- 1Master of Science Program in Applied Microbiology (International Program), Faculty of Science, Chiang Mai University, Chiang Mai, Thailand
- 2Department of Biology, Faculty of Science, Chiang Mai University, Chiang Mai, Thailand
- 3Center of Excellence in Microbial Diversity and Sustainable Utilization, Chiang Mai University, Chiang Mai, Thailand
- 4Academy of Science, The Royal Society of Thailand, Bangkok, Thailand
Members of the family Herpotrichiellaceae are distributed worldwide and can be found in various habitats including on insects, plants, rocks, and in the soil. They are also known to be opportunistic human pathogens. In this study, 12 strains of rock-inhabiting fungi that belong to Herpotrichiellaceae were isolated from rock samples collected from forests located in Lamphun and Sukhothai provinces of northern Thailand during the period from 2021 to 2022. On the basis of the morphological characteristics, growth temperature, and multi-gene phylogenetic analyses of a combination of the internal transcribed spacer, the large subunit, and the small subunit of ribosomal RNA, beta tubulin and the translation elongation factor 1-a genes, the new genus, Petriomyces gen. nov., has been established to accommodate the single species, Pe. obovoidisporus sp. nov. In addition, three new species of Cladophialophora have also been introduced, namely, Cl. rupestricola, Cl. sribuabanensis, and Cl. thailandensis. Descriptions, illustrations, and a phylogenetic trees indicating the placement of these new taxa are provided. Here, we provide updates and discussions on the phylogenetic placement of other fungal genera within Herpotrichiellaceae.
Introduction
Herpotrichiellaceae is a family within order Chaetothyriales (class Eurotiomycetes), which was introduced by Munk (1953) to accommodate Herpotrichiella, with H. moravica as the type species. Accordingly, Herpotrichiella moravica is now considered a synonym of Capronia moravica based on the teleomorph–anamorph connection of Herpotrichiellaceae (Müller et al., 1987). Likewise, the morphology of Capronia moravica and Ca. pilosella has been found to be identical; thus, Ca. moravica is now synonymized under Ca. pilosella (Untereiner, 1997). Capronia is the older name that was introduced by Saccardo (1883); therefore, Capronia has been retained over Herpotrichiella (Untereiner, 1997; Quan et al., 2020). Herpotrichiellaceae is characterized by having setose, ostiolate ascomata, and bitunicate asci with endotunica; by appearing greenish gray to brown; and by having didymosporous, phragmosporous, or dictyosporous ascospores (Müller et al., 1987; Untereiner et al., 1995; Untereiner, 1997; Tian et al., 2021). The asexual morphs of the family are characterized as dematiaceous hyphomycetes, and commonly referred to as black yeast (Müller et al., 1987; Untereiner et al., 1995; Tian et al., 2021). This family is well-known to consist of a sexual morph as a genus Capronia and several asexual morph genera, such as Cladophialophora, Exophiala, Fonsecaea, Phialophora, and Rhinocladiella (de Hoog et al., 2011; Phukhamsakda et al., 2022). Due to the similar morphological features of these asexual genera, the morphological identification of these asexual genera can be challenging (Badali et al., 2008; Yang et al., 2021). Therefore, a combination of morphological and molecular data is needed to identify the species in this family and to clarify their phylogenetic placement. Twenty genera are currently accepted in Herpotrichiellaceae; namely, Aculeata, Atrokylindriopsis, Brycekendrickomyces, Capronia, Cladophialophora, Exophiala, Fonsecaea, Marinophialophora, Melanoctona, Metulocladosporiella, Minimelanolocus, Neosorocybe, Phialophora, Pleomelogramma, Rhinocladiella, Sorocybe, Thysanorea, Uncispora, Valentiella, and Veronaea (Wijayawardene et al., 2020; Tian et al., 2021; Bezerra et al., 2022; Wijayawardene et al., 2022). The members of Herpotrichiellaceae have been collected from several sources worldwide. These consist of air, insects, lichens, plants, rocks, and soil (Diederich et al., 2013; Feng et al., 2014; Das et al., 2019; Libert et al., 2019; Lima et al., 2020; Sun et al., 2020; Thitla et al., 2022). In addition, numerous studies have reported that some herpotrichiellaceous fungi are opportunistic fungi in humans and other animals (Badali et al., 2008; de Hoog et al., 2011; Li et al., 2017). Several herpotrichiellaceous fungi were reported to survive under extreme conditions (high and low temperatures, desiccation, lack of nutrients, and UV radiation), such as the species within genera Exophiala and Cladophialophora (Seyedmousavi et al., 2011; Gostinčar et al., 2022; Tesei, 2022). The accumulation of melanin in the cell walls of these fungi is one of the key factors to survival in extreme conditions (de Hoog et al., 2011; Li et al., 2017; Coleine and Selbmann, 2021).
Cladophialophora is one of the genera in the family Herpotrichiellaceae and has been reported in many regions of the world including Africa, Asia, Europe, North America, Oceania, and South America (de Hoog et al., 1995; de Hoog et al., 2007; Badali et al., 2008; Badali et al., 2011; Madrid et al., 2016; Sun et al., 2020; Nascimento et al., 2021). The genus was introduced by Borelli (1980) with Cl. ajelloi Borelli as the type species [currently named Cl. carrionii (Badali et al., 2008; Boonmee et al., 2021). Presently, there are 53 records of Cladophialophora in the Index Fungorum database (Index Fungorum, 2023). Of those, some species may belong to different genera. For example, Cl. brevicatenata, Cl. hachijoensis, and Cl. kellermaniana were subsequently transferred to Tyrannosorus hanlinianus, Pseudocladosporium hachijoense, and Chalastospora gossypii, respectively (Braun, 1998; Crous et al., 2009; Shen et al., 2020). However, 49 Cladophialophora species have currently been validated and accepted.
Rocks are one of the interesting natural habitats of fungi. Rock-inhabiting fungi were divided into two groups on the basis of their ecology and taxonomy. The first group comprises hyphomycetes of soil and epiphytic fungi (such as Aureobasidium or Phoma), whereas the second group includes melanized cell-walled fungi that exhibit slow growth, meristematic growth, or the production of yeast-like cells, typically belonging to the orders Capnodoales, Chaetothyriales, and Dothideales (Wollenzien et al., 1995; Gorbushina, 2007; Coleine and Selbmann, 2021). There are numerous reports on the biotechnological capabilities of rock-inhabiting fungi in astrobiology, radioprotection, biomedical, and bioremediation fields (Aureli et al., 2020; Lin and Xu, 2020; Tran-Ly et al., 2020; Coleine and Selbmann, 2021; Mattoon et al., 2021; Cassaro et al., 2022; Liu et al., 2022). Several species of herpotrichiellaceous fungi were previously isolated from rocks, such as Cladophialophora nyingchiensis, Cl. tumulicola, Exophiala bonariae, Ex. clavispora, and Ex. siamensis (Isola et al., 2016; Kiyuna et al., 2018; Sun et al., 2020; Thitla et al., 2022). In addition, numerous prior reports have highlighted Thailand as a hot spot for discovering new fungal species (Hyde et al., 2018; Khuna et al., 2022). Nonetheless, information on the rock-inhabiting fungi in Thailand is still limited. Thus, the main objective of this study is to study the diversity of rock-inhabiting fungi in Thailand. During our investigation, we identified 12 herpotrichiellaceous fungi, viz., 10 strains of Cladophialophora and two strains of unrecognized fungal taxa. Morphology, growth temperature, and multi-gene phylogenetic analyses indicate that four herpotrichiellaceous fungi are novel in Herpotrichiellaceae. In addition, we have updated the existing reference data on the members of Herpotrichiellaceae.
Materials and methods
Sample collection and fungal isolation
Rock samples appearing black fungal mycelia were collected from a dipterocarp forests in Lamphun (18°32′13″N 99°07′31″E, elevation at 432 m; and 18°32′11″N 99°07′22″E, elevation at 444 m) and Sukhothai (17°32′58″N 99°29′49″E, elevation at 153 m) provinces, Thailand, in 2021−2022. During the collection period, Lamphun province had daily rainfall of 6.5 mm, whereas Sukhothai province received daily rainfall of 2.3 mm. Temperatures in Lamphun province ranged from 22°C to 36°C, whereas temperatures in Sukhothai province ranged from 24°C to 38°C. Rock samples were obtained following the method described by Thitla et al. (2022). Fungi were isolated using an adaptation of the technique reported by Selbmann et al. (2014). Rock samples were cleaned in 1% sodium hypochlorite for 10 min before being rinsed five times with sterile and deionized water to eliminate any trace of detergent. To obtain the fungal strain, pounding rock samples and seeding rock shards were sprinkled onto malt extract agar (MEA; Gibco, Life Technologies Corporation, USA) and dichloran-rose bengal agar (DRBC; Difco, Becton, Dickinson and Company, USA) supplemented with chloramphenicol 100 mg/L. Plates were incubated at 25°C for 4 weeks, with daily inspections. Dark mycelia fungi were aseptically transferred to a MEA plates. Pure fungal strains were kept in 15% glycerol and deposited in the Culture Collection of Sustainable Development of Biological Resources Laboratory (SDBR), Faculty of Science, Chiang Mai University, Chiang Mai, Thailand. In addition, new fungal taxa were registered in the MycoBank database (MycoBank, 2023).
Morphology and growth temperature
Macro-morphologies, including colony characterization, pigment production, and colony diameter, were investigated on potato dextrose agar (PDA; Condalab, Laboratorios Conda S.A., Spain), MEA, and oatmeal agar (OA; Difco, Becton, Dickinson and Company, USA) at 25°C in the dark for 28 days. To examine the fungal growth at different temperatures, the colony diameter was measured on MEA at 4°C, 10°C, 15°C, 20°C, 25°C, 28°C, 30°C, 35°C, 37°C, and 40°C for 4 weeks in darkness. Three replicates were performed for each fungal strain at each temperature. Micromorphological features were investigated using a light microscope (Nikon Eclipse Ni-U, Japan). The Tarosoft (R) Image was used to calculate the size of fungal structures (e.g., hyphae, conidiophore, conidiogenous cell, conidia, and chlamydospore).
DNA extraction, amplification, and sequencing
Fungal genomic DNA of each fungal strain was extracted from mycelia grown on MEA at 25°C for 3 weeks, using s DNA extraction kit (FAVORGEN, Ping-Tung, Taiwan). The internal transcribed spacer (ITS), large subunit (nrLSU), and small subunit (nrSSU) of ribosomal RNA, beta tubulin gene (tub2), and the translation elongation factor 1-α (tef1-α) genes were amplified by polymerase chain reaction (PCR) using ITS5/ITS4 (White et al., 1990), LROR/LR5 (Vilgalys and Hester, 1990; Rehner and Samuels, 1994), NS1/NS4 (White et al., 1990), Bt2a/Bt2b (Glass and Donaldson, 1995), and EF1-728F/EF1-986R (Carbone and Kohn, 1999) primers, respectively. PCR amplifications were performed using 20 µL of reaction mixtures, consisting of 1 µL of genomic DNA, 1 µL of each primer, 10 µL of Quick TaqTM HS Dye-Mix (TOYOBO, Japan), and 7 µL of deionized water. The conditions of PCR reactions consisted of a first denaturation step performed at 95°C for 5 min and denaturation step at 95°C for 30 s; then, to amplify the ITS region, an annealing step was performed at 55°C for 30 s, an annealing step of nrSSU and nrLSU region was performed at 52°C for 45 s, whereas annealing step of the tub2 gene was performed at 52°C for 30 s; an elongation step were performed at 72°C for 1 min; lastly to amplify the tef1-α gene, annealing step at 57°C for 1 min and an elongation step at 72°C for 1.30 min were was performed. Cycles were performed 35 times, with a final extension at 72°C for 10 min on a peqSTAR thermal cycler (PEQLAB Ltd., Fareham, UK). PCR products were checked on 1% agarose gel electrophoresis and measured quantity with NanoDrop OneC (Thermo Scientific, USA). Then, the PCR clean-up Gel Extraction NucleoSpin® Gel and the PCR Clean-up Kit (Macherey-Nagel, Düren, Germany) were used to purify PCR products. Purified PCR products were sequenced by 1st BASE Company (Kembangan, Malaysia).
Sequence alignment and phylogenetic analyses
The ITS, nrLSU, nrSSU, tub2, and tef1-α sequence data were assembled using the software Sequencher 5.4.6 (Nishimura, 2000). The consensus sequences were blasted in the BLAST search tool via NCBI website. Two datasets were prepared to construct the phylogenetic trees for clarifying the family Herpotrichiellaceae (Table 1) and genus Cladophialophora (Table 2). Multiple sequence alignment was performed by MUSCLE using MEGA 6 (Edgar, 2004) and adjusted manually in BioEdit v.7.2.5 (Hall, 2004). Phylogenetic analysis of the family Herpotrichiellaceae (analysis I) was carried out on the basis of only the ITS, nrLSU, and SSU sequences because the amount of available sequence data in the tub2 and tef1-α genes is practically limited. To construct a phylogenetic tree of Cladophialophora (analysis II), five gene datasets (ITS, nrLSU, nrSSU, tub2, and tef1-α) were used. Maximum likelihood (ML) analysis was generated with 25 categories and 1,000 bootstrap (BS) replications under the GTRCAT model using the RAxML-HPC2 on XSEDE (v.8.2.12) in the CIPRES web portal (Felsenstein, 1985; Stamatakis, 2006; Miller et al., 2009). Bayesian inference (BI) analysis was performed using MrBayes v.3.2.6 (Ronquist and Huelsenbeck, 2003). Bayesian posterior probability (PP) was determined by Markov chain Monte Carlo (MCMC) sampling. Six simultaneous Markov chains were run for 5 million generations for analysis I and 2 million generations for analysis II with random initial trees, wherein every 100th generations were sampled. The first 20% of generated trees representing the burn-in phase of the analysis were discarded, whereas the remaining trees were used for calculating PP in the majority-rule consensus tree. Branches with BS support and PP values of more than or equal to 75% and 0.95, respectively, were deemed to have been substantially supported. The tree topologies were visualized in FigTree v1.4.0 (Rambaut, 2019).
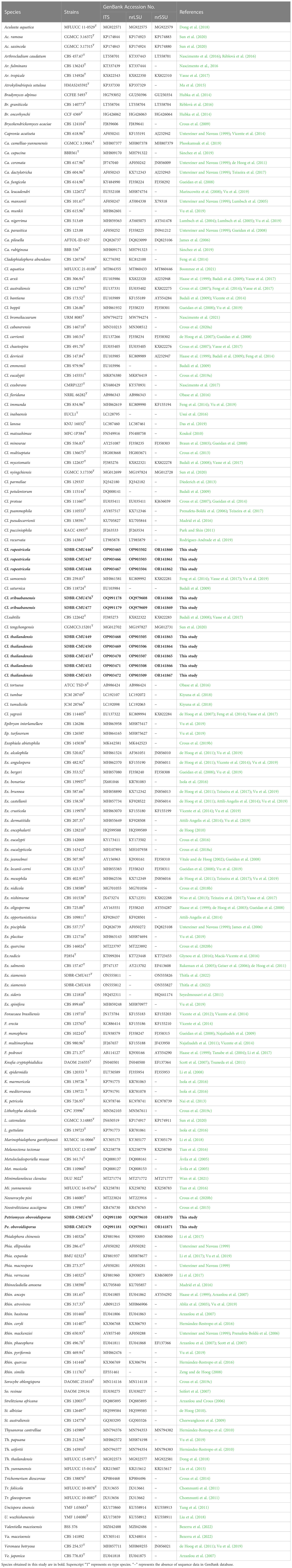
Table 1 GenBank accession numbers of herpotrichiellaceous fungi used in the molecular phylogenetic analysis.
