- 1Lanzhou Institute of Husbandry and Pharmaceutical Sciences of Chinese Academy of Agricultural Sciences (CAAS), Lanzhou, China
- 2Key Laboratory of New Animal Drug Project of Gansu Province, Lanzhou, China
- 3Key Laboratory of Veterinary Pharmaceutical Development, Ministry of Agriculture, Lanzhou, China
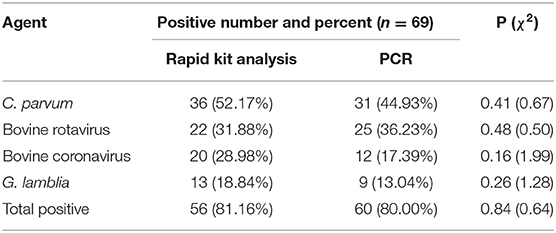
Neonatal calf diarrhea (NCD) is one of the most serious health challenges facing the livestock industry and has caused substantial economic losses due to increased morbidity and mortality rates. The present study investigated the main infectious pathogens causing NCD among cattle in Yangxin County, China. Sixty-nine fecal samples were collected from diarrheic newborn cattle and tested for infectious agents, including bovine rotavirus, bovine coronavirus, Escherichia coli K99, Cryptosporidium parvum, and Giardia lamblia, that cause NCD, as determined by rapid kit analysis and polymerase chain reaction (PCR) amplification. The PCR results showed that the percentages of samples that were positive for C. parvum, bovine rotavirus A, bovine coronavirus, and G. lamblia were 44.93, 36.23, 17.39, and 13.04%, respectively. The rapid kit analysis results showed that the prevalence of C. parvum, rotavirus, coronavirus, and G. lamblia was 52.17, 31.88, 28.98, and 18.84%, respectively. No E. coli K99 was detected by either method. The total positivity of the samples, as determined by PCR and rapid kit analysis, was 80.00 and 81.16%, respectively. No significant difference between the two methods was observed. The results of this study may help to establish a foundation for future research investigating the epidemiology of NCD in cattle and may facilitate the implementation of measures to control NCD transmission to cattle in Yangxin County, Shandong Province, China.
Introduction
Neonatal calf diarrhea (NCD) is one of the most serious diseases worldwide among newborn calves (<1 month old). NCD causes notable levels of morbidity and mortality through several complications, such as dehydration, acidosis, and solution depletion (1–3). At present, NCD continues to be a major cause of fruitfulness reduction and economic loss in Bos taurus herds around the world (4), despite advances in the cattle industry regarding herd management, animal facilities and care, feeding and nutrition, and timely use of biopharmaceuticals. This disease may be triggered by both infectious and non-infectious factors (the intrinsic characteristics of the calf, its organic processes, veterinary treatment, management of the herd, and environmental factors) (5). Among these factors, infectious agents are the leading cause of death. Among the numerous infectious agents causing NCD, rotavirus, coronavirus, Escherichia coli enterotoxin K99/F5, and Cryptosporidium parvum spp. are recognized as the four most essential pathogens (6). C. parvum and rotavirus are often detected in fecal samples (2). In past research, most analyses centered on individual pathogens, but recent studies have urged that concurrent infection of a number of pathogens may be necessary to model the pathophysiology of gastrointestinal diseases (7). A study reported that the rate of coinfection was 55% in fecal samples obtained from diarrheic calves. Notably, the rate of coinfection in healthy calves was only 3% (8). Cryptosporidiosis is a worldwide zoonotic disease, and cattle are recognized as a major parasite reservoir and contributor to zoonotic infection (9). Therefore, research into the pathogens that infect cattle and cause NCD is vital for farmers and staff members in regions with intensive cattle farming. On the other hand, antibiotics are currently measured through empirical observation based on when the symptom of loose stools occurs, though a proper etiological diagnosis is rarely made, and it may not be clear what the cause is. This approach leads to the unnecessary and excessive use of antibiotics in food animal species and the potential development of antibiotic-resistant bacteria. This phenomenon has prompted a ban on the subtherapeutic usage of antibiotics in several countries (10).
