- 1Department of Medical Genetics, Shahid Beheshti University of Medical Sciences, Tehran, Iran
- 2Urology and Nephrology Research Center, Shahid Beheshti University of Medical Sciences, Tehran, Iran
Melanoma is the utmost fatal kind of skin neoplasms. Molecular changes occurring during the pathogenic processes of initiation and progression of melanoma are diverse and include activating mutations in BRAF and NRAS genes, hyper-activation of PI3K/AKT pathway, inactivation of p53 and alterations in CDK4/CDKN2A axis. Moreover, several miRNAs have been identified to be implicated in the biology of melanoma through modulation of expression of genes being involved in these pathways. In the current review, we provide a summary of the bulk of information about the role of miRNAs in the pathobiology of melanoma, their possible application as biomarkers and their emerging role as therapeutic targets for this kind of skin cancer.
Introduction
Arising from unrestrained proliferation of melanocytes, melanoma is the utmost fatal kind of skin neoplasm (1). Though melanoma encompasses less than 5% of all skin cancers, it accounts for most of skin neoplasms mortalities (2). When the cancer is diagnosed in early stages, surgical resection of the tumor is the appropriate therapeutic options for enhancement of survival of patients. Yet, based on the metastatic potential of melanoma, surgery is not satisfactory in advanced stages of melanoma (3). Although the mortality rate of primary melanoma is about 11%, metastatic melanoma has a poor prognosis resulting from inefficiency of conventional therapies (4, 5). Meanwhile, novel therapeutic option might offer efficient methods for these patients. For instance, immunotherapeutic approaches such as administration of Anti-PD1 (nivolumab, pembrolizumab) alone, or the combination of anti-PD1 with anti-cytotoxic T lymphocyte-associated protein 4 (CTLA4) ipilimumab has raised the survival of patients who suffer from advanced stages of melanoma (6).
Targeted therapies, like combinations of BRAF inhibitors (Dabrafenib) and MEK inhibitors (vemurafenib) are also frequently used on BRAFV600E mutant melanomas. Superficial spreading, nodular, lentigo maligna and acral lentiginous melanomas represent the main types of melanoma with the first one being the most frequent type (4). Ultraviolet radiation and melanocytic nevi are two main risk factors for development of this kind of skin cancer (4). Molecular changes occurring during the pathogenic processes of initiation and progression of melanoma are diverse and include activating mutations in BRAF and NRAS genes, hyper-activation of PI3K/AKT pathway, inactivation of p53 and alterations in CDK4/CDKN2A axis (4). In addition, several studies have shown the critical role of microRNAs (miRNAs) both in the initiation and in the progression of melanoma (7). These transcripts have sizes around 22 nucleotides and are generated through a multi-step process from DNA sequences into primary, precursor and mature miRNAs, respectively. As a general rule, they regulate gene expression through binding with complementary sequences in the 3′ untranslated region (3′ UTR) of mRNAs and subsequently lead to degradation and suppression of translation of the target transcript. Less frequently, they interact with the 5′ UTR, coding or promoter regions (8). Moreover, there are some reports of activation of translation of certain genes by miRNAs in some situations. For instance, let-7 family of miRNAs can induce translation when cell cycle is arrested in spite of their inhibitory effects on translation during cell proliferation (9). Therefore, miRNAs are regarded as important mediators of gene expression. Besides, their presence in extracellular vesicles provides them the opportunity to module communication between various cells (8). In the current paper, we summarize the bulk of information about the role of miRNAs in the pathobiology of melanoma, their possible application as biomarkers and their emerging role as therapeutic targets for this kind of skin cancer.
Dysregulated miRNAs in Melanoma
Expression pattern of miRNAs in melanoma cell lines and clinical specimens has been assessed by both high throughput and candidate gene approaches. An example of the former types of studies is the study conducted by Zhang et al. (10). They reported DNA copy number changes in miRNA coding genes in the majority of the assessed melanoma samples. Notably, miRNA copy alterations have been correlated with miRNA expression. Moreover, they reported copy number alterations in genes contributing in the biogenesis or function of miRNAs in tumor samples (10). Through a microarray-based technique, Aksenenko et al. have identified differential expression of 143 miRNAs between melanoma samples and adjacent skin tissues. Among the dysregulated miRNAs has been the up-regulated miRNA hsa-miR-146a-5p which has been predicted to be associated with Toll-like receptor, NF-κB and ErB pathways. Moreover, this miRNA has been shown to target one of the most recurrently mutated genes in melanoma i.e., the NRAS gene (11).
miRNA also affect activity of melanoma-related signaling pathways. Figure 1 depicts the functional association between two miRNAs and AKT and NF-κB signaling pathways.
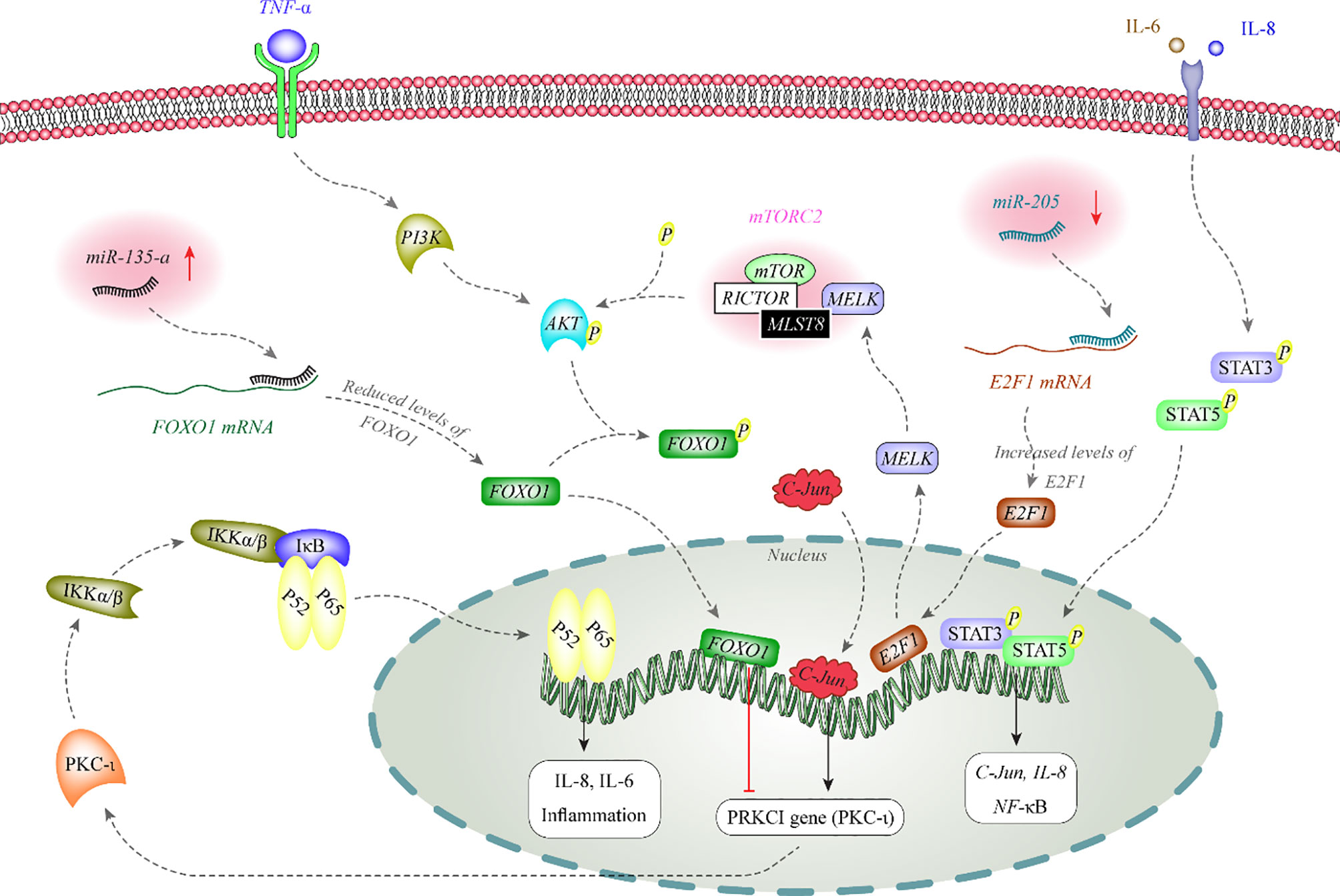
Figure 1 AKT phosphorylates FOXO1 to inhibit its nuclear translocation. FOXO1 has a role in the suppression of expression of PKC-iota in the nucleus. PKC-iota is an inducer of NF-κB which enhances expression of inflammatory genes in the nucleus. Expression of miR-135-a is increased in melanoma. This miRNA binds with the 3’ UTR of FOXO1 to decrease its expression (12, 13). On the other hand, miR-205 is decreased in melanoma. This miRNA inhibits expression of E2F1 through binding with its 3’ UTR. E2F1 increases expression of MELK. MELK activates mTORC2 through binding with MLST8. mTORC2 has a role in phosphorylation and activation of AKT (14).
Expression profiling of miRNAs in melanocytes and melanoma cells originated from primary or metastatic melanoma cells has provided valuable data about the role of miRNAs in each phase of cancer development. A panel of miRNAs including miR-133a, miR-199b, miR-453, miR-520f, miR-521, and miR-551b has been found consistently up-regulated in the course of cancer development from melanocytes to primary cancerous cell and from primary to metastatic melanomas. On the other hand, miR-190 had the opposite trend during this course. Furthermore, expressions of miR-126, miR-29c, miR-506, miR-507, and miR-520d* have been found to be increased during the early progression of melanoma and have been decreased in the metastatic phase. Two other miRNAs including miR-489 and miR-527 had the opposite pattern of expression (15). Levati et al. have demonstrated up-regulation of miR-17-5p, miR-18a, miR-20a, and miR-92a while down-regulation of miR-146a, miR-146b and miR-155 in most of assessed melanoma cell lines compared with melanocytes (16).
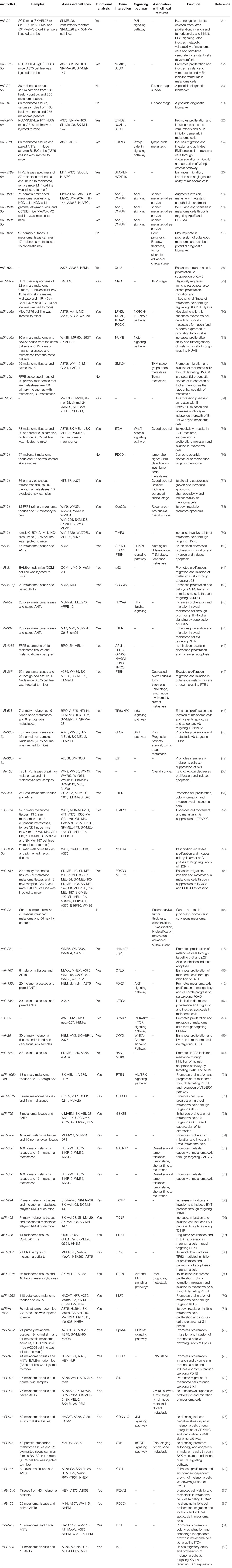
Other studies have reported dysregulation of several other miRNAs in the melanoma samples. Among up-regulated miRNAs are miR-221 and miR-222 which induce malignant features through decreasing expression of c-KIT receptor and p27Kip. Both miRNAs promote epithelial-mesenchymal transition (17, 18). Moreover, expression of miR-210 has been demonstrated to be elevated in several cancer types including melanoma. Its expression has been correlated with metastatic potential of melanoma tumors. Up-regulation of miR-210 in cancer cell lines facilitates evasion from hypoxia-induced cell cycle arrest and partly upturned the hypoxic gene expression profile. This miRNA has been revealed to target a known MYC antagonist namely MNT. Therefore, miR-210 has been shown to modulate the hypoxia response in cancer cells via regulating an important transcriptional suppressor of the MYC-MAX axis (19). In an attempt to detect the miRNAs that are regulated by BRAFV600E mutation via the ERK pathway, Vitiello et al. have conducted RNA sequencing on A375 cell line and a vemurafenib-resistant clone. Their experiments have led to identification of miR-204 and miR-211 as the utmost over-expressed miRNAs by vemurafenib. In spite of belonging to an identical miRNA family, miR-204 and miR-211 have distinguishing characteristics. miR-204 is regulated by STAT3 and its transcript levels are increased in amelanotic melanoma cells, where it functions as a mediator of anti-migratory effects of vemurafenib by modulating expression of AP1S2. On the contrary, miR-211, as a direct target of MITF, is over-expressed in melanotic melanoma cells. miR-211 regulates expression of EDEM1 and subsequently weakens the destruction of Tyrosinase. Thus, miR-211 is a facilitator of pro-pigmentation function of vemurafenib (20). Table 1 displays the list of over-expressed miRNAs in melanoma.
Numerous tumor suppressor miRNAs have been down-regulated in melanoma samples. For instance, while miR-34a is constantly detected in normal melanocytes, it is not expressed in uveal melanoma cells. Forced over-expression of this miRNA in uveal melanoma cells remarkably diminishes their growth and migratory abilities. Mechanistically, this miRNA inhibits expression of c-Met protein and decreases the levels of phosphorylated Akt and cell cycle-related proteins (83). Besides, miR-34b, miR-34c, and miR-199a* have been shown to down-regulate MET expression, suppressing the invasive growth features in the melanoma cells (84). Furthermore, expressions of the let-7 miRNAs have been shown to be decreased in primary melanomas compared with benign nevus samples. Forced up-regulation of let-7b in melanoma cells has led to significant decrease in the expression of cyclins D1, D3, and A, and CDK4. The functional interaction between let-7b and cyclin D1 has been verified through in vitro experiments (85). The inhibitory effect of let-7a on expression of integrin beta 3 has been verified in another study (86). In addition, functional studies have shown the role of miR-155 in the suppression of proliferation of a number of melanoma cell lines and induction of apoptosis in these cells (16). Table 2 lists the down-regulated miRNAs in melanoma.
Diagnostic/prognostic miRNAs in melanoma
Hanniford et al. have introduced a miRNA panel consisting of miR-150-5p, miR-15b-5p, miR-16-5p, and miR-374b-3p whose expression levels could predict the possibility of brain metastasis of melanoma tumors along with clinical stage. Moreover, Kaplan-Meier analysis showed the significance of this miRNA panel in determination of brain-metastasis-free and overall survival of patients with melanoma (273). Stark et al. have assessed expression levels of 17 miRNAs in both melanoma tissues and serum samples of these patients compared with cancer-free individuals. Expression levels of these miRNAs in melanoma samples have been shown to predict stage, recurrence, and survival of patients. Notably, serum expression of a seven-miRNA panel could distinguish melanoma patients from control subjects with 93% sensitivity and more than 82% specificity if at least 4 miRNAs were expressed. Based on the superiority of this miRNA panel above the conventional serological biomarkers for melanoma, it has been suggested as a tool for monitoring disease course in early metastatic melanoma cases to identify relapse after tumor excision or adjuvant therapy (23). Worley et al. have used a high throughput technique to identify the miRNAs whose expression profile could predict the metastatic potential of uveal melanomas. Their approach led to identification of let-7b and miR-199a as the most robust discriminators. Notably, expression profile of six miRNAs could differentiate low and high risk groups with optimal sensitivity and specificity values (274). Table 3 shows the role of miRNAs in the prediction of prognosis of melanoma using Kaplan-Meier or Cox regression analyses.