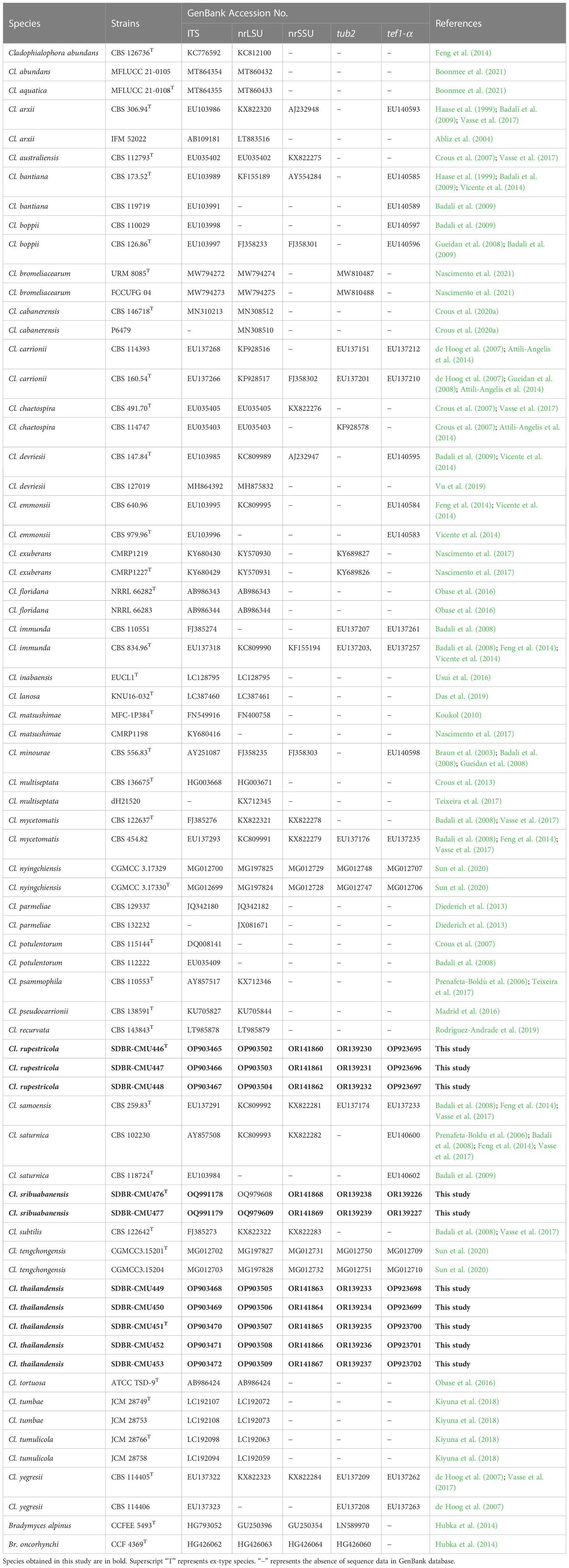
Table 2 GenBank accession numbers of Cladophialophora in the family Herpotrichiellaceae used in the molecular phylogenetic analysis.
Results
Fungal isolation, morphological study, and growth temperature
A total of 12 fungal strains were obtained from different rock samples. Ten fungal strains (SDBR-CMU446, SDBR-CMU447, SDBR-CMU448, SDBR-CMU449, SDBR-CMU450, SDBR-CMU451, SDBR-CMU452, SDBR-CMU453, SDBR-CMU476, and SDBR-CMU477) exhibited similar characteristics by appearing one-celled and hyaline and by forming conidial chains. Initially, these 10 fungal strains were identified as Cladophialophora species according to their micromorphological features. However, they were divided into three different groups on the basis of their colony characteristics on culture media, micromorphological, and growth temperature profiles (Table 3). In addition, two fungal strains (SDBR-CMU478 and SDBR-CMU479) appeared similar in their characteristics with produced sympodial conidial formation on conidiogenous loci and by appearing one-celled, hyaline to subhyaline, and obovoidal conidia. On the basis of morphological characteristics, these two fungal strains could not be assigned to any genera. Therefore, multi-gene phylogenetic analyses were used to identify their species-level and phylogenetic placement.
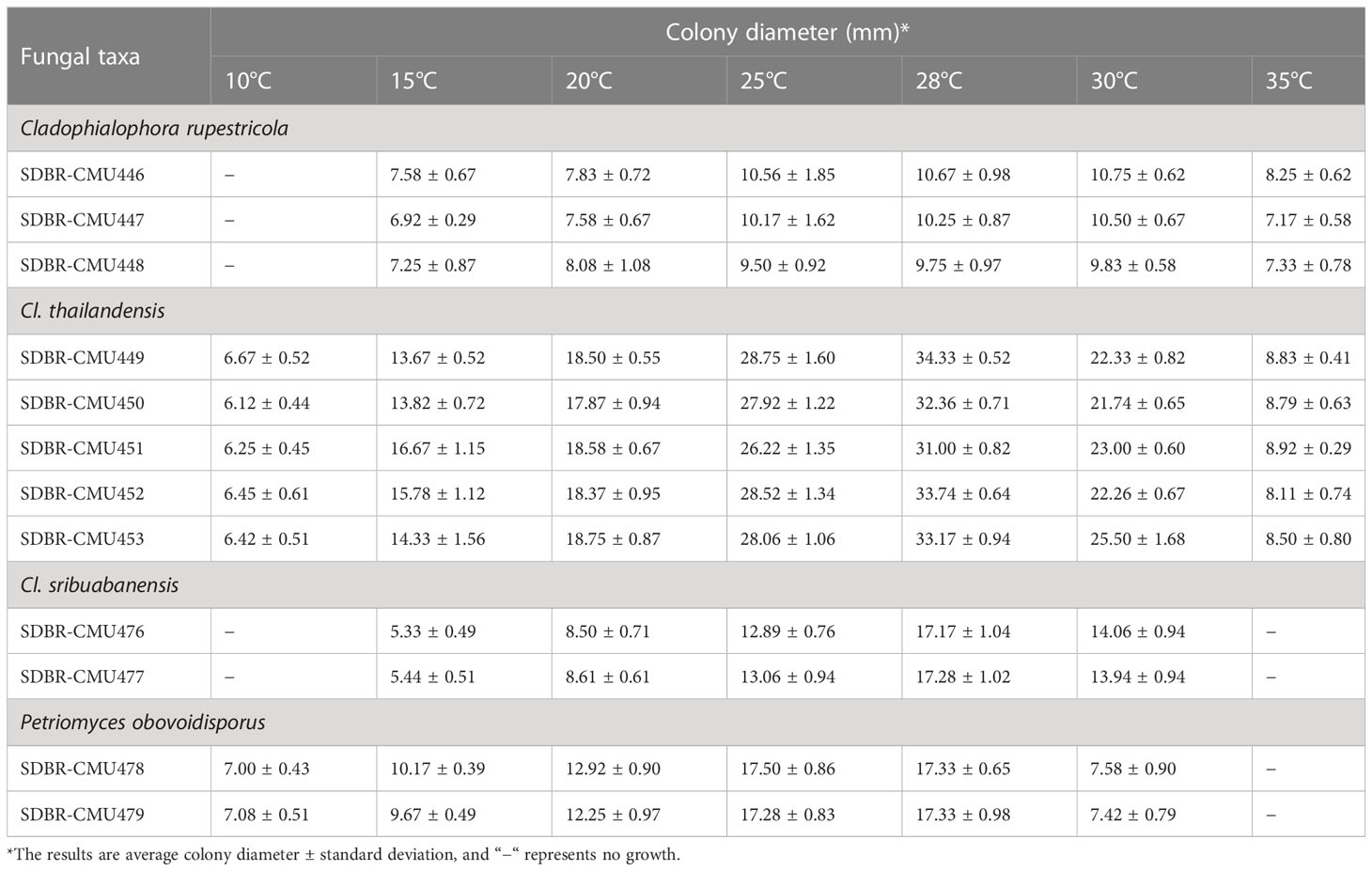
Table 3 Colony diameter of obtained fungi cultured on MEA at various temperatures for 28 days in the dark.
The observation of fungal growth at various temperatures (4°C−40°C) revealed that temperature had a strong influence on the fungal growth of the obtained fungi. The average colony diameter of each fungal strain is presented in Table 3. The results indicate that all 12 fungal strains could not grow at 4°C, 37°C, and 40°C. Three strains (SDBR-CMU446, SDBR-CMU447, and SDBR-CMU448) could not grow at 10°C but grew at temperatures ranging from 25°C to 30°C. Five strains (SDBR-CMU449, SDBR-CMU450, SDBR-CMU451, SDBR-CMU452, and SDBR-CMU453) exhibited the highest average colony diameter at 28°C. Two strains (SDBR-CMU476 and SDBR-CMU477) could not grow at 10°C and 35°C, yet they showed the greatest average colony diameter at 28°C. Two fungal strains (SDBR-CMU478 and SDBR-CMU479) grew at temperatures ranging from 10°C to 30°C and grew well at 28°C.
Phylogenetic study
For analysis I, a phylogenetic tree of the family Herpotrichiellaceae was constructed from a combined ITS, nrLSU, and nrSSU sequence dataset. The dataset comprised 159 sequence strains from representatives in the families Herpotrichiellaceae and Trichomeriaceae, including the new strains that were proposed in this study. Epibryon interlamellare CBS 126286 and Ep. turfosorum CBS 126587 (famliy Epibryaceae) were selected as the outgroup. The concatenated dataset comprised 3,022 positions (ITS, 1−928 base pair (bp); nrLSU, 929−1,842 bp; and nrSSU, 1,843−3,022 bp) including gaps. RAxML analysis of the integrated dataset yielded the best scoring tree with a final ML optimization likelihood value of −45,154.8753. The matrix contained 1,610 distinct alignment patterns with 38.67% undetermined characters or gaps. The estimated base frequencies were recorded as follows: A = 0.2716, C = 0.2009, G = 0.2559, and T = 0.2714; whereas the substitution rates were established as AC = 0.9883, AG = 1.7181, AT = 0.9613, CG = 0.7406, CT = 2.9999, and GT = 1.0000. The gamma distribution shape parameter alpha value was equal to 0.1267, whereas the tree length was equal to 17.4438. At the end of the total MCMC generations, the final average standard deviation of the split frequencies was calculated to be 0.009866 through BI analysis. According to the topological results, ML and BI phylogenetic analyses produced similar topologies. Therefore, only the phylogenetic tree constructed from the ML analysis is shown in Figure 1. The phylogenetic tree clearly separates the family Herpotrichiellaceae from the family Trichomeriaceae with strong support (100% BS and 1.00 PP). In this study, all fungal strains obtained belonged to the family Herpotrichiellaceae and were separated from the previously known species (Figure 1). Cladophiophora rupestricola (SDBR-CMU446, SDBR-CMU447, and SDBR-CMU448) forms a distinct lineage sister clade to Cl. sribuabanensis (SDBR-CMU476 and SDBR-CMU477) with 100% BS and 1.00 PP support values. Accordingly, five fungal strains (SDBR-CMU449, SDBR-CMU450, SDBR-CMU451, SDBR-CMU452, and SDBR-CMU453) of Cl. thailandensis form a distinct lineage closely related to Cl. inabaensis (EUCL1) and Cl. lanosa (KNU 16032). In addition, two fungal strains of Petriomyces obovoidisporus (SDBR-CMU478 and SDBR-CMU479) formed a well-resolved clade (100% BS and 1.00 PP; Figure 1) in Herpotrichiellaceae, with Atrokylindriopsis setulose (HMAS245592) and Exophiala siamensis (SDBR-CMU417) as the sister clade.
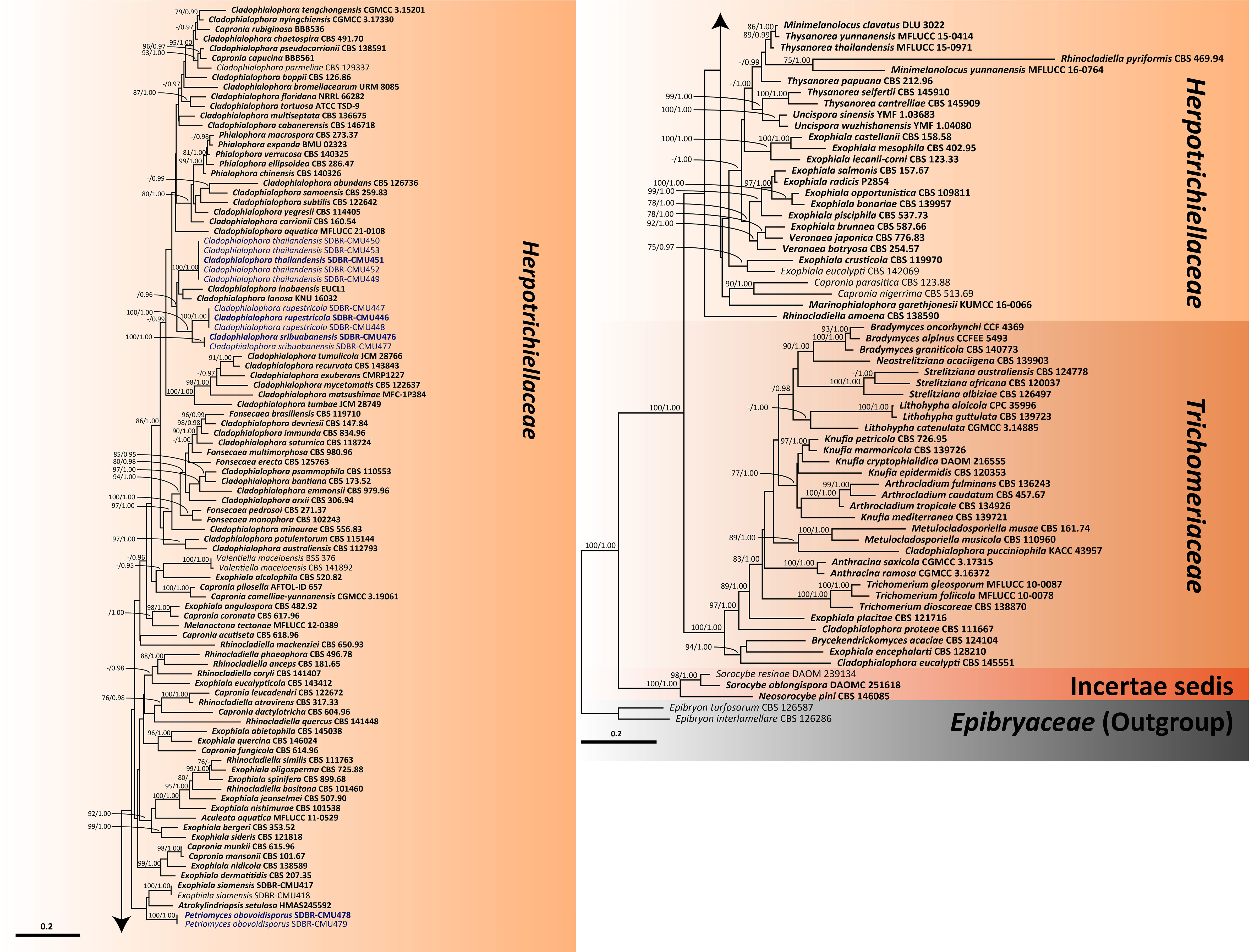
Figure 1 Phylogenetic tree generated from maximum likelihood analysis of 159 fungal strains based on a combined ITS, nrLSU, and nrSSU sequence dataset. Epibryon interlamellare CBS 126286 and Ep. turfosorum CBS 126587 were used as the outgroup. The numbers above branches show bootstrap percentages (left) and Bayesian posterior probabilities (right). Bootstrap values ≥75% and Bayesian posterior probabilities ≥0.95 are shown. The scale bar reflects the estimated number of nucleotide substitutions per site. The newly generated sequences are in blue. The ex-type species are in bold.
In our phylogenetic tree for clarifying the family Herpotrichiellaceae (Figure 1), 15 previously known herpotrichiellaceous genera (Aculeata, Atrokylindriopsis, Capronia, Cladophialophora, Exophiala, Fonsecaea, Marinophialophora, Melanoctona, Minimelanolocus, Phialophora, Rhinocladiella, Thysanorea, Uncispora, Valentiella, and Veronaea) and a new genus (Petriomyces) identified in this study were assigned to the Herpotrichiellaceae clade. The genera Brycekendrickomyces and Metulocladosporiella were placed in the Trichomeriaceae clade. The clade of genera Neosorocybe and Sorocybe was clearly separated from the clade of the families Herpotrichiellaceae and Trichomeriaceae with strong support of 100% BS and 1.00 PP (Figure 1).
