- State Key Laboratory of Veterinary Etiological Biology, Key Laboratory of Veterinary Parasitology of Gansu Province, Lanzhou Veterinary Research Institute, CAAS, Lanzhou, China
Hydatigera taeniaeformis (H. taeniaeformis) is one of the most robust of tapeworm parasites that is widely distributed around the world. Information of proteins of H. taeniaeformis has not previously been reported. Using liquid chromatography tandem-mass spectrometry (LC–MS/MS) analysis, the proteome of H. taeniaeformis metacestode was profiled and a total of 408 proteins were identified. Of these, 26.5% (108/408) were annotated to be associated with metabolic pathways. Consistently, Gene Ontology analysis showed that those identified proteins were mainly classified into metabolic process of the biological processes. A set of metabolic enzymes, including Fructose-1,6-bisphosphate aldolase, enolase, Glucan phosphorylase, and phosphoenolpyruvate carboxykinase, were abundant in H. taeniaeformis metacestodes. In addition, some rare but interesting proteins like antigens (such as tegument protein and Antigen B) were identified. The two recombinant proteins of tegument protein and Antigen B were well-recognized by the sera from the H. taeniaeformis-infected mice. The H. taeniaeformis metacestode proteome might help to find new candidates for the immunodiagnosis and vaccine development and provide valuable information on H. taeniaeformis biology.
Keypoints
• A total of 408 proteins were identified in T. taeniaeformis metacestodes.
• Identified proteins were classified mainly to be involved in metabolic process.
• The two recombinant proteins of tegument protein and Antigen B were well-recognized by the sera from H. taeniaeformis-infected mice.
Introduction
Hydatigera taeniaeformis (also called Taenia taeniaeformis) is a parasitic helminth belonging to a family of Taenidae. The natural life cycle of H. taeniaeformis alternates between a definitive host (cats or dogs) and an intermediate host (mostly rodents and less frequently lagomorphs and humans) (1, 2). Adult tapeworms inhabit the small intestine of definitive cats or dogs and shed the terminal segments into the environment. Rodents which serve as the intermediate hosts are infected by ingesting the feed or water contaminated eggs or gravid proglottids (3, 4). Oncosphere larvae hatched from eggs in the small intestine. The larvae were passed through the digestive tract and transported passively though blood or lymph vessels to the livers, where the oncosphere larvae develop into metacestode larvae. Cysticercus fasciolaris is a metacestode stage of H. taeniaeformis. Recently, there are many reports of H. taeniaeformis infection in wild rodents (5–7). Our recent results highlight the prevalence of H. taeniaeformis in wild rodents and a risk of opportunistic parasite infection in human populations, especially those who have close contact with pets (7).
To date, proteomic studies of several cestodes have been reported, including Echinococcus spp. (8, 9), Taenia solium (10), Taenia hydatigena (11), and Taenia ovis (12). However, information of proteins of H. taeniaeformis has not previously been reported. This study is the first to describe the analysis of H. taeniaeformis metacestode protein extracts by Liquid chromatography and tandem mass spectrometry (LC-MS/MS). Identification and characterization of proteins from H. taeniaeformis metacestode might help to find new candidates for the immunodiagnosis and vaccine development and provide valuable information on H. taeniaeformis biology.
Materials and Methods
Parasites
By surveying on parasites in wild rodents in Xiji County, we obtained cysts in mouse livers. Fresh metacestodes were carefully dissected from the livers of naturally wild rodents as previously described (7). Combined with the morphological characteristics and host preference, the isolated parasites were suspected to be H. taeniaeformis (Figure S1). Then parasites were further identified using mitochondrial cox1 genes as a molecular marker. Sequence analysis showed that the cox1 nucleotide sequences were 1620 bp in length and shared 99% identity with that of H. taeniaeformis. Phylogenetic tree was constructed by the maximum likelihood method based on sequences of mitochondrial cytochrome oxidase I (cox1). In the phylogenetic tree based on cox-1 sequences, both Miaoping (MP) and Wangping (WP) isolates were clustered together with H. taeniaeformis isolated from China (FJ886784, FJ597547, and NC_014768), Germany (JQ663994.1), and Japan (AB221484.1) with high probabilities (Figure S2). The dissected parasites were washed with ice-cold PBS. Afterwards, parasites were immediately frozen in liquid nitrogen and then stored at −80°C until use.
LC–MS/MS Analysis and Protein Identification
Hydatigera taeniaeformis metacestodes were grounded into powder and homogenized in 100 μL of lysis buffer (containing 0.5 mM PMSF and 1 × proteinase inhibitor), and then added 6 M urea to the mixture. After centrifugation, the supernatant proteins (approximately 2 mg) were digested with 40 μL trypsin buffer (3 μg trypsin (Promega, USA) in 40 μL (25 mM) NH4HCO3) in a 37 °C water bath for 16–18 h. This digest was transferred to clean ultrafiltration tubes fitted with 10 kDa membranes, and centrifuged at 14,000 × g for 10 min. Peptide sequencing was conducted using high performance liquid chromatography coupled tandem mass spectrometry (LC-MS/MS) in Q-Exactive (Thermo Fisher Scientific) as described previously (12). The raw LC-MS/MS data was analyzed with the Mascot search engine (version 2.3) against H. taeniaeformis protein database retrieved from WormBase Parasite1 (http://parasite.wormbase.org/Hydatigera_taeniaeformis_prjeb534/Info/Index/) with following parameters: tryptic-specific peptides, maximum of one missed cleavages, a peptide mass tolerance of 20 ppm and a MS/MS tolerance of 0.5 Da. Label-free quantitation of proteins were calculated by intensity-based absolute quantification (iBAQ). Gene Ontology (GO) terms and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathways for the identified proteins were annotated.