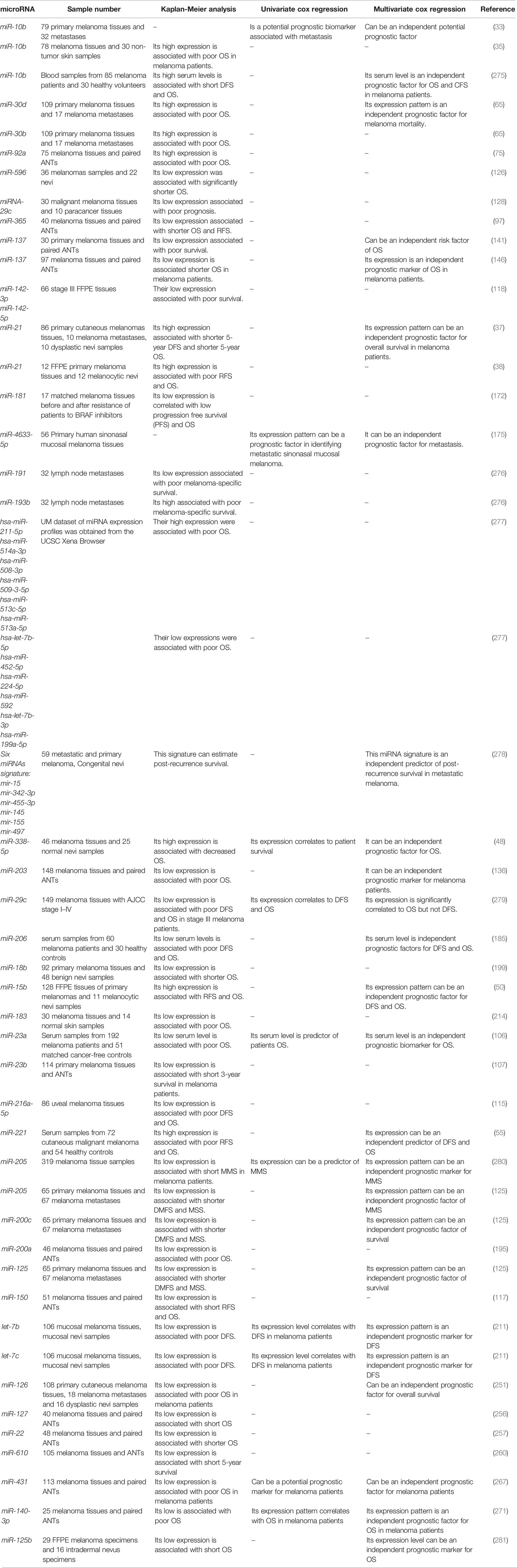

Table 3 Role of melanoma in prediction of prognosis of melanoma (DMFS, distant metastasis free survival; OS, overall survival; DFS, disease-free survival; RFS, relapse-free survival; MSS, melanoma specific survival).
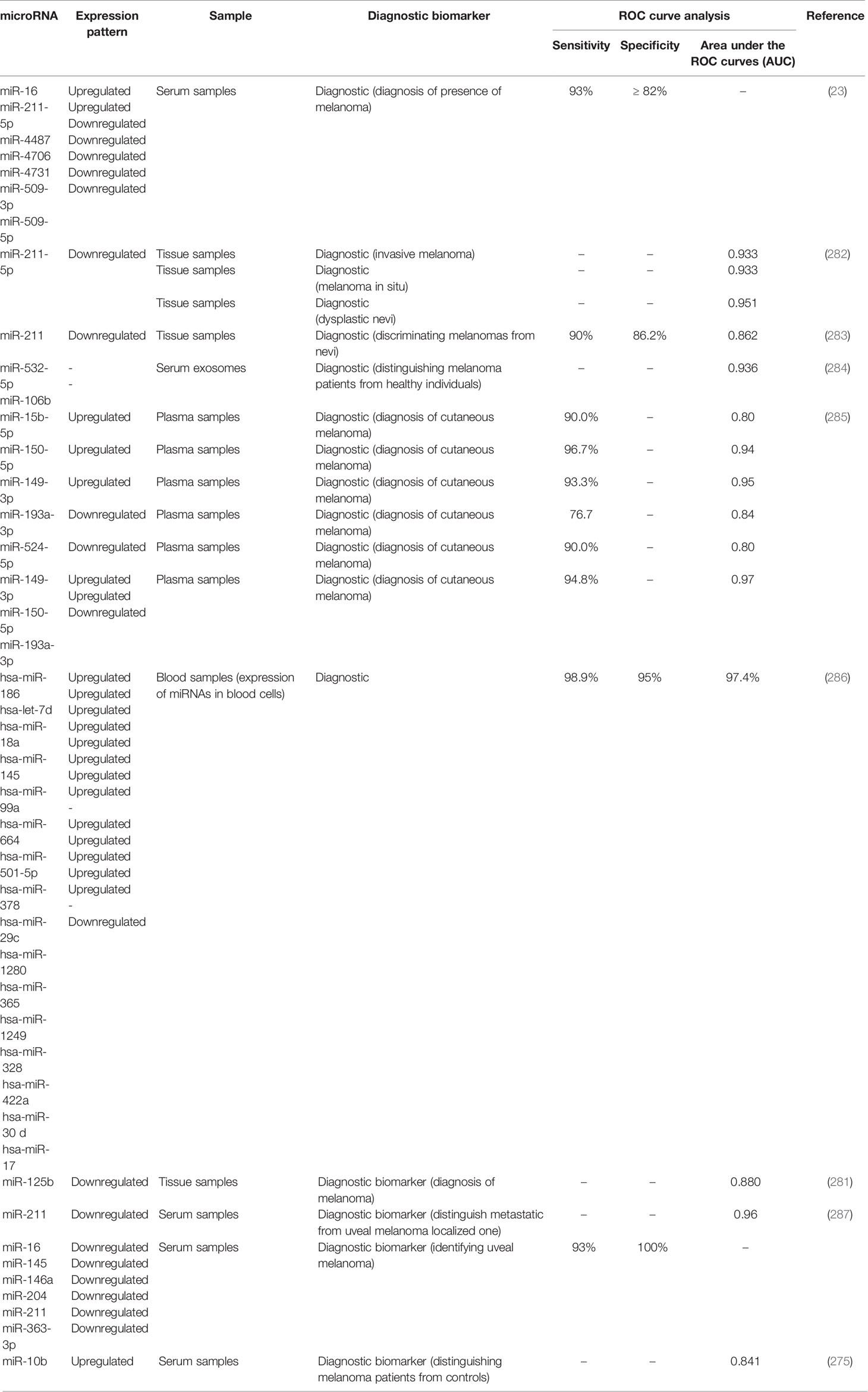
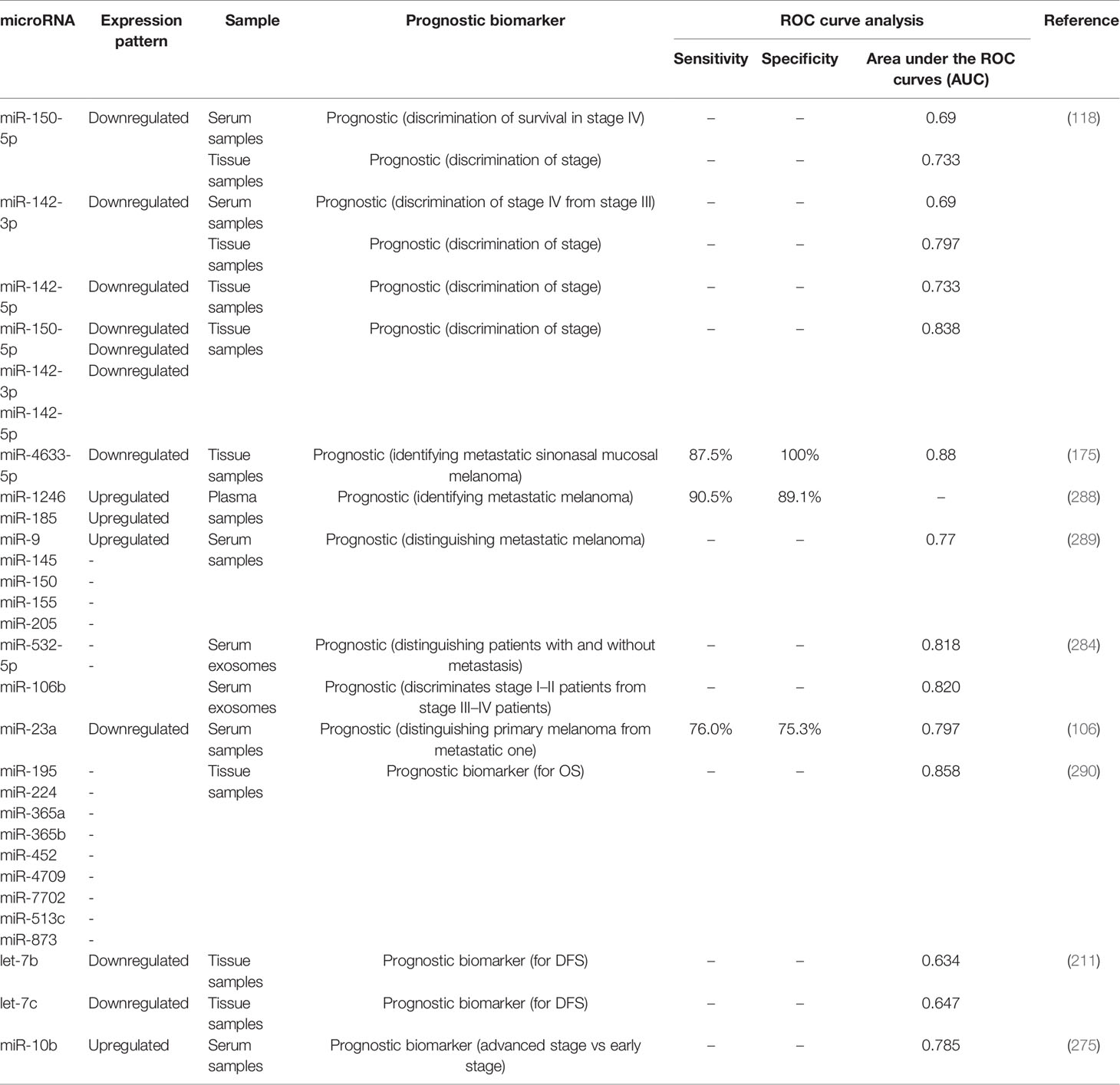
Receiver operating characteristic (ROC) curves have been used to assess the diagnostic or prognostic values of miRNAs in melanoma. Based on the area under curve (AUC) values, several miRNAs can be suggested as appropriate biomarkers for this kind of cancer. In the field of miRNA application in melanoma diagnosis, these curves depict the diagnostic capability of expression level of a miRNA as a binary classifier system for detection of melanoma cases as its discrimination threshold is changed. In other words, these curves are generated by plotting the true positive rate against the false positive rate at different threshold points. Notably, serum expression levels of several miRNAs have high sensitivity and specificity values for differentiating between melanoma patients and healthy subjects or between metastatic and non-metastatic melanomas. Tables 4, 5 list the miRNAs whose application as diagnostic or prognostic markers has been evaluated using ROC curve analysis, respectively.
Implications of miRNAs in the Treatment of Melanoma
miRNAs are implicated in the therapeutic effects of several anti-cancer agents. For instance, Genistein, the isoflavone extracted from soybean, has been shown to suppress proliferation of human uveal melanoma cells possibly through modulating expression of miR-27a and its target gene ZBTB10 (291).
miRNAs are also involved in conferring resistance to immunotherapeutic modalities. For instance, expression of miR-222 has been shown to be higher in melanoma samples obtained from patients who did not respond to ipilimumab compared with those benefitting from this option (292). Mechanistically, the ADAR1/miR-222/ICAM1 axis has been reported to be involved in this process (292). Other miRNAs such as miR-488-3p, miR-195 and miR-211 participate in the regulation of response to the chemotherapeutic agent cisplatin (130, 162, 163)
Application of miRNAs in the therapeutic settings is limited by target specificity issues (293). However, some miRNAs are currently being tested in some diseases. Among these therapeutic modalities are miR-122/miravirsen and miR-92/MRG 110 which have been manufactured by Roche/Santaris and Regulus Therapeutics, respectively (293).
Association Between Polymorphisms Within miRNAs and Risk of Melanoma
Theoretically, polymorphisms with miRNA coding genes can alter their expression or function. Although such polymorphisms are predicted to influence the risk of different cancers such as melanoma, this field has not been vastly explored. Few studies have assessed association between a certain polymorphism within miR-146a namely the rs2910164 G/C and melanoma risk. In spite of the proposed role for allele C of this polymorphism in conferring risk of melanoma (294, 295), cell line studies have shown that G allele confers high proliferative capacity to melanoma cells (296). Table 6 summarizes the results of these studies.
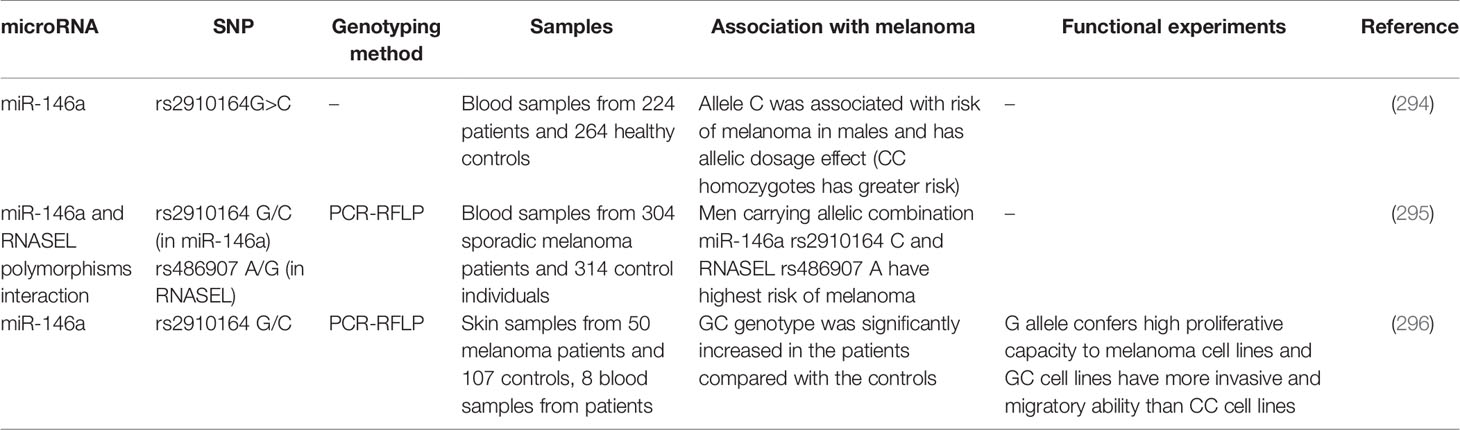

Table 6 Summary of studies which assessed association between miRNA polymorphisms and risk of melanoma.
Discussion
Dysregulation of miRNAs in melanoma samples and cell line have been reported by several studies. The functional consequences of such dysregulation on cell behavior have also been appraised. However, the underlying mechanism of such dysregulation is not clarified completely. Copy number variations in miRNA-coding genes or genes associated with the biogenesis or function of miRNAs may be responsible for the observed dysregulation of miRNAs in melanoma and other types of cancers (10). Moreover, the role of epigenetic factors in this process should not be ignored. For instance, CpG methylation of the miR-34a promoter has been suggested as an underlying mechanism for down-regulation of this miRNA in primary melanoma samples and melanoma cell lines (297). Another possible mediators of miRNA dysregulation in the melanoma are melanoma-inducing transcription factors such as MITF whose role in the expression of a number of miRNAs has been verified (298). As several miRNAs are implicated in the modulation of skin response to ultraviolet radiation (299), this environmental carcinogen might also affect expression of miRNAs which are involved in the melanomagenesis.
