- 1National Reference Laboratory for Animal Brucellosis, China Institute of Veterinary Drug Control, Beijing, China
- 2South China Agricultural University, Guangzhou, China
- 3Zhaoqing Institute of Biotechnology Co., Ltd, Zhaoqing, China
- 4China Animal Disease Control Center, Beijing, China
Brucellosis is one of the most common zoonotic epidemics with a serious threat to public health and livestock development in many countries across the world. Vaccination is a key control strategy toward preventing brucellosis in high-prevalence regions. Recently, a rough-type Brucella melitensis mutant strain (RM57) induced from a B. melitensis strain M1981 showed protective effects in guinea pigs indicating that it is a good vaccine candidate. In this study, stress response assays were performed to reveal the mechanisms underlying virulence attenuation of RM57. In addition, a genome-wide transcriptome profile of RM57 was analyzed relative to the parent strain M1981 in order to reveal genetic factors controlling the phenotypes. Our results indicated a similar sensitivity to various stress conditions in RM57 owing to a lack of significant differences from its parent strain. Transcriptome analysis showed that a total of 1,205 genes were differentially expressed between RM57 and M1981 with gene ontology terms revealing that these genes are involved in energy production and conversion, translation, ribosomal structure, and biogenesis. Pathway enrichment analysis revealed that genes involved in oxidative phosphorylation, ribosome, nitrogen metabolism, tyrosine metabolism, and two-component system were significantly affected. As a result of these differences at the molecular level, the function of type IV secretion system in RM57 was found to be affected leading to reduced virulence of the RM57 mutant strain in both macrophage and mice infection models.
Introduction
Brucellosis, an infectious disease caused by Brucella spp. and spreads through unpasteurized dairy products, remains one of the most serious bacterial zoonosis across the world. In animals, this pathogen mainly causes abortion in females and male infertility although various other clinical symptoms including fever, night sweats, anorexia, polyarthritis, meningitis, and pneumonia have been reported in humans. Humans usually are infected by Brucella spp. through consumption of contaminated dairy products, inhalation of bacterial aerosols as well as contact with wounds or mucosal tissues (1). Due to the serious economic losses and public health risks caused by this disease, regulatory authorities across many countries are engaged in measures toward prevention, control, and elimination of brucellosis in domestic animals thereby guaranteeing safety of dairy products (2).
So far, vaccination has been successful as a control strategy for preventing exposure to brucellosis in regions where the disease is prevalent (3). Consequently, various studies have demonstrated that live Brucella vaccines such as S19, RB51, S2, and Rev.1 are able to effectively offer immune protection against brucellosis and these are therefore widely used around the world (4). However, negative implications of these vaccines including diagnostic interference, residual virulence in the host, and pathogenic effects to human health cast doubts to this control strategy (3, 5–7).
To overcome these drawbacks and build an efficient and reliable vaccine strategy against the disease, a rough attenuated vaccine candidate Brucella melitensis RM57 (Accession: SAMN12827002), free of antibiotic resistance, was developed from its parent strain B. melitensis M1981 (Accession: SAMN12827001). As RM57 was attenuated in mice and guinea pig infection models and showed promise as an ideal vaccine candidate, we aimed to unravel the mechanism of virulence attenuation of the RM57 strain through phenotypic characterization and transcriptome analysis. In this study, we therefore assessed RM57 phenotypes that are pertinent to infection and virulence using macrophages and mice models and further performed RNA sequencing analysis to study the genetic factors controlling the observed phenotypes.
Materials and Methods
Ethics Statement
The female BALB/c mice were handled in strict accordance with the Experimental Animal Regulation Ordinances defined by the China National Science and Technology Commission. The study was approved by the animal ethics committee of China Institute of Veterinary Drug Control under permit number (CIVDC 2019-000672).
Bacteria Strains and Culture Conditions
The B. melitensis M1981 strain was isolated in China in 2007. For induction of the RM57 rough strain, the M1981 strain was first induced with tryptic soy broth (TSB, Becton, Dickinson and Company, Sparks, MD, USA) medium containing chloramphenicol (1 μg/ml) for 40 generations. The rough-type colony was selected and incubated with anti-B. melitensis serum at 37°C for 2 h and 4°C for 12 h. The rough-type colony of the non-agglutinated bacteria culture medium was selected and induced with anti-B. melitensis serum again as described above. After the rough-type strain was induced by the antiserum for 45 generations, it was continuously passaged 12 generations in guinea pigs to finally obtain the RM57 strain, which was stable in both phenotype and bacteria virulence.
The Brucella strains were routinely grown in TSB at 37°C or on tryptic soy agar (TSA, Becton, Dickinson and Company, Sparks, MD, USA) medium incubated at 37°C. All the bacterial strains were frozen at −80°C and supplemented with 25% (v/v) glycerol.
For the growth curve assay, Brucella strains RM57 and M1981 (initial density of 1 × 106 CFU/ml) were grown in TSB medium at 37°C with continuous shaking. The CFUs of RM57 and M1981 were measured by plate count at different time points.
Lipopolysaccharide Extraction and Silver Staining
The parent strain M1981 and rough type strain RM57 were cultured in TSB medium then grown to the stationary phase at 37°C with continuous shaking. Bacterial cells were collected by centrifugation then LPS was extracted using an LPS Extraction Kit (iNtRON, Seoul, Korea) following the manufacturer's instructions. Samples were loaded on a 12% polyacrylamide gel for SDS-PAGE analysis followed by the silver staining assay as previously described (8).
