- Department of Nuclear Medicine, School of Medicine, Kyungpook National University and Hospital, Daegu, South Korea
Extracellular vesicles (EVs) are currently being considered as promising drug delivery vehicles. EVs are naturally occurring vesicles that exhibit many characteristics favorable to serve as drug delivery vehicles. In addition, EVs have inherent properties for treatment of cancers and other diseases. For research and clinical translation of use of EVs as drug delivery vehicles, in vivo tracking of EVs is essential. The latest molecular imaging techniques enable the tracking of EVs in living animals. However, each molecular imaging technique has its certain advantages and limitations for the in vivo imaging of EVs; therefore, understanding the molecular imaging techniques is essential to select the most appropriate imaging technology to achieve the desired imaging goal. In this review, we summarize the characteristics of EVs as drug delivery vehicles and the molecular imaging techniques used in visualizing and monitoring EVs in in vivo environments. Furthermore, we provide a perceptual vision of EVs as drug delivery vehicles and in vivo monitoring of EVs using molecular imaging technologies.
Introduction
Extracellular vesicles (EVs) are naturally occurring nanovesicles released by different types of cells, including reticulocytes (Johnstone et al., 1991), platelets (Brisson et al., 2017), mesenchymal stem cells (Rajendran et al., 2017), T cells (Karlsson et al., 2001; Ludwig et al., 2017), B lymphocytes (Raposo et al., 1996), NK cells (Shoae-Hassani et al., 2017; Zhu et al., 2017), dendritic cells (DCs) (Lu et al., 2017), and some tumor cells (Aharon et al., 2017; Baumgart et al., 2017; Schillaci et al., 2017); these nanovesicles can be detected in human biological fluids (Cappello et al., 2017). EVs include exosomes (small membranous vesicles) and microvesicles (large membranous vesicles) shed by cells (Di Rocco et al., 2016; Gangadaran et al., 2017a). In this article, we use the term “EVs” to refer to both exosomes and microvesicles.
EVs have recently gained attention as mediators of cellular communication (Srivastava et al., 2015). Several studies suggest that EVs are not merely secreted to provide a degradation route for unwanted biological materials (Johnstone et al., 1991) but are equipped to withstand lysis by the complement system to perform vital extracellular functions (Clayton et al., 2003).
EVs can carry various biological constituents such as lipids, proteins, and nucleic acids (Yáñez-Mó et al., 2015). These molecules are packed into EVs from the host cell cytoplasm through endosomal sorting complexes (Gangadaran et al., 2017a). These molecules and other contents of the EVs act on target cells. EVs from several cells induce apoptosis in tumors and induce anti-tumor immune responses (Filipazzi et al., 2012; Kalimuthu et al., 2016; Zhu et al., 2017). Moreover, serum-derived EVs and mesenchymal stem cell (MSC)-derived EVs increase the angiogenic activities of endothelial cells (Cavallari et al., 2017; Rajendran et al., 2017).
Accumulating studies suggest the importance of EVs in long distance cell-cell communication because the secreted EVs can enter the circulation and pass through additional biological barriers (Jiang and Gao, 2017; Sarko and McKinney, 2017). EVs are used as carriers for anticancer drugs (Agrawal et al., 2017), miRNA (Yang et al., 2017), and siRNA (Vader et al., 2017), and are now considered as promising drug transporters.
To utilize EVs as drug delivery vehicles, in vivo tracking of EVs to target organs is warranted. Non-invasive imaging modalities can provide accurate in vivo distribution and kinetics of the EVs and provide better understanding of the in vivo therapeutic effects of EVs. Recent advances in molecular imaging modalities allow us to well-recognize both cellular and subcellular biological processes within living subjects (Lee et al., 2016; Li et al., 2016). However, labeling procedures are necessary for accurate in vivo visualization of certain biomaterials in an animal model. EVs can be directly labeled using various agents such as, lipophilic tracer dyes (Ohno et al., 2013; Grange et al., 2014), radionuclides (Hwang et al., 2015), or magnetic particles (Piffoux et al., 2017) and indirectly labeled with a reporter gene (luciferase or fluoresce) by transducing the gene in originating cells (Koumangoye et al., 2011; Lai et al., 2014; Hoshino et al., 2015; Gangadaran et al., 2017b).
In this paper, we will discuss EVs as drug delivery vehicles, labeling techniques used for molecular imaging of EVs to evaluate their biodistribution and lesion targeting, merits and demerits of the labeling methods, and EV-based targeted drug delivery in diseases. Furthermore, we review the technology developments and strategies that led to the current state-of-the-art techniques used for EV visualization with specific in vitro and in vivo examples.
Biogenesis of EVs
EVs are nanosized membrane vesicles released by cells into the extracellular space and are found in various body fluids such as blood, urine, and central nervous system fluids. EVs are classified into exosomes and microvesicles; exosomes (50–200 nm) are membrane vesicles released by multi-vesicular bodies, whereas microvesicles (50–1,000 nm) are released from the cell membrane via the budding process and they are larger than exosomes (Figures 1A,B; Gangadaran et al., 2017a).
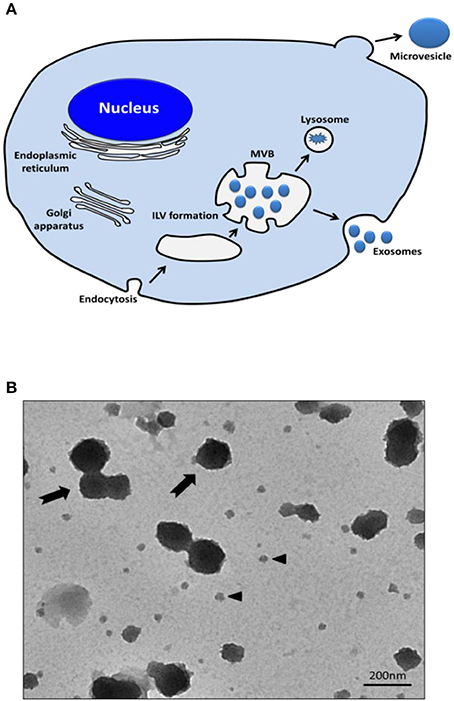
Figure 1. Release of exosomes and microvesicles. (A) Exosomes are represented by small vesicles of different sizes, released from MVB whereas microvesicles bud directly from the plasma membrane. (B) TEM images showing the typical structure of exosomes (black arrow head) and microvesicles (black arrows). MVB, Multi-vesicular bodies; ILV, intraluminal vesicles; TEM, transmission electron microscopy.
Several exosome production pathways have been identified. The endosomal sorting complex required for transport (ESCRT) and related proteins such as programmed cell death 6 interacting protein (PDCD6IP; also known as ALIX) and tumor susceptibility gene 101 (TSG101) protein are associated with the cargo sorting of exosomes. Moreover, ESCRT-independent mechanisms, such as ceramide-related pathway, also operate to generate exosomes of certain biochemical compositions (Trajkovic et al., 2008; Bobrie et al., 2011). Then, the exosomes are released from the cells by membrane fusion of the exosome-containing multivesicular body with the cell membrane. In contrast, microvesicles are formed by the outward budding and fission of the cell membrane. These processes are controlled by membrane lipid microdomains and regulatory proteins such as ADP-ribosylation factor 6 (Muralidharan-Chari et al., 2009). The membrane composition of microvesicles reflects that of the parent cell more closely than that of exosomes because of their different biogenesis mechanisms. Both exosomes and microvesicles contain various biological materials, but microvesicles are a relatively heterogenous population of vesicles, compared to exosomes (Théry et al., 2009; EL Andaloussi et al., 2013).
Recently, extracellular vesicle mimetics (EVMs), also called artificial nanovesicles, are being considered as new drug delivery vehicles. Large quantities of cells and culture medium are needed to obtain the desirable EVs; however, EVM production is less time-consuming and laborious (Kim et al., 2017b). EVMs were prepared by breaking down the cells through serial extrusion using nanosized filters with diminishing pore sizes and were isolated using density gradient ultracentrifugation (Jang et al., 2013).
EVs can carry various biological materials like lipids, proteins, mRNA, miRNA, and extra-chromosomal DNA. A recent study revealed that EVs contain 4,563 proteins, 194 lipids, 1,639 mRNAs, and 764 miRNAs (Mathivanan et al., 2012). The protein content of EVs is related to their originating cell type and their biogenesis. Recent proteomic study using EVs originated from DCs showed that exosomes are additionally characterized by the presence of the tetraspanins (CD9, CD63, and CD81), the ESCRT protein TSG101, and syntenin. In contrast, a number of factors such as class II major histocompatibility complex (MHCII), flotillin, or heat shock 70 kDa proteins were found in both exosomes and microvesicles (Kowal et al., 2016). EVM showed similar protein markers, such as CD9, ALIX, and TSG101. Membrane lipids of EVM were more similar to those of exosomes than those of their parent cells (Goh et al., 2017).
Until 1973, EVs were considered as a garbage materials released by the cells (Nolte-‘t Hoen et al., 2016). However, recent studies demonstrated that EVs are vital cell-to-cell communication messengers between distant cells and that EVs can attach to a cell surface or enter into recipient cells (Escrevente et al., 2011). EVs serve as a carrier and can transfer information from the parent cells to their target cells (Kalimuthu et al., 2016; Gangadaran et al., 2017c; Zhu et al., 2017).
Cells internalize EVs by various endocytic pathways, including clathrin-dependent endocytosis (Escrevente et al., 2011), caveolin-mediated uptake, micropinocytosis (Fitzner et al., 2011), phagocytosis (Hemler, 2003), and lipid raft-mediated internalization (Svensson et al., 2013; Figure 2A). EVs may enter the target cell via more than one route. The uptake mechanism may depend on the proteins and glycoproteins present on the surface of both EVs and the target cell (Mulcahy et al., 2014).