Mechanistically, several melanoma-associated miRNAs function upstream or downstream of known oncogenes in melanoma. For instance, miR-137 and miR-182 are among miRNAs that target MITF oncogene (54, 300). Moreover, expressions of several miRNAs such as a number of let-7 family members, miR-221/222, miR-17-92 and miR-106-363 clusters, miR-29, miR-146a, miR-148b, and miR-125b have been shown to be modulated by MITF (298). Moreover, several miRNAs such as miR-7, miR-23a and miR-596 have functional interactions with MAPK/ERK and PI3K/PTEN/Akt signaling pathways in the context of melanoma. A number of miRNAs such as miR-378, miR-10b, miR-25, miR-485-5p, miR-708, miR-136, miR-488-5p, miR-29a, miR-22 and miR-140-5p have interactions with Wnt/β-catenin pathway. Finally, miR-21, miR-7-5p, miR-23b, miR-145-5p, miR-9, miR-29a, miR-377 and miR-140-5p interacts with NF-κB signaling in the context of melanoma development. Thus, a number of miRNAs provide functional links between cancer-related pathways in this context.
miRNAs have functions both in the paternal cell in which they are produced as well as in the adjoining cells. These transcripts can modulate characteristics of adjacent melanoma cells or directly affect tumor niche by modifying extracellular matrix and function of resident cells in this environment including fibroblasts and endothelial or immune cells. This activity of miRNAs potentiates them as contributors of melanoma metastatic potential through affecting intravasation of cancer cells into vessels, viability of tumor cells in the circulation, their leakage in the target tissues, and establishment of the pre-metastatic milieu in remote organs (301).
Several miRNAs have been shown to differentiate melanoma patients from healthy subjects or distinguish between metastatic and non-metastatic melanoma patients. The prognostic assays founded on miRNAs signature can enhance the efficacy of conventional staging systems in predicting patients’ prognosis and their management in the clinical settings in the terms of choosing adjuvant therapies or clinical trial enrolment. Therefore, these miRNAs are potential biomarkers for this kind of skin cancer.
Numerous miRNAs have been dysregulated in tumor samples or peripheral blood of patients with melanoma. Such dysregulation can be used as biomarker for early detection of melanoma or follow-up of patients after initial treatments to uncover any possible tumor recurrence. Blood-based biomarkers are expected to substitute invasive methods of cancer diagnosis in future. Based on the heterogeneous pattern of miRNAs expression in tumor samples and the varied expressions among affected individuals, multi-miRNA panels are more promising in the diagnostic approaches compared with individual miRNAs.
Finally, miRNAs might be implicated in the anti-cancer effects of a number of therapeutic agents including both chemical and herbal medicines. Evidence for supporting this idea has come from several studies including a study which revealed the role of miR-27a in mediating the anti-proliferative effects of Genistein in human uveal melanoma cells (291). Moreover, the observed up-regulation of miR-222 in melanoma samples obtained from patients who did not respond to ipilimumab compared with those benefitting from this option (292) implies its contribution in resistance to this agent. Therefore, miRNAs are promising targets for modulation of response of melanoma cells to a wide range of therapeutic options.
Perspectives and Future Directions
Assessment of expression pattern of miRNAs in cohorts of melanoma patients from different ethnicities and uncovering their association with genetic polymorphisms would facilitate design of prognostic/diagnostic panels. The relationship between aberrant miRNA profile and response to therapeutic regimens should be unraveled. Such kinds of approaches pave the way for design of personalized methods of treatment of melanoma. Therapeutic targeting of miRNAs can influence melanoma course and enhance sensitivity to both conventional therapies and immunotherapeutic approaches. Yet, safety and bioavailability issues remained to be solved before implementation of these techniques in the clinical settings.
Author Contributions
SG-F and MT wrote the draft and revised it. MG collected the tables and designed it. All authors contributed to the article and approved the submitted version.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
References
1. Linos E, Swetter SM, Cockburn MG, Colditz GA, Clarke CA. Increasing Burden of Melanoma in the United States. J Invest Dermatol (2009) 129(7):1666–74. doi: 10.1038/jid.2008.423
2. Noone A, Howlader N, Krapcho M, Miller D, Brest A, Yu M, et al. SEER Cancer Statistics Review, 1975-2015 Vol. 4. Bethesda, MD: National Cancer Institute (2018).
3. Erdei E, Torres SM. A New Understanding in the Epidemiology of Melanoma. Expert Rev Anticancer Ther (2010) 10(11):1811–23. doi: 10.1586/era.10.170
4. Liu Y, Sheikh MS. Melanoma: Molecular Pathogenesis and Therapeutic Management. Mol Cell Pharmacol (2014) 6(3):228.
5. Safa A, Gholipour M, Dinger ME, Taheri M, Ghafouri-Fard S. The Critical Roles of LncRNAs in the Pathogenesis of Melanoma. Exp Mol Pathol (2020)117:104558. doi: 10.1016/j.yexmp.2020.104558
6. Gellrich FF, Schmitz M, Beissert S, Meier F. Anti-PD-1 and Novel Combinations in the Treatment of Melanoma—an Update. J Clin Med (2020) 9(1):223. doi: 10.3390/jcm9010223
8. O’Brien J, Hayder H, Zayed Y, Peng C. Overview of MicroRNA Biogenesis, Mechanisms of Actions, and Circulation. Front Endocrinol (2018) 9:402. doi: 10.3389/fendo.2018.00402
9. Vasudevan S, Steitz JA. AU-rich-element-mediated Upregulation of Translation by FXR1 and Argonaute 2. Cell (2007) 128(6):1105–18. doi: 10.1016/j.cell.2007.01.038
10. Zhang L, Huang J, Yang N, Greshock J, Megraw MS, Giannakakis A, et al. MicroRNAs Exhibit High Frequency Genomic Alterations in Human Cancer. Proc Natl Acad Sci (2006) 103(24):9136–41. doi: 10.1073/pnas.0508889103
11. Aksenenko M, Palkina N, Komina A, Tashireva L, Ruksha T. Differences in MicroRNA Expression Between Melanoma and Healthy Adjacent Skin. BMC Dermatol (2019) 19(1):1–9. doi: 10.1186/s12895-018-0081-1
12. Ren J-W, Li Z-J, Tu C. MiR-135 Post-Transcriptionally Regulates FOXO1 Expression and Promotes Cell Proliferation in Human Malignant Melanoma Cells. Int J Clin Exp Pathol (2015) 8(6):6356.
13. Ratnayke WS, Apostolatos CA, Breedy S, Apostolatos AH, Acevedo-Duncan M. FOXO1 Regulates Oncogenic PKC-ι Expression in Melanoma Inversely to c-Jun in an Autocrine Manner Via IL-17E and ICAM-1 Activation. World Acad Sci J (2019) 1(1):25–38. doi: 10.3892/wasj.2018.1
14. Dar AA, Majid S, de Semir D, Nosrati M, Bezrookove V, Kashani-Sabet M. MiRNA-205 Suppresses Melanoma Cell Proliferation and Induces Senescence Via Regulation of E2F1 Protein. J Biol Chem (2011) 286(19):16606–14. doi: 10.1074/jbc.M111.227611
15. Mueller DW, Rehli M, Bosserhoff AK. MiRNA Expression Profiling in Melanocytes and Melanoma Cell Lines Reveals MiRNAs Associated With Formation and Progression of Malignant Melanoma. J Invest Dermatol (2009) 129(7):1740–51. doi: 10.1038/jid.2008.452
16. Levati L, Alvino E, Pagani E, Arcelli D, Caporaso P, Bondanza S, et al. Altered Expression of Selected MicroRNAs in Melanoma: Antiproliferative and Proapoptotic Activity of Mirna-155. Int J Oncol (2009) 35(2):393–400. doi: 10.3892/ijo_00000352
17. Felicetti F, Errico MC, Segnalini P, Mattia G, Carè A. MicroRNA-221 and-222 Pathway Controls Melanoma Progression. Expert Rev Anticancer Ther (2008) 8(11):1759–65. doi: 10.1586/14737140.8.11.1759
18. Igoucheva O, Alexeev V. MicroRNA-dependent Regulation of cKit in Cutaneous Melanoma. Biochem Biophys Res Commun (2009) 379(3):790–4. doi: 10.1016/j.bbrc.2008.12.152
19. Zhang Z, Sun H, Dai H, Walsh R, Imakura M, Schelter J, et al. MicroRNA miR-210 Modulates Cellular Response to Hypoxia Through the MYC Antagonist MNT. Cell Cycle (2009) 8(17):2756–68. doi: 10.4161/cc.8.17.9387
20. Vitiello M, Tuccoli A, D’Aurizio R, Sarti S, Giannecchini L, Lubrano S, et al. Context-Dependent miR-204 and miR-211 Affect the Biological Properties of Amelanotic and Melanotic Melanoma Cells. Oncotarget (2017) 8(15):25395. doi: 10.18632/oncotarget.15915
21. Sahoo A, Sahoo SK, Joshi P, Lee B, Perera RJ. MicroRNA-211 Loss Promotes Metabolic Vulnerability and BRAF Inhibitor Sensitivity in Melanoma. J Invest Dermatol (2019) 139(1):167–76. doi: 10.1016/j.jid.2018.06.189
22. Díaz-Martínez M, Benito-Jardón L, Alonso L, Koetz-Ploch L, Hernando E, Teixidó J. miR-204-5p and miR-211-5p Contribute to BRAF Inhibitor Resistance in Melanoma. Cancer Res (2018) 78(4):1017–30. doi: 10.1158/0008-5472.CAN-17-1318
23. Stark MS, Klein K, Weide B, Haydu LE, Pflugfelder A, Tang YH, et al. The Prognostic and Predictive Value of Melanoma-Related MicroRNAs Using Tissue and Serum: A MicroRNA Expression Analysis. EBioMedicine (2015) 2(7):671–80. doi: 10.1016/j.ebiom.2015.05.011
24. Sun M, Ma X, Tu C, Wang X, Qu J, Wang S, et al. MicroRNA-378 Regulates Epithelial–Mesenchymal Transition and Metastasis of Melanoma by Inhibiting FOXN3 Expression Through the Wnt/β-Catenin Pathway. Cell Biol Int (2019) 43(10):1113–24. doi: 10.1002/cbin.11027
25. Tupone MG, D’Aguanno S, Di Martile M, Valentini E, Desideri M, Trisciuoglio D, et al. MicroRNA-378a-5p is a Novel Positive Regulator of Melanoma Progression. Oncogenesis (2020) 9(2):1–13. doi: 10.1038/s41389-020-0203-6
26. Pencheva N, Tran H, Buss C, Huh D, Drobnjak M, Busam K, et al. Convergent Multi-MiRNA Targeting of Apoe Drives LRP1/LRP8-dependent Melanoma Metastasis and Angiogenesis. Cell (2012) 151(5):1068–82. doi: 10.1016/j.cell.2012.10.028
27. Lin N, Zhou Y, Lian X, Tu Y. Expression of MicroRNA-106b and Its Clinical Significance in Cutaneous Melanoma. Genet Mol Res (2015) 14(4):16379–85. doi: 10.4238/2015.December.9.6
28. Wang J, Li H, Zhang J, Zhang C, Hou X. Suppression of Connexin 43 Expression by miR-106a Promotes Melanoma Cell Proliferation. Eur Rev Med Pharmacol Sci (2019) 23(3):965–71. doi: 10.26355/eurrev_201902_16983
29. Mastroianni J, Stickel N, Andrlova H, Hanke K, Melchinger W, Duquesne S, et al. miR-146a Controls Immune Response in the Melanoma Microenvironment. Cancer Res (2019) 79(1):183–95. doi: 10.1158/0008-5472.CAN-18-1397
30. Raimo M, Orso F, Grassi E, Cimino D, Penna E, De Pittà C, et al. miR-146a Exerts Differential Effects on Melanoma Growth and Metastatization. Mol Cancer Res (2016) 14(6):548–62. doi: 10.1158/1541-7786.MCR-15-0425-T
31. Forloni M, Dogra SK, Dong Y, Conte D, Ou J, Zhu LJ, et al. miR-146a Promotes the Initiation and Progression of Melanoma by Activating Notch Signaling. Elife (2014) 3:e01460. doi: 10.7554/eLife.01460
32. Pu W, Shang Y, Shao Q, Yuan X. miR-146a Promotes Cell Migration and Invasion in Melanoma by Directly Targeting SMAD4. Oncol Lett (2018) 15(5):7111–7. doi: 10.3892/ol.2018.8172
33. Saldanha G, Elshaw S, Sachs P, Alharbi H, Shah P, Jothi A, et al. MicroRNA-10b is a Prognostic Biomarker for Melanoma. Modern Pathol (2016) 29(2):112–21. doi: 10.1038/modpathol.2015.149
34. Datar I, Kalpana G, Choi J, Basuroy T, Trumbly R, Chaitanya Arudra SK, et al. Critical Role of miR-10b in B-RafV600E Dependent Anchorage Independent Growth and Invasion of Melanoma Cells. PloS One (2019) 14(4):e0204387. doi: 10.1371/journal.pone.0204387
35. Wang S, Wu Y, Xu Y, Tang X. miR-10b Promoted Melanoma Progression Through Wnt/β-Catenin Pathway by Repressing ITCH Expression. Gene (2019) 710:39–47. doi: 10.1016/j.gene.2019.05.043
