- Key Laboratory of Animal Parasitology of Ministry of Agriculture, Shanghai Veterinary Research Institute, Chinese Academy of Agricultural Sciences, Shanghai, China
Thioredoxin peroxidases (TPxs) are ubiquitous cysteine-based peroxidases that reduce peroxides as part of antioxidant defenses and redox signaling and are essential for Babesia microti protection against adverse environment agents like reactive oxygen species (ROS) and reactive nitrogen species (RNS). To better systematically understand TPxs, we identified a novel 2-Cys peroxiredoxin-Q (BmTPx-Q) of B. microti. The full-length BmTPx-Q gene is 653 bp that consists of an intact open reading frame of 594 bp that encodes a 197-amino acid protein. The predicted protein has a molecular weight of 22.3 kDa and an isoelectric point of 9.18. Moreover, BmTPx-Q showed low identity at the amino acid level to other peroxiredoxins (Prxs) among the currently known subfamilies. The recombinant BmTPx-Q protein (rBmTPx-Q) was expressed in Escherichia coli and purified with beads. The native protein BmTPx-Q was detected using mouse anti-BmTPx-Q polyclonal serum with western blotting and indirect immunofluorescence assay (IFA). In addition, enzyme activity was observed using nicotinamide adenine dinucleotide phosphate (NADPH) as substrate and triggered the NADPH-dependent reduction of the Trx/TrxR system. It was also discovered that BmTPx-Q mainly exists as a monomer whether under its native or functional states. In addition, when incubated with Chloroquine diphosphate salt for 24 h in vitro, the expression of BmTPx-Q showed a marked downward trend with the increase of drug concentration. These results suggest that B. microti uses BmTPx-Q to reduce and detoxify hydrogen peroxides to survive and proliferate inside the host. Furthermore, BmTPx-Q showed the lowest identity with host enzymes and could be a potential drug target for the development of novel strategies to control B. microti infection.
Introduction
Babesiosis is a serious disease caused by infection with protozoan parasite Babesia microti that is transmitted to humans via the bite of an infected tick or a contaminated blood transfusion. There have been many reports of cases from Europe and the USA in recent years (1–3). Babesiosis has a huge impact on elderly, splenectomized or immunocompromised patients, leading to anemia, fatigue, and fever hematuria (4). Although babesiosis can be controlled by treatment with antiparasitic drugs, many drugs have safety issues (5). Therefore, identification of new drug targets is needed to develop novel therapeutic strategies and overcome setbacks, such as drug resistance.
It is well-known that Babesia has a complex life cycle, including both arthropod vectors and mammalian hosts, and that it replicates in the host's red blood cells. Since it is surrounded by oxygen-rich environments, the parasite is likely to counteract the toxic effects of reactive oxygen species (ROS) and reactive nitrogen species (RNS) that could induce oxidative DNA damage and lipid peroxidation (6, 7). The ROS and RNS are highly active compounds during normal cell metabolism. Therefore, to avoid the deleterious effects of ROS, various defense mechanisms have been adapted, such as non-enzymatic elements, which include glutathione (GSH) and vitamin C, and antioxidant enzymatic elements [i.e., catalase (CAT), superoxide dismutase (SOD), as well as peroxidase] (8). The parasite has developed a wide range of antioxidant systems, including peroxiredoxins (Prxs) (9), to keep their redox balance while living inside host erythrocytes (10, 11). Prxs have been a research topic of interest as a family of thiol-specific non-heme antioxidant peroxidases detoxifying hydrogen peroxide (H2O2), alkyl peroxides, and peroxynitrite (12). Moreover, Prxs are expressed at high levels in cells of almost all organisms, which protects cells against toxicity from ROS by reducing and detoxifying hydroperoxides, highlighting the importance of this protein family.
The Prx proteins have been divided into those that contain a single catalytic cysteine (Cys) residue and those that have an additional conserved residue (13). However, a new classification system has been suggested by Nelson and colleagues (14). Under this classification, Prxs are divided into six subfamilies based on the abundant sequence homology and structural similarity analyses, namely: Tpx, PrxQ-BCP, Prx1/AhpC, Prx5, Prx6, and AhpE (15–17). Alternatively, on the basis of the presence or relative locations of the resolving cysteine (Cr) residue, Prxs are classified into three types based on the distinct catalytic mechanisms: 1-Cys Prxs (Prx6 and AhpE), the typical 2-Cys Prx (Prx1/AhpC), and atypical 2-Cys (Tpx, TPx-Q, and Prx5) (18).
Prxs are collectively called thioredoxin peroxidases (TPxs) and constitute a large family of thiol-dependent peroxidases (18). Prxs are also potential drug targets. The studies demonstrated that these poorly cope with oxidative stress in Prx knockout strains of Plasmodium falciparum (19, 20). Recently, several TPxs of malaria parasites were identified, and the functional properties of the enzymes were considered key factors for the development of new drugs (21–24). Prx has also been reported to play key roles in innate immunity and inflammation (25) besides cellular redox signaling (26). Among these Prxs, the TPx-Q subfamily members have been proposed to be the most ancestral, most complex and the least systematically characterized (27). The TPx-Q proteins are thiol-based peroxidases, and are important for maintaining redox homeostasis in several organisms. TPx-Q is an atypical Prx that has already been identified in bacteria, parasites, and some lower eukaryotes but is not found in mammals (28–31).
TPx-Q, which is a cysteine-based peroxidase, has been detected in various protozoan parasites. TPx-Q-mediated resistance to major stresses largely relies on ROS degradation. TPx-Qs usually occurs at the monomeric state, with an intramolecular disulfide bond that are reduced by thioredoxins (Trx) (32). It has been shown that over-expression of TPx-Qs can cope with oxidative stresses in cyanobacteria. TPx-Q B from Mycobacterium tuberculosis is monomeric under reduced and oxidized states, and it is a thioredoxin-dependent and highly efficient fatty acid hydroperoxide reductase (33). Regardless of the mode of classification of these proteins, the catalytic mechanism of the enzyme remains central and relies on the redox-active cys, which is highly conserved in its amino acid sequence (34).
Although Prxs have been extensively studied, especially Tpx and Prx1 in Plasmodium, little is known about the Trx peroxidases-Q (TPx-Q) of B. microti. Sequencing of the full genome for B. microti has been completed (35), but the currently existing results are unilateral and unsystematic. In our previous study, we have done some basic research about the 2-Cys Trx peroxidases-1 and peroxidases-2 of B. microti (BmTPx-1 and BmTPx-2) (36, 37) and found evidence suggesting that the B. microti possesses at least two Prx subfamilies (Tpx and PrxQ). In this study, we identified and characterized a novel thioredoxin peroxidase (BmTPx-Q) from a strain of B. microti, analyzed the activity and assessed the BmTPx-Q expression after treatment with antiparasitic agents. All results suggest that B. microti can use BmTPx-Q to reduce and detoxify H2O2 for survival. Additionally, our new investigations on BmTPx-Q, a member of Prxs, provide new insights into the structure and function of Prx. We demonstrated that BmTPx-Q might act as an oxidative stress defensive molecule as well as drug target in B. microti.
Materials and Methods
Parasite Culture in vivo
The ATCC® PRA-99TM strain of B. microti was obtained from the American Type Culture Collection (ATCC, USA) and maintained by serial passage in BALB/C mice (SLAC Laboratory Animal Co., Ltd., China) using a method described previously (36). The parasites were isolated until the erythrocyte infection rate reached 30%−40%, which was confirmed with Giemsa-stained thin-blood smears.
Parasite Culture and Treatment in vitro
For one independent experiment, B. microti was obtained from at least two mice; 3 days after infection (according to our protocol). The blood of the infected mice was carefully collected under aseptic conditions and added to bacteria-free anticoagulant. Then the blood was passed through a 27G needle three to five times. The blood cell debris was removed with a 5 μm syringe filter, and most of the parasites were isolated. Subsequently, the parasites were centrifuged at 2,000 rpm for 5 min with a horizontal centrifuge. The supernatant was discarded and the pellets were re-suspended with sterile culture medium. The pellets used for incubation experiments were then washed three times with phosphate-buffered saline (PBS, pH 7.2) containing 50 μg/mL gentamycin sulfate (Sigma) under aseptic conditions. Healthy red blood cells were taken from 21-days-old normal mice and were distributed in a 12-well culture plate. Subsequently, each well was cultured in 5% CO2 at 37°C in RPMI 1640 (Gibco, USA) supplemented with 1% penicillin/streptomycin (Gibco, USA) and 40% fetal bovine serum (Gibco, USA). The incubations were performed for 12 and 24 h at 37°C in a 5% CO2 incubator.