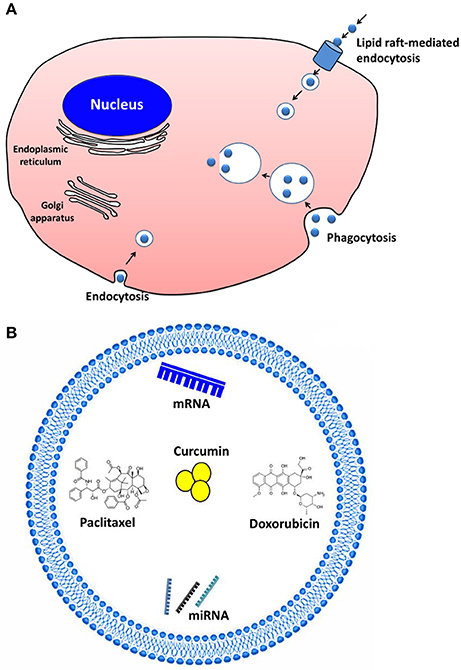
Figure 2. Extracellular vesicles (EVs) as drug delivery vehicles and their communication. (A) Cells internalize EVs by various endocytic pathways, including clathrin-dependent endocytosis, caveolin-mediated uptake, micropinocytosis, phagocytosis, and lipid raft–mediated internalization. (B) The therapeutic cargo can contain different types of interfering RNAs, mRNA, or even therapeutics (e.g., paclitaxel, doxorubicin, curcumin) to treat cancer and inflammatory diseases.
EVs as Drug Delivery Vehicles
Recently, the most studied drug delivery platform are liposomes and polymeric nanoparticles. Liposomes are tiny vesicles composed of fatty acid-containing bilayers surrounding an aqueous core, and they have established their roles as carriers for therapeutic drugs (Sercombe et al., 2015). Polymeric nanoparticles are drug delivery platform that help in the encapsulation, entrapment, or attaching the drug molecules (Kamaly et al., 2016). Both liposome and polymeric nanoparticles have been used to deliver various drug molecules, such as anti-cancer drugs (Hofheinz et al., 2005; Masood, 2016). However, synthesis of non-toxic liposomes with high stability, ability to circulate for a long time, and with the ability to evade the host immune system, remains a concern (Sercombe et al., 2015). Polymeric nanoparticles showed a better stability than liposomal systems but their biocompatibility needs to be evaluated (Li et al., 2015).
Several recent reports have shown the advantages of using EVs for drug delivery: (i) EVs are small and can penetrate into deep tissues (Gangadaran et al., 2017b); (ii) they possesses negative zeta potential for long circulation (Malhotra et al., 2016); (iii) EVs have membrane structure similar to that of cells (Hood and Wickline, 2012); (iv) EVs exhibit an increased ability to escape degradation (Luan et al., 2017); (v) EVs can evade the immune system (Anderson et al., 2016; Kamerkar et al., 2017). In addition, few human clinical trials performed using EVs from DCs for cancer therapy, reported positive results with respect to the feasibility and safety of EVs (Escudier et al., 2005; Morse et al., 2005; Besse et al., 2016). Overall, EVs are clinically applicable, excellent, natural carriers mainly because of their inherent biocompatibility.
However, to qualify as drug delivery vehicles, EVs should be able to carry a substantial amount of therapeutics. A variety of cargos have now been shown to exhibit therapeutic effect after EV-based delivery (Sun et al., 2010; Banizs et al., 2014; Pascucci et al., 2014; Tian et al., 2014; Figure 2B). EVs were reported to deliver the anti-inflammatory agent curcumin to activated myeloid cells in a mouse model. The curcumin delivered by EVs was more stable in vivo and more highly concentrated in the mouse blood (Sun et al., 2010). Other studies reported that EVs can be used for the delivery of pharmaceutical drugs like paclitaxel and doxorubicin to inhibit tumor growth both in vitro and in animal models (Pascucci et al., 2014; Tian et al., 2014). Furthermore, EVs were suggested to be useful as nanocarriers for exogenous siRNA to control gene expression in the recipient cells (Wahlgren et al., 2012; Banizs et al., 2014). Using EVs, an antitumor miRNA (let-7a miRNA) was delivered to EGFR-expressing xenograft breast cancer cells in mice; this delivery system efficiently inhibited the tumor growth (Ohno et al., 2013). In mouse models of pancreatic cancer, exosomes containing siRNA or shRNA specific to oncogenic KRAS showed increased therapeutic effects, compare to the controls including liposomes containing siRNA or shRNA (Kamerkar et al., 2017).
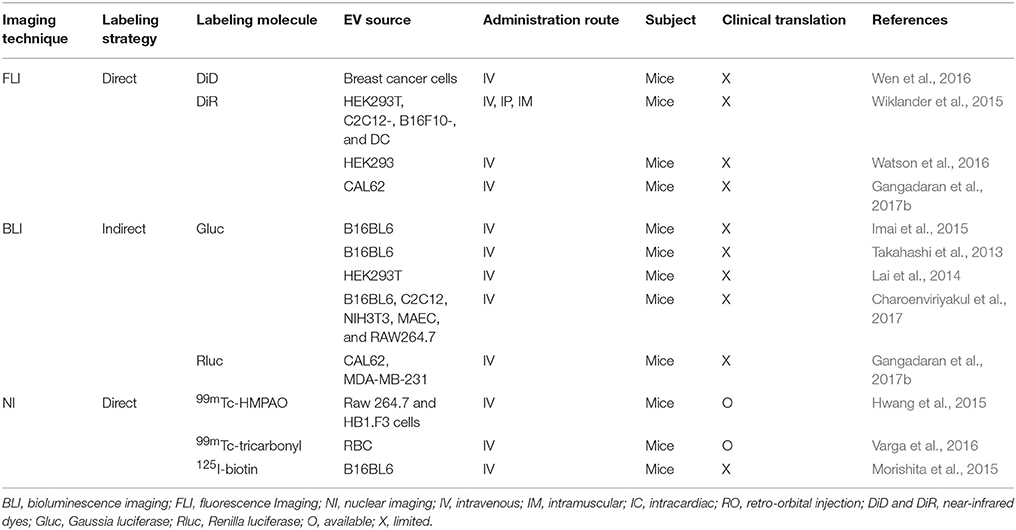
Although the use of EVs as systems to deliver therapeutic materials has been widely studied (Table 1), the effectiveness of EV-based therapy depends on the targetability of EVs to tumor or another desired cell in vivo. Non-invasive imaging modalities might provide clear view on the in vivo distribution of EVs and provide accurate targetability of EVs to the desired cell/tissue and would be useful in the development of EVs as drug delivery vehicles.
Investigating EVs by Molecular Imaging
Several molecular imaging strategies [optical, nuclear, and magnetic resonance imaging (MRI); Figures 3, 4] have been employed for in vitro, in vivo, and ex vivo tracking of EVs to determine their biodistribution in animal models and targeting of certain EVs via various delivery routes in small animal models for different diseases (Table 2). However, EVs derived from tumors or other cells influence the tumor itself as well as the tumor microenvironment, and the EVs are able to accelerate or inhibit growth and metastasis of the tumor (Hoshino et al., 2015; Liu and Cao, 2016; Schillaci et al., 2017; Wang et al., 2017). Therefore, a better understanding of EV biodistribution after administration is warranted for the safe and effective clinical application of EV-based therapies for various diseases (Lener et al., 2015).
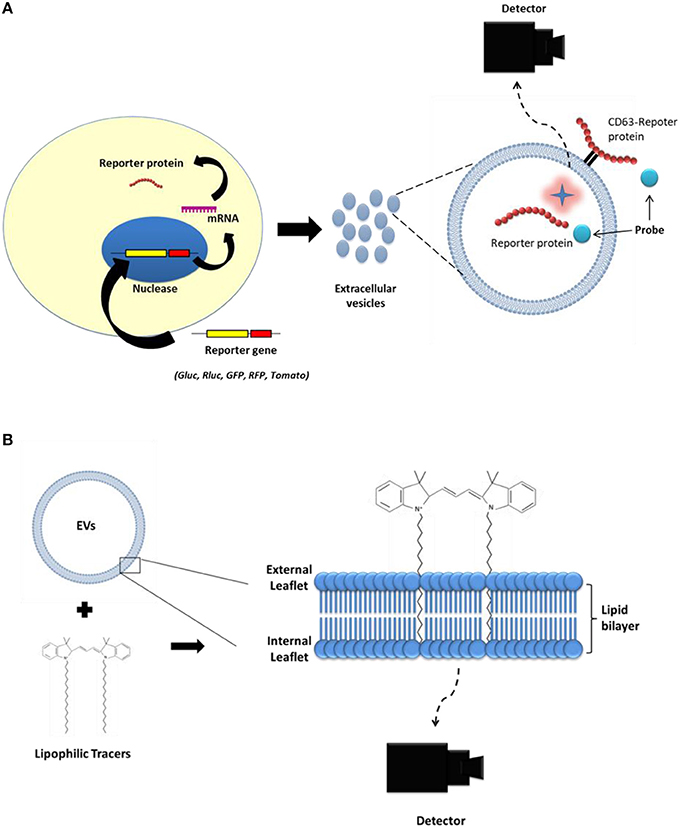
Figure 3. Strategies to label extracellular vesicles (EVs) for optical imaging. (A) First, reporter genes (Gluc, Rluc, GFP, RFP, tdTomato) are transduced into the parent cell line. Then, EVs produced from the parent cells expressing the reporter protein carry the reporter protein inside their lumen or on their membrane. (B) Lipophilic imaging agents (such as, DiD and DiR) could bind to the membrane of the EVs.
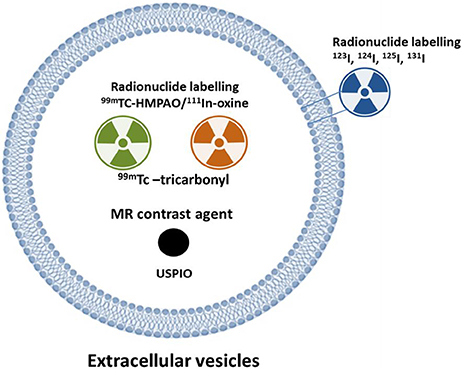
Figure 4. Strategy to label extracellular vesicles (EVs) for nuclear or MR imaging. Complex of 111In-oxine and 99mTc-HMPAO are lipophilic and penetrate the membrane of cells or extracellular vesicles (EVs). Inside the EVs, 111In-oxine attaches to cytoplasmic components (such as lactoferrin). 99mTc-HMPAO reacts with glutathione inside the EVs and it is converted to hydrophilic form. 99mTc-tricarbonyl binds to amino acids (such as histidine, methionine, and cysteine) of the EV membrane. EVs expressing lactadherin-streptavidin fusion protein on the membrane bind with the radioiodine-labeled biotin. USPIO was loaded to EVs using electroporation or incubation. HMPAO: hexamethylpropyleneamineoxime; USPIO: ultra-small super paramagnetic iron oxide.
