- 1Department of Physiology and Pharmacology, Schulich School of Medicine & Dentistry, The University of Western Ontario, London, ON, Canada
- 2Children’s Health Research Institute, Lawson Health Research Institute, London, ON, Canada
The derivation of canine embryonic stem cells and generation of canine-induced pluripotent stem cells are significant achievements that have unlocked the potential for developing novel cell-based disease models, drug discovery platforms, and transplantation therapies in the dog. A progression from concept to cure in this clinically relevant companion animal will not only help our canine patients but also help advance human regenerative medicine. Nevertheless, many issues remain to be resolved before pluripotent cells can be used clinically in a safe and reproducible manner.
Canine Embryonic Stem Cells
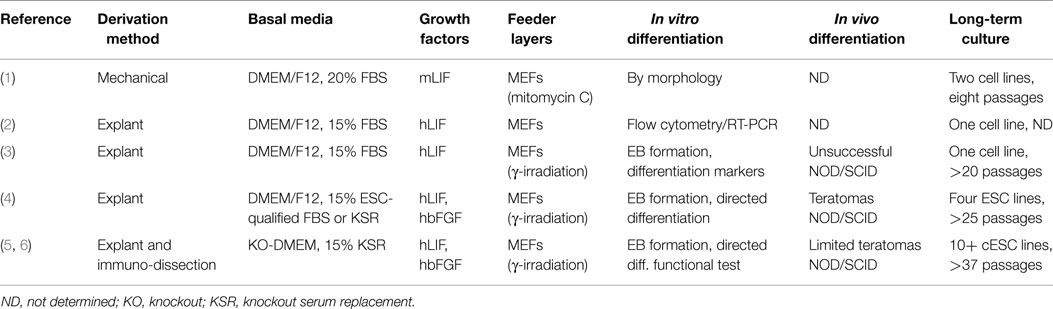
We were among a number of research groups between 2007 and 2009 who derived numerous canine embryonic stem cell (cESC) lines from blastocyst stage embryos (1–6) (Table 1). These cESC lines expressed the core pluripotency markers (Oct4, Sox2, and Nanog) and were capable of differentiating into representative lineages of all three germ layers in vitro. Limited proliferative potential was observed for several cESC lines tested by a few research groups (1, 2), while only the cESC lines established by the Hough and Betts labs were capable of forming modest teratomas in vivo (4, 5).
Canine ESC colonies seem to exhibit two phenotypically discrete morphologies, with some colonies having distinct borders and a flattened appearance similar to human ESCs (hESCs), whereas others were three-dimensional, round, tightly packed colonies that resemble mouse ESCs (mESCs). Interestingly, our mESC-like cESCs could not be successfully propagated long-term in KnockOut serum (KSR)-containing medium supplemented with leukemia inhibitory factor (LIF) and basic fibroblast growth factor (bFGF) (5), while the cESCs derived by Hayes et al. could not be cultured long-term in medium containing LIF only, but formed loosely adhered colonies interspersed with feeder cells (3). The hESC-like cESCs generated differed in growth factor dependency, as prolonged maintenance required supplementation with either soley LIF or LIF and bFGF (4, 5). Morphologically distinct cESC may represent multiple and perhaps metastable pluripotent states (e.g., naïve and primed) that have been recently characterized for both mouse and human pluripotent stem cells (PSCs) (7–9). It appears as though naïve and primed pluripotency may be at least partly conserved within other Eutherian mammals because PSC lines with naïve-like characteristics have been derived from rabbits (10), pigs (11), cows (12), and perhaps the dog (13). Many of these lines must be established and/or maintained in the presence of LIF and inhibitors of glycogen synthase kinase 3β and the MAP kinase pathway (2i), which suppress pro-differentiation stimuli (14). In depth “omics” examination of the molecular signatures underlying cESCs and preimplantation dog embryos will allow a greater understanding of the most relevant embryonic counterparts of differing metastable pluripotent states and will help define and optimize specific culture conditions for their unlimited self-renewal and directed differentiation into therapeutically desirable cell types.