Cloning and Sequence Analysis of B. microti TPx-Q (BmTPx-Q)
To find more novel genes from B. microti. Total RNA sequencing was performed to characterize all transcriptional activity. The brief method of preparing total RNA and cDNA for library construction and sequencing of the samples was described in our previous study (36). The full-length cDNA of the putative BmTPx-Q was generated by transcriptome analysis. The sequence was analyzed as described in our previous investigation (37). The Genetyx software (Software Development Co., Ltd., Tokyo, Japan) was used for the analysis of BmTPx-Q nucleotide and amino acid sequences. To further understand the relationships between BmTPx-Q and other TPx-Q genes, the inferred amino acid sequence of the BmTPx-Q was compared with other protein sequences (namely, the homologous TPx-Q proteins from different species) retrieved from the GenBank database using the National Center for Biotechnology Information (NCBI) Basic Local Alignment Search Tool (BLAST) (http://www.ncbi.nlm.nih.gov/blast/). The protein domain was identified using BLAST (http://www.ncbi.nlm.nih.gov/Structure/cdd/wrpsb.cgi). The signal peptide was predicted with the SignalP 4.1 server (http://www.cbs.dtu.dk/services/SignalP/).
Expression and Purification of Recombinant BmTPx-Q (rBmTPx-Q)
The recombinant proteins (rBmTPx-Q) were produced as described previously (36). Briefly, the open reading frame (ORF) of BmTPx-Q was amplified by PCR with the following primer pair (forward) 5′-TT CAT ATG TTC AAA ATA CTG AAT TCA CGG-3′ and (reverse) 5′-TT CTCGAG CAG TTT ATC AAT AAA TTC-3′ (the underlined sequences contain the NdeI and XhoI restriction sites). The PCR product was inserted into the expression vector pET-30a (Novagen, USA). The recombinant plasmids harboring the BmTPx-Q (BmTPx-Q/pET-30a) coding sequence were transformed into E. coli (strain BL21). Induction of rBmTPx-Q Histidine-tag expression was performed using 1 mM isopropyl thio-b-D-galactoside (IPTG), followed by purification using Ni-NTA agarose beads (Merck-Millipore Corporation, USA). Purified rBmTPx-Q was evaluated a sample in 12% sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS–PAGE) using reducing conditions and stained it with Coomassie Brilliant blue R-250. Protein concentration was determined with the BCA assay (Thermo Fisher Scientific, USA).
Antioxidant Activity Assay
To evaluate the antioxidant activity of rBmTPx-Q applying a mixed-function oxidation (MFO) assay (21, 29, 38). First, rBmTPx-Q (250, 500, and 1,000 μg/mL) was added to the reaction mixture, followed by incubation at 37°C for 1 h. Then, 500 ng of pBluescript SK(-) (Stratagene, USA) plasmid DNA was added to the reaction mixture, followed by incubation for an additional 2.5 h. Plasmid nicking was determined by using the MFO assay and analysis by 1% agarose gel electrophoresis, which was stained with DuRed dye (Fanbo Biochemicals Co. Ltd., China). Subsequently, to investigate whether rBmTPx-Q possesses peroxidase activity using a ferrithiocyanate system (39). The reaction mixtures (200 μL) containing rBmTPx-Q (1.5 μg protein) and 85 μL of buffer (0.5% glycerol/5 mM DTT/0.03 × PBS) was pre-incubated at room temperature (RT) for 2 min. H2O2 (1 mM) was used to start the reaction and was terminated with 40 μL of 26% trichloroacetic acid, which was added at 2-min intervals. The disappearance of H2O2 was monitored to assess rBmTPx-Q activity. In the reaction mixture, the remaining peroxide content was allowed to react with ~40 μL of 10 mM (NH4)2Fe(SO4)2 and 20 μL of 2.5 M KSCN, which formed a ferric thiocyanate complex that was red in color. The color intensity was measured at a wavelength of 475 nm using a microplate reader (SpectraMax M5; Molecular Devices, USA). In the presence of Trx and Trx reductase (TrxR), oxidation of nicotinamide adenine dinucleotide phosphate (NADPH) coupled with rBmTPx-Q to the reduction of H2O2 was examined using the method described by Kang et al. (40). NADPH oxidation was monitored in A340 in a 200-μL reaction mixture (0.14 μM TR, 6.4 μM Trx, 0.375 mM NADPH, 500 μg/mL rBmTPx-Q protein, 250 μM H2O2, 50 mM HEPES). The negative control (-)rBmTPx-Q indicates the absence of BmTPx-Q in reaction mixture.
Western Blotting for the Native BmTPx-Q Protein
To identify the native BmTPx-Q protein in the lysate of B. microti, B. microti-infected red blood cells (iRBCs) were prepared from Kunming mice at 4, 5, 6, 7, and 8 days post-infection, and non-infected erythrocytes were used as a negative control. First, the infected blood samples were centrifuged in low speed and hemolyzed by red cell lysis buffer (Tiangen Biotech, China) after discarding the supernatant. The soluble fractions (20 mg per lane) were separated on a 12% SDS-PAGE. The Western blot analysis was performed as described previously (36). Briefly, the proteins were electrophoretically transferred to polyvinylidene difluoride (PVDF) membranes (Merck-Millipore Corporation, USA), and was blocked with 5% skim milk diluted in PBS/0.05% Tween (PBST) for 2 h at 37°C. Then, the membranes were incubated overnight at 4°C in mouse anti-rBmTPx-Q serum diluted to 1:200 in PBST. The blotted membranes were washed using PBST, followed by incubation in the presence of goat anti-mouse IgG antibody (horseradish peroxidase-conjugated; dilution, 1:2,000; Bethyl Laboratories, Inc., USA) for 1 h at 37°C. A washing step took place with PBST before applying an enhanced DAB chromogenic substrate kit (Tiangen Biotech, China) to visualize the bands in accordance with the manufacturer's guidelines.
Expression Analysis of BmTPx-Q Post-infection
Assessment of BmTPx-Q expression post-infection was performed according to a method described elsewhere (41, 42). Briefly, we collected blood from mice infected with 1 × 108 iRBCs at 1–10 days post-injection, from which total RNAs were extracted using TRIzol. The cDNA was constructed by a PrimeScript RT reagent kit with gDNA eraser (TaKaRa, Japan) and used in quantitative real-time PCR (qRT-PCR) analysis. The qRT-PCR was performed with SYBR® Premix Ex Taq TM II (TaKaRa, Japan) and a StepOne Plus PCR system (Applied Biosystems). The primer pair of BmTPx-Q were BmTPxQ-qRT-F: ACAAGCACAATCTCCCATAC (forward) and BmTPxQ-qRT-R: TCTCCAGCACTAACTCCC (reverse). The 18s ribosomal RNA of B. microti (Bm18S) (GenBank: XM_021481625.1) was used as an internal control. The primer pair of Bm18S were Bm18S-qRT-F: GTTATAGTTTATTTGATGTTCGTTT (forward) and Bm18S-qRT-R: AAGCCATGCGATTCGCTAAT (reverse). The parasitemia was calculated at each day post-infection. Analysis was performed using the 2−ΔΔct method, and experimental values were expressed as relative amounts (43).
Indirect Immunofluorescent Antibody Test
To determine the intracellular location of BmTPx-Q in B. microti, the test was performed as described previously (36). Briefly, thin smears of iRBCs were prepared and fixed in 50% acetone-50% methanol solution for 10 min at −30°C. Mouse antiserum against rBmTPx-Q was used as the primary antibody (dilution, 1:200), incubated at 37°C for 45 min. After it was washed with PBS, Alexa-Fluor® 488-conjugated goat anti-mouse IgG (Life Technologies Corporation, USA) was added as a secondary antibody (dilution, 1:2,000) and incubated at 37°C for 45 min. Then, the slides were washed using PBS and incubated with 0.5 μM 4′,6′-diamidino-2-phenylindole (Molecular Probes, Inc., USA) at RT for 20 min. The slides were later washed and mounted with Cytomation fluorescent mounting medium (Dako Corporation, USA) and visualized under a confocal laser-scanning microscope (Zeiss LSM 880, Germany).