Cloning, Expression, Purification, and Serological Evaluation of the Recombinant Proteins
The coding sequences of tegument protein and Antigen B genes were amplified with their specific primes. PCR fragments were digested and subcloned into prokaryotic expression plasmid pET-28a. The recombinant plasmids were transformed into Escherichia coli BL21 (DE3) cells. For the expression and purification of the recombinant proteins, bacteria were induced with 0.5 M isopropyl-b-D-thiogalactoside (IPTG) and the recombinant proteins were purified by Ni Sepharose 6 Fast Flow (GE Healthcare) and checked by 15% sodium dodecyl sulfate-polyacrylamide gel electrophoresis SDS-PAGE gel.
Western Blotting Analysis
The recombinant proteins were separated by 12% SDS-PAGE and transferred to a PVDF membranes using a Semi-Dry electrophoretic transfer cell (BioRad) for 30 min. The membranes were incubated with 5% (w/v) non-fat for 1 h and then incubated overnight at 4°C with mouse ant-sera against H. taeniaeformis (1:1,000, v/v dilution). Finally, they were washed and incubated with a horseradish peroxidase (HRP)-conjugated goat anti-mouse IgG (1:1,000, v/v dilution; Boster, Wuhan, China) for 1 h. Signals were visualized by exposing X-ray films using Pierce ECL Western Blotting Substrate (ThermoFisher Scientific, USA).
Results and Discussion