For analysis II, the phylogenetic placement of the genus Cladophialophora within Herpotrichiellaceae was established by combining five gene sequence datasets (ITS, nrLSU, nrSSU, tub2, and tef1-α) from a total of 71 taxa. The concatenated dataset comprised 3,931 positions including gaps (ITS, 1−692 bp; nrLSU, 693−1,512 bp; nrSSU, 1,513−3,185 bp; tub2, 3,186−3,698; and tef1-α, 3,699−3,931 bp). RAxML analysis of the integrated dataset yielded the best scoring tree with a final ML optimization likelihood value of −21,115.8911. The matrix contained 1,300 distinct alignment patterns with 48.15% undetermined characters or gaps. The estimated base frequencies were recorded as follows: A = 0.2256, C = 0.2775, G = 0.2336, and T = 0.2632; whereas the substitution rates were established as AC = 1.0546, AG = 2.0969, AT = 0.9937, CG = 0.5923, CT = 3.1988, and GT = 1.0000. The gamma distribution shape parameter alpha value was equal to 0.4693, whereas the tree length was equal to 4.6134. The final average standard deviation of split frequencies at the end of total MCMC generations was calculated as 0.007796 through the BI analysis. The phylogram demonstrated that all three new species of Cladophialophora discovered in this study were distinctly separate from the previously known species. Cladophialophora rupestricola and Cl. sribuabanensis remained a sister group with strong support (100% BS and 1.00 PP), which was consistent with the findings of analysis I. Furthermore, Cl. thailandensis was a distinct lineage that is still related to Cl. inabaensis and Cl. lanosa.
Taxonomic descriptions
New genus
Petriomyces, T. Thitla and N. Suwannarach, gen. nov.
MycoBank number: MB849334
Etymology: “Petriomyces” refers to the fungus that dwells in rocks.
Mycelium: hyaline, smooth, branched, and septate hyphae. Conidiophores: semi-micronematous, arising vertically from creeping hyphae at right angles, straight, branched, subhyaline to pale brown, smooth, thin-walled, and septate. Conidiogenous cells: integrated, intercalary or terminal, sympodial, polyblastic, subconspicuous to conspicuous conidiogenous loci, subcylindrical, subdenticulate, smooth, and thin-walled. Conidia: obovoid or pyriform, aseptate, hyaline to subhyaline, smooth, and hilum conspicuous.
Habitat and distribution: sandstone on natural forest; known from Sukhothai province, Thailand.
Type species: Petriomyces obovoidisporus, T. Thitla, J. Kumla, and N. Suwannarach
Petriomyces obovoidisporus, T. Thitla, J. Kumla, and N. Suwannarach, sp. nov. (Figure 2)
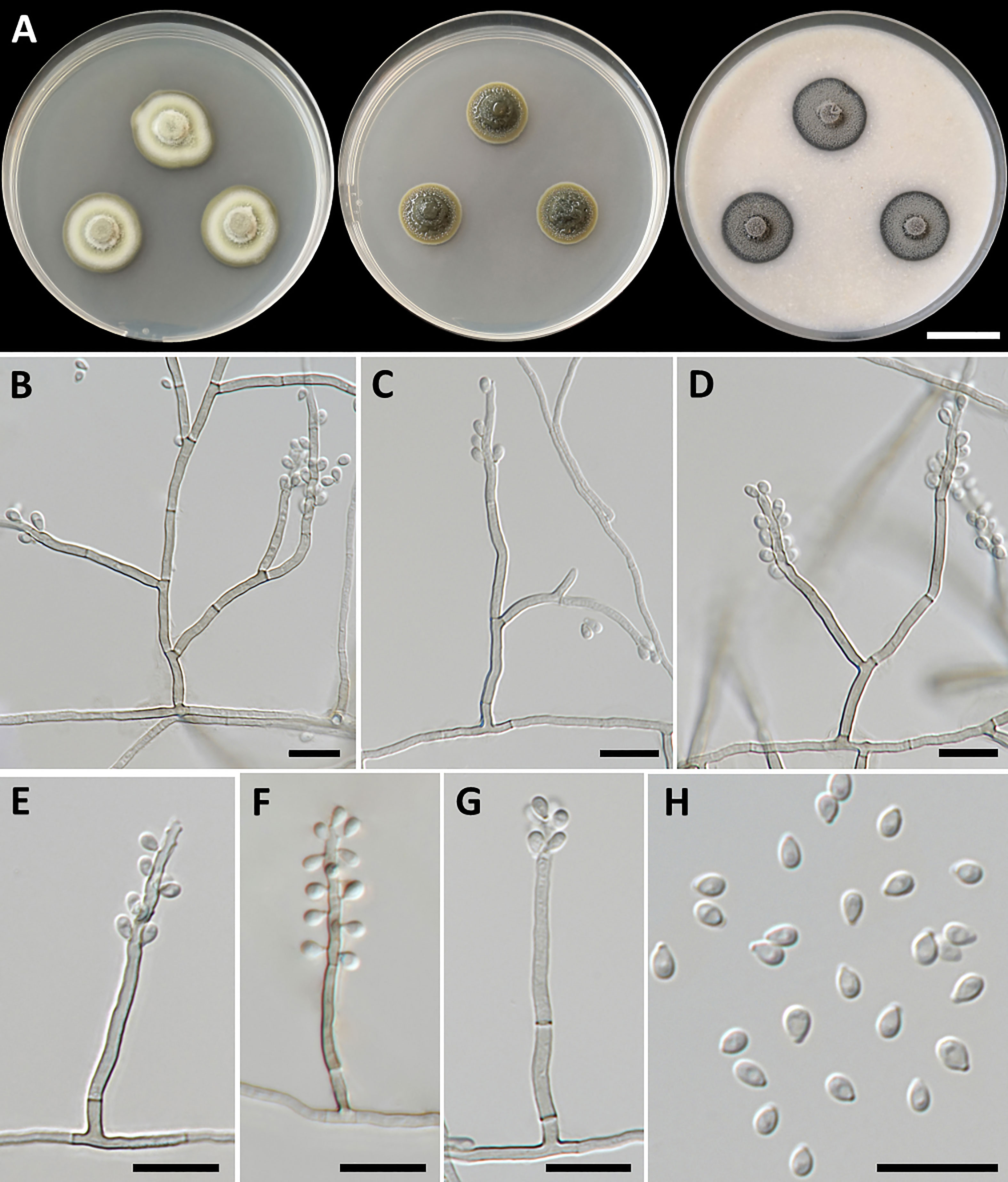
Figure 2 Petriomyces obovoidisporus (SDBR-CMU478, ex-type). (A) Colonies at 25°C for 28 days on PDA, MEA, and OA, respectively. (B–G) Conidiophore, conidiogenous cells, and conidia. (B–H) Conidia. Scale bars: (A) 2 cm and (B–G) 10 µm.
MycoBank number: MB849367
Etymology: “obovoidisporus” refers to the obovoid shape of the conidia.
Holotype: Thailand, Sukhothai province, Si Satchanalai District, 17°32′58″N 99°29′49″E, elevation at 153 m, isolated from the sandstone of natural forest, June 2021, T. Thitla; dried culture, SDBR-SKT3-6; ex-type culture, SDBR-CMU478.
GenBank number: OQ991180 (ITS), OQ979610 (nrLSU), OR141870 (nrSSU), OR139228 (tef1-α), and OR139240 (tub2)
Culture characteristics: Colonies on PDA at 25°C: attaining 20-mm to 25-mm diameter in 28 days, restricted, circular, flat, slimy, and grayish green to dull green with grayish green and entire margin; dark green in reverse; on MEA: attaining 16-mm to 20-mm diameter, restricted, circular, convex, slimy, and dark green with grayish green and entire margin; reverse grayish green to deep green; and on OA: attaining 18-mm to 22-mm diameter, restricted, circular, flat, velvety, and greenish gray with black and entire margin; reverse greenish gray. No diffusible pigment produced on any media. Submerged hyphae smooth, thin-walled, hyaline to light brown, 1-µm to 2-µm wide, and aerial hyphae subhyaline to pale brown with branched hyphae. Conidiophores: semi-micronematous, arising vertically from hyphae at right angles, straight, sometimes branched, thin-walled, subhyaline to pale brown, 1−3 septate, cylindrical, and 8−35.5 µm × 1−2 µm (mean = 18.3 µm × 1.5 µm, n = 30). Conidiogenous cell: integrated, intercalary or terminal, polyblastic, subconspicuous to conspicuous conidiogenous loci, subcylindrical, subdenticulate, smooth, thin-walled, cylindrical, slightly paler than conidiophore, and 13−65 µm × 1−2 µm (mean = 27.3 µm × 1.5 µm, n = 30). Conidia: one-celled, smooth, hyaline to subhyaline, obovoid or pyriform, 2−4 µm × 1−2 µm (mean = 2.8 µm × 1.7 µm, n = 50), and hilum conspicuous. Chlamydospores and sexual morph were not produced on media.
Growth temperature: minimum at 10°C, optimum at the range of 25°C−28°C, maximum at 30°C, and no growth at 4°C and 35°C
Additional specimens examined: Thailand, Sukhothai province, Si Satchanalai District, 17°32′58″N 99°29′49″E, elevation at 153 m, isolated from the rock of natural forest, June 2021, T. Thitla: SDBR-CMU479 [GenBank number: OQ991181 (ITS), OQ979611 (nrLSU), OR141871 (nrSSU),OR139229 (tef1-α), and OR139241 (tub2)].
Habitat and distribution: sandstone; collected from Sukhothai province, Thailand.
Notes: Some herpotrichiellacious fungi are known to be rock-inhabiting fungi, including Cladophialophora, Exophiala, Phialophora, and Rhinocladiella (Coleine and Selbmann, 2021; Liu et al., 2022). Cladophialophora is characterized by the production of branched or unbranched chains of one-cell conidia (Badali et al., 2008). Exophiala produced annellidic conidiogenesis and yeast-like states (de Hoog et al., 2011). Phialophora exhibited the basic morphological characteristics of conidial production through large phialidic conidiogenesis (Li et al., 2017). Rhinocladiella and Petriomyces share morphological characteristics of being polyblastic, as well as sympodial conidial formation on conidiogenous loci and aseptate conidia (Arzanlou et al., 2007). In addition, the conidia of Rhinocladiella appears in various shapes, including subglobose, ellipsoidal, obovoid, and subcylindrical to clavate, and is similar to Petriomyces, which appears obovoid or pyriform conidia. However, Rhinocladiella commonly produces thick-walled and brown conidiophores (Arzanlou et al., 2007), whereas Petriomyces has semi-micronematous, thin-walled, subhyaline, and pale brown conidiophores.
A multi-gene phylogenetic study revealed that Petriomyces formed a unique monophyletic clade in Herprotrichiellaceae with strong support values (100% BS and 1.00 PP) and formed a sister clade to Atrokylindriopsis setulose (HMAS245592) and Exophiala siamensis (SDBR-CMU417) (Figure 1). Atrokylindriopsis setulose was isolated from dead branches of an unidentified broadleaf tree and produced monophialidic conidiogenesis with setulate conidia (Ma et al., 2015). On the other hand, Petriomyces was isolated from rocks and produced a sympodial conidial formation with obovoidal conidia. Petriomyces obovoidisporus and Ex. siamensis were also isolated from rocks. However, Ex. siamensis produces short conidiophores, subspherical conidia, and pale brown chlamydospores, whereas Pe. obovoidisporus produces long conidiophores and obovoidal conidia and does not produce chlamydospores (Thitla et al., 2022). In addition, a pairwise nucleotide comparison of ITS and nrLSU sequencing data between Petriomyces obovoidisporus and Atrokylindriopsis setulose revealed different values of 12.5% (76/607 bp) and 2.7% (15/548 bp) including gaps. Notably, Pe. obovoidisporus and Ex. siamensis revealed pairwise nucleotide comparison values of 12.3% (75/609 bp) for ITS, 0.7% (7/1,007 bp) for nrSSU, 26.4% (58/220 bp) for tef1-α, and 25.8% (132/512 bp) for tub2 genes.
New species
Cladophialophora rupestricola, T. Thitla, J. Kumla, and N. Suwannarach, sp. nov. (Figure 3)
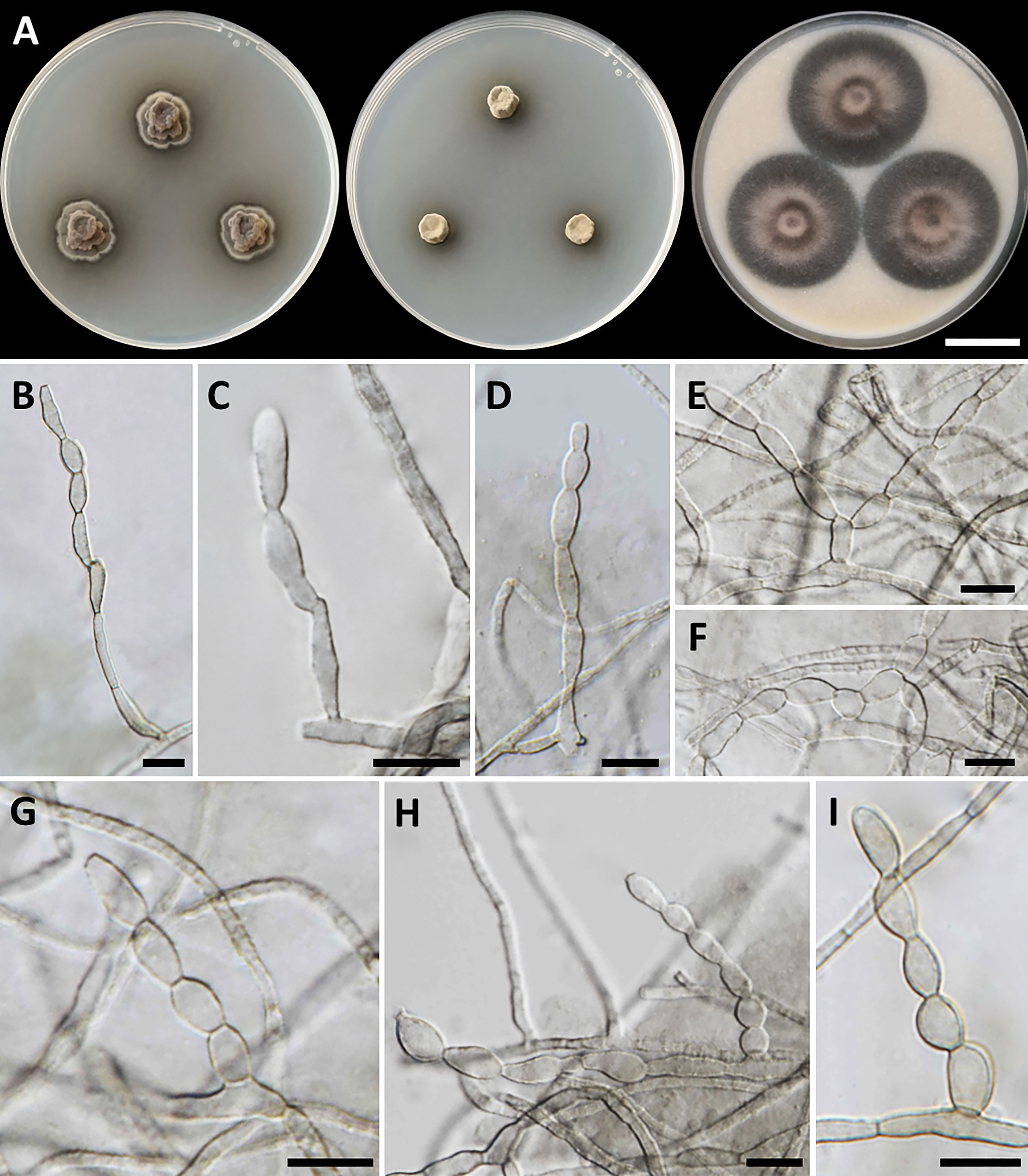
Figure 3 Cladophialophora rupestricola (SDBR-CMU446, ex-type). (A) Colonies at 25°C for 28 days on PDA, MEA, and OA, respectively. (B−D) Conidiophore and conidial chains. (E−I) Conidial chains on hyphae. Scale bars: (A) 2 cm and (B−I) 10 µm.
MycoBank number: MB846945
Etymology: “rupestricola” refers to the rock, where the habitat of the fungus.
Holotype: Thailand, Lamphun province, Mueang Lamphun District, Sribuaban Subdistrict, 18°32′11″N 99°07′22″E, elevation at 444 m, isolated from the sandstone of dipterocarp forest, July 2021, T. Thitla; dried culture, CMULPN5-5; ex-type culture, SDBR-CMU446.