Canine-Induced Pluripotent Stem Cells
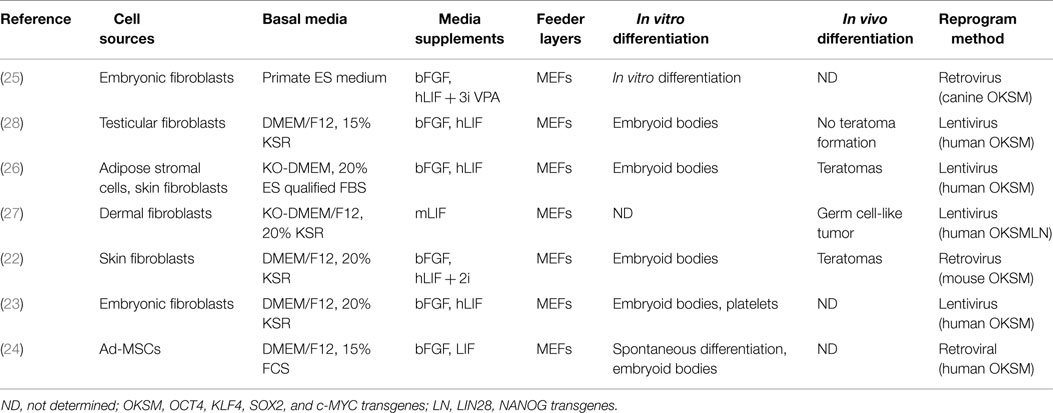
In 2006, Takahashi and Yamanaka (15) demonstrated that mouse embryonic fibroblasts could be reprogramed to a pluripotent state by the over-expression of exogenous pluripotency transcription factors (Oct-4, Klf4, Sox2) and c-Myc (OKSM). These so-called induced pluripotent cells (iPSCs) exhibit indefinite proliferative capacity and were deemed pluripotent by the expression of ESC-specific genes and the formation of embryoid bodies and teratomas. It was also shown that they could contribute to chimeric embryos with germline competency, demonstrating that iPSCs and ESCs exhibit similar in vivo differentiation capabilities (16). This initial publication has spurred a whole new research field and subsequently iPSCs have now been generated in a number of different mammals including the dog (17–19). However, random infection and integration of viral transgenes introduces both population heterogeneity and asynchrony to iPSC generation. Presumptive iPSC clones should be extensively screened with surface markers that best correlate with active endogenous pluripotency circuitry (20, 21). Unfortunately, there are discrepancies with regards to the antigen profile of ciPSCs and not all laboratories have access to canine embryos to properly control for antibody specificity (22–24). To date, there have been seven reports on the production of canine (c)iPSCs (Table 2). Two studies reprogramed canine embryonic fibroblasts (23, 25), while three used adult skin fibroblasts of various ages as the source of parental cells (22, 26, 27) by either retro- or lentiviral transduction of dog (25), human (23, 26, 27), or mouse (22) pluripotency transcription factor homologs. Two studies reported the generation of ciPSCs from adipose multipotent stromal cells (24, 26), whereas the Cibelli laboratory reprogramed from testicular fibroblasts (28). The ability to reprogram cells from multiple tissues accessible during routine procedures minimizes unnecessary harm to the patient, but it is currently unknown why different somatic cell types achieve dissimilar reprograming efficiencies (26).
Like their cESC counterparts, the majority of generated ciPSCs favor dual-factor culture of both bFGF and LIF for proliferation in the undifferentiated state (22–26, 28), but with only a few reports of in vivo teratoma formation (22, 26). A LIF-dependent ciPSC line was established using the standard “Yamanaka” factors (OCT4, KLF4, SOX2, and c-MYC) plus LIN28 and NANOG, however germ cell-like tumors were formed upon engraftment into NOD/SCID mice (27). It is presently unclear if species-specificity of reprograming factors is important for proper pluripotency induction in the dog because most of the ciPSCs were generated by retroviral transduction with human and murine homologous sequences that remain expressed in late passage ciPSCs (22, 23, 27, 28). However, these retroviral methodologies are not favorable for prospective clinical transplantation therapies because of their biased, non-random integration of transgenic sequences in promoter and coding regions, which result in dysregulation of endogenous gene expression leading to possible tumorigenesis and/or immunogenicity problems (29, 30). Alternatively, non-integrating reprograming systems have been developed and utilized in other species including episomal vectors, mini circle DNAs, plasmid vectors, small molecules, mRNAs, recombinant proteins, and transposons (31). Utilization of either the non-integrating Sendai viral-based or Cre-excisable lentiviral-based reprograming systems along with small molecular facilitators of DNA demethylation on canine fibroblasts should facilitate the generation of transgene integration-free iPS cells in the dog.