Transcript Changes of BmTPx-Q After Treatment With Antiparasitic Agents
To assess the mRNA relative expression profile of BmTPx-Q after treatment with antiparasitic agents, a short-term culture system of B. microti iRBCs was established in vitro. This study was performed as described in previous research (41). Briefly, B. microti-infected mice blood (~30% parasitemia) was collected and iRBCs were washed with PBS. A 12-well flat-bottom plate (Thermo, USA) was used for drug screening. The B. microti iRBCs (~2 × 107) were cultured in RPMI 1640 (Life Technologies, USA) supplemented with 25 mM HEPES (Life Technologies, USA) and 40% fetal bovine serum (Life Technologies, USA) at 37°C in a 5% CO2 (44, 45). To evaluate the effect of drugs on BmTPx-Q gene expression, the iRBCs were grown in vitro for a short period before exposure to the three antiparasitic drugs (Quinine monohydrochloride dehydrate, Dihydroartemisinin, and Chloroquine diphosphate salt) (Sigma Aldrich, USA) was assessed. Quinine and Dihydroartemisinin were dissolved into dimethylsulfoxide (DMSO), while Chloroquine was dissolved in PBS. iRBCs were treated with various concentrations (20, 50, or 100 μM) of Quinine, Dihydroartemisinin, and Chloroquine at different timepoints (12, 24, and 36 h), meanwhile the controls were treated with DMSO or PBS. Relative BmTPx-Q transcript levels were assessed as previously described, by Reverse Transcription (RT)-PCR. Briefly, total RNA was isolated from infected RBCs using TRI solution (Life Technologies Corporation, USA), and RT-PCR was carried out applying 1 μg of total RNA and specific primers by a PrimeScript™ One-Step RT-PCR Kit (Takara, China). Conditions for the PCRs were as follows: 95°C for 30 s; 95°C for 5 s, 60°C for 35 s, for 40 cycles. The experiment was repeated in triplicate.
Statistical Analysis
A GraphPad PRISM 5 software (GraphPad Software Inc., CA, USA) was used for the data analysis. The mean ± standard deviation (SD) of each group was calculated. The differences between groups were assessed using two-tailed t-tests. P < 0.05 was considered significant and P < 0.01 was considered highly significant.
Results
Identification and Characterization of the BmTPx-Q Gene
The identified full-length cDNA of BmTPx-Q has 653 bp in which includes a single ORF of 594 bp. Sequence analysis indicated that the ORF of BmTPx-Q gene encodes a protein of 197 amino acids with a theoretical molecular weight and isoelectric point of 22.3 kDa and 9.18, respectively. The amino acid sequence was deduced from the cDNA sequence of BmTPx-Q (Figure 1A). SignalP server indicated that BmTPx-Q has a signal peptide. The BmTPx-Q protein sequence was 98% identical to that of B. microti Prx Q (RI strain, XP_012647890.1), showing 35% sequence similarity with Prx Q of Babesia bigemina (XP_012767998.1), 35% with Prx Q of Blastomyces dermatitidis (EEQ83458.1), 39% with Prx Q of Babesia sp. Xinjiang (XP_028870106.1), and 43% with Ahp/TSA family-related protein, putative of Theileria annulata (XP_954500.1) (Figure 1B). The sequence alignment showed that the conserved peroxidatic cysteine (Cp) of BmTPx-Q is located at position 95 in a PxxxTxxC-motif, and an additional conserved cysteine at position 100 in BmTPx-Q (Figure 1B). The BmTPx-Q sequence contains a Thioredoxin-dependent hydroperoxide peroxidase activity of bacterioferritin comigratory protein (PRX-BCP) domain, the PRX-BCP is a new member of the thiol-specific antioxidant protein (TSA)/Alkyl hydroperoxide peroxidase C (AhpC) family (Figure 1C). The BmTPx-Q belonging to the Thioredoxin-like superfamily, it also contains a catalytic triad on conserved domain PRX_BCP, 3 of 3 of the residues that compose this conserved feature (Figure 1C).
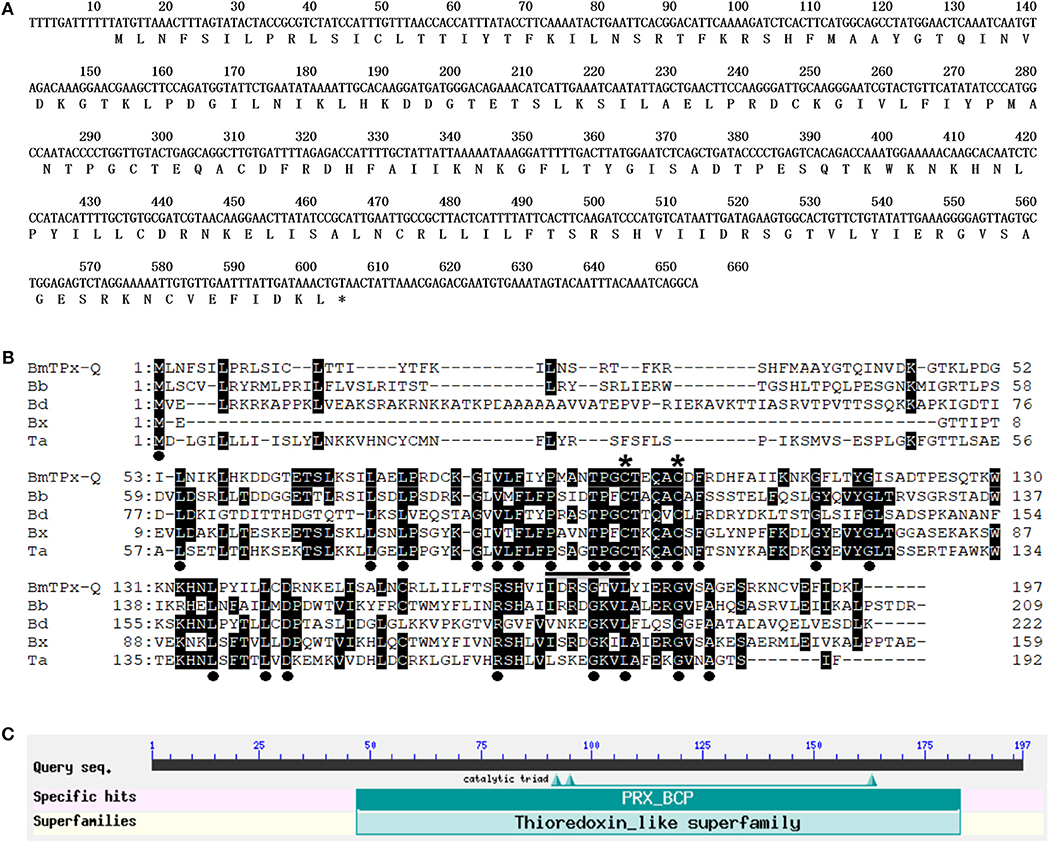
Figure 1. Analysis of the sequence and primary structure of BmTPx-Q. (A) Nucleotide and deduced amino acid sequences of BmTPx-Q. (B) BmTPx-Q multiple sequence alignment analysis. B. microti (BmTPx-Q); Babesia bigemina (Bb: XP_012767998.1); Blastomyces dermatitidis (Bd: EEQ83458.1); Babesia sp. Xinjiang (Bx: XP_028870106.1); Theileria annulata (Ta: XP_954500.1). The dot indicates the identical amino acids in all sequences, an asterisk indicates the conserved cysteine residue, and the conserved PxxxTxxC-motif around the active site is underlined. (C) Analysis of the active domains in the BmTPx-Q amino acid sequence as identified by NCBI blast.
Expression and Purification of the Recombinant BmTPx-Q
The PCR product was cloned into the pET30a vector and the recombinant protein was successfully expressed in E. coli BL21 (DE3) as a his-tagged protein. The rBmTPx-Q protein was purified with Ni-NTA agarose beads and was analyzed by SDS/PAGE. As shown in Figure 2, the purified protein showed a single band (~21 kDa) by 12% SDS/PAGE in the presence of DTT (reducing).