GenBank number: OP903465 (ITS), OP903502 (nrLSU), OR141860 (nrSSU), OP923695 (tef1-α), and OR139230 (tub2)
Culture characteristics: Colonies on PDA, MEA, and OA were described after 28 days of incubation at 25°C in darkness: on PDA: attaining 15-mm to 17-mm diameter, restricted, circular, convex, velvety, gray in the center with black, and undulate margin; black in reverse; on MEA: attaining 8-mm to 13-mm diameter, restricted, circular, velvety, and dull green with dark green and entire margin; reverse dark green; on OA: attaining 36-mm to 40-mm diameter, restricted, circular, raised centrally, and gray aerial mycelium with greenish gray and entire margin; reverse dark green. A soluble olive and dull green pigment were observed around the fungal colonies on PDA and MEA, respectively. Hyphae smooth-walled, septate, hyaline to brown, 1.5-µm to 3-µm wide. Conidiophores reduced to conidiogenous cells, lateral or terminal on undifferentiated hyphae, smooth, hyaline to pale olivaceous, cylindrical, and 2.5-µm to 5-µm wide. Conidia spherical to ellipsoidal, aseptate, smooth, subhyaline to hyaline, forming branched acropetal chains at hyphae and conidiophore, 6−13.5 µm × 3−6 µm (mean = 9.5 µm × 4.6 µm, n = 50); Chlamydospores absent. Sexual morph was not produced on media.
Growth temperature: minimum at 15°C, optimum at the range of 25°C−30°C, maximum at 35°C, and no growth at 10°C and 37°C.
Additional specimens examined: Thailand, Lamphun province, Mueang Lamphun District, Sribuaban Subdistrict, 18°32′11″N 99°07′22″E, elevation at 444 m, isolated from the rock of dipterocarp forest, July 2021, isolated by T. Thitla: SDBR-CMU447 [GenBank number: OP903466 (ITS), OP903503 (nrLSU), OR141861 (nrSSU), OP923696 (tef1-α), and OR139231 (tub2)] and SDBR-CMU448 [GenBank: OP903467 (ITS), OP903504 (nrLSU), OR141862 (nrSSU), OP923697 (tef1-α), and OR139232 (tub2)].
Habitat and distribution: sandstone; collected from Lamphun province, Thailand.
Notes: The colony characteristics of Cl. rupestricola resembled those of Cl. chaetospira and Cl. floridana that were isolated from the decaying bamboo and sclerotia of Cenococcum geophilum, respectively. However, the conidial size of Cl. rupestricola differs from that of Cl. chaetospira and Cl. floridana. Specifically, the conidia of Cl. rupestricola (6−13.5 µm × 3−6 µm) were significantly shorter than that of Cl. chaetospira (20−45 µm × 3−5 µm) and slightly broader than that of Cl. floridana (3.5−8 µm × 2.0−3µm) (Crous et al., 2007; Obase et al., 2016). In addition, these three species also exhibit distinct conidiophore and conidial characteristics. Cladophialophora floridana displayed narrower conidiophores (2-µm to 3-µm wide) than Cl. repestricola, which produced broader cylindrical, 2.5-µm- to 5-µm-wide conidiophores (Obase et al., 2016). In terms of conidial characteristics, Cl. chaetospira produced 1−3 septate of conidia that is in contrast to Cl. repestricola, which produced aseptate conidia (Crous et al., 2007). The multi-gene phylogenetic analyses indicated that Cl. rupestricola is a distinct species in Cladophialophora and it is closely related to Cl. sribuabanensis (Figure 4). Cladophialophora rupestricola differs from Cl. sribuabanensis in terms of the temperature growth profile. Cladophialophora rupestricola has a maximum temperature of 35°C, whereas Cl. sribuabanensis has a maximum temperature of 30°C. In addition, Cl. rupestricola produced a soluble pigment on PDA and MEA, whereas Cl. sribuabanensis did not produce any pigment. Furthermore, a pairwise nucleotide comparison of Cl. rupestricola and Cl. sribuabanensis resulted in differences of 7.0% (45/642 bp) in ITS, 0.2% (2/903 bp) in nrLSU, 0.3% (3/1037 bp) in nrSSU, 9.8% (20/205 bp) in tef1-α, and 10.6% (54/509 bp) in tub2 sequences including gaps.
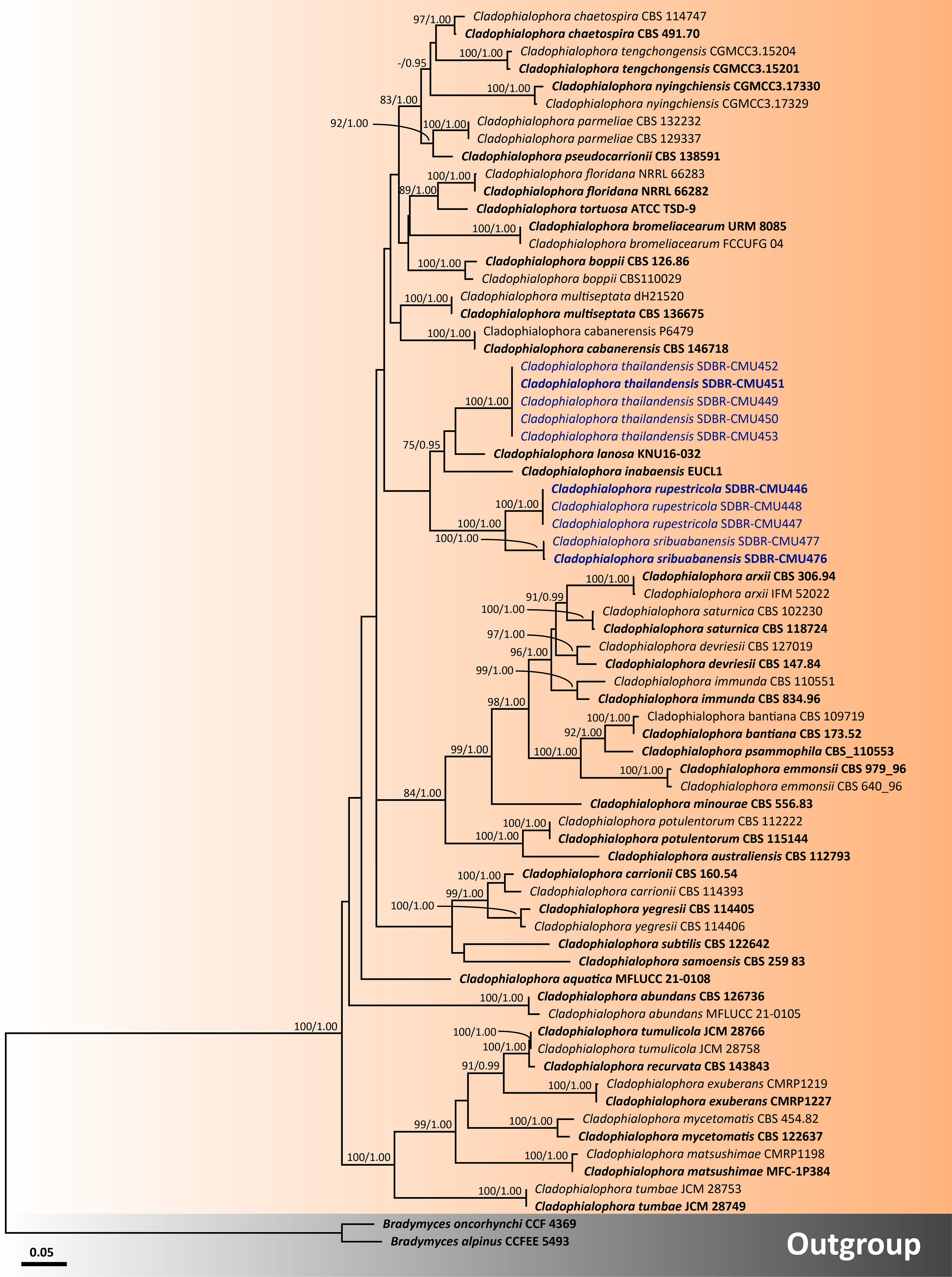
Figure 4 Phylogenetic relationships of Cladophialophora within the family Herpotrichiellaceae reconstructed by maximum likelihood analysis based on a combined dataset of ITS, nrLSU, nrSSU, tub2, and tef1-α genes. Bradymyces alpinus CCFEE 5493 and Br. oncorhynchi CCF 4369 were used as the outgroup. The values presented above branches represent the bootstrap percentages (left) and Bayesian posterior probabilities (right). Bootstrap values ≥75% and Bayesian posterior probabilities ≥0.95 are displayed. The scale bar indicates the estimated number of nucleotide substitutions per site. The newly generated sequences are in blue, whereas the ex-type species are indicated in bold.
Cladophialophora sribuabanensis, T. Thitla, N. Suwannarach, and S. Lumyong, sp. nov. (Figure 5).
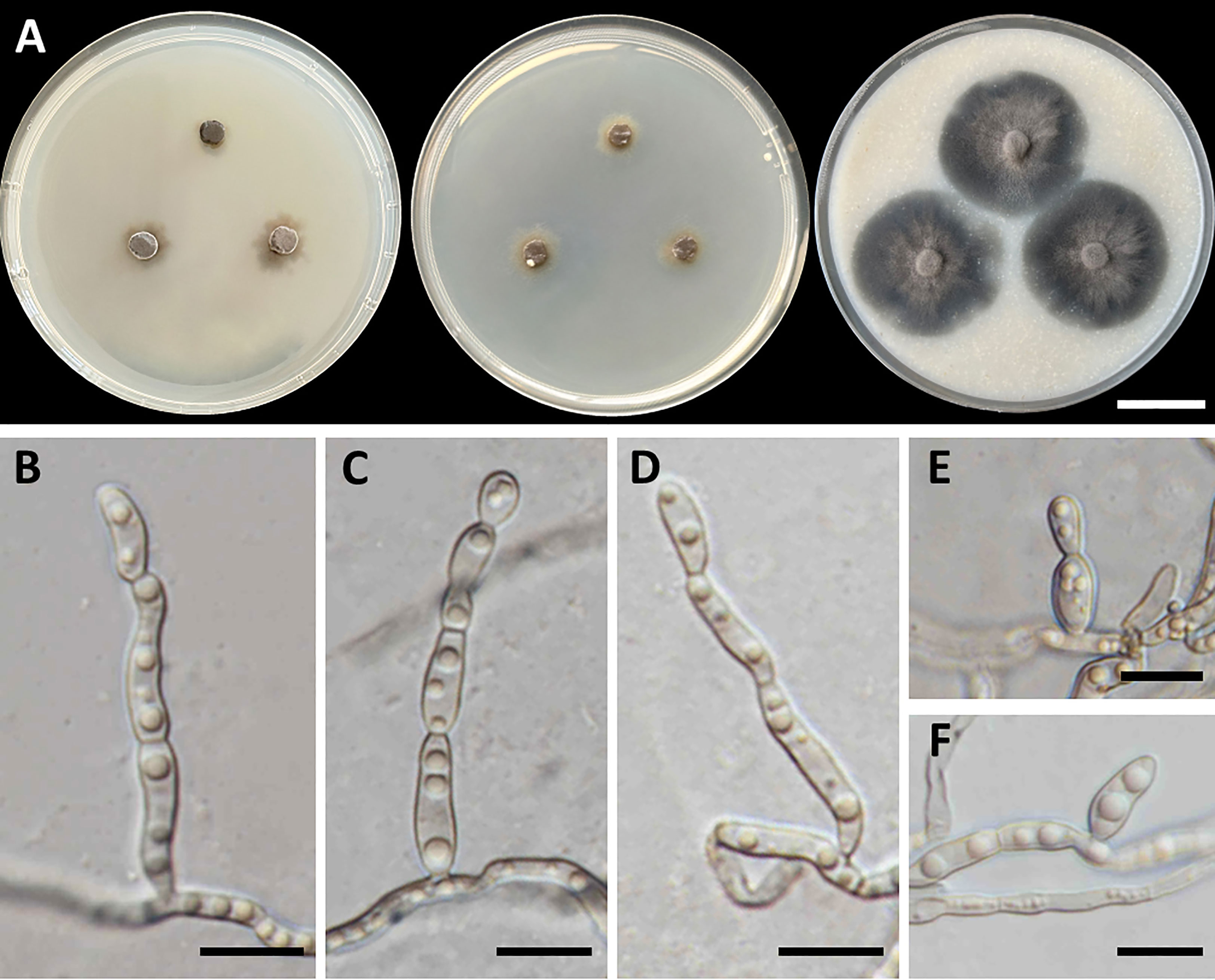
Figure 5 Cladophialophora sribuabanensis (SDBR-CMU446, ex-type). (A) Colonies at 25°C for 28 days on PDA, MEA, and OA, respectively. (B–D) Conidiophore and conidia. (E−F) Conidia on hyphae. Scale bars: (A) 2–cm and (B–F) 10 µm.
MycoBank number: MB849333
Etymology: “sribuabanensis” refers to Sribuaban Subdistrict, Lamphun province, Thailand, where the rock samples were collected.
Holotype: Thailand, Lamphun province, Mueang Lamphun District, Sribuaban Subdistrict, 18°32′11″N 99°07′22″E, elevation at 444 m, isolated from the sandstone of dipterocarp forest, July 2021, T. Thitla; dried culture, CMULPN5-9; ex-type culture, SDBR-CMU476.
GenBank number: OQ991178 (ITS), OQ979608 (nrLSU), OR141868 (nrSSU), OR139226 (tef1-α), and OR139238 (tub2)
Culture characteristics: Colonies on PDA and MEA at 25°C extremely slow growing: on PDA: attaining 8-mm to 15-mm diameter; and on MEA: attaining 12-mm to 14-mm diameter in 28 days. The colony character on PDA spreading, circular, flat, olive brown, and undulate margin; brown in reverse; on MEA: circular, spreading, flat, velvety, and grayish green to olive with entire margin; reverse grayish green; on OA: attaining 32-mm to 38-mm diameter, restricted, circular, flat, velvety, and olive brown to olive gray with entire margin; reverse olive gray. No diffusible pigment produced on any media. Hyphae: thin and smooth-walled, hyaline to light brown, 2-µm to 3-µm wide. Conidiophores: reduced to conidiogenous cells, semi-micronematous, lateral or terminal, smooth, thin-walled, hyaline to pale brown, sub-cylindrical, and 2-µm to 4-µm wide. Conidia: subglobose to fusiform, aseptate, guttules, smooth, thin-walled, hyaline to pale brown, forming to unbranched conidial chain at hyphae and conidiophore, and 7−16 µm × 3−6 µm (mean = 11.2 µm × 3.8 µm, n = 50). Chlamydospores and sexual structures were not produced on any media.
Growth temperature: minimum at 15°C, optimum at the range of 28°C, maximum at 30°C, and no growth at 10°C and 35°C.
Additional specimens examined: Thailand, Lamphun province, Mueang Lamphun District, Sribuaban Subdistrict, 18°32′11″N 99°07′22″E, elevation at 444 m, isolated from the rock of dipterocarp forest, July 2021, isolated by T. Thitla: SDBR-CMU477 [GenBank number: OQ991179 (ITS), OQ979609 (nrLSU), OR141869 (nrSSU), OR139227 (tef1-α), and OR139239 (tub2)].
Habitat and distribution: sandstone; collected from Lamphun province, Thailand.
Notes: Cladophialophora sribuabanensis was isolated from rock samples along with several other species, including Cl. nyingchiensis, Cl. rupestricola, Cl. tengchongensis, Cl. thailandensis, Cl. tumbae, and Cl. tumulicola, all of which were also isolated from rocks (Kiyuna et al., 2018; Sun et al., 2020). However, the conidial size of Cl. sribuabanensis (7−16 µm × 3−6 µm) was obviously larger than that of Cl. tumbae (4.5−6 µm × 2−2.5 µm) and longer than that of Cl. tumulicola (5−7 µm × 3−4 µm). Furthermore, Cl. nyingchiensis produced septate conidia, which differs from Cl. sribuabanensis, which produced aseptate conidia. Cladophialophora sribuabanensis showed an exceptionally slow growth rate, reaching only 8−15 mm and 12−14 mm in diameter at 25°C for 28 days on PDA and MEA, respectively. In contrast, both Cl. tumbae and Cl. tumulicola achieved significantly larger diameters of 15−16 mm and 22−25 mm on PDA, respectively, within just 14 days at 25°C (Kiyuna et al., 2018). Cladophialophora nyingchiensis, Cl. tengchongensis, and Cl. thailandensis also displayed faster growth rates on MEA, attaining diameters of 21 mm, 28 mm, and 23–30 mm, respectively, compared to Cl. sribuabanensis at the same temperature (Sun et al., 2020). Cladophialophora sribuabanensis also differs from Cl. thailandensis, which does not produce chlamydospores on any media, whereas Cl. thailandensis produces subspherical chlamydospores (7.5−17 µm × 4.5−14 µm). The temperature profile of growth revealed that Cl. sribuabanensis has a minimum growth temperature of 15°C, whereas Cl. nyingchiensis and Cl. tengchongensis could grow at 4°C. In addition, the maximum growth temperature of Cl. sribuabanensis is 30°C, whereas Cl. rupestricola has a maximum growth temperature of 35°C. Phylogenetically, Cl. sribuabanensis forms a sister clade to Cl. rupestricola (Figure 4). Cladophialophora sribuabanensis differs from Cl. rupestricola at different maximum growth temperatures (as mentioned above). Moreover, Cl. sribuabanensis does not produce any pigments in contrast to Cl. rupestricola, which produces olive and dull green pigments on PDA and MEA, respectively. Also, a pairwise nucleotide comparison of Cl. rupestricola and Cl. sribuabanensis revealed differences of 7.0% (45/642 bp) in the ITS, 0.2% (2/903 bp) in nrLSU, 0.3% (3/1,037 bp) in nrSSU, 9.8% (20/205 bp) in tef1-α, and 10.6% (54/509 bp) in tub2 sequences including gaps.