Considerations for Clinical Translation of Canine Pluripotent Stem Cell Technologies
The generation of canine pluripotent stem cells (cPSC) with disease-specific alleles and the derivation of cell types afflicted by the disease promises the development of novel cellular disease models, drug screening platforms, and potential regenerative therapies. It seems that cPSC technologies may be on the verge of clinical translation in the dog (6, 23, 26, 32), however to facilitate these cPSC-based treatments a number of hurdles need to be overcome. Canine iPSC generation efficiencies have not been routinely included in publications, but have been reported as low as 7.0 × 10−4% (27). This poses a fundamental question regarding the optimal trophic and physicochemical requirements of cPSCs. To date, we are unsure of how or when key signaling pathways are critical for the growth and epigenetic resetting of transduced canine cells. Despite of this fact, two canine somatic cell reprograming protocols have applied the transforming growth factor β (TGFβ) receptor inhibitor A-83-01 (25, 27), presumably to facilitate epithelialization of transduced cells.
Current differentiation strategies for PSCs produce cell types that correspond to immature, but lineage-committed cells from embryonic or fetal sources. Subsequently, PSC derivatives are matured with sequential growth factor exposure within a microenvironmental niche that favors the desired cell type (33). We have established synaptically competent neurons from cESCs, but were only functional when co-cultured with primary neurons derived from canine fetuses (6). Nishimura and colleagues differentiated ciPSCs into mature megakaryocytes that released functional platelets in vitro (23), while Deanne Whitworth has recently derived highly proliferative mesenchymal stromal cells (MSCs) from ciPSCs that undergo robust differentiation into the osteo-, chondro- and adipogenic cell lineages (32). Excitingly, Joseph Wu’s group at Stanford has demonstrated the preclinical potential of ciPSCs by treating immunodeficient mouse models of myocardial infarction and hindlimb ischemia with transplanted endothelial cells derived from ciPSCs (26).
All cPSC lines have been established and differentiated in culture systems that expose the canine cells to proteins or feeder cells from another species. Xeno-contamination is an important biosafety concern for graft recipients and can be limited if chemically defined culture media (i.e., serum free with recombinant factors) can be adapted for the derivation and maintenance of cPSCs (34). Moreover, technologies that enable the characterization of desired cell fates in individual cells (e.g., flow-based quantification of marker expression, RNA-sequencing, chromatin state, etc.) and the production of highly purified cell types with adult-like functional properties in vivo (e.g., small molecule inhibitors/activators of signaling pathways, three-dimensional scaffolds, etc.) need to be further developed (33).
Combining patient-specific ciPSC generation with targeted genome editing technologies (35) will enable correction of genetic defects, thereby offering potential treatment of some inherited canine diseases (36). Initially, genome editing will allow for the genetic alteration of existing cESC lines or gene correction of ciPSC lines that have been reprogramed from diseased canine cells to produce isogenic comparisons that can be utilized for disease modeling. In the long term, diseased and allele-corrected ciPSCs will serve as provocative tools within pharmacological screening platforms for efficient and predictive drug discovery and toxicity studies for the treatment of various diseases in the dog (36, 37). It is anticipated over the next few years that the CRISPR/Cas9 genome editing system will be rapidly implemented as a means to conduct loss-of-function gene mutation studies in cESCs and/or to correct genetic mutations in ciPSCs generated from fibroblasts with a disease-causing allele, as a proof of principle study.