A total number of 4,024 spectra were detected and 2,946 unique peptides were obtained by LC-MS/MS in H. taeniaeformis metacestodes. In total, 408 proteins were identified by at least two unique peptides (Table S1). Similarly, 633 proteins with at least 2 unique spectra were identified from Taenia ovis metacestodes (12). The identified proteins include many high-abundant house-keeping proteins, such as actin, paramyosin, heat shock protein 70 (HSP70), and phosphoenolpyruvate carboxykinase, but also some rare but interesting proteins like antigens (tegument and immunogenic proteins) and receptors. The biological processes category showed enrichment of proteins involved in the metabolic process (36.6%) (Figure 1A), whereas molecular function showed a significant enrichment in catalytic activity (46.7%) and binding (44.2%) (Figure 1B). Within the cellular component category, the most enriched components were associated with structural proteins (40.2%) (Figure 1C). GO term enrichment analysis of identified proteins were shown in Figure S3. KEGG analysis showed that 408 proteins were annotated into 241 pathways, predominantly metabolic pathways (Figure 2).
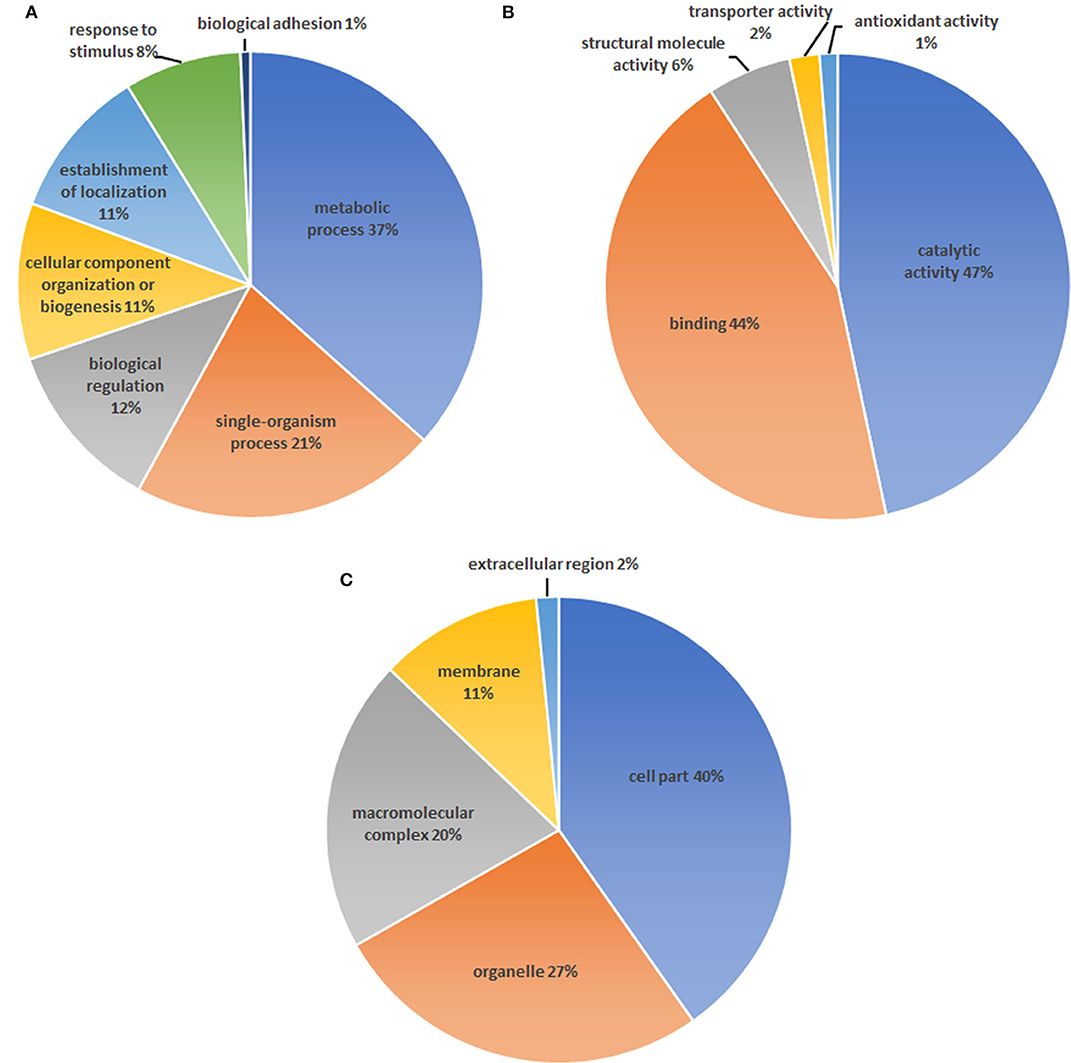
Figure 1. Gene ontology analysis of proteins identified in the Hydatigera taeniaeformis metacestodes. (A) biological processes. (B) Molecular function. (C) Cellular component.
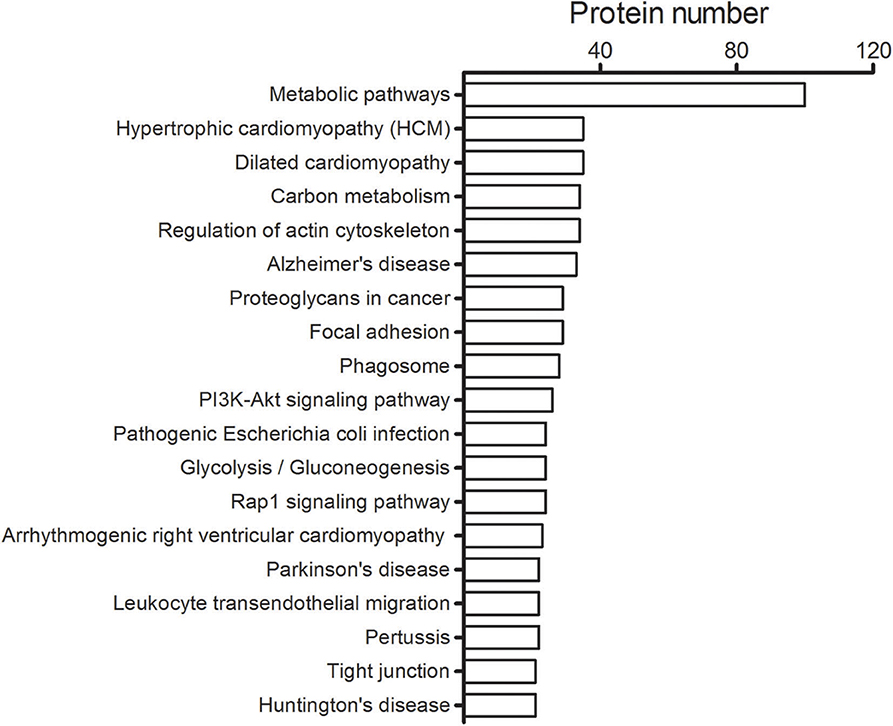
Figure 2. Top 20 pathways identified for the H. taeniaeformis metacestodes. The number of proteins involved in each of pathways is shown beside individual columns.