Cladophialophora thailandensis, T. Thitla, J. Kumla, and N. Suwannarach, sp. nov. (Figure 6)
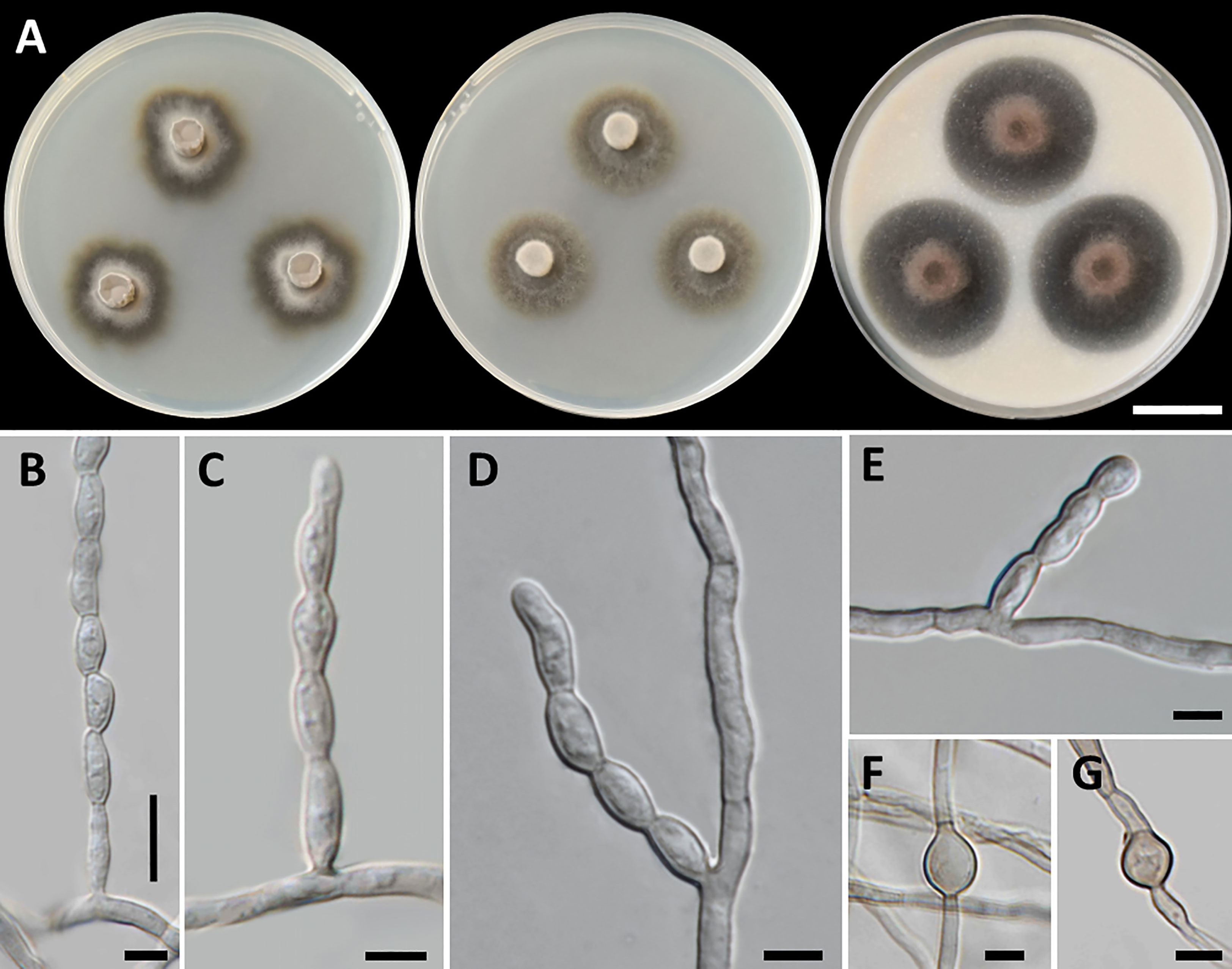
Figure 6 Cladophialophora thailandensis (SDBR-CMU451, ex-type), (A) Colonies at 25°C for 28 days on PDA, MEA, and OA, respectively. (B−E) Conidiophore and conidial chains. (F, G) Chlamydospores. Scale bars: (A) 2 cm and (B−G) 5 µm.
MycoBank number: MB846946
Etymology: “thailandensis” referring to Thailand, where the new species was found.
Holotype: Thailand, Lamphun province, Mueang Lamphun District, Sribuaban Subdistrict, 18°32′13″N 99°07′31″E, elevation at 432 m, isolated from the sandstone of dipterocarp forest, July 2021, T. Thitla; dried culture, CMULPN13-12; ex-type culture, SDBR-CMU451.
GenBank number: OP903470 (ITS), OP903507 (nrLSU), OR141865 (nrSSU), OP923700 (tef1-α), and OR139235 (tub2).
Culture characteristics: Colonies on PDA, MEA, and OA were described at 25°C in the dark after 28 days: on PDA: attaining 28-mm to 34-mm diameter, irregularly circular, flat, center velvety gray to black with grayish green and entire to undulate margin; black to grayish green in reverse; on MEA: attaining 23-mm to 30-mm diameter, spreading, circular, velvety, and greenish gray to dark green with grayish green and entire margin; reverse black to grayish green; on OA: attaining 32-mm to 39-mm diameter, restricted, circular, convex, and olive brown aerial mycelium with dark green and entire margin; reverse dark green to dull green. No diffusible pigment produced on any media. Hyphae smooth-walled, septate, hyaline to pale brown, 1.5-µm to 4-µm wide. Conidiophores: reduced to conidiogenous cells, lateral on undifferentiated hyphae, producing long chains conidia, smooth, pale olivaceous, cylindrical to subcylindrical, and 2-µm to 4-µm wide. Conidia: subspherical to fusiform, aseptate, subhyaline to hyaline, smooth, acropetal, catenulate, and 5−11 µm × 3−4 µm (mean = 7.3 µm × 3.3 µm, n = 50). Chlamydospores: subspherical, smooth, light to dark brown, and 7.5−17 µm × 4.5−14 µm (mean = 10.7 µm × 7.2 µm, n = 20). Sexual morph was not produced on media.
Growth temperature: minimum at 10°C, optimum at 28°C, maximum at 35°C, and no growth at 4°C and 37°C.
Additional specimens examined: Thailand, Lamphun province, Mueang Lamphun District, Sribuaban Subdistrict, 18°32′13″N 99°07′31″E, elevation at 432 m, isolated from the rock of dipterocarp forest, July 2021, isolated by T. Thitla SDBR-CMU449 [GenBank number: OP903468 (ITS), OP903505 (nrLSU), OR141863 (nrSSU), OP923698 (tef1-α), and OR139233 (tub2)], SDBR-CMU450 [GenBank number: OP903469 (ITS), OP903506 (nrLSU), OR141864 (nrSSU), OP923699 (tef1-α), and OR139234 (tub2)], and SDBR-CMU452 [GenBank number: OP903471 (ITS), OP903508 (nrLSU), OR141866 (nrSSU), OP923701 (tef1-α), and OR139236 (tub2)]; rock from dipterocarp forest, August 2022, isolated by T. Thitla SDBR-CMU453 [GenBank number: OP903472 (ITS), OP903509 (nrLSU), OR141867 (nrSSU), OP923702 (tef1-α), and OR139237 (tub2)].
Habitat and distribution: sandstone; known from Lamphun province, Thailand.
Notes: Cladophialophora thailandensis produced a greenish gray colony on MEA. This outcome was similar to that of Cl. eucalypti, Cl. immunda, Cl. mycetomatis, and Cl. pseudocarrionii. Nevertheless, Cl. thailandensis has distinct habitats from these species. Cladophialophora eucalypti was isolated from leaves of Eucalyptus dunnii, whereas Cl. pseudocarrionii originated from soil, and both Cl. immunda and Cl. mycetomatis were isolated from humans (Badali et al., 2008; Madrid et al., 2016; Crous et al., 2019a). Furthermore, the conidial size of Cl. thailandensis (5−11 µm × 3−4 µm) is shorter than that of Cl. eucalypti (8−20 µm × 2.5−3 µm) (Crous et al., 2019a) but longer than that of Cl. immunda (3−4.5 µm × 2.5−4 µm) and Cl. mycetomatis (2.5−3 µm × 2−3 µm) (Badali et al., 2008). Cladophialophora thailandensis can be distinguished from Cl. pseudocarrionii by having a subspherical chlamydospore, whereas the chlamydospore production of Cl. pseudocarrionii was not observed (Madrid et al., 2016). In terms of the growth temperature, Cl. thailandensis also differs from Cl. immunda, Cl. mycetomatis, and Cl. pseudocarrionii, as Cl. thailandensis is unable to grow at 37°C (Badali et al., 2008; Madrid et al., 2016).
The multi-gene phylogenetic analyses indicated that Cl. thailandensis formed a distinct monophyletic clade within Cladophialophora and was closely related to Cl. inabaensis and Cl. lanosa isolated from soil samples (Figure 4). However, the absence of chlamydospore formation in Cl. inabaensis is differentiated from Cl. thailandensis (Usui et al., 2016). In contrast to Cl. thailandensis, which exhibited a maximum growth temperature of 35°C, Cl. inabaensis could grow at 37°C, indicating a higher tolerance to elevated temperatures (Usui et al., 2016). Moreover, the conidia of Cl. thailandensis are longer than those of Cl. lanosa (2−4 µm × 2−3 µm) (Das et al., 2019). A pairwise nucleotide comparison between the ITS sequence data of Cl. thailandensis to Cl. inabaensis and Cl. lanosa indicated base pair differences of 8.8% (45/510 bp) and 6.9% (36/515 bp), respectively. In addition, a pairwise nucleotide comparison of nrLSU data also indicated that Cl. thailandensis differs from Cl. lanosa by 0.5% (4/764 bp) including gaps.
Discussion
Herpotrichiellaceae is one of the well-known families in the order Chaetothyriales (Wijayawardene et al., 2020; Tian et al., 2021; Bezerra et al., 2022; Wijayawardene et al., 2022). Presently, four genera in Herpotrichiellaceae have been discovered on rocks, viz., Cladophialophora, Exophiala, Phialophora, and Rhinocladiella (Sun et al., 2020; Liu et al., 2022; Thitla et al., 2022). In this study, we introduced new taxa of rock-inhabiting fungi in the family Herpotrichiellaceae that were collected from northern Thailand. These taxa comprised a novel genus, named Petriomyces and three new species of Cladophialophora. The genus Petriomyces has been introduced on the basis of its fungal habitat, morphological characteristics, growth temperature, and phylogenetic analysis. Consequently, the number of rock-inhabiting fungal genera in the family Herpotrichiellaceae has increased to five genera.
Prior to this study, 20 genera (Aculeata, Atrokylindriopsis, Brycekendrickomyces, Capronia, Cladophialophora, Exophiala, Fonsecaea, Marinophialophora, Melanoctona, Metulocladosporiella, Minimelanolocus, Neosorocybe, Phialophora, Pleomelogramma, Rhinocladiella, Sorocybe, Thysanorea, Uncispora, Valentiella, and Veronaea) have been accepted into the family Herpotrichiellaceae (Wijayawardene et al., 2020; Tian et al., 2021; Bezerra et al., 2022; Wijayawardene et al., 2022). However, the number of genera in Herpotrichiellaceae has been a subject of debate in previous studies due to the variability of their morphological characteristics (Quan et al., 2020; Tian et al., 2021). Therefore, molecular phylogeny has been recognized as a valuable tool for researchers in the identification of fungi within Herpotrichiellaceae. Consequently, the aim of this study was to inspect and update the members of this family through molecular phylogeny using combined ITS, nrLSU, and nrSSU sequences. In our phylogenetic tree (Figure 1) indicated that species of herpotrichiellaceous fungi are paraphyletic and polyphyletic within the family Herpotrichiellaceae and Trichomeriaceae and this is concurred with previous studies (Quan et al., 2020; Wan et al., 2021).
A new genus (Petriomyces) obtained from this study formed within Herpotrichiellaceae (Figure 1). Regarding species Brycekendrickomyces and Metulocladosporiella, which were previously placed in the family Herpotrichiellaceae (Crous et al., 2006; Crous et al., 2009; Wijayawardene et al., 2020; Tian et al., 2021; Bezerra et al., 2022; Wijayawardene et al., 2022), our phylogenetic analysis placed their type sequences within the family Trichomeriaceae. Morphologically, these two species were reported from asexual characters, whereas the members of family Trichomeriaceae are predominantly known for their sexual characters. Although, there have been reports of asexual reproduction in certain species within family Trichomeriaceae, the specific asexual characters vary within the family. Because of the variability of asexual characters within the family Trichomeriaceae, we have tentatively transferred genera Brycekendrickomyces and Metulocladosporiella to the family Trichomeriaceae based solely on their phylogenetic placement. However, further studies, including additional collections and the linkage of sexual and asexual morphs for these two species, are necessary to confirm their phylogenetic placement and to gain a better understanding of the diversity of asexual characters within the family Trichomeriaceae. Furthermore, Neosorocybe and Sorocybe were also previously classified into Herpotrichiellaceae (Crous et al., 2020b; Wijayawardene et al., 2022). However, the phylogenetic analysis in this study revealed that these genera formed a unique clade to be clearly distinct from Herpotrichiellaceae and Trichomeriaceae (Figure 1). These two genera should be excluded from the family Herpotrichiellaceae. Therefore, we propose that the family Herpotrichiellaceae should contain 17 fungal genera (Table 4). However, there is still a need for the molecular data of the genus Pleomelogramma (Spegazzini, 1909) to confirm its phylogenetic placement. Most herpotrichiellaceous fungi still have a broadly paraphyletic and polyphyletic relationship. Therefore, further resolutions and characterizations are required to achieve a better understanding of their taxonomic classification within Herpotrichiellaceae.
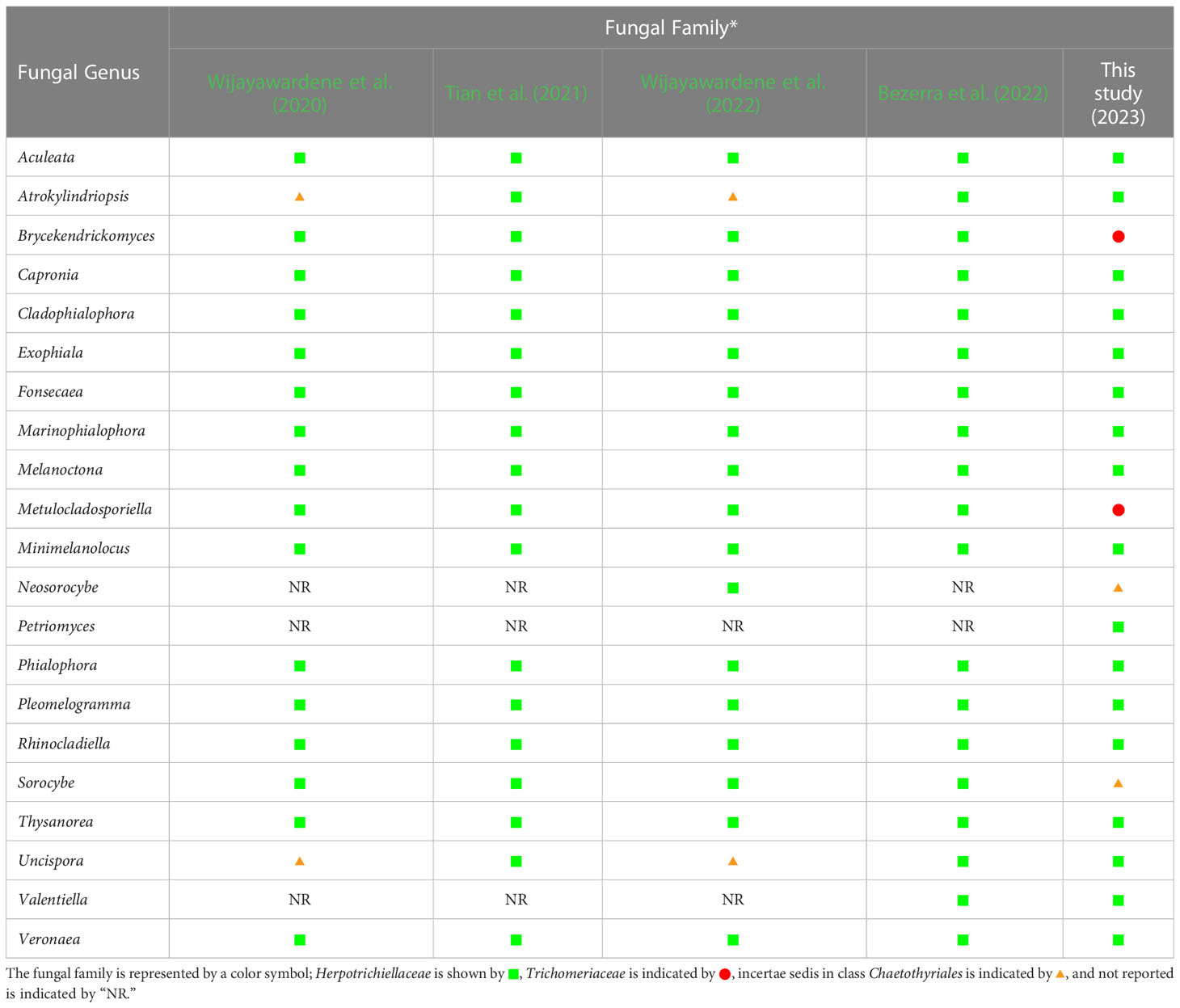
Table 4 Comparison of fungal genera belonging to the family Herpotrichiellaceae from previous reports and this study. .