The aim of this study was to detect the types of pathogens causing NCD and to determine the prevalence of this disease in two intensive cattle herds in Yangxin County, Shandong Province, China, through diagnosis in order to design rational and efficacious protocols for the prevention and treatment of diarrhea.
Materials and Methods
Sampling
The study area was conducted in Yangxin County, Shandong Province, eastern China. This area is located between north latitudes 37°58′ and 37°66′ and east longitudes 117°47′ and 117°77′. A small geographical map of the farm is shown in Figure 1. This area is one of the most important cattle and dairy farming areas in the country. A total of 69 fresh stool samples were collected on October 8–10, 2019. At the beginning of October, during the NCD outbreak on the farm, the clinical signs were depression, anorexia, dehydration, and watery diarrhea. When the calves were administered broad-spectrum antibiotics (such as florfenicol), the animals were unresponsive to these medicines. Death occurred in four calves 3–5 days after the onset of diarrhea. To confirm the causes of the infectious agents, fresh stool specimens were obtained from calves for <1 month. Fifteen calves with no abnormal clinical signs were employed as controls, and 54 diarrheic calves were employed as the experimental group.
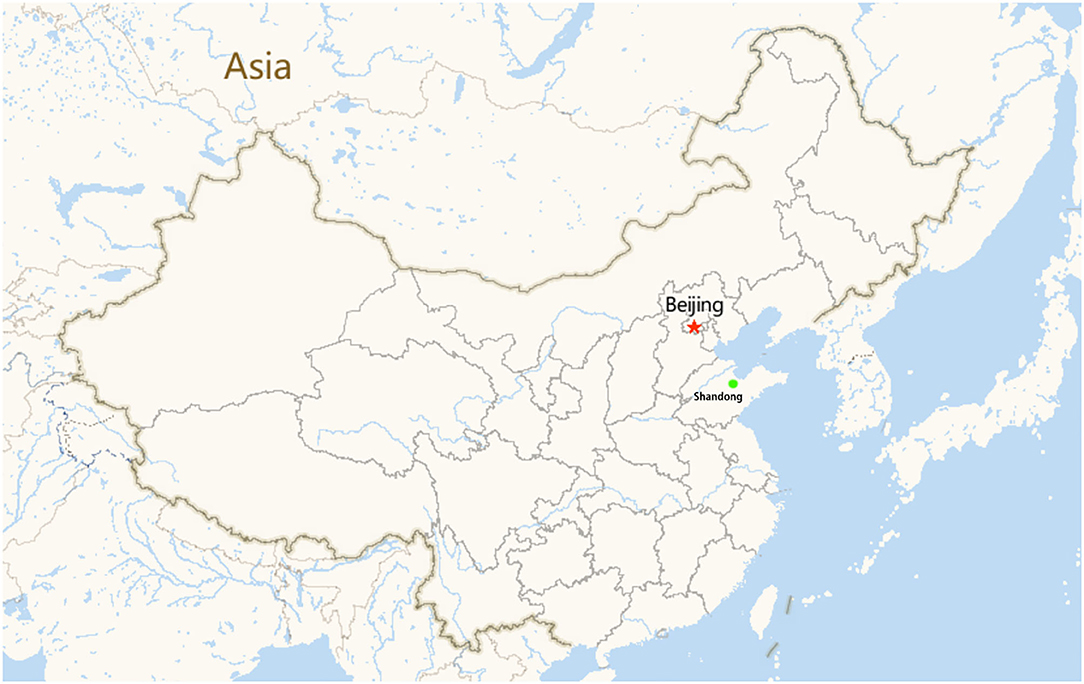
Figure 1. Small geographical map of the farms (the green dots represent the area where the cattle farms are located, the red star represent the capital of China, Beijing).
The female cows are transferred to the maternity barn after delivery, and the newborn calves are transferred to the calving barn in time. The colostrum is milked within 6 h after delivery, checked for quality, pasteurized, and then frozen. All calves were fed 4 L of colostrum within 12 h after birth. Calves were housed centrally in housing with ventilation, natural temperature, and unheated conditions. The study was conducted at two dairy cow farms (the average number of cows and breeding heifers is 800, with an average milk yield of 25 kg per cow per day). Heifers were not vaccinated against coronavirus and rotavirus. The cows were housed in a cattle-dense feedlot with all feed provided in a feed bunk and in individual pens. The amount of pasteurized whole milk offered to calves per day was about 6 L. Fecal samples were collected using disposable sterile plastic straw trimmed into the shape of a shovel and temporarily stored thereafter in an aseptic plastic sampling tube. To ensure that samples were not disturbed by microbes in the environment, swabs were utilized to take parts of the fecal center when sampling. All samples were stored in liquid nitrogen and transported to the laboratory for research. The numbers and ages of the animals were noted on each sampling tube.
