. Targeted therapy in non-small cell lung cancer: a focus on epidermal growth factor receptor mutations. Chin Clin Oncol. (2015) 4:47. doi: 10.3978/j.issn.2304-3865.2015.12.04
6. Gallant JN, Sheehan JH, Shaver TM, Bailey M, Lipson D, Chandramohan R. EGFR kinase domain duplication (EGFR-KDD) is a novel oncogenic driver in lung cancer that is clinically responsive to afatinib. Cancer Discov. (2015) 5:1155–63. doi: 10.1158/2159-8290.CD-15-0654
7. Ozer BH, Wiepz GJ, Bertics PJ. Activity and cellular localization of an oncogenic glioblastoma multiforme-associated EGF receptor mutant possessing a duplicated kinase domain. Oncogene. (2010) 29:855–64. doi: 10.1038/onc.2009.385
8. Baik CS, Wu D, Smith C, Martins RG, Pritchard CC. Durable response to tyrosine kinase inhibitor therapy in a lung cancer patient harboring epidermal growth factor receptor tandem kinase domain duplication. J Thorac Oncol. (2015) 10:e97–9. doi: 10.1097/JTO.0000000000000586
9. Zhu YC, Wang WX, Xu CW, Tan QH, Li JY, Zhuang W. Lung adenocarcinoma patient with an EGFR kinase domain duplication (KDD) and the response to icotinib. J Thorac Dis. (2018) 10:E359–63. doi: 10.21037/jtd.2018.04.162
10. Gonzalez De Castro D, Clarke PA, Al-Lazikani B, Workman P. Personalized cancer medicine: molecular diagnostics, predictive biomarkers, and drug resistance. Clin Pharmacol Ther. (2013) 93:252–9. doi: 10.1038/clpt.2012.237
Keywords: non-small cell lung cancer, epidermal growth factor receptor, kinase domain, next-generation sequencing, afatinib
Citation: Chen D, Li X, Wu B, Zheng X, Wang W, Chen H, Dong Y, Xu C and Fang M (2020) A Novel Oncogenic Driver in a Lung Adenocarcinoma Patient Harboring an EGFR-KDD and Response to Afatinib. Front. Oncol. 10:867. doi: 10.3389/fonc.2020.00867
Received: 12 December 2019; Accepted: 04 May 2020;
Published: 16 June 2020.
Edited by:
Massimo Di Maio, University of Turin, ItalyReviewed by:
Timothy F. Burns, University of Pittsburgh, United StatesGiulia Mazzaschi, University Hospital of Parma, Italy
Copyright © 2020 Chen, Li, Wu, Zheng, Wang, Chen, Dong, Xu and Fang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Wen-xian Wang, aGVsZW4tMDQwN0AxNjMuY29t; Yi-yu Dong, ZHl5MjAwNkAyMWNuLmNvbQ==; Chun-wei Xu, eHVjaHVud2VpYmJiQDE2My5jb20=
†These authors have contributed equally to this work
 Dong Chen1†
Dong Chen1† Chun-wei Xu
Chun-wei Xu