In this study, the phylogenetic results indicate that Cladophialophora is a polyphyletic group, as is consistent with the findings of previous studies that reported the polyphyletic nature of the Cladophialophora species (Badali et al., 2008; Teixeira et al., 2017; Quan et al., 2020). Our three new species of Cladophialophora (Cl. rupestricola, Cl. sribuabanensis, and Cl. thailandensis) were established on the basis of comprehensive analyses of morphological characteristics, growth temperature, and multi-gene phylogeny. Furthermore, the nucleotide comparisons of the ITS, tef1-α, or tub2 genes with phylogenetically related species revealed nucleotide differences that were greater than 1.5%. This is in line with the suggestion made by Jeewon and Hyde (2016) that nucleotide differences above 1.5% are necessary to justify the recognition of a new species. Therefore, Cl. rupestricola, Cl. sribuabanensis, and Cl. thailandensis can be considered new species, whereas the total number of accepted Cladophialophora species globally has increased to 52. On the basis of our phylogenetic analysis, a total of 38 Cladophialophora species were classified as members of Herpotrichiellaceae. Three species (Cl. eucalypti, Cl. proteae, and Cl. pucciniophila) were placed in Trichomeriaceae, which is consistent with the study of Quan et al. (2020). Moreover, Quan et al. (2020) reported that Cl. humicola, Cl. minutissima, and Cl. sylvestris were placed in Epibryaceae. Accordingly, Cl. behniae, Cl. modesta, Cl. scillae, and Cl. hostae have been assigned in an incertae sedis clade Chaetothyriales (Quan et al., 2020; Crous et al., 2021). However, the phylogenetic placements of the remaining four species (Cl. cladoniae, Cl. hawksworthii, Cl. megalosporae, and Cl. normandinae) are uncertain and the acquisition of further molecular data will be necessary.
An investigation of fungal grown at variant temperatures indicates that our fungi could grow under a wide range of temperature conditions. Three new species of Cladophialophora could thrive at temperatures ranging from 10°C to 35°C. Moreover, a new genus Pe. obovoidisporus could thrive at temperatures ranging between 10°C and 30°C. Accordingly, these temperatures were within a range of 10°C−38°C in the dipterocarp forest in Lamphun and Sukhothai provinces, Thailand, respectively. As has become evident, both fungi have adapted to survive in their respective environments. Remarkably, growth temperature is one of the factors that can limit fungal ability to infect human and animal bodies due to their high body temperatures (Garcia-Solache and Casadevall, 2010). Some herpotrichiellaceous fungi that are known to infect humans and animals can grow at high temperatures. For example, Cl. arxii, Cl. carrionii, Phia. chinensis, and Phia. expanda can grow at temperatures of 37°C or higher (de Hoog et al., 1995; Li et al., 2017). The growth temperature of herpotrichiellaceous fungi is often associated with clinical predilections with the species growing at 40°C often causing systemic infections (de Hoog et al., 2000; Badali et al., 2008). Three new species of Cladophialophora and Petriomyces identified in this study may not be the source pathogen in humans and animals because they cannot grow at temperatures above 37°C. However, their ability to be the causal agents of infections in humans and animals needs to be further investigated. Currently, global temperatures are rising because of climate change, which can affect all aspects of a given ecosystem (Gadre et al., 2022). As a result, this environment may have an impact on the global diversity and distribution of herpotrichiellaceous fungi. Some herpotrichiellace fungi can survive and grow at high temperatures, but others cannot survive at higher temperatures and die.
In conclusion, we investigated the rock-inhabiting fungi belonged to the family Herpotrichiellaceae collected from northern Thailand. Three new species of Cladophialophora (Cl. rupestricola, Cl. sribuabanensis, and Cl. thailandensis) and a new genus (Petriomyces) were identified on the basis of the relevant morphological characteristics, growth temperature, and multi-gene phylogeny. Furthermore, the relevant data on the genera in Herpotrichiellaceae have been updated. In this study, we propose that Herpotrichiellaceae consists of 17 genera, including Aculeata, Atrokylindriopsis, Capronia, Cladophialophora, Exophiala, Fonsecaea, Marinophialophora, Melanoctona, Minimelanolocus, Petriomyces, Phialophora, Pleomelogramma, Rhinocladiella, Thysanorea, Uncispora, Valentiella, and Veronaea. However, the classification of several genera in this family is still problematic due to variabilities in some of the morphological characteristics, their existing polyphyletic relationships, and the limited amount of molecular data, especially protein coding genes. Therefore, further studies involving morphology, ecology, and molecular investigations, as well as in-depth evolutionary studies, should be conducted to further clarify the family Herpotrichiellaceae.
Data availability statement
The datasets presented in this study can be found in online repositories. The names of the repository/repositories and accession number(s) can be found in the article/supplementary material.
Author contributions
NS and SL: conceptualization in this study. TT, JK, and NS: methodology. TT, JK, SH, CS, and NS: software. TT, JK, and NS: validation. TT, SK, and JK: formal analysis. TT, JK, and NS: investigation. TT, SK, JK, and NS: resources. TT and JK: data curation. TT, JK, and NS: writing—original draft preparation. TT, JK, SH, CS, SK, SL, and NS: writing—review and editing. JK, SH, and NS: visualization. SL and NS: supervision. NS and JK: project administration. SL and NS: funding acquisition. All authors have read and agreed to the published version of the manuscript.
Funding
This study was supported by Fundamental Fund 2023 (grant number: FF66/065), Chiang Mai University, Thailand. Saisamorn Lumyong thanks National Research Council of Thailand for the support (grant number: N42A650198).
Acknowledgments
The authors gratefully acknowledge the TA & RA Scholarship, graduate school, Chiang Mai University, Thailand. The authors are grateful to Russell Kirk Hollis for the English correction of this manuscript.
Conflict of interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Publisher's note
All claims expressed in this article are solely those of the authors and do not necessarily represent those of their affiliated organizations, or those of the publisher, the editors and the reviewers. Any product that may be evaluated in this article, or claim that may be made by its manufacturer, is not guaranteed or endorsed by the publisher.
References
Abliz, P., Fukushima, K., Takizawa, K., Nieda, N., Miyaji, M., Nishimura, K. (2003). Rapid identification of the genus Fonsecaea by PCR with specific oligonucleotide primers. J. Clin. Microbiol. 41, 873–876. doi: 10.1128/JCM.41.2.873-876.2003
Abliz, P., Fukushima, K., Takizawa, K., Nishimura, K. (2004). Specific oligonucleotide primers for identification of Cladophialophora carrionii, a causative agent of chromoblastomycosis. J. Clin. Microbiol. 42, 404–407. doi: 10.1128/JCM.42.1.404-407.2004
Arzanlou, M., Crous, P. W. (2006). “Strelitziana africana,” in Fungal Planet No. 8 (Utrecht, The Netherlands: CBS-KNAW Fungal Biodiversity Centre).
Arzanlou, M., Groenewald, J. Z., Gams, W., Braun, U., Shin, H. D., Crous, P. W. (2007). Phylogenetic and morphotaxonomic revision of Ramichloridium and allied genera. Stud. Mycol. 58, 57–93. doi: 10.3114/sim.2007.58.03
Attili-Angelis, D., Duarte, A. P. M., Pagnocca, F. C., Nagamoto, N. S., de Vries, M., Stielow, J. B., et al. (2014). Novel Phialophora species from leaf-cutting ants (tribe Attini). Fungal Divers. 65, 65–75. doi: 10.1007/s13225-013-0275-0
Aureli, L., Pacelli, C., Cassaro, A., Fujimori, A., Moeller, R., Onofri, S. (2020). Iron ion particle radiation resistance of dried colonies of Cryomyces antarcticus embedded in Martian Regolith Analogues. Life 10, 306. doi: 10.3390/life10120306
Ávila, A., Groenewald, J. Z., Trapero, A., Crous, P. W. (2005). Characterisation and epitypification of Pseudocercospora cladosporioides, the causal organism of Cercospora leaf spot of olives. Mycol. Res. 109, 881–888. doi: 10.1017/S0953756205003503
Badali, H., Carvalho, V. O., Vicente, V., Attili-Angelis, D., Kwiatkowski, I. B., Gerrits Van Den Ende, A. H., et al. (2009). Cladophialophora saturnica sp. nov., a new opportunistic species of Chaetothyriales revealed using molecular data. Med. Mycol. 47, 51–62. doi: 10.1080/13693780802291452
Badali, H., Gueidan, C., Najafzadeh, M. J., Bonifaz, A., Gerrits van den Ende, A. H. G., de Hoog, G. S. (2008). Biodiversity of the genus cladophialophora. Stud. Mycol. 61, 175191. doi: 10.3114/sim.2008.61.18
Badali, H., Prenafeta-Boldu, F. X., Guarro, J., Klaassen, C. H., Meis, J. F., de Hoog, G. S. (2011). Cladophialophora psammophila, a novel species of Chaetothyriales with a potential use in the bioremediation of volatile aromatic hydrocarbons. Fungal Biol. 115, 1019–1029. doi: 10.1016/j.funbio.2011.04.005
Bezerra, J. D. P., Navarro, H. M. C., Almeida, J. H., Félix, C. R., Landell, M. F. (2022). Valentiella maceioensis gen. et sp. nov. (Herpotrichiellaceae, Chaetothyriales), a new black yeast-like fungus isolated from bromeliads in Brazil. Mycol. Prog. 21, 30. doi: 10.1007/s11557-022-01783-3
Boonmee, S., Wanasinghe, D. N., Calabon, M. S., Huanraluek, N., Chandrasiri, S. K. U., Jones, G. E. B., et al. (2021). Fungal diversity notes 1387−1511: taxonomic and phylogenetic contributions on genera and species of fungal taxa. Fungal Divers. 111, 1–335. doi: 10.1007/s13225-021-00489-3
Borelli, D. (1980). “Causal agents of chromoblastomycosis (Chromomycetes),” in Proceedings of the 5th International Conference on Mycoses: Superficial, cutaneous and subcutaneous infections, Scientific Publication No. 396, Pan American Health Organization, Caracas.
Braun, U. (1998). A monograph of Cercosporella, Ramularia and allied genera (phytopathogenic hyphomycetes) Vol. 2 (IHW-Verlag, Eching, Germany). 493 pp.
Braun, U., Crous, P. W., Dugan, F., Groenewald, J. Z., de Hoog, G. S. (2003). Phylogeny and taxonomy of Cladosporium-like hyphomycetes, including Davidiella gen. nov., the teleomorph of Cladosporium s. str. Mycol. Prog. 2, 3–18. doi: 10.1007/s11557-006-0039-2
Carbone, I., Kohn, L. M. (1999). A method for designing primer sets for speciation studies in filamentous ascomycetes. Mycologia 91, 553–556. doi: 10.1080/00275514.1999.12061051
Cassaro, A., Pacelli, C., Onofri, S. (2022). Survival, metabolic activity, and ultrastructural damages of Antarctic black fungus in perchlorates media. Front. Microbiol. 13. doi: 10.3389/fmicb.2022.992077
Cheewangkoon, R., Groenewald, J. Z., Summerell, B. A., Hyde, K. D., To-Anun, C., Crous, P. W. (2009). Myrtaceae, a cache of fungal biodiversity. Persoonia 23, 55–85. doi: 10.3767/003158509X474752
Chomnunti, P., Schoch, C. L., Aguirre-Hudson, B., Ko-Ko, T. W., Hongsanan, S., Jones, E. B., et al. (2011). Capnodiaceae. Fungal Divers. 51, 103–134. doi: 10.1007/s13225-011-0145-6
Coleine, C., Selbmann, L. (2021). “2.1 Black fungi inhabiting rock surfaces,” in Life at Rock Surfaces: Challenged by Extreme Light, Temperature and Hydration Fluctuations. Eds. Büdel, B., Friedl, T. (Berlin, Boston: De Gruyter), 57–86. doi: 10.1515/9783110646467-003
Crous, P. W., Braun, U., Wingfield, M. J., Wood, A. R., Shin, H. D., Summerell, B. A., et al. (2009). Phylogeny and taxonomy of obscure genera of microfungi. Persoonia 22, 139–161. doi: 10.3767/003158509X461701
Crous, P. W., Carnegie, A. J., Wingfield, M. J., Sharma, R., Mughini, G., Noordeloos, M. E., et al. (2019a). Fungal Planet description sheets: 868–950. Persoonia 42, 291–473. doi: 10.3767/persoonia.2019.42.11
Crous, P. W., Cowan, D. A., Maggs-Kölling, G., Yilmaz, N., Thangavel, R., Wingfield, M. J., et al. (2021). Fungal planet description sheets: 1182-1283. Persoonia 46, 313–528. doi: 10.3767/persoonia.2021.46.11
Crous, P. W., Schroers, H. J., Groenewald, J. Z., Braun, U., Schubert, K. (2006). Metulocladosporiella gen. nov. for the causal organism of Cladosporium speckle disease of banana. Mycol. Res. 110, 264–275. doi: 10.1016/j.mycres.2005.10.003
Crous, P. W., Schubert, K., Braun, U., de Hoog, G. S., Hocking, A. D., Shin, H. D., et al. (2007). Opportunistic, human-pathogenic species in the Herpotrichiellaceae are phenotypically similar to saprobic or phytopathogenic species in the Venturiaceae. Stud. Mycol. 58, 185–217. doi: 10.3114/sim.2007.58.07
Crous, P. W., Schumacher, R. K., Akulov, A., Thangavel, R., Hernández-Restrepo, M., Carnegie, A. J., et al. (2019b). New and interesting fungi. 2. Fungal Syst. Evol. 3, 57–134. doi: 10.3114/fuse.2019.03.06
Crous, P. W., Schumacher, R. K., Wingfield, M. J., Akulov, A., Denman, S., Roux, J., et al. (2018a). New and interesting fungi. 1. Fungal Syst. Evol. 1, 169–216. doi: 10.3114/fuse.2018.01.08
Crous, P. W., Wingfield, M. J., Burgess, T. I., Hardy, G. E., Crane, C., Barrett, S., et al. (2016). Fungal Planet description sheets: 469−557. Persoonia 37, 218–403. doi: 10.3767/003158516X694499
Crous, P. W., Wingfield, M. J., Burgess, T. I., Hardy, G. E. S. J., Gené, J., Guarro, J., et al. (2018b). Fungal Planet description sheets: 716−784. Persoonia 40, 240–393. doi: 10.3767/persoonia.2018.40.10
Crous, P. W., Wingfield, M. J., Chooi, Y. H., Gilchrist, C. L. M., Lacey, E., Pitt, J. I., et al. (2020a). Fungal Planet description sheets: 1042–1111. Persoonia 44, 301–459. doi: 10.3767/persoonia.2020.44.11
Crous, P. W., Wingfield, M. J., Guarro, J., Cheewangkoon, R., van der Bank, M., Swart, W. J., et al. (2013). Fungal Planet description sheets: 154–213. Persoonia 31, 188–296. doi: 10.3767/003158513X675925
Crous, P. W., Wingfield, M. J., Guarro, J., Hernández-Restrepo, M., Sutton, D. A., Acharya, K., et al. (2015). Fungal Planet description sheets: 320–370. Persoonia 34, 167–266. doi: 10.3767/003158515X688433
Crous, P. W., Wingfield, M. J., Lombard, L., Roets, F., Swart, W. J., Alvarado, P., et al. (2019c). Fungal Planet description sheets: 951–1041. Persoonia 43, 223–425. doi: 10.3767/persoonia.2019.43.06
Crous, P. W., Wingfield, M. J., Schumacher, R. K., Akulov, A., Bulgakov, T. S., Carnegie, A. J., et al. (2020b). New and interesting fungi. 3. Fungal Syst. Evol. 6, 157–231. doi: 10.3114/fuse.2020.06.09
Crous, P. W., Wingfield, M. J., Schumacher, R. K., Summerell, B. A., Giraldo, A., Gené, J., et al. (2014). Fungal Planet description sheets: 281–319. Persoonia 33, 212–289. doi: 10.3767/003158514X685680
Das, K., Lee, S. Y., Jung, H. Y. (2019). Cladophialophora lanosa sp. nov., a new species isolated from soil. Mycobiology 47, 173–179. doi: 10.1080/12298093.2019.1611242
de Hoog, G. S. (2010). Persoonial Reflections, Beware of your dishwasher! Persoonia 25, 117–159. doi: 10.3767/003158510X552636
de Hoog, G. S., Guarro, J., Gené, J. (2000). Atlas of Clinical Fungi. 2nd edn (Washington, DC: American Society for Microbiology Press).