Realizing the therapeutic potential of PSCs for clinical applications remain a central goal for the veterinary and scientific communities. Since at least half of all heritable canine diseases are known to have human equivalents (38), the dog will increasingly become an unrivaled translational animal model for developing stem cell-based therapies in humans as well. Thus, key components for any successful cPSC-based treatment will be the ability to efficiently produce ciPSCs without off-target genetic alterations or karyotypic abnormalities (39), to produce a pure population of cPSC derivatives in a scalable and good manufacturing practice (GMP)-cooperative manner, and the selection of appropriate diseases to target for regenerative medicine in the dog (33, 36). The relatively relaxed legal and ethical regulation of veterinary stem cell research compared to human medicine has facilitated the development and clinical application of a number of unproven cell-based therapies in large animals including the dog (40). The Food and Drug Administration (FDA) recently issued a draft guidance for industry on cell-based products for animal use (Draft Guidance #218) that clarifies how existing FDA regulations apply to cell-based products and encourages stakeholders to communicate and interact with the FDA early in the developmental process for each stem cell product. These guidelines and future FDA regulations of veterinary stem cell-based therapies may spearhead the much-needed double-blinded, randomized, and controlled clinical trials to properly evaluate the safety, utility, and efficacy of cPSC-based therapies in the dog.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
This commentary is an updated and revised review of a paper presented at the 2015 North American Veterinary Regenerative Medicine Association (NAVRMA) Meeting held June 28–July 1 at Asilomar Conference Hotel, Monterey, CA, USA. DB and IT thank Dr. Thomas Koch for his careful review of the manuscript. The financial support of the Natural Sciences and Engineering Research Council of Canada (NSERC) is gratefully acknowledged.
References
1. Hatoya S, Torii R, Kondo Y, Okuno T, Kobayashi K, Wijewardana V, et al. Isolation and characterization of embryonic stem-like cells from canine blastocysts. Mol Reprod Dev (2006) 73:298–305. doi: 10.1002/mrd.20392
2. Schneider MR, Adler H, Braun J, Kienzle B, Wolf E, Kolb HJ. Canine embryo-derived stem cells – toward clinically relevant animal models for evaluating efficacy and safety of cell therapies. Stem Cells (2007) 25:1850–1. doi:10.1634/stemcells.2006-0357
3. Hayes B, Fagerlie SR, Ramakrishnan A, Baran S, Harkey M, Graf L, et al. Derivation, characterization, and in vitro differentiation of canine embryonic stem cells. Stem Cells (2008) 26:465–73. doi:10.1634/stemcells.2007-0640
4. Vaags AK, Rosic-Kablar S, Gartley CJ, Zheng YZ, Chesney A, Villagomez DA, et al. Derivation and characterization of canine embryonic stem cell lines with in vitro and in vivo differentiation potential. Stem Cells (2009) 27:329–40. doi:10.1634/stemcells.2008-0433
5. Wilcox JT, Semple E, Gartley C, Brisson BA, Perrault SD, Villagomez DA, et al. Characterization of canine embryonic stem cell lines derived from different niche microenvironments. Stem Cells Dev (2009) 18:1167–78. doi:10.1089/scd.2008.0336
6. Wilcox JT, Lai JK, Semple E, Brisson BA, Gartley C, Armstrong JN, et al. Synaptically-competent neurons derived from canine embryonic stem cells by lineage selection with EGF and Noggin. PLoS One (2011) 6:e19768. doi:10.1371/journal.pone.0019768
7. Nichols J, Smith A. Naive and primed pluripotent states. Cell Stem Cell (2009) 4:487–92. doi:10.1016/j.stem.2009.05.015