36. Jiao J, Fan Y, Zhang Y. Expression and Clinicopathological Significance of MicroRNA-21 and Programmed Cell Death 4 in Malignant Melanoma. J Int Med Res (2015) 43(5):672–8. doi: 10.1177/0300060515583707
37. Jiang L, Lv X, Li J, Li J, Li X, Li W, et al. The Status of MicroRNA-21 Expression and Its Clinical Significance in Human Cutaneous Malignant Melanoma. Acta Histochem (2012) 114(6):582–8. doi: 10.1016/j.acthis.2011.11.001
38. Satzger I, Mattern A, Kuettler U, Weinspach D, Niebuhr M, Kapp A, et al. Micro RNA-21 is Upregulated in Malignant Melanoma and Influences Apoptosis of Melanocytic Cells. Exp Dermatol (2012) 21(7):509–14. doi: 10.1111/j.1600-0625.2012.01510.x
39. Del Campo SEM, Latchana N, Levine KM, Grignol VP, Fairchild ET, Jaime-Ramirez AC, et al. MiR-21 Enhances Melanoma Invasiveness Via Inhibition of Tissue Inhibitor of Metalloproteinases 3 Expression: in Vivo Effects of MiR-21 Inhibitor. PloS One (2015) 10(1):e0115919. doi: 10.1371/journal.pone.0115919
40. Mao XH, Chen M, Wang Y, Cui PG, Liu SB, Xu ZY. MicroRNA-21 Regulates the ERK/NF-κb Signaling Pathway to Affect the Proliferation, Migration, and Apoptosis of Human Melanoma A375 Cells by Targeting SPRY1, PDCD4, and PTEN. Mol Carcinogen (2017) 56(3):886–94. doi: 10.1002/mc.22542
41. Wang Y-C, Yang X, Wei W-B, Xu X-L. Role of MicroRNA-21 in Uveal Melanoma Cell Invasion and Metastasis by Regulating p53 and Its Downstream Protein. Int J Ophthalmol (2018) 11(8):1258. doi: 10.18240/ijo.2018.08.03
42. Yang Z, Liao B, Xiang X, Ke S. miR-21-5p Promotes Cell Proliferation and G1/s Transition in Melanoma by Targeting CDKN2C. FEBS Open Bio (2020) 10(5):752–60. doi: 10.1002/2211-5463.12819
43. Xia Z, Yang C, Yang X, Wu S, Feng Z, Qu L, et al. MiR-652 Promotes Proliferation and Migration of Uveal Melanoma Cells by Targeting HOXA9. Med Sci Monit (2019) 25:8722. doi: 10.12659/MSM.917099
44. Ling J, Lu P, Zhang Y, Jiang S, Zhang Z. miR-367 Promotes Uveal Melanoma Cell Proliferation and Migration by Regulating PTEN. Genet Mol Res (2017) 16(3). doi: 10.4238/gmr16039067
45. Komina A, Palkina N, Aksenenko M, Tsyrenzhapova S, Ruksha T. Antiproliferative and Pro-Apoptotic Effects of MiR-4286 Inhibition in Melanoma Cells. PloS One (2016) 11(12):e0168229. doi: 10.1371/journal.pone.0168229
46. Long J, Luo J, Yin X. miR−367 Enhances the Proliferation and Invasion of Cutaneous Malignant Melanoma by Regulating Phosphatase and Tensin Homolog Expression. Mol Med Rep (2018) 17(5):6526–32. doi: 10.3892/mmr.2018.8663
47. Bhattacharya A, Schmitz U, Raatz Y, Schönherr M, Kottek T, Schauer M, et al. miR-638 Promotes Melanoma Metastasis and Protects Melanoma Cells From Apoptosis and Autophagy. Oncotarget (2015) 6(5):2966. doi: 10.18632/oncotarget.3070
48. Long J, Luo J, Yin X. MiR-338-5p Promotes the Growth and Metastasis of Malignant Melanoma Cells Via Targeting CD82. Biomed Pharmacother (2018) 102:1195–202. doi: 10.1016/j.biopha.2018.03.075
49. Hao T, Li C, Ding X, Xing X. MicroRNA-363-3p/p21 (Cip1/Waf1) Axis is Regulated by HIF-2α in Mediating Stemness of Melanoma Cells. Neoplasma (2019) 66(3):427–36. doi: 10.4149/neo_2018_180828N655
50. Satzger I, Mattern A, Kuettler U, Weinspach D, Voelker B, Kapp A, et al. MicroRNA-15b Represents an Independent Prognostic Parameter and is Correlated With Tumor Cell Proliferation and Apoptosis in Malignant Melanoma. Int J Cancer (2010) 126(11):2553–62. doi: 10.1002/ijc.24960
51. Sun L, Wang Q, Gao X, Shi D, Mi S, Han Q. MicroRNA-454 Functions as an Oncogene by Regulating PTEN in Uveal Melanoma. FEBS Lett (2015) 589(19):2791–6. doi: 10.1016/j.febslet.2015.08.007
52. Penna E, Orso F, Cimino D, Tenaglia E, Lembo A, Quaglino E, et al. MicroRNA-214 Contributes to Melanoma Tumour Progression Through Suppression of TFAP2C. EMBO J (2011) 30(10):1990–2007. doi: 10.1038/emboj.2011.102
53. Li J, Zhao R, Fang R, Wang J. miR-122-5p Inhibits the Proliferation of Melanoma Cells by Targeting NOP14. J South Med Univ (2018) 38(11):1360–5. doi: 10.12122/j.issn.1673-4254.2018.11.14
54. Segura MF, Hanniford D, Menendez S, Reavie L, Zou X, Alvarez-Diaz S, et al. Aberrant miR-182 Expression Promotes Melanoma Metastasis by Repressing FOXO3 and Microphthalmia-Associated Transcription Factor. Proc Natl Acad Sci (2009) 106(6):1814–9. doi: 10.1073/pnas.0808263106
55. Li P, He Q-Y, Luo C-Q, Qian L-Y. Circulating miR-221 Expression Level and Prognosis of Cutaneous Malignant Melanoma. Med Sci Monit (2014) 20:2472. doi: 10.12659/MSM.891327
56. Zhang K, Guo L. MiR-767 Promoted Cell Proliferation in Human Melanoma by Suppressing CYLD Expression. Gene (2018) 641:272–8. doi: 10.1016/j.gene.2017.10.055
57. Hu Y, Wang Q, Zhu X-H. MiR-135b is a Novel Oncogenic Factor in Cutaneous Melanoma by Targeting LATS2. Melanoma Res (2019) 29(2):119–25. doi: 10.1097/CMR.0000000000000524
58. Jiang Q-Q, Liu W-B. miR-25 Promotes Melanoma Progression by Regulating RNA Binding Motif Protein 47. Médecine/Sciences (2018) 34:59–65. doi: 10.1051/medsci/201834f111
59. Huo J, Zhang Y, Li R, Wang Y, Wu J, Zhang D. Upregulated MicroRNA-25 Mediates the Migration of Melanoma Cells by Targeting DKK3 Through the WNT/β-Catenin Pathway. Int J Mol Sci (2016) 17(11):1124. doi: 10.3390/ijms17111124
60. Koetz-Ploch L, Hanniford D, Dolgalev I, Sokolova E, Zhong J, Díaz-Martínez M, et al. Micro RNA-125a Promotes Resistance to BRAF Inhibitors Through Suppression of the Intrinsic Apoptotic Pathway. Pigment Cell Melanoma Res (2017) 30(3):328–38. doi: 10.1111/pcmr.12578
61. Chen X-E, Chen P, Chen S-S, Ma T, Shi G, Zhou Y, et al. miR−106b−5p Promotes Cell Cycle Progression of Malignant Melanoma by Targeting PTEN. Oncol Rep (2018) 39(1):331–7. doi: 10.3892/or.2017.6099
62. Zhang L, He X, Li F, Pan H, Huang X, Wen X, et al. The miR-181 Family Promotes Cell Cycle by Targeting CTDSPL, a Phosphatase-Like Tumor Suppressor in Uveal Melanoma. J Exp Clin Cancer Res (2018) 37(1):1–13. doi: 10.1186/s13046-018-0679-5
63. Qiu H-J, Lu X-H, Yang S-S, Weng C-Y, Zhang E-K, Chen F-C. MiR-769 Promoted Cell Proliferation in Human Melanoma by Suppressing GSK3B Expression. Biomed Pharmacother (2016) 82:117–23. doi: 10.1016/j.biopha.2016.04.052
64. Zhou J, Jiang J, Wang S, Xia X. Oncogenic Role of MicroRNA−20a in Human Uveal Melanoma. Mol Med Rep (2016) 14(2):1560–6. doi: 10.3892/mmr.2016.5433
65. Gaziel-Sovran A, Segura MF, Di Micco R, Collins MK, Hanniford D, de Miera EV-S, et al. miR-30b/30d Regulation of GalNAc Transferases Enhances Invasion and Immunosuppression During Metastasis. Cancer Cell (2011) 20(1):104–18. doi: 10.1016/j.ccr.2011.05.027
66. Knoll S, Fürst K, Kowtharapu B, Schmitz U, Marquardt S, Wolkenhauer O, et al. E2F1 Induces miR-224/452 Expression to Drive EMT Through TXNIP Downregulation. EMBO Rep (2014) 15(12):1315–29. doi: 10.15252/embr.201439392
67. Ohira T, Naohiro S, Nakayama Y, Osaki M, Okada F, Oshimura M, et al. miR-19b Regulates HTERT MRNA Expression Through Targeting PITX1 MRNA in Melanoma Cells. Sci Rep (2015) 5(1):1–9. doi: 10.1038/srep08201
68. Lankenau MA, Patel R, Liyanarachchi S, Maharry SE, Hoag KW, Duggan M, et al. MicroRNA-3151 Inactivates TP53 in BRAF-mutated Human Malignancies. Proc Natl Acad Sci (2015) 112(49):E6744–E51. doi: 10.1073/pnas.1520390112
69. Cui L, Li Y, Lv X, Li J, Wang X, Lei Z, et al. Expression of MicroRNA-301a and Its Functional Roles in Malignant Melanoma. Cell Physiol Biochem (2016) 40(1–2):230–44. doi: 10.1159/000452540
70. Zhang D, Li Z, Zhang Y, Tu C, Huo J, Liu Y. miR-4262 Promotes the Proliferation of Human Cutaneous Malignant Melanoma Cells Through KLF6-mediated EGFR Inactivation and p21 Upregulation. Oncol Rep (2016) 36(6):3657–63. doi: 10.3892/or.2016.5190
71. Prasad R, Katiyar SK. Down-Regulation of MiRNA-106b Inhibits Growth of Melanoma Cells by Promoting G1-Phase Cell Cycle Arrest and Reactivation of P21/WAF1/Cip1 Protein. Oncotarget (2014) 5(21):10636. doi: 10.18632/oncotarget.2527
72. Hua K-T, Hong J-B, Sheen Y-S, Huang H-Y, Huang Y-L, Chen J-S, et al. miR-519d Promotes Melanoma Progression by Downregulating Epha4. Cancer Res (2018) 78(1):216–29. doi: 10.1158/0008-5472.CAN-17-1933
73. Wei S, Ma W. MiR-370 Functions as Oncogene in Melanoma by Direct Targeting Pyruvate Dehydrogenase B. Biomed Pharmacother (2017) 90:278–86. doi: 10.1016/j.biopha.2017.03.068
74. Bai X, Yang M, Xu Y. MicroRNA−373 Promotes Cell Migration Via Targeting Salt−Inducible Kinase 1 Expression in Melanoma. Exp Ther Med (2018) 16(6):4759–64. doi: 10.3892/etm.2018.6784
75. Sun H, Yang G, Wang S, Zhang Y, Ding J, Zhang X. MicroRNA-92a Regulates the Development of Cutaneous Malignant Melanoma by Mediating FOXP1. Eur Rev Med Pharmacol Sci (2019) 23(20):8991–9. doi: 10.26355/eurrev_201910_19299
76. Yang C, Yan Z, Hu F, Wei W, Sun Z, Xu W. Silencing of MicroRNA-517a Induces Oxidative Stress Injury in Melanoma Cells Via Inactivation of the JNK Signaling Pathway by Upregulating CDKN1C. Cancer Cell Int (2020) 20(1):32. doi: 10.1186/s12935-019-1064-y
77. Tang H, Xu X, Xiao W, Liao Y, Xiao X, Li L, et al. Silencing of MicroRNA-27a Facilitates Autophagy and Apoptosis of Melanoma Cells Through the Activation of the SYK-dependent MTOR Signaling Pathway. J Cell Biochem (2019) 120(8):13262–74. doi: 10.1002/jcb.28600
78. Qiu H, Yuan S, Lu X. miR−186 Suppressed CYLD Expression and Promoted Cell Proliferation in Human Melanoma. Oncol Lett (2016) 12(4):2301–6. doi: 10.3892/ol.2016.5002
79. Yu Y, Yu F, Sun P. MicroRNA-1246 Promotes Melanoma Progression Through Targeting Foxa2. OncoTargets Ther (2020) 13:1245. doi: 10.2147/OTT.S234276
80. Wan J, Yang J, Huang Y, Deng L. MicroRNA−150 Inhibitors Enhance Cell Apoptosis of Melanoma by Targeting PDCD4. Oncol Lett (2018) 15(2):1475–82. doi: 10.3892/ol.2017.7445
81. Sun M-X, An Q, Chen L-M, Guo L. MIR-520f Regulated Itch Expression and Promoted Cell Proliferation in Human Melanoma Cells. Dose-Response (2020) 18(2):1559325820918450. doi: 10.1177/1559325820918450
82. Wang Z, Liu Y. MicroRNA−633 Enhances Melanoma Cell Proliferation and Migration by Suppressing KAI1. Oncol Lett (2021) 21(2):1–. doi: 10.3892/ol.2020.12349
83. Yan D, Zhou X, Chen X, Hu D-N, Da Dong X, Wang J, et al. MicroRNA-34a Inhibits Uveal Melanoma Cell Proliferation and Migration Through Downregulation of C-Met. Invest Ophthalmol Visual Sci (2009) 50(4):1559–65. doi: 10.1167/iovs.08-2681
84. Migliore C, Petrelli A, Ghiso E, Corso S, Capparuccia L, Eramo A, et al. MicroRNAs Impair MET-mediated Invasive Growth. Cancer Res (2008) 68(24):10128–36. doi: 10.1158/0008-5472.CAN-08-2148
85. Schultz J, Lorenz P, Gross G, Ibrahim S, Kunz M. MicroRNA let-7b Targets Important Cell Cycle Molecules in Malignant Melanoma Cells and Interferes With Anchorage-Independent Growth. Cell Res (2008) 18(5):549–57. doi: 10.1038/cr.2008.45
86. Müller D, Bosserhoff A-K. Integrin β 3 Expression is Regulated by let-7a MiRNA in Malignant Melanoma. Oncogene (2008) 27(52):6698–706. doi: 10.1038/onc.2008.282
87. Huang D, Wang F, Wu W, Lian C, Liu E. MicroRNA-429 Inhibits Cancer Cell Proliferation and Migration by Targeting the AKT1 in Melanoma. Cancer Biomark (2019) 26(1):63–8. doi: 10.3233/CBM-190289
88. Sheng H, Guo Y, Cao D, Li X, Zhao Y, Ding H, et al. MiR-429-5p Attenuates the Migration and Invasion of Malignant Melanoma by Targeting LIMK1. Eur Rev Med Pharmacol Sci (2020) 24(5):2625–31. doi: 10.26355/eurrev_202003_20531
89. Kang K, Zhang J, Zhang X, Chen Z. MicroRNA-326 Inhibits Melanoma Progression by Targeting KRAS and Suppressing the AKT and ERK Signalling Pathways. Oncol Rep (2018) 39(1):401–10. doi: 10.3892/or.2017.6074
90. Dong F, Lou D. MicroRNA-34b/c Suppresses Uveal Melanoma Cell Proliferation and Migration Through Multiple Targets. Mol Vis (2012) 18:537.