The study obtained consent from the dairy farmers when collecting samples.
Detection of Infectious Agents by Anigen Test Kit
Stool samples were tested using the Anigen Rapid BoviD-5 Ag Test Kit (BioNote, Korea) within a short period after collection on the farm. The Anigen Rapid BoviD-5 Ag Test Kit employs a solid-phase immunochromatographic assay for the rapid and qualitative detection of C. parvum antigen, bovine rotavirus antigen, bovine coronavirus antigen, E. coli K99 antigen, and Giardia lamblia antigen in feces. The testing process was performed in accordance with the manufacturer's instructions.
DNA Extraction
The stool genomic DNA of feces was extracted using the E.Z.N.A. Stool DNA Kit (Omega, Georgia, USA). The genomic DNA was subsequently utilized to detect G. lamblia, C. parvum, and E. coli K99 by polymerase chain reaction (PCR) amplification.
RNA Extraction
The clinical fecal samples were diluted with phosphate-buffered saline (PBS), vortexed, and centrifuged at 13.8 g at 4°C for 5 min. The supernatant was collected and used to extract viral RNA. Total RNA from feces was extracted using a Viral RNA Kit (OMEGA, Georgia, USA) according to the instruction of the manufacturer. The viral RNA was subsequently used as a template for reverse transcription (RT)-PCR.
Confirmation of Infectious Agents by PCR Sequencing and Analysis
Primers for C. parvum, G. lamblia, E. coli, bovine rotavirus, and bovine coronavirus were used to detect infectious agents, as shown in Table 1.
Nested PCR was performed to detect C. parvum and Giardia using 2× Taq Master Mix (Vazyme, Nanjing, China). PCR amplification of E. coli was carried out using 2× ExTaq DNA polymerase (Vazyme, Nanjing, China).
Fragments of the 16S rRNA gene were amplified. A nested PCR protocol was used with the first-round primers Gia2029 and Gia2150 and the secondary primers RH11 and RH4; the β-giardin gene was amplified by nested PCR using G7/G759/G376 primers and PCR by GGL/GGR primers; the gdh gene was amplified by nested PCR using GDHeF/GDHiRm/GDHiF primers.
All PCR products were visualized on a 1.5% agarose gel. All positive samples were purified and submitted for Sanger sequencing (Tianqigene Bio Com, Lanzhou, China). The sequence results were aligned in GenBank for each possible C. parvum species, G. lamblia, and E. coli.
One-step RT-PCR was performed to confirm bovine rotavirus and bovine coronavirus using the PrimeScript One Step RT-PCR Kit, Ver. 2 (TaKaRa, Dalian, China). The reaction was performed with an initial reverse transcription step at 50°C for 30 min, followed by PCR amplification at 94°C for 2 min and 30 cycles (98°C for 10 s and 68°C for 1 min). PCR products were visualized on 1.0% agarose gels, and positive samples were subsequently selected for sequencing. The sequence results were compared using BLAST in GenBank to confirm the virus species.
Statistical Analysis
Statistical analysis was performed with R software, version 3.5.3. Chi-square tests were performed at a 5% level of significance.