Transcriptome Analysis
To understand genetic regulation of virulence in the Brucella strains under this study, we performed a transcriptome analysis for M1981 and RM57. Briefly, 3 samples of each Brucella strain from a single colony was grown in tubes with TSB at 37°C until the log phase was reached. Total RNA was then isolated from the bacterial pellets using the TRIzol method (Invitrogen, Carlsbad, CA, USA) according to the manufacturer's instructions. Residual DNA in the RNA samples was removed using DNase I (Thermo Scientificm, USA). RNA concentration and purity were determined using a nanodrop spectrophotometer (ND 1000 spectrophotometer Thermo Scientific, Wilmington, USA).
The sequencing library of each RNA sample was prepared by using NEBNext Ultra Directional RNA Library Prep Kit for Illumina as recommended by the manufacturer. In brief, RNA fragments were reverse-transcribed and amplified to double-stranded cDNA and then ligated with an adaptor. The amplified cDNA was purified using magnetic bead based method, and the molar concentration was determined for each cDNA library. The HiSeq 4000 platform was used to perform the transcriptome sequencing. Sequencing and subsequent bioinformatics analysis were completed at Novel Bioinformatics Co., Ltd, Shanghai, China.
Quantitative Real Time Polymerase Chain Reaction
To validate gene expression, a quantitative real time polymerase chain reaction (qRT-PCR) was performed as previously described (9) targeting differentially expressed genes earlier identified in the RNAseq analysis. The genes targeted as well as primer sequences used for qRT-qPCR are provided in Table S1. Real time PCR was run in triplicates in a 20 μl reaction volume containing 10 μl 2×SYBR® Premix Ex TaqTM II (TAKARA), 100 nM forward and reverse primers, and 1 μl appropriately diluted cDNA as the template. To normalize the expression, 16S rRNA, which is constantly transcribed in bacteria, was used as an internal amplification control. qRT-PCR analysis was performed in an Applied Biosystems 7500 Real-Time PCR System and the relative transcription levels were determined by the 2−ΔΔCt method.
Determination of Cell Viability
To investigate intracellular viability of M1981 and RM57 strains in vitro, we obtained RAW264.7 cells from the Cell Resource Center, IBMS, CAMS/PUMC (Beijing, China) and performed cell infection assays as previously described (10). Briefly, cells were cultured in 24-well plates (Corning, NY, USA) then infected with Brucella strains at a multiplicity of infection (MOI) of 100. After 1 h of incubation, plates containing the cells were washed three times using PBS and then incubated in a medium containing gentamycin (50 μg/ml) to kill extracellular bacteria. At 1, 12, 24, 48, and 72 h post-infection (hpi), the cultures were washed and lysed with 500 μl 0.1% (v/v) Triton X-100-water solution, then the number of surviving bacterial cells was determined using the plating technique on TSA. The experiment was performed in triplicates and repeated at least twice.
Infection Assays Using Animal Models
A total of 45 female 4–6 week-old BALB/c mice were used in the infection assay. The test animals were acquired from the Beijing Vital River Laboratory Animal Technology Co., Ltd then randomly divided into three groups. In each group, mice were inoculated intraperitoneally with 100 μl (~1 × 105 CFUs per animal) of RM57 mutant strain and the parental strain M1981. For the control group, PBS solution was used. Infected and control animals (n = 5 per group) were euthanized at 14, 28, and 42 days post infection (dpi) via carbon dioxide asphyxiation, sacrificed then their spleens analyzed for level of infection. First, the spleen was carefully detached from the rest of the body, weighed and homogenized in 1 ml of PBS. Bacterial colonies in each sample were then enumerated by TSA plate count followed by calculation of CFUs per spleen.
Sensitivity of Brucella Strains to the Bactericidal Action of Non-immune Serum
We evaluated the effect of bactericidal action in non-immune serum on sensitivity in Brucella strains according to a previously reported protocol (11). Briefly, cultures of RM57 and M1981 strains were adjusted to 104 CFUs/ml using PBS then dispensed in microtiter plates (at a volume of 30 μl/well) containing 60 μl of new-born bovine serum. Cultures were incubated for 90 min at 37°C with gentle agitation then complement action was blocked by adding 150 μl/well of TSB medium. The TSB medium was mixed well with the bacterial suspension then 75 μl was plated on TSA plates in triplicates. The plated cultures were incubated at 37°C for 3 days followed by an analysis of sensitivity. The results were expressed as the percentage of CFU recovered with respect to control samples where new-born bovine serum was substituted by PBS.
Determination of Brucella Strain Susceptibility Under Stress Conditions
To test susceptibility of Brucella strains under stress conditions, an assay was modified from a previously reported study (10).
In stress assay, initially both Brucella strains (with an initial density of 1 × 107 CFU/ml) were cultured in TSB (5% EtOH, pH4.5, 2.5 mM H2O2) for 2 h. Afterwards, the concentration of bacteria was measured by plate count, and the survival rate (%) was calculated as surviving bacteria relative to the TSB control.