91. Hou Q, Han S, Yang L, Chen S, Chen J, Ma N, et al. The Interplay of MicroRNA-34a, LGR4, EMT-associated Factors, and MMP2 in Regulating Uveal Melanoma Cells. Invest Ophthalmol Visual Sci (2019) 60(13):4503–10. doi: 10.1167/iovs.18-26477
92. Liu R, Xie H, Luo C, Chen Z, Zhou X, Xia K, et al. Identification of FLOT2 as a Novel Target for MicroRNA-34a in Melanoma. J Cancer Res Clin Oncol (2015) 141(6):993–1006. doi: 10.1007/s00432-014-1874-1
93. Greenberg E, Hershkovitz L, Itzhaki O, Hajdu S, Nemlich Y, Ortenberg R, et al. Regulation of Cancer Aggressive Features in Melanoma Cells by Micrornas. PloS One (2011) 6(4):e18936. doi: 10.1371/journal.pone.0018936
94. Yan D, Da Dong X, Chen X, Yao S, Wang L, Wang J, et al. Role of MicroRNA-182 in Posterior Uveal Melanoma: Regulation of Tumor Development Through MITF, BCL2 and Cyclin D2. PloS One (2012) 7(7):e40967. doi: 10.1371/journal.pone.0040967
95. Venza I, Visalli M, Beninati C, Benfatto S, Teti D, Venza M. Il-10rα Expression is Post-Transcriptionally Regulated by miR-15a, miR-185, and miR-211 in Melanoma. BMC Med Genomics (2015) 8(1):1–9. doi: 10.1186/s12920-015-0156-3
96. Zhu Y, Wen X, Zhao P. MicroRNA-365 Inhibits Cell Growth and Promotes Apoptosis in Melanoma by Targeting BCL2 and Cyclin D1 (CCND1). Med Sci Monit (2018) 24:3679. doi: 10.12659/MSM.909633
97. Bai J, Zhang Z, Li X, Liu H. MicroRNA-365 Inhibits Growth, Invasion and Metastasis of Malignant Melanoma by Targeting NRP1 Expression. Cancer Biomark (2015) 15(5):599–608. doi: 10.3233/CBM-150500
98. Wu J, Li J, Ren J, Zhang D. MicroRNA-485-5p Represses Melanoma Cell Invasion and Proliferation by Suppressing Frizzled7. Biomed Pharmacother (2017) 90:303–10. doi: 10.1016/j.biopha.2017.03.064
99. Zhu Y, H-l Z, Wang Q-Y, Chen M-J, Liu L-B. Overexpression of MicroRNA-612 Restrains the Growth, Invasion, and Tumorigenesis of Melanoma Cells by Targeting Espin. Molec Cells (2018) 41(2):119. doi: 10.14348/molcells.2018.2235
100. Giles KM, Brown RA, Ganda C, Podgorny MJ, Candy PA, Wintle LC, et al. MicroRNA-7-5p Inhibits Melanoma Cell Proliferation and Metastasis by Suppressing Rela/NF-κb. Oncotarget (2016) 7(22):31663. doi: 10.18632/oncotarget.9421
101. Giles KM, Brown RA, Epis MR, Kalinowski FC, Leedman PJ. MiRNA-7-5p Inhibits Melanoma Cell Migration and Invasion. Biochem Biophys Res Commun (2013) 430(2):706–10. doi: 10.1016/j.bbrc.2012.11.086
102. Sun X, Li J, Sun Y, Zhang Y, Dong L, Shen C, et al. miR-7 Reverses the Resistance to BRAFi in Melanoma by Targeting EGFR/IGF-1R/CRAF and Inhibiting the MAPK and PI3K/AKT Signaling Pathways. Oncotarget (2016) 7(33):53558. doi: 10.18632/oncotarget.10669
103. Zeng HF, Yan S, Wu SF. MicroRNA-153-3p Suppress Cell Proliferation and Invasion by Targeting SNAI1 in Melanoma. Biochem Biophys Res Commun (2017) 487(1):140–5. doi: 10.1016/j.bbrc.2017.04.032
104. Fang W, Fan Y, Fa Z, Xu J, Yu H, Li P, et al. MicroRNA-625 Inhibits Tumorigenicity by Suppressing Proliferation, Migration and Invasion in Malignant Melanoma. Oncotarget (2017) 8(8):13253. doi: 10.18632/oncotarget.14710
105. Lu S, Xu Q. MicroRNA-23a Inhibits Melanoma Cell Proliferation, Migration, and Invasion in Mice Through a Negative Feedback Regulation of Sdcbp and the MAPK/ERK Signaling Pathway. IUBMB Life (2019) 71(5):587–600. doi: 10.1002/iub.1979
106. Guo W, Wang H, Yang Y, Guo S, Zhang W, Liu Y, et al. Down-Regulated miR-23a Contributes to the Metastasis of Cutaneous Melanoma by Promoting Autophagy. Theranostics (2017) 7(8):2231. doi: 10.7150/thno.18835
107. Lv R, Yu J, Sun Q. Anti-Angiogenic Role of MicroRNA-23b in Melanoma by Disturbing NF-κb Signaling Pathway Via Targeted Inhibition of NAMPT. Future Oncol (2020) 16(10):541–458. doi: 10.2217/fon-2019-0699
108. Ma M, Dai J, Tang H, Xu T, Yu S, Si L, et al. MicroRNA-23a-3p Inhibits Mucosal Melanoma Growth and Progression Through Targeting Adenylate Cyclase 1 and Attenuating CAMP and MAPK Pathways. Theranostics (2019) 9(4):945. doi: 10.7150/thno.30516
109. Alderman C, Sehlaoui A, Xiao Z, Yang Y. MicroRNA-15a Inhibits the Growth and Invasiveness of Malignant Melanoma and Directly Targets on CDCA4 Gene. Tumor Biol (2016) 37(10):13941–50. doi: 10.1007/s13277-016-5271-z
110. Panza E, Ercolano G, De Cicco P, Armogida C, Scognamiglio G, Botti G, et al. MicroRNA-143-3p Inhibits Growth and Invasiveness of Melanoma Cells by Targeting Cyclooxygenase-2 and Inversely Correlates With Malignant Melanoma Progression. Biochem Pharmacol (2018) 156:52–9. doi: 10.1016/j.bcp.2018.08.008
111. Nabipoorashrafi SA, Shomali N, Sadat-Hatamnezhad L, Mahami-Oskouei M, Mahmoudi J, Sandoghchian Shotorbani B, et al. miR-143 Acts as an Inhibitor of Migration and Proliferation as Well as an Inducer of Apoptosis in Melanoma Cancer Cells in Vitro. IUBMB Life (2020) 72(9):2034–44. doi: 10.1002/iub.2345
112. Lu H-J, Yan J, Jin P-Y, Zheng G-H, Zhang H-L, Bai M, et al. Mechanism of MicroRNA-708 Targeting BAMBI in Cell Proliferation, Migration, and Apoptosis in Mice With Melanoma Via the Wnt and TGF-β Signaling Pathways. Technol Cancer Res Treat (2018) 17:1533034618756784. doi: 10.1177/1533034618756784
113. Song X-F, Wang Q-H, Huo R. Effects of MicroRNA-708 on Epithelial-Mesenchymal Transition, Cell Proliferation and Apoptosis in Melanoma Cells by Targeting LEF1 Through the Wnt Signaling Pathway. Pathol Oncol Res (2019) 25(1):377–89. doi: 10.1007/s12253-017-0334-z
114. Sun M, Wang X, Tu C, Wang S, Qu J, Xiao S. MicroRNA-216b Inhibits Cell Proliferation and Migration in Human Melanoma by Targeting FOXM1 in Vitro and in Vivo. Cell Biol Int (2017) 41(12):1272–82. doi: 10.1002/cbin.10754
115. Liu Y, Huo Y, Wang D, Tai Y, Li J, Pang D, et al. MiR-216a-5p/Hexokinase 2 Axis Regulates Uveal Melanoma Growth Through Modulation of Warburg Effect. Biochem Biophys Res Commun (2018) 501(4):885–92. doi: 10.1016/j.bbrc.2018.05.069
116. Yang X, Zhao H, Yang J, Ma Y, Liu Z, Li C, et al. MiR-150-5p Regulates Melanoma Proliferation, Invasion and Metastasis Via SIX1-mediated Warburg Effect. Biochem Biophys Res Commun (2019) 515(1):85–91. doi: 10.1016/j.bbrc.2019.05.111
117. Sun X, Zhang C, Cao Y, Liu E. miR-150 Suppresses Tumor Growth in Melanoma Through Downregulation of MYB. Oncol Res (2019) 27(3):317. doi: 10.3727/096504018X15228863026239
118. Tembe V, Schramm SJ, Stark MS, Patrick E, Jayaswal V, Tang YH, et al. MicroRNA and MRNA Expression Profiling in Metastatic Melanoma Reveal Associations With BRAF Mutation and Patient Prognosis. Pigment Cell Melanoma Res (2015) 28(3):254–66. doi: 10.1111/pcmr.12343
119. Wang J-J, Li Z-F, Li X-J, Han Z, Zhang L, Liu Z-J. Effects of MicroRNA-136 on Melanoma Cell Proliferation, Apoptosis, and Epithelial–Mesenchymal Transition by Targetting PMEL Through the Wnt Signaling Pathway. Biosci Rep (2017) 37(5):BSR20170743. doi: 10.1042/BSR20170743
120. Prabhakar K, Rodrίguez CI, Jayanthy AS, Mikheil DM, Bhasker AI, Perera RJ, et al. Role of miR-214 in Regulation of β-Catenin and the Malignant Phenotype of Melanoma. Mol Carcinogen (2019) 58(11):1974–84. doi: 10.1002/mc.23089
121. Zhang J, Na S, Liu C, Pan S, Cai J, Qiu J. MicroRNA-125b Suppresses the Epithelial–Mesenchymal Transition and Cell Invasion by Targeting ITGA9 in Melanoma. Tumor Biol (2016) 37(5):5941–9. doi: 10.1007/s13277-015-4409-8
122. Zhang J, Lu L, Xiong Y, Qin W, Zhang Y, Qian Y, et al. MLK 3 Promotes Melanoma Proliferation and Invasion and is a Target of Micro RNA-125b. Clin Exp Dermatol (2014) 39(3):376–84. doi: 10.1111/ced.12286
123. Kappelmann M, Kuphal S, Meister G, Vardimon L, Bosserhoff A. MicroRNA miR-125b Controls Melanoma Progression by Direct Regulation of c-Jun Protein Expression. Oncogene (2013) 32(24):2984–91. doi: 10.1038/onc.2012.307
124. Essa S, Denzer N, Mahlknecht U, Klein R, Collnot E, Tilgen W, et al. VDR MicroRNA Expression and Epigenetic Silencing of Vitamin D Signaling in Melanoma Cells. J Steroid Biochem Mol Biol (2010) 121(1–2):110–3. doi: 10.1016/j.jsbmb.2010.02.003
125. Sánchez-Sendra B, Martinez-Ciarpaglini C, González-Muñoz JF, Murgui A, Terrádez L, Monteagudo C. Downregulation of Intratumoral Expression of miR-205, miR-200c and miR-125b in Primary Human Cutaneous Melanomas Predicts Shorter Survival. Sci Rep (2018) 8(1):1–14. doi: 10.1038/s41598-018-35317-3
126. Liu S-M, Lin C-H, Lu J, Lin I-Y, Tsai M-S, Chen M-H, et al. miR-596 Modulates Melanoma Growth by Regulating Cell Survival and Death. J Invest Dermatol (2018) 138(4):911–21. doi: 10.1016/j.jid.2017.11.016
127. Yu H, Ma M, Wang X, Zhou Z, Li R, Guo Q. Propofol Suppresses Proliferation, Invasion, and Migration of Human Melanoma Cells Via Regulating MicroRNA-137 and Fibroblast Growth Factor 9. J Cell Physiol (2019) 234(12):23279–88. doi: 10.1002/jcp.28896
128. Yang H, Shen C. MicroRNA-29c Induces G1 Arrest of Melanoma by Targeting CDK6. J BUON (2019) 24:819–25.
129. Arnold J, Engelmann JC, Schneider N, Bosserhoff AK, Kuphal S. miR-488-5p and Its Role in Melanoma. Exp Mol Pathol (2020) 112:104348. doi: 10.1016/j.yexmp.2019.104348
130. Li N, Ma Y, Ma L, Guan Y, Ma L, Yang D. MicroRNA-488-3p Sensitizes Malignant Melanoma Cells to Cisplatin by Targeting PRKDC. Cell Biol Int (2017) 41(6):622–9. doi: 10.1002/cbin.10765
131. Liu K, Jin J, Rong K, Zhuo L, Li P. MicroRNA−675 Inhibits Cell Proliferation and Invasion in Melanoma by Directly Targeting Metadherin. Mol Med Rep (2018) 17(2):3372–9. doi: 10.3892/mmr.2017.8264
132. Dietrich P, Kuphal S, Spruss T, Hellerbrand C, Bosserhoff AK. Micro RNA-622 is a Novel Mediator of Tumorigenicity in Melanoma by Targeting Kirsten Rat Sarcoma. Pigment Cell Melanoma Res (2018) 31(5):614–29. doi: 10.1111/pcmr.12698
133. Shidal C, Singh NP, Nagarkatti P, Nagarkatti M. MicroRNA-92 Expression in CD133+ Melanoma Stem Cells Regulates Immunosuppression in the Tumor Microenvironment Via Integrin-Dependent Activation of Tgfβ. Cancer Res (2019) 79(14):3622–35. doi: 10.1158/0008-5472.CAN-18-2659
134. Noguchi S, Kumazaki M, Mori T, Baba K, Okuda M, Mizuno T, et al. Analysis of MicroRNA-203 Function in CREB/MITF/RAB27a Pathway: Comparison Between Canine and Human Melanoma Cells. Vet Comp Oncol (2016) 14(4):384–94. doi: 10.1111/vco.12118
135. Chang X, Sun Y, Han S, Zhu W, Zhang H, Lian S. MiR-203 Inhibits Melanoma Invasive and Proliferative Abilities by Targeting the Polycomb Group Gene BMI1. Biochem Biophys Res Commun (2015) 456(1):361–6. doi: 10.1016/j.bbrc.2014.11.087
136. Wang K, Zhang Z-W. Expression of miR-203 is Decreased and Associated With the Prognosis of Melanoma Patients. Int J Clin Exp Pathol (2015) 8(10):13249.