Results
Detection of Infectious Agents by Rapid Kit Analysis
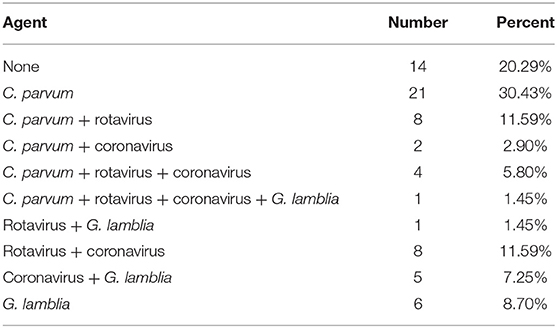
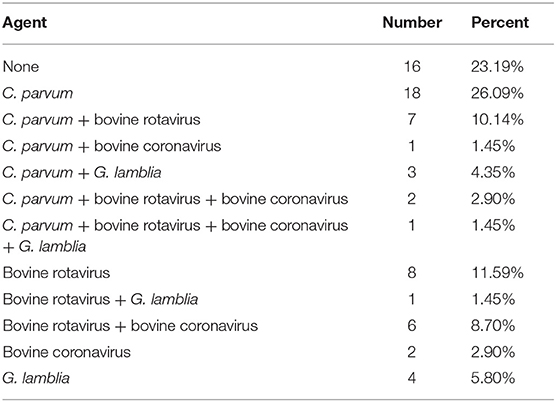
In this study, four etiological agents were tested, namely, C. parvum, rotavirus, coronavirus, and G. lamblia. From 69 fecal samples, 56 samples (81.16%) were positive for infectious agents. The results of the experiment are shown in Tables 2, 3 and Supplementary Figures 1, 2. The incidences of a single etiological agent with positive samples were as follows: C. parvum in 30.43% and G. lamblia in 8.70%. Negative samples were 20.29% (14/69). The rest of the samples exhibited coinfection.
The prevalence of coinfection among samples was as follows: C. parvum and rotavirus in 11.59%; coronavirus and rotavirus in 11.59%; coronavirus and G. lamblia in 7.25%; C. parvum, coronavirus, and rotavirus in 5.80%; C. parvum and coronavirus in 2.90%; rotavirus and G. lamblia in 1.45%; and C. parvum, coronavirus, rotavirus, and G. lamblia in 1.45%.
As seen from the data in Table 2, of the four major pathogens tested, C. parvum exhibited a significant preponderance of infection rates relative to the other three and was the predominant pathogen in this infection (P < 0.05). There were no significant differences in the proportions among the other three pathogens.
Detection of Infectious Agent by PCR
C. parvum
Subgenotyping data of 69 SSU rRNA sequences was employed to identify C. parvum. No novel genotype was obtained by variant BLAST searches. Among 69 samples, 31 (44.93%) were positive for C. parvum. Of these samples, 26.09% were infected by C. parvum alone, whereas 18.84% were mixed infections. In all mixed infections, C. parvum and bovine rotavirus coinfection was predominant (10.14%), followed by coinfection with C. parvum and G. lamblia (4.35%).
Giardia
The primers for β-giardin GGL-F/GGL-R (one-step PCR) and the gdh gene GDHeF/GDHiRm/GDHiF (two-step nested PCR) failed to amplify all the fecal samples in this study. However, the two sets of β-giardin primers G7/G376/G759 (two-step nested PCR) and Gia2029/Gia2150 and RH11/RH4 (two-step nested PCR) successfully amplified five and four samples, respectively. The PCR results showed that all Giardia pathogens belong to G. lamblia. The percentage of samples positive for G. lamblia was 13.04%. The percentage of samples only infected by G. lamblia was 5.80%, whereas the percentage of samples coinfected with this species was 7.24%.
Bovine Rotavirus
Sequencing identified the bovine rotavirus as bovine rotavirus A. The percentage of samples determined by PCR to be infected with bovine rotavirus was 36.23%. The percentage of samples determined to be coinfected with this virus was 24.64%, whereas the percentage of single-infected samples was 11.59%. Details of mixed infections are shown in Table 4.
Bovine Coronavirus
The overall positive rate of bovine coronavirus by PCR was 17.39% (12/69), and the rate of coinfection was 14.49%. Details of mixed infections are shown in Table 4.
E. coli
PCR results for E. coli were obtained from 0 (0%) out of 69 fecal samples.
Discussion
This study investigated and characterized the pathogens causing NCD on cattle farms in Yangxin County, Shandong Province, China. Two rural sample groups were studied, and although a larger sample size would have improved the validity of the results, the sample size for this study was limited by time.