8. Chen Y, Lai D. Pluripotent states of human embryonic stem cells. Cell Reprogram (2015) 17:1–6. doi:10.1089/cell.2014.0061
9. Wu J, Okamura D, Li M, Suzuki K, Luo C, Ma L, et al. An alternative pluripotent state confers interspecies chimaeric competency. Nature (2015) 521:316–21. doi:10.1038/nature14413
10. Osteil P, Tapponnier Y, Markossian S, Godet M, Schmaltz-Panneau B, Jouneau L, et al. Induced pluripotent stem cells derived from rabbits exhibit some characteristics of naive pluripotency. Biol Open (2013) 2:613–28. doi:10.1242/bio.20134242
11. Fujishiro SH, Nakano K, Mizukami Y, Azami T, Arai Y, Matsunari H, et al. Generation of naive-like porcine-induced pluripotent stem cells capable of contributing to embryonic and fetal development. Stem Cells Dev (2013) 22:473–82. doi:10.1089/scd.2012.0173
12. Verma V, Huang B, Kallingappa PK, Oback B. Dual kinase inhibition promotes pluripotency in finite bovine embryonic cell lines. Stem Cells Dev (2013) 22:1728–42. doi:10.1089/scd.2012.0481
13. Hall V, Hinrichs K, Lazzari G, Betts DH, Hyttel P. Early embryonic development, assisted reproductive technologies, and pluripotent stem cell biology in domestic mammals. Vet J (2013) 197:128–42. doi:10.1016/j.tvjl.2013.05.026
14. Ying QL, Wray J, Nichols J, Batlle-Morera L, Doble B, Woodgett J, et al. The ground state of embryonic stem cell self-renewal. Nature (2008) 453:519–23. doi:10.1038/nature06968
15. Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell (2006) 126:663–76. doi:10.1016/j.cell.2006.07.024
16. Okita K, Ichisaka T, Yamanaka S. Generation of germline-competent induced pluripotent stem cells. Nature (2007) 448:313–7. doi:10.1038/nature05934
17. Ezashi T, Telugu BP, Alexenko AP, Sachdev S, Sinha S, Roberts RM. Derivation of induced pluripotent stem cells from pig somatic cells. Proc Natl Acad Sci U S A (2009) 106:10993–8. doi:10.1073/pnas.0905284106
18. Liu J, Balehosur D, Murray B, Kelly JM, Sumer H, Verma PJ. Generation and characterization of reprogrammed sheep induced pluripotent stem cells. Theriogenology (2012) 77: 338–346.e331. doi:10.1016/j.theriogenology.2011.08.006
19. Talluri TR, Kumar D, Glage S, Garrels W, Ivics Z, Debowski K, et al. Derivation and characterization of bovine induced pluripotent stem cells by transposon-mediated reprogramming. Cell Reprogram (2015) 17:131–40. doi:10.1089/cell.2014.0080
20. Chan EM, Ratanasirintrawoot S, Park IH, Manos PD, Loh YH, Huo H, et al. Live cell imaging distinguishes bona fide human iPS cells from partially reprogrammed cells. Nat Biotechnol (2009) 27:1033–7. doi:10.1038/nbt.1580
21. Buganim Y, Faddah DA, Cheng AW, Itskovich E, Markoulaki S, Ganz K, et al. Single-cell expression analyses during cellular reprogramming reveal an early stochastic and a late hierarchic phase. Cell (2012) 150:1209–22. doi:10.1016/j.cell.2012.08.023
22. Koh S, Thomas R, Tsai S, Bischoff S, Lim JH, Breen M, et al. Growth requirements and chromosomal instability of induced pluripotent stem cells generated from adult canine fibroblasts. Stem Cells Dev (2013) 22:951–63. doi:10.1089/scd.2012.0393
23. Nishimura T, Hatoya S, Kanegi R, Sugiura K, Wijewardana V, Kuwamura M, et al. Generation of functional platelets from canine induced pluripotent stem cells. Stem Cells Dev (2013) 22:2026–35. doi:10.1089/scd.2012.0701
24. Baird A, Barsby T, Guest DJ. Derivation of canine induced pluripotent stem cells. Reprod Domest Anim (2015) 50(4):669–76. doi:10.1111/rda.12562
25. Shimada H, Nakada A, Hashimoto Y, Shigeno K, Shionoya Y, Nakamura T. Generation of canine induced pluripotent stem cells by retroviral transduction and chemical inhibitors. Mol Reprod Dev (2010) 77:2. doi:10.1002/mrd.21117