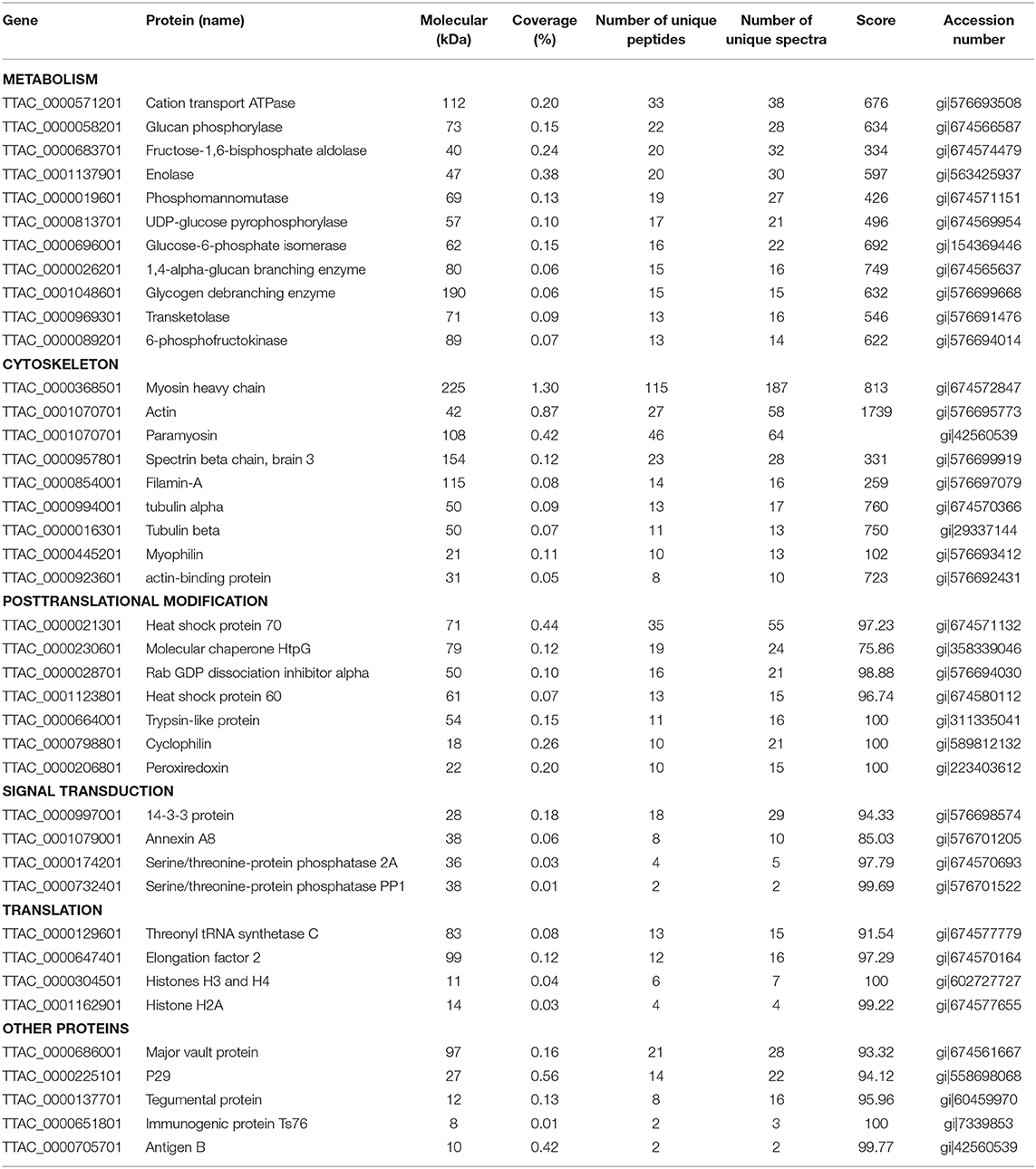
According to cluster of orthologous group (COG) functional classification, 287 identified proteins were mainly grouped into six categories (metabolism, cytoskeletal proteins, post-translational modification, signal transduction, translation, and other proteins) (Table S1). Approximately, 29.7% (121 proteins) of all identified proteins could not be assigned to any functional category (annotated as uncharacterized proteins) due to many proteins whose function were uncharacterized. Most of the identified H. taeniaeformis proteins are related to the metabolism (Table S1). Three metabolic enzymes were most abundant in the metacestodes of H. taeniaeformis, including glucan phosphorylase, fructose-bisphosphate aldolase (FBA), and enolase (Table 1). Higher abundance of three metabolic enzymes suggest that the sugar metabolism is an important event in the metacestodes of H. taeniaeformis. Whether the high expression of fructose-bisphosphate aldolase and enolase is beneficial for parasite's growth and development needs to be investigated in future. Previous studies showed enolase has primarily functioned as glycolytic enzymes in the cytoplasm (13–15). Besides the cytoplasmic localization, enolase was detected in the tegument of protoscolexes and adult worms, suggesting potentially participated in host–parasite molecular crosstalk (13). The absence of a typical N-terminal signal peptide indicates that both FBA and enolase does not transit through the classical secretory pathway but it possibly enters an exosome-mediated secretion. Both FBA and enolase has been identified in the hydatid fluid of various helminth parasites, and excretory–secretory (ES) products and exosome-like vesicles from in vitro-cultured protoscoleces (13, 16–18), showing potential as antigenic candidate antigens for the diagnosis of parasitic infections.
A group of cytoskeletal proteins appeared to be very abundant in the H. taeniaeformis metacestodes stage, including actin, paramyosin, actin and related proteins (Table 1). These proteins have also been identified in protoscoleces from other tapeworm parasites like Echinococcus multilocularis (9). Several studies showed that helminth cytoskeletal proteins could be potential vaccine candidates (e.g., paramyosin and actin-binding protein) or targets of parasite motility (19, 20). Of them, paramyosin, a 97 kDa myofibrillar protein, can induce effective anti-infective immune protection (21). Previous study showed monoclonal anti-paramyosin antibody conferred good protection against Schistosoma japonicum in mice (22). A recent study showed that the recombinant paramyosin protein also conferred significant levels of protection against S. japonicum infection in water buffalo (19). Therefore, the high-abundant paramyosin may render it as a promising drug target or vaccine antigen against H. taeniaeformis. Two heat shock proteins (HSP70 and HSP60), related to posttranslational modification, were very abundant in the H. taeniaeformis metacestodes (Table 1). Several studies showed flatworm HSP is considered as vaccine and diagnostic targets for parasitic diseases (23–25).