Regarding the characteristics of the farm, the facilities, variety, and number of employees were similar between the two farms. The average size of the two farms was 800 cows, consisting of 80–90% adult cows, making them large-scale dairy farms. Klein-Jobstl believes that the occurrence of diarrhea is significantly associated with herd size (21). These two cattle farms were therefore at greater risk of diarrhea.
The maternity barn is believed to be strongly associated with the health of the calf. Regular cleaning of the calving barns can reduce the occurrence of calf diseases. The time calves spend in the maternity barn with their dams is also a risk factor. In particular, maternity barns have a poor environment with sick animals using maternity areas (22). Most enteric pathogens are transmitted between cows and calves, either by colostrum or by the fecal–oral route of environmental transmission. Even healthy cows carry a large number of pathogens during the perinatal period (23). In addition, adequate intake of colostrum is recognized to improve the immunity of calves and reduce the development of infectious intestinal diseases in calves (24). On the two farms that we investigated, all newborn calves were allowed to suckle colostrum from their dams; therefore, no colostrum-related diarrhea was observed in this investigation.
Barrington recommended housing calves separately, as this approach may result in a lower pathogen exposure (25). In a separate barn, calves are fed on individual basis in line with their specific desires, and it is easier to regulate the animals and identify certain abnormalities. In our survey, calves on both farms were kept in groups indoors with ventilation systems, but environmental hygiene issues were often overlooked, and although relevant disinfection and hygiene protocols were in place on the farms, they were not rigorously followed by the workers.
After calves are weaned from colostrum, their nutrition determines the level of immunity (26). The farms surveyed both used feeding to buy surplus waste milk from elsewhere. For calves, waste milk has a better nutritional composition than concentrated feed and can better enhance the growth of the animal. However, unpasteurized whole milk poses a high risk of intestinal infection if it is not consumed immediately (27). Both farms surveyed established pasteurization units for purchased waste milk to ensure the biosecurity of liquid feed.
A study by Gutzwiller et al. noted that diarrhea was considerably more likely to occur in calves during colder seasons, such as in autumn and winter, than in spring and summer, and that this phenomenon was not related to inadequate levels of immunoglobulin ingested by the calves (24). Our study was conducted in late autumn, and temperatures had started to drop, especially at night. While the change in weather affects calf diarrhea, it also contributes to another important factor—the onset of respiratory disease. There have been many studies demonstrating a close association between respiratory illness and diarrhea (27–29). For herds with >200 cows, the number of routine veterinary visits to the farm and the incidence of respiratory disease in calves accounted for 64.2% of the incidence of calf diarrhea (22). However, due to the short sampling time of the current survey, it was not possible to properly determine the risk of respiratory illness. However, we found that self-grooming of calves can be a risk factor for diarrhea (27). The pathogens become attached to the skin surface of the calf when it makes contact with objects in the barn, such as bars and floors, that may be contaminated with pathogenic bacteria, and then the pathogens from the environment enter the calf's body through licking the skin during self-grooming, leading to diarrhea.
In this investigation, the use of a rapid test kit to detect the four main pathogens of diarrhea demonstrated that C. parvum was detected most frequently, which is consistent with the results obtained by Bartels (30). The detection rate of C. parvum in the feces of diarrheic calves in our survey was 52.17%. Bartels et al. obtained a detection rate of ~43% for C. parvum in 1- to 2-week-old calves with diarrhea and an increase in shedding associated with the use of conventional antibiotics (30). Other studies have reported that over 60% of diarrheic calves are infected with C. parvum, either alone or in combination (31, 32). The same study also noted that rotavirus is often mixed with C. parvum (33), which is also very similar to the results of the current study (30). Coronaviruses are conditional pathogens that are rarely detected when good management conditions are ensured but can invade the body when environmental conditions become poor. Therefore, coronaviruses generally have a very low detection rate (29, 34). G. lamblia, although also a major cause of diarrhea in calves, was detected at a low rate in this survey, as well as in some other surveys (30). In general, E. coli is shed in a shorter period, resulting in a lower incidence; therefore, E. coli K99 was not detected in this survey (30). Although the use of rapid test kits is convenient and quick, the results obtained with these kits were validated using PCR analyses to verify their accuracy and specificity. The results presented in Table 2 show that there were no significant differences (P > 0.05) between the rapid test kit group and the PCR group when they were compared using the chi-square test. This result suggests the feasibility of employing rapid test kit for clinical use in calf diarrhea. Further research to perform pathogenic analysis of diarrheic cattle and ensure the rational use of drugs may be effective in reducing mortality.