137. Noguchi S, Kumazaki M, Yasui Y, Mori T, Yamada N, Akao Y. MicroRNA-203 Regulates Melanosome Transport and Tyrosinase Expression in Melanoma Cells by Targeting Kinesin Superfamily Protein 5b. J Invest Dermatol (2014) 134(2):461–9. doi: 10.1038/jid.2013.310
138. Noguchi S, Mori T, Otsuka Y, Yamada N, Yasui Y, Iwasaki J, et al. Anti-Oncogenic MicroRNA-203 Induces Senescence by Targeting E2F3 Protein in Human Melanoma Cells. J Biol Chem (2012) 287(15):11769–77. doi: 10.1074/jbc.M111.325027
139. Bu P, Yang P. MicroRNA−203 Inhibits Malignant Melanoma Cell Migration by Targeting Versican. Exp Ther Med (2014) 8(1):309–15. doi: 10.3892/etm.2014.1708
140. Wu S, Chen H, Han N, Zhang C, Yan H. Long Noncoding RNA PVT1 Silencing Prevents the Development of Uveal Melanoma by Impairing MicroRNA-17-3p–Dependent MDM2 Upregulation. Invest Ophthalmol Visual Sci (2019) 60(14):4904–14. doi: 10.1167/iovs.19-27704
141. Luan W, Zhou Z, Zhu Y, Xia Y, Wang J, Xu B. miR-137 Inhibits Glutamine Catabolism and Growth of Malignant Melanoma by Targeting Glutaminase. Biochem Biophys Res Commun (2018) 495(1):46–52. doi: 10.1016/j.bbrc.2017.10.152
142. Chen X, Wang J, Shen H, Lu J, Li C, Hu D-N, et al. Epigenetics, MicroRNAs, and Carcinogenesis: Functional Role of MicroRNA-137 in Uveal Melanoma. Invest Ophthalmol Visual Sci (2011) 52(3):1193–9. doi: 10.1167/iovs.10-5272
143. Lv N, Hao S, Luo C, Abukiwan A, Hao Y, Gai F, et al. miR-137 Inhibits Melanoma Cell Proliferation Through Downregulation of GLO1. Sci China Life Sci (2018) 61(5):541–9. doi: 10.1007/s11427-017-9138-9
144. Peres J, Kwesi-Maliepaard EM, Rambow F, Larue L, Prince S. The Tumour Suppressor, miR-137, Inhibits Malignant Melanoma Migration by Targetting the TBX3 Transcription Factor. Cancer Lett (2017) 405:111–9. doi: 10.1016/j.canlet.2017.07.018
145. Chang X, Zhang H, Lian S, Zhu W. miR-137 Suppresses Tumor Growth of Malignant Melanoma by Targeting Aurora Kinase a. Biochem Biophys Res Commun (2016) 475(3):251–6. doi: 10.1016/j.bbrc.2016.05.090
146. Li N. Low Expression of Mir-137 Predicts Poor Prognosis in Cutaneous Melanoma Patients. Med Sci Monit (2016) 22:140. doi: 10.12659/MSM.895207
147. Hao S, Luo C, Abukiwan A, Wang G, He J, Huang L, et al. miR-137 Inhibits Proliferation of Melanoma Cells by Targeting PAK 2. Exp Dermatol (2015) 24(12):947–52. doi: 10.1111/exd.12812
148. Deng Y, Deng H, Bi F, Liu J, Bemis LT, Norris D, et al. MicroRNA-137 Targets Carboxyl-Terminal Binding Protein 1 in Melanoma Cell Lines. Int J Biol Sci (2011) 7(1):133. doi: 10.7150/ijbs.7.133
149. Luo C, Tetteh PW, Merz PR, Dickes E, Abukiwan A, Hotz-Wagenblatt A, et al. miR-137 Inhibits the Invasion of Melanoma Cells Through Downregulation of Multiple Oncogenic Target Genes. J Invest Dermatol (2013) 133(3):768–75. doi: 10.1038/jid.2012.357
150. Qi J, Ww W, Chen W, Lu WY, Shang AQ. Mechanism of miR-137 Regulating Migration and Invasion of Melanoma Cells by Targeting PIK3R3 Gene. J Cell Biochem (2019) 120(5):8393–400. doi: 10.1002/jcb.28124
151. Liu E, Sun X, Li J, Zhang C. miR−30a−5p Inhibits the Proliferation, Migration and Invasion of Melanoma Cells by Targeting SOX4. Mol Med Rep (2018) 18(2):2492–8. doi: 10.3892/mmr.2018.9166
152. Wei Y, Du Y, Chen X, Li P, Wang Y, Zang W, et al. Expression Patterns of MicroRNA-218 and Its Potential Functions by Targeting CIP2A and BMI1 Genes in Melanoma. Tumor Biol (2014) 35(8):8007–15. doi: 10.1007/s13277-014-2079-6
153. Chen L, Cao Y, Rong D, Wang Y, Cao Y. MicroRNA-605 Functions as a Tumor Suppressor by Targeting INPP4B in Melanoma. Oncol Rep (2017) 38(2):1276–86. doi: 10.3892/or.2017.5740
154. Xiao Y, Diao Q, Liang Y, Peng Y, Zeng K. MicroRNA−24−1−5p Promotes Malignant Melanoma Cell Autophagy and Apoptosis Via Regulating Ubiquitin D. Mol Med Rep (2017) 16(6):8448–54. doi: 10.3892/mmr.2017.7614
155. Ge X, Niture S, Lin M, Cagle P, Li PA, Kumar D. MicroRNA-205-5p Inhibits Skin Cancer Cell Proliferation and Increase Drug Sensitivity by Targeting TNFAIP8. Sci Rep (2021) 11(1):1–14. doi: 10.1038/s41598-021-85097-6
156. Valentini V, Zelli V, Gaggiano E, Silvestri V, Rizzolo P, Bucalo A, et al. MiRNAs as Potential Prognostic Biomarkers for Metastasis in Thin and Thick Primary Cutaneous Melanomas. Anticancer Res (2019) 39(8):4085–93. doi: 10.21873/anticanres.13566
157. Liu S, Tetzlaff MT, Liu A, Liegl-Atzwanger B, Guo J, Xu X. Loss of MicroRNA-205 Expression is Associated With Melanoma Progression. Lab Invest (2012) 92(7):1084–96. doi: 10.1038/labinvest.2012.62
158. Li Y, Luo J-T, Liu Y-M, Wei W-B. MiRNA-145/miRNA-205 Inhibits Proliferation and Invasion of Uveal Melanoma Cells by Targeting NPR1/CDC42. Int J Ophthalmol (2020) 13(5):718. doi: 10.18240/ijo.2020.05.04
159. Liu S, Gao G, Yan D, Chen X, Yao X, Guo S, et al. Effects of miR-145-5p Through NRAS on the Cell Proliferation, Apoptosis, Migration, and Invasion in Melanoma by Inhibiting MAPK and PI 3K/AKT Pathways. Cancer Med (2017) 6(4):819–33. doi: 10.1002/cam4.1030
160. Jin C, Wang A, Liu L, Wang G, Li G, Han Z. miR-145-5p Inhibits Tumor Occurrence and Metastasis Through the NF-κb Signaling Pathway by Targeting TLR4 in Malignant Melanoma. J Cell Biochem (2019) 120(7):11115–26. doi: 10.1002/jcb.28388
161. Yang J-Y, Li Y, Wang Q, Zhou W-J, Yan Y-N, Wei W-B. MicroRNA-145 Suppresses Uveal Melanoma Angiogenesis and Growth by Targeting Neuroblastoma RAS Viral Oncogene Homolog and Vascular Endothelial Growth Factor. Chin Med J (2020) 133(16):1922. doi: 10.1097/CM9.0000000000000875
162. Cirilo PDR, de Sousa Andrade LN, Corrêa BRS, Qiao M, Furuya TK, Chammas R, et al. MicroRNA-195 Acts as an Anti-Proliferative MiRNA in Human Melanoma Cells by Targeting Prohibitin 1. BMC Cancer (2017) 17(1):1–12. doi: 10.1186/s12885-017-3721-7
163. Li N, Liu Y, Pang H, Lee D, Zhou Y, Xiao Z. Methylation-Mediated Silencing of MicroRNA-211 Decreases the Sensitivity of Melanoma Cells to Cisplatin. Med Sci Monit (2019) 25:1590. doi: 10.12659/MSM.911862
164. Mazar J, Qi F, Lee B, Marchica J, Govindarajan S, Shelley J, et al. MicroRNA 211 Functions as a Metabolic Switch in Human Melanoma Cells. Mol Cell Biol (2016) 36(7):1090–108. doi: 10.1128/MCB.00762-15
165. Mazar J, DeYoung K, Khaitan D, Meister E, Almodovar A, Goydos J, et al. The Regulation of MiRNA-211 Expression and Its Role in Melanoma Cell Invasiveness. PloS One (2010) 5(11):e13779. doi: 10.1371/journal.pone.0013779
166. Yu H, Yang W. MiR-211 is Epigenetically Regulated by DNMT1 Mediated Methylation and Inhibits EMT of Melanoma Cells by Targeting RAB22A. Biochem Biophys Res Commun (2016) 476(4):400–5. doi: 10.1016/j.bbrc.2016.05.133
167. Sakurai E, Maesawa C, Shibazaki M, Yasuhira S, Oikawa H, Sato M, et al. Downregulation of MicroRNA-211 is Involved in Expression of Preferentially Expressed Antigen of Melanoma in Melanoma Cells. Int J Oncol (2011) 39(3):665–72. doi: 10.3892/ijo.2011.1084
168. Boyle GM, Woods SL, Bonazzi VF, Stark MS, Hacker E, Aoude LG, et al. Melanoma Cell Invasiveness is Regulated by miR-211 Suppression of the BRN2 Transcription Factor. Pigment Cell Melanoma Res (2011) 24(3):525–37. doi: 10.1111/j.1755-148X.2011.00849.x
169. Bell RE, Khaled M, Netanely D, Schubert S, Golan T, Buxbaum A, et al. Transcription Factor/MicroRNA Axis Blocks Melanoma Invasion Program by miR-211 Targeting NUAK1. J Invest Dermatol (2014) 134(2):441–51. doi: 10.1038/jid.2013.340
170. Levy C, Khaled M, Iliopoulos D, Janas MM, Schubert S, Pinner S, et al. Intronic miR-211 Assumes the Tumor Suppressive Function of Its Host Gene in Melanoma. Mol Cell (2010) 40(5):841–9. doi: 10.1016/j.molcel.2010.11.020
171. Du M, Zhang Z, Gao T. Piceatannol Induced Apoptosis Through Up-Regulation of MicroRNA-181a in Melanoma Cells. Biol Res (2017) 50. doi: 10.1186/s40659-017-0141-8
172. Barbato A, Iuliano A, Volpe M, D’Alterio R, Brillante S, Massa F, et al. Integrated Genomics Identifies Mir-181/TFAM Pathway as a Critical Driver of Drug Resistance in Melanoma. Int J Mol Sci (2021) 22(4):1801. doi: 10.3390/ijms22041801
173. Mazar J, DeBlasio D, Govindarajan SS, Zhang S, Perera RJ. Epigenetic Regulation of MicroRNA-375 and Its Role in Melanoma Development in Humans. FEBS Lett (2011) 585(15):2467–76. doi: 10.1016/j.febslet.2011.06.025
174. Li J-R, Wang J-Q, Gong Q, Fang R-H, Guo Y-L. MicroRNA-328 Inhibits Proliferation of Human Melanoma Cells by Targeting TGFB2. Asian Pac J Cancer Prev (2015) 16(4):1575–9. doi: 10.7314/APJCP.2015.16.4.1575
175. Zou B, Zhu W, Liu H, Wang S, Zhu H. Identification and Functional Evaluation of miR-4633-5p as a Biomarker and Tumor Suppressor in Metastatic Melanoma. Cell Physiol Biochem (2018) 49(4):1364–79. doi: 10.1159/000493414
176. Wang H, Yu L, Shan X. Expression Levels of MicroRNA−455 and Its Potential Functions by Targeting IGF−1R in Melanoma. Mol Med Rep (2017) 15(6):3852–8. doi: 10.3892/mmr.2017.6468
177. Noguchi S, Mori T, Hoshino Y, Yamada N, Nakagawa T, Sasaki N, et al. Comparative Study of Anti-Oncogenic MicroRNA-145 in Canine and Human Malignant Melanoma. J Vet Med Sci (2011) 74(1):1108090601. doi: 10.1292/jvms.11-0264
178. Yang L, Qiming H, Xuehui S, Xiang J, Li S, Xiaolin X, et al. MicroRNA 145 May Play an Important Role in Uveal Melanoma Cell Growth by Potentially Targeting Insulin Receptor Substrate-1. Chin Med J (2014) 127(8):1410–6. doi: 10.3760/cma.j.issn.0366-6999.20133206
179. Dynoodt P, Speeckaert R, De Wever O, Chevolet I, Brochez L, Lambert J, et al. miR-145 Overexpression Suppresses the Migration and Invasion of Metastatic Melanoma Cells. Int J Oncol (2013) 42(4):1443–51. doi: 10.3892/ijo.2013.1823
180. Long J, Menggen Q, Wuren Q, Shi Q, Pi X. MiR-219-5p Inhibits the Growth and Metastasis of Malignant Melanoma by Targeting BCL-2. BioMed Res Int (2017) 2017(9032502):7. doi: 10.1155/2017/9032502
181. Asangani IA, Harms PW, Dodson L, Pandhi M, Kunju LP, Maher CA, et al. Genetic and Epigenetic Loss of MicroRNA-31 Leads to Feed-Forward Expression of EZH2 in Melanoma. Oncotarget (2012) 3(9):1011. doi: 10.18632/oncotarget.622
182. Chen X, He D, Da Dong X, Dong F, Wang J, Wang L, et al. MicroRNA-124a is Epigenetically Regulated and Acts as a Tumor Suppressor by Controlling Multiple Targets in Uveal Melanoma. Invest Ophthalmol Visual Sci (2013) 54(3):2248–56. doi: 10.1167/iovs.12-10977
183. Zhang D, Han Y, Xu L. Upregulation of miR-124 by Physcion 8-O-β-Glucopyranoside Inhibits Proliferation and Invasion of Malignant Melanoma Cells Via Repressing RLIP76. Biomed Pharmacother (2016) 84:166–76. doi: 10.1016/j.biopha.2016.09.022
184. Yang P, Bu P, Li C. miR−124 Inhibits Proliferation, Migration and Invasion of Malignant Melanoma Cells Via Targeting Versican. Exp Ther Med (2017) 14(4):3555–62. doi: 10.3892/etm.2017.4998
185. Tian R, Liu T, Qiao L, Gao M, Li J. Decreased Serum MicroRNA-206 Level Predicts Unfavorable Prognosis in Patients With Melanoma. Int J Clin Exp Pathol (2015) 8(3):3097.