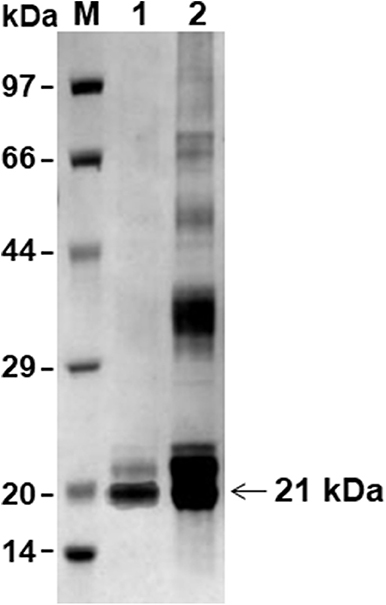
Figure 2. SDS-PAGE analysis of the rBmTPx-Q-His fusion protein. Analysis of expression and purification of rBmTPx-Q by reducing SDS-PAGE (12%). Lane M: standard protein molecular weight marker; Lane 1: purified rBmTPx-Q; Lane 2: induced cell lysates. Purified rBmTPx-Q-His was ~21 kDa and was used in subsequent antibody production and antioxidant activity assays.
Antioxidant Activity of rBmTPx-Q
The activity was evaluated using the MFO assay (Figure 3A). In this assay, FeCl3 and DTT generate hydroxyl radicals in the reaction mixture induced nicks in the supercoiled plasmid DNA, thereby altering the mobility of DNA during agarose electrophoresis. Both FeCl3 and DTT generated nicks in the DNA in the absence of rBmTPx-Q. Thus, there was an apparent increase in DNA size (Figure 3A, lane 4). However, nicks in the plasmid DNA were detected after purified rBmTPx-Q was added to the reaction mixtures at 1,000, 500, and 250 μg/mL concentrations (Figure 3A, lanes 5–7). The results indicate rBmTPx-Q has antioxidant activity. TPx-Q proteins are thiol-based peroxidases that catalyze the reduction of H2O2. The ability of the rBmTPx-Q to remove H2O2 was evaluated utilizing a ferrithiocyanate system. The rBmTPx-Q was examined by monitoring oxidation of NADPH in the E. coli Trx/TrxR system in the presence of DTT. As demonstrated in Figure 3B, rBmTPx-Q showed peroxidase activity in the presence of the Trx system (Trx, TR, and NADPH) with a concentration-dependent manner. These results indicate that the atypical 2-Cys TPx-Q peroxidases use thioredoxin as a reductant. The results showed a decrease at A340 nm in the presence of rBmTPx-Q, possibly due to the oxidation of NADPH after the addition of H2O2. Within 10 min, 1.5 μg of rBmTPx-Q destroyed nearly about half of 1 mM H2O2. This result suggested that rBmTPx-Q can be described as a thioredoxin peroxidase.
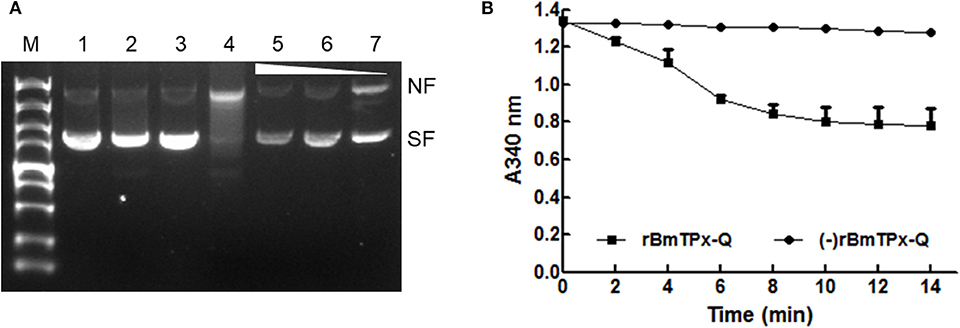
Figure 3. Antioxidant activity of the BmTPx-Q protein. (A) The pBluescript plasmid DNA was incubated with reaction mixture including the rBmTPx-Q, and nicking of the supercoiled plasmids by MFO was evaluated on 1% agarose gels stained with ethidium bromide. M, DNA marker; 1, DNA + H2O; 2, DNA + DTT; 3, 180 ng DNA + FeCl3; 4, DNA + MFO; 5, 1,000 μg/mL rBmTPx-Q + MFO + DNA; 6, 500 μg/mL rBmTPx-Q + MFO + DNA; and 7, 250 μg/mL rBmTPx-Q + MFO + DNA. The nicked form (NF) and supercoiled form (SF) of the plasmids are indicated on the right, and the triangle shows the increasing concentration of the recombinant proteins. (B) Peroxidase activity of rBmTPx-Q. NADPH oxidation coupled by rBmTPx-Q (solid line) to reduction of H2O2 in the presence of the E. coli Trx/Trx R system. NADPH oxidation was monitored as the decrease in A 340 in a 200-μL reaction mixture (0.14 μM TR, 6.4 μM Trx, 0.375 mM NADPH, 500 μg/mL rBmTPx-Q, 250 μM H2O2, 50 mM HEPES). The dotted line indicates the absence of rBmTPx-Q in reaction mixture. Data are presented as the means of three independent experiments.
Expression Analysis of BmTPx-Q in B. microti Post-infection
To identify the native BmTPx-Q protein in B. microti, western blot analysis was performed using the mouse anti-rBmTPx-Q serum. The results showed that an ~22 kDa band was detected in the iRBC lysates 7 and 8 days post-infection (Figure 4A, lines 3 and 4), which revealed the predicted monomeric size, whereas no specific bands were detected in the non-infected control (Figure 4A, line 6). The size of the recombinant BmTPx-Q band was similar to that of the native BmTPx-Q. The results suggested that BmTPx-Q protein has high immunogenicity and can induce the host's immune system and also BmTPx-Q exists primarily in a monomeric form without the formation of intermolecular disulfide bonds. Moreover, total RNA of iRBCs from different days post-infection was used to investigate the expression profile of BmTPx-Q by qRT-PCR analysis. BmTPx-Q expression peaked twice on 4 and 8 days post-infection and declined suddenly on 5 and 9 days post-infection (Figure 4B). Blood smears were stained with Giemsa to assess the infection by calculating the ratio of iRBCs. B. microti was detected in iRBCs on the first day post-infection. The parasitemia ratio increased until 5 days post-infection, and started to decline on 6 days post-infection until reaching undetectable levels by day 10 post-infection (Figure S1).
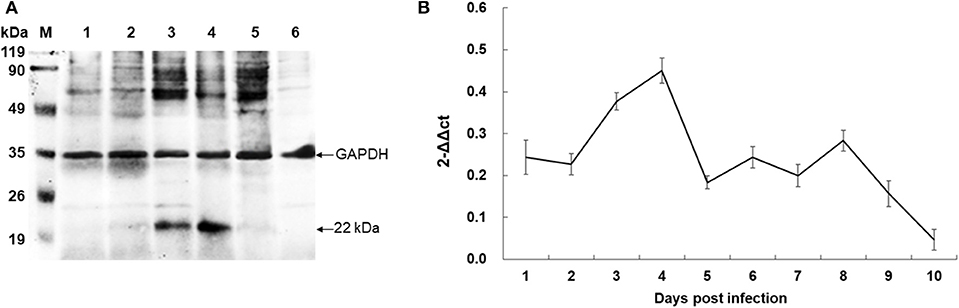
Figure 4. Expression analysis of BmTPx-Q in B. microti post-infection. (A) Western blot analysis of the native BmTPx-Q. Lane M: standard protein molecular weight marker; Lanes 1–5: B. microti infected mouse erythrocyte lysates on 5th, 6th, 7th, 8th, and 9th day post-infection; Lane 6: uninfected mouse erythrocyte lysate; mouse anti-rBmTPx-Q serum was used as primary antibody in this Western blot analysis and revealed a band of ~22 kDa. (B) Relative expression analysis of BmTPx-Q at different days post-infection. The x-axis refers to the day of infection of mouse RBCs with B. microti; the y-axis refers to the relative BmTPx-Q expression level in mouse RBCs on different days post-infection by using the 2−ΔΔCT mean.