de Hoog, G. S., Guého, E., Masclaux, F., Gerrits van den Ende, A. H. G., Kwon-Chung, K. J., McGinnis, M. R. (1995). Nutritional physiology and taxonomy of human-pathogenic Cladosporium-Xylohypha species. J. Med. Vet. Mycol. 33, 339–347. doi: 10.1080/02681219580000661
de Hoog, G. S., Nishikaku, A. S., Fernandez-Zeppenfeldt, G., Padín-González, C., Burger, E., Badali, H., et al. (2007). Molecular analysis and pathogenicity of the Cladophialophora carrionii complex, with the description of a novel species. Stud. Mycol. 58, 219–234. doi: 10.3114/sim.2007.58.08
de Hoog, G. S., Vicente, V., Caligiorne, R. B., Kantarcioglu, S., Tintelnot, K., Gerrits van den Ende, A. H., et al. (2003). Species diversity and polymorphism in the Exophiala spinifera clade containing opportunistic black yeast-like fungi. J. Clin. Microbiol. 41, 4767–4778. doi: 10.1128/JCM.41.10.4767-4778.2003
de Hoog, G. S., Vicente, V. A., Najafzadeh, M. J., Harrak, M. J., Badali, H., Seyedmousavi, S. (2011). Waterborne Exophiala species causing disease in cold-blooded animals. Persoonia 27, 46–72. doi: 10.3767/003158511X614258
Diederich, P., Ertz, D., Lawrey, J. D., Sikaroodi, M., Untereiner, W. A. (2013). Molecular data place the hyphomycetous lichenicolous genus Sclerococcum close to Dactylospora (Eurotiomycetes) and S. parmeliae in Cladophialophora (Chaetothyriales). Fungal Divers. 58, 61–72. doi: 10.1007/s13225-012-0179-4
Dong, W., Hyde, K. D., Bhat, D. J., Zhang, H. (2018). Introducing Aculeata aquatica gen. et sp. nov., Minimelanolocus Thailandensis sp. nov. and Thysanorea aquatica sp. nov. (Herpotrichiellaceae, Chaetothyriales) from freshwater in northern Thailand. Mycol. Prog. 17, 617–629. doi: 10.1007/s11557-018-1389-2
Edgar, R. C. (2004). MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 5, 113. doi: 10.1186/1471-2105-5-113
Felsenstein, J. (1985). Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791. doi: 10.2307/2408678
Feng, P. Y., de Hoog, G. S., Najafzadeh, M. J., Gerrits van den Ende, A. H. G., Stielow, J. B., Badali, H., et al. (2014). Cladophialophora abundans, a novel species of Chaetothyriales isolated from the natural environment. Mycol. Prog. 13, 381–391. doi: 10.1007/s11557-013-0924-4
Gadre, A., Enbiale, W., Andersen, L. K., Coates, S. J. (2022). The effects of climate change on fungal diseases with cutaneous manifes-tations: a report from the international society of dermatology climate change committee. J. Clim. Change Health 6, 100156. doi: 10.1016/j.joclim.2022.100156
Garcia-Solache, M. A., Casadevall, A. (2010). Global warming will bring new fungal diseases for mammals. MBio 1, 00061–00010. doi: 10.1128/mBio.00061-10
Geiser, D. M., Gueidan, C., Miadlikowska, J., Lutzoni, F., Kauff, F., Hofstetter, V., et al. (2006). Eurotiomycetes: eurotiomycetidae and chaetothyriomycetidae. Mycologia 98, 1053–1064. doi: 10.3852/mycologia.98.6.1053
Glass, N. L., Donaldson, G. C. (1995). Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Appl. Environ. Microbiol. 61, 1323–1330. doi: 10.1128/aem.61.4.1323-1330.1995
Glynou, K., Ali, T., Buch, A. K., Haghi Kia, S., Ploch, S., Xia, X., et al. (2016). The local environment determines the assembly of root endophytic fungi at a continental scale. Environ. Microbiol. 18, 2418–2434. doi: 10.1111/1462-2920.13112
Gorbushina, A. A. (2007). Life on the rocks. Environ. Microbiol. 9, 1613–1631. doi: 10.1111/j.1462-2920.2007.01301.x
Gostinčar, C., Zalar, P., Gunde-Cimerman, N. (2022). No need for speed: slow development of fungi in extreme environments. Fungal Biol. Rev. 39, 1–14. doi: 10.1016/j.fbr.2021.11.002
Gueidan, C., Aptroot, A., da Silva Cáceres, M. E., Badali, H., Stenroos, S. (2014). A reappraisal of orders and families within the subclass Chaetothyriomycetidae (Eurotiomycetes, Ascomycota). Mycol. Prog. 13, 990. doi: 10.1007/s11557-014-0990-2
Gueidan, C., Villaseñor, C. R., de Hoog, G. S., Gorbushina, A. A., Untereiner, W. A., Lutzoni, F. (2008). A rock-inhabiting ancestor for mutualistic and pathogen-rich fungal lineages. Stud. Mycol. 61, 111–119. doi: 10.3114/sim.2008.61.11
Haase, G., Sonntag, L., Melzer-Krick, B., de Hoog, G. S. (1999). Phylogenetic inference by SSU gene analysis of members of the Herpotrichiellaceae, with special reference to human pathogenic species. Stud. Mycol. 43, 80–97.
Hall, T. (2004) Bioedit Version 6.0.7. Available at: http://www.mbio.ncsu.edu/bioedit/bioedit (Accessed 28 March 2023).
Hernández-Restrepo, M., Giraldo, A., van Doorn, R., Wingfield, M. J., Groenewald, J. Z., Barreto, R. W., et al. (2010). The genera of fungi - G6: Arthrographis, Kramasamuha, Melnikomyces, Thysanorea, and Verruconis. Fungal Syst. Evol. 6, 1–24. doi: 10.3114/fuse.2020.06.01
Hernández-Restrepo, M., Schumacher, R. K., Wingfield, M. J., Ahmad, I., Cai, L., Duong, T. A., et al. (2016). Fungal systematics and evolution: FUSE 2. Sydowia 68, 193–230. doi: 10.12905/0380.sydowia68-2016-0193
Hubka, V., Réblová, M., Rehulka, J., Selbmann, L., Isola, D., de Hoog, S. G., et al. (2014). Bradymyces gen. nov. (Chaetothyriales, Trichomeriaceae), a new ascomycete genus accommodating poorly differentiated melanized fungi. Antonie van Leeuwenhoek 106, 979–992. doi: 10.1007/s10482-014-0267-4
Hyde, K. D., Norphanphoun, C., Chen, J., Dissanayake, A. J., Doilom, M., Hongsanan, S., et al. (2018). Thailand’s amazing diversity: up to 96% of fungi in northern Thailand may be novel. Fungal Divers. 93, 215–239. doi: 10.1007/s13225-018-0415-7
Index Fungorum. (2023). Available at: http://www.indexfungorum.org (Accessed 20 February 2023).
Isola, D., Zucconi, L., Onofri, S., Caneva, G., de Hoog, G. S., Selbmann, L. (2016). Extremotolerant rock inhabiting black fungi from Italian monumental sites. Fungal Divers. 76, 75–96. doi: 10.1007/s13225-015-0342-9
James, T. Y., Kauff, F., Schoch, C. L., Matheny, P. B., Hofstetter, V., Cox, C. J., et al. (2006). Reconstructing the early evolution of fungi using a six-gene phylogeny. Nature 443, 818–822. doi: 10.1038/nature05110
Jeewon, R., Hyde, K. D. (2016). Establishing species boundaries and new taxa among fungi: recommendations to resolve taxonomic ambiguities. Mycosphere 7, 1669–1677. doi: 10.5943/mycosphere/7/11/4
Khuna, S., Kumla, J., Thitla, T., Nuangmek, W., Lumyong, S., Suwannarach, N. (2022). Morphology, molecular identification, and pathogenicity of two novel Fusarium species associated with postharvest fruit rot of cucurbits in northern Thailand. J. Fungi 8, 1135. doi: 10.3390/jof8111135
Kiyuna, T., An, K. D., Kigawa, R., Sano, C., Sugiyama, J. (2018). Two new Cladophialophora species, C. tumbae sp. nov. and C. tumulicola sp. nov., and chaetothyrialean fungi from biodeteriorated samples in the Takamatsuzuka and Kitora Tumuli. Mycoscience 59, 7584. doi: 10.1016/j.myc.2017.08.008
Koukol, O. (2010). Revision of “Septonema ochraceum“ revealed three new species of Venturiaceae and Herpotrichiellaceae. Mycol. Prog. 9, 369–378. doi: 10.1007/s11557-009-0645-x
Li, D. M., de Hoog, G. S., Saunte, D. M., van den Ende, A. H., Chen, X. R. (2008). Coniosporium epidermidis sp. nov., a new species from human skin. Stud. Mycol. 61, 131–136. doi: 10.3114/sim.2008.61.13
Li, J., Jeewon, R., Phookamsak, R., Bhat, D. J., Mapook, A., Chukeatirote, E., et al. (2018). Marinophialophora garethjonesii gen. et sp. nov.: a new hyphomycete associated with Halocyphina from marine habitats in Thailand. Phytotaxa 345, 001–012. doi: 10.11646/phytotaxa.345.1.1
Li, Y., Xiao, J., de Hoog, G. S., Wang, X., Wan, Z., Yu, J., et al. (2017). Biodiversity and human-pathogenicity of Phialophora verrucosa and relatives in Chaetothyriales. Persoonia 38, 1–19. doi: 10.3767/003158517X692779
Libert, X., Chasseur, C., Packeu, A., Bureau, F., Roosens, N. H., De Keersmaecker, S. C. J. (2019). Exploiting the advantages of molecular tools for the monitoring of fungal indoor air contamination: first detection of Exophiala jeanselmei in indoor air of air-conditioned offices. Microorganisms 7, 674. doi: 10.3390/microorganisms7120674
Lima, B. J. F. S., Voidaleski, M. F., Gomes, R. R., Fornari, G., Soares, J. M. B., Bombassaro, A., et al. (2020). Selective isolation of agents of chromoblastomycosis from insect-associated environmental sources. Fungal Biol. 124, 194–204. doi: 10.1016/j.funbio.2020.02.002
Lin, L., Xu, J. (2020). Fungal pigments and their roles associated with human health. J. Fungi 6, 280. doi: 10.3390/jof6040280
Liu, L. N., Abdul, R., Atri, N. S., Bau, T., Belbahri, L., Bouket, A. C., et al. (2018). Fungal systematics and evolution: FUSE 4. Sydowia 70, 211–286. doi: 10.12905/0380.sydowia70-2018-0211
Liu, B., Fu, R., Wu, B., Liu, X., Xiang, M. (2022). Rock-inhabiting fungi: terminology, diversity, evolution and adaptation mechanisms. Mycology 13, 131. doi: 10.1080/21501203.2021.2002452
Liu, X. Y., Udayanga, D., Luo, Z. L., Chen, L. J., Zhou, D. Q., Su, H. Y., et al. (2015). Backbone tree for Chaetothyriales with four new species of Minimelanolocus from aquatic habitats. Fungal Biol. 119, 1046–1062. doi: 10.1016/j.funbio.2015.08.005
Lumbsch, H. T., Mangold, A., Lücking, R., García, M. A., Martín, M. P. (2004). Phylogeneticposition of the genera Nadvornikia and Pyrgillus (Ascomycota) based on molecular data. Symb. Bot. Ups. 34, 9–17.
Lumbsch, H. T., Schmitt, I., Lindemuth, R., Miller, A., Mangold, A., Fernandez, F., et al. (2005). Performance of four ribosomal DNA regions to infer higher-level phylogenetic relationships of inoperculate euascomycetes (Leotiomyceta). Mol. Phylogenet. Evol. 34, 512–524. doi: 10.1016/j.ympev.2004.11.007
Ma, Y. R., Xia, J. W., Gao, J. M., Li, X. Y., Castañeda Ruíz, R. F., Zhang, X. G., et al. (2015). Atrokylindriopsis, a new genus of hyphomycetes from Hainan, China, with relationship to Chaetothyriales. Mycol. Prog. 14, 77. doi: 10.1007/s11557-015-1071-x
Maciá-Vicente, J. G., Glynou, K., Piepenbring, M. (2016). A new species of Exophiala associated with roots. Mycol. Prog. 15, 18. doi: 10.1007/s11557-016-1161-4
Madrid, H., Hernández-Restrepo, M., Gené, J., Cano, J., Guarro, J., Silva, V. (2016). New and interesting chaetothyrialean fungi from Spain. Mycol. Prog. 15, 1179–1201. doi: 10.1007/s11557-016-1239-z
Marincowitz, S., Crous, P. W., Groenewald, J. Z., Wingfield, M. J. (2008). Microfungi Occurring on Proteaceae in the Fynbos (Utrecht, the Netherlands: CBS Fungal Biodiversity Centre). 166 pp.
Mattoon, E. R., Cordero, R. J. B., Casadevall, A. (2021). Fungal melanins and applications in healthcare, bioremediation and industry. J. Fungi 7, 488. doi: 10.3390/jof7060488
Miller, M. A., Holder, M. T., Vos, R., Midford, P. E., Liebowitz, T., Chan, L., et al. (2009) The CIPRES Portals. Available at: http://www.phylo.org/sub_sections/portal (Accessed 12 June 2023).
Müller, E., Petrini, O., Fisher, P. J., Samuels, G. J., Rossman, A. Y. (1987). Taxonomy and anamorphs of the Herpotrichiellaceae with notes on generic synonymy. Trans. Br. Mycol. Soc 88, 63–74. doi: 10.1016/S0007-1536(87)80186-9
Munk, A. (1953). The system of the pyrenomycetes. Dansk Botanisk Arkiv Udgivet af Dansk Botanisk Forening 15, 1–163.
Mycobank Database. Available at: https://www.mycobank.org/ (Accessed 29 June 2023).