Numerous preclinical studies involving molecular imaging modalities, either alone or multiple modalities, are needed to improve our knowledge of EV distribution for better clinical translation of the EV-based therapeutic approaches. Here, we discuss the highlights of using molecular imaging approaches for monitoring EVs in animal models. Furthermore, factors that affect detection sensitivity, such as methods of EV labeling, efficiency of EV labeling, toxicity, and limits of detection of imaging modalities as well as the future prospects of the use of EVs have been discussed.
Molecular Imaging Techniques for in Vivo Monitoring of EVs
Optical imaging is a powerful tool for cell tracking in small animals over the desired time periods without sacrificing the subjects (Kim et al., 2015; Li et al., 2016). Optical imaging consists of two main types: (i) Fluorescence protein imaging that involves the use of endogenous or exogenous molecules or materials that emit light when activated by an external light source such as a laser (Figures 3A,B); and (ii) Bioluminescent imaging that involves the use of a natural light-emitting protein that is activated by a chemical reaction, such as luciferase to trace the movement of certain cells or to identify the location of specific chemical reactions within the body (Figures 3A,B).
Nuclear imaging using radionuclides has been widely used for imaging human and mouse models. Radionuclides emit radiation that can be detected in living animals using a specific camera. Nuclear imaging provides excellent sensitivity and good tissue penetration (Ahn, 2014) such that it enables the visualization of deep organs such as the liver and spleen (Gangadaran et al., 2017a). MRI was conventionally used for anatomical imaging with high resolution and good tissue contrasts. The recent advancements in development of MR agents make it applicable to visualize the in vivo localization of EVs.
Monitoring in Vivo Biodistribution of EVs by Fluorescence Imaging
Fluorescence imaging involves the use of an external light source and a low-light camera with appropriate filters to collect fluorescence emission lights from samples (Kim et al., 2015). Recent advances in this system allow real-time visualization of EVs in an animal model (Grange et al., 2014; Lai et al., 2015; Smyth et al., 2015). Near-infrared (NIR) dyes are suitable for non-invasive in vivo applications because of their high signal-to-noise ratio, low auto fluorescence of biological tissue in the 700–900 nm spectral range, and deep tissue penetration of the NIR light. In particular, DiR (1,1′-Dioctadecyl-3,3,3′,3′-Tetramethylindotricarbocyanine Iodide) is a lipophilic dye, weakly fluorescent in H2O but fluorescent and photo-stable when incorporated into lipid membranes (of cells or EVs).
Fluorescent proteins were applied to visualize endogenous and exogenous EVs and track cell-to-cell communication in animal models. However, these proteins show low tissue penetration and this technique does not allow non-invasive in vivo imaging because of low resolution and the imaging can be performed only after the animal is sacrificed or surgically exposed (Hoshino et al., 2015; Lai et al., 2015; Zomer et al., 2015). Although fluorescent proteins fused with membrane proteins are extensively expressed in EVs, only a small population of EVs express the fluorescent proteins and the signal intensity was variable, according to the amount of reporter protein expression (Choi and Lee, 2016). Compared to lipophilic dyes, fluorescent reporter protein imaging systems are more specific to EVs. However, these systems require genetically engineered cells, which may change the characteristics of the parent cells and these changes can occur in the EVs as well.
Labeling of EVs by direct fluorescence has been widely used to investigate non-invasive in vivo behavior of exogenous EVs in small animal models (Ohno et al., 2013; Grange et al., 2014; Smyth et al., 2015). The labeling process is simple and lipophilic dyes are suitable for real-time monitoring of EVs in their native environments by NIR fluorescence imaging. Lipophilic dyes, including PKH, DiI, DiD, cy7, and DiR, are commonly used and yield stable fluorescent signals in vitro as well as in vivo (Grange et al., 2014; Watson et al., 2016; Jung et al., 2017).
A recent study used direct labeling of EVs with a dye and reported the biodistribution of breast cancer-derived EVs in mice. EVs derived from different breast cancer cells showed different in vivo and ex vivo distribution in after 24 h of administration (Wen et al., 2016). Wiklander et al. studied the biodistribution of EVs in mice after a systemic delivery; EVs were isolated from three different murine cell sources, including DCs and labeled with a NIR lipophilic dye. The route-of-administration and the dose and cell source of EVs influenced the biodistribution pattern, as demonstrated by in vivo imaging of DiR-labeled EVs (Wiklander et al., 2015). Watson et al. showed the efficient production of engineered EVs from HEK293 cells and labeled them with DiR. Further, live animal imaging showed dramatically lesser liver uptake of EV and increased EVs in blood by pre-treatment with scavenger receptor-A blocker (Watson et al., 2016). Gangadaran et al. recently used an NIR dye (DiR) to study the biodistribution of thyroid cancer (CAL62)-derived EVs in mouse models. Intravenously injected EVs were predominantly distributed to the liver and the spleen followed by the lung and the kidney (Gangadaran et al., 2017b).
Dye-based optical imaging is limited to exogenous EVs and the labeled fluorescent dyes stays in tissues even after EVs are degraded or internalized by the cells. As lipid labeling is non-specific for intact EVs, fluorescence signals can be emitted by cells in which EVs were internalized or attached to cell surface (Wiklander et al., 2015; Choi and Lee, 2016). The major limitation, however, is that the labeling with lipophilic dyes promotes clumping of EVs and may give rise to artifacts, especially during in vivo imaging (Grange et al., 2014). Moreover, the repeated washing steps required to remove the free dye residues might end up in significant EV damage. The lipophilic dye remains in the animal tissues even after the clearance of labeled EVs from the system because of several days of estimated in vivo half-life of the dye (Lai et al., 2014). In addition, a recent study compared the dye-based direct labeling with the Renilla luciferase (Rluc)-based indirect labeling of EVs derived from thyroid cancer cells. This study revealed that labeling with the dye can affect the normal distribution of EVs in an animal model; EV organotropism, which occurs due to the integrins present in the EV membrane, could be influenced by the dye attached onto the EV surface membrane, thus leading to different in vivo distribution (Gangadaran et al., 2017b).
Monitoring in Vivo Biodistribution of EVs by Bioluminescence Imaging
Bioluminescence is produced by a chemical reaction between bioluminescent proteins and their appropriate substrates (firefly luciferase and D-luciferin, Renilla or Gaussia luciferases-coelenterazine) (Wilson and Hastings, 1998; Li et al., 2016; Kalimuthu et al., 2017). Bioluminescence imaging has enabled the real-time visualization of EVs in an in vivo animal model and helped study the biodistribution of EVs. Furthermore, bioluminescence imaging offers sensitivity as well as a broad dynamic range for in vivo quantification (Takahashi et al., 2013; Imai et al., 2015). Compared to fluorescent-based imaging, bioluminescent imaging has an extremely high signal-to-noise ratio, because the auto-luminescence in mammalian tissue is negligible. Bioluminescence imaging has very low background emission and is independent from an excitation source to emit light. Therefore, bioluminescence imaging has been extensively used to determine cellular distribution, survival, proliferation, and differentiation after transplantation in the development of cell-based therapies (Kim et al., 2015). Takahashi et al. generated a fusion protein be made up of Gaussia luciferase (Gluc) and a lactadherin. This genetically engineered EV revealed the spatio-temporal distribution of EVs in a quantifying manner (Takahashi et al., 2013).
Lai et al. combined Gluc and biotinylation to create a EV reporter for multi-modal in vivo imaging. They monitored EVs in various major organs and body fluids (blood and urine) after administration of the bioluminescent EVs. Furthermore, they revealed that the EVs first undergo a fast distribution followed by an extended elimination via hepatic and renal routes within 6 h (Lai et al., 2014). Imai et al. used Gluc-lactadherin EVs and revealed that macrophages play significant roles in the clearance of intravenously injected EVs from the circulation (Imai et al., 2015). In another study, the fusion protein of Gluc-lactadherin was used to evaluate the pharmacokinetics of EVs from five different cells (Charoenviriyakul et al., 2017). Recently, Gangadaran et al. generated cancer cells expressing the bioluminescent reporter gene Rluc to study the biodistribution in nude mice. EVs were isolated from cancer cells (thyroid cancer: CAL62 and breast cancer: MDA-MB-231) expressing RLUC, and injected intravenously and imaged. CAL62-derived EVs mostly distributed to the lung, followed by the liver, spleen, and kidney, whereas MDA-MB-231-derived EVs distributed to the liver, followed by the lung, kidney, and spleen (Gangadaran et al., 2017b).
However, there are some limitations in using the bioluminescent system. Bioluminescent signal can be reduced when the EVs are located in deep internal organs (Ahn, 2014). Injection of substrates is required to generate optical signals and these substrates might be toxic to the animal and long-term sequential imaging might be technically limited by the multiple injection procedure involved (Gangadaran and Ahn, 2017). Procedures of bioluminescent labeling are complicated, compared with those of fluorescence dyes, because cells undergo a bioluminescent gene transduction, which might modify the natural behavior of the transduced cells (Gangadaran et al., 2017a). In addition, the transduction process is a time consuming.
Monitoring in Vivo Biodistribution of EVs by Nuclear Imaging
Nuclear imaging could be a good option for tracking EVs and evaluating their biodistribution. In this method, three-dimensional images are obtained using single-photon emission computed tomography (SPECT) or positron emission tomography (PET). Furthermore, nuclear imaging combined with anatomical imaging, such as computed tomography (CT) or MRI is also available, and this combined imaging technology provides a better understanding of the localization of the EVs (Figure 4).