In order to determine the sensitivity of RM57 mutant to heat shock stress, iron deficiency and hypertonic environments, the bacteria were cultured in TSB medium and treated with 42°C (heat shock stress), 10 mM 2′2- dipyridyl (iron deficiency), and 500 mM NaCl (hypertonic environment), respectively. The bacteria were cultured in this medium at the same initial density (1 × 107 CFU/ml) and the CFUs were determined at 24 and 48 h post-treatment for both strains. The survival rate (%) was calculated as surviving bacteria relative to the 37°C as optimum temperature control.
Effector Translocation Assay
To determine whether the type IV secretion function in RM57 strain was affected, an effector translocation assay was performed as previously described (12, 13). Briefly, a TEM1-VceC fusion plasmid based on the original pZL1790-TME1 was constructed then introduced into M1981 and RM57 by electroporation. To perform the effector translocation assay, 5 × 104 RAW264.7 macrophages were seeded into 96-well plates and infected with M1981 and RM57 expressing the TEM1-VceC fusion protein at a multiplicity of infection of 1,000:1. Cells were incubated for 30 min at 37°C in 5% CO2 then each plate washed twice with PBS and 0.1 ml DMEM with 0.5 mM IPTG. The plates were incubated at 37°C in 5% CO2, washed once with Hank's balanced salt solution (Gibco, Grand Island, NY) 15 h post-infection then loaded with a solution containing the fluorescent substrate CCF4/AM at a final concentration of 1 mM. The cultures were left to stand for 2 h at room temperature according to the standard protocol (Invitrogen, Carlsbad, CA) then the translocation of TEM1-VceC fusion protein into the macrophages was analyzed by fluorescence microscopy.
Statistical Analysis
Basic statistical analyses were performed using SPSS 16.0 (SPSS Inc., Chicago, IL, USA). In the virulence assay of RM57 strain in mice, an unpaired Student's t-test was performed at each time point. In the group analysis, Analysis of Variance (ANOVA) method was used for data analyses. Differences were considered significant at P-values of <0.05.
Results
Growth and Lipopolysaccharide (LPS) Characterization of RM57 Mutant Strain
To characterize the growth rate of Brucella mutant strains, RM57 and M1981 were cultured in TSB medium at 37°C following a similar initial optical density. We observed that the growth rate of RM57 mutant strain was almost similar as its parent strain M1981 (Figure 1). On the other hand, findings from the LPS silver staining showed that the rough type Brucella RM57 had a modified pattern of O-antigen compared to its parent (Figure 2).
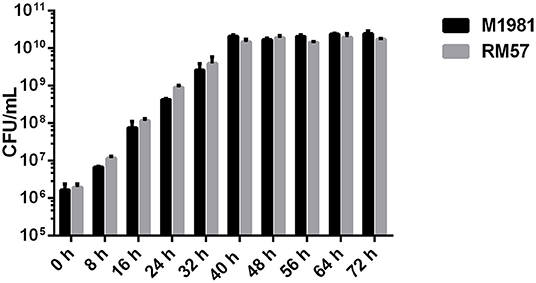
Figure 1. Growth rate of RM57 mutant and its parent strain M1981 in TSB medium. Brucella strains (initial density of 1 × 106 CFU/ml) were grown in TSB at 37°C with continuous shaking for 72 h. The CFUs of RM57 and M1981 were measured at different time points.
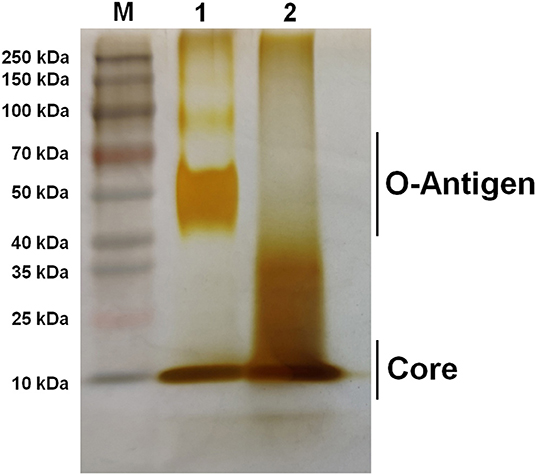
Figure 2. Silver staining of bacterial LPS. Lane M: Prestain page ruler (Thermo Scientific, USA); Lane 1: M1981 LPS; Lane 2: RM57 LPS.
RM57 Was Attenuated in Macrophage and Mice Infection Models
To assess virulence of the RM57 strain, its ability to multiply within a cultured macrophage RAW264.7 was tested. It was observed that intracellular bacterial load of the RM57 strain was almost similar to that of M1981 during the first 24 h post infection. However, RM57 load significantly reduced at 48 (P < 0.05) and 72 h (P < 0.05) in murine macrophages compared to the parent line M1981 (Figure 3).
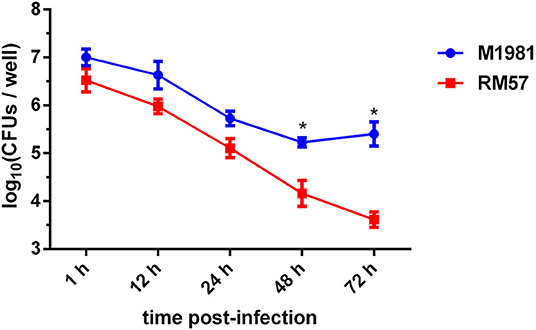
Figure 3. Profiles of virulence attenuation in RM57 in macrophage infection models. Multiplication of RM57 and M1981 strains in RAW264.7 macrophages over 72 h. Values represent the means of three experiments performed in duplicate while error bars indicate the standard deviation (SD) of the mean. *p < 0.05.