Immunofluorescence Assays
A thin blood smear of B. microti-infected RBCs (~20% parasitemia) was used by indirect immunofluorescence assay (IFA) with the mouse anti-rBmTPx-Q serum. The blue fluorescence indicates the nucleus of B. microti, whereas the green fluorescence shows BmTPx-Q located within the nucleus of B. microti merozoites in iRBCs (Figure 5a). Additionally, co-localization of anti-BmTPx-Q signal (green) with DAPI (blue) indicates that most of BmTPx-Q is present in the cytoplasm of the parasites (Figure 5d, Merged). In the control sample, no green fluorescence was detected in the iRBCs which were incubated with serum collected from uninfected mice (Figure 5e). Therefore, our results show that BmTPx-Q is expressed in B. microti merozoite cytoplasm.
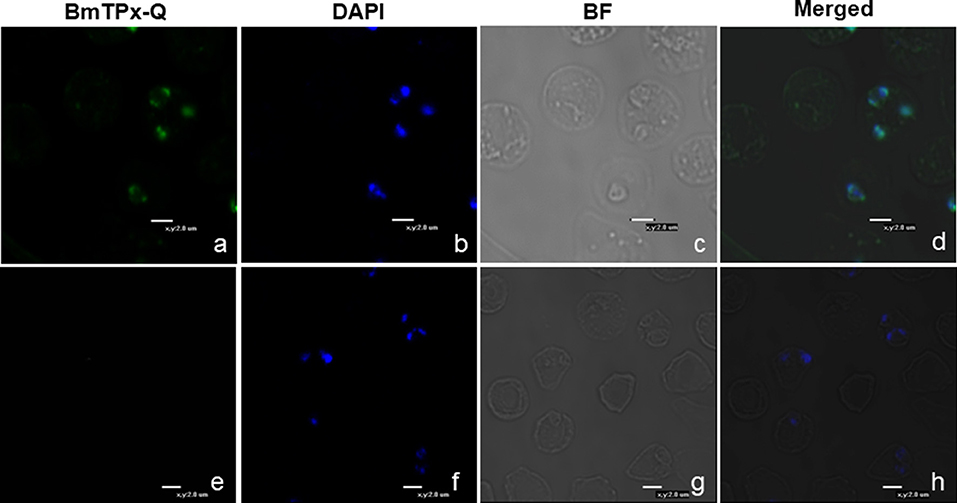
Figure 5. Immunofluorescence microscopy analysis of the cellular localization of TPx-Q in B. microti-infected RBCs. Antiserum against BmTPx-Q (a–d) or normal mouse serum (e–h) was incubated with acetone-fixed B. microti-infected mice erythrocytes and stained with Alexa Fluor 488-conjugated goat anti-mouse IgG antibody. (a,e) Incubation with anti-rBmTPx-Q or normal mouse serum staining; (b,f) DAPI staining; (c,g) brightfield; and (d,h) merged images. Scale bar indicates 2.0 μm.
Transcript Analysis of BmTPx-Q After Treatment With Antiparasitic Agents
The RT-PCR was conducted using various cDNA templates isolated samples treated with different doses of Quinine monohydrochloride dehydrate, Dihydroartemisinin, or Chloroquine at 12, 24, and 36 h. As shown in Figures 6A,B, BmTPx-Q was expressed in three different time points, although there was no significant difference, but a decreasing trend was observed as time went by in the three different concentrations under Quinine and Dihydroartemisninin compared to the control group (we have subtracted the control group data when analyzing). In addition, BmTPx-Q was expressed at the highest level in 20 and 50 μM at 24 h under Chloroquine diphosphate salt (Figure 6C). High BmTPx-Q expression in the parasites was due to adverse environmental factors, which increases gradually to peak at 24 h before decreasing sharply in the subsequent time points. The growth status of B. microti after that was severely inhibited at 36 h, which was also confirmed through Giemsa-stained thin-blood films (Figure S2). That can explain the low expression level of BmTPx-Q in the subsequent stages, also indicating that B. microti might be inhibited by these three drugs. These findings suggest that BmTPx-Q might be implicated by different mechanisms in the response of B. microti to Quinine, Dihydroartemisninin, and Chloroquine. Although Prxs play a role in protection against oxidative damage in parasites and ensure a certain degree of defense, the effects of external factors can be irreversible.
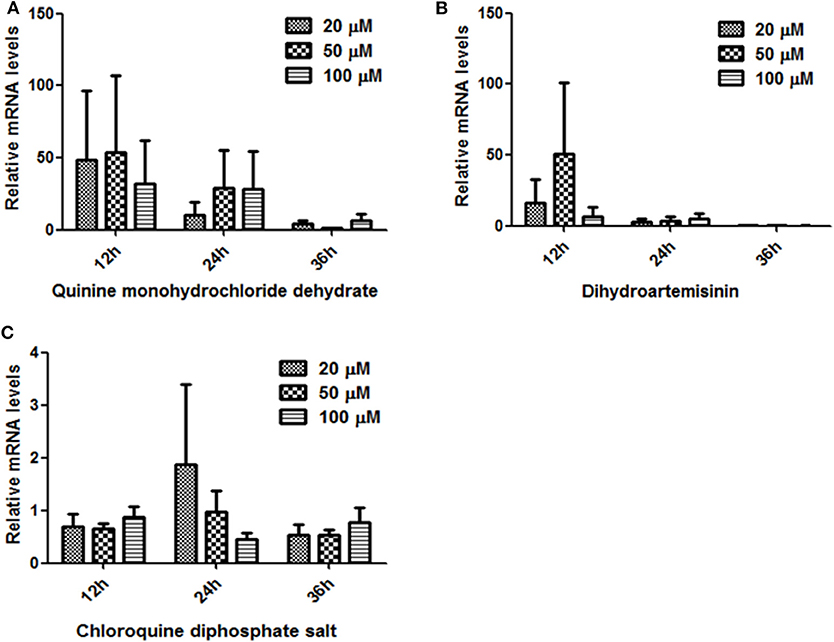
Figure 6. Relative expression analysis of BmTrx-Q in iRBCs exposed to anti-parasitic agents for 12, 24, and 36 h. The iRBCs were treated with different concentrations (20, 50, 100 μM) of Quinine (A), Dihydroartemisinin (B), and Chloroquine (C).
Discussion
Prxs reduce peroxides using a peroxidatic cysteine residue, and the Cys are essential for enzyme activity. In atypical 2-Cys Prxs, which are also named Type II Prxs, the position of one of two cysteine residues is not conserved (8, 46, 47). The typical 2-Cys Prxs have physiological functions as peroxidase, 1-Cys Prx and atypical 2-Cys act as monomers, whereas typical 2-Cys Prxs act as dimers (48, 49).
To better systematically understand TPxs, we previously have identified the Prxs (BmTPx-1 and BmTPx-2) from B. microti (36, 37). Herein, we extended these efforts to select additional members of this group. A BLAST search revealed that the BmTPx-Q has low sequence similarities with those of Prxs of host species. This may indicate that this protein may have a good prospect in terms of drug targets. As expected, like all 2-Cys-containing Prxs, BmTPx-Q possesses the PxxxTxxC-motif and beside two conserved cysteine residues Cp and Cr that are essential for peroxidase activity. The Cp is located at the N-terminal end of the protein and Cr is usually four residues or 30 amino acids away from Cp (50). In our study, BmTPx-Q as a 2-Cys Prx, Cys95 being Cp, which is essential for peroxidase activity, whereas Cys100 acts as Cr. For 2-Cys Prxs, the Cp is oxidated by a peroxide substrate to generate a Cys sulfenic acid (Cys-SOH) intermediate, which reacts with the Cr to form an monomer or dimmer (18, 51). The disulfide bond is reduced to reform the reduced Cp by an external reducing substrate of Trx/TrxR system with an atypical 2-Cys catalytic mechanism (32). Afterwards, the generated TPx-Q is ready for another catalytic cycle. Su et al. (30) indicated that the two cysteine residues are essential for enzymatic activity by mutation analysis. Furthermore, they proved that C. glutamicum TPx-Q catalytically eliminates peroxides by exclusively receiving electrons from Trx/TrxR system (30). Unlike the typical 2-Cys Prx proteins, TPx-Q may exist as monomers, dimers, or a mixture (52). In this study, BmTPx-Q may exist mainly in monomeric form with an intramolecular disulfide bond. Moreover, it has been documented that the three forms of TPx-Q contain a wealth of information useful to the field (27).