Proteins of signaling transduction group were dominated by 14-3-3 and Annexin A8 (Table 1). The 14-3-3 proteins as adaptor molecular are implicated in many signaling mechanisms due to their interaction with various target proteins (26). A more extensive function of 14–3-3 proteins have been studied in Schistosoma spp. and Echinococcus spp (27–29). Multiple isoforms of 14-3-3 proteins exist in different parasitic organisms. For example, seven different 14–3-3 isoforms were found in S. mansoni and S. japonicum while five different 14-3-3 isoforms were detected in genus Echinococcus (28, 30). Interesting, 14–3-3 were detected in excretion/secretion products and exosome-like vesicles from in vitro-cultured protoscoleces (31). The important roles of 14–3-3 proteins in parasite proliferation and survival suggested that these proteins have potential as vaccine candidates.
Three typical nuclear protein homologs grouped into translation group were identified, including an elongation factor EF-2, an adenine nucleotide translocator, and Histones H3/4. The presence of elongation factors has been detected in Taenia saginata and Taenia asiatica (32). Echinococcus granulosus elongation factor-1 β/δ (EgEF-1 β/δ) is an allergenic molecule of potential clinical importance of the intensity of echinococcosis immune response (33). Histones H3 have been identified as a biomarker for neoblasts in E. multilocularis metacestode (34). Several identified proteins were previously characterized as antigenic or involved in mechanisms of host immune evasion, such as Antigen B (AgB) (35), tegumental protein (TegP) (36), and putative major vault protein (37). Early studies have proposed that protective antibodies in serum may only be effective on the very early stages of H. taeniaeformis (38, 39). A successful immunization of mice against H. taeniaeformis using solubilized metacestode proteins suggested that more effective functional antigens appear to be present in the somatic antigens extracted from the metacestodes (40, 41). It would be interesting to screen functional antigens from the metacestodes and evaluate their protective roles against H. taeniaeformis infection. In present study, we selected and evaluated the immunoreactivity of two antigens (TegP and AgB) by western blot. The recombinant proteins of HtTegP and HtAtB were about 18 and 16 kDa on the SDS-PAGE, respectively (Figure 3). Western blot assays revealed that they could be reacted with mouse anti-sera against H. taeniaeformis, but not by the negative control serum, indicating the good immunoreactivity of recombinant proteins (Figure 3). Tegumental proteins (TegPs) are a group of proteins that coat on worm's entire outer surface (36). A growing number of studies have shown that parasite TegPs are associated with host immune responses during infection (42, 43). Recently study showed that CsTegP may be a potential druggable target (44). In E. multilocularis, TegP11 was primarily localized in the surface of protoscoleces and has an immunoregulatory capacity on RAW246.7 macrophages (36). Antigen B has received more attention due to its high specificity and good immunogenicity (45). Several studies showed Antigen B has great potential potentiality for diagnosis of human cystic echinococcosis (46, 47). Further studies will be evaluated for their immunodiagnostic value and immune protection against H. taeniaeformis infection.
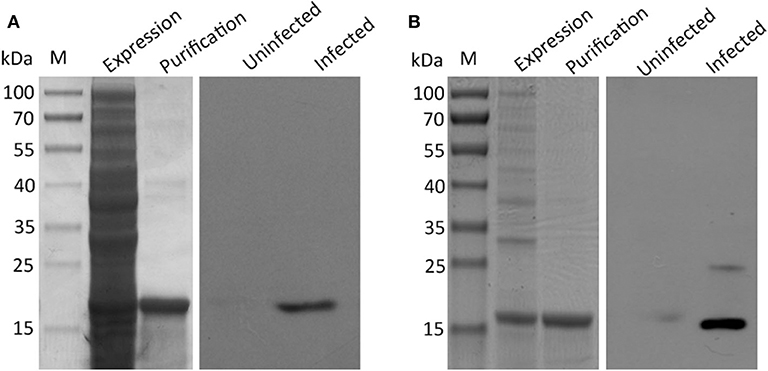
Figure 3. SDS-PAGE and western blot analysis of recombinant proteins of tegument protein (A) and Antigen B (B). The expression and purification of the recombinant proteins were run on 12% SDS-PAGE, respectively. Western blot analysis of recombinant proteins using sera from H. taeniaeformis-infected and uninfected mice.
Conclusions
A total of 408 proteins were identified in T. taeniaeformis metacestodes, most of which were classified mainly to be involved in metabolic process. The two recombinant proteins of tegument protein and Antigen B were well-recognized by the sera from H. taeniaeformis-infected mice. Identification and characterization of proteins from H. taeniaeformis metacestode might help to find new candidates for the immunodiagnosis and vaccine development.
Data Availability Statement
The raw data supporting the conclusions of this article will be made available by the authors, without undue reservation, to any qualified researcher.