A similar pattern was observed when mice models were used. Analysis of spleen weight following infection showed that mice infected by the RM57 mutant strain had a significantly lower weight than those infected with M1981 at 2, 4, and 6 weeks post infection (P < 0.001). On the other hand, spleens recovered from mice infected with RM57 were significantly heavier than those from the negative control group (P < 0.01) at 2 weeks post infection while at 4 and 6 weeks post infection, spleens from mice infected with RM57 and PBS group showed no significant difference (P > 0.05) (Figure 4A). In addition, the RM57 mutant recovered from the spleens of mice was significantly reduced at 2 week post infection (P < 0.01) with no mutant strain recovered from the spleens of mice at 4 and 6 weeks post infection (Figure 4B).
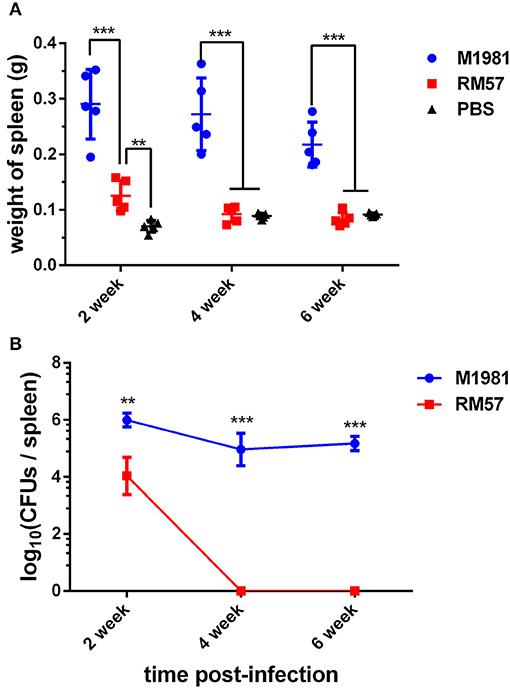
Figure 4. Virulence of RM57 attenuation in mice infection models. (A) Spleen weight of RM57 and M1981 infected mice at 2, 4, and 6 weeks post infection. (B) Splenic CFUs of RM57 and M1981 infected mice at 2, 4, and 6 weeks post infection. Values represent the means of spleen weight or splenic CFUs at each time point, and the error bars indicate the standard deviation (SD) of the mean. *p < 0.01 and ***p < 0.001.
RM57 Shows No Sensitivity to Stress Conditions
Assaying sensitivity of Brucella strains against a hypertonic environment, iron deficiency medium, and heat stress conditions revealed no significant effect of these conditions to the bacteria. Particularly, survival rate of the RM57 mutant strain showed normal growth in TSB medium as well as when challenged with H2O2, and 5% EtOH (Figure 5A). Similarly, there was no observable difference in survival ratio of the two strains when challenged with acidic environment (Figure 5A). Previously, the rough type Brucella strains have been described as being more sensitive to the bactericidal action of normal serum than their smooth counterparts (11). However, after incubation for 90 min in non-immune serum, we observed that the survival rate of RM57 was not significantly reduced compared to the parent strain M1981 (Figure 5A). Similarly, exposure to the other stress conditions of hypertonic environment, iron deficiency medium and heat stress conditions revealed no differences in survival rates (Figures 5B–D).
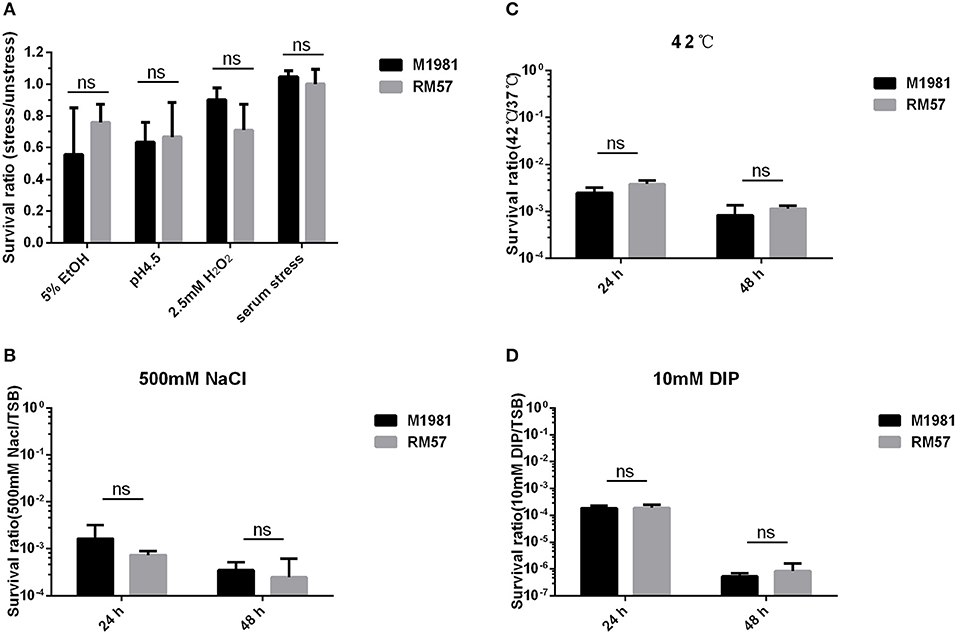
Figure 5. (A) The survival ratio of RM57 and M1981 cultured in TSB (5% EtOH, pH4.5, 2.5 mM H2O2, and non-immune serum) for 2 h. (B) The survival ratio of RM57 and M1981 cultured in TSB at 42°C (heat shock stress). (C) The survival ratio of RM57 and M1981 cultured in TSB with 500 mM NaCl (hypertonic environment). (D) The survival ratio of RM57 and M1981 cultured in TSB with 10 mM 2′2-dipyridyl (iron deficiency).