Conclusion
Many factors were determined to be similar between the two farms investigated, including milk management, housing, and feeding procedures. Therefore, no important effects of those factors on the symptom of calves diarrhea were found during this study. In contrast, environmental conditions were shown to be considerably related to calves diarrhea on the farms. Variables that were strongly related to calves diarrhea were farm size, cleanliness of the calving barns during the birth of calves, placement of individual barns, and presence of respiratory disorders. The possible influence of those factors on calves diarrhea on farms should thus be considered. Optimization of these factors may cause a reduction in calves diarrhea, which may thus reduce morbidity and mortality, which would be of economic importance. The use of rapid test kits for the detection of pathogens in diarrheic calves and the administration of appropriate treatment may serve to reduce mortality due to diarrhea on large dairy cattle farms.
Data Availability Statement
The original contributions presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding author/s.
Ethics Statement
The experiment is approved by Research Ethics Committee of Lanzhou Institute of Husbandry and Pharmaceutical Science of CAAS, Lanzhou, PR China. All fecal samples were collected from cattles with diarrhea with the consent of the owners.
Author Contributions
XW proposed the test scheme. ZD wrote the manuscript and was responsible for revising it. WW and BL performed the laboratory experiments. XZ, FC, and XW collected the samples. JZ provided the funding resource. All authors contributed to the article and approved the submitted version.
Funding
This study was supported by the Research on the Key Technologies of the Creation and Industrialization of Basic Operating Expenses of the Institute (Grant Number 1610322017014) and Drug Development and Clinical Drug Use Posts of the National Beef Yak Industry Technical System (Grant Number CARS-37).
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
The authors acknowledge and are grateful for the help provided by staff members on the cattle farms. The authors would like to thank the American Journal Experts for their linguistic corrections to the manuscript.
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fvets.2020.589126/full#supplementary-material
Supplementary Figure 1. Rapid kit for positive results.
Supplementary Figure 2. Rapid kit for negative results.
References
1. Trefz FM, Lorch A, Feist M, Sauter-Louis C, Lorenz I. Construction and validation of a decision tree for treating metabolic acidosis in calves with neonatal diarrhea. BMC Vet Res. (2012) 8:238. doi: 10.1186/1746-6148-8-238
2. Meganck V, Hoflack G, Piepers S, Opsomer G. Evaluation of a protocol to reduce the incidence of neonatal calf diarrhoea on dairy herds. Prev Vet Med. (2015) 118:64–70. doi: 10.1016/j.prevetmed.2014.11.007
3. Golbeck L, Cohrs I, Leonhard-Marek S, Grunberg W. Effect of dehydration and acidemia on the potassium content of muscle tissue and erythrocytes in calves with neonatal diarrhea. J Dairy Sci. (2018) 101:9339–49. doi: 10.3168/jds.2018-14790
4. Picco NY, Alustiza FE, Bellingeri RV, Grosso MC, Motta CE, Larriestra AJ, et al. Molecular screening of pathogenic Escherichia coli strains isolated from dairy neonatal calves in Cordoba province, Argentina. Rev Argent Microbiol. (2015) 47:95–102. doi: 10.1016/j.ram.2015.01.006
5. Bendali F, Sanaa M, Bichet H, Schelcher F. Risk factors associated with diarrhoea in newborn calves. Vet Res. (1999) 30:509–22.