Ethics Statement
Animal experiments were approved by Animal Ethics Committee of Lanzhou Veterinary Research Institute, Chinese Academy of Agricultural Sciences. All animal experiments in the study were handled in strict accordance with good animal practice of Animal Ethics Procedures and Guidelines of the People's Republic of China.
Author Contributions
XG performed study design, data analysis, and preparation of the manuscript.
Funding
This study was financially supported by grants from the National Natural Science Foundation of China (No. 31702224).
Conflict of Interest
The author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fvets.2020.00474/full#supplementary-material
References
1. Lavikainen A, Iwaki T, Haukisalmi V, Konyaev SV, Casiraghi M, Dokuchaev NE, et al. Reappraisal of Hydatigera taeniaeformis (Batsch, 1786) (Cestoda: Taeniidae) sensu lato with description of Hydatigera kamiyai n. sp. Int J Parasitol. (2016) 46:361–74. doi: 10.1016/j.ijpara.2016.01.009
2. Haukisalmi V, Lehtinen M, Henttonen H, Oksanen A, Meri S. A phylogeny of members of the family Taeniidae based on the mitochondrial cox1 and nad1 gene data. Parasitology. (2008) 135:1457–67. doi: 10.1017/S003118200800499X
3. Soulsby EJL. Helminths arthropods and protozoa of domesticated animals. Arthropod Parasites Domestic Anim. (1982) 55:249. doi: 10.2307/3277384
4. Catalano S, Bâ K, Diouf ND, Léger E, Verocai GG, Webster JP. Rodents of Senegal and their role as intermediate hosts of Hydatigera spp. (Cestoda: Taeniidae). Parasitology. (2018) 146:299–304. doi: 10.1017/S0031182018001427
5. Sinniah B, Narasiman M, Habib S, Gaik BO. Prevalence of Calodium hepaticum and Cysticercus fasciolaris in Urban rats and their histopathological reaction in the livers. J Vet Med. (2014) 2014:172829. doi: 10.1155/2014/172829
6. Gholipoury M, Rezai HR, Namroodi S, Khazaeli FA. Zoonotic and non-zoonotic parasites of wild rodents in turkman sahra, Northeastern Iran. Iran J Parasitol. (2016) 11:350–7.
7. He W, Ke L, Guo X, Chen Y. A survey on parasites in wild rodents in Xiji County, a northwestern part of China. Tropical Biomed. (2017) 34:449–52. Available online at: https://www.cabdirect.org/cabdirect/abstract/20173312184
8. Hidalgo C, García MP, Stoore C, Ramírez JP, Monteiro KM, Hellman U, et al. Proteomics analysis of Echinococcus granulosus protoscolex stage. Vet Parasitol. (2016) 218:43–5. doi: 10.1016/j.vetpar.2015.12.026
9. Wang Y, Cheng Z, Lu X, Tang C. Echinococcus multilocularis: proteomic analysis of the protoscoleces by two-dimensional electrophoresis and mass spectrometry. Exp Parasitol. (2009) 123:162–7. doi: 10.1016/j.exppara.2009.06.014
10. Santivañez SJ, Hernández-González A, Chile N, Oleaga A, Arana Y, Palma S, et al. Proteomic study of activated Taenia solium oncospheres. Mol Biochem Parasitol. (2010) 171:32–9. doi: 10.1016/j.molbiopara.2010.01.004
11. Zheng Y. Proteomic analysis of Taenia hydatigena cyst fluid reveals unique internal microenvironment. Acta Tropica. (2017) 176:224–7. doi: 10.1016/j.actatropica.2017.08.015
12. Zheng Y. Proteomic analysis of Taenia ovis metacestodes by high performance liquid chromatography-coupled tandem mass spectrometry. Vet Parasitol. (2017) 236:113–6. doi: 10.1016/j.vetpar.2017.01.024
13. Lorenzatto KR, Monteiro KM, Paredes R, Paludo GP, da Fonsêca MM, Galanti N, et al. Fructose-bisphosphate aldolase and enolase from Echinococcus granulosus: genes, expression patterns and protein interactions of two potential moonlighting protein. Gene. (2012) 506:76–84. doi: 10.1016/j.gene.2012.06.046
14. Wang Y, Zhang J, Yuan ZY. Cloning, expression and immunodiagnostic evaluation of enolase from Echinococcus granulosus. Zhongguo Xue Xi Chong Bing Fang Zhi Za Zhi. (2012) 24:549–52. doi: 10.16250/j.32.1374.2012.05.019
15. Mcmanus DP, Smyth JD. Isoelectric focusing of some enzymes from Echinococcus granulosus (horse sheep strains) E. multilocularis. Trans R Soc Trop Med Hyg. (1979) 73:259–65. doi: 10.1016/0035-9203(79)90079-8
16. Li WH, Qu ZG, Zhang NZ, Yue L, Jia WZ, Luo JX, et al. Molecular characterization of enolase gene from Taenia multiceps. Res Vet Sci. (2015) 102:53–8. doi: 10.1016/j.rvsc.2015.06.013