Identification of Differentially Expressed Genes in Brucella Strains
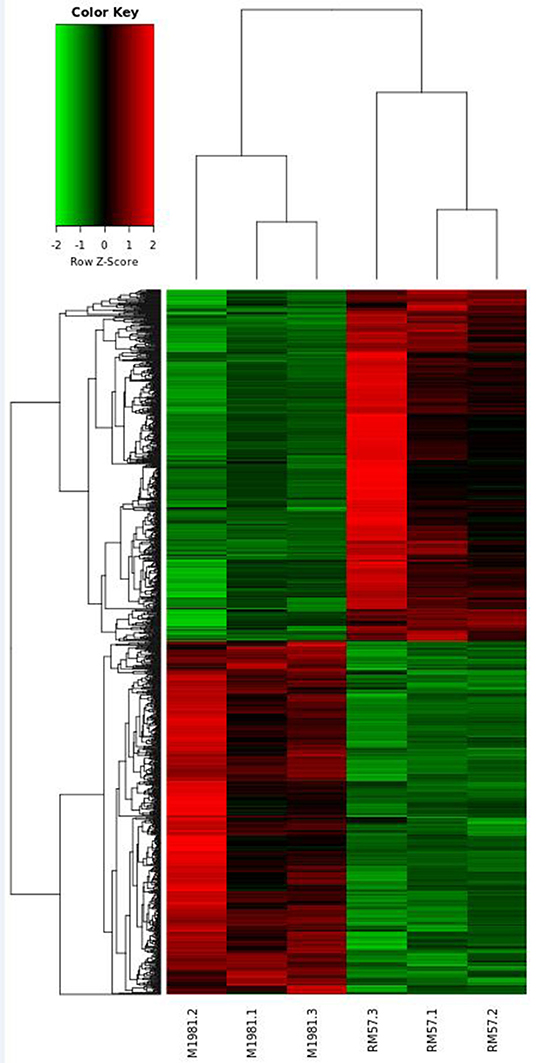
To reveal genetic factors regulating virulence attenuation of RM57 relative to the parent strain, changes in transcriptional profiles of gene expression in the two strains were determined using transcriptomic analysis following RNA sequencing. The transcriptome dataset has been submitted to the GEO repository and is available using accession number GSE131394. Transcriptome data showed that a total of 1,205 genes exhibited significant differences (log2FC > 1 or <−1 and FDR < 0.05) and among these, 597 were significantly up-regulated while 608 were significantly down-regulated in RM57 compared to its parent strain M1981 (Figure 6). Among these, the highest ranked upregulated genes included BMEI1041, BMEII0642, BMEII1003, BMEI1040, BMEII0987, BMEII0105, BMEI1362, BMEII0988, BMEII0353, and BMEII0321 while BMEI0427, BMEI1313, BMEI0877, BMEII0948, BMEII0759, BMEI0422, BMEI0386, BMEII0758, BMEI0454, and BMEII0844 were highly down-regulated.
With regard to gene ontology terms, the differentially expressed genes mainly comprised those involved in energy production and conversion and translation, ribosomal structure and biogenesis (Figure 7A). KEGG pathway analysis further revealed that the genes which exhibited significant differences were primarily enriched in oxidative phosphorylation, ribosome, nitrogen metabolism, tyrosine metabolism, and two-component system (Figure 7B).
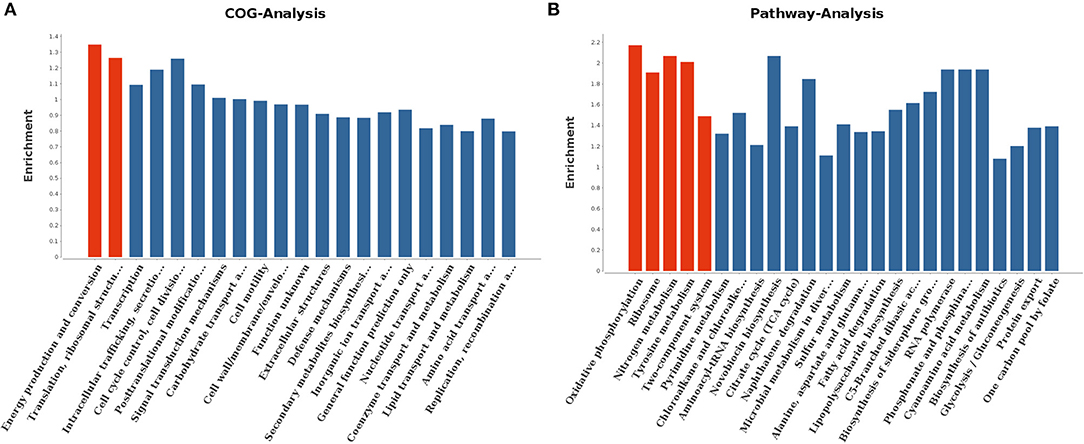
Figure 7. COG analysis (A) and KEGG pathway analysis (B) for differentially expressed genes. The red columns indicated these categories or pathways were significantly enriched (p < 0.05) while the blue columns means these categories or pathways were not significantly enriched.