Recently, 111In-oxine and 99mTc-hexamethylpropyleneamineoxime (HMPAO) were used for tracking EVs and EVMs (Hwang et al., 2015; Smyth et al., 2015). These materials were widely used for direct radionuclide labeling of leukocytes. Indium arranges an uncharged pseudo-octahedral complex with three molecules of 8-hydroxyquinoline (oxine). As this complex is neutral and lipid-soluble, it can penetrate the lipid bilayer of EVs easily and indium becomes decisively attached to cytoplasmic components such as lactoferrin (Roca et al., 2010). Smyth et al. labeled exosomes with 111In-oxine and then analyzed their biodistribution (Smyth et al., 2015).
Hwang et al. labeled EVMs with 99mTc-HMPAO and successfully imaged the EVMs using SPECT/CT (Hwang et al., 2015). As 99mTc-HMPAO is a lipophilic agent, it can penetrate the lipid bilayer of EV/EVMs. 99mTc-HMPAO reacts with reducing agents such as glutathione inside the EV/EVMs and is then converted to its hydrophilic form. Therefore, 99mTc could remain inside the EV/EVMs (de Vries et al., 2010). Compared to 111In, 99mTc is much cheaper and provides better image quality on gamma camera imaging; however, 111In is preferable for delayed imaging owing to its long half-life (2.8 days). 99mTc-HMPAO-labeled EV/EVMs are well-visualized using a gamma camera or SPECT (Hwang et al., 2015; Gangadaran et al., 2017a). As glutathione plays a major role in 99mTc-HMPAO labeling, glutathione concentration of the EVs might be important. Although most cells present glutathione, its concentration varies with the cell type (Gamcsik et al., 2012). The efficiency of labeling EVs with 99mTc-HMPAO might vary with the parent cells of EVs.
Varga et al. used 99mTc-tricarbonyl for labeling EVs. 99mTc-tricarbonyl binds to numerous amino acids, such as histidine, cysteine, and methionine, that might be bound to the surface of EVs (Varga et al., 2016). 99mTc-tricarbonyl showed relatively higher labeling efficiency in RBC-derived EVs (38.8%) with 98% radiochemical purity. However, they only performed 1-h imaging and image-based analysis. Therefore, further studies are needed to validate the efficiency of this labeling method.
Radioiodine (123I, 124I, 125I, and 131I) could be an option for labeling EVs (Gangadaran et al., 2017a). 123I and 131I can be used for gamma camera or SPECT imaging, and 124I can be used for PET imaging. Although each radioiodine isotope has different physical properties (such as, half-life, and emitting energy), these isotopes have identical chemical properties; therefore, same labeling methods can be used for these isotopes. Morishita et al. generated exosomes expressing streptavidin-lactadherin fusion protein and then labeled these exosomes with 125I-biotin using the streptavidin-biotin system (Morishita et al., 2015). They showed good stability of labeled radioiodine and well-demonstrated the biodistribution of these labeled exosomes. However, image acquisition was not possible because they used 125I. In their previous study, they synthesized 123I-biotin and performed gamma camera imaging, but the radiochemical yield of 123I-biotin is lower than that of 125I-biotin (29% vs. 65%) (Kudo et al., 2009). To apply this labeling method to other EVs, the desired gene must be delivered into the target cells and the expression of the protein in EVs must be determined. One big limitation of this transduction technology is the possibility of altering EV characteristics by the transduction procedure. Furthermore, for nuclear imaging, well-trained personnel are needed for the safe handling of the radionuclide. High cost and regulatory policies for the use of radioactive molecules are the other hurdles for using nuclear imaging.
Monitoring in Vivo Biodistribution of EVs by MRI
In recent studies, EVs were loaded with MRI contrasts, and the location of these EVs was visualized using MRI. Hu et al. loaded melanoma exosomes with ultra-small super paramagnetic iron oxide (USPIO) (Hu et al., 2014). They loaded EVs with 5 nm-sized nanoparticles, which show low signal intensity in T2-weighted images, by electroporation (54.9 μg iron per 100 μg EV protein). After injection of these EVs into the feet of the mice, MRI successfully revealed the migration of these EVs to the draining lymph nodes. As exosome aggregation or fusion could occur during electroporation, recent studies used a biocompatible trehalose-based electroporation pulse media; using this unique pulse media, melanoma exosomes were loaded with 5 nm USPIO while minimizing the electroporation-induced aggregation effect (Hu et al., 2014). Busato et al. loaded parent cells (adipose stem cells) with USPIO and collected the EVs from these cells (Busato et al., 2016). As they did not perform electroporation or the other manipulation in the EVs, the integrity of the EV membranes could be preserved. However, the amount of USPIO in EVs was much lower than that reported in previous studies (Hood et al., 2014; Busato et al., 2016). For tracking EVs via MRI, a large amount of USPIO-loaded EVs are needed because of the inherent low sensitivity of MRI technology. Only the EVs remaining at the injection sites after intramuscular injection were observed in this study (Busato et al., 2016).
Tracking EVs and Monitoring Tumor and Metastatic Behaviors Using Molecular Imaging
In the last decade, major developments have been made in characterizing the cellular source and role of EVs. The finding that they are natural carriers of miRNA, mRNA, and proteins led to the hypothesis that they can be used as therapeutic agents and vehicles for the delivery of therapeutic cargoes (exogenous) to tumors. Furthermore, the role of EVs in tumor metastasis would help researchers to understand how a tumor prepares the metastatic site at distal organs. Ohno et al. successfully showed that injected exosomes by intravenously delivered let-7a miRNA to an EGFR-expressing tumor by DiR labeling of the EVs in mice and inhibited tumor growth (Ohno et al., 2013). Smyth et al. observed a comparable fast clearance and limited tumor accumulation of injected EVs labeled with DiR and 111In-oxine by intravenous route (Smyth et al., 2015). Bellavia et al. engineered HEK293T to express the EV protein Lamp2b, fused to a portion of interleukin 3 (IL3) to target specific cancer cells (chronic myelogenous leukemia); DiR was used to label the HEK293T-derived EVs (Bellavia et al., 2017). Watson et al. showed efficient enhanced tumor delivery of engineered EVs by using DiR-labeled EVs, and living animal imaging showed dramatically reduced liver uptake of EVs and increased EVs circulating in blood; the EVs were eventually targeted to the tumor by pre-treatment with scavenger receptor-A blocker (Watson et al., 2016). Lai et al. combined Gluc and biotinylation to create an EV reporter for multi-modal in vivo imaging. They administered bioluminescent EVs and monitored their targeting to the tumor. Furthermore, they revealed that the EVs first undergo a quick distribution followed by targeting to tumor within 60 min (Lai et al., 2014).
Hoshino et al. directly labeled the EVs with PKH67 (green) or PKH26 (red) membrane dye, and the fluorescently labeled EVs were systemically injected (the tail vein, retro-orbital venous sinus, or intracardially) into nude mice. They quantified EV distribution and uptake in organs by NIR and confocal microscopic imaging, and demonstrated that integrins of EVs could be exploited to predict the organ-specific metastasis of tumors (Hoshino et al., 2015). Zomer et al. demonstrated that in vivo imaging of EVs revealed metastatic behavior similar to that observed using fluorescent protein with high-resolution intravital imaging on surgically exposed imaging site in a mouse model (Zomer et al., 2015). Hu et al. showed sentinel lymph node using MRI after injection of EVs loaded with USPIO into the foot pad (Hu et al., 2014; Table 3).
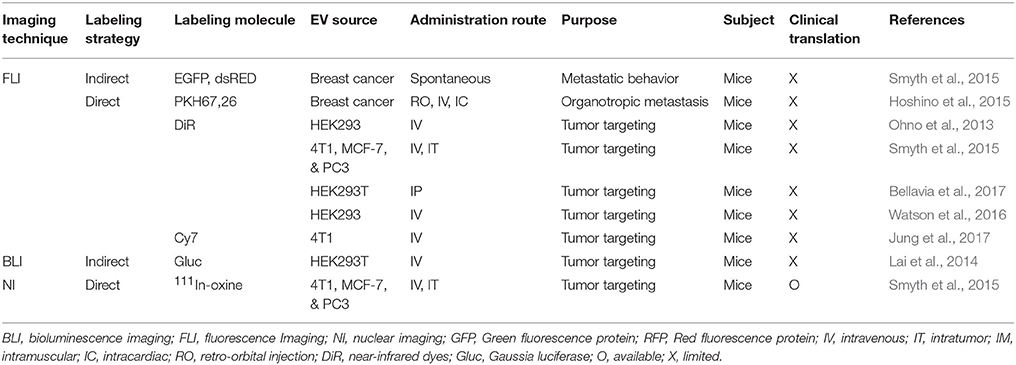
Table 3. Tumor targeting and tumor metastatic behavior of extracellular vesicles, as assessed by molecular imaging strategies.
Molecular Imaging for Monitoring Targeted EVs in Non-Cancerous Diseases
EVs are not only used in targeting tumors (Lai et al., 2014; Watson et al., 2016) but also in other diseases that can be monitored by molecular imaging (Table 4; Grange et al., 2014; Gangadaran et al., 2017c). DiD-labeled EVs were derived from MSCs pre-incubated with the DiD dye, and the isolated EVs were directly labeled with the DiD dye. Further, the labeled EVs were found to be accumulated predominately in the mice kidneys with acute kidney injury, and directly labeled EVs showed a prolonged signal in the animal model (Grange et al., 2014). Recently, Gangadaran et al. used fluoresce imaging to monitor EVs derived from MSCs and demonstrated prolonged in vivo retention of the EVs by mixing with scaffold in an ischemic hindlimb mouse model (Gangadaran et al., 2017c). Rajendran et al. used a NIR dye (DiR) to determine the treatment interval duration for EVs derived from MSCs at an intradermal site. EVs remained at the injection site (intradermal) for 48 h and were then distributed to the internal organs (lungs, liver, and kidneys) by 72 h; no signals were then observed at the injection site (Rajendran et al., 2017). Lai et al. reported successful visualization of EV-mediated communication between cells by imaging using GFP and tdTomato. Here, they transduced the cell with the fluorescent proteins and then isolated the EVs from the cells; they used multiphoton intravital microscopy to analyze the EV-RNA cargo delivery (Lai et al., 2015).