17. Zhang S, Guo A, Zhu X, You Y, Hou J, Wang Q, et al. Identification and functional characterization of alpha-enolase from Taenia pisiformis metacestode. Acta Tropica. (2015) 144:31–40. doi: 10.1016/j.actatropica.2015.01.007
18. Bernal D, de la Rubia JE, Carrasco-Abad AM, Toledo R, Mas-Coma S, Marcilla A. Identification of enolase as a plasminogen-binding protein in excretory–secretory products of Fasciola hepatica. FEBS Lett. (2004) 563:203–6. doi: 10.1016/S0014-5793(04)00306-0
19. Wu HW, Fu ZQ, Lu K, Pondtor S, Meng R, Hong Y, et al. Vaccination with recombinant paramyosin in Montanide ISA206 protects against Schistosoma japonicum infection in water buffalo. Vaccine. (2017) 35:3409–15. doi: 10.1016/j.vaccine.2017.05.007
20. Vogt CM, Armða-Fernà ndez MT, Tobler K, Hilbe M, Aguilar C, Ackermann M, et al. Oral application of recombinant Bacillus subtilis spores to dogs results in a humoral response against specific Echinococcus granulosus paramyosin and tropomyosin antigens. Infect Immun. (2017) 86:IAI.00495–17. doi: 10.1128/IAI.00495-17
21. Kalinna BH, Mcmanus DP. A vaccine against the Asian schistosome, Schistosoma japonicum: an update on paramyosin as a target of protective immunity. Int J Parasitol. (1997) 27:1213–9. doi: 10.1016/S0020-7519(97)00119-7
22. Kojima S, Janecharut T, Hata H, Niimura M. Role of a mouse monoclonal IgE antibody in passive transfer of immunity to Schistosoma japonicum infection. Memórias Do Instituto Oswaldo Cruz. (1987) 82:237. doi: 10.1590/S0074-02761987000800045
23. Vacirca D, Perdicchio M, Campisi E, Delunardo F, Ortona E, Margutti P, et al. Favourable prognostic value of antibodies anti-HSP20 in patients with cystic echinococcosis: a differential immunoproteomic approach. Parasite Immunol. (2011) 33:193–8. doi: 10.1111/j.1365-3024.2010.01264.x
24. Ming MD, Rui MX, Yuan CX, Yun YL, Liu Q, Guo FC, et al. SjHSP70, a recombinant Schistosoma japonicum heat shock protein 70, is immunostimulatory and induces protective immunity against cercarial challenge in mice. Parasitol Res. (2015) 114:3415–29. doi: 10.1007/s00436-015-4567-z
25. Caffrey CR, Rohwer A, Oellien F, Braschi S, Oliveira G, Mckerrow JH, et al. A comparative chemogenomics strategy to predict potential drug targets in the metazoan pathogen, Schistosoma mansoni. PLoS ONE. (2009) 4:e4413. doi: 10.1371/journal.pone.0004413
26. Siles-Lucas Mdel M, Gottstein B. The 14-3-3 protein: a key molecule in parasites as in other organisms. Trends Parasitol. (2003) 19:575–81. doi: 10.1016/j.pt.2003.10.003
27. Nunes CP, Zaha A, Gottstein B, Müller N, Siles-Lucas MDM. 14-3-3gene characterization description of a second 14-3-3 isoform in both Echinococcus granulosus E. multilocularis. Parasitol Res. (2004) 93:403–9. doi: 10.1007/s00436-004-1147-z
28. Teichmann A, Vargas DM, Monteiro KM, Meneghetti BV, Dutra CS, Paredes R, et al. Characterization of 14-3-3 isoforms expressed in the Echinococcus granulosus pathogenic larval stage. J Proteome Res. (2015) 14:1700–15. doi: 10.1021/pr5010136
29. Mcgonigle S, Loschiavo M, Pearce EJ. 14-3-3 Proteins in Schistosoma mansoni; identification of a second epsilon isoform. Int J Parasitol. (2002) 32:685–93. doi: 10.1016/S0020-7519(01)00323-X
30. Siles-Lucas M, Nunes CP, Zaha A. Comparative analysis of the 14-3-3 gene and its expression in Echinococcus granulosus and Echinococcus multilocularis metacestodes. Parasitology. (2001) 122:281–93. doi: 10.1017/S0031182001007405
31. Sileslucas M, Nunes CP, Zaha A, Breijo M. The 14-3-3 protein is secreted by the adult worm of Echinococcus granulosus. Parasite Immunol. (2000) 22:521–8. doi: 10.1046/j.1365-3024.2000.00334.x
32. Shen P, Wu X, Huang J, Hu X, Yu X, Bao H, et al. Prokaryotic expression of elongation factor 1 gene of Taenia saginata asiatica and analysis of purification and immunogenicity of the recombinant protein. J Xian Jiaotong Univ. (2008) 29:378–82. Avialable online at: http://www.google.com/url?sa=t&rct=j&q=&esrc=s&source=web&cd=&ved=2ahUKEwiU8cvZ0vvqAhWayDgGHep0Bz4QFjAIegQIChAB&url=http%3A%2F%2Fyxxb.xjtu.edu.cn%2Foa%2Fpdfdow.aspx%3FSid%3D200804005&usg=AOvVaw0oQwmZh2meH72xYcXdQQGx