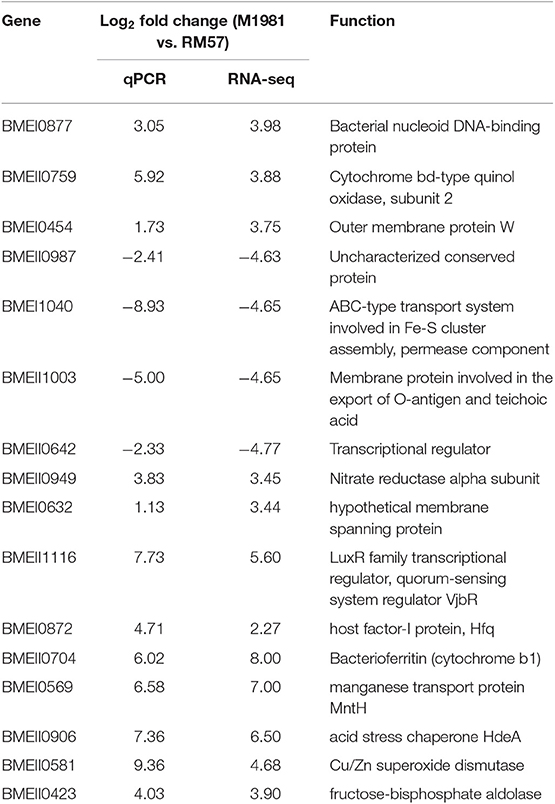
To validate the RNA-seq result, 16 genes were chosen and quantified via qRT-PCR. Expression levels of the 16 selected genes resulted in a similar profile using both qRT-PCR and RNA-seq and this is outline in Table 1.
Expression Profiles of Genes Encoding Virulence Factors Show a Variation Between the Mutant and Parent Strain
Based on the result of RNA-seq, it was found that the expression levels of several genes encoding virulence factors were significantly reduced in the mutant strain relative to the parent line. Particularly, BMEII1116 (encoding quorum-sensing system regulator VjbR) and BMEI0872 (encoding Hfq protein) were both significantly down-regulated in RM57 relative to M1981. A similar profile was observed in the qRT-PCR result (Table 1). In addition, expression of four genes in the virB operon (BMEII0029, BMEII0030, BMEII0034, and BMEII0035) which encodes the type IV secretion system showed a downregulation recording a reduction in the fold change from 2.2 to 3.7 fold.
Type IV Secretion System Is Affected in RM57 Mutant Strain
To determine whether the function of T4SS in RM57 was affected, M1981 and RM57 expressing the TEM1-VceC fusion protein were used to infect RAW264.7 macrophages to determine the translocation of the VceC protein (an effector of the Brucella Type IV secretion system translocated into macrophages) (12, 13). We observed the translocation of TEM1-VceC protein into RAW264.7 macrophages infected by M1981, and the percentage of blue cells was ~10.25% at 15 h post-infection (Figure 8A). However, about 1.69% of the cells were blue when infected with RM57 (Figure 8B). These results indicated that the function of T4SS was affected in the RM57 mutant strain.
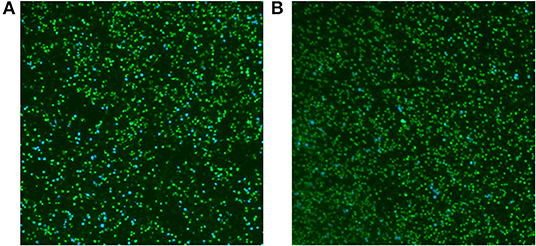
Figure 8. Translocation of the effector VceC into mouse macrophages RAW264.7 infected by M1981 (A) and RM57 mutant strain (B). Cells in which translocation of the fusion protein has occurred appear as blue. The result was a representative individual experiment repeated three times.
Discussion
The current study describes phenotypic characterization and transcriptome analysis of two strains of B. melitensis and further reveals sensitivity to various stress conditions. RM57, a rough mutant strain was previously induced from a parent line B. melitensis isolate M1981 and has been shown to be attenuated in both mice and guinea pig infection models (14). Results of the current study corroborated this earlier report indicating that RM57 was attenuated in both microphage and mice infection models. Previous studies have demonstrated that LPS of Brucella played a key role in virulence as it could impair normal recognition by the innate immune system thereby delaying an organism's immune response (11). According to our LPS silver staining, we observed that O-antigen in RM57 was modified and this was consistent with the increased sensitivity to SDS and polymyxin B in the RM57 mutant strain as previously reported (14). It is therefore possible that the defective LPS was involved in the reduced virulence of RM57 mutant strain relative to the parent line M1981.
Facultative intracellular pathogens, such as B. melitensis, encounter considerable environmental stresses during their interactions with host cells including exposure to reactive oxygen and nitrogen species, unfavorable pH ranges as well as nutritional deprivation (15). In line with this, we carried out stress response assays for the Brucella mutant strains under induced stresses of heat, iron deficiency, and hypertonic environment and compared the findings with those from the parent line. Our data indicated that RM57 was not sensitive to the aforementioned stress conditions.