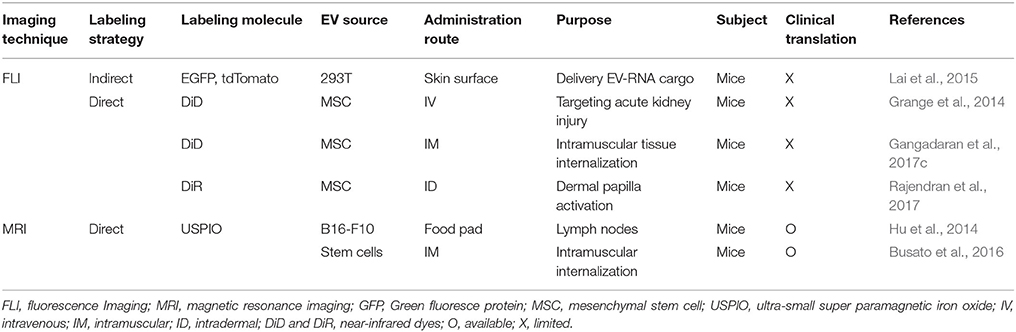
Table 4. Tracking extracellular vesicles for target/delivering to non-cancerous diseases using molecular imaging strategies.
Future Directions
Development of safe and effective drug delivery systems to desired target sites is receiving increasing attention recently. Number of studies are now shifting their focus from synthetic carriers to biological carriers that can achieve better efficacy and safety (Kim et al., 2017b). EVs have many pathophysiological functions, which might be helpful for drug delivery to and treatment of target lesions. In this aspect, EVs might have superiority over other synthetic carriers.
Bioengineering of parent cells might be helpful for enhancing the favorable characteristics of EVs as drug delivery vehicles. It can increase the targetability of EVs to lesions or enhance their therapeutic effect by loading useful biomaterials (Gujrati et al., 2014; Kim et al., 2017a). However, these bioengineering methods could be another hurdle for clinical translation.
EVs can be labeled or loaded with therapeutic radionuclides such as alpha-emitting (e.g., 211At, 213Bi) or beta-emitting (e.g., 131I, 90Y, 177Lu) radionuclides, which are widely used for radionuclide therapy (Ahn, 2014). As USPIOs can mediate magnetic hyperthermia, EVs may be used as theranostic nanocarriers to simultaneously detect and treat tumor microenvironments (Hood, 2016).
As drugs loaded into EVs show different pharmacokinetics, we can find new application of withdrawn drugs, which have good therapeutic effects but fail to reach the target tissue or have adverse effects in non-target tissues. Furthermore, insoluble drug candidates could be loaded into the EVs and effectively delivered to target tissues.
However, various components and low production yield of EVs are obstacles for clinical translation. EVMs derived from cells could be a good solution for the low production yield of EVs (Jang et al., 2013). EVMs could be a good substitute for EVs as drug delivery vehicles. Both EVs and EVMs have complex components, which need to be assessed for toxicity. To overcome this issue, self-derived EVs and EVMs from patients can be used (Escudier et al., 2005; Morse et al., 2005; Besse et al., 2016).
Several different labels and imaging techniques have been explored for labeling and tracking EVs in animal models to understand cancer biology and to develop EV-based targeted therapies. Direct labeling of EVs with lipophilic dye is easier than the indirect labeling methods and the safety profiles of direct labeling with dyes are relatively better than those of indirect labeling. However, the disadvantage of direct cell labeling is that the dye itself is detected rather than the EVs of interest (Choi and Lee, 2016). Although there are many reporter luciferase gene-based cell labeling studies available (Kim et al., 2015), unfortunately the luciferase reporter gene-based EV labeling studies are very few (Gangadaran et al., 2017a). Although luciferases can enable in vivo long-term monitoring of the cells in a quantitative manner in small animal models (Li et al., 2016), such long-term monitoring cannot be performed for EVs (Lai et al., 2014). Therefore, care must be taken when interpreting the experimental results, and rigorous validation is certainly needed to obtain more robust and reliable data.
Although there are some obstacles to use radionuclides in experiments, nuclear imaging has major advantage of no depth limitation compared to the optical imaging. After injection, most of the EVs are visualized in the liver and spleen; therefore, optical imaging is not sufficient for tracking EVs. Nuclear imaging can provide three-dimensional images by SPECT or PET, and the combined use of nuclear imaging and CT or MRI enhances the anatomical localization of EVs. Nuclear imaging can also provide semi-quantitative parameters. To the best of our knowledge, no literature data are available for the in vivo PET imaging of EVs; however, indirect labeling of EVs with positron-emitting radionuclides will be developed for the best nuclear in vivo imaging of EVs (Ahn, 2014).
Although MRI contrast agents have shown relatively lower sensitivity than that of optical imaging or nuclear imaging (Ahn, 2014), technological advances might provide better imaging contrast agents and machines. Nowadays, promising novel imaging sequences, such as SWIFT and UTE, provide positive contrast for SPIO, and it could be possible to characterize the distribution of exosomes using MRI (Hu et al., 2014).
Safety of labeled EVs is always an immense concern in potential clinical studies, as the introduction of foreign materials such as dye, radionucleotides, magnetic parties, and reporter proteins may cause unpredictable results in patients. As radionuclides are already used in clinics, there is less ethical and legal obstacles in clinical translation compared to those observed when using the other new probes. Recently, some clinical trials were performed to treat advanced melanoma and lung cancer by EVs (Escudier et al., 2005; Morse et al., 2005; Besse et al., 2016); however, there is no report on the kinetics of EVs in the human body. Molecular imaging techniques should be helpful to elucidate the in vivo kinetics of EVs used as drug delivery vehicles in humans.
Conclusion
In recent years, EVs have become an enthusiastic subject as drug delivery vehicles. Nonetheless, action mechanism, biodistribution, and pharmacokinetics of exogenously administered EVs are not well-studied, and the possibility of targeted delivery of drugs using EVs has not been fully assessed. In vivo molecular imaging of EVs would contribute to the understanding of the pathophysiological influence of EVs and accelerate the development of EV therapeutics in clinical fields. Application of optimal molecular imaging technology is needed to ensure the efficient use of EVs in various specific study situations.
Author Contributions
PG, CH, and B-CA: Contributed to the conception, writing, and discussion of this review; PG and CH: Wrote the initial draft of the manuscript. The final version was approved by all the authors.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
The reviewer DL and handling Editor declared their shared affiliation.
Acknowledgments
This work was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2016R1D1A1A02936968). PG would like to acknowledge Kyungpook National University for International Graduate Scholarship.
Abbreviations
CT, computed tomography; DCs, dendritic cells; ESCRT, endosomal sorting complex required for transport; EVM, extracellular vesicle mimetics; EVs, extracellular vesicles; HMPAO, 99mTc-hexamethylpropyleneamineoxime; MRI, magnetic resonance imaging; MSC, mesenchymal stem cell; NIR, Near-infrared; PDCD6IP, programmed cell death 6 interacting protein; PET, positron emission tomography; SPECT, single-photon emission computed tomography; TSG101, tumor susceptibility gene 101; USPIO, ultra-small super paramagnetic iron oxide.
References
Agrawal, A. K., Aqil, F., Jeyabalan, J., Spencer, W. A., Beck, J., Gachuki, B. W., et al. (2017). Milk-derived exosomes for oral delivery of paclitaxel. Nanomed. Nanotechnol. Biol. Med. 13, 1627–1636. doi: 10.1016/j.nano.2017.03.001
Aharon, A., Sabbah, A., Ben-Shaul, S., Berkovich, H., Loven, D., Brenner, B., et al. (2017). Chemotherapy administration to breast cancer patients affects extracellular vesicles thrombogenicity and function. Oncotarget 8, 63265–63280. doi: 10.18632/oncotarget.18792
Ahn, B.-C. (2014). Requisites for successful theranostics with radionuclide-based reporter gene imaging. J. Drug Target 22, 295–303. doi: 10.3109/1061186X.2013.878940
Alvarez-Erviti, L., Seow, Y., Yin, H., Betts, C., Lakhal, S., and Wood, M. J. (2011). Delivery of siRNA to the mouse brain by systemic injection of targeted exosomes. Nat. Biotechnol. 29:341. doi: 10.1038/nbt.1807
Anderson, M. R., Kashanchi, F., and Jacobson, S. (2016). Exosomes in viral disease. Neurotherapeutics 13, 535–546. doi: 10.1007/s13311-016-0450-6
Banizs, A. B., Huang, T., Dryden, K., Berr, S. S., Stone, J. R., Nakamoto, R. K., et al. (2014). In vitro evaluation of endothelial exosomes as carriers for small interfering ribonucleic acid delivery. Int. J. Nanomedicine 9, 4223–4230. doi: 10.2147/IJN.S64267
Baumgart, S., Hölters, S., Ohlmann, C.-H., Bohle, R., Stöckle, M., Ostenfeld, M. S., et al. (2017). Exosomes of invasive urothelial carcinoma cells are characterized by a specific miRNA expression signature. Oncotarget 8, 58278–58291. doi: 10.18632/oncotarget.17619