To assess the enzymatic activity and generate specific antibodies against BmTPx-Q, the recombinant proteins were expressed as a his-tagged protein in E. coli purified by agarose beads (Figure 2). It is now known that Prxs are prevalent in prokaryotic and eukaryotic organisms and are essential against various oxidative stresses through catalyzing the reduction of H2O2 and organic hydroperoxides to keep the cellular redox balance (53). In addition, Prxs are more than just simple peroxide-eliminating enzymes. They are localized to various subcellular compartments and function as regulators of local H2O2 levels. The studies of transcription revealed that the Prx Q has a function in oxidant defense (54). Therefore, the antioxidant activities of rBmTPx-Q were evaluated by the MFO assay (Figure 3A). In MFO assay, FeCl3 and DTT generate hydroxyl radicals that produced nicks in the supercoiled DNA in the reaction mixture, which could be monitored by changing the running behavior of the DNA in electrophoresis (38). Therefore, the presence of rBmTPx-Q in the reaction mixtures with 250, 500, and 1,000 μg/mL concentration prevented the damage of DNA (Figure 3A), suggesting the antioxidant activities of rBmTPx-Q. Data shown in Figure 3B demonstrated that the peroxidase activity of rBmTPx-Q acts as an antioxidant enzyme, the results were consistent that Prx as a peroxidase to reduce H2O2 via the parasitic Trx system in previous study (55).
Although Prxs are primarily as peroxidases, these also function as chaperones or phospholipases and are involved in redox signaling (48, 56). The chaperone or the phospholipase activity of Prxs is independent with the catalytic cysteine residues. Molecular chaperones can assist the covalent folding of proteins and prevent the protein aggregation (57). Su et al. (30) found that TPx-Q functions as molecular chaperone and peroxidase. Cho et al. (58) first reported that Prx Q of Deinococcus radiodurans R1 is a monomeric atypical 2-Cys Prx and has dual activity as peroxidase and a molecular chaperone. However, the molecular chaperones function of BmTPx-Q needs to be further studied.
Meanwhile, we collected the B. microti infected mouse erythrocyte lysates post-infection and detected the native BmTPx-Q protein in the lysate of B. microti using the mouse anti-rBmTPx-Q serum. Western blotting showed an ~22 kDa band in the iRBC lysates 7 and 8 days post-infection (Figure 4A, lines 3 and 4). This indicates that the native BmTPx-Q was expressed in large quantities that could be detected on the 7th and 8th day after infection. Moreover, the expression of BmTPx-Q peaked at both 4 and 8 days post-infection by qRT-PCR analysis (Figure 4B). Based on the growth curve of parasitemia, B. microti was undergoing a rapid propagation process in RBCs 4 days post-infection. During the erythrocytic stages, B. microti are exposed to oxygen-rich environments and must secrete large amounts of antioxidants, such as Trx or Prx to act against oxidative stress and protect the parasites (23). Furthermore, the native B. microti Prx protects DNA from oxidative damage (23, 59). From a rational point of view, high expression levels of BmTPx-Q at 4 days post-infection helped B. microti to cope with the oxidative stress during the rapid propagation process. The level of BmTPx-Q expression reached a peak, whereas B. microti parasitemia decreased to low levels at 8 days post-infection, similar results were also observed for B. microti other antioxidant enzymes, such as peroxiredoxin 2, thioredoxin 2, and thioredoxin 3 in previous studies (37, 41, 42). The molecular immune mechanisms of high expression levels of these antioxidant molecules need to be further investigated.
In addition, previous research has reported that Prxs could induce protective immunity against Leishmania major infections and microfilaria Brugiamalayi infections in mice and Fasciola hepatica infections in goats (60). Since Prxs have potential as candidate vaccines for parasite species, the enzymatic activity may be related to the protective efficacy of these antigens. Our laboratory will evaluate the potential of this vaccine candidate in future studies.
IFA remains the most widely used method to localize proteins in organisms' intact cells. Figure 5 shows the expression patterns of BmTPx-Q in the red blood cells. The green fluorescence surrounding merozoite nuclei indicates enzyme expression in the cytoplasm of the parasite (Figure 5a). Based on a previous report, 1-Cys-Prx is also upregulated in the cytoplasm at the trophozoite stage, which is the metabolically active phase of P. falciparum (61). Unlike thioredoxin peroxidase-1 (TPx-1) of B. microti, BmTPx-Q exhibits a dot-like pattern expression in the parasite. Further investigation into whether BmTPx-Q and BmTPx-1 or BmTPx-2 co-exist in the peroxidase active organelles is required.
There was no significant difference regarding the mRNA relative expression profile of BmTPx-Q in the groups treated with the antiparasitic agents, compared to the control group, but a decreasing trends were observed against time (Figure 6). Three drugs managed to inhibit B. microti growth, but there was no marked effect on BmTPx-Q expression or activity. It is necessary to find specific inhibitors or drugs able to target BmTPx-Q, which will further confirm the importance of this molecule for B. microti survival. Moreover, it is necessary to confirm the TPx-Q whether as an oxidative stress defensive molecule as well as drug target in other strain or species of Babesia.
Conclusions
We characterized a BmTPx-Q in B. microti, and our results suggest that TPx-Q acts as a thioredoxin-dependent monomeric peroxidase that contributes to the resistance against oxidative in B. microti. BmTPx-Q is a monomeric form with two Cys that form an intramolecular disulfide bond. Since BmTPx-Q has an antioxidant activity, apparently it has a crucial role in the reduction of ROS. Concerning other aspects, Babesia parasites have other antioxidant proteins, such as SOD and Gpx. Thus, it is necessary to study the correlations between BmTPx-Q and these antioxidant proteins. Our data on BmTPx-Q can be used to investigate the precise role and biological functions of BmTPx-Q in the parasite. These findings may be relevant to the field of parasitological research and lead to the development of an anti-Babesiosis drug.
Data Availability Statement
All datasets generated for this study are included in the article/Supplementary Material.
Ethics Statement
The Institutional Animal Care and Use Committee of the Shanghai Veterinary Research Institute (IACUC Approval Number SHVRI-mo-2017070806) approved and the Animal Ethical Committee of Shanghai Veterinary Research Institute authorized this investigation.
Author Contributions
HZ and JZ conceived and designed this study. HZ and ZW performed the experiments. JH conducted the molecular analysis. JC performed the serology. YZ conducted the statistical analysis. All authors have read and approved the final version of this manuscript.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors thank the National Natural Science Foundation of China (Grant No. 31801974) for providing financial support.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fvets.2020.00076/full#supplementary-material
Figure S1. Giemsa-stained thin-blood film of the parasitemia. D0: Uninfected mouse RBCs; D1–D10: Days post-infection with B. microti of mouse RBCs. Scale bar indicates 5.0 μm.
Figure S2. The growth status of B. microti after treatment with antiparasitic agents at 36 h. iRBCs were treated with various concentrations (20, 50, or 100 μM) of Quinine, Dihydroartemisinin, and Chloroquine at 36 h, the controls were treated with DMSO. Thin-blood film was stained with Giemsa. Scale bar indicates 10.0 μm.