Nai, C., Wong, H. Y., Pannenbecker, A., Broughton, W. J., Benoit, I., de Vries, R. P., et al. (2013). Nutritional physiology of a rock-inhabiting, model microcolonial fungus from an ancestral lineage of the Chaetothyriales (Ascomycetes). Fungal Genet. Biol. 56, 54–66. doi: 10.1016/j.fgb.2013.04.001
Najafzadeh, M. J., Gueidan, C., Badali, H., Van Den Ende, A. H., Xi, L., de Hoog, G. S. (2009). Genetic diversity and species delimitation in the opportunistic genus Fonsecaea. Med. Mycol. 47, 17–25. doi: 10.1080/13693780802527178
Najafzadeh, M. J., Vicente, V. A., Sun, J., Meis, J. F., de Hoog, G. S. (2011). Fonsecaea multimorphosa sp. nov., a new species of Chaetothyriales isolated from a feline cerebral abscess. Fungal Biol. 115, 1066–1076. doi: 10.1016/j.funbio.2011.06.007
Nascimento, S. S., Ferro, L. O., Freire, K. T. L. S., Barbosa, R. N., Paiva, L. M., Souza-Motta, C. M., et al. (2021). Cladophialophora bromeliacearum (Herpotrichiellaceae, Chaetothyriales), a novel endophytic species from the Brazilian tropical dry forest. Phytotaxa 509, 211–224. doi: 10.11646/phytotaxa.509.2.4
Nascimento, M. M. F., Selbmann, L., Sharifynia, S., Al-Hatmi, A. M. S., Voglmayr, H., Vicente, V. A., et al. (2016). Arthrocladium, an unexpected human opportunist in Trichomeriaceae (Chaetothyriales). Fungal Biol. 120, 207–218. doi: 10.1016/j.funbio.2015.08.018
Nascimento, M. M. F., Vicente, V. A., Bittencourt, J. V. M., Gelinski, J. M. L., Prenafeta-Boldú, F. X., Romero-Güiza, M., et al. (2017). Diversity of opportunistic black fungi on babassu coconut shells, a rich source of esters and hydrocarbons. Fungal Biol. 121, 488–500. doi: 10.1016/j.funbio.2017.01.006
Nishimura, D. (2000). Sequencher 3.1.1. Biotech softw. Internet Rep. 1, 24–30. doi: 10.1089/152791600319231
Obase, K., Douhan, G. E., Matsuda, Y., Smith, M. E. (2016). Cladophialophora floridana and Cladophialophora tortuosa, new species isolated from sclerotia of Cenococcum geophilum in forest soils of Florida, USA. Mycoscience 57, 26–34. doi: 10.1016/j.myc.2015.07.005
Park, M. J., Shin, H. D. (2011). Cladophialophora pucciniophila, a new hyphomycete parasitizing a rust fungus. Mycotaxon 116, 449–456. doi: 10.5248/116.449
Phookamsak, R., Hyde, K. D., Jeewon, R., Bhat, D. J., Jones, E. B. G., Maharachchikumbura, S. S. N., et al. (2019). Fungal diversity notes 929–1035: taxonomic and phylogenetic contributions on genera and species of fungi. Fungal Divers. 95, 1–273. doi: 10.1007/s13225-019-00421-w
Phukhamsakda, C., Nilsson, R. H., Bhunjun, C. S., de Farias, A. R. G., Sun, Y. R., Wijesinghe, S. N., et al. (2022). The numbers of fungi: contributions from traditional taxonomic studies and challenges of metabarcoding. Fungal Divers. 114, 327–386. doi: 10.1007/s13225-022-00502-3
Prenafeta-Boldú, F. X., Summerbell, R., de Hoog, G. S. (2006). Fungi growing on aromatic hydrocarbons: biotechnology's unexpected encounter with biohazard? FEMS Microbiol. Rev. 30, 109–130. doi: 10.1111/j.1574-6976.2005.00007.x
Quan, Y., Muggia, L., Moreno, L. F., Wang, M., Al-Hatmi, A. M. S., Menezes, N. D. S., et al. (2020). A re-evaluation of the Chaetothyriales using criteria of comparative biology. Fungal Divers. 103, 47–85. doi: 10.1007/s13225-020-00452-8
Rakeman, J. L., Bui, U., Lafe, K., Chen, Y. C., Honeycutt, R. J., Cookson, B. T. (2005). Multilocus DNA sequence comparisons rapidly identify pathogenic molds. J. Clin. Microbiol. 43, 3324–3333. doi: 10.1128/JCM.43.7.3324-3333.2005
Rambaut, A. (2019) FigTree tree figure drawing tool version 131; Institute of Evolutionary 623 Biology (Edinburgh, Scotland: University of Edinburgh). Available at: http://treebioedacuk/software/figtree/ (Accessed 12 June 2023).
Réblová, M., Hubka, V., Thureborn, O., Lundberg, J., Sallstedt, T., Wedin, M., et al. (2016). From the tunnels into the treetops: new lineages of black yeasts from biofilm in the Stockholm metro system and their relatives among ant-associated fungi in the Chaetothyriales. PloS One 11, 163396. doi: 10.1371/journal.pone.0163396
Rehner, S. A., Samuels, G. J. (1994). Taxonomy and phylogeny of gliocladium analyzed from nuclear large subunit ribosomal DNA sequences. Mycol. Res. 98, 625–634. doi: 10.1016/S0953-7562(09)80409-7
Rodríguez-Andrade, E., Stchigel, A. M., Guarro, J., Cano-Lira, J. F. (2019). Fungal diversity of deteriorated sparkling wine and cork stoppers in Catalonia, Spain. Microorganisms 8, 12. doi: 10.3390/microorganisms8010012
Ronquist, F., Huelsenbeck, J. P. (2003). MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19, 1572–1574. doi: 10.1093/bioinformatics/btg180
Saccardo, P. A. (1883). “Sylloge Pyrenomycetum, Vol. II,” in Sylloge Fungorum, vol. 2, Johnson Reprint Corperation, New York.1–288.
Sánchez, R. M., Miller, A. N., Bianchinotti, M. V. (2019). New species of Capronia (Herpotrichiellaceae, Ascomycota) from Patagonian forests, Argentina. Plant Fungal Syst. 64, 81–90. doi: 10.2478/pfs-2019-0009
Scott, J. A., Untereiner, W. A., Ewaze, J. O., Wong, B., Doyle, D. (2007). Baudoinia, a new genus to accommodate Torula compniacensis. Mycologia 99, 592–601. doi: 10.3852/mycologia.99.4.592
Seifert, K. A., Hughes, S. J., Boulay, H., Louis-Seize, G. (2007). Taxonomy, nomenclature and phylogeny of three cladosporium-like hyphomycetes, Sorocybe resinae, Seifertia azaleae and the Hormoconis anamorph of Amorphotheca resinae. Stud. Mycol. 58, 235–245. doi: 10.3114/sim.2007.58.09
Selbmann, L., Isola, D., Egidi, E., Zucconi, L., Gueidan, C., de Hoog, G. S., et al. (2014). Mountain tips as reservoirs for new rock-fungal entities: Saxomyces gen. nov. and four new species from the Alps. Fungal Divers. 65, 167–182. doi: 10.1007/s13225-013-0234-9
Seyedmousavi, S., Badali, H., Chlebicki, A., Zhao, J., Prenafeta-Boldú, F. X., de Hoog, G. S. (2011). Exophiala sideris, a novel black yeast isolated from environments polluted with toxic alkyl benzenes and arsenic. Fungal Biol. 115, 1030–1037. doi: 10.1016/j.funbio.2011.06.004
Shen, M., Zhang, J. Q., Zhao, L. L., Groenewald, J. Z., Crous, P. W., Zhang, Y. (2020). Venturiales. Stud. Mycol. 96, 185–308. doi: 10.1016/j.simyco.2020.03.001
Spegazzini, C. (1909). Mycetes argentinenses. Series IV. Anales Mus. Nac. Hist. Nat. Buenos Aires 3, 257–458.
Stamatakis, A. (2006). RAxML-VI-HPC: maximum likelihood-based phylogenetic analyses with thousands of taxa and mixed models. Bioinformatics 22, 2688–2690. doi: 10.1093/bioinformatics/btl446
Sun, W., Su, L., Yang, S., Sun, J., Liu, B., Fu, R., et al. (2020). Unveiling the hidden diversity of rock-inhabiting fungi: Chaetothyriales from China. J. Fungi 6, e187. doi: 10.3390/jof6040187
Tanabe, H., Kawasaki, M., Mochizuki, T., Ishizaki, H. (2004). Species identification and strain typing of Fonsecaea pedrosoi using ribosomal RNA gene internal transcribed spacer regions. Jpn. J. Med. Mycol. 45, 105–112. doi: 10.3314/jjmm.45.105
Teixeira, M. M., Moreno, L. F., Stielow, B. J., Muszewska, A., Hainaut, M., Gonzaga, L., et al. (2017). Exploring the genomic diversity of black yeasts and relatives (Chaetothyriales, Ascomycota). Stud. Mycol. 86, 1–28. doi: 10.1016/j.simyco.2017.01.001
Tesei, D. (2022). Black fungi research: out-of-this-world implications. Encyclopedia 2, 212–229. doi: 10.3390/encyclopedia2010013
Thitla, T., Kumla, J., Khuna, S., Lumyong, S., Suwannarach, N. (2022). Species diversity, distribution, and phylogeny of Exophiala with the addition of four new species from Thailand. J. Fungi 8, 766. doi: 10.3390/jof8080766
Tian, Q., Chomnunti, P., Lumyong, S., Liu, J. K., Hyde, K. D. (2021). Phylogenetic relationships and morphological reappraisal of Chaetothyriales. Mycosphere 12, 1157–1261. doi: 10.5943/mycosphere/12/1/15
Tian, Q., Doilom, M., Luo, Z., Chomnunti, P., Bhat, J. D., Xu, J., et al. (2016). Introducing Melanoctona tectonae gen. et sp. nov. and Minimelanolocus yunnanensis sp. nov. (Herpotrichiellaceae, Chaetothyriales). Cryptogam. Mycol. 37, 477–492. doi: 10.7872/crym/v37.iss4.2016.477
Tran-Ly, A. N., Reyes, C., Schwarze, F. W. M. R., Ribera, J. (2020). Microbial production of melanin and its various applications. World J. Microbiol. Biotechnol. 36, 170. doi: 10.1007/s11274-020-02941-z
Tsuneda, A., Hambleton, S., Currah, R. S. (2011). The anamorph genus Knufia and its phylogenetically allied species in Coniosporium, Sarcinomyces, and Phaeococcomyces. Botany 89, 523–536. doi: 10.1139/b11-041
Untereiner, W. A. (1997). Taxonomy of selected members of the ascomycete genus Capronia with notes on anamorph-teleomorph connections. Mycologia 89, 120–131. doi: 10.2307/3761181
Untereiner, W. A., Naveau, F. A. (1999). Molecular systematics of the Herpotrichiellaceae with an assessment of the phylogenetic positions of Exophiala dermatitidis and Phialophora americana. Mycologia 91, 67–83. doi: 10.1080/00275514.1999.12060994
Untereiner, W. A., Straus, N. A., Malloch, D. (1995). A molecular-morphotaxonomic approach to the systematics of the Herpotrichiellaceae and allied black yeasts. Mycol. Res. 99, 897–913. doi: 10.1016/s0953-7562(09)80748-x
Usui, E., Takashima, Y., Narisawa, K. (2016). Cladophialophora inabaensis sp. nov., a new species among the dark septate endophytes from a secondary forest in Tottori, Japan. Microbes Environ. 31, 357–360. doi: 10.1264/jsme2.ME16016
Vasse, M., Voglmayr, H., Mayer, V., Gueidan, C., Nepel, M., Moreno, L., et al. (2017). A phylogenetic perspective on the association between ants (Hymenoptera: Formicidae) and black yeasts (Ascomycota: Chaetothyriales). Proc. R. Soc B 284, 20162519. doi: 10.1098/rspb.2016.2519
Vicente, V. A., Najafzadeh, M. J., Sun, J., Gomes, R. R., Robl, D., Marques, S. G., et al. (2014). Environmental siblings of black agents of human chromoblastomycosis. Fungal Divers. 65, 47–63. doi: 10.1007/s13225-013-0246-5
Vicente, V. A., Orélis-Ribeiro, R., Najafzadeh, M. J., Sun, J., Guerra, R. S., Miesch, S., et al. (2012). Black yeast-like fungi associated with Lethargic Crab Disease (LCD) in the mangrove-land crab, Ucides cordatus (Ocypodidae). Vet. Microbiol. 158, 109–122. doi: 10.1016/j.vetmic.2012.01.031
Vilgalys, R., Hester, M. (1990). Rapid genetic identification and mapping of enzymatically amplified ribosomal DNA from several Cryptococcus species. J. Bacteriol. 172, 4238–4246. doi: 10.1128/jb.172.8.4238-4246.1990
Vitale, R. G., de Hoog, G. S. (2002). Molecular diversity, new species and antifungal susceptibilities in the Exophiala spinifera clade. Med. Mycol. 40, 545–556. doi: 10.1080/mmy.40.6.545.556
Vu, D., Groenewald, M., de Vries, M., Gehrmann, T., Stielow, B., Eberhardt, U., et al. (2019). Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Stud. Mycol. 92, 135–154. doi: 10.1016/j.simyco.2018.05.001
Wan, Y., Bao, D. F., Luo, Z. L., Bhat, D. J., Xu, Y. X., Su, H. Y., et al. (2021). Two new species of Minimelanolocus (Herpotrichiellaceae, Chaetothyriales) from submerged wood in Yunnan, China. Phytotaxa 480, 45–56. doi: 10.11646/phytotaxa.480.1.4
White, T. J., Bruns, T., Lee, S., Taylor, J. W. (1990). “Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics,” in PCR Protocols: A Guide to Methods and Applications. Eds. Innes, M. A., Gelfand, D. H., Sninsky, J. J., White, T. J. (San Diego, CA, USA: Academic Press), 315–322.
Wijayawardene, N. N., Hyde, K. D., Al-Ani, L. K. T., Tedersoo, L., Haelewaters, D., Rajeshkumar, K. C., et al. (2020). Outline of Fungi and fungus-like taxa. Mycosphere 11, 1060–1456. doi: 10.5943/mycosphere/11/1/8
Wijayawardene, N. N., Hyde, K. D., Dai, D. Q., Sánchez-García, M., Goto, B. T., Saxena, R. K., et al. (2022). Outline of Fungi and fungus-like taxa – 2021. Mycosphere 13, 53–453. doi: 10.5943/mycosphere/13/1/2
Wollenzien, U., de Hoog, G. S., Krumbein, W. E., Urzí, C. (1995). On the isolation of microcolonial fungi occurring on and in marble and other calcareous rocks. Sci. Total Environ. 167, 1–3. doi: 10.1016/0048-9697(95)04589-S
Woo, P. C., Ngan, A. H., Tsang, C. C., Ling, I. W., Chan, J. F., Leung, S. Y., et al. (2013). Clinical spectrum of exophiala infections and a novel Exophiala species, Exophiala hongkongensis. J. Clin. Microbiol. 51, 260–267. doi: 10.1128/JCM.02336-12
Yang, X. Q., Feng, M. Y., Yu, Z. F. (2021). Exophiala pseudooligosperma sp. nov., a novel black yeast from soil in southern China. Int. J. Syst. Evol. Microbiol. 71, 5116. doi: 10.1099/ijsem.0.005116
Yang, G. Z., Lu, J., Yu, Z., Zhang, K., Qiao, M. (2011). Uncispora sinensis, a new species from China. Mycotaxon 116, 171–174. doi: 10.5248/116.171
Keywords: herpotrichiellaceous fungi, new taxa, rock-inhabiting fungi, taxonomy, tropical area
Citation: Thitla T, Kumla J, Hongsanan S, Senwanna C, Khuna S, Lumyong S and Suwannarach N (2023) Exploring diversity rock-inhabiting fungi from northern Thailand: a new genus and three new species belonged to the family Herpotrichiellaceae. Front. Cell. Infect. Microbiol. 13:1252482. doi: 10.3389/fcimb.2023.1252482
Received: 03 July 2023; Accepted: 01 August 2023;
Published: 25 August 2023.
Edited by:
Yusufjon Gafforov, Academy of Sciences Republic of Uzbekistan (UzAS), UzbekistanReviewed by:
Saowaluck Tibpromma, Qujing Normal University, ChinaLei Su, Chinese Academy of Medical Sciences and Peking Union Medical College, China
Copyright © 2023 Thitla, Kumla, Hongsanan, Senwanna, Khuna, Lumyong and Suwannarach. This is an open-access article distributed under the terms of the Creative Commons Attribution License (CC BY). The use, distribution or reproduction in other forums is permitted, provided the original author(s) and the copyright owner(s) are credited and that the original publication in this journal is cited, in accordance with accepted academic practice. No use, distribution or reproduction is permitted which does not comply with these terms.
*Correspondence: Nakarin Suwannarach, c3V3YW5fNDYxQGhvdG1haWwuY29t
 Tanapol Thitla
Tanapol Thitla Jaturong Kumla
Jaturong Kumla Sinang Hongsanan
Sinang Hongsanan Chanokned Senwanna
Chanokned Senwanna Surapong Khuna
Surapong Khuna Saisamorn Lumyong
Saisamorn Lumyong Nakarin Suwannarach
Nakarin Suwannarach