Bellavia, D., Raimondo, S., Calabrese, G., Forte, S., Cristaldi, M., Patinella, A., et al. (2017). Interleukin 3- receptor targeted exosomes inhibit in vitro and in vivo Chronic Myelogenous Leukemia cell growth. Theranostics 7, 1333–1345. doi: 10.7150/thno.17092
Besse, B., Charrier, M., Lapierre, V., Dansin, E., Lantz, O., Planchard, D., et al. (2016). Dendritic cell-derived exosomes as maintenance immunotherapy after first line chemotherapy in NSCLC. Oncoimmunology 5:e1071008. doi: 10.1080/2162402X.2015.1071008
Bobrie, A., Colombo, M., Raposo, G., and Théry, C. (2011). Exosome secretion: molecular mechanisms and roles in immune responses. Traffic Cph. Den. 12, 1659–1668. doi: 10.1111/j.1600-0854.2011.01225.x
Brisson, A. R., Tan, S., Linares, R., Gounou, C., and Arraud, N. (2017). Extracellular vesicles from activated platelets: a semiquantitative cryo-electron microscopy and immuno-gold labeling study. Platelets 28, 263–271. doi: 10.1080/09537104.2016.1268255
Busato, A., Bonafede, R., Bontempi, P., Scambi, I., Schiaffino, L., Benati, D., et al. (2016). Magnetic resonance imaging of ultrasmall superparamagnetic iron oxide-labeled exosomes from stem cells: a new method to obtain labeled exosomes. Int. J. Nanomedicine 11, 2481–2490. doi: 10.2147/IJN.S104152
Cappello, F., Logozzi, M., Campanella, C., Bavisotto, C. C., Marcilla, A., Properzi, F., et al. (2017). Exosome levels in human body fluids: a tumor marker by themselves? Eur. J. Pharm. Sci. 96, 93–98. doi: 10.1016/j.ejps.2016.09.010
Cavallari, C., Ranghino, A., Tapparo, M., Cedrino, M., Figliolini, F., Grange, C., et al. (2017). Serum-derived extracellular vesicles (EVs) impact on vascular remodeling and prevent muscle damage in acute hind limb ischemia. Sci. Rep. 7:8180. doi: 10.1038/s41598-017-08250-0
Charoenviriyakul, C., Takahashi, Y., Morishita, M., Matsumoto, A., Nishikawa, M., and Takakura, Y. (2017). Cell type-specific and common characteristics of exosomes derived from mouse cell lines: yield, physicochemical properties, and pharmacokinetics. Eur. J. Pharm. Sci. 96, 316–322. doi: 10.1016/j.ejps.2016.10.009
Choi, H., and Lee, D. S. (2016). Illuminating the physiology of extracellular vesicles. Stem Cell Res. Ther. 7:55. doi: 10.1186/s13287-016-0316-1
Clayton, A., Harris, C. L., Court, J., Mason, M. D., and Morgan, B. P. (2003). Antigen-presenting cell exosomes are protected from complement-mediated lysis by expression of CD55 and CD59. Eur. J. Immunol. 33, 522–531. doi: 10.1002/immu.200310028
de Vries, E. F. J., Roca, M., Jamar, F., Israel, O., and Signore, A. (2010). Guidelines for the labelling of leucocytes with (99m)Tc-HMPAO. Inflammation/infection taskgroup of the european association of nuclear medicine. Eur. J. Nucl. Med. Mol. Imaging 37, 842–848. doi: 10.1007/s00259-010-1394-4
Di Rocco, G., Baldari, S., and Toietta, G. (2016). Towards therapeutic delivery of extracellular vesicles: strategies for in vivo tracking and biodistribution analysis. Stem Cells Int. 2016:5029619. doi: 10.1155/2016/5029619
EL Andaloussi, S., Mäger, I., Breakefield, X. O., and Wood, M. J. (2013). Extracellular vesicles: biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 12, 347–357. doi: 10.1038/nrd3978
Escrevente, C., Keller, S., Altevogt, P., and Costa, J. (2011). Interaction and uptake of exosomes by ovarian cancer cells. BMC Cancer 11:108. doi: 10.1186/1471-2407-11-108
Escudier, B., Dorval, T., Chaput, N., André, F., Caby, M.-P., Novault, S., et al. (2005). Vaccination of metastatic melanoma patients with autologous dendritic cell (DC) derived-exosomes: results of thefirst phase I clinical trial. J. Transl. Med. 3:10. doi: 10.1186/1479-5876-3-10
Filipazzi, P., Bürdek, M., Villa, A., Rivoltini, L., and Huber, V. (2012). Recent advances on the role of tumor exosomes in immunosuppression and disease progression. Semin. Cancer Biol. 22, 342–349. doi: 10.1016/j.semcancer.2012.02.005
Fitzner, D., Schnaars, M., van Rossum, D., Krishnamoorthy, G., Dibaj, P., Bakhti, M., et al. (2011). Selective transfer of exosomes from oligodendrocytes to microglia by macropinocytosis. J. Cell Sci. 124, 447–458. doi: 10.1242/jcs.074088
Gamcsik, M. P., Kasibhatla, M. S., Teeter, S. D., and Colvin, O. M. (2012). Glutathione levels in human tumors. Biomarkers 17, 671–691. doi: 10.3109/1354750X.2012.715672
Gangadaran, P., and Ahn, B.-C. (2017). Molecular imaging: a useful tool for the development of natural killer cell-based immunotherapies. Front. Immunol. 8:1090. doi: 10.3389/fimmu.2017.01090
Gangadaran, P., Hong, C. M., and Ahn, B.-C. (2017a). Current perspectives on in vivo noninvasive tracking of extracellular vesicles with molecular imaging. Biomed Res. Int. 2017:9158319. doi: 10.1155/2017/9158319
Gangadaran, P., Li, X. J., Lee, H. W., Oh, J. M., Kalimuthu, S., Rajendran, R. L., et al. (2017b). A new bioluminescent reporter system to study the biodistribution of systematically injected tumor-derived bioluminescent extracellular vesicles in mice. Oncotarget 8, 109894–109914. doi: 10.18632/oncotarget.22493
Gangadaran, P., Rajendran, R. L., Lee, H. W., Kalimuthu, S., Hong, C. M., Jeong, S. Y., et al. (2017c). Extracellular vesicles from mesenchymal stem cells activates VEGF receptors and accelerates recovery of hindlimb ischemia. J. Control. Release 264, 112–126. doi: 10.1016/j.jconrel.2017.08.022
Goh, W. J., Zou, S., Ong, W. Y., Torta, F., Alexandra, A. F., Schiffelers, R. M., et al. (2017). Bioinspired cell-derived nanovesicles versus exosomes as drug delivery systems: a cost-effective alternative. Sci. Rep. 7:14322. doi: 10.1038/s41598-017-14725-x
Grange, C., Tapparo, M., Bruno, S., Chatterjee, D., Quesenberry, P. J., Tetta, C., et al. (2014). Biodistribution of mesenchymal stem cell-derived extracellular vesicles in a model of acute kidney injury monitored by optical imaging. Int. J. Mol. Med. 33, 1055–1063. doi: 10.3892/ijmm.2014.1663
Gujrati, V., Kim, S., Kim, S.-H., Min, J. J., Choy, H. E., Kim, S. C., et al. (2014). Bioengineered bacterial outer membrane vesicles as cell-specific drug-delivery vehicles for cancer therapy. ACS Nano 8, 1525–1537. doi: 10.1021/nn405724x
Haney, M. J., Klyachko, N. L., Zhao, Y., Gupta, R., Plotnikova, E. G., He, Z., et al. (2015). Exosomes as drug delivery vehicles for Parkinson's disease therapy. J. Control. Release 207, 18–30. doi: 10.1016/j.jconrel.2015.03.033
Hemler, M. E. (2003). Tetraspanin proteins mediate cellular penetration, invasion, and fusion events and define a novel type of membrane microdomain. Annu. Rev. Cell Dev. Biol. 19, 397–422. doi: 10.1146/annurev.cellbio.19.111301.153609
Hofheinz, R. D., Gnad-Vogt, S. U., Beyer, U., and Hochhaus, A. (2005). Liposomal encapsulated anti-cancer drugs. Anticancer. Drugs 16, 691–707. doi: 10.1097/01.cad.0000167902.53039.5a
Hood, J. L. (2016). Post isolation modification of exosomes for nanomedicine applications. Nanomedicine 11, 1745–1756. doi: 10.2217/nnm-2016-0102
Hood, J. L., and Wickline, S. A. (2012). A systematic approach to exosome-based translational nanomedicine. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 4, 458–467. doi: 10.1002/wnan.1174
Hood, J. L., Scott, M. J., and Wickline, S. A. (2014). Maximizing exosome colloidal stability following electroporation. Anal. Biochem. 448, 41–49. doi: 10.1016/j.ab.2013.12.001
Hoshino, A., Costa-Silva, B., Shen, T.-L., Rodrigues, G., Hashimoto, A., Tesic Mark, M., et al. (2015). Tumour exosome integrins determine organotropic metastasis. Nature 527, 329–335. doi: 10.1038/nature15756
Hu, L., Wickline, S. A., and Hood, J. L. (2014). Magnetic resonance imaging of melanoma exosomes in lymph nodes. Magn. Reson. Med. 74, 266–271. doi: 10.1002/mrm.25376
Hwang, D. W., Choi, H., Jang, S. C., Yoo, M. Y., Park, J. Y., Choi, N. E., et al. (2015). Noninvasive imaging of radiolabeled exosome-mimetic nanovesicle using (99m)Tc-HMPAO. Sci. Rep. 5:15636. doi: 10.1038/srep15636
Imai, T., Takahashi, Y., Nishikawa, M., Kato, K., Morishita, M., Yamashita, T., et al. (2015). Macrophage-dependent clearance of systemically administered B16BL6-derived exosomes from the blood circulation in mice. J. Extracell. Vesicles 4:26238. doi: 10.3402/jev.v4.26238
Jang, S. C., Kim, O. Y., Yoon, C. M., Choi, D.-S., Roh, T.-Y., Park, J., et al. (2013). Bioinspired exosome-mimetic nanovesicles for targeted delivery of chemotherapeutics to malignant tumors. ACS Nano 7, 7698–7710. doi: 10.1021/nn402232g
Jiang, X.-C., and Gao, J.-Q. (2017). Exosomes as novel bio-carriers for gene and drug delivery. Int. J. Pharm. 521, 167–175. doi: 10.1016/j.ijpharm.2017.02.038