186. Georgantas RW II, Streicher K, Luo X, Greenlees L, Zhu W, Liu Z, et al. Micro RNA-206 Induces G 1 Arrest in Melanoma by Inhibition of CDK 4 and C Yclin D. Pigment Cell Melanoma Res (2014) 27(2):275–86. doi: 10.1111/pcmr.12200
187. Su B-B, Zhou S-W, Gan C-B, Zhang X-N. MiR-186 Inhibits Cell Proliferation and Invasion in Human Cutaneous Malignant Melanoma. J Cancer Res Ther (2018) 14(8):60. doi: 10.4103/0973-1482.157340
188. Li M, Long C, Yang G, Luo Y, Du H. MiR-26b Inhibits Melanoma Cell Proliferation and Enhances Apoptosis by Suppressing TRAF5-mediated MAPK Activation. Biochem Biophys Res Commun (2016) 471(3):361–7. doi: 10.1016/j.bbrc.2016.02.021
189. Mueller DW, Bosserhoff AK. MicroRNA miR-196a Controls Melanoma-Associated Genes by Regulating HOX-C8 Expression. Int J Cancer (2011) 129(5):1064–74. doi: 10.1002/ijc.25768
190. Braig S, Mueller DW, Rothhammer T, Bosserhoff A-K. MicroRNA miR-196a is a Central Regulator of HOX-B7 and BMP4 Expression in Malignant Melanoma. Cell Mol Life Sci (2010) 67(20):3535–48. doi: 10.1007/s00018-010-0394-7
191. Chen J, Feilotter HE, Paré GC, Zhang X, Pemberton JG, Garady C, et al. MicroRNA-193b Represses Cell Proliferation and Regulates Cyclin D1 in Melanoma. Am J Pathol (2010) 176(5):2520–9. doi: 10.2353/ajpath.2010.091061
192. Chen J, Zhang X, Lentz C, Abi-Daoud M, Paré GC, Yang X, et al. miR-193b Regulates Mcl-1 in Melanoma. Am J Pathol (2011) 179(5):2162–8. doi: 10.1016/j.ajpath.2011.07.010
193. Liu S, Tetzlaff MT, Cui R, Xu X. miR-200c Inhibits Melanoma Progression and Drug Resistance Through Down-Regulation of BMI-1. Am J Pathol (2012) 181(5):1823–35. doi: 10.1016/j.ajpath.2012.07.009
194. Bustos MA, Ono S, Marzese DM, Oyama T, Iida Y, Cheung G, et al. MiR-200a Regulates CDK4/6 Inhibitor Effect by Targeting CDK6 in Metastatic Melanoma. J Invest Dermatol (2017) 137(9):1955–64. doi: 10.1016/j.jid.2017.03.039
195. Chen W, Xu Y, Zhang X. Targeting GOLM1 by MicroRNA-200a in Melanoma Suppresses Cell Proliferation, Invasion and Migration Via Regulating PI3K/Akt Signaling Pathway and Epithelial-Mesenchymal Transition. Eur Rev Med Pharmacol Sci (2019) 23(16):6997–7007. doi: 10.26355/eurrev_201908_18740
196. Peng J, Liu H, Liu C. MiR-155 Promotes Uveal Melanoma Cell Proliferation and Invasion by Regulating NDFIP1 Expression. Technol Cancer Res Treat (2017) 16(6):1160–7. doi: 10.1177/1533034617737923
197. Levati L, Pagani E, Romani S, Castiglia D, Piccinni E, Covaciu C, et al. MicroRNA-155 Targets the SKI Gene in Human Melanoma Cell Lines. Pigment Cell Melanoma Res (2011) 24(3):538–50. doi: 10.1111/j.1755-148X.2011.00857.x
198. Li H, Song J, Chen H, Wang Q, Meng L, Li Y. MiR-155 Inhibits Proliferation, Invasion and Migration of Melanoma Via Targeting CBL. Eur Rev Med Pharmacol Sci (2019) 23(21):9525–34. doi: 10.26355/eurrev_201911_19447
199. Dar AA, Majid S, Rittsteuer C, de Semir D, Bezrookove V, Tong S, et al. The Role of miR-18b in MDM2-p53 Pathway Signaling and Melanoma Progression. J Natl Cancer Inst (2013) 105(6):433–42. doi: 10.1093/jnci/djt003
200. Chen Y, Zhang Z, Luo C, Chen Z, Zhou J. MicroRNA-18b Inhibits the Growth of Malignant Melanoma Via Inhibition of HIF-1α-Mediated Glycolysis. Oncol Rep (2016) 36(1):471–9. doi: 10.3892/or.2016.4824
201. Reuland SN, Smith SM, Bemis LT, Goldstein NB, Almeida AR, Partyka KA, et al. MicroRNA-26a is Strongly Downregulated in Melanoma and Induces Cell Death Through Repression of Silencer of Death Domains (SODD). J Invest Dermatol (2013) 133(5):1286–93. doi: 10.1038/jid.2012.400
202. Fu X, Meng Z, Liang W, Tian Y, Wang X, Han W, et al. miR-26a Enhances MiRNA Biogenesis by Targeting Lin28B and Zcchc11 to Suppress Tumor Growth and Metastasis. Oncogene (2014) 33(34):4296–306. doi: 10.1038/onc.2013.385
203. Qian H, Yang C, Yang Y. MicroRNA-26a Inhibits the Growth and Invasiveness of Malignant Melanoma and Directly Targets on MITF Gene. Cell Death Discovery (2017) 3(1):1–9. doi: 10.1038/cddiscovery.2017.28
204. Liu S, Kumar SM, Lu H, Liu A, Yang R, Pushparajan A, et al. MicroRNA-9 Up-Regulates E-Cadherin Through Inhibition of NF-κb1–Snail1 Pathway in Melanoma. J Pathol (2012) 226(1):61–72. doi: 10.1002/path.2964
205. Zhao G, Li Q, Wang A, Jiao J. YY1 Regulates Melanoma Tumorigenesis Through a miR-9~ RYBP Axis. J Exp Clin Cancer Res (2015) 34(1):1–11. doi: 10.1186/s13046-015-0177-y
206. Liu N, Sun Q, Chen J, Li J, Zeng Y, Zhai S, et al. MicroRNA-9 Suppresses Uveal Melanoma Cell Migration and Invasion Through the NF-κb1 Pathway. Oncol Rep (2012) 28(3):961–8. doi: 10.3892/or.2012.1905
207. Xu D, Chen X, He Q, Luo C. MicroRNA-9 Suppresses the Growth, Migration, and Invasion of Malignant Melanoma Cells Via Targeting NRP1. OncoTargets Ther (2016) 9:7047. doi: 10.2147/OTT.S107235
208. Bu P, Luo C, He Q, Yang P, Li X, Xu D. MicroRNA-9 Inhibits the Proliferation and Migration of Malignant Melanoma Cells Via Targeting Sirituin 1. Exp Ther Med (2017) 14(2):931–8. doi: 10.3892/etm.2017.4595
209. Lu N-H, Wei C-Y, Qi F-Z, Gu J-Y. Hsa-let-7b Suppresses Cell Proliferation by Targeting UHRF1 in Melanoma. Cancer Invest (2020) 38(1):52–60. doi: 10.1080/07357907.2019.1709482
210. Zhou Y, Zhang L, Fan J, Jia R, Song X, Xu X, et al. Let-7b Overexpression Leads to Increased Radiosensitivity of Uveal Melanoma Cells. Melanoma Res (2015) 25(2):119–26. doi: 10.1097/CMR.0000000000000140
211. Tang H, Ma M, Dai J, Cui C, Si L, Sheng X, et al. miR-let-7b and miR-let-7c Suppress Tumourigenesis of Human Mucosal Melanoma and Enhance the Sensitivity to Chemotherapy. J Exp Clin Cancer Res (2019) 38(1):1–14. doi: 10.1186/s13046-019-1190-3
212. Yao Y, Zuo J, Wei Y. Targeting of TRX2 by miR-330-3p in Melanoma Inhibits Proliferation. Biomed Pharmacother (2018) 107:1020–9. doi: 10.1016/j.biopha.2018.08.058
213. Su B-B, Zhou S-W, Gan C-B, Zhang X-N. MiR-330-5p Regulates Tyrosinase and PDIA3 Expression and Suppresses Cell Proliferation and Invasion in Cutaneous Malignant Melanoma. J Surg Res (2016) 203(2):434–40. doi: 10.1016/j.jss.2016.03.021
214. Sun Y, Cheng H, Wang G, Yu G, Zhang D, Wang Y, et al. Deregulation of miR-183 Promotes Melanoma Development Via LncRNA MALAT1 Regulation and ITGB1 Signal Activation. Oncotarget (2017) 8(2):3509. doi: 10.18632/oncotarget.13862
215. Sun L, Bian G, Meng Z, Dang G, Shi D, Mi S. MiR-144 Inhibits Uveal Melanoma Cell Proliferation and Invasion by Regulating c-Met Expression. PloS One (2015) 10(5):e0124428. doi: 10.1371/journal.pone.0124428
216. Amaro A, Croce M, Ferrini S, Barisione G, Gualco M, Perri P, et al. Potential Onco-Suppressive Role of miR122 and miR144 in Uveal Melanoma Through ADAM10 and C-Met Inhibition. Cancers (2020) 12(6):1468. doi: 10.3390/cancers12061468
217. Zhao G, Wei Z, Guo Y. MicroRNA-107 is a Novel Tumor Suppressor Targeting POU3F2 in Melanoma. Biol Res (2020) 53(1):1–10. doi: 10.1186/s40659-020-00278-3
218. Wang X, Hu Y, Cui J, Zhou Y, Chen L. Coordinated Targeting of MMP-2/MMP-9 by Mir-296-3p/FOXCUT Exerts Tumor-Suppressing Effects in Choroidal Malignant Melanoma. Mol Cell Biochem (2018) 445(1):25–33. doi: 10.1007/s11010-017-3248-x
219. Rang Z, Yang G, Y-w W, Cui F. miR-542-3p Suppresses Invasion and Metastasis by Targeting the Proto-Oncogene Serine/Threonine Protein Kinase, PIM1, in Melanoma. Biochem Biophys Res Commun (2016) 474(2):315–20. doi: 10.1016/j.bbrc.2016.04.093
220. Zhang H, Feng C, Zhang M, Zeng A, Si L, Yu N, et al. Mir-625-5p/PKM2 Negatively Regulates Melanoma Glycolysis State. J Cell Biochem (2019) 120(3):2964–72. doi: 10.1002/jcb.26917
221. Weber CE, Luo C, Hotz-Wagenblatt A, Gardyan A, Kordaß T, Holland-Letz T, et al. miR-339-3p is a Tumor Suppressor in Melanoma. Cancer Res (2016) 76(12):3562–71. doi: 10.1158/0008-5472.CAN-15-2932
222. Mou K, Ding M, Han D, Zhou Y, Mu X, Liu W, et al. miR−590−5p Inhibits Tumor Growth in Malignant Melanoma by Suppressing YAP1 Expression. Oncol Rep (2018) 40(4):2056–66. doi: 10.3892/or.2018.6633
223. Jiang C, Croft A, Tseng H, Guo S, Jin L, Hersey P, et al. Repression of MicroRNA-768-3p by MEK/ERK Signalling Contributes to Enhanced MRNA Translation in Human Melanoma. Oncogene (2014) 33(20):2577–88. doi: 10.1038/onc.2013.237
224. Babapoor S, Fleming E, Wu R, Dadras SS. A Novel miR-451a isomiR, Associated With Amelanotypic Phenotype, Acts as a Tumor Suppressor in Melanoma by Retarding Cell Migration and Invasion. PloS One (2014) 9(9):e107502. doi: 10.1371/journal.pone.0107502
225. Mishra PJ, Mishra PJ, Merlino G. Integrated Genomics Identifies Mir-32/MCL-1 Pathway as a Critical Driver of Melanomagenesis: Implications for miR-replacement and Combination Therapy. PloS One (2016) 11(11):e0165102. doi: 10.1371/journal.pone.0165102
226. Cui A, Jin Z, Gao Z, Jin M, Zhu L, Li L, et al. Downregulation of miR-493 Promoted Melanoma Proliferation by Suppressing IRS4 Expression. Tumor Biol (2017) 39(5):1010428317701640. doi: 10.1177/1010428317701640
227. Hanniford D, Segura MF, Zhong J, Philips E, Jirau-Serrano X, Darvishian F, et al. Identification of Metastasis-Suppressive MicroRNAs in Primary Melanoma. J Natl Cancer Inst (2015) 107(3):dju494. doi: 10.1093/jnci/dju494
228. Guo B, Hui Q, Zhang Y, Chang P, Tao K. miR-194 is a Negative Regulator of GEF-H1 Pathway in Melanoma. Oncol Rep (2016) 36(4):2412–20. doi: 10.3892/or.2016.5020
229. Bai M, Zhang M, Long F, Yu N, Zeng A, Zhao R. Circulating MicroRNA-194 Regulates Human Melanoma Cells Via PI3K/AKT/FoxO3a and p53/p21 Signaling Pathway. Oncol Rep (2017) 37(5):2702–10. doi: 10.3892/or.2017.5537
230. Song H, Tao Y, Ni N, Zhou X, Xiong J, Zeng X, et al. miR-128 Targets the CC Chemokine Ligand 18 Gene (CCL18) in Cutaneous Malignant Melanoma Progression. J Dermatol Sci (2018) 91(3):317–24. doi: 10.1016/j.jdermsci.2018.06.011
231. Sun V, Zhou WB, Nosrati M, Majid S, Thummala S, De Semir D, et al. Antitumor Activity of miR-1280 in Melanoma by Regulation of Src. Mol Ther (2015) 23(1):71–8. doi: 10.1038/mt.2014.176
232. Wang H-F, Chen H, Ma M-W, Wang J-A, Tang T-T, Ni L-S, et al. miR-573 Regulates Melanoma Progression by Targeting the Melanoma Cell Adhesion Molecule. Oncol Rep (2013) 30(1):520–6. doi: 10.3892/or.2013.2451
233. Zhang Z, Yang N. MiR-33a-5p Inhibits the Growth and Metastasis of Melanoma Cells by Targeting SNAI2. Neoplasma (2020) 67(4):813–24. doi: 10.4149/neo_2020_190823N811
234. Cao K, Li J, Chen J, Qian L, Wang A, Chen X, et al. MicroRNA-33a-5p Increases Radiosensitivity by Inhibiting Glycolysis in Melanoma. Oncotarget (2017) 8(48):83660. doi: 10.18632/oncotarget.19014
235. Zhou J, Xu D, Xie H, Tang J, Liu R, Li J, et al. miR-33a Functions as a Tumor Suppressor in Melanoma by Targeting HIF-1α. Cancer Biol Ther (2015) 16(6):846–55. doi: 10.1080/15384047.2015.1030545
236. Tian F, Wei H, Tian H, Qiu Y, Xu J. miR-33a is Downregulated in Melanoma Cells and Modulates Cell Proliferation by Targeting PCTAIRE1. Oncol Lett (2016) 11(4):2741–6. doi: 10.3892/ol.2016.4321
237. Zhao Y, Wu C, Li L. MicroRNA−33b Inhibits Cell Proliferation and Glycolysis by Targeting Hypoxia−Inducible Factor−1α in Malignant Melanoma. Exp Ther Med (2017) 14(2):1299–306. doi: 10.3892/etm.2017.4702
238. Li F, Li X-J, Qiao L, Shi F, Liu W, Li Y, et al. miR-98 Suppresses Melanoma Metastasis Through a Negative Feedback Loop With Its Target Gene IL-6. Exp Mol Med (2014) 46(10):e116–e. doi: 10.1038/emm.2014.63
239. Liu P, Hu Y, Ma L, Du M, Xia L, Hu Z. miR-425 Inhibits Melanoma Metastasis Through Repression of PI3K-Akt Pathway by Targeting IGF-1. Biomed Pharmacother (2015) 75:51–7. doi: 10.1016/j.biopha.2015.08.010
240. Xiao W, Yao E, Zheng W, Tian F, Tian L. miR-337 Can be a Key Negative Regulator in Melanoma. Cancer Biol Ther (2017) 18(6):392–9. doi: 10.1080/15384047.2017.1323581
241. Zhang J, Liu W-L, Zhang L, Ge R, He F, Gao T-Y, et al. MiR-637 Suppresses Melanoma Progression Through Directly Targeting P-REX2a and Inhibiting PTEN/AKT Signaling Pathway. Cell Mol Biol (2018) 64(11):50–7. doi: 10.14715/cmb/2018.64.11.10
242. Mo Y, Fang RH, Wu J, Si Y, Jia SQ, Li Q, et al. MicroRNA-329 Upregulation Impairs the HMGB2/β-Catenin Pathway and Regulates Cell Biological Behaviors in Melanoma. J Cell Physiol (2019) 234(12):23518–27. doi: 10.1002/jcp.28920
243. Fattore L, Mancini R, Acunzo M, Romano G, Laganà A, Pisanu ME, et al. miR-579-3p Controls Melanoma Progression and Resistance to Target Therapy. Proc Natl Acad Sci (2016) 113(34):E5005–13. doi: 10.1073/pnas.1607753113
244. Luo C, Merz PR, Chen Y, Dickes E, Pscherer A, Schadendorf D, et al. MiR-101 Inhibits Melanoma Cell Invasion and Proliferation by Targeting MITF and EZH2. Cancer Lett (2013) 341(2):240–7. doi: 10.1016/j.canlet.2013.08.021
245. Ding Z, Jian S, Peng X, Liu Y, Wang J, Zheng L, et al. Loss of MiR-664 Expression Enhances Cutaneous Malignant Melanoma Proliferation by Upregulating PLP2. Medicine (2015) 94(33):e1327. doi: 10.1097/MD.0000000000001327
246. Xiong Y, Liu L, Qiu Y, Liu L. MicroRNA-29a Inhibits Growth, Migration and Invasion of Melanoma A375 Cells in Vitro by Directly Targeting BMI1. Cell Physiol Biochem (2018) 50(1):385–97. doi: 10.1159/000494015
247. Liu S-M, Lu J, Lee H-C, Chung F-H, Ma N. miR-524-5p Suppresses the Growth of Oncogenic BRAF Melanoma by Targeting BRAF and ERK2. Oncotarget (2014) 5(19):9444. doi: 10.18632/oncotarget.2452
248. Meng F, Zhang Y, Li X, Wang J, Wang Z. Clinical Significance of miR-138 in Patients With Malignant Melanoma Through Targeting of PDK1 in the PI3K/AKT Autophagy Signaling Pathway. Oncol Rep (2017) 38(3):1655–62. doi: 10.3892/or.2017.5838
249. Chen Y, Cao K, Wang S, Chen J, He B, He G, et al. MicroRNA-138 Suppresses Proliferation, Invasion and Glycolysis in Malignant Melanoma Cells by Targeting HIF-1α. Exp Ther Med (2016) 11(6):2513–8. doi: 10.3892/etm.2016.3220
250. Qiu H, Chen F, Chen M. MicroRNA-138 Negatively Regulates the Hypoxia-Inducible Factor 1α to Suppress Melanoma Growth and Metastasis. Biol Open (2019) 8(8). doi: 10.1242/bio.042937
251. Lin N, Zhou Y, Lian X, Tu Y. Down-Regulation of Tissue MicroRNA-126 was Associated With Poor Prognosis in Patients With Cutaneous Melanoma. Int J Clin Exp Med (2015) 8(3):4297.