6. Kongsted H, Jonach B, Haugegaard S, Angen O, Jorsal SE, Kokotovic B, et al. Microbiological, pathological and histological findings in four Danish pig herds affected by a new neonatal diarrhoea syndrome. BMC Vet Res. (2013) 9:206. doi: 10.1186/1746-6148-9-206
7. Jang JY, Kim S, Kwon MS, Lee J, Yu DH, Song RH, et al. Rotavirus-mediated alteration of gut microbiota and its correlation with physiological characteristics in neonatal calves. J Microbiol. (2019) 57:113–21. doi: 10.1007/s12275-019-8549-1
8. Al Mawly J, Grinberg A, Prattley D, Moffat J, Marshall J, French N. Risk factors for neonatal calf diarrhoea and enteropathogen shedding in New Zealand dairy farms. Vet J. (2015) 203:155–60. doi: 10.1016/j.tvjl.2015.01.010
9. Roy SS, Pramanik AK, Batabyal S, Sarkar S, Das P. Cryptosporidiosis an important zoonotic disease: a review article. Intas Polivet. (2006) 11:681–6. doi: 10.14202/vetworld.2018.681-686
10. Bossi P. Resistance to antibiotics and the prudent use of antibiotics in veterinary medicine. Equine Vet Educ. (2000) 12:108–12. doi: 10.1111/j.2042-3292.2000.tb01773.x
11. Tiangtip R, Jongwutiwes S. Molecular analysis of Cryptosporidium species isolated from HIV-infected patients in Thailand. Trop Med Int Health. (2002) 7:357–64. doi: 10.1046/j.1365-3156.2002.00855.x
12. Hopkins RM, Meloni BP, Groth DM, Wetherall JD, Thompson JaRCA. Ribosomal RNA sequencing reveals differences between the genotypes of Giardia isolates recovered from humans and dogs living in the same locality. J Parasitol. (1997) 83:44–51. doi: 10.2307/3284315
13. Appelbee AJ, Frederick LM, Heitman TL, Olson ME. Prevalence and genotyping of Giardia duodenalis from beef calves in Alberta, Canada. Vet Parasitol. (2003) 112:289–94. doi: 10.1016/S0304-4017(02)00422-3
14. Mahbubani MH, Bej AK, Perlin MH, Schaefer FW, Jakubowski W. Differentiation of Giardia duodenalis from other Giardia spp. by using polymerase chain reaction and gene probes. J Clin Microbiol. (1992) 30:74–8. doi: 10.1128/JCM.30.1.74-78.1992
15. Cacciò SM, Giacomo MD, Pozio E. Sequence analysis of the beta-giardin gene and development of a polymerase chain reaction-restriction fragment length polymorphism assay to genotype Giardia duodenalis cysts from human faecal samples. Int J Parasitol. (2002) 32:1023–30. doi: 10.1016/S0020-7519(02)00068-1
16. Read CM, Monis PT, Thompson RC. Discrimination of all genotypes of Giardia duodenalis at the glutamate dehydrogenase locus using PCR-RFLP. Infec Genet Evol. (2004) 4:125–30. doi: 10.1016/j.meegid.2004.02.001
17. Baker DA, Holberton DV, Marshall J. Sequence of a giardin subunit cDNA from G. lamblia. Nucl Acids Res. (1988) 16:7177. doi: 10.1093/nar/16.14.7177
18. Ewers C, Li G, Wilking H, Kieβling S, Alt K, Antáo E-M, et al. Avian pathogenic, uropathogenic, and newborn meningitis-causing Escherichia coli: how closely related are they? Int J Med Microbiol. (2007) 297:163–76. doi: 10.1016/j.ijmm.2007.01.003
19. Luan JJ, Yang SH, Zhang WJ, Gao YD, Zhong JF, Zhao HK. Rapid detection of bovine rotavirus with semi-nested RT-PCR assay. Chinese J Zoonoses. (2006) 22:671–3.
20. Cui X. Epidemiological Survey of Rotavirus and Coronavirus in Some Large-Scale Dairy Farms in Southern Xinjiang. (2019). (Master dissertation). Tarim: Tarim University.