To further unravel the genetic factors regulating the related phenotypes and virulence attenuation of RM57 relative to the parental line, we generated a transcriptome profile of RM57 and M1981 and determined differentially expressed genes. We observed that up to 1,205 genes were differentially expressed (>2-fold in RM57) thereby indicating that RM57 exhibits transcriptional changes relative to M1981. Among the differentially expressed genes mutant and the parent, we focused on the down-regulated genes in mutant strain. Here, several genes involved in Brucella virulence on a host were significantly down-regulated in the mutant including BMEII1116, BMEI0872, and virB operon. The BMEII1116 gene encodes a LuxR-type regulator VjbR which is a well-known virulence factor of Brucella spp. with various studies demonstrating the major role played by this gene in modulating expression of several Brucella virulence factors such as VirB operon, VirB effectors, flagellar components, and cell surface structures (12, 16).
Similarly, previous transcriptomic and proteomic analyses have implicated VjbR indirect or indirect expression of hundreds of genes (17, 18). In the current study, expression levels of vjbR was reduced more than 5-fold, pointing toward a possible reduction in virulence of RM57. In addition, as VjbR could directly up-regulate the expression of virB operon in Brucella strains, the low-expression level of VjbR might not completely induce the expression of virB operon in RM57 (12).
Our results further showed that, expression levels of hfq were significantly reduced in the mutant strain relative to the parent. Hfq protein, encoded by the BMEI0872 gene is an important RNA chaperone involved in stress resistance and Brucella virulence in both macrophage and mice infection models (19). Furthermore, recent studies have demonstrated that this chaperone can regulate virB operon expression and plays a key role in the interaction between sRNAs and their target mRNAs (20, 21). Downregulation of this gene in our mutant strain therefore indicates a reduction in virulence relative to the parental line. We also observed a marked downregulation of type IV secretion system encoding genes in RM57 relative to the parent line. Type IV secretion system is important in Brucella spp. for evading fusion of endosome vesicles with lysosomes through secretion of effector protein. This system, which is encoded by virB operon, also plays a key role in Brucella virulence. Brucella mutant strains that lack a functional type IV secretion system are reported to result in high attenuation in both macrophages and mouse infection models (12).
Furthermore, we observed that the function of type IV secretion system in the mutant strain was affected relative to its parent strain thereby indicating that an abnormal type IV secretion system could have been the cause of the observed reduction in bacterial virulence.
In summary, although the mutant Brucella strain was not sensitive to various stress responses at the phenotypic level, comparative transcriptome analysis and expression profiles of various important virulence factors revealed changes between the mutant and parent line. These changes at the molecular level could be playing a key role in the attenuation of RM57.
Conclusion
This work provides insights into the molecular regulation of LPS deficiency in RM57, which was unknown, and further generates knowledge on the differences between this mutant and its parental line.
Data Availability Statement
The data in this study can be found in the GEO database (https://www.ncbi.nlm.nih.gov/geo/), with the accession number: GSE131394.
Ethics Statement
The animal study was reviewed and approved by Animal ethics committee of China Institute of Veterinary Drug Control.
Author Contributions
JD, XP, and HD designed the experiments. YL, HJ, KN, QG, and JS performed the experiments. HD and YQ prepared the manuscript. YF analyzed the data. All authors have read and approved the final version of this manuscript.
Funding
This work was supported by the National Natural Science Foundation of China (Nos. 31602017 and 31602055), and the National Key Research and Development Program of China (No. 2016YFD0500902). Major Applied Agricultural Technology Innovation Projects in Shandong province.
Conflict of Interest
YL was employed by company Zhaoqing Institute of Biotechnology Co., Ltd.
The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Novel Bioinformatics Ltd., Co for the support of bioinformatics analysis with their Novel Bio Cloud Analysis Platform.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fvets.2019.00459/full#supplementary-material
References
1. Corbel MJ. Brucellosis: an overview. Emerg Infect Dis. (1997) 3:213–21. doi: 10.3201/eid0302.970219
2. Olsen SC, Stoffregen WS. Essential role of vaccines in brucellosis control and eradication programs for livestock. Expert Rev Vaccines. (2005) 4:915–28. doi: 10.1586/14760584.4.6.915
3. Hou H, Liu X, Peng Q. The advances in brucellosis vaccines. Vaccine. (2019) 37:3981–8. doi: 10.1016/j.vaccine.2019.05.084