References
1. Herwaldt BL, Caccio S, Gherlinzoni F, Aspöck H, Slemenda SB, Piccaluga P, et al. Molecular characterization of a non-Babesia divergens organism causing zoonotic babesiosis in Europe. Emerg Infect Dis. (2003) 9:942–8. doi: 10.3201/eid0908.020748
2. Obiegala A, Pfeffer M, Pfister K, Karnath C, Silaghi C. Molecular examinations of Babesia microti in rodents and rodent-attached ticks from urban and sylvatic habitats in Germany. Ticks Tick Borne Dis. (2015) 6:445–9. doi: 10.1016/j.ttbdis.2015.03.005
3. Vannier E, Krause PJ. Human babesiosis. N Engl J Med. (2012) 366:2397–407. doi: 10.1056/NEJMra1202018
4. Homer MJ, Aguilar-Delfin I, Telford SR III, Krause PJ, Persing DH. Babesiosis. Clin Microbiol Rev. (2000) 13:451–69. doi: 10.1128/CMR.13.3.451
5. Vial HJ, Gorenflot A. Chemotherapy against babesiosis. Vet Parasitol. (2006) 138:147–60. doi: 10.1016/j.vetpar.2006.01.048
6. Gray J, Zintl A, Hildebrandt A, Hunfeld KP, Weiss L. Zoonotic babesiosis: overview of the disease and novel aspects of pathogen identity. Ticks Tick Borne Dis. (2010) 1:3–10. doi: 10.1016/j.ttbdis.2009.11.003
7. Robinson MW, Hutchinson AT, Dalton JP, Donnelly S. Peroxiredoxin: acentral player in immune modulation. Parasite Immunol. (2010) 32:305–13. doi: 10.1111/j.1365-3024.2010.01201.x
8. Reuter S, Gupta SC, Chaturvedi MM, Aggarwal BB. Oxidative stress, inflammation, and cancer: how are they linked? Free Radic Biol Med. (2010) 49:1603–16. doi: 10.1016/j.freeradbiomed.2010.09.006
9. Kawazu S, Komaki-Yasuda K, Oku H, Kano S. Peroxiredoxins in malaria parasites: parasitologic aspects. Parasitol Int. (2008) 57:1–7. doi: 10.1016/j.parint.2007.08.001
10. Bosch SS, Kronenberger T, Meissner KA, Zimbres FM, Stegehake D, Izui NM, et al. Oxidative stress control by apicomplexan parasites. Biomed Res Int. (2015) 2015:351289. doi: 10.1155/2015/351289
11. Poole LB, Hall A, Nelson KJ. Overview of peroxiredoxins in oxidant defense and redox regulation. Curr Protoc Toxicol. (2011) Chapter 7:Unit7.9. doi: 10.1002/0471140856.tx0709s49
12. Fujii J, Ikeda Y. Advances in our understanding of peroxiredoxin, a multifunctional, mammalian redox protein. Redox Rep. (2002) 7:123–30. doi: 10.1179/135100002125000352
13. Nickel C, Trujillo M, Rahlfs S, Deponte M, Radi R, Becker K. Plasmodium falciparum 2-Cys peroxiredoxin reacts with plasmoredoxin and peroxynitrite. Biol Chem. (2005) 386:1129–36. doi: 10.1515/BC.2005.129
14. Nelson KJ, Knutson ST, Soito L, Klomsiri C, Poole LB, Fetrow JS. Analysis of the peroxiredoxin family: using active-site structure and sequence information for global classification and residue analysis. Proteins. (2011) 79:947–64. doi: 10.1002/prot.22936
15. Chae HZ, Robison K, Poole LB, Church G, Storz G, Rhee SG. Cloning and sequencing of thiol specific antioxidant from mammalian brain: alkyl hydroperoxide reductase and thiol-specific antioxidant define a large family of antioxidant enzymes. Proc Natl Acad Sci USA. (1994) 91:7017–21. doi: 10.1073/pnas.91.15.7017
16. Karplus PA. A primer on peroxiredoxin biochemistry. Free Radic Biol Med. (2015) 80:183–90. doi: 10.1016/j.freeradbiomed.2014.10.009
17. Perkins A, Nelson KJ, Parsonage D, Poole LB, Karplus PA. Peroxiredoxins: guardians against oxidative stress and modulators of peroxide signaling. Trends Biochem Sci. (2015) 40:435–45. doi: 10.1016/j.tibs.2015.05.001
18. Hall A, Karplus PA, Poole LB. Typical 2-Cys peroxiredoxins-structures, mechanisms and functions. FEBS J. (2009) 276:2469–77. doi: 10.1111/j.1742-4658.2009.06985.x
19. Krnajski Z, Gilberger TW, Walter RD, Cowman AF, Müller S. Thioredoxin reductase is essential for the survival of Plasmodium falciparum erythrocytic stages. J Biol Chem. (2002) 277:25970–5. doi: 10.1074/jbc.M203539200
20. Sarma GN, Nickel C, Rahlfs S, Fischer M, Becker K, Karplus PA. Crystal structure of a novel Plasmodium falciparum 1-Cys peroxiredoxin. J Mol Biol. (2005) 346:1021–34. doi: 10.1016/j.jmb.2004.12.022
21. Hakimi H, Asada M, Angeles JM, Inoue N, Kawazu S. Cloning and characterization of Plasmodium vivax thioredoxin peroxidase-1. Parasitol Res. (2012) 111:525–9. doi: 10.1007/s00436-012-2864-3
22. Hakimi H, Suganuma K, Usui M, Masuda-Suganuma H, Angeles JM, Asada M, et al. Plasmodium knowlesi thioredoxin peroxidase 1 binds to nucleic acids and has RNA chaperone activity. Parasitol Res. (2014) 113:3957–62. doi: 10.1007/s00436-014-4060-0
23. Hakimi H, Goto Y, Suganuma K, Angeles JM, Kawai S, Inoue N, et al. Development of monoclonal antibodies against Plasmodium falciparum thioredoxin peroxidase 1 and its possible application for malaria diagnosis. Exp Parasitol. (2015) 154:62–6. doi: 10.1016/j.exppara.2015.04.018
24. Usui M, Masuda-Suganuma H, Fukumoto S, Angeles JM, Hakimi H, Inoue N, et al. Effect of thioredoxin peroxidase-1 gene disruption on the liver stages of the rodent malaria parasite Plasmodium berghei. Parasitol Int. (2015) 64:290–4. doi: 10.1016/j.parint.2014.09.013
25. Knoops B, Argyropoulou V, Becker S, Ferté L, Kuznetsova O. Multiple roles of peroxiredoxins in inflammation. Mol Cells. (2016) 39:60–4. doi: 10.14348/molcells.2016.2341
26. Hampton MB, O'Connor KM. Peroxiredoxins and the regulation of cell death. Mol Cells. (2016) 39:72–6. doi: 10.14348/molcells.2016.2351
27. Perkins A, Gretes MC, Nelson KJ, Poole LB, Karplus PA. Mapping the active site helix-to-strand conversion of CxxxxC peroxiredoxin Q enzymes. Biochemistry. (2012) 51:7638–50. doi: 10.1021/bi301017s
28. Buchko GW, Perkins A, Parsonage D, Poole LB, Karplus PA. Backbone chemical shift assignments for Xanthomonas campestris peroxiredoxin Q in the reduced and oxidized states: a dramatic change in backbone dynamics. Biomol NMR Assign. (2016) 10:57–61. doi: 10.1007/s12104-015-9637-8
29. Masatani T, Asada M, Ichikawa-Seki M, Usui M, Terkawi MA, Hayashi K, et al. Cloning and characterization of a 2-Cys peroxiredoxin from Babesia gibsoni. J Vet Med Sci. (2014) 76:139–43. doi: 10.1292/jvms.13-0274
30. Su T, Si M, Zhao Y, Liu Y, Yao S, Che C, et al. A thioredoxin-dependent peroxiredoxin Q from Corynebacterium glutamicum plays an important role in defense against oxidative stress. PLoS ONE. (2018) 13:e0192674. doi: 10.1371/journal.pone.0192674
31. Tailor V, Ballal A. Novel molecular insights into the function and the antioxidative stress response of a Peroxiredoxin Q protein from cyanobacteria. Free Radic Biol Med. (2017) 106:278–87. doi: 10.1016/j.freeradbiomed.2017.01.031
32. Kong W, Shiota S, Shi Y, Nakayama H, Nakayama K. A novel peroxiredoxin of the plant Sedum lineare is a homologue of Escherichia coli bacterioferritin co-migratory protein (Bcp). Biochem J. (2000) 351:107–14. doi: 10.1042/bj3510107
33. Reyes AM, Vazquez DS, Zeida A, Hugo M, Piñeyro MD, De Armas MI, et al. TPx-Q B from Mycobacterium tuberculosis is a monomeric, thioredoxin-dependent and highly efficient fatty acid hydroperoxide reductase. Free Radic Biol Med. (2016) 101:249–60. doi: 10.1016/j.freeradbiomed.2016.10.005