21. Klein-Jobstl D, Iwersen M, Drillich M. Farm characteristics and calf management practices on dairy farms with and without diarrhea: a case-control study to investigate risk factors for calf diarrhea. J Dairy Sci. (2014) 97:5110–9. doi: 10.3168/jds.2013-7695
22. Frank NA, Kaneene JB. Management risk factors associated with calf diarrhea in michigan dairy herds. J Dairy Sci. (1993) 76:1313–23. doi: 10.3168/jds.S0022-0302(93)77462-7
23. Pelan-Mattocks LS, Kehrli ME, Casey TA, Goff JP. Fecal shedding of coliform bacteria during the periparturient period in dairy cows. Am J Vet Res. (2000) 61:1636–8. doi: 10.2460/ajvr.2000.61.1636
24. Gutzwiller A. Effect of colostrum intake on diarrhoea incidence in new-born calves. Schweiz Arch Tierheilkd. (2002) 144:59–64. doi: 10.1024/0036-7281.144.2.59
25. Barrington GM, Gay JM, Evermann JF. Biosecurity for neonatal gastrointestinal diseases. Vet Clin North Am Food Anim Pract. (2002) 18:7–34. doi: 10.1016/S0749-0720(02)00005-1
26. Nonnecke BJ, Foote MR, Smith JM, Pesch BA, Amburgh MEV. Composition and functional capacity of blood mononuclear leukocyte populations from neonatal calves on standard and intensified milk replacer diets. J Dairy Sci. (2003) 86:3592–604. doi: 10.3168/jds.S0022-0302(03)73965-4
27. Mcguirk SM. Disease management of dairy calves and heifers. Vet Clin North Am Food Anim Pract. (2008) 24:139–53. doi: 10.1016/j.cvfa.2007.10.003
28. Woolums AR, Berghaus RD, Smith DR, White BJ, Waggoner ER. Producer survey of herd-level risk factors for nursing beef calf respiratory disease. J Am Vet Med Asso. (2013) 243:538–47. doi: 10.2460/javma.243.4.538
29. Reiten M, Rousing T, Thomsen PT, Otten ND, Forkman B, Houe H, et al. Mortality, diarrhea and respiratory disease in Danish dairy heifer calves: effect of production system and season. Prev Vet Med. (2018) 155:21–6. doi: 10.1016/j.prevetmed.2018.04.007
30. Bartels CJM, Holzhauer M, Jorritsma R, Swart WaJM, Lam TJGM. Prevalence, prediction and risk factors of enteropathogens in normal and non-normal faeces of young Dutch dairy calves. Prev Vet Med. (2010) 93:162–9. doi: 10.1016/j.prevetmed.2009.09.020
31. Sischo WM, Atwill ER, Lanyon LE, George J. Cryptosporidia on dairy farms and the role these farms may have in contaminating surface water supplies in the northeastern United States. Pre Vet Med. (2000) 43:253–67. doi: 10.1016/S0167-5877(99)00107-5
32. Trotz-Williams LA, Wayne Martin S, Leslie KE, Duffield T, Nydam DV, Peregrine AS. Association between management practices and within-herd prevalence of C. parvum shedding on dairy farms in southern Ontario. Prev Vet Med. (2008) 83:11–23. doi: 10.1016/j.prevetmed.2007.03.001
33. Castrucci G, Ferrari M, Frigeri F, Traldi V, Angelillo V. A study on neonatal calf diarrhea induced by rotavirus. Comp Immunol Microbiol Infect Dis. (1994) 17:321–31. doi: 10.1016/0147-9571(94)90051-5
Keywords: neonatal calf diarrhea, etiological agents, China, cattle, Yangxin County
Citation: Wei X-j, Wang W-w, Dong Z, Cheng F-s, Zhou X-z, Li B and Zhang J-y (2021) Detection of Infectious Agents Causing Neonatal Calf Diarrhea on Two Large Dairy Farms in Yangxin County, Shandong Province, China. Front. Vet. Sci. 7:589126. doi: 10.3389/fvets.2020.589126
Received: 31 July 2020; Accepted: 31 December 2020;
Published: 05 February 2021.
Edited by:
Beate Pinior, University of Veterinary Medicine Vienna, AustriaReviewed by:
László Ózsvári, University of Veterinary Medicine Budapest, HungaryMonika Dzieciol, University of Veterinary Medicine Vienna, Austria
Copyright © 2021 Wei, Wang, Dong, Cheng, Zhou, Li and Zhang. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Jiyu Zhang, aW5memp5QHNpbmEuY29t
†These authors have contributed equally to this work
 Xiaojuan Wei
Xiaojuan Wei Weiwei Wang1,2,3
Weiwei Wang1,2,3 Zhen Dong
Zhen Dong Jiyu Zhang
Jiyu Zhang