26. Lee AS, Xu D, Plews JR, Nguyen PK, Nag D, Lyons JK, et al. Preclinical derivation and imaging of autologously transplanted canine induced pluripotent stem cells. J Biol Chem (2011) 286:32697–704. doi:10.1074/jbc.M111.235739
27. Whitworth DJ, Ovchinnikov DA, Wolvetang EJ. Generation and characterization of LIF-dependent canine induced pluripotent stem cells from adult dermal fibroblasts. Stem Cells Dev (2012) 21:2288–97. doi:10.1089/scd.2011.0608
28. Luo J, Suhr ST, Chang EA, Wang K, Ross PJ, Nelson LL, et al. Generation of leukemia inhibitory factor and basic fibroblast growth factor-dependent induced pluripotent stem cells from canine adult somatic cells. Stem Cells Dev (2011) 20:1669–78. doi:10.1089/scd.2011.0127
29. Hacein-Bey-Abina S, Von Kalle C, Schmidt M, McCormack MP, Wulffraat N, Leboulch P, et al. LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science (2003) 302:415–9. doi:10.1126/science.1088547
30. Zhao T, Zhang ZN, Rong Z, Xu Y. Immunogenicity of induced pluripotent stem cells. Nature (2011) 474:212–5. doi:10.1038/nature10135
31. Kumar D, Talluri TR, Anand T, Kues WA. Induced pluripotent stem cells: mechanisms, achievements and perspectives in farm animals. World J Stem Cells (2015) 7:315–28. doi:10.4252/wjsc.v7.i2.315
32. Whitworth DJ, Frith JE, Frith TJ, Ovchinnikov DA, Cooper-White JJ, Wolvetang EJ. Derivation of mesenchymal stromal cells from canine induced pluripotent stem cells by inhibition of the TGFbeta/activin signaling pathway. Stem Cells Dev (2014) 23:3021–33. doi:10.1089/scd.2013.0634
33. Tabar V, Studer L. Pluripotent stem cells in regenerative medicine: challenges and recent progress. Nat Rev Genet (2014) 15:82–92. doi:10.1038/nrg3563
34. Desai N, Rambhia P, Gishto A. Human embryonic stem cell cultivation: historical perspective and evolution of xeno-free culture systems. Reprod Biol Endocrinol (2015) 13:9. doi:10.1186/s12958-015-0005-4
35. Li M, Suzuki K, Kim NY, Liu GH, Izpisua Belmonte JC. A cut above the rest: targeted genome editing technologies in human pluripotent stem cells. J Biol Chem (2014) 289:4594–9. doi:10.1074/jbc.R113.488247
36. Cebrian-Serrano A, Stout T, Dinnyes A. Veterinary applications of induced pluripotent stem cells: regenerative medicine and models for disease? Vet J (2013) 198:34–42. doi:10.1016/j.tvjl.2013.03.028
37. Ebert AD, Liang P, Wu JC. Induced pluripotent stem cells as a disease modeling and drug screening platform. J Cardiovasc Pharmacol (2012) 60:408–16. doi:10.1097/FJC.0b013e318247f642
38. Starkey MP, Scase TJ, Mellersh CS, Murphy S. Dogs really are man’s best friend – canine genomics has applications in veterinary and human medicine! Brief Funct Genomic Proteomic (2005) 4:112–28. doi:10.1093/bfgp/4.2.112
39. Koh S, Piedrahita JA. From “ES-like” cells to induced pluripotent stem cells: a historical perspective in domestic animals. Theriogenology (2014) 81:103–11. doi:10.1016/j.theriogenology.2013.09.009
Keywords: canine, dog, embryonic stem cells, genome editing, induced pluripotent stem cells, transplantation therapy
Citation: Betts DH and Tobias IC (2015) Canine pluripotent stem cells: are they ready for clinical applications? Front. Vet. Sci. 2:41. doi: 10.3389/fvets.2015.00041
Received: 09 July 2015; Accepted: 21 September 2015;
Published: 07 October 2015
Edited by:
Dominique J. Griffon, Western University of Health Sciences, USAReviewed by:
Ruchi Sharma, Axol Biosciences, UKWanda J. Gordon-Evans, Wisconsin Veterinary Referral Center, USA
Copyright: © 2015 Betts and Tobias. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Dean H. Betts, Department of Physiology and Pharmacology, Schulich School of Medicine & Dentistry, The University of Western Ontario, London, ON N6A 5C1, Canada,ZGVhbi5iZXR0c0BzY2h1bGljaC51d28uY2E=
 Dean H. Betts
Dean H. Betts Ian C. Tobias
Ian C. Tobias