4. Zhu L, Feng Y, Zhang G, Jiang H, Zhang Z, Wang N, et al. Brucella suis strain 2 vaccine is safe and protective against heterologous Brucella spp. infections. Vaccine. (2016) 34:395–400. doi: 10.1016/j.vaccine.2015.09.116
5. Truong QL, Cho Y, Park S, Kim K, Hahn TW. Brucella abortusΔcydCΔcydD and ΔcydCΔpurD double-mutants are highly attenuated and confer long-term protective immunity against virulent Brucella abortus. Vaccine. (2016) 34:237–44. doi: 10.1016/j.vaccine.2015.11.030
6. Schurig GG, Sriranganathan N, Corbel MJ. Brucellosis vaccines: past, present and future. Vet Microbiol. (2002) 90:479–96. doi: 10.1016/S0378-1135(02)00255-9
7. Ashford DA, di Pietra J, Lingappa J, Woods C, Noll H, Neville B, et al. Adverse events in humans associated with accidental exposure to the livestock brucellosis vaccine RB51. Vaccine. (2004) 22:3435–9. doi: 10.1016/j.vaccine.2004.02.041
8. Bao Y, Tian M, Li P, Liu J, Ding C, Yu S. Characterization of Brucella abortus mutant strain Δ22915, a potential vaccine candidate. Vet Res. (2017) 48:17. doi: 10.1186/s13567-017-0422-9
9. Liu W, Dong H, Liu W, Gao X, Zhang C, Wu Q. OtpR regulated the growth, cell morphology of B. melitensis and tolerance to beta-lactam agents. Vet Microbiol. (2012) 159:90–8. doi: 10.1016/j.vetmic.2012.03.022
10. Liu Y, Dong H, Peng X, Gao Q, Jiang H, Xu G, et al. RNA-seq reveals the critical role of Lon protease in stress response and Brucella virulence. Microb Pathog. (2019) 130:112–9. doi: 10.1016/j.micpath.2019.01.010
11. Salvador-Bescos M, Gil-Ramirez Y, Zuniga-Ripa A, Martinez-Gomez E, de Miguel MJ, Munoz PM, et al. WadD, a new brucella lipopolysaccharide core glycosyltransferase identified by genomic search and phenotypic characterization. Front Microbiol. (2018) 9:2293. doi: 10.3389/fmicb.2018.02293
12. de Jong MF, Sun YH, den Hartigh AB, van Dijl JM, Tsolis RM. Identification of VceA and VceC, two members of the VjbR regulon that are translocated into macrophages by the Brucella type IV secretion system. Mol Microbiol. (2008) 70:1378–96. doi: 10.1111/j.1365-2958.2008.06487.x
13. Wang Z, Liu W, Wu T, Bie P, Wu Q. RNA-seq reveals the critical role of CspA in regulating Brucella melitensis metabolism and virulence. Sci China Life Sci. (2016) 59:417–24. doi: 10.1007/s11427-015-4981-6
14. Feng Y, Peng X, Jiang H, Peng Y, Zhu L, Ding J. Rough brucella strain RM57 is attenuated and confers protection against Brucella melitensis. Microb Pathog. (2017) 107:270–5. doi: 10.1016/j.micpath.2017.03.045
15. Roop RN, Gaines JM, Anderson ES, Caswell CC, Martin DW. Survival of the fittest: how Brucella strains adapt to their intracellular niche in the host. Med Microbiol Immunol. (2009) 198:221–38. doi: 10.1007/s00430-009-0123-8
16. Delrue RM, Deschamps C, Leonard S, Nijskens C, Danese I, Schaus JM, et al. A quorum-sensing regulator controls expression of both the type IV secretion system and the flagellar apparatus of Brucella melitensis. Cell Microbiol. (2005) 7:1151–61. doi: 10.1111/j.1462-5822.2005.00543.x
17. Uzureau S, Lemaire J, Delaive E, Dieu M, Gaigneaux A, Raes M, et al. Global analysis of quorum sensing targets in the intracellular pathogen Brucella melitensis 16 M. J Proteome Res. (2010) 9:3200–17. doi: 10.1021/pr100068p
18. Weeks JN, Galindo CL, Drake KL, Adams GL, Garner HR, Ficht TA. Brucella melitensis VjbR and C12-HSL regulons: contributions of the N-dodecanoyl homoserine lactone signaling molecule and LuxR homologue VjbR to gene expression. BMC Microbiol. (2010) 10:167. doi: 10.1186/1471-2180-10-167
19. Robertson GT, Roop RJ. The Brucella abortus host factor I (HF-I) protein contributes to stress resistance during stationary phase and is a major determinant of virulence in mice. Mol Microbiol. (1999) 34:690–700. doi: 10.1046/j.1365-2958.1999.01629.x
20. Caswell CC, Gaines JM, Roop RN. The RNA chaperone Hfq independently coordinates expression of the VirB type IV secretion system and the LuxR-type regulator BabR in Brucella abortus 2308. J Bacteriol. (2012) 194:3–14. doi: 10.1128/JB.05623-11
Keywords: transcriptome analysis, stress response, Brucella, gene expression, virulence
Citation: Peng X, Liu Y, Qin Y, Jiang H, Feng Y, Sun J, Niu K, Gao Q, Dong H and Ding J (2020) Comparative Transcriptome Analysis of Artificially Induced Rough-Mutant Brucella Strain RM57 and Its Parent Strain Brucella melitensis M1981. Front. Vet. Sci. 6:459. doi: 10.3389/fvets.2019.00459
Received: 25 September 2019; Accepted: 28 November 2019;
Published: 10 January 2020.
Edited by:
Michael Kogut, United States Department of Agriculture, United StatesReviewed by:
Gary Splitter, University of Wisconsin-Madison, United StatesMd. Aminul Islam, Tohoku University, Japan
Copyright © 2020 Peng, Liu, Qin, Jiang, Feng, Sun, Niu, Gao, Dong and Ding. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Hao Dong, dHVuZ2hhb0AxNjMuY29t; Jiabo Ding, ZGluZ2ppYWJvQDEyNi5jb20=
†These authors share first authorship
 Xiaowei Peng1†
Xiaowei Peng1† Kai Niu
Kai Niu Hao Dong
Hao Dong Jiabo Ding
Jiabo Ding