Johnstone, R. M., Mathew, A., Mason, A. B., and Teng, K. (1991). Exosome formation during maturation of mammalian and avian reticulocytes: evidence that exosome release is a major route for externalization of obsolete membrane proteins. J. Cell. Physiol. 147, 27–36. doi: 10.1002/jcp.1041470105
Jung, K. O., Youn, H., Lee, C.-H., Kang, K. W., and Chung, J.-K. (2017). Visualization of exosome-mediated miR-210 transfer from hypoxic tumor cells. Oncotarget 8, 9899–9910. doi: 10.18632/oncotarget.14247
Kalimuthu, S., Gangadaran, P., Li, X. J., Oh, J. M., Lee, H. W., Jeong, S. Y., et al. (2016). In vivo therapeutic potential of mesenchymal stem cell-derived extracellular vesicles with optical imaging reporter in tumor mice model. Sci. Rep. 6:30418. doi: 10.1038/srep30418
Kalimuthu, S., Oh, J. M., Gangadaran, P., Zhu, L., Lee, H. W., Rajendran, R. L., et al. (2017). In vivo tracking of chemokine receptor CXCR4-engineered mesenchymal stem cell migration by optical molecular imaging. Stem Cells Int. 2017:8085637. doi: 10.1155/2017/8085637
Kamaly, N., Yameen, B., Wu, J., and Farokhzad, O. C. (2016). Degradable controlled-release polymers and polymeric nanoparticles: mechanisms of controlling drug release. Chem. Rev. 116, 2602–2663. doi: 10.1021/acs.chemrev.5b00346
Kamerkar, S., LeBleu, V. S., Sugimoto, H., Yang, S., Ruivo, C. F., Melo, S. A., et al. (2017). Exosomes facilitate therapeutic targeting of oncogenic KRAS in pancreatic cancer. Nature 546, 498–503. doi: 10.1038/nature22341
Karlsson, M., Lundin, S., Dahlgren, U., Kahu, H., Pettersson, I., and Telemo, E. (2001). “Tolerosomes” are produced by intestinal epithelial cells. Eur. J. Immunol. 31, 2892–2900. doi: 10.1002/1521-4141(2001010)31:10<2892::AID-IMMU2892>3.0.CO;2-I
Kim, J. E., Kalimuthu, S., and Ahn, B.-C. (2015). In vivo cell tracking with bioluminescence imaging. Nucl. Med. Mol. Imaging 49, 3–10. doi: 10.1007/s13139-014-0309-x
Kim, M. S., Haney, M. J., Zhao, Y., Mahajan, V., Deygen, I., Klyachko, N. L., et al. (2016). Development of exosome-encapsulated paclitaxel to overcome MDR in cancer cells. Nanomedicine 12, 655–664. doi: 10.1016/j.nano.2015.10.012
Kim, O. Y., Dinh, N. T. H., Park, H. T., Choi, S. J., Hong, K., and Gho, Y. S. (2017a). Bacterial protoplast-derived nanovesicles for tumor targeted delivery of chemotherapeutics. Biomaterials 113, 68–79. doi: 10.1016/j.biomaterials.2016.10.037
Kim, O. Y., Lee, J., and Gho, Y. S. (2017b). Extracellular vesicle mimetics: novel alternatives to extracellular vesicle-based theranostics, drug delivery, and vaccines. Semin. Cell Dev. Biol. 67, 74–82. doi: 10.1016/j.semcdb.2016.12.001
Koumangoye, R. B., Sakwe, A. M., Goodwin, J. S., Patel, T., and Ochieng, J. (2011). Detachment of breast tumor cells induces rapid secretion of exosomes which subsequently mediate cellular adhesion and spreading. PLoS ONE 6:e24234. doi: 10.1371/journal.pone.0024234
Kowal, J., Arras, G., Colombo, M., Jouve, M., Morath, J. P., Primdal-Bengtson, B., et al. (2016). Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proc. Natl. Acad. Sci. U.S.A. 113, E968–E977. doi: 10.1073/pnas.1521230113
Kudo, T., Ueda, M., Kuge, Y., Mukai, T., Tanaka, S., Masutani, M., et al. (2009). Imaging of HIF-1-active tumor hypoxia using a protein effectively delivered to and specifically stabilized in HIF-1-active tumor cells. J. Nucl. Med. 50, 942–949. doi: 10.2967/jnumed.108.061119
Lai, C. P., Kim, E. Y., Badr, C. E., Weissleder, R., Mempel, T. R., Tannous, B. A., et al. (2015). Visualization and tracking of tumour extracellular vesicle delivery and RNA translation using multiplexed reporters. Nat. Commun. 6:7029. doi: 10.1038/ncomms8029
Lai, C. P., Mardini, O., Ericsson, M., Prabhakar, S., Maguire, C. A., Chen, J. W., et al. (2014). Dynamic biodistribution of extracellular vesicles in vivo using a multimodal imaging reporter. ACS Nano 8, 483–494. doi: 10.1021/nn404945r
Lee, H. W., Gangadaran, P., Kalimuthu, S., and Ahn, B.-C. (2016). Advances in molecular imaging strategies for in vivo tracking of immune cells. Biomed Res. Int. 2016:1946585. doi: 10.1155/2016/1946585
Lener, T., Gimona, M., Aigner, L., Börger, V., Buzas, E., Camussi, G., et al. (2015). Applying extracellular vesicles based therapeutics in clinical trials - an ISEV position paper. J. Extracell. Vesicles 4:30087. doi: 10.3402/jev.v4.30087
Li, C., Zhang, J., Zu, Y.-J., Nie, S.-F., Cao, J., Wang, Q., et al. (2015). Biocompatible and biodegradable nanoparticles for enhancement of anti-cancer activities of phytochemicals. Chin. J. Nat. Med. 13, 641–652. doi: 10.1016/S1875-5364(15)30061-3
Li, X. J., Gangadaran, P., Kalimuthu, S., Oh, J. M., Zhu, L., Jeong, S. Y., et al. (2016). Role of pulmonary macrophages in initiation of lung metastasis in anaplastic thyroid cancer. Int. J. Cancer 139, 2583–2592. doi: 10.1002/ijc.30387
Liu, Y., and Cao, X. (2016). Organotropic metastasis: role of tumor exosomes. Cell Res. 26, 149–150. doi: 10.1038/cr.2015.153
Lu, Z., Zuo, B., Jing, R., Gao, X., Rao, Q., Liu, Z., et al. (2017). Dendritic cell-derived exosomes elicit tumor regression in autochthonous hepatocellular carcinoma mouse models. J. Hepatol. 67, 739–748. doi: 10.1016/j.jhep.2017.05.019
Luan, X., Sansanaphongpricha, K., Myers, I., Chen, H., Yuan, H., and Sun, D. (2017). Engineering exosomes as refined biological nanoplatforms for drug delivery. Acta Pharmacol. Sin. 38, 754–763. doi: 10.1038/aps.2017.12
Ludwig, S., Floros, T., Theodoraki, M.-N., Hong, C.-S., Jackson, E. K., Lang, S., et al. (2017). Suppression of lymphocyte functions by plasma exosomes correlates with disease activity in patients with head and neck cancer. Clin. Cancer Res. 23, 4843–4854. doi: 10.1158/1078-0432.CCR-16-2819
Malhotra, H., Sheokand, N., Kumar, S., Chauhan, A. S., Kumar, M., Jakhar, P., et al. (2016). Exosomes: tunable nano vehicles for macromolecular delivery of transferrin and lactoferrin to specific intracellular compartment. J. Biomed. Nanotechnol. 12, 1101–1114. doi: 10.1166/jbn.2016.2229
Masood, F. (2016). Polymeric nanoparticles for targeted drug delivery system for cancer therapy. Mater. Sci. Eng. C 60, 569–578. doi: 10.1016/j.msec.2015.11.067
Mathivanan, S., Fahner, C. J., Reid, G. E., and Simpson, R. J. (2012). ExoCarta 2012: database of exosomal proteins, RNA and lipids. Nucleic Acids Res. 40, D1241–D1244. doi: 10.1093/nar/gkr828
Momen-Heravi, F., Bala, S., Bukong, T., and Szabo, G. (2014). Exosome-mediated delivery of functionally active miRNA-155 inhibitor to macrophages. Nanomedicine Nanotechnol. Biol. Med. 10, 1517–1527. doi: 10.1016/j.nano.2014.03.014
Morishita, M., Takahashi, Y., Nishikawa, M., Sano, K., Kato, K., Yamashita, T., et al. (2015). Quantitative analysis of tissue distribution of the B16BL6-derived exosomes using a streptavidin-lactadherin fusion protein and iodine-125-labeled biotin derivative after intravenous injection in mice. J. Pharm. Sci. 104, 705–713. doi: 10.1002/jps.24251
Morse, M. A., Garst, J., Osada, T., Khan, S., Hobeika, A., Clay, T. M., et al. (2005). A phase I study of dexosome immunotherapy in patients with advanced non-small cell lung cancer. J. Transl. Med. 3:9. doi: 10.1186/1479-5876-3-9
Mulcahy, L. A., Pink, R. C., and Carter, D. R. (2014). Routes and mechanisms of extracellular vesicle uptake. J. Extracell. Vesicles 3:24641. doi: 10.3402/jev.v3.24641
Muralidharan-Chari, V., Clancy, J., Plou, C., Romao, M., Chavrier, P., Raposo, G., et al. (2009). ARF6-regulated shedding of tumor cell-derived plasma membrane microvesicles. Curr. Biol. 19, 1875–1885. doi: 10.1016/j.cub.2009.09.059
Nolte-‘t Hoen, E., Cremer, T., Gallo, R. C., and Margolis, L. B. (2016). Extracellular vesicles and viruses: are they close relatives? Proc. Natl. Acad. Sci. U.S.A. 113, 9155–9161. doi: 10.1073/pnas.1605146113
Ohno, S., Takanashi, M., Sudo, K., Ueda, S., Ishikawa, A., Matsuyama, N., et al. (2013). Systemically injected exosomes targeted to EGFR deliver antitumor MicroRNA to breast cancer cells. Mol. Ther. 21, 185–191. doi: 10.1038/mt.2012.180
Pascucci, L., Coccè, V., Bonomi, A., Ami, D., Ceccarelli, P., Ciusani, E., et al. (2014). Paclitaxel is incorporated by mesenchymal stromal cells and released in exosomes that inhibit in vitro tumor growth: a new approach for drug delivery. J. Control. Release 192, 262–270. doi: 10.1016/j.jconrel.2014.07.042
Piffoux, M., Gazeau, F., Wilhelm, C., and Silva, A. K. A. (2017). “Imaging and therapeutic potential of extracellular vesicles,” in Design and Applications of Nanoparticles in Biomedical Imaging, eds J. Bulte and M. Modo (Cham: Springer), 43–68.