252. Felli N, Felicetti F, Lustri AM, Errico MC, Bottero L, Cannistraci A, et al. miR-126&126* Restored Expressions Play a Tumor Suppressor Role by Directly Regulating ADAM9 and MMP7 in Melanoma. PloS One (2013) 8(2):e56824. doi: 10.1371/journal.pone.0056824
253. Zehavi L, Schayek H, Jacob-Hirsch J, Sidi Y, Leibowitz-Amit R, Avni D. MiR-377 Targets E2F3 and Alters the NF-kB Signaling Pathway Through MAP3K7 in Malignant Melanoma. Mol Cancer (2015) 14(1):1–16. doi: 10.1186/s12943-015-0338-9
254. Yang C, Xia Z, Zhu L, Li Y, Zheng Z, Liang J, et al. MicroRNA-139-5p Modulates the Growth and Metastasis of Malignant Melanoma Cells Via the PI3K/AKT Signaling Pathway by Binding to IGF1R. Cell Cycle (2019) 18(24):3513–24. doi: 10.1080/15384101.2019.1690881
255. Shi Q, He Q, Wei J. MicroRNA-342 Prohibits Proliferation and Invasion of Melanoma Cells by Directly Targeting Zinc-Finger E-Box-Binding Homeobox 1. Oncol Res (2018) 26(9):1447. doi: 10.3727/096504018X15193823766141
256. Tian P, Tao L, Wang Y, Han X. MicroRNA-127 Inhibits the Progression of Melanoma by Downregulating Delta-Like Homologue 1. BioMed Res Int (2020) 2020(8523465):10. doi: 10.1155/2020/8523465
257. Shi L, Huo J, Chen S, Xue J, Gao W, Li X, et al. MicroRNA-22 Targets FMNL2 to Inhibit Melanoma Progression Via the Regulation of the Wnt/β-Catenin Signaling Pathway and Epithelial-Mesenchymal Transition. Eur Rev Med Pharmacol Sci (2019) 23(12):5332–42. doi: 10.26355/eurrev_201906_18200
258. Palkina N, Komina A, Aksenenko M, Moshev A, Savchenko A, Ruksha T. miR−204−5p and miR−3065−5p Exert Antitumor Effects on Melanoma Cells. Oncol Lett (2018) 15(6):8269–80. doi: 10.3892/ol.2018.8443
259. Luan W, Qian Y, Ni X, Bu X, Xia Y, Wang J, et al. miR-204-5p Acts as a Tumor Suppressor by Targeting Matrix Metalloproteinases-9 and B-Cell Lymphoma-2 in Malignant Melanoma. OncoTargets Ther (2017) 10:1237. doi: 10.2147/OTT.S128819
260. Zhang G, Ai D, Yang X, Ji S, Wang Z, Feng S. MicroRNA-610 Inhibits Tumor Growth of Melanoma by Targeting LRP6. Oncotarget (2017) 8(57):97361. doi: 10.18632/oncotarget.22125
261. Zhu L, Liu Z, Dong R, Wang X, Zhang M, Guo X, et al. MicroRNA-3662 Targets ZEB1 and Attenuates the Invasion of the Highly Aggressive Melanoma Cell Line A375. Cancer Manage Res (2019) 11:5845. doi: 10.2147/CMAR.S200540
262. Chen L, Ma G, Cao X, An X, Liu X. MicroRNA-331 Inhibits Proliferation and Invasion of Melanoma Cells by Targeting Astrocyte-Elevated gene-1. Oncol Res (2018) 26(9):1429. doi: 10.3727/096504018X15186047251584
263. Chen W, Zhang J, Xu H, Dai J, Zhang X. The Negative Regulation of miR-149-5p in Melanoma Cell Survival and Apoptosis by Targeting LRIG2. Am J Trans Res (2017) 9(9):4331.
264. Zhang C, Li H, Wang J, Zhang J, Hou X. MicroRNA−338−3p Suppresses Cell Proliferation, Migration and Invasion in Human Malignant Melanoma by Targeting MACC1. Exp Ther Med (2019) 18(2):997–1004. doi: 10.3892/etm.2019.7644
265. Zhou H, Rao Y, Sun Q, Liu Y, Zhou X, Chen Y, et al. MiR-4458/human Antigen R (HuR) Modulates PBX3 MRNA Stability in Melanoma Tumorigenesis. Arch Dermatol Res (2020) 312:665–73. doi: 10.1007/s00403-020-02051-8
266. Yang X, Zhu X, Yan Z, Li C, Zhao H, Ma L, et al. Mir-489-3p/SIX1 Axis Regulates Melanoma Proliferation and Glycolytic Potential. Mol Ther-Oncolyt (2020) 16:30–40. doi: 10.1016/j.omto.2019.11.001
267. Sun Y, Li X, Wang H, Wu J. MiR-431 is a Prognostic Marker and Suppresses Cell Growth, Migration and Invasion by Targeting NOTCH2 in Melanoma. Eur Rev Med Pharmacol Sci (2019) 23(9):3876–84. doi: 10.26355/eurrev_201905_17815
268. Li Y, Fu Y, Gao Y, Li H, Ma L, Shu C, et al. MicroRNA-134 Inhibits Melanoma Growth and Metastasis by Negatively Regulating Collagen Triple Helix Repeat Containing-1 (CTHRC1). Int J Clin Exp Pathol (2018) 11(9):4319.
269. Li J, Liu X, Li C, Wang W. miR-224-5p Inhibits Proliferation, Migration, and Invasion by Targeting PIK3R3/AKT3 in Uveal Melanoma. J Cell Biochem (2019) 120(8):12412–21. doi: 10.1002/jcb.28507
270. Zhao G, Yin Y, Zhao B. miR-140-5p is Negatively Correlated With Proliferation, Invasion, and Tumorigenesis in Malignant Melanoma by Targeting SOX4 Via the Wnt/β-Catenin and NF-κb Cascades. J Cell Physiol (2020) 235(3):2161–70. doi: 10.1002/jcp.29122
271. He Y, Yang Y, Liao Y, Xu J, Liu L, Li C, et al. miR-140-3p Inhibits Cutaneous Melanoma Progression by Disrupting AKT/p70S6K and JNK Pathways Through ABHD2. Mol Ther-Oncolyt (2020) 17:83–93. doi: 10.1016/j.omto.2020.03.009
272. Zhang X, Xin Z. MiR-135b-5p Inhibits the Progression of Malignant Melanoma Cells by Targeting RBX1. Eur Rev Med Pharmacol Sci (2020) 24(3):1309–15. doi: 10.21203/rs.3.rs-16546/v3
273. Hanniford D, Zhong J, Koetz L, Gaziel-Sovran A, Lackaye DJ, Shang S, et al. A MiRNA-based Signature Detected in Primary Melanoma Tissue Predicts Development of Brain Metastasis. Clin Cancer Res (2015) 21(21):4903–12. doi: 10.1158/1078-0432.CCR-14-2566
274. Worley LA, Long MD, Onken MD, Harbour JW. Micro-RNAs Associated With Metastasis in Uveal Melanoma Identified by Multiplexed Microarray Profiling. Melanoma Res (2008) 18(3):184–90. doi: 10.1097/CMR.0b013e3282feeac6
275. Bai M, Zhang H, Si L, Yu N, Zeng A, Zhao R. Upregulation of Serum miR-10b is Associated With Poor Prognosis in Patients With Melanoma. J Cancer (2017) 8(13):2487. doi: 10.7150/jca.18824
276. Caramuta S, Egyházi S, Rodolfo M, Witten D, Hansson J, Larsson C, et al. MicroRNA Expression Profiles Associated With Mutational Status and Survival in Malignant Melanoma. J Invest Dermatol (2010) 130(8):2062–70. doi: 10.1038/jid.2010.63
277. Falzone L, Romano GL, Salemi R, Bucolo C, Tomasello B, Lupo G, et al. Prognostic Significance of Deregulated MicroRNAs in Uveal Melanomas. Mol Med Rep (2019) 19(4):2599–610. doi: 10.3892/mmr.2019.9949
278. Segura MF, Belitskaya-Lévy I, Rose AE, Zakrzewski J, Gaziel A, Hanniford D, et al. Melanoma MicroRNA Signature Predicts Post-Recurrence Survival. Clin Cancer Res (2010) 16(5):1577–86. doi: 10.1158/1078-0432.CCR-09-2721
279. Nguyen T, Kuo C, Nicholl MB, Sim M-S, Turner RR, Morton DL, et al. Downregulation of MicroRNA-29c is Associated With Hypermethylation of Tumor-Related Genes and Disease Outcome in Cutaneous Melanoma. Epigenetics (2011) 6(3):388–94. doi: 10.4161/epi.6.3.14056
280. Hanna JA, Hahn L, Agarwal S, Rimm DL. In Situ Measurement of miR-205 in Malignant Melanoma Tissue Supports Its Role as a Tumor Suppressor Microrna. Lab Invest (2012) 92(10):1390–7. doi: 10.1038/labinvest.2012.119
281. Yan J, Jiang Q, Lu H, Na S, Long S, Xin Y, et al. Association Between MicroRNA−125b Expression in Formalin−Fixed Paraffin−Embedded Tumor Tissues and Prognosis in Patients With Melanoma. Oncol Lett (2019) 18(2):1856–62. doi: 10.3892/ol.2019.10506
282. Kozubek J, Ma Z, Fleming E, Duggan T, Wu R, Shin D-G, et al. In-Depth Characterization of MicroRNA Transcriptome in Melanoma. PloS One (2013) 8(9):e72699. doi: 10.1371/journal.pone.0072699
283. Babapoor S, Horwich M, Wu R, Levinson S, Gandhi M, Makkar H, et al. MicroRNA in Situ Hybridization for miR-211 Detection as an Ancillary Test in Melanoma Diagnosis. Modern Pathol (2016) 29(5):461–75. doi: 10.1038/modpathol.2016.44
284. Tengda L, Shuping L, Mingli G, Jie G, Yun L, Weiwei Z, et al. Serum Exosomal MicroRNAs as Potent Circulating Biomarkers for Melanoma. Melanoma Res (2018) 28(4):295–303. doi: 10.1097/CMR.0000000000000450
285. Fogli S, Polini B, Carpi S, Pardini B, Naccarati A, Dubbini N, et al. Identification of Plasma MicroRNAs as New Potential Biomarkers With High Diagnostic Power in Human Cutaneous Melanoma. Tumor Biol (2017) 39(5):1010428317701646. doi: 10.1177/1010428317701646
286. Leidinger P, Keller A, Borries A, Reichrath J, Rass K, Jager SU, et al. High-Throughput MiRNA Profiling of Human Melanoma Blood Samples. BMC Cancer (2010) 10(1):1–11. doi: 10.1186/1471-2407-10-262
287. Stark MS, Gray ES, Isaacs T, Chen FK, Millward M, McEvoy A, et al. A Panel of Circulating MicroRNAs Detects Uveal Melanoma With High Precision. Trans Vision Sci Technol (2019) 8(6):12–. doi: 10.1167/tvst.8.6.12
288. Armand-Labit V, Meyer N, Casanova A, Bonnabau H, Platzer V, Tournier E, et al. Identification of a Circulating MicroRNA Profile as a Biomarker of Metastatic Cutaneous Melanoma. Acta Dermato-Venereolog (2016) 96(1):29–34. doi: 10.2340/00015555-2156
289. Shiiyama R, Fukushima S, Jinnin M, Yamashita J, Miyashita A, Nakahara S, et al. Sensitive Detection of Melanoma Metastasis Using Circulating MicroRNA Expression Profiles. Melanoma Res (2013) 23(5):366–72. doi: 10.1097/CMR.0b013e328363e485
290. Xin X, Zhang Y, Ling F, Wang L, Sheng X, Qin L, et al. Identification of a Nine-MiRNA Signature for the Prognosis of Uveal Melanoma. Exp Eye Res (2019) 180:242–9. doi: 10.1016/j.exer.2019.01.004
291. Sun Q, Cong R, Yan H, Gu H, Zeng Y, Liu N, et al. Genistein Inhibits Growth of Human Uveal Melanoma Cells and Affects MicroRNA-27a and Target Gene Expression. Oncol Rep (2009) 22(3):563–7. doi: 10.3892/or_00000472
292. Galore-Haskel G, Nemlich Y, Greenberg E, Ashkenazi S, Hakim M, Itzhaki O, et al. A Novel Immune Resistance Mechanism of Melanoma Cells Controlled by the ADAR1 Enzyme. Oncotarget (2015) 6(30):28999. doi: 10.18632/oncotarget.4905
293. Varrone F, Caputo E. The MiRNAs Role in Melanoma and in Its Resistance to Therapy. Int J Mol Sci (2020) 21(3):878. doi: 10.3390/ijms21030878
294. Gomez-Lira M, Ferronato S, Orlandi E, Dal Molin A, Malerba G, Frigerio S, et al. Association of Micro RNA 146a Polymorphism rs2910164 and the Risk of Melanoma in an Italian Population. Exp Dermatol (2015) 24(10):794–5. doi: 10.1111/exd.12778
295. Sangalli A, Orlandi E, Poli A, Maurichi A, Santinami M, Nicolis M, et al. Sex-Specific Effect of RNASEL rs486907 and miR-146a rs2910164 Polymorphisms’ Interaction as a Susceptibility Factor for Melanoma Skin Cancer. Melanoma Res (2017) 27(4):309–14. doi: 10.1097/CMR.0000000000000360
296. Yamashita J, Iwakiri T, Fukushima S, Jinnin M, Miyashita A, Hamasaki T, et al. The rs2910164 G> C Polymorphism in MicroRNA-146a is Associated With the Incidence of Malignant Melanoma. Melanoma Res (2013) 23(1):13–20. doi: 10.1097/CMR.0b013e32835c5b30
297. Lodygin D, Tarasov V, Epanchintsev A, Berking C, Knyazeva T, Körner H, et al. Inactivation of miR-34a by Aberrant CpG Methylation in Multiple Types of Cancer. Cell Cycle (2008) 7(16):2591–600. doi: 10.4161/cc.7.16.6533
298. Ozsolak F, Poling LL, Wang Z, Liu H, Liu XS, Roeder RG, et al. Chromatin Structure Analyses Identify MiRNA Promoters. Genes Dev (2008) 22(22):3172–83. doi: 10.1101/gad.1706508
299. Syed D N, Imran Khan M, Shabbir M, Mukhtar H. MicroRNAs in Skin Response to UV Radiation. Curr Drug Targets (2013) 14(10):1128–34. doi: 10.2174/13894501113149990184
300. Bemis LT, Chen R, Amato CM, Classen EH, Robinson SE, Coffey DG, et al. MicroRNA-137 Targets Microphthalmia-Associated Transcription Factor in Melanoma Cell Lines. Cancer Res (2008) 68(5):1362–8. doi: 10.1158/0008-5472.CAN-07-2912
Keywords: miRNA, melanoma, biomarker, expression, polymorphism
Citation: Ghafouri-Fard S, Gholipour M and Taheri M (2021) MicroRNA Signature in Melanoma: Biomarkers and Therapeutic Targets. Front. Oncol. 11:608987. doi: 10.3389/fonc.2021.608987
Received: 22 September 2020; Accepted: 30 March 2021;
Published: 22 April 2021.
Edited by:
Giuseppe Palmieri, National Research Council (CNR), ItalyReviewed by:
Georg Wondrak, University of Arizona, United StatesEva Hernando, New York University, United States
Copyright © 2021 Ghafouri-Fard, Gholipour and Taheri. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Mohammad Taheri, bW9oYW1tYWRfODIzQHlhaG9vLmNvbQ==
 Soudeh Ghafouri-Fard
Soudeh Ghafouri-Fard Mahdi Gholipour
Mahdi Gholipour Mohammad Taheri
Mohammad Taheri