34. Rhee SG, Kil IS. Multiple functions and regulation of mammalian peroxiredoxins. Annu Rev Biochem. (2017) 86:749–75. doi: 10.1146/annurev-biochem-060815-014431
35. Cornillot E, Kaddour KH, Dassouli A, Noel B, Ranwez V, Vacherie B. Sequencing of the smallest Apicomplexan genome from the human pathogen Babesia microti. Nucleic Acids Res. (2012) 40:9102–14. doi: 10.1093/nar/gks700
36. Zhang H, Wang Z, Gong H, Cao J, Zhou Y, Zhou J. Identification and functional study of a novel 2-cys peroxiredoxin (BmTPx-1) of Babesia microti. Exp Parasitol. (2016) 170:21–7. doi: 10.1016/j.exppara.2016.08.005
37. Hai X, Zhang H, Wang Z, Gong H, Cao J, Zhou Y, et al. Identification of 2-Cys peroxiredoxin (BmTPx-2) as antioxidant active molecule from Babesia microti. Front Microbiol. (2017) 8:1959. doi: 10.3389/fmicb.2017.01959
38. Sauri H, Butterfield L, Kim A, Shau H. Antioxidant function of recombinant natural killer enhancing factor. Biochem Biophys Res Commun. (1995) 208:964–9. doi: 10.1006/bbrc.1995.1428
39. Thurman RG, Ley HG, Scholz R. Hydrogen peroxide formation and the role of catalase. Eur J Biochem. (1972) 25:420–30. doi: 10.1111/j.1432-1033.1972.tb01711.x
40. Kang SW, Baines I, Rhee SG. Characterization of a mammalian peroxiredoxin that contains one conserved cysteine. J Biol Chem. (1998) 273:6303–11. doi: 10.1074/jbc.273.11.6303
41. Huang J, Xiong K, Zhang H, Zhao Y, Cao J, Gong H, et al. Molecular characterization of Babesia microti thioredoxin (BmTrx2) and its expression patterns induced by antiprotozoal drugs. Parasit Vectors. (2018) 11:38. doi: 10.1186/s13071-018-2619-9
42. Huang J, Xiong K, Zhang H, Zhao Y, Cao J, Zhou Y, et al. Babesia microti thioredoxin 3 is an effective antioxidant and involved in the response to antiprotozoal drugs. Ticks Tick Borne Dis. (2018) 9:645–53. doi: 10.1016/j.ttbdis.2018.01.021
43. Pfaffl MW. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. (2001) 29:e45. doi: 10.1093/nar/29.9.e45
44. Blasa M, Angelino D, Gennari L, Ninfali P. The cellular antioxidant activity in red blood cells (CAA-RBC): a new approach to bioavailability and synergy of phytochemicals and botanical extracts. Food Chem. (2011) 125:685–91. doi: 10.1016/j.foodchem.2010.09.065
45. Fredriksson K, Stridh H, Lundahl J, Rennard SI, Skold CM. Red blood cells inhibit proliferation and stimulate apoptosis in human lung fibroblasts in vitro. Scand J Immunol. (2004) 59:559–65. doi: 10.1111/j.1365-3083.2004.01433.x
46. Bréhélin C, Meyer EH, de Souris JP, Bonnard G, Meyer Y. Resemblance and dissemblance of Arabidopsis type II peroxiredoxins: similar sequences for divergent gene expression, protein localization, and activity. Plant Physiol. (2003) 132:2045–57. doi: 10.1104/pp.103.022533
47. Rouhier N, Jacquot JP. The plant multigenic family of thiol peroxidases. Free Radic Biol Med. (2005) 38:1413–21. doi: 10.1016/j.freeradbiomed.2004.07.037
48. Banerjee M, Chakravarty D, Ballal A. Redox-dependent chaperone/peroxidase function of 2-Cys-Prx from the cyanobacterium Anabaena PCC7120: role in oxidative stress tolerance. BMC Plant Biol. (2015) 15:60. doi: 10.1186/s12870-015-0444-2
49. Lee EM, Lee SS, Tripathi BN, Jung HS, Cao GP, Lee Y, et al. Site-directed mutagenesis substituting cysteine for serine in 2-Cys peroxiredoxin (2-Cys Prx A) of Arabidopsis thaliana effectively improves its peroxidase and chaperone functions. Ann Bot. (2015) 116:713–25. doi: 10.1093/aob/mcv094
50. Clarke DJ, Ortega XP, Mackay CL, Valvano MA, Govan JR, Campopiano DJ, et al. Subdivision of the bacterioferritin comigratory protein family of bacterial peroxiredoxins based on catalytic activity. Biochemistry. (2010) 49:1319–30. doi: 10.1021/bi901703m
51. Rhee SG, Woo HA. Multiple functions of peroxiredoxins: peroxidases, sensors and regulators of the intracellular messenger H2O2, and protein chaperones. Antioxid Redox Signal. (2011) 15:781–94. doi: 10.1089/ars.2010.3393
52. Dietz KJ. Peroxiredoxins in plants and cyanobacteria. Antioxid Redox Signal. (2011) 15:1129–59. doi: 10.1089/ars.2010.3657
53. Okazaki S, Naganuma A, Kuge S. Peroxiredoxin-mediated redox regulation of the nuclear localization of Yap1, a transcription factor in budding yeast. Antioxid Redox Signal. (2005) 7:327–34. doi: 10.1089/ars.2005.7.327
54. Horling F, Lamkemeyer P, König J, Finkemeier I, Kandlbinder A, Baier M, et al. Divergent light-, ascorbate-, and oxidative stress-dependent regulation of expression of the peroxiredoxin gene family in Arabidopsis. Plant Physiol. (2003) 131:317–25. doi: 10.1104/pp.010017
55. Chae HZ, Uhm TB, Rhee SG. Dimerization of thiol-specific antioxidant and the essential role of cysteine 47. Biochemistry. (1994) 91:7022–26. doi: 10.1073/pnas.91.15.7022
56. König J, Galliardt H, Jütte P, Schäper S, Dittmann L, Dietz KJ. The conformational bases for the two functionalities of 2-cysteine peroxiredoxins as peroxidase and chaperone. J Exp Bot. (2013) 64:3483–97. doi: 10.1093/jxb/ert184
57. Kim YE, Hipp MS, Bracher A, Hayer-Hartl M, Hartl FU. Molecular chaperone functions in protein folding and proteostasis. Annu Rev Biochem. (2013) 82:323–55. doi: 10.1146/annurev-biochem-060208-092442
58. Cho C, Lee GW, Hong SH, Kaur S, Jung KW, Jung JH, et al. Novel functions of peroxiredoxin Q from Deinococcus radiodurans R1 as peroxidase and a molecular chaperone. FEBS Lett. (2019) 593:219–29. doi: 10.1002/1873-3468.13302
59. Koncarevic S, Rohrbach P, Deponte M, Krohne G, Prieto JH, Yates J III, et al. The malarial parasite Plasmodium falciparum imports the human protein peroxiredoxin 2 for peroxide detoxification. Proc Natl Acad Sci USA. (2009) 106:13323–8. doi: 10.1073/pnas.0905387106
60. Mendes RE, Pérez-Ecija RA, Zafra R, Buffoni L, Martínez-Moreno A, Dalton JP, et al. Evaluation of hepatic changes and local and systemic immune responses in goats immunized with recombinant Peroxiredoxin (Prx) and challenged with Fasciola hepatica. Vaccine. (2010) 28:2832–40. doi: 10.1016/j.vaccine.2010.01.055
Keywords: Babesia microti, thioredoxin, peroxidase-Q, antioxidant activity, drug target
Citation: Zhang H, Wang Z, Huang J, Cao J, Zhou Y and Zhou J (2020) A Novel Thioredoxin-Dependent Peroxiredoxin (TPx-Q) Plays an Important Role in Defense Against Oxidative Stress and Is a Possible Drug Target in Babesia microti. Front. Vet. Sci. 7:76. doi: 10.3389/fvets.2020.00076
Received: 19 November 2019; Accepted: 29 January 2020;
Published: 18 February 2020.
Edited by:
Lise Roy, Paul Valéry University, Montpellier III, FranceReviewed by:
Xiangye Liu, Xuzhou Medical University, ChinaXiaokai Song, Nanjing Agricultural University, China
Copyright © 2020 Zhang, Wang, Huang, Cao, Zhou and Zhou. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jinlin Zhou, amlubGluemhvdUBzaHZyaS5hYy5jbg==
†These authors have contributed equally to this work
 Houshuang Zhang
Houshuang Zhang Zhonghua Wang
Zhonghua Wang Jingwei Huang
Jingwei Huang Jie Cao
Jie Cao Yongzhi Zhou
Yongzhi Zhou Jinlin Zhou
Jinlin Zhou