Rajendran, R. L., Gangadaran, P., Bak, S. S., Oh, J. M., Kalimuthu, S., Lee, H. W., et al. (2017). Extracellular vesicles derived from MSCs activates dermal papilla cell in vitro and promotes hair follicle conversion from telogen to anagen in mice. Sci. Rep. 7:15560. doi: 10.1038/s41598-017-15505-3
Raposo, G., Nijman, H. W., Stoorvogel, W., Liejendekker, R., Harding, C. V., Melief, C. J., et al. (1996). B lymphocytes secrete antigen-presenting vesicles. J. Exp. Med. 183, 1161–1172. doi: 10.1084/jem.183.3.1161
Roca, M., de Vries, E. F., Jamar, F., Israel, O., and Signore, A. (2010). Guidelines for the labelling of leucocytes with (111)In-oxine. Inflammation/infection taskgroup of the european association of nuclear medicine. Eur. J. Nucl. Med. Mol. Imaging 37, 835–841. doi: 10.1007/s00259-010-1393-5
Saari, H., Lázaro-Ibáñez, E., Viitala, T., Vuorimaa-Laukkanen, E., Siljander, P., and Yliperttula, M. (2015). Microvesicle- and exosome-mediated drug delivery enhances the cytotoxicity of Paclitaxel in autologous prostate cancer cells. J. Control. Release 220, 727–737. doi: 10.1016/j.jconrel.2015.09.031
Sarko, D. K., and McKinney, C. E. (2017). Exosomes: origins and therapeutic potential for neurodegenerative disease. Front. Neurosci. 11:82. doi: 10.3389/fnins.2017.00082
Schillaci, O., Fontana, S., Monteleone, F., Taverna, S., Di Bella, M. A., Di Vizio, D., et al. (2017). Exosomes from metastatic cancer cells transfer amoeboid phenotype to non-metastatic cells and increase endothelial permeability: their emerging role in tumor heterogeneity. Sci. Rep. 7:4711. doi: 10.1038/s41598-017-05002-y
Sercombe, L., Veerati, T., Moheimani, F., Wu, S. Y., Sood, A. K., and Hua, S. (2015). Advances and challenges of liposome assisted drug delivery. Front. Pharmacol. 6:286. doi: 10.3389/fphar.2015.00286
Shoae-Hassani, A., Hamidieh, A. A., Behfar, M., Mohseni, R., Mortazavi-Tabatabaei, S. A., and Asgharzadeh, S. (2017). NK cell-derived exosomes from NK cells previously exposed to neuroblastoma cells augment the antitumor activity of cytokine-activated NK cells. J. Immunother. 40, 265–276. doi: 10.1097/CJI.0000000000000179
Shtam, T. A., Kovalev, R. A., Varfolomeeva, E. Y., Makarov, E. M., Kil, Y. V., and Filatov, M. V. (2013). Exosomes are natural carriers of exogenous siRNA to human cells in vitro. Cell Commun. Signal. 11:88. doi: 10.1186/1478-811X-11-88
Smyth, T., Kullberg, M., Malik, N., Smith-Jones, P., Graner, M. W., and Anchordoquy, T. J. (2015). Biodistribution and delivery efficiency of unmodified tumor-derived exosomes. J. Control. Release 199, 145–155. doi: 10.1016/j.jconrel.2014.12.013
Srivastava, A., Filant, J., Moxley, K. M., Sood, A., McMeekin, S., and Ramesh, R. (2015). Exosomes: a role for naturally occurring nanovesicles in cancer growth, diagnosis and treatment. Curr. Gene Ther. 15, 182–192. doi: 10.2174/1566523214666141224100612
Sun, D., Zhuang, X., Xiang, X., Liu, Y., Zhang, S., Liu, C., et al. (2010). A novel nanoparticle drug delivery system: the anti-inflammatory activity of curcumin is enhanced when encapsulated in exosomes. Mol. Ther. 18, 1606–1614. doi: 10.1038/mt.2010.105
Svensson, K. J., Christianson, H. C., Wittrup, A., Bourseau-Guilmain, E., Lindqvist, E., Svensson, L. M., et al. (2013). Exosome uptake depends on ERK1/2-heat shock protein 27 signaling and lipid Raft-mediated endocytosis negatively regulated by caveolin-1. J. Biol. Chem. 288, 17713–17724. doi: 10.1074/jbc.M112.445403
Takahashi, Y., Nishikawa, M., Shinotsuka, H., Matsui, Y., Ohara, S., Imai, T., et al. (2013). Visualization and in vivo tracking of the exosomes of murine melanoma B16-BL6 cells in mice after intravenous injection. J. Biotechnol. 165, 77–84. doi: 10.1016/j.jbiotec.2013.03.013
Théry, C., Ostrowski, M., and Segura, E. (2009). Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 9, 581–593. doi: 10.1038/nri2567
Tian, Y., Li, S., Song, J., Ji, T., Zhu, M., Anderson, G. J., et al. (2014). A doxorubicin delivery platform using engineered natural membrane vesicle exosomes for targeted tumor therapy. Biomaterials 35, 2383–2390. doi: 10.1016/j.biomaterials.2013.11.083
Trajkovic, K., Hsu, C., Chiantia, S., Rajendran, L., Wenzel, D., Wieland, F., et al. (2008). Ceramide triggers budding of exosome vesicles into multivesicular endosomes. Science 319, 1244–1247. doi: 10.1126/science.1153124
Vader, P., Mäger, I., Lee, Y., Nordin, J. Z., Andaloussi, S. E. L., and Wood, M. J. A. (2017). Preparation and isolation of siRNA-loaded extracellular vesicles. Methods Mol. Biol. 1545, 197–204. doi: 10.1007/978-1-4939-6728-5_14
Varga, Z., Gyurkó, I., Pálóczi, K., Buzás, E. I., Horváth, I., Hegedus, N., et al. (2016). Radiolabeling of extracellular vesicles with (99m)Tc for quantitative in vivo imaging studies. Cancer Biother. Radiopharm. 31, 168–173. doi: 10.1089/cbr.2016.2009
Wahlgren, J., De L Karlson, T., Brisslert, M., Vaziri Sani, F., Telemo, E., Sunnerhagen, P., et al. (2012). Plasma exosomes can deliver exogenous short interfering RNA to monocytes and lymphocytes. Nucleic Acids Res. 40:e130. doi: 10.1093/nar/gks463
Watson, D. C., Bayik, D., Srivatsan, A., Bergamaschi, C., Valentin, A., Niu, G., et al. (2016). Efficient production and enhanced tumor delivery of engineered extracellular vesicles. Biomaterials 105, 195–205. doi: 10.1016/j.biomaterials.2016.07.003
Wang, J., Wu, Y., Guo, T., Fei, Y., Yu, L., Ma, S., et al. (2017). Adipocyte-derived exosomes promote lung cancer metastasis by increasing MMP9 activity via transferring MMP3 to lung cancer cells. Oncotarget 8, 81880–81891. doi: 10.18632/oncotarget.18737
Wen, S. W., Sceneay, J., Lima, L. G., Wong, C. S., Becker, M., Krumeich, S., et al. (2016). the biodistribution and immune suppressive effects of breast cancer-derived exosomes. Cancer Res. 76, 6816–6827. doi: 10.1158/0008-5472.CAN-16-0868
Wiklander, O. P., Nordin, J. Z., O'Loughlin, A., Gustafsson, Y., Corso, G., Mäger, I., et al. (2015). Extracellular vesicle in vivo biodistribution is determined by cell source, route of administration and targeting. J. Extracell. Vesicles 4:26316. doi: 10.3402/jev.v4.26316
Wilson, T., and Hastings, J. W. (1998). Bioluminescence. Annu. Rev. Cell Dev. Biol. 14, 197–230. doi: 10.1146/annurev.cellbio.14.1.197
Yáñez-Mó, M., Siljander, P. R., Andreu, Z., Zavec, A. B., Borràs, F. E., Buzas, E. I., et al. (2015). Biological properties of extracellular vesicles and their physiological functions. J. Extracell. Vesicles 4:27066. doi: 10.3402/jev.v4.27066
Yang, J., Zhang, X., Chen, X., Wang, L., and Yang, G. (2017). Exosome mediated delivery of miR-124 promotes neurogenesis after ischemia. Mol. Ther. Nucleic Acids 7, 278–287. doi: 10.1016/j.omtn.2017.04.010
Yang, T., Martin, P., Fogarty, B., Brown, A., Schurman, K., Phipps, R., et al. (2015). Exosome delivered anticancer drugs across the blood-brain barrier for brain cancer therapy in Danio rerio. Pharm. Res. 32, 2003–2014. doi: 10.1007/s11095-014-1593-y
Zhang, D., Lee, H., Zhu, Z., Minhas, J. K., and Jin, Y. (2017). Enrichment of selective miRNAs in exosomes and delivery of exosomal miRNAs in vitro and in vivo. Am. J. Physiol.-Lung Cell. Mol. Physiol. 312, L110–L121. doi: 10.1152/ajplung.00423.2016
Zhu, L., Kalimuthu, S., Gangadaran, P., Oh, J. M., Lee, H. W., Baek, S. H., et al. (2017). Exosomes derived from natural killer cells exert therapeutic effect in melanoma. Theranostics 7, 2732–2745. doi: 10.7150/thno.18752
Zhuang, X., Xiang, X., Grizzle, W., Sun, D., Zhang, S., Axtell, R. C., et al. (2011). Treatment of brain inflammatory diseases by delivering exosome encapsulated anti-inflammatory drugs from the nasal region to the brain. Mol. Ther. 19, 1769–1779. doi: 10.1038/mt.2011.164
Keywords: extracellular vesicles, drug delivery vehicles, molecular imaging, in vivo distribution, labeling
Citation: Gangadaran P, Hong CM and Ahn B-C (2018) An Update on in Vivo Imaging of Extracellular Vesicles as Drug Delivery Vehicles. Front. Pharmacol. 9:169. doi: 10.3389/fphar.2018.00169
Received: 27 November 2017; Accepted: 15 February 2018;
Published: 28 February 2018.
Edited by:
Ruggero De Maria, Università Cattolica del Sacro Cuore, ItalyReviewed by:
Donatella Lucchetti, Università Cattolica del Sacro Cuore, ItalyMaria Felice Brizzi, Università degli Studi di Torino, Italy
Copyright © 2018 Gangadaran, Hong and Ahn. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Byeong-Cheol Ahn, YWJjMjAwMEBrbnUuYWMua3I=
 Prakash Gangadaran
Prakash Gangadaran Chae Moon Hong
Chae Moon Hong Byeong-Cheol Ahn
Byeong-Cheol Ahn