- 1Bacteriology, Animal and Plant Health Agency Weybridge, Addlestone, United Kingdom
- 2Department of Zoology, University of Oxford, Oxford, United Kingdom
- 3NIHR Health Protection Research Unit in Gastrointestinal Infections, University of Oxford, Oxford, United Kingdom
- 4Veterinary Surveillance, Animal and Plant Health Agency Weybridge, Addlestone, United Kingdom
- 5Department of Biology and Biotechnology, The Milner Centre for Evolution, University of Bath, Bath, United Kingdom
A corrigendum on
Genome Reduction for Niche Association in Campylobacter Hepaticus, A Cause of Spotty Liver Disease in Poultry
by Petrovska, L., Tang, Y., Jansen van Rensburg, M. J., Cawthraw, S., Nunez, J., Sheppard, S. K., et al. (2017). Front. Cell. Infect. Microbiol. 7:354. doi: 10.3389/fcimb.2017.00354
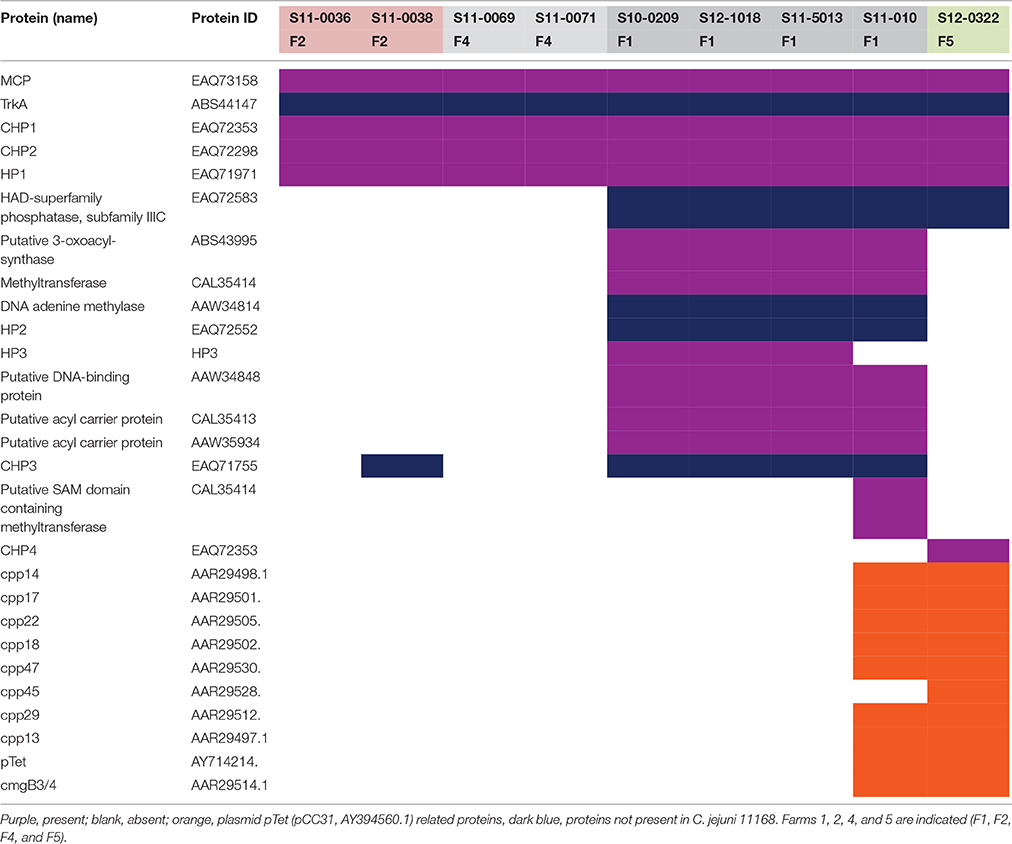
In the original article, there was a mistake in Table 3 as published. Table 3 had additional genes inserted for isolates S11-0036, S11-0038, S11-0069, and S12-0071. Isolate S12-002 should not be included in Table 3.
Additionally, there was an incorrect sentence. Incorrect sentence describing the number of RNA coding sequences and the GC content. A correction has been made to Results, C. hepaticus Isolates Have Reduced Genomes, Paragraph Number One and appears below.
The C. hepaticus isolates had a lower number (average of 44) of RNA coding sequences and a lower GC content (average of 28.4%) in comparison to the C. jejuni reference genomes (average of 52.4 and 30.5%, respectively).
Similarly, there was an incorrect sentence. Incorrect sentence describing the genes related to pathogenicity of C. hepaticus. A correction has been made to Results, genes related to the Pathogenicity of C. hepaticus, Paragraph Number One and appears below.
The UK C. hepaticus isolates contained relatively few genes linked to pathogenesis: 5 were identified in the genomes of S11-0036, S11-0069, S11-0071 and (from farms 2, and 4); 6 in S11-0038 (farm 2); 15 in S10-0209, S12-1018, S11-5013, and S11-010, (farm 1); and 7 in isolate S12-0322 (farm 5; Table 3). The cpp and cmgB3/4 genes, both components of the pTet plasmid (Batchelor et al., 2004), and a complete pTet plasmid (Batchelor et al., 2004) sequences were identified in isolates S11-010, and S12-0322 (Table 3).
Finally, in incorrect spelling of metabolism was used, we omitted “the” and misspelled “rich.” A correction has been made to Discussion, Paragraph Number Four and appears below.
Furthermore, Stahl and co-workers found that the ability to metabolize L-fucose in vivo provided C. jejuni with competitive advantage during colonization of the piglet infection model. Similar was not observed in the chick commensal model (Stahl et al., 2011), suggesting potential niche specific advantage for colonization in the L-fucose rich environment in the pig small intestine and cecum.
The authors apologize for these errors and state that this does not change the scientific conclusions of the article in any way.
Conflict of Interest Statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Keywords: Campylobacter hepaticus, spotty liver disease, poultry, genome reduction, niche adaptation
Citation: Petrovska L, Tang Y, Jansen van Rensburg MJ, Cawthraw S, Nunez J, Sheppard SK, Ellis RJ, Whatmore AM, Crawshaw TR and Irvine RM (2017) Corrigendum: Genome Reduction for Niche Association in Campylobacter Hepaticus, A Cause of Spotty Liver Disease in Poultry. Front. Cell. Infect. Microbiol. 7:480. doi: 10.3389/fcimb.2017.00480
Received: 21 August 2017; Accepted: 03 November 2017;
Published: 14 November 2017.
Edited and reviewed by: Alain Stintzi, University of Ottawa, Canada
Copyright © 2017 Petrovska, Tang, Jansen van Rensburg, Cawthraw, Nunez, Sheppard, Ellis, Whatmore, Crawshaw and Irvine. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) or licensor are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Liljana Petrovska, bGlsamFuYS5wZXRyb3Zza2FAYXBoYS5nc2kuZ292LnVr
†These authors have contributed equally to this work.
 Liljana Petrovska
Liljana Petrovska Yue Tang1†
Yue Tang1† Melissa J. Jansen van Rensburg
Melissa J. Jansen van Rensburg Shaun Cawthraw
Shaun Cawthraw Richard J. Ellis
Richard J. Ellis Adrian M. Whatmore
Adrian M. Whatmore Tim R. Crawshaw
Tim R. Crawshaw