33. Ortona E, Margutti P, Vaccari S, Riganò R, Profumo E, Buttari B, et al. Elongation factor 1 beta/delta of Echinococcus granulosus and allergic manifestations in human cystic echinococcosis. Clin Exp Immunol. (2010) 125:110–6. doi: 10.1046/j.1365-2249.2001.01569.x
34. Zhang XY, Yang JY, Gao ZQ, Lou ZZ, Jia WZ, Luo XN, et al. Identification and tissue distribution characterization of neoblasts in Echinococcus multilocularis metacestode. Chin Vet Sci. (2014) 44:45–9. doi: 10.16656/j.issn.1673-4696.2014.01.018
35. Riganò R, Profumo E, Bruschi F, Carulli G, Azzarà A, Ioppolo S, et al. Modulation of human immune response by Echinococcus granulosus antigen B and its possible role in evading host defenses. Infect Immun. (2001) 69:288–96. doi: 10.1128/IAI.69.1.288-296.2001
36. Zheng Y, Guo X, Su M, Chen X, Jin X, Ding J, et al. Identification of emu-TegP11, an EF-hand domain-containing tegumental protein of Echinococcus multilocularis. Vet Parasitol. (2018) 255:107–13. doi: 10.1016/j.vetpar.2018.04.006
37. Reis EV, Pereira RV, Gomes M, Jannotti-Passos LK, Baba EH, Coelho PMZ, et al. Characterisation of major vault protein during the life cycle of the human parasite Schistosoma mansoni. Parasitol Int. (2014) 63:120–6. doi: 10.1016/j.parint.2013.10.005
38. Musoke AJ, Williams JF. The immunological response of the rat to infection with Taenia taeniaeformis. V. sequence of appearance of protective immunoglobulins and the mechanism of action of 7Sgamma2a antibodies. Immunology. (1975) 29:855–66.
39. Cook RW, Trapp AL, Williams JF. Pathology of Taenia taeniaformis infection in the rat: hepatic, lymph node and thymic changes. J Comp Pathol. (1981) 91:219–26. doi: 10.1016/0021-9975(81)90026-8
40. Rajasekariah GR, Rickard MD, Mitchell GF. Immunization of mice against infection with Taenia taeniaeformis using various antigens prepared from eggs, oncospheres, developing larvae and strobilocerci. Int J Parasitol. (1980) 10:315–24. doi: 10.1016/0020-7519(80)90013-2
41. Lightowlers MW, Mitchell GF, Bowtell DD, Anders RF, Rickard MD. Immunization against Taenia taeniaeformis in mice: studies on the characterization of antigens from oncospheres. Int J Parasitol. (1984) 14:321–33. doi: 10.1016/0020-7519(84)90084-5
42. Yan H, Zhenwen Z, Xuchu H, Quande W, Jin X, Zhongdao W, et al. A novel tegumental protein 31.8 kDa of Clonorchis sinensis: sequence analysis, expression, and immunolocalization. Parasitol Res. (2007) 102:77–81. doi: 10.1007/s00436-007-0728-z
43. Kim Y-J, Yoo WG, Lee M-R, Kim D-W, Lee W-J, Kang J-M, et al. Identification and characterization of a novel 21.6-kDa tegumental protein from Clonorchis sinensis. Parasitol Res. (2012) 110:2061–2066. doi: 10.1007/s00436-011-2681-0
44. Kim Y-J, Yoo WG, Lee M-R, Kang J-M, Na B-K, Cho S-H, et al. Molecular and structural characterization of the tegumental 20.6-kDa protein in clonorchis sinensis as a potential druggable target. Int J Mol Sci. (2017) 18:557. doi: 10.3390/ijms18030557
45. Zhang W, Li J, McManus DP. Concepts in immunology and diagnosis of hydatid disease. Clin Microbiol Rev. (2003) 16:18–36. doi: 10.1128/CMR.16.1.18-36.2003
46. Abdi J, Kazemi B, Mohebali M, Bandehpour M, Rahimi MT, Rokni MB. Gene cloning, expression and serological evaluation of the 12-kDa antigen-B subunit from Echinococcus granulosus. Ann Trop Med Parasitol. (2010) 104:399–407. doi: 10.1179/136485910X12743554760261
Keywords: Hydatigera taeniaeformis, metacestode, tegument protein, Antigen B, proteome
Citation: Guo X (2020) Proteomics Analysis of Hydatigera taeniaeformis Metacestode Stage. Front. Vet. Sci. 7:474. doi: 10.3389/fvets.2020.00474
Received: 19 December 2019; Accepted: 25 June 2020;
Published: 06 August 2020.
Edited by:
Jerome Depaquit, Université de Reims Champagne-Ardenne, FranceReviewed by:
Juan Pedro Laclette, National Autonomous University of Mexico, MexicoHoushuang Zhang, Chinese Academy of Agricultural Sciences, China
Copyright © 2020 Guo. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Xiaola Guo, Z3VveGlhb2xhQGNhYXMuY24=
 Xiaola Guo
